Abstract
Perchlorate is a widespread drinking water contaminant with regulatory standards ranging from 2 to 18 μg/L. The hydrogen-based membrane-biofilm reactor (MBfR) can effectively reduce perchlorate, but it is challenging to achieve low-µg/L levels. We explored chlorate addition to increase the abundance of perchlorate-reducing bacteria (PRB) and improve removals. MBfR reactors were operated with and without chlorate addition. Results show that chlorate doubled the abundance of putative PRB (e.g., Rhodocyclales) and improved perchlorate reduction to 23 ± 17 µg/L, compared to 53 ± 37 µg/L in the control. Sulfate reduction was substantially inhibited during chlorate addition, but quickly recovered once suspended. Our results suggest that chlorate addition can enhance perchlorate reduction by providing a selective pressure for PRB. It also decreases net sulfate reduction.
Key points
• Chlorate increased the abundance of perchlorate-reducing bacteria
• Chlorate addition improved perchlorate removal
• Chlorate appeared to suppress sulfate reduction
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Perchlorate (ClO4−) is a ubiquitous water contaminant that is highly soluble and stable. It can inhibit thyroid function (Srinivasan and Sorial 2009), and European and US state-level standards for ClO4− in drinking water range from 2 to 15 μg/L (ANSES 2012; CA DEP 2007; EFSA CONTAM Panel 2014; MA DEP 2006). Reverse osmosis (RO) and ion exchange (IX) can remove ClO4−, but produce high-strength ClO4− concentrates that requires further treatment or disposal. Microbial reduction of ClO4− by perchlorate-reducing bacteria (PRB) is a more promising strategy, as it reduces ClO4− to innocuous chloride (Cl−).
PRB sequentially reduce ClO4− to chlorate (ClO3−) and chlorite (ClO2−) via the perchlorate reductase (pcr) enzyme. Then ClO2− is transformed into chloride (Cl−) and oxygen (O2) by the chlorite dismutase (cld) enzyme. The produced O2 is further utilized by PRB as an electron acceptor (Coates and Achenbach 2004; Ma et al. 2016; Sijimol et al. 2015; Srinivasan and Sorial 2009; Ye et al. 2012). Many PRB use organic electron donors, such as acetate and lactate, but some also use and inorganic electron donors, such as hydrogen and reduced sulfur compounds. Inorganic electron donors are often considered more environmentally friendly, due to their low carbon footprint (Choe et al. 2013).
Biofilm processes are ideal for biological water treatment, as they retain high amounts of biomass in the reactor and minimize downstream treatment. In particular, the hydrogen-based membrane-biofilm reactor (H2-MBfR) has been successfully applied for denitrification (Martin and Nerenberg 2012) and ClO4− reduction (Nerenberg et al. 2008; Ontiveros-Valencia et al. 2013, 2014a). In this type of biofilm reactor, the control of the biomass accumulation and management of the microbial community is necessary to achieve successful ClO4− reduction (Ma et al. 2016).
PRB are ubiquitous in the natural environment, even in pristine areas without anthropogenic perchlorate contamination (Coates et al. 1999). This is probably because PRB have a versatile metabolism, using oxygen, nitrate, perchlorate, and chlorate as electron acceptors, which are widespread compounds. Also, perchlorate and chlorate occur naturally and are ubiquitous in the environment at trace concentrations (Dasgupta et al. 2005; Zhang et al. 2021). However, ClO4− is a highly energetic electron acceptor and can provide a strong selective pressure for PRB. For example, Nerenberg et al. (2008) found that the relative abundance of PRB in a denitrifying H2-MBfR was around 13% when the bulk ClO4− was approximately 10 µg/L. However, the relative abundance increased to nearly 50% when the bulk ClO4− concentration was around 5 mg/L.
Biological perchlorate reduction has several challenges. Since the influent ClO4− concentrations are low, typically less than 100 µg/L, and usually need to be removed to below 15 µg/L, there is a weak selective pressure for PRB. In part, this is because the ClO4− reduction rates are significantly lower at such low concentrations (Nerenberg et al. 2006). Also, O2, nitrate (NO3−), and sulfate (SO42−) are commonly present along with perchlorate. O2 and NO3− are preferred acceptors and inhibit microbial ClO4− reduction by PRB; thus, low NO3− and O2 levels are needed (Bardiya and Bae 2011; Tang et al. 2012). However, low levels of O2 and NO3− favor sulfate-reducing bacteria (SRB) (Ontiveros-Valencia et al. 2013, 2014a). Reduction of SO42− is undesirable, since SRB do not reduce perchlorate, and also produce H2S—a toxic, corrosive, and malodorous compound.
Studies of H2-MBfRs for ClO4− reduction in the presence of NO3− and SO42− showed SO42− reduction from 10 to 80%, and also difficulties in achieving low effluent ClO4− concentrations in a one-stage H2-MBfR when the influent ClO4− was above 1 mg/L (Ontiveros-Valencia et al. 2013, 2014a). These studies, which showed incomplete ClO4− reduction, suggest that SRB have an ecological advantage over PRB, as the higher sulfate concentrations can fully penetrate the biofilm. This allows SRB to grow in the inner biofilm layers, far from the detachment zone, separate from faster growing denitrifiers, and closer to the H2 source (Ontiveros-Valencia et al. 2013).
In order to enhance ClO4− reduction, the microbial community can be manipulated to favor PRB. Increasing the abundance of PRB should reduce the effluent ClO4− concentration. Adding ClO4− to the reactor influent would help select for PRB, but this would have to be done while the reactor is off-line, and there still would be concerns about adding a toxic compound that would require special handling and control of any discharges. A better strategy would be adding a less toxic compound that selects for PRB.
ClO3− addition is a potential option. It is an intermediate in the ClO4− reduction pathway and transformed by the pcr enzyme. Furthermore, PRB grow faster on ClO3− than on ClO4− (Bardiya and Bae 2011). For example, Nerenberg et al. (2006) found that, for a pure culture of PRB, Dechloromonas sp. PC1, the specific growth rate for ClO3− was double that for ClO4−. Also, the half saturation constant, K, for ClO3− was below 0.014 mg/L, compared to 0.14 mg/L for ClO4−, meaning that high growth rates could be obtained at very low ClO3− concentrations (Nerenberg et al. 2006). Furthermore, the recommended level for ClO3− in drinking water is 0.7 mg/L, around two orders of magnitude higher than that for perchlorate (WHO 2016).
Based on the above, ClO3− may provide a strong selective pressure for PRB. In addition, ClO3− is known as an inhibitor of SRB (Carlson et al. 2015; Engelbrektson et al. 2014, 2018). Thus, ClO3− could both help select for PRB and inhibit SRB. However, the effect of ClO3− on perchlorate-reducing biofilms has not been studied. Also, a potential concern with ClO3− is that it could select for chlorate-reducing bacteria (CRB), which are distinct from PRB and do not reduce ClO4− (Bardiya and Bae 2011).
The goal of this study was to analyze the effects of ClO3− addition on the development of the microbial community during the reduction of ClO4 in the presence of O2, NO3−, and SO42− at relevant drinking water concentrations. Two types of reactors were used. One type included a bundle of hollow fiber membranes, mimicking an actual MBfR reactor. The other had a single membrane, where the effluent concentration was essentially equal to the influent. This allowed the bulk substrate concentrations to be maintained at a desired level.
Materials and methods
Reactor configuration and culture medium
We operated two identical H2-MBfRs: reactors S1 and S2 (single membrane), and reactors M1 and M2 (multiple membranes). Reactors S1 and S2 were used to determine the effect of ClO3− on PRB development, whereas reactors M1 and M2 were used to determine the effect of ClO3− on ClO4− reduction in a system mimicking a real treatment system.
Reactors S1 and S2 consisted of two glass tubes with a single membrane (Mitsubishi MHF200TL) joined together with Norprene tubing and connectors. One tube was considered the main membrane and the other a “coupon” for biofilm samples. The membranes had a combined total length of 70 cm and were connected to the H2 source at one end and open at the other. After cutting a piece for biofilm sampling, the coupon membrane remained with an open end. To prevent excessive gas venting, a solenoid valve connected to a timer (ChronTrol) opened the H2 inlet for 30 s every 29.5 min. The H2 pressure was set to 1 psi for the duration of the experiment, except from days 33 to 36, when the pressure increased to 6 psi.
Reactors M1 and M2 consisted of two glass tubes connected with Norprene tubing. One glass tube contained a bundle of 31 membranes (Mitsubishi MHF200TL) (“main bundle”) and the other had three membranes (Mitsubishi MHF200TL) to allow biofilm sample collection (“coupon”). The membranes in both tubes were 23.5 cm long and were connected to the H2 line at one end and sealed at the other. A schematic of the setup is shown in Fig. 1 and operational parameters are summarized in Table 1.
Schematic of the experimental systems. a Reactors S1 and S2 consisted of two glass tubes with single membrane. One of these membranes was used for biofilm sampling (“coupon”). Hydrogen was supplied from one end of the membranes and the other end was opened. b M1 and M2 reactors, each consisted of two glass tubes. One tube contained a bundle of 31 membranes (“main bundle”) and the other tube had three membranes for biofilm sampling (“coupon”). Hydrogen was supplied from one end of the membranes, and the other end was sealed
The influent media for reactors S1 and S2 contained the following (g/L): 0.434 Na2HPO4; 0.128 KH2PO4; 0.026 MgSO4 × 7H2O; 0.14 MgCl2 × 6H2O; 0.2 NaHCO3; 0.001 FeSO4 × 7H2O; 0.001 CaCl2; and 1 mL of trace mineral solution. The trace mineral solution consisted of (mg/L): 100 ZnSO4 × 7H2O; 30 MnCl2 × 4H2O; 300 H3BO3; 200 CoCl2 × 6H2O; 10 CuCl2 × 2H2O; 10 NiCl2 × 6H2O; 30 Na2MoO4 × 2H2O; and 30 Na2SeO3 (Nerenberg et al. 2008). The pH of the media was adjusted to 7.5. The influent media for reactors M1 and M2 was the same, except that Mg salts (MgSO4 × 7H2O and MgCl2 × 6H2O) were replaced with 0.2 mg/L of MgSO4 × 7H2O.
Startup and stage experiments
All the reactors were inoculated with 1 mL of activated sludge from the South Bend Wastewater Treatment Plant, South Bend, Indiana, USA. This inoculum was previously divided in aliquots with 20% glycerol and stored at − 80 °C. After inoculation, the reactors were run in recirculation mode for 16 h to allow the biofilm to become established. The operational conditions are described in Table 2. The high recirculation flow rate created well-mixed conditions in the reactor (Nerenberg et al. 2008).
Reactors S1 and S2 were operated in two stages, as described in Table 2. In the first stage, reactors S1 (control) and S2 (experimental) were supplied with an influent of 0.04 mg/L ClO4−, 0.1 mg N/L NO3−, and 10 mg/L SO42− for 22 days. In the second stage, 5 mg/L ClO3− was added continuously to reactor S2 for 14 days. Given the small membrane area, the bulk liquid and effluent concentrations of these reactors were essentially equal to influent. The influent concentrations of NO3− and SO42− were selected to obtain a typical bulk concentrations found in actual MBfR reactors (Nerenberg et al. 2008; Ontiveros-Valencia et al. 2014a, b). The high concentration of ClO3−, relative to of NO3− and SO42−, was expected to exert a selective pressure for PRB. The influent medium of both reactors was purged with N2 gas for 30 min to remove any residual O2.
Reactors M1 (control) and M2 (experimental) were operated in three stages, as described in Table 2. In the first stage, the reactors were supplied with 5 mg N/L NO3−, 80 mg/L SO42−, and 8.4 mg/L O2 for 24 days. ClO4− was supplied at 0.1 mg/L during the first 9 days, and then was increased to 0.6 mg/L. These concentrations were based on the values typically found in drinking water sources and on previous MBfR pilot studies addressing water remediation (Nerenberg et al. 2008; Ontiveros-Valencia et al. 2014b; US EPA 2009; WHO 2004, 2011). Unlike the single membrane reactors, the effluent concentrations in most cases were substantially lower than the influent. In the second stage, after NO3− and ClO4− microbial reduction reached steady state, ClO3− was continuously supplied at 5 mg/L to reactor M2 for 39 days. Steady state was reached when the variation in the effluent concentrations of a particular compound were less than 10% for three or more HRTs. Reactor M1 continued with the same influent media. In the last stage, ClO3− addition was discontinued and reactors M1 and M2 were operated for 16 days.
Analytical methods
The reactor’s effluent was sampled daily, and the influent every 3 days. Samples were filtered with 0.2-µm polyethersulfone (PES) syringe filters before storing at 4 °C. The concentrations of NO3−, ClO4−, ClO3−, and SO42− were analyzed by ion chromatography (DIONEX ICS-2500) with a 4-mm AG20 guard column and a 4-mm AS20 analytical column. Sodium hydroxide (NaOH) was used as eluent at 1 mL/min with a gradient program ranging from 5 to 55 mM NaOH. For samples from reactors S1 and S2, the detection limit for NO3−, ClO3−, and SO42− was 0.015 mg N/L, 50 µg/L, and 0.1 mg/L, respectively. ClO4− was not detected below 50 µg/L. For samples from reactors M1 and M2, the detection limit for NO3−, ClO4−, ClO3−, and SO42− was 0.06 mg N/L, 12 µg/L, 50 µg/L, and 0.6 mg/L, respectively. The influent concentrations for days without sampling were the average of the values from the adjacent sampling dates.
The pH was measured by using a pH meter (Accument® AB250, Fisher Scientific), and dissolved oxygen (DO) in the influent was measured with a DO probe (LOD101, Hach). The DO in the effluent was assumed to be below the detection limit.
Biofilm sampling and DNA extraction
Biofilm samples were collected from Reactors S1 and S2 at the following:
-
i)
Day 22, just before adding ClO3− to S2 (t1)
-
ii)
Day 36, which was after 14 days of ClO3− addition (t2)
For t1 samples, we cut a 21–23-cm long section of the coupon membrane and maintained the membrane with open end. To ensure the sample from the second stage would be representative of the evolution of the biofilm since the beginning; including the ClO3− addition, the t2 sample was taken in duplicate. One sample corresponded to the whole main membrane (~ 35 cm), and the other corresponded to the remaining piece of the coupon membrane.
Biofilm samples from reactors M1 and M2 were collected at the following:
-
i)
Day 21, and just before adding ClO3− to the M2 (t1)
-
ii)
Day 60, and during ClO3− addition and shortly before stopping it (t2)
-
iii)
Day 78, which was 15 days after stopping ClO3− addition (t3)
To sample the biofilm, a 2- to 4-cm section was cut from two coupon membranes (duplicates), and the remaining sections were sealed with a knot to prevent gas leakage.
To separate the biofilm from the membrane, the sampled membrane section was placed in a 2-mL centrifuge tube, vortexed at maximum intensity for 10 min, and centrifuged it at maximum speed for 10 min. DNA was extracted from the obtained pellet with UltraClean™ Microbial DNA Isolation kit (MO BIO Laboratories) according to the manufacturer’s instructions. The concentrate absorbance was measured with a spectrophotometer (NanoDrop 2000, Thermo Scientific) to ensure sufficient DNA quantity and quality. The extracted DNA was analyzed via 16S rRNA sequencing as detailed below. DNA was stored at − 20 °C after extraction and until analysis.
High-throughput sequencing and taxonomic analysis
The microbial community structure was determined by high-throughput sequencing of regions V4 and V5 of the 16S rRNA, using the primers 515F (5′-GTGYCAGCMGCCGCGGTAA-3′) and 926R (5′-CCGYCAATTYMTTTRAGTTT-3′). These are the most reliable regions to represent full length 16S rRNA (Yang et al. 2016). Samples were sent to DNA Services Facility at the University of Illinois at Chicago for sequencing with MiSeq Illumina technology. Taxonomic analysis was performed in QIIME v 1.8 (Caporaso et al. 2011). The lowest number of sequences found was 50,500. These sequences were clustered into operational taxonomic units (OTU) with 97% identity against the Greengenes rRNA database (DeSantis et al. 2006). More details are described in Ontiveros-Valencia et al. (2014a). After the taxonomic analysis, the relative abundances of the duplicate samples—sample at t2 of reactors S1 and S2, and samples at each sampling time of reactors M1 and M2—were averaged. Raw sequences are available at the Sequence Read Archive (SRA) of the National Center for Biotechnology Information (NCBI) under accession number PRJNA606597.
Statistical analysis
The statistical significance of the effect of ClO3− addition on the removal rates of ClO4− and SO42− was determined in stage 2 of reactors M1 and M2 with a one-way repeated measured analysis of variance (ANOVA). We employed SPSS Statistics v.25 to determine the statistical significance of ClO3− addition (reactors M1 and M2) on ClO4− and SO42− reduction. The data from stage 2 (days 24 to 63) from both reactors was analyzed with a one-way repeated measures analysis of variance (ANOVA) using the software SPSS Statistics v.25. The data set from stage 2 excluded data from the days 31 to 34, day 42, and days 48 and 49, as the performance results were affected by leaking and sloughing events.
Results
Reactors S1 and S2—single membrane setup
ClO3 − suppressed SO4 2 − reduction
After 8 to 9 days of operation, and before ClO3− addition to reactor S2 (stage 1), both reactors averaged an overall NO3− effluent concentration of 0.04 mg N/L (~ 65% removal) (Fig. 2). SO42− reduction was always near zero in both H2-MBfRs during both stages except for the end of stage 2 in reactor S1, which is likely explained by the increase on H2 pressure from 1 to 6 psi.
Reduction of NO3 − , SO42 − , and ClO3 − . a Reactor S1 (control). b Reactor S2 (experimental, ClO3 − addition). In the first stage, both reactors were run without ClO3 − . From days 22 to 36, ClO3 − was added to reactor S2. The arrows indicate the days in which the biological samples were taken. From day 33, H2 pressure increased from 1 to 6 psi
ClO3− was reduced from 5 mg/L to an average concentration of 2.4 mg/L in the first week of its addition to reactor S2 (stage 2). From days 33 to 36, ClO3− reduction improved, with the effluent dropping to 0.4 mg/L, which is also explained by the increase on H2 pressure. NO3− reduction decreased at stage 2 in both reactors, probably due to the large amount of membrane removed for biofilm sampling before stage 2. Although it seems that ClO3− affected NO3− reduction, this might be related to its high concentration of e− equivalents compared to NO3− (Table 2). At this stage, NO3− reduction averaged 68% in reactor S1, while it averaged 60% in reactor S2. Moreover, once ClO3− reduction improved to 90%, NO3− reduction dropped to 40%.
Microbial community structure
Reactors S1 and S2 were operated to test if chlorate would select for PRB. Biofilms from S1 and S2 had 80–90% heterotrophic and autotrophic phylotypes that include members as denitrifying bacteria (DNB) and PRB (Fig. 3). The dominant orders were Rhodocyclales, Bulkholderiales, and Rhizobiales, representing 65–70% relative abundance in both reactors. These three orders are very diverse, with members capable of denitrification and ClO4− reduction (Coates et al. 2001; Oren 2014; Willems 2014; Zhao et al. 2011).
Microbial community structure at order level in S1 and S2. Biological samples were taken at 2 times: (i) day 22, before adding chlorate to reactor B (t1); (ii) day 34, after 14 days of chlorate addition (t2). In each reactor, the results at t2 are an average of two samples. The sum of the less abundant phylotypes (< 1%) is classified as “Other”
Both S1 and S2 had similar abundance of Rhodocyclales by the end of stage 1 (17–19%) (Fig. 3). However, after 2 weeks of ClO3− addition to S2 in stage 2 (t2), Rhodocyclales doubled their abundance compared to stage 1 (t1). Moreover, Dechloromonas, one of the two genera of this order present in the microbial community, increased almost four times at stage 2 of S2, while it only increased 1.5 times in S1 (Fig. S1). In addition, it seems that ClO3− slightly enriched for Rhodobacterales, changing from 1% in stage 1 to 3% in stage 2. This order only presented the genus Rhobodacter which is a denitrifying bacteria (Kraft et al. 2014; Zhang et al. 2016), although Roldán et al. (1994) reported species from the genus Rhodobacter capable of ClO3− reduction.
Reactors M1 and M2—multiple fibers setup
Effect of ClO3 − on ClO4 − and SO42 − reduction
In both reactors, NO3− was reduced to below detection after 5 days (Fig. 4). ClO4− was reduced to approximately 12 μg/L (i.e., 95% reduction) on day 9 in M1 and on day 6 in M2. The delay and lower perchlorate reduction in M1 may be due to a leak in the recirculation tubing during the first 5 days of operation.
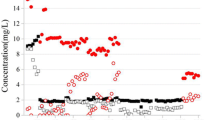
Reduction of NO3 − , SO42 − , ClO4 − and ClO3 − . a Control reactor M1. b Experimental reactor M2 (with ClO3 − addition). In the first stage, both reactors were run identically, without ClO3 − . From days 24 to 63, ClO3 − was added to M2. Both reactors continued to operate for 16 days, without ClO3 − . The dashed lines indicate the limits between each stage (Table 2), and the arrows indicate the days when biofilm samples were collected
SO42− reduction was detected on day 17 after an imposed batch period while the reactors had recirculation problems. During the incident, SO42− effluent concentration values were 71 mg/L and 74 mg/L on day 22 in reactors M1 and M2, respectively. This was detrimental to ClO4− reduction, as effluent concentrations increased up to 150 μg/L (77% ClO4− removal) in M1 and 45 μg/L (94% ClO4− removal) in M2. These results agree with previous studies which showed that SO42− reduction can inhibit ClO4− reduction (Ontiveros-Valencia et al. 2013, 2014a).
In stage 2, and once ClO3− was added to M2 (day 24), ClO3− was immediately reduced and reached 98% reduction (effluent of 120 µg/L) within 2 days (day 26). For the rest of stage 2, effluent ClO3− concentrations were around 90 ± 49 μg/L, except for day 30 and days 48–49, where leaks and biofilm sloughing caused some variations. Despite the low effluent ClO3− concentration during stage 2, which would exert a low selective pressure, ClO4− reduction improved and reached > 96% reduction, with an average effluent concentration of 23 ± 17 μg/L, and SO42− reduction diminished to < 5% and averaged 80 ± 3 mg/L in the effluent. In contrast, the average ClO4− reduction decreased to 92% in M1, with an effluent of 53 ± 37 μg/L—the double compared to M2—and the average effluent SO42− concentration was 73 ± 3 mg/L (~ 10% of reduction). This suggests that ClO3− improved ClO4− reduction.
Once ClO3− addition was discontinued, SO42− reduction quickly resumed and reached levels higher than before ClO3− addition, and also higher than M1 (Fig. 4). In stage 3, we also observed a decrease in ClO4− removals to below 95% for M2. ANOVA tests indicated that reduction of ClO4− and SO42− during stage 2 in M2 was significantly different than in M1 (F-ratio = 17.21 and p = 0.0005 for ClO4−, and F-ratio = 78.82 and p = 0.0005 for SO42−).
Assessment of the microbial community with 16S rRNA sequencing
The microbial community in reactor M1 was dominated by phylotypes related to Rhodocyclales (35%) and Bulkhoderiales (28%) at the end of the first stage (t1) (Fig. 5, Fig. S2). Then, the community was enriched with phylotypes related to SRB at the second and third stages, represented by the order Desulfovibrionales (Muyzer and Stams 2008). This order was 3% of the microbial community initially, but it was 52% at the end of stage 3. As the presence of SRB increased, the phylotypes Rhodocyclales and Bulkhoderiales decreased from 63 to 22%.
Microbial community structure at order level in the control (M1) and experimental (M2) reactors. Biological samples were taken at three time points: (i) day 21, before adding chlorate to the experimental reactor (t1); (ii) day 60, after 36 days of chlorate addition (t2); and (iii) day 78, after 15 days of stopping chlorate addition (t3). The presented results are the average of two samples at each time in each reactor. The sum of the less abundant phylotypes (< 1%) is classified as “Other”
In M2, the community in stage 1 was also dominated by the orders Rhodocyclales and Burkholderiales (71% in total) (Fig. 5, Fig. S2). During ClO3− addition in stage 2 (t2), phylotypes related to SRB Desulfovibrionales increased from 7 to 41%, and Rhodocyclales and Burkholderiales decreased to 32% in total, even more than in M1. Interestingly, without ClO3− (t3), Desulfovibrionales decreased to 17% and Rhodocyclales and Burkholderiales increased to 47%.
Both reactors presented sulfur-oxidizing bacteria (SOB) Thiobacterales and Campylobacterales at t2 and t3 (Fig. 5). SOB can oxidize H2S to S0 or SO42− coupled with O2 or NO3− or ClO4− (Bomberg et al. 2016; Kersters et al. 2006; Shao et al. 2010). Although initially SOB only represented 3–4% of the total relative abundance, once SO42− was reduced and produced H2S, these bacteria increased in abundance in the biofilm.
Discussion
Chlorate effect in the microbial communities
In both types of reactors (i.e., S1, S2, M1, and M2), the microbial community was dominated by Rhodocyclales and Burkholderiales. These types of bacteria are capable of denitrification and ClO4− reduction. They have also been detected in other studies reducing NO3− and ClO4− (Nerenberg et al. 2008; Ontiveros-Valencia et al. 2014a, b; Zhao et al. 2011).
The high increment of Dechloromonas (Rhodocyclales) species in S2 (19 to 42%) compared to S1 (17 to 21%) at stage 2 suggests it was the main organism responsible for ClO3− reduction. This would also explain the higher ClO3− reduction, and the inhibition of NO3− reduction, when H2 availability was higher (day 32) (Fig. 2). Moreover, Dechloromonas outcompeted Bulkholderiales during ClO3− addition (stage 2) in S2. This agrees with previous studies indicating that members of Bulkholderiales (i.e., Hydrogenophaga) in reactors with NO3− and ClO4− have been mainly responsible for NO3− reduction but not for ClO4− reduction (Liu et al. 2019; Willems and Gillis 2015; Zhao et al. 2011). This shows that ClO3− addition for 2 weeks can enrich for PRB, but they are not exclusively ClO4− reducers.
Similarly to reactors S1 and S2, the reduction of ClO4− and NO3− in reactors M1 and M2 was likely performed by Dechloromonas, which was present in both reactors, even with the low influent ClO4− concentration (Fig. 4). This was suggested by Nerenberg et al. (2008), who proposed that NO3− can serve as a primary electron acceptor substrate for PRB. Also, as ClO3− was immediately reduced, it indicates the microbial community already had the ability to reduce ClO3−. Since the community presumably contained PRB prior to ClO3− addition, it is likely that PRB were responsible for the initial reduction, rather than CRB. The increase of ClO3− reduction during the first 4 days of addition suggests a further enrichment of PRB.
As ClO3− decreased SO42− reduction in M2, we expected a lower abundance of SRB than in M1 at the end of the stage 2 (day 60). However, 16S rRNA results at t2 showed a larger presence of Desulfovibrionales in M2 than in M1 (41% compared to 25%) (Fig. 5). Also, once the influent ClO3− was removed (stage 3, Fig. 4), SO42− reduction was lower, but SRB abundance was higher (Fig. 5). A potential explanation for the higher abundance of SRB is that the microbial abundance of SRB might not be completely related to their activity, as the biological samples may contain DNA of bacteria that are inactive (dead) but still persist in the biofilm (Li et al. 2017). This can be particularly true for SRB in these reactors (M1 and M2), as they might proliferate closer to the membranes, where they have more access to H2 and are more protected from detachment (further explanation in the following section). It is also possible that H2S formed by SRB could serve as an electron donor in the outer biofilm and be re-oxidized to SO42−, obscuring the SO42− reduction. H2S profiles obtained with microsensors in a H2-MBfR with the same conditions as M1 showed that H2S was both formed and consumed within the biofilm (Fig. S3), supporting this explanation.
Chlorate addition as a mean to enrich PRB in MBfRs
As a strategy to enrich PRB, ClO3− addition to H2-MBfRs worked in both types of H2MBfR, but its effect in the microbial community was clearer in the single membrane reactors, where ClO3− loading was 10 times higher than in the multiple membrane reactors, allowing higher bulk ClO3− concentrations. Also, the addition of ClO3− clearly helped inhibit SO42− reduction, but it apparently did not eliminate SRB, given the fast recovery of SO42− reduction after discontinuing ClO3− supply in reactor M2. This indicates the strategy of adding ClO3− has to be permanent and that the ClO3− concentration in the bulk must be kept above a certain level, around 0.1 mg/L in our experiments, in order to maintain ClO4− reduction at higher levels (i.e., > 96%) and to prevent SO42− reduction.
In reactors M1 and M2, the membranes were potted together at both ends, forming a bundle, and membranes may have clumped together. This configuration allows excessively growth of biomass, which entails problems such as a decrease in the reduction activities in the external part of the biofilm, due to a limitation of H2 (Martin et al. 2015). In the inner part of the biofilm, limitations of NO3−, ClO3−, and ClO4− may occur, but not SO42−. As SO42− is generally present at much higher concentrations than the other electron acceptors, it more readily diffuses through the biofilm and reaches the membrane surface, where maximum H2 concentrations are found. This provides perfect conditions for SRB to grow, and also gives them more protection from detachment (Martin and Nerenberg 2012; Tang et al. 2013). Profiles of H2S in a bundle showed that H2S was accumulated in the center of the bundle, where SRB are more likely to be located (Fig. S3).
In the single membrane reactors, the biofilms were thinner and reducing the likelihood of SRB proliferation. These reactors can represent a “real” H2-MBfR with multiple “single membranes” (membranes separated from each other) at steady state, as we are using influent concentrations closer to the effluent concentrations in other H2-MBfR studies—except for ClO3−—(Martin and Nerenberg 2012; Nerenberg et al. 2008; Ontiveros-Valencia et al. 2014b). In S2 the ClO3− concentration in the bulk remained close to 2 mg/L and the shifts in the microbial community were relevant as compared to S1. However, despite the small membrane area, an increment in H2 availability allowed ClO3− to be reduced to lower levels and competed with NO3−. Furthermore, single membrane reactors showed that SO42− reduction was not detected until the increment in H2 pressure (Fig. 2, day 33), but ClO3− was able to inhibit it, despite that the acceptor loading in mEq/m2-day (Table 2) for SO42− was almost four times higher than ClO3−. H2 pressure is a key operational parameter for the performance of H2-MBfRs (Ontiveros-Valencia et al. 2014a).
One concern with the ClO3− addition strategy is that it could enrich for CRB, which reduce ClO3−, but not ClO4−, using the specialized chlorate reductase enzyme. This enzyme differs from the perchlorate reductases (Steinberg et al. 2005). Most known CRB belong to the γ-proteobacteria (Youngblut et al. 2016). However, as explained before, our results showed that ClO3− addition enriched for PRB, and no members of γ-proteobacteria were present in the microbial community (Figs. 3 and 4). Also, it should be noted that the effluent ClO3− concentrations were 0.09 mg/L in reactors M1 and M2, which is well below the ClO3− standard of 0.7 mg/L, and did not represent a contamination problem.
To our best knowledge, this is the first time ClO3− addition was tested to enhance ClO4− reduction. ClO3− addition increased the abundance of PRB in H2-MBfRs, improved perchlorate reduction, and suppressed SO42− reduction. ClO3− effect did not last long after it was removed from the reactors, indicating that intermittent ClO3− addition would be needed.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
ANSES: French Agency for Food, Environmental and Health Occupational & Safety (2012) ANSES opinion concerning epidemiological studies on associations between exposure to perchlorate in drinking water and thyroid function in specific populations., Vol. 2017. Retrieved 16 May, 2017, from https://www.anses.fr/en/system/files/EAUX2012sa0119EN.pdf
Bardiya N, Bae J-H (2011) Dissimilatory perchlorate reduction: a review. Microbiol Res 166(4):237–254
Bomberg M, Lamminmäki T, Itävaara M (2016) Microbial communities and their predicted metabolic characteristics in deep fracture groundwaters of the crystalline bedrock at Olkiluoto, Finland. Biogeosciences 13(21):6031–6047
CA DEPT (California Department of Public Health) (2007) State adoption of a perchlorate standard. Retrieved 16 May, 2017 from: https://www.waterboards.ca.gov/drinking_water/certlic/drinkingwater/documents/perchlorate/AdoptionMemotoWaterSystems-10-2007.pdf
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci 108(Supplement 1):4516–4522
Carlson HK, Kuehl JV, Hazra AB, Justice NB, Stoeva MK, Sczesnak A, Mullan MR, Iavarone AT, Engelbrektson A, Price MN, Deutschbauer AM, Arkin AP, Coates JD (2015) Mechanisms of direct inhibition of the respiratory sulfate-reduction pathway by (per)chlorate and nitrate. ISME J 9(6):1295–1305
Choe JK, Mehnert MH, Guest JS, Strathmann TJ, Werth CJ (2013) Comparative assessment of the environmental sustainability of existing and emerging perchlorate treatment technologies for drinking water. Environ Sci Technol 47(9):4644–4652
Coates JD, Achenbach LA (2004) Microbial perchlorate reduction: rocket-fuelled metabolism. Nat Rev Microbiol 2(7):569–580
Coates JD, Chakraborty R, Lack JG, O’Connor SM, Cole KA, Bender KS, Achenbach LA (2001) Anaerobic benzene oxidation coupled to nitrate reduction in pure culture by two strains of Dechloromonas. Nature 411:1039
Coates JD, Michaelidou U, Bruce RA, O’Connor SM, Crespi JN, Achenbach LA (1999) Ubiquity and diversity of dissimilatory (per)chlorate-reducing bacteria. Appl Environ Microbiol 65(12):5234–5241. https://doi.org/10.1128/AEM.65.12.5234-5241
Dasgupta PK, Martinelango PK, Jackson WA, Anderson TA, Tian K, Tock RW, Rajagopalan S (2005) The origin of naturally occurring perchlorate: the role of atmospheric processes. Environ Sci Technol 39(6):1569–1575. https://doi.org/10.1021/es048612x
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a Chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72(7):5069–5072
EFSA Panel on Contaminants in the Food Chain (EFSA CONTAM Panel) (2014) Scientific opinion on the risks to public health related to the presence of perchlorate in food, in particular fruits and vegetables. EFSA J 12(10):3869
Engelbrektson A, Hubbard CG, Tom LM, Boussina A, Jin YT, Wong H, Piceno YM, Carlson HK, Conrad ME, Anderson G, Coates JD (2014) Inhibition of microbial sulfate reduction in a flow-through column system by (per)chlorate treatment. Front Microbiol 5:315
Engelbrektson AL, Cheng Y, Hubbard CG, Jin YT, Arora B, Tom LM, Hu P, Grauel A-L, Conrad ME, Andersen GL, Ajo-Franklin JB, Coates JD (2018) Attenuating sulfidogenesis in a soured continuous flow column system with perchlorate treatment. Front Microbiol 9:1575–1575
Kersters K, De Vos P, Gillis M, Swings J, Vandamme P, Stackebrandt E (2006) Introduction to the proteobacteria. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes: Volume 5: Proteobacteria: alpha and beta subclasses. Springer, New York, pp 3–37
Kraft B, Tegetmeyer HE, Meier D, Geelhoed JS, Strous M (2014) Rapid succession of uncultured marine bacterial and archaeal populations in a denitrifying continuous culture. Environ Microbiol 16(10):3275–3286
Li R, Tun HM, Jahan M, Zhang Z, Kumar A, Dilantha Fernando WG, Farenhorst A, Khafipour E (2017) Comparison of DNA-, PMA-, and RNA-based 16S rRNA Illumina sequencing for detection of live bacteria in water. Sci Rep 7(1):5752
Liu W, Lian J, Guo J, Zhang C, Guo Y, Niu Y, Duan L (2019) Perchlorate reduction by anaerobic granular sludge under different operation strategies: performance, extracellular polymeric substances and microbial community. Bioresour Technol Rep 8:100312. https://doi.org/10.1016/j.biteb.2019.100312
Ma H, Bonnie NA, Yu M, Che S, Wang Q (2016) Biological treatment of ammonium perchlorate-contaminated wastewater: a review. J Water Reuse Desalination 7(4):82–107
MA DEPT. (Massachussets Department of Environmental Protection) (2006) Update to “Perchlorate Toxicological Profile and Health Assessment”. Available: https://www.mass.gov/doc/310-cmr-2200-the-massachusetts-drinking-water-regulations/download
Martin KJ, Nerenberg R (2012) The membrane biofilm reactor (MBfR) for water and wastewater treatment: principles, applications, and recent developments. Bioresour Technol 122:83–94
Martin KJ, Picioreanu C, Nerenberg R (2015) Assessing microbial competition in a hydrogenbased membrane biofilm reactor (MBfR) using multidimensional modeling. Biotechnol Bioeng 112(9):1843–1853. https://doi.org/10.1002/bit.25607
Muyzer G, Stams AJM (2008) The ecology and biotechnology of sulphate-reducing bacteria. Nat Rev Microbiol 6:441
Nerenberg R, Kawagoshi Y, Rittmann BE (2006) Kinetics of a hydrogen-oxidizing, perchlorate-reducing bacterium. Water Res 40(17):3290–3296
Nerenberg R, Kawagoshi Y, Rittmann BE (2008) Microbial ecology of a perchlorate-reducing, hydrogen-based membrane biofilm reactor. Water Res 42(4–5):1151–1159
Ontiveros-Valencia A, Tang Y, Krajmalnik-Brown R, Rittmann BE (2013) Perchlorate reduction from a highly contaminated groundwater in the presence of sulfate-reducing bacteria in a hydrogen-fed biofilm. Biotechnol Bioeng 110(12):3139–3147
Ontiveros-Valencia A, Tang Y, Krajmalnik-Brown R, Rittmann BE (2014a) Managing the interactions between sulfate- and perchlorate-reducing bacteria when using hydrogen-fed biofilms to treat a groundwater with a high perchlorate concentration. Water Res 55:215–224
Ontiveros-Valencia A, Tang Y, Zhao H-P, Friese D, Overstreet R, Smith J, Evans P, Rittmann BE, Krajmalnik-Brown R (2014b) Pyrosequencing analysis yields comprehensive assessment of microbial communities in pilot-scale two-stage membrane biofilm reactors. Environ Sci Technol 48(13):7511–7518
Oren A (2014) The Family Xanthobacteraceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The Prokaryotes: Alphaproteobacteria and Betaproteobacteria. Springer, Berlin Heidelberg, pp 709–726
Roldán MD, Reyes F, Moreno-Vivián C, Castillo F (1994) Chlorate and nitrate reduction in the phototrophic bacteria Rhodobacter capsulatus and Rhodobacter sphaeroides. Curr Microbiol 29(4):241–245
Shao M-F, Zhang T, Fang H (2010) Sulfur-driven autotrophic denitrification: diversity, biochemistry, and engineering applications. Appl Microbiol Biotechnol 88:1027–1042
Sijimol MR, Jyothy S, Pradeepkumar AP, Chandran MSS, Ghouse SS, Mohan M (2015) Review on Fate, Toxicity, and Remediation of Perchlorate. Environ Forensics 16(2):125–134
Srinivasan R, Sorial GA (2009) Treatment of perchlorate in drinking water: a critical review. Sep Purif Technol 69(1):7–21
Steinberg LM, Trimble JJ, Logan BE (2005) Enzymes responsible for chlorate reduction by Pseudomonas sp. Are different from those used for perchlorate reduction by Azospira sp. FEMS Microbiol Lett 247(2):153–159. https://doi.org/10.1016/j.femsle.2005.05.003
Tang Y, Zhao H, Marcus AK, Krajmalnik-Brown R, Rittmann BE (2012) A Steady-State Biofilm Model for Simultaneous Reduction of Nitrate and Perchlorate, Part 2: Parameter Optimization and Results and Discussion. Environ Sci Technol 46(3):1608–1615
Tang Y, Ontiveros-Valencia A, Feng L, Zhou C, Krajmalnik-Brown R, Rittmann BE (2013) A biofilm model to understand the onset of sulfate reduction in denitrifying membrane biofilm reactors. Biotechnol Bioeng 110(3):763–772
US Environmental Protection Agency (US EPA) (2009) Revised Assessment Guidance for Pechlorate. Memorandum to Regional Administrators. US EPA Office of Solid Waste and Emergency Response, Washington, DC
Willems A (2014) The Family Comamonadaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The prokaryotes: Alphaproteobacteria and Betaproteobacteria. Springer, Berlin Heidelberg, pp 777–851
Willems A., Gillis, M. 2015. Hydrogenophaga. in: Bergey’s manual of systematics of archaea and bacteria, (Ed.) W. B. Whitman, F. Rainey, P. Kämpfer, M. Trujillo, J. Chun, P. DeVos, B. Hedlund and S. Dedysh.
World Health Organization (WHO) (2004) Sulfate in drinking-water. WHO Press
World Health Organization (WHO) (2011) Nitrate and nitrite in drinking-water. WHO Press
World Health Organization (WHO) (2016) Chlorine dioxide, chlorite and chlorate in drinking-water. WHO Press
Yang B, Wang Y, Qian P-Y (2016) Sensitivity and correlation of hypervariable regions in 16S rRNA genes in phylogenetic analysis. BMC Bioinformatics 17(1):135. https://doi.org/10.1186/s12859-016-0992-y
Ye L, You H, Yao J, Su H (2012) Water treatment technologies for perchlorate: a review. Desalination 298:1–12
Youngblut MD, Wang O, Barnum TP, Coates JD (2016) (Per)chlorate in biology on Earth and beyond. Annu Rev Microbiol 70:435–457
Zhang S, Pang S, Wang P, Wang C, Guo C, Addo FG, Li Y (2016) Responses of bacterial community structure and denitrifying bacteria in biofilm to submerged macrophytes and nitrate. Sci Rep 6:36178
Zhang M, Li X, Cheng X, Wang X, Song M, Wang X, Ma X (2021) Isotopic tracing of perchlorate sources in the environment. J Chem 2021:9978489. https://doi.org/10.1155/2021/9978489
Zhao HP, Van Ginkel S, Tang Y, Kang DW, Rittmann B, Krajmalnik-Brown R (2011) Interactions between perchlorate and nitrate reductions in the biofilm of a hydrogen-based membrane biofilm reactor. Environ Sci Technol 45(23):10155–10162
Acknowledgements
The authors gratefully acknowledge the Center for Environmental Science and Technology (CEST) at Notre Dame for their support with analytical equipment.
Funding
This research was supported by the University of Notre Dame-Pontificia Universidad Católica Collaborative Seed Fund, the CEDEUS (Centro de Desarrollo Urbano Sustentable) CONICYT/FONDAP/15110020, FONDECYT 1201134, and National Scholarship for PhD (CONICYT, Chile) 21161573.
Author information
Authors and Affiliations
Contributions
RN, ITV, and MV conceived and designed the research. MV and AOV conducted the experiments and analyzed the data. MV wrote the manuscript. RN, AOV, and ITV revised the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Vega, M., Ontiveros-Valencia, A., Vargas, I.T. et al. Chlorate addition enhances perchlorate reduction in denitrifying membrane-biofilm reactors. Appl Microbiol Biotechnol 106, 4341–4350 (2022). https://doi.org/10.1007/s00253-022-11976-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-11976-1