Abstract
Squalene, a naturally occurring linear triterpene formed via MVA or MEP biosynthetic pathway, is widely distributed in microorganisms, plants and animals. At present, squalene is used extensively in the food, cosmetic and medicine industries because of its antioxidant, antistatic and anti-carcinogenic properties. Increased consumer demand has led to the development of microbial bioprocesses for the commercial production of squalene, in addition to the traditional methods of isolating squalene from the liver oils of deep-sea sharks and plant seed oils. As knowledge of the biosynthetic enzymes and of regulatory mechanisms modulating squalene production increases, opportunities arise for the genetic engineering of squalene production in hosts. In this review, we present the various strategies used up to date to improve and/or engineer squalene production in microbes and analyze yields.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Squalene, a dehydro-triterpenic hydrocarbon, is a basic intermediate in the biosynthesis of sterols, hopanoids and triterpenes (Kohno et al. 1995). It was not until around a hundred years after it was first described (Tsujimoto 1916) to be isolated from the sea shark’s (Squalus spp) liver oil (Gershbein and Singh 1969) that it was formally named. The amount of squalene varies in plant seeds (Berger et al. 2003; Martirosyan et al. 2007; Maguire et al. 2009), oils (Singhal and Kulkarni 1990; De Leonardis et al. 1998; Huang et al. 2009; Xiao et al. 2016) and freshwater fish (Ackman et al. 1968; KopiCoVá and VaVreiNoVá 2007). In humans, it is the principal hydrocarbon in surface lipids, making up approximately 10 % of the total surface fat (Liu et al. 1976). Currently, squalene is used widely in the food, cosmetics and medicine industries due to its multiple functions. Up until now, the liver oil of deep sea sharks (Hernández-Pérez et al. 1997) and plant oils (He et al. 2002) have been the major commercial source of squalene. However, the continuous supply and future availability of these sources are uncertain due to the endangerment of the shark species (Lack and Sant 2009) and the unpredictable effect of region and seasonal variation on crop production (Salvador et al. 2003). Therefore, in recent years, many studies have tried to develop microbial strains for squalene production using fermentation processes (Ghimire et al. 2016). This review focuses on these current methods of microbial fermentation and explores the impact that our improving knowledge of squalene biosynthesis may have on future production.
Structure and functions of squalene
Squalene (2, 6, 10, 15, 19, 23-hexamethyltetracosa-2, 6, 10, 14, 18, 22-hexaene,), an unsaponifiable lipid, is a dehydro-triterpenic hydrocarbon (C30H50) with six double bonds (Fig. 1). Squalene acts as an natural antioxidant that protects cells from free radicals and reactive oxygen species (Kohno et al. 1995; Obulesu et al. 2015). It has been proven to have radioprotective (Storm et al. 1993) and cardioprotective (Aguilera et al. 2005) functions. Squalene also participates in reducing serum cholesterol levels (Miettinen and Vanhanen 1994; Chan et al. 1996) and improving cellular and non-specific immune functions (Ohkuma et al. 1983; Kelly 1999). Various studies have demonstrated that it is an effective inhibitor of chemically induced tumorigenesis in the colon, lung and skin (Budiyanto et al. 2000; Ronco and De Stéfani 2013; Fang 2014). Since 1997, over 22 million squalene-containing flu vaccines have been administered for their potential capability of increasing immune responses (Kaya et al. 2011).
Recently, it has been found that squalene can modulate fatty acid metabolism (Ravi Kumar et al. 2016) and improve DSS-induced acute colitis by down-regulating signaling pathways (Sánchez-Fidalgo et al. 2015).
Squalene biosynthesis in natural microbial producers
Due to their fast and massive growth, microorganisms are considered a promising alternative to other sources for squalene production. Squalene isolation has been reported in Saccharomyces cerevisiae (Kamimura et al. 1994; Socaciu et al. 1995; Mantzouridou and Tsimidou 2010; Naziri et al. 2011; Garaiová et al. 2014; Hull et al. 2014), Torulaspora delbrueckii (Bhattacharjee et al. 2001), Synechocystis (Englund et al. 2014), Pseudomonas (Uragami and Koga 1986), Candida (Tsujiwaki et al. 1995; Lee et al. 2014), the microalgae Traustochytrium (Dessi et al. 2002), Chlamydomonas reinhardtii (Kajikawa et al. 2015) and Schizochytrium mangrovei (Jiang et al. 2004; Yue and Jiang 2009; Hoang et al. 2016), Rhodopseudomonas palustris (Welander et al. 2009; Xu et al. 2016), and Botryococcus braunii (Banerjee et al. 2002). In these microorganisms, each molecule of squalene is formed by fusing two molecules of farnesyl diphosphate (FPP), an isoprenoid compound (Spanova and Daum 2011). Herein, the squalene biosynthetic pathway is re-cast into two modules: the isoprenoid pathway and the fusion pathway.
Isoprenoid pathway
All isoprenoids are synthesized from common isopentenyl units including isopentenyl diphosphate (IPP) and its isomer dimethylallyl diphosphate (DMAPP) (Lange et al. 2000) via either the classical mevalonate (MVA) pathway or 2-C-methyl-d-erythritol 4-phosphate (MEP) pathway using non-homologous enzymes (Lange et al. 2000). The MEP pathway most commonly occurs in prokaryotes such as E.coli and the plastids of photosynthetic organisms (Rohmer et al. 1993; Orihara et al. 1998; Kuzuyama 2002; Dellas et al. 2013), although some parasitic prokaryotic microorganisms such as Rickettsia and Mycoplasma appear to simply obtain their isoprenoids from the host cells (Fraser et al. 1995; Andersson et al. 1998).
Isoprenoid biosynthesis begins with the formation of 1-deoxy-d-xylulose- 5-phosphate (DXP) catalyzed by DXP synthase (DXS), a rate-limiting enzyme of the MEP pathway (Kim et al. 2006; Lee et al. 2007). DXP then undergoes rearrangement and forms reduced MEP by DXP reductoismerase DXR or DRL encoded by the dxr or drl genes respectively (Sangari et al. 2010). A series of other enzymes are used in the following sequential reactions to convert MEP into the building blocks of IPP and DMAPP which are further isomerized via the enzyme isopentanyl diphosphate isomerase (IDI). Farnesyl diphosphate synthase (FPS) catalyzes the sequential 1′-4 coupling of IPP with DMAPP and GPP, resulting in the formation of FPP (Fig. 2).
Metabolic pathway involved in the synthesis of squalene. AACT acetoacetyl- CoA thiolase, DMAPP dimethylallyl diphosphate, DXS 1-deoxy-xylulose-5-phosphate synthase, DXP 1-deoxy-xylulose -5-phosphate, DXR DXP reductoisomerase, DRL DXR-like protein, FPS farnesyl diphosphate (FPP) synthase, GA3P D-glyceraldehyde-3-phosphate, GPP geranyl diphosphate, HMGS 3-hydroxy-3- methylglutaryl-CoA (HMG-CoA) synthase, HMGR HMG-CoA reductase, HMGR HSQS (R)-12-hydroxysqualene (HSQ) synthase, IDI isopentenyl diphosphate isomerase, IPP isopentenyl diphosphate, IspH IPP synthase, PSPPS presqualene diphosphate (PSPP) synthase, SQS squalene synthase
Eukaryotic microorganisms such as fungi and yeasts lack the MEP pathway and rely almost entirely on the MVA pathway. The biosynthesis of IPP using the MVA pathway begins with the conversions of three molecules of acetyl-CoQ to MVA sequentially through acetoacetyl-CoA and 3-hydroxy-3-methylglutaryl-CoA (HMG-CoA), catalyzed by acetoacetyl-CoA synthsae (AAS) and HMG-CoA synthase (HMGS) (Maury et al. 2005). MVA is then converted to MVA 5-diphosphate by two phosphorylations that are mediated by MVA kinase and phospho-MVA kinase, respectively. MVA 5-diphosphate undergoes dehydration-decarboxylation in the presence of adenosine triphosphate (ATP) to form IPP. An IPP isomerase (IDI) catalyzes the isomerization of IPP to DMAPP. Finally, GPP, formed by the condensation of IPP with DMAPP, is condensed with another molecule of IPP to create FPP (Fig. 2).
Fusion pathway
The studies in animals (Shechter et al. 1992; Thompson et al. 1998), plants (Nakashima et al. 1995), fungi (Jennings et al. 1991) and a few bacteria (Lee and Poulter 2008; Ohtake et al. 2014) show that squalene synthase (SQS), catalyzes the fusion of two FPP molecules into one molecule of squalene in two consecutive steps—coupling of two molecules of FPP into presqualene diphosphate (PSPP) and NADOH-dependent reductive rearrangement of PSPP to squalene (Epstein and Rilling 1970). Recently, a three-step pathway using a set of three enzymes including PSPP synthase (PSPPS), HSQ synthase (HSQS) and squalene synthase (SQS) has been discovered in bacteria (Pan et al. 2015) (Fig. 2).
Strategies for increasing squalene production in natural producers
Microorganisms have a great potential to become the sources for squalene production. Current investigations are focused on increasing the content of squalene in microorganisms. Multiple strategies, including seeking alternative sources, optimizing fermentation and using metabolic engineering as a tool, have been applied to improve the squalene production.
Exploration of squalene producing sources
Natural producers with high content of squalene are of importance for squalene production. Therefore, much effort has been directed towards exploring these new sources. Marine microorganisms such as Pseudozyma sp. JCC207 (Chang et al. 2008), Schizochytrium sp. CCTCCM209059 (Ren et al. 2010), Schizochytrium mangrovei (Jiang et al. 2004), Rubritalea sabuli sp. nov. (Yoon et al. 2008) and Rubritalea squalenifaciens sp. nov (Kasai et al. 2007) have been characterized as suitable squalene producers. Thraustochytrids Aurantiochytrium strains have a high potential for commercial production (Aasen et al. 2016). Kaya et al. successfully isolated the 18W-13a strain of Aurantiochytrium sp. from a mangrove area. It was found that the strain produced 198 mg/g dry cell weight (DCW) and 1.29 ± 0.13 g/L of squalene after 4 days of culture (Kaya et al. 2011).
Recently, Nakazawa et al. reported that 176 strains of thraustochytrids isolated from various regions in Asia were screened using thin layer chromatography (TLC) for squalene contents. A total of 38 strains were rated as “+” (high), 29 as “±” (medium), and 109 as “−” (low). The strain with the highest squalene content (317.74 mg/g DCW) was Aurantiochytrium sp. Yonez 5-1 (Nakazawa et al. 2014).
Optimization of fermentation conditions
Mantzouridou et al. examined the growth dynamics of two wild-type strains of S. cerevisiae EGY48 and BY4741 for sterols production through investigation of the impact of oxygen supply, inoculums, and fermentation period on yield. The highest squalene yield were 3.13 mg/L for EGY48 and 2.97 mg/L for BY4741 respectively (Mantzouridou et al. 2009). The production peak was achieved by treating the strain EGY48 with terbinafine (0.05–0.55 mM) and methyl jasmonate (0–1.0 mM). Maximum squalene content (10.02 mg/g DCW) and yield (20.70 mg/L) were reached at 0.442 mM terbinafine plus 0.044 mM methyl jasmonate after 28 h of culture (Naziri et al. 2011). Bhattacharjee et al. used both S. cerevisiae and Torulaspora delbrueckii for fermentative squalene production under anaerobic conditions. It was demonstrated that the production of squalene from Torulaspora delbrueckii (237.25 μg/g DCW) was much higher than that from S. cerevisiae (41.16 μg/g DCW), suggesting Torulaspora delbrueckii could be a potential suitable alternative source (Bhattacharjee et al. 2001).
Fan et al. increased the squalene content and yield in the thraustochytrid, Aurantiochytrium mangrovei FB3, through medium optimization and terbinafine treatment to inhibit squalene monooxygenase in the sterol biosynthetic pathway. The highest biomass concentration of 21.2 g/L was found in media with a glucose concentration at 60 g/L. Although addition of terbinafine led to a slight inhibition of cell growth, an increase in squalene content was observed. Compared to the control, the squalene content increased 36 and 40 % in the presence of 10 and 100 mg/L of terbinafine respectively (Fan et al. 2010). For squalene production by microalga Aurantiochytrium sp.BR-MP4-A1 in heterotrophic cultures, Li et al. optimized the nitrogen sources. Monosodium glutamate, yeast extract and tryptone were founded to be able to enhance cell growth and squalene production during screenings. By using the optimal concentrations of the three nitrogen sources, the squalene content and squalene yield reached 0.72 mg/g DCW and 5.90 mg/L, respectively (Li et al. 2009). Nakazawa et al. optimized the culture conditions for Aurantiochytrium sp. 18W-13a, including media ingredients and temperature. With 25–50 % seawater, 2–6 % glucose and 25 °C, the highest squalene content and yield reached 171 mg/g DCW and 900 mg/L, indicating that 18W-13a strain is a potential source for commercial squalene (Nakazawa et al. 2012).
Recently, Englund et al. (2014) used the unicellular cyanobacterium Synechocystis sp. PCC 6803 as a host for the production of squalene. Investigation of the impacts of light intensity and growth stage on squalene accumulation in the mutant strain found that normal light conditions (50 μmol photons m−2 s−2) were better for squalene accumulation, compared to low and high light conditions (Englund et al. 2014). Drozdíková et al. (2015) tested the squalene production in Kluyveromyces lactis. By partial inhibition of squalene epoxidase with terbinafine and additional nutrients such as commercially available glucose diary wastes containing lactose, the level of produced squalene was high. In addition, this study shows a promising future for squalene production in K. lactis (Drozdíková et al. 2015).
Metabolic engineering
With better understanding of biosynthetic enzymes and regulatory mechanisms during squalene production, metabolic engineering in microbes becomes practical. Kamimura et al. (1994) was the first group that successfully increased squalene production in S. cerevisiae, by disrupting a gene involved in the conversion of squalene to ergosterol. The yield of squalene reached 5 mg/g DCW when culturing the mutant strains in aerobic condition. When grass juice was added as the feedstock together with 0.025 μg/mL of doxycycline to treat the recombinant S. cerevisiae YUG37-EGR1, the level of squalene reached 18.0 ± 4.18 mg/L (Kamimura et al. 1994). The ERG1 gene encodes an enzyme (squalene epoxidase) that catalyzes the rate-limiting step in the squalene biosynthetic pathway. It has been proven that targeted genetic manipulation of ERG1 gene is a promising tool for increasing squalene production in yeast. In the study from Garaiová et al. (2014), the highest squalene level (over 1000 μg/109 cells) was induced by specific point mutations in the ERG1 gene that were capable of reducing the activity of squalene epoxidase and inducing terbinafine hypersensitivity, without disturbing their cell growth. Treatment with squalene epoxidase inhibitor terbinafine revealed a limit of 700 μg/109 cells in squalene accumulation, which was attributed to pronounced growth defects (Garaiová et al. 2014). Mantzouridou and Tsimidou (2010) studied the capacity of squalene accumulation in S. cerevisiae via constructing the strains of AM63 and AM64. AM63 had an extra copy of the HMG2 gene with a K6R stabilizing mutation in Hmg2p under the control of the inducible galactose promoter, while AM64 was derived from AM63 with an additional deletion of the ERG6 gene. In AM63, the squalene content was 20-fold higher than that in the wild-type EGY48 parental strain. However, in AM64, the combined Hmg2p stabilization and ERG6 deletion did not further enhance squalene accumulation. It was believed that the lack of ergosterol feedback inhibition led to an elevated transfer of surplus squalene into C27 sterols (Mantzouridou and Tsimidou 2010).
Recently, Englund et al. (2014) engineered Synechocystis sp. PCC 6803 by deleting the slr2089 gene encoding squalene hopene cyclase (Shc), an enzyme converting squalene into hopene. The accumulated squalene in the recombinant strain was over 70-times higher than that in wild-type cells, reaching 3.16 mg/g DCW (Englund et al. 2014). Similarly, our study (2016) on Rhodopseudomonas palustris showed that the deletion of the shc gene resulted in a squalene production of 3.8 mg/g DCW, which was 27-times higher than that in the wild-type strain. Moreover, it was found that the fusion expression of the two genes, crtE and hpnD, was better than the co-expression method for squalene production. Finally, the titer of squalene reached 15.8 mg/g DCW by fusing the two genes and co-expressing the dxs gene in the shc-deficient strain, corresponding to a 112-fold increase compared to that in the wild-type strain (Xu et al. 2016). Kajikawa et al. (2015) investigated the accumulation of squalene in microalgal Chlamydomonas reinhardtii on the basis of the characterization of squalene synthease (CrSQS) and squalene epoxidase (CrSQE). It was found that the overexpression of CrSQS increased the rate of conversion of C14-labeled farnesyl pyrophosphate into squalene, but not squalene content. However, in the CrSQE-deficient strain, the titer of squalene was increased significantly (0.9-1.1 μg/mg DCW), indicating that partially knocking down CrSQE is a promising approach for increasing squalene production in C. reinhardtii cells (Kajikawa et al. 2015). The squalene production levels among different microorganisms are presented in Table 1.
Recombinant engineering in host Escherichia Coli
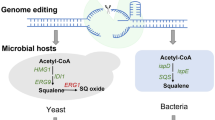
Many efforts on recombinant techniques are focused on E. coli because this organism is suitable for genetic modification and large-scale fermentation. For instance, Ghimire et al. (2009) used E. coli for heterologous production of squalene up to 4.1 mg/L via expression of the Streptomyces peucetius-originated genes (hopA, hopB and hopD). Moreover, E. coli endogenous dxs and idi genes encoding 1-deoxy-d-xylulose 5-phosphate synthase and isopentenyl diphosphate isomerase were co-expressed, leading to the production up to 11.8 mg/L of squalene (Ghimire et al. 2009). Recently, E. coli was engineered successfully by introducing a three-step pathway for squalene synthesis using a set of three enzymes viz. PSPPS, HSQS and SQS (Pan et al. 2015). To establish an efficient E. coli-based system for squalene production, Katabami et al. (2015) tested two different squalene synthases and their mutants in combination with precursor pathways. By co-expressing a chimeric mevalonate pathway with human or Thermosynechococcus squalene synthase, E. coli was able to produce squalene up to 230 mg/L or 55 mg/g DCW in flask culture (Katabami et al. 2015). These studies suggest that E. coli is a promising host for squalene production. Metabolic engineering approaches to enhance squalene production are represented in Fig. 3.
Conclusion and future prospects
The biosynthesis pathway and biotechnological production of squalene have been of interest to many researchers because of its benefit for human health, as well as its applications in the pharmaceutical and cosmetics industries. So far, the isolation from natural squalene producers with optimized media has shown to be the most successful approach to increase squalene yield. Strategies need to be developed to explore alternative squalene producers with high squalene levels.
Using microorganisms such as S. cerevisiae and E. coli as hosts enables specific genes to be targetted and drives squalene production through specific biochemical pathways. Despite the promising results currently obtained, the yields are not sufficient for commercial production. Increasing the production of precursors by introducing foreign pathways (Zahiri et al. 2006) or blocking branch pathways (Xu et al. 2014) and enhancing the yield of target-soluble proteins via stress-minimization techniques (Wyre and Overton 2014) may be potential approaches to increase metabolic engineering efficiency for squalene production. Important questions that are still outstanding are whether the yields are limited by the carbon flux through the pathways or if there are some additional, yet unidentified, physiological factors that may limit squalene accumulation in the engineered strains. Further analysis of strains naturally producing high levels of squalene might be of help.
References
Aasen IM, Ertesvåg H, Heggeset TMB, Liu B, Brautaset T, Vadstein O, Ellingsen TE (2016) Thraustochytrids as production organisms for docosahexaenoic acid (DHA), squalene, and carotenoids. Appl Microbiol Biotechnol 100:4309–4321
Ackman R, Addison R, Eaton C (1968) Unusual occurrence of squalene in a fish, the Eulachon Thaleichthys pacificus. Nature 220:1033–1034
Aguilera Y, Dorado ME, Prada FA, Martínez JJ, Quesada A, Ruiz-Gutiérrez V (2005) The protective role of squalene in alcohol damage in the chick embryo retina. Exp Eye Res 80:535–543
Andersson SG, Zomorodipour A, Andersson JO, Sicheritz-Pontén T, Alsmark UCM, Podowski RM, Näslund AK, Eriksson A-S, Winkler HH, Kurland CG (1998) The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature 396:133–140
Banerjee A, Sharma R, Chisti Y, Banerjee U (2002) Botryococcus braunii: a renewable source of hydrocarbons and other chemicals. Crit Rev Biotechnol 22:245–279
Berger A, Gremaud G, Baumgartner M, Rein D, Monnard I, Kratky E, Geiger W, Burri J, Dionisi F, Allan M (2003) Cholesterol-lowering properties of amaranth grain and oil in hamsters. Int J Vitam Nutr Res 73:39–47
Bhattacharjee P, Shukla V, Singhal R, Kulkarni P (2001) Studies on fermentative production of squalene. World J Microbiol Biotechnol 17:811–816
Budiyanto A, Ahmed NU, Wu A, Bito T, Nikaido O, Osawa T, Ueda M, Ichihashi M (2000) Protective effect of topically applied olive oil against photocarcinogenesis following UVB exposure of mice. Carcinogenesis 21:2085–2090
Chan P, Tomlinson B, Lee CB, Lee YS (1996) Effectiveness and safety of low-dose pravastatin and squalene, alone and in combination, in elderly patients with hypercholesterolemia. J Clin Pharmacol 36:422–427
Chang MH, Kim HJ, Jahng KY, Hong SC (2008) The isolation and characterization of Pseudozyma sp. JCC 207, a novel producer of squalene. Appl Microbiol Biotechnol 78:963–972
Chen G, Fan K-W, Lu F-P, Li Q, Aki T, Chen F, Jiang Y (2010) Optimization of nitrogen source for enhanced production of squalene from thraustochytrid Aurantiochytrium sp. New Biotechnol 27:382–389
De Leonardis A, Macciola V, De Felice M (1998) Rapid determination of squalene in virgin olive oils using gas–liquid chromatography. Ital J Food Sci 10:75–80
Dellas N, Thomas ST, Manning G, Noel JP (2013) Discovery of a metabolic alternative to the classical mevalonate pathway. Elife 2:e00672
Dessi MA, Deiana M, Day BW, Rosa A, Banni S, Corongiu FP (2002) Oxidative stability of polyunsaturated fatty acids: effect of squalene. Eur J Lipid Sci Technol 104:506–512
Drozdíková E, Garaiová M, Csáky Z, Obernauerová M, Hapala I (2015) Production of squalene by lactose-fermenting yeast Kluyveromyces lactis with reduced squalene epoxidase activity. Lett Appl Microbiol 61:77–84
Englund E, Pattanaik B, Ubhayasekera SJK, Stensjö K, Bergquist J, Lindberg P (2014) Production of squalene in Synechocystis sp. PCC 6803. PLoS One 9:e90270
Epstein W, Rilling H (1970) Studies on the mechanism of squalene biosynthesis the structure of presqualene pyrophosphate. J Biol Chem 245:4597–4605
Fan KW, Aki T, Chen F, Jiang Y (2010) Enhanced production of squalene in the thraustochytrid Aurantiochytrium mangrovei by medium optimization and treatment with terbinafine. World J Microbiol Biotechnol 26:1303–1309
Fang H (2014) Frontier and future development of information technology in medicine and education. Lect Notes Electr Eng 269:1699–1705
Fraser CM, Gocayne JD, White O, Adams MD (1995) The minimal gene complement of Mycoplasma genitalium. Science 270:197
Garaiová M, Zambojová V, Šimová Z, Griač P, Hapala I (2014) Squalene epoxidase as a target for manipulation of squalene levels in the yeast Saccharomyces cerevisiae. FEMS Yeast Res 14:310–323
Gershbein LL, Singh EJ (1969) Hydrocarbons of dogfish and cod livers and herring oil. J Am Oil Chem Soc 46:554–557
Ghimire GP, Lee HC, Sohng JK (2009) Improved squalene production via modulation of the methylerythritol 4-phosphate pathway and heterologous expression of genes from Streptomyces peucetius ATCC 27952 in Escherichia coli. Appl Environ Microbiol 75:7291–7293
Ghimire GP, Thuan NH, Koirala N, Sohng JK (2016) Advances in biochemistry and microbial production of squalene and its derivatives. J Microbiol Biotechnol 26:441–451
He H-P, Cai Y, Sun M, Corke H (2002) Extraction and purification of squalene from amaranthus grain. J Agric Food Chem 50:368–372
Hernández-Pérez M, Gallego RMR, Alayón PJP, Hernández AB (1997) Squalene content in livers from deep-sea sharks caught in Canary Island waters. Mar Freshw Res 48:573–576
Hoang TMH, Nguyen CH, Le TT, Hoang THQ, Ngo THT, Hoang TLA, Dang DH (2016) Squalene isolated from Schizochytrium mangrovei is a peroxisome proliferator-activated receptor-α agonist that regulates lipid metabolism in HepG2 cells. Biotechnol Lett 38:1065–1071
Huang Z-R, Lin Y-K, Fang J-Y (2009) Biological and pharmacological activities of squalene and related compounds: potential uses in cosmetic dermatology. Molecules 14:540–554
Hull CM, Loveridge EJ, Rolley NJ, Donnison IS, Kelly SL, Kelly DE (2014) Co-production of ethanol and squalene using a Saccharomyces cerevisiae ERG1 (squalene epoxidase) mutant and agro-industrial feedstock. Biotechnol Biofuels 7:1
Jennings SM, Tsay YH, Fisch TM, Robinson GW (1991) Molecular cloning and characterization of the yeast gene for squalene synthetase. Proc Natl Acad Sci 88:6038–6042
Jiang Y, Fan K-W, Tsz-Yeung Wong R, Chen F (2004) Fatty acid composition and squalene content of the marine microalga Schizochytrium mangrovei. J Agric Food Chem 52:1196–1200
Kajikawa M, Kinohira S, Ando A, Shimoyama M, Kato M, Fukuzawa H (2015) Accumulation of squalene in a microalga Chlamydomonas reinhardtii by genetic modification of squalene synthase and squalene epoxidase genes. PLoS One 10:e0120446
Kamimura N, Hidaka M, Masaki H, Uozumi T (1994) Construction of squalene-accumulating Saccharomyces cerevisiae mutants by gene disruption through homologous recombination. Appl Microbiol Biotechnol 42:353–357
Kasai H, Katsuta A, Sekiguchi H, Matsuda S, Adachi K, Shindo K, Yoon J, Yokota A, Shizuri Y (2007) Rubritalea squalenifaciens sp. nov., a squalene-producing marine bacterium belonging to subdivision 1 of the phylum ‘Verrucomicrobia’. Int J Syst Evol Microbiol 57:1630–1634
Katabami A, Li L, Iwasaki M, Furubayashi M, Saito K, Umeno D (2015) Production of squalene by squalene synthases and their truncated mutants in Escherichia coli. J Biosci Bioeng 119:165–171
Kaya K, Nakazawa A, Matsuura H, Honda D, Inouye I, Watanabe MM (2011) Thraustochytrid Aurantiochytrium sp. 18W-13a accummulates high amounts of squalene. Biosci Biotechnol Biochem 75:2246–2248
Kelly GS (1999) Squalene and its potential clinical uses. Altern Med Rev J Clin Ther 4:29–36
Kim S-J, Kim M-D, Choi J-H, Kim S-Y, Ryu Y-W, Seo J-H (2006) Amplification of 1-deoxy-d-xyluose 5-phosphate (DXP) synthase level increases coenzyme Q (10) production in recombinant Escherichia coli. Appl Microbiol Biotechnol 72:982–985
Kohno Y, Egawa Y, Itoh S, S-i Nagaoka, Takahashi M, Mukai K (1995) Kinetic study of quenching reaction of singlet oxygen and scavenging reaction of free radical by squalene in n-butanol. Biochim Biophys Acta (BBA) Lipids Lipid Metab 1256:52–56
KopiCoVá Z, VaVreiNoVá S (2007) Occurrence of squalene and cholesterol in various species of Czech freshwater fish. Czech J Food Sci 25:195
Kuzuyama T (2002) Mevalonate and nonmevalonate pathways for the biosynthesis of isoprene units. Biosci Biotechnol Biochem 66:1619–1627
Lack M, Sant G (2009) Trends in global shark catch and recent developments in management. In: TRAFFIC International, vol 33
Lange BM, Rujan T, Martin W, Croteau R (2000) Isoprenoid biosynthesis: the evolution of two ancient and distinct pathways across genomes. Proc Natl Acad Sci 97:13172–13177
Lee S, Poulter CD (2008) Cloning, solubilization, and characterization of squalene synthase from Thermosynechococcus elongatus BP-1. J Bacteriol 190:3808–3816
Lee J-K, Oh D-K, Kim S-Y (2007) Cloning and characterization of the dxs gene, encoding 1-deoxy-d-xylulose 5-phosphate synthase from Agrobacterium tumefaciens, and its overexpression in Agrobacterium tumefaciens. J Biotechnol 128:555–566
Lee PY, Yong VC, Rosli R, Gam LH, Chong PP (2014) Cloning, expression and purification of squalene synthase from Candida tropicalis in Pichia pastoris. Protein Expr Purif 94:15–21
Li Q, Chen G-Q, Fan K-W, Lu F-P, Aki T, Jiang Y (2009) Screening and characterization of squalene-producing thraustochytrids from Hong Kong mangroves. J Agric Food Chem 57:4267–4272
Liu G, Ahrens E, Schreibman PH, Crouse JR (1976) Measurement of squalene in human tissues and plasma: validation and application. J Lipid Res 17:38–45
Maguire L, O’sullivan S, Galvin K, O’connor T, O’brien N (2009) Fatty acid profile, tocopherol, squalene and phytosterol content of walnuts, almonds, peanuts, hazelnuts and the macadamia nut. Int J Food Sci Nutr 55:171–178
Mantzouridou F, Tsimidou MZ (2010) Observations on squalene accumulation in Saccharomyces cerevisiae due to the manipulation of HMG2 and ERG6. FEMS Yeast Res 10:699–707
Mantzouridou F, Naziri E, Tsimidou MZ (2009) Squalene versus ergosterol formation using Saccharomyces cerevisiae: combined effect of oxygen supply, inoculum size, and fermentation time on yield and selectivity of the bioprocess. J Agric Food Chem 57:6189–6198
Martirosyan DM, Miroshnichenko LA, Kulakova SN, Pogojeva AV, Zoloedov VI (2007) Amaranth oil application for coronary heart disease and hypertension. Lipids Health Dis 6:1
Maury J, Asadollahi MA, Møller K, Clark A, Nielsen J (2005) Microbial isoprenoid production: an example of green chemistry through metabolic engineering. Biotechnology for the Future. Springer, Berlin, pp 19–51
Miettinen TA, Vanhanen H (1994) Serum concentration and metabolism of cholesterol during rapeseed oil and squalene feeding. Am J Clin Nutr 59:356–363
Nakashima T, Inoue T, Oka A, Nishino T, Osumi T, Hata S (1995) Cloning, expression, and characterization of cDNAs encoding Arabidopsis thaliana squalene synthase. Proc Natl Acad Sci 92:2328–2332
Nakazawa A, Matsuura H, Kose R, Kato S, Honda D, Inouye I, Kaya K, Watanabe MM (2012) Optimization of culture conditions of the thraustochytrid Aurantiochytrium sp. strain 18W-13a for squalene production. Bioresour Technol 109:287–291
Nakazawa A, Kokubun Y, Matsuura H, Yonezawa N, Kose R, Yoshida M, Tanabe Y, Kusuda E, Van Thang D, Ueda M (2014) TLC screening of thraustochytrid strains for squalene production. J Appl Phycol 26:29–41
Naziri E, Mantzouridou F, Tsimidou MZ (2011) Enhanced squalene production by wild-type Saccharomyces cerevisiae strains using safe chemical means. J Agric Food Chem 59:9980–9989
Obulesu T, Anandan R, Mathew S, Ganesan B, Krishna G, Lakra W, Asha K (2015) Antioxidant defence of dietary squalene supplementation on n-3 poly unsaturated fatty acids (PUFA)-mediated oxidative stress in young and aged rats. Fish Technol 52:48–52
Ohkuma T, Otagiri K, Tanaka S, Ikekawa T (1983) Intensification of host’s immunity by squalene in sarcoma 180 bearing ICR mice. J Pharmacobiodyn 6:148–151
Ohtake K, Saito N, Shibuya S, Kobayashi W, Amano R, Hirai T, Sasaki S, Nakano C, Hoshino T (2014) Biochemical characterization of the water-soluble squalene synthase from Methylococcus capsulatus and the functional analyses of its two DXXD (E) D motifs and the highly conserved aromatic amino acid residues. FEBS J 281:5479–5497
Orihara N, Kuzuyama T, Takahashi S, Furihata K, Seto H (1998) Studies on the biosynthesis of terpenoid compounds produced by actinomycetes. 3. Biosynthesis of the isoprenoid side chain of novobiocin via the non-mevalonate pathway in Streptomyces niveus. J Antibiot 51:676–678
Pan J-J, Solbiati JO, Ramamoorthy G, Hillerich BS, Seidel RD, Cronan JE, Almo SC, Poulter CD (2015) Biosynthesis of squalene from farnesyl diphosphate in bacteria: three steps catalyzed by three enzymes. ACS Cent Sci 1:77–82
Ravi Kumar S, Yamauchi I, Narayan B, Katsuki A, Hosokawa M, Miyashita K (2016) Squalene modulates fatty acid metabolism: enhanced EPA/DHA in obese/diabetic mice (KK-Ay) model. Eur J Lipid Sci Technol. doi:10.1002/ejlt.201600006
Ren LJ, Ji XJ, Huang H, Qu L, Feng Y, Tong QQ, Ouyang P-K (2010) Development of a stepwise aeration control strategy for efficient docosahexaenoic acid production by Schizochytrium sp. Appl Microbiol Biotechnol 87:1649–1656
Rohmer M, Knani M, Simonin P, Sutter B, Sahm H (1993) Isoprenoid biosynthesis in bacteria: a novel pathway for the early steps leading to isopentenyl diphosphate. Biochem J 295:517–524
Ronco AL, De Stéfani E (2013) Squalene: a multi-task link in the crossroads of cancer and aging. Funct Foods Health Dis 3:462–476
Salvador M, Aranda F, Gomez-Alonso S, Fregapane G (2003) Influence of extraction system, production year and area on Cornicabra virgin olive oil: a study of five crop seasons. Food Chem 80:359–366
Sánchez-Fidalgo S, Villegas I, Rosillo MÁ, Aparicio-Soto M, de la Lastra CA (2015) Dietary squalene supplementation improves DSS-induced acute colitis by downregulating p38 MAPK and NFkB signaling pathways. Mol Nutr Food Res 59:284–292
Sangari FJ, Pérez-Gil J, Carretero-Paulet L, García-Lobo JM, Rodríguez-Concepción M (2010) A new family of enzymes catalyzing the first committed step of the methylerythritol 4-phosphate (MEP) pathway for isoprenoid biosynthesis in bacteria. Proc Natl Acad Sci 107:14081–14086
Shechter I, Klinger E, Rucker M, Engstrom R, Spirito J, Islam M, Boettcher B, Weinstein D (1992) Solubilization, purification, and characterization of a truncated form of rat hepatic squalene synthetase. J Biol Chem 267:8628–8635
Singhal RS, Kulkarni P (1990) Effect of puffing on oil characteristics of Amaranth (Rajgeera) seeds. J Am Oil Chem Soc 67:952–954
Socaciu C, Faye M, Salin F, Pauly G, Gleizes M (1995) In vitro yeast (Saccharomyces cerevisiae) presqualene and squalene synthesis related to substrate and cofactor availability. C rendus de l’Acad Sci Ser III Sci de la vie 318:919–926
Spanova M, Daum G (2011) Squalene-biochemistry, molecular biology, process biotechnology, and applications. Eur J Lipid Sci Technol 113:1299–1320
Storm HM, Oh SY, Kimler BF, Norton S (1993) Radioprotection of mice by dietary squalene. Lipids 28:555–559
Thompson JF, Danley DE, Mazzalupo S, Milos PM, Lira ME, Harwood HJ (1998) Truncation of human squalene synthase yields active, crystallizable protein. Arch Biochem Biophys 350:283–290
Tsujimoto M (1916) A highly unsaturated hydrocarbon in shark liver oil. Ind Eng Chem 8:889–896
Tsujiwaki G, Yamamoto H, Minami K (1995) Manufacture of squalene with Candida famata. Japan Kokai Tokkyo Koho JP 07, 289, 272 (C.A.-124: 230184)
Uragami S, Koga H (1986) Bacterial production of squalene. Japan Kokai Tokkyo Koho JP 61:212–290
Welander PV, Hunter RC, Zhang L, Sessions AL, Summons RE, Newman DK (2009) Hopanoids play a role in membrane integrity and pH homeostasis in Rhodopseudomonas palustris TIE-1. J Bacteriol 191:6145–6156
Wyre C, Overton TW (2014) Use of a stress-minimisation paradigm in high cell density fed-batch Escherichia coli fermentations to optimise recombinant protein production. J Ind Microbiol Biotechnol 41:1391–1404
Xiao H, Yao Z, Peng Q, Ni F, Sun Y, Zhang C, Zhong Z (2016) Extraction of squalene from camellia oil by silver ion complexation. Sep Purif Technol 169:196–201
Xu W, Yang S, Zhao J, Su T, Zhao L, Liu J (2014) Improving coenzyme Q (8) production in Escherichia coli employing multiple strategies. J Ind Microbiol Biotechnol 41:1297–1303
Xu W, Chai C, Shao L, Yao J, Wang Y (2016) Metabolic engineering of Rhodopseudomonas palustris for squalene production. J Ind Microbiol Biotechnol 43:719–725
Yoon J, Matsuo Y, Matsuda S, Adachi K, Kasai H, Yokota A (2008) Rubritalea sabuli sp. nov., a carotenoid-and squalene-producing member of the family Verrucomicrobiaceae, isolated from marine sediment. Int J Syst Evol Microbiol 58:992–997
Yue C-J, Jiang Y (2009) Impact of methyl jasmonate on squalene biosynthesis in microalga Schizochytrium mangrovei. Process Biochem 44:923–927
Zahiri HS, Yoon SH, Keasling JD, Lee SH, Kim SW, Yoon SC, Shin YC (2006) Coenzyme Q (10) production in recombinant Escherichia coli strains engineered with a heterologous decaprenyl diphosphate synthase gene and foreign mevalonate pathway. Metab Eng 8:406–416
Acknowledgments
We thank Professor Arno Müllbacher (Australian National University) for critical reading of the manuscript. This paper was supported by the Shaanxi Science and Technology Innovation Project 2016KTCQ03-07 (to Y. Wang). X. Ma was supported by a Research Support Grant (2015NQ03) from the Xi’an Medical University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xu, W., Ma, X. & Wang, Y. Production of squalene by microbes: an update. World J Microbiol Biotechnol 32, 195 (2016). https://doi.org/10.1007/s11274-016-2155-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-016-2155-8