Abstract
Previous work documented complete perchlorate reduction in a membrane biofilm reactor (MBfR) using methane as the sole electron donor and carbon source. This work explores how the biofilm’s microbial community evolved as the biofilm stage-wise reduced different combinations of perchlorate, nitrate, and nitrite. The initial inoculum, carrying out anaerobic methane oxidation coupled to denitrification (ANMO-D), was dominated by uncultured Anaerolineaceae and Ferruginibacter sp. The microbial community significantly changed after it was inoculated into the CH4-based MBfR and fed with a medium containing perchlorate and nitrite. Archaea were lost within the first 40 days, and the uncultured Anaerolineaceae and Ferruginibacter sp. also had significant losses. Replacing them were anoxic methanotrophs, especially Methylocystis, which accounted for more than 25 % of total bacteria. Once the methanotrophs became important, methanol-oxidizing denitrifying bacteria, namely, Methloversatilis and Methylophilus, became important in the biofilm, probably by utilizing organic matter generated by the metabolism of methanotrophs. When methane consumption was equal to the maximum-possible electron-donor supply, Methylomonas, also an anoxic methanotroph, accounted for >10 % of total bacteria and remained a major part of the community until the end of the experiments. We propose that aerobic methane oxidation coupled to denitrification and perchlorate reduction (AMO-D and AMO-PR) directly oxidized methane and reduced NO3 − to NO2 − or N2O under anoxic condition, producing organic matter for methanol-assimilating denitrification and perchlorate reduction (MA-D and MA-PR) to reduce NO3 −. Simultaneously, bacteria capable of anaerobic methane oxidation coupled to denitrification and perchlorate reduction (ANMO-D and ANMO-PR) used methane as the electron donor to respire NO3 − or ClO4 − directly.

ᅟ
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Perchlorate (ClO4 −) causes serious human health problems by interfering with the production of thyroid hormones needed for growth and development (Coates and Achenbach 2004), and California is planning to lower its drinking water maximum contaminant level (MCL) to 1 μg/L (Water Resource Foundation 2015). The main environmental sources of perchlorate are rocket fuel, explosives, and certain fertilizers (Coates and Achenbach 2004). ClO4 − contamination often is accompanied by pollution from nitrate (NO3 −; USEPA 2001), which has an MCL of 10 mg N/L because it causes methemoglobinemia in infants (USEPA 2015).
Extensive studies have proven that ClO4 − can be biologically reduced using organic substrates (e.g., Dechloromonas, Azospira, and Dechlorspirillum bacteria use acetate and lactate) (Coates and Achenbach 2004) and inorganic substrates as electron donors (e.g., Dechloromonas bacteria use hydrogen gas (H2)) (Logan et al. 2001; Nerenberg et al. 2002; Coates et al. 1999; Rikken et al. 1996). We recently reported complete ClO4 − reduction using anaerobic methane oxidation coupled to denitrification (ANMO-D), in which methane (CH4) was the sole electron donor and carbon source (Luo et al. 2015). We described the process as anaerobic methane oxidation coupled to perchlorate reduction (ANMO-PR). The ANMO-D biofilm reduced up to 5 mg/L of ClO4 − to a non-detectable concentration when CH4 delivery was sufficient; NO3 − was completely reduced as well when its surface loading (SL) was ≤0.302 g N/m2 day. When CH4 delivery was insufficient, NO3 − inhibited ClO4 − reduction by competing for the scarce electron donor. Nitrite (NO2 −) inhibited ClO4 − reduction at a lower SL (0.10 g N/m2 day), probably due to cellular toxicity. Similarly, although some bacteria, e.g., Azospira suillum and Dechloromonas, were reported to use both nitrate and perchlorate as electron acceptors (Chaudhuri et al. 2002; Nerenberg et al. 2008), Ziv-El and Rittmann reported that, when hydrogen was supplied as electron donor and in a limited condition, the nitrate had a clear H2 utilization priority than perchlorate.
Aerobic methane oxidation coupled to denitrification (AMO-D) is performed by the combined actions of methane oxidizers (types I, II, and X methanotrophs) and denitrifiers (Eisentraeger et al. 2001; Knowles 2005; Yao et al. 2013; Modin et al. 2007; Osaka et al. 2006). When methanotrophs oxidize methane using oxygen (O2) for the initial mono-oxygenation step, they release soluble organic compounds such as methanol, acetate, and citrate. These organics can be oxidized by denitrifiers to reduce NO3 − or NO2 − (Eisentraeger et al. 2001; Modin et al. 2010; Hu et al. 2009).
ANMO-D involves two processes conducted by two distinct microbial groups. Archaea phylogenetically positioned between Methanosaeta and ANME-II (candidate Methanoperedens nitroreducens) carry out reverse methanogenesis to generate H2 for themselves to reduce NO3 − to NO2 − (Raghoebarsing et al. 2006; Haroon et al. 2013). NO2 − is then reduced by denitrifying bacteria such as NC10 (Candidate division) using methane as the electron donor through an “intra-aerobic” pathway (Ettwig et al. 2010; Wu et al. 2011). During the intra-aerobic pathway, Methylomirabilis oxyfera stepwise reduces NO2 − to NO using nitrate reductase (narGHJI) and nitrite reductase (nirSJFD/GH/L); then, it intracellularly dismutates NO to O2 and N2. The O2 is used by a membrane-bound particulate methane mono-oxygenase (pMMO) for methane oxidation.
AMO-D and ANMO-D have three distinct differences. First, ANMO-D microorganisms are much slower growers than are the AMO-D bacteria (He et al. 2015). Second, AMO-D bacteria grow with a dissolved oxygen (DO) concentration ≤ 0.5 mg/L (Sun et al. 2013), while the ANMO-D microorganisms work with much lower dissolved oxygen concentration, since the oxygen is disproportionated intracellularly from NO (Ettwig et al. 2010). Third, the ratios of methane consumption to nitrate reduction (CH4/NO3 −) are significantly different between the two types of methane consumers, 5/4 mol CH4/mol NO3 − for AMO-D (according to Eq. 3) versus 5/8 mol CH4/mol NO3 − for ANMO-D (according to Eq. 5) (Modin et al. 2007, 2010; Osaka et al. 2006; Hu et al. 2009; Raghoebarsing et al. 2006). For aerobic methane oxidation, a half of the electrons from CH4 eventually flow to O2 (shown in Eqs. 1, 2, and 3) and the other half flow to NO3 − (shown in Eq. 3). For intra-aerobic-type anaerobic methane oxidation, the oxygen which is used to activate methane comes from the dismutation of electron acceptors; thus, all the electrons from CH4 eventually flow to NO3 (shown in Eq. 5). Consequently, for AMO-D, 1-mol NO3 − reduction needs 5/4 mol CH4 and 5/4 mol O2, while for ANMO-D, only 5/8 mol CH4 is needed. The electron flow pattern for ClO4 − is the same as that for NO3 −. Based on Eqs. 1–6 and electron acceptors’ concentrations in the membrane biofilm reactor (MBfR) influent and effluent, we calculated the DO demand for two pathways.
Perchlorate reduction process, especially chlorite dismutation, is similar to ANMO-D, since they both involve a step to produce oxygen. During ClO4 − reduction, perchlorate-reducing bacteria (PRB) reduce ClO4 − to ClO2 − using perchlorate reductase (pcrA). Disproportion of ClO2 − to Cl− and O2 is carried out by a chlorite dismutase enzyme (cld) (Bender et al. 2005).
Here, we evaluate which microorganisms were responsible for reducing NO3 − and ClO4 − and their likely metabolic pathways. Specifically, we use high-throughput sequencing and principal component analysis (PCoA) to identify the key microorganisms responsible for ANMO-D and ANMO-PR and important relationships between the community structure and the fluxes of the different electron acceptors. To understand the likely metabolic pathways, we first review pathways of methane oxidation coupled to denitrification or perchlorate reduction.
Materials and methods
Inoculum
The inoculum was from an ANMO-D culture donated by Dr. Wei Xiang Wu at Zhejiang University (China) and maintained anoxic (dissolved oxygen concentration ≤0.2 mg/L) in a sequencing batch reactor in which the inoculum was able to reduce NO3 − at 7 mg N/L day. Using this inoculum, Luo et al. (2015) achieved complete perchlorate reduction in a CH4-based MBfR.
MBfR configuration and performance
Detailed information about the ANMO-PR MBfR is given by Luo et al (2015). In brief, the MBfR was composed of two glass tubes that held bundles of composite hollow fibers (composite polyethylene fibers, 280 μm o.d., and a 180 μm i.d., pore size 0.1–0.15 μm) manufactured by Mitsubishi Rayon (Model MHF-200TL, Mitsubishi, Ltd., Japan). One tube held the main bundle of 32 fibers, and the second tube held 10 fibers used for biofilm sampling. The liquid contents (65 mL) were completely mixed by recirculation with a peristaltic pump (Longer Pump, model 1515X, Longer Precision Pump Co., Ltd., China) at 100 mL/min. The influent feeding rate was 0.5 mL/min, giving a hydraulic residence time (HRT) of 130 min. Influent and effluent samples were assayed for NO3 −, NO2 −, ClO4 −, and dissolved O2 by the methods reported in Luo et al. (2015).
To accumulate biomass, we fed the MBfR with 2 mg N/L of NO2 − continuously for 40 days, when complete NO2 − reduction was achieved. The MBfR was then operated for eight experimental stages: stages 1 and 7 were fed with ClO4 − and NO2 −; stages 3, 4, 5, and 8 were fed with ClO4 − and NO3 −; and stages 2 and 6 were fed with ClO4 − only. The actual influent and effluent concentrations of all stages were in Luo et al (2015); we list the surface loading and fluxes of all stages in Table 1. We moved to a new stage once acceptor-removal rates reached steady state (effluent concentrations having <10 % variation for a minimum of three HRTs). Actually, it took less than 2 days for all the stages to reach a steady state, except stage 4, which took almost 2 weeks to reach a steady state. The feed medium was de-gassed with N2 for at least 15 min to maintain an anaerobic condition (DO ≤ 0.2 mg/L, actually anoxic), after which the pH was adjusted to 7.0 ± 0.2 with hydrochloric acid. The CH4 pressure was 10 psig (1.7-atm absolute pressure) for stages 1–4 and 15 psig (2 atm absolute) for stages 5–8. The DO concentration was ~0.2 mg/L for the influent and ≤0.1 mg/L for the effluent. The experiment was conducted at a temperature of 29 ± 1 °C for all stages.
Biofilm microbial ecology analyses
We collected biofilm samples when the reactor reached steady state for all stages except stage 2. We cut off one ~10-cm-long section from one of the coupon fiber, then sealed the remaining fiber by tying the end into a knot. N2 gas was sparged at the sampling point to minimize any effects of O2 exposure during sampling. We then extracted DNA using the DNeasy Blood and Tissue Kit (Qiagen, USA) as previously described by Zhao et al. (2011).
DNA samples were sent to Shanghai Majorbio Technology (Shanghai, China) to perform amplicon pyrosequencing with standard Illumina MiSeq sequencing protocols. We used primers 338F (5′-ACTCCTACGGGAGGCAGC-3′) and 806R (5′-CCGTCAATTCMTTTRAGTTT-3′) to target the conserved V3–V5 regions of the bacteria 16S ribosomal ribonucleic acid (rRNA) gene. We processed the data analysis using QIIME (version 1.17) pipeline as described previously (Lai et al. 2014). Operational units (OTUs) with 97 % similarity cutoff were clustered using UPARSE (version 7.1) after chimeric sequences were removed by HCHIME (Edgar 2010). We analyzed the phylogenetic affiliation of representative sequences by RDP Classifier against the silva (SSU115)16S rRNA database using confidence threshold of 70 % (DeSantis et al. 2006; Caporaso et al. 2010). All sequences shorter than 200 bps, having homopolymers of 6 bps and primer mismatches, and a quality score lower than 25 were removed. Since all of the DNA samples were treated in same condition at the same time, we used the sequence numbers of each OTUs, with a total of 260,099 high-quality sequences for the 16S rRNA gene for all stages, to quantify the relative abundance of microbial communities in all the stages. But, the 16S rRNA gene sequence numbers for different species are various; thus, the quantification only represents estimation.
We evaluated the overall community composition using the unweighted UniFrac distance matrix and the relationships among samples with Cytoscape and PCoA (Lozupone et al. 2006; Shannon et al. 2003). The PCoA is a scaling or ordination method that starts with a matrix of similarities (close) or dissimilarities (distance) between a set of individuals and produce a low-dimensional graphical, in which the distances between points in the plot are close to original dissimilarities. We used MEGA 6 to align functional gene nucleotide sequences and the neighbor-joining algorithm of ARB and MEGA 6 program packages to generate the phylogenetic trees based on distance analysis for 16S rRNA gene afterward (Tamura et al. 2013). The robustness of inferred topologies was tested by bootstrap re-sampling using the same distance model (1000 replicates).
All sequencing data were deposited into NCBI with accession numbers SAMO03468106 for stage 0, SAMO03468114 for stage 1, SAMO03470077 for stage 3, SAMO03470079-80 for stages 4 and 5, SAMO03470084 for stage 6, and SAMO03470086-87 for stages 7 and 8.
Results and discussion
Summary of removal performance of electron acceptors
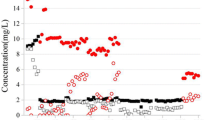
Table 1 summarizes the performance of the CH4-based MBfR. In brief, the CH4-delivery capacity exceeds the CH4 demand in stages 1, 2, 3, 5, 6, and 7. In stages 2 and 6, perchlorate was the only electron acceptor, and the biofilm was able to reduce up to ~5 mg/L of ClO4 − to a non-detectable level (Luo et al. 2015). Stages 1and 7 demonstrated that the presence of ≥0.10 g N/m2 day of NO2 − inhibited ClO4 − reduction, although NO2 − itself was completely reduced. They also demonstrated that NO3 − had no effect on ClO4 − reduction when its loading was <0.302 g N/m2 day. When the CH4-delivery capacity was not much larger than the CH4 demand (stages 4 and 8), the presence of NO3 − inhibited ClO4 − reduction due to electron-donor competition (Luo et al. 2015).
Microbial community change
We used pyrosequencing targeting the V3–V5 regions of the 16S rRNA gene to analyze the diversity and structure of the MBfR bacterial communities in the MBfR biofilm samples. Pyrosequencing of the 16S rRNA gene yielded a total of 658,054 sequences with a median length of 468 bp for all biofilm samples of all stages.
Figure 1 shows the unweighted UniFrac analysis of the biofilm samples for all stages except stage 2. Stages having similar influent loadings are marked with the same color stage label. Once ClO4 − was added in the first stage, the biofilm community structure significantly changed, since stage 0 was very distinct from stage 1 and all other stages. Clearly, the ANMO-D microbial community was greatly changed in the biofilm after the addition of ClO4 −.
Clustering based on the unweighted UniFrac analysis of the microbial community structure at the class level (relative abundances of dominant microbial phylotypes). Stage 0 was the inoculum. Stages 1 and 7 were fed with ClO4 − and NO2 −; stages 3, 4, 5, and 8 were fed with ClO4 − and NO3 −, and stage 6 was fed with ClO4 − only
Figure 2 shows the unweighted PCoA based on the absence or presence of phylotypes. The most important trends is that samples for stages 1 to 8 grouped together, having much lower PC2 values compared to stage 0. The stages 5 and 6 were grouped comparatively distant because of much higher perchlorate loading in stage 6 than stage 5. Thus, the PCoA analysis reinforces that introducing ClO4 − had the highest impact on shaping the microbial community structure. Lai et al. reported that an autotrophic nitrate-reducing community was significantly changed after the introduction of selenite (Lai et al. 2014). Ontiveros-Valencia et al. found that high SO4 2− flux reshaped the original NO3 −-reducing microbial community in a H2-based MBfR (Ontiveros-Valencia et al. 2014).
Principle coordinate analysis (PcoA) based on the unweighted UniFrac analysis showing the microbial community groupings. The red oval is for all MBfR biofilm samples, and the blue oval is for biofilm samples with the highest total electron acceptors loadings (stages 4–8 refer to the total surface loadings in Table 1). The inoculum community (stage 0) was very different from all other stages
The biofilm samples from stages 4 to 8 were close to each other along the PC1 vector, while the biofilm samples from stages 1 and 3 were distant. Compared to the latter stages, stages 1 and 3 had much lower average total acceptor loadings (<8 mmol CH4/m2 day) so that competition for the electron donor was not strong (Table 1). Therefore, the availability of the electron donor significantly affected the microbial community. Probably, only the bacteria that were competitive in an environment that had limited electron donor survived in stages 4 to 8 (>25 mmol CH4/m2 day, except for stage 6).
Another significant difference between the inoculum (stage 0) and the stages with perchlorate addition is with Archaea. Archaea contain the functional methylcoenzyme M reductase (coded by mcrA gene) that conducts the reverse methanogenesis supporting ANMO-D. Figure S1 shows that Archaea were present in stage 0 at low abundance compared to bacteria but Archaea were absent in latter stages. The result is consistent with our finding that the mcrA gene (very low abundance) had no correlation with genes associated with respiration of NO3 − or ClO4 − (Luo et al. 2015).
The phylogenetic tree analysis
Figure 3 shows the phylogenetic tree constructed from the bacterial communities in the MBfR biofilm and selected known perchlorate-reducing bacteria (PRBs), methanol-assimilating denitrifiers (MADs), methanotrophs containing denitrifying genes (AMO-D), and nitrite-dependent anaerobic methane oxidizing bacteria (n-DAMO).
Phylogenetic analysis of the predominant bacterial genera in the MBfR biofilm (stages 1 to 8) (black), known PRBs (blue, most of them reduce NO3 − as well), bacteria from phylum NC10 (red), known methanol-assimilating denitrifers (green), and methanotrophs known to have denitrifying gene content (orange). An MBfR genus with bold and underline was among the relatively most abundant (≥10 %) in one of the stages; detailed relative abundance was shown in the supporting information Fig. S2
Detected genuses Methylomonas and Methylocystis are phylogenetically close to the known AMO-D Methylomonas methanica, Methylomonas koyamae, and Methylocystis trichosporium. Methylomonas was almost absent in stages 0 to 3 but became important from stage 4 (~10 %), when electron-acceptor fluxes greatly increased, but were >15 % of bacterial abundance in stages 5 and 8, which had higher CH4 pressure and total acceptor flux. Methylocystis was remarkably enriched in stage 1, started to decrease in stage 4, but remained important through stage 8 (relative abundance range from 4 to 10 %). The enrichment of AMO-D Methylomonas and Methylocystis in the MBfR probably was caused by the significant delivery of CH4 through the MBfR fibers; they initiated methane oxidation using the limited DO in the influent and respired NO3 − for energy generation (Luo et al. 2015; Tang et al. 2012). Dam et al. reported that Methylocystis sp. strain SC2 was able to reduce NO3 − to N2 with DO-limiting (0.4 mg/L) condition (Dam et al. 2013). Very recently, Dimitri et al. reported that Methylomonas denitrificans sp. nov., strain FJG1T coupled methane oxidation to reduction of NO3 − to N2O with very low DO (1.6 μg/L) concentration (Dimitri et al. 2015).
Methylophilus are phylogenetically close to known methanol-assimilation denitrifiers (MA-D, green color in Fig. 3), such as Methylophilus quaylei, Methylotenera mobilis, Methyloversatilis universalis, and to selected known PRBs (blue color in Fig. 3), such as Dechloromonas agitate, Dechloromonas aromatic, and Azospira restricta. While Methylophilus was almost absent in stage 0, it became important in stage 1 after perchlorate was introduced, reached ~30 % of bacterial abundance in stage 5, and then decreased sharply. The increase of the Methylophilus abundance in the first five stages may have been due to the accumulation of methanotrophs that provided organic electron donors, such as methanol, acetate, and amino acids. Ginige et al. and Doronina et al. reported that Methylophilus can use simple organics, such as methanol and methylamine, to reduce NO3 − in anoxic conditions, but they cannot grow on methane (Ginige et al. 2004; Doronina et al. 2005).
Pelomonas started to grow in stage 3, and became dominant from stages 4 to 8, consistently following the development of Methylomonas. The genera of Pelomonas is phylogenetically close to known PRBs, e.g., Dechloromonas agitate CKB and Dechloromonas aromatic RCB. The accumulation of Pelomonas may have been due the increasing flux of perchlorate and organic donors produced by methanotrophs. Dechloromonas is the main genera known to reduce NO3 − and ClO4 − in the MBfR (Nerenberg et al. 2008), where its abundance increased with increasing ClO4 − loading relative to NO3 −.
Uncultured Chloroflexi existed in all the stages and were significantly enriched after stage 6. The phylotype of uncultured Chloroflexi is phylogenetically close to bacteria in the NC10 phylum, e.g., M. oxyfera. Therefore, the uncultured Chloroflexi may work similarly like NC10 phylum bacteria in the ANMO-D processes. In fact, Ettwig et al. (2009) reported that Chloroflexi existed in M. oxyfera enrichment culture that performed n-DAMO. Beal et al. and Hu et al. found that Chloroflexi present during anaerobic methane oxidation were coupled to manganese and iron-reducing culture and a reactor containing ammonium oxidation (anammox) and ANMO-D processes (Beal et al. 2009; Hu et al. 2015).
Metabolic pathways based on the DO and methane/N ratio analysis
Although the AMO-D (AMO-PR) and MA-D (MA-PR) bacteria were important in the biofilm samples, they may not be the only methane oxidation manners in MBfR. Figure 4 shows the DO demand for both computed aerobic-type and intra-aerobic-type anaerobic methane oxidation coupled to denitrification and perchlorate reduction. Clearly, the actual DO concentrations in the MBfR system were much lower than the theoretical demand for aerobic methane oxidation. Ettwig et al. reported that during the intra-aerobic-type ANMO-D pathway, the oxygen produced by M. oxyfera intracellularly for methane mono-oxygenation, was not detectable (Ettwig et al. 2010). Considering the conflict between actual DO concentrations and theoretical demand, we propose that beside AMO-D (AMO-PR) and MA-D (MA-PR), the ANMO-D (ANMO-PR) were also important in all the stages except stage 2 in the MBfR system (Fig. 5). To quantify the comprehensive processes of different electron acceptor/donor utilizations, further studies on the effect of oxygen and microbial community are required.
The calculated oxygen required for different metabolic pathways. Max methane supply is the theoretical maximum methane supply through the MBfR (decrease because considering the membrane surface area loss due sampling); it gradually declines by stage due to volume loss through sample. Methane demand for aerobic methanotroph is the methane required for aerobic methane oxidation coupled to denitrification and perchlorate reduction. Oxygen demand for aerobic methanotrophs is the oxygen required for aerobic methane oxidation coupled to denitrification and perchlorate reduction. The dashed line represents the actual average DO concentration along all stages
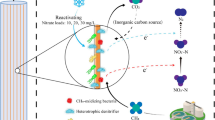
Proposed microbial community functions in the CH4-based MBfR. MA-D methanol-assimilating denitrification reduction, MA-PR methanol-assimilating perchorate reduction, AMO-D aerobic methane oxidation coupled to denitrification perchlorate reduction, AMP-PR aerobic methane oxidation coupled to perchlorate reduction, ANMO-D anaerobic methane oxidation coupled to denitrification, and ANMO-PR anaerobic methane oxidation coupled to perchlorate reduction
Besides the DO concentration, we analyzed the CH4/NO3 − mole ratios through all stages. When CH4 delivery was limited at stages 4 and 8, the calculated C/N ratios values were of 1.52 and 1.15, respectively. Both of the two observed C/N ratios were much lower than the minimum ration for AMO-D. Thus, the ANMO-D (ANMO-PR) might also play important roles in the MBfR.
Overall, we propose that in the MBfR system, the AMO-D (AMO-PR) directly oxidize methane and reduce NO3 − to NO2 − or N2O under anoxic condition, producing organic matters for MA-D (MA-PR) to reduce NO3 −. Simultaneously, the ANMO-D (ANMO-PR) bacteria use methane as electron donor to respire NO3 − or ClO4 − directly. The metabolic intermediates of the AMO-D (AMO-PR) may improve the metabolism of ANMO-D (ANMO-PR). For example, Wu et al. 2015 reported that the genome of “M. oxyfera” lacks known pyrroloquinoline quinone (PQQ) biosynthesis machinery, which is necessary during the methane metabolism. However, the PQQ biosynthesis exists in aerobic methanotrophs, so AMO-D (AMO-PR) could possibly supply PQQ for ANMO-D (ANMO-PR).
In summary, we found that the microbial community of an ANMO-D culture significantly changed after it was inoculated to the CH4-based MBfR reducing perchlorate, nitrate, and nitrite. Original, the ANMO-D culture was dominated by uncultured Anaerolineaceae and Ferruginibacter. After being introduced to the anoxic MBfR fed with nitrite and perchlorate initially, all Archaea were removed out, the abundances of both uncultured Anaerolineaceae and Ferruginibacter significantly decreased, while the anoxic methanotrophs greatly increased, especially Methylocystis. With the development of methanotrophs, the methanol-assimilating denitrifying bacteria, e.g., Methloversatilis and Methylophilus, became important in the biofilm. When nitrate loading increased to 0.75 g N/m2 day in stage 4, the genus Pelomonas accounted ~15 % of total bacteria and remained >10 % to the end, while another methanotrophs Methylomonas accounted >10 % of total bacteria. The genera Azospirillum significantly increased in stage 6 when perhchlorate loading was up to 0.39 mg/m2 day. Based on the DO and C/N ratio analyses, we propose that beside anoxic AMO-D, AMO-PR processes, MA-D, and MA-PR processes, the anaerobic ANMO-D and ANMO-PR processes played important roles in the methane oxidation coupled to multiple electron acceptors reduction in the MBfR.
References
Beal EJ, House CH, Orphan VJ (2009) Manganese- and iron-dependent marine methane oxidation. Science 325:184–187
Bender KS, Shang C, Chakraborty R, Belchik SM, Coates JD, Achenbach LA (2005) Identification, characterization, and classification of genes encoding perchlorate reductase. J Bacteriol 187:5090–5096
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R (2010) Pynast: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26:266–267
Chaudhuri SK, O’Connor SM, Gustavson RL, Achenbach LA, Coates JD (2002) Environmental factors that control microbial perchlorate reduction. Appl Environ Microbiol 68:4425–4430
Coates JD, Achenbach LA (2004) Microbial perchlorate reduction: rocket-fuelled metabolism. Nat Rev 2:569–580
Coates JD, Michaelidou U, Bruce RA, O’Connor SM, Crespi JN, Achenbach LA (1999) Ubiquity and diversity of dissimilatory (per)chlorate-reducing bacteria. Appl Environ Microbiol 65:5234–5241
Dam B, Dam S, Blom J, Liesack W (2013) Genome analysis coupled with physiological studies reveals a diverse nitrogen metabolism in Methylocystis sp. strain SC2. Plos One. doi:10.1371/journal.pone.0074767
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16s rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
Dimitri Kits K, Klotz MG, Stein LY (2015) Methane oxidation coupled to nitrate reduction under hypoxia by the Gammaproteobacteria Methylomonas denitrificans sp. nov., type strain FJG1. Environ Microbiol. doi:10.1111/1462-2920.12772
Doronina N, Ivanova E, Trotsenkoa Y, Pshenichnikovab A, Kalininab E, Shvetsb V (2005) Methylophilus quaylei sp. nov., a new aerobic obligately methylotrophic bacterium. Syst Appl Microbiol 28:303–309
Edgar RC (2010) Search and clustering orders of magnitude faster than blast. Bioinformatics 26:2460–2461
Eisentraeger A, Klag P, Vansbotter B, Heymann E, Dott W (2001) Denitrification of groundwater with methane as sole hydrogen donor. Water Res 35:2261–2267
Ettwig KF, van Alen T, van de Pas-Schoonen KT, Jetten MSM, Strous M (2009) Enrichment and molecular detection of denitrifying methanotrophic bacteria of the NC10 phylum. Appl Environ Microbiol 75:3656–3662
Ettwig KF, Butler MK, Le Paslier D, Pelletier E, Mangenot S, Kuypers MMM, Schreiber F, Dutilh BE, Zedelius J, de Beer D, Gloerich J, Wessels HJCT, van Alen T, Luesken F, Wu ML, van de Pas-Schoonen KT, Op den Camp HJM, Janssen-Megens EM, Francoijs KJ, Stunnenberg H, Weissenbach J, Jetten MSM, Strous M (2010) Nitrite-driven anaerobic methane oxidation by oxygenic bacteria. Nature 464:544–548
Ginige MP, Hugenholtz P, Daims H, Wagner M, Keller J, Blackall LL (2004) Use of stable-isotope probing, full-cycle rRNA analysis, and fluorescence in situ hybridization-microautoradiography to study a methanol-fed denitrifying microbial community. Appl Environ Microbiol 70:588–596
Haroon MF, Hu SH, Shi Y, Imelfort M, Keller J, Hugenholtz P, Yuan ZG (2013) Anaerobic oxidation of methane coupled to nitrate reduction in anovel archaeal lineage. Nature 500:567–570
He ZF, Geng S, Shen LD, Lou LP, Zheng P, Xu XY, Hu BL (2015) The short- and long-term effects of environmental conditions on anaerobic methane oxidation coupled to nitrite reduction. Water Res 68:554–562
Hu S, Zeng RJ, Burow LC, Lant P, Keller J, Yuan ZG (2009) Enrichment of denitrifying anaerobic methane oxidizing microorganisms. Environ Microbiol Rep 1:845–854
Hu SH, Raymond JZ, Mohamed FH, Jurg K, Paul AL, Gene WT, Yuan ZG (2015) A laboratory investigation of interactions between denitrifying anaerobic methane oxidation (DAMO) and anammox processes in anoxic environments. Sci Rep 5:8706. doi:10.1038/srep08706
Knowles R (2005) Denitrifiers associated with methanotrophs and their potential impact on the nitrogen cycle. Ecol Eng 24:441–446
Lai CY, Yang XE, Tang YN, Rittmann BE, Zhao HP (2014) Nitrate shaped the selenate-reducing microbial community in a hydrogen-based biofilm reactor. Environ Sci Technol 48:3395–3402
Logan BE, Zhang H, Mulvaney P, Milner MG, Head IM, Unz RF (2001) Kinetics of perchlorate- and chlorate-respiring bacteria. Appl Environ Microbiol 67:2499–2506
Lozupone C, Hamady M, Knight R (2006) UniFrac—an online tool for comparing microbial community diversity in a phylogenetic context. BMC Bioinf 7:371–385
Luo YH, Chen R, Wen LL, Meng F, Lai CY, Rittmann BE, Zhao HP, Zhen P (2015) Complete perchlorate reduction using methane as the sole electron donor and carbon source. Environ Sci Technol 49:2341–2349
Modin O, Fukushi K, Yamamoto K (2007) Denitrification with methane as external carbon source. Water Res 41:2726–2738
Modin O, Fukushi K, Nakajima F, Yamamoto K (2010) Nitrate removal and biofilm characteristics in methanotrophic membrane biofilm reactors with various gas supply regimes. Water Res 44:85–96
Nerenberg R, Rittmann BE, Najm I (2002) Perchlorate reduction in a hydrogen-based membrane-biofilm reactor. J Am Water Works Assoc 94:103–114
Nerenberg R, Kawagoshi Y, Rittmann BE (2008) Microbial ecology of a perchlorate-reducing, hydrogen-based membrane biofilm reactor. Water Res 42:1151–1159
Ontiveros-Valencia A, Tang YN, Zhao HP, Friese D, Ryan O, Smith J, Evans P, Rittmann BE, Krajmalnik-Brown R (2014) Pyrosequencing analysis yields comprehensive assessment of microbial communities in pilot-scale two-stage membrane biofilm reactors. Environ Sci Technol 48:7511–7518
Osaka T, Yoshie S, Tsuneda S, Hirata A, Iwami N, Inamori Y (2006) Identification of acetate- or methanol-assimilating bacteria under nitrate-reducing conditions by stable-isotope probing. Microb Ecol 52:253–266
Raghoebarsing AA, Pol A, van de Pas-Schoonen KT, Smolders AJP, Ettwig KF, Rijpstra WIC, Schouten S, Sinninghe Damste JS, Op den Camp HJM, Jetten MSM, Strous M (2006) A microbial consortium couples anaerobic methane oxidation to denitrification. Nature 440:918–921
Rikken GB, Kroon AGM, van Ginkel CG (1996) Transformation of (per)chlorate into chloride by a newly isolated bacterium: reduction and dismutation. Appl Microbiol Biotechnol 45:420–426
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Sun FY, Dong WY, Shao MF, Lv XM, Li J, Peng LY, Wang HJ (2013) Aerobic methane oxidation coupled to denitrification in a membrane biofilm reactor: treatment performance and the effect of oxygen ventilation. Bioresour Technol 146:2–9
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tang Y, Zhao HP, Marcus A, Krajmalnik-Brown R, Rittmann BE (2012) A steady-state-biofilm model for simultaneous reduction of nitrate and perchlorate—part 2: parameter optimization and results and discussion. Environ Sci Technol 46:1608–1615
United States Environmental Protection Agency (2001) Record of decision of the Western http://www.epa.gov/superfund/sites/rods/fulltext/r0901535.pdf
United States Environmental Protection Agency (2015) Basic information about nitrate in drinking water. http://water.epa.gov/drink/contaminants/basicinformation/nitrate.cfm
Water Resource Foundation (2015). Perchlorate in drinking water regulatory update and treatment options. http://www.waterrf.org/resources/stateofthesciencereports/perchlorate_stateofthescience.pdf
Wu ML, Ettwig KF, Jetten MSM, Strous M, Keltjens JT, van Niftrik LA (2011) New intra-aerobic metabolism in the nitrite-dependent anaerobic methane-oxidizing bacterium Candidatus Methylomirabilis oxyfera. Biochem Soc Trans 39:243–248
Wu ML, Wessels HJCT, Pol A, den Camp JMO, Jetten MSM, von Niftrik L, Keltjens JT (2015) Xoxf-Type methanol dehydrogenase from the anaerobic methanotroph “Candidatus Methylomirabilis oxygera”. Appl Environ Microb 81:1442–1451
Yao S, Ni J, Zhao S, Qiang C, Zhang H, Wang S (2013) COD and nitrogen removal in facilitated transfer membrane-aerated biofilm reactor (FT-MABR). J Membr Sci 389:257–264
Zhao HP, Van Ginkel S, Tang Y, Kang DW, Rittmann BE, Krajmalnik-Brown R (2011) Interactions between perchlorate and nitrate reductions in the biofilm of a hydrogen-based membrane biofilm reactor. Environ Sci Technol 45:10155–10162
Acknowledgments
Authors greatly thank the “National Key Technology R&D Program (2014ZX07101-012),” the “National Natural Science Foundation of China (Grant Nos. 21377109 and 21577123),” and the “National High Technology Research and Development Program of China (2013AA06A205)” for their financial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Gerald Thouand
Ran Chen and Yi-Hao Luo contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 155 kb)
Rights and permissions
About this article
Cite this article
Chen, R., Luo, YH., Chen, JX. et al. Evolution of the microbial community of the biofilm in a methane-based membrane biofilm reactor reducing multiple electron acceptors. Environ Sci Pollut Res 23, 9540–9548 (2016). https://doi.org/10.1007/s11356-016-6146-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-016-6146-y