Abstract
The LATERAL ORGAN BOUNDARIES (LOB) DOMAIN (LBD) proteins are plant-specific transcriptional factors (TFs) functioning in the growth and development of Arabidopsis and other plant species. However, the involvement of LBD TFs in regulating the ripening of economically important fruits is largely unknown. In the present study, four full-length LBD genes, designated as MaLBD1–MaLBD4, were isolated and characterized from banana fruit. Expressions of MaLBD1–MaLBD4 in fruit with three ripening characteristics revealed that MaLBD1/2/3 were ethylene-inducible, and their transcript levels obviously increased during fruit ripening, while MaLBD4 changed slightly. Moreover, the activities of MaLBD1/2/3 promoters were activated after ethylene treatment, further supporting their involvement in fruit ripening. Subcellular localization showed that MaLBD1/2/3 were nuclear proteins, and a transactivation assay in protoplasts demonstrated that MaLBD1/2/3 had transactivation activity. More importantly, a transient expression assay further indicated that MaLBD1/2/3 were transcriptional activators that regulated ripening-related MaEXP1/2 expressions by directly binding to MaEXP1/2 promoters. These results suggest that MaLBDs are involved in regulating fruit ripening, in part by transcriptional activation of the EXPANSIN expression related to cell wall modification. Taken together, our findings provide some new information about the functions of LBD TFs involved in the regulation of fruit ripening.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fruit ripening represents a complex and highly coordination of physiological and biochemical processes, accompanying with changes of thousand genes associated with color, flavor, aroma, texture, and nutritional value of the flesh (Klee and Giovannoni 2011; Gapper et al. 2013), which are influenced by internal and external cues regulated by many critical transcription factors (TFs) (Martel et al. 2011). A major breakthrough in dissecting the transcriptional control of fruit ripening was the identification of several genes underlying rare mutations that completely abolish the normal ripening process of tomato, including ripening inhibitor (RIN), Colorless nonripening (CNR), and nonripening (NOR) (Seymour et al. 2013). These three mutants loci that all harbor TFs belong to MADS, NAC, and SBP TFs family, respectively (Seymour et al. 2013). Other ripening TFs, such as HD-zip protein LeHB1, TAGL1, SlAP2a, and SlERF6, also have been identified in tomato (Gapper et al. 2013). In addition, several TFs, like ERFs and EIN3-like, are suggested to be involved in the ripening of other fruits, such as apples, kiwifruit, plums, and longan fruit (Wang et al. 2007; El-Sharkawy et al. 2009; Yin et al. 2010; Kuang et al. 2012). Together, these studies highlight the dedicated transcriptional control of the fruit ripening process.
The LATERAL ORGAN BOUNDARIES (LOB) DOMAIN (LBD) (also known as ASL for ASYMMETRIC LEAVES2-LIKE) proteins are plant-specific TFs, characterized by a typical N-terminal LOB/AS2 domain with a CX2CX6CX3C motif and a Leu zipper-like sequence (Husbands et al. 2007; Majer and Hochholdinger 2011). LBD proteins exhibit varied expression patterns ranging from temporal to tissue differences, indicating that they may function in diverse processes (Shuai et al. 2002). Functional characterization of several LBD members in Arabidopsis, rice, maize, and poplar reveals that LBD proteins play critical roles in defining lateral organ boundaries and in regulating many aspects of plant development, including root, leaf, inflorescence, and embryo development (Liu et al. 2005; Soyano et al. 2008; Yordanov et al. 2010; Majer and Hochholdinger 2011; Lee et al. 2013; Wang et al. 2013). In addition to developmental processes, LBD proteins also have been found to be directly involved in the metabolic regulation in response to nitrogen deficiency (Rubin et al. 2009), anthocyanin synthesis (Rubin et al. 2009), auxin signal cascades (Okushima et al. 2007; Lee et al. 2009; Rast and Simon 2012), and jasmonate signaling (Thatcher et al. 2012a). However, LBD TFs have not been previously implicated in the regulation of fruit ripening.
Bananas are the world’s fourth most important food crop after rice, wheat, and maize, with a vast majority of the crop grown and consumed in the tropical and subtropical countries (Cruz-Hernández and Paredes-lópez 2012). Banana is a typical climacteric fruit, characterized by a peak in ethylene production that orchestrates ripening-associated processes, which results in a short postharvest life of 10–15 days at ambient temperature (Bapat et al. 2010; Shan et al. 2012; Xiao et al. 2013). Accordingly, it is important to gain a better understanding of the mechanism of banana fruit ripening, in order to explore effective technologies to maintain the quality and storage potential of this fruit. Very recently, several ripening TFs, including MADS-box, NAC, and ERF, have been identified in banana fruit, as well as their transcriptional control of ripening (Elitzur et al. 2010; Choudhury et al. 2012; Shan et al. 2012; Xiao et al. 2013). However, it is clear that multiple TFs influence the ripening process (Klee and Giovannoni 2011), indicating that the full complexity of transcriptional regulatory mechanism remains elusive. Moreover, recently, the whole-genome sequence for banana is reported (D’Hont et al. 2012), which has given researchers access to a new source of vast numbers of TFs. In this study, four LBD TFs were isolated and characterized from banana fruit. Their expression patterns in association with three different ripening treatments were analyzed. Moreover, the interactions of the ripening-related MaLBDs with cell wall-modifying genes MaEXPs encoding for EXPANSIN protein were investigated. Our findings suggest that MaLBDs may be involved in banana fruit ripening via in part by transcriptional activation of cell wall-loosening-related genes MaEXPs.
Materials and Methods
Plant materials and Treatments
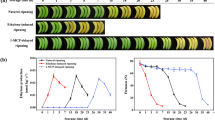
Pre-climacteric banana (Musa acuminata, AAA group, cv. Cavendish) fruits at 75–80 % maturation were obtained from a local commercial plantation near Guangzhou, China. Three postharvest treatments, including a control (natural ripening), ethylene-induced ripening (100 μl l−1 ethylene, 18 h), and 1-methylcyclopropene (1-MCP)-delayed ripening (0.5 μl l−1 1-MCP, 18 h) were performed to create three different ripening characteristics, and samples were taken as described previously (Shan et al. 2012). All assessments were conducted using three biological replicates.
RNA Extraction, Gene Isolation, and Sequence Analysis
Total RNA was extracted using the hot borate method of Wan and Wilkins (1994). Total RNA extract was treated with DNAse I (Promega, Madison, WI, USA), and the resulting DNA-free total RNA was used as the template for reverse transcription PCR (RT-PCR). The first-strand cDNA of the product was subjected to PCR amplification.
According to gene annotation and bioinformatics analysis, four full-length LBD genes with complete start and stop codes, termed MaLBD1-MaLBD4, were selected randomly from banana whole-genome sequence (D’Hont et al. 2012), and the sequences were verified by further cloning from banana fruit pulp and sequencing. Alignments were carried out on ClustalX (version 1.83) and GeneDoc software, and a phylogenetic tree was constructed using the minimum evolution test method with the MEGA5 program.
Quantitative Real-time PCR Analysis
Synthesis of first-strand cDNA and all RT-qPCR analysis were performed as described previously (Chen et al. 2011; Shan et al. 2012). The sequences of all primers used for RT-qPCR analysis are listed in Supplementary Table S1. MaRPS4 (ribosomal protein 2) was selected as a reference gene according to our previous study on the selection of reliable reference genes under different experimental conditions (Chen et al. 2011). All RT-qPCR reactions were normalized using Ct value corresponding to the reference gene. The relative expression levels of target gene were calculated with the formula 2−ΔΔCT. Three independent biological replicates were used in the analysis.
Promoter Activity Assay
Genomic DNA was extracted from banana leaves using the DNeasy Plant Mini Kit (Qiagen). The promoter of MaLBDs gene was isolated using the GenomeWalker Kit (Clontech) with nest PCR according to the manufacturer’s instructions. Two gene-specific primers for the nest PCR are listed in Supplementary Table S1. The amplification product was cloned into pGEM-Teasy vector (Promega, Madison, WI, USA) and sequenced. Conserved cis-element motifs of promoter were predicted using PLACE (http://www.dna.affrc.go.jp/PLACE/signalscan.html) and Plant-CARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/) databases.
For promoter activity assay, the MaLBD promoter region was amplified by PCR using the specific primers listed in Supplementary Table S1. The PCR product was inserted into the pGreenII 0800-LUC double reporter vector (Hellens et al. 2005) at the HindIII and BamHI sites to fuse it with the Firefly luciferase (LUC) reporter gene (MaLBDs pro-LUC); a Renilla (REN) luciferase under the control of the 35S promoter at the same vector was used as a positive control. The construct CaMV35S-REN/MaLBDs pro-LUC was transformed into tobacco BY-2 protoplasts by polyethylene glycol (PEG) methods as described previously (Abel and Theologis 1994). The promoter activity is indicated by the ratio of LUC to REN. The transformed protoplasts were subjected to 0 mM (control) or 0.8 mM of ethylene releaser (ethrel) treatment and then incubated at 23 °C for 14 h, and LUC and REN luciferase activities were assayed using the dual luciferase assay kits (Promega). The analysis was carried out using the Luminoskan Ascent Microplate Luminometer (Thermo) according to the manufacturer’s instructions, with a 5-s delay and 15-s integrated measurements. At least six assay measurements were included for each.
Subcellular Localization of MaLBDs
The coding sequence of MaLBDs without the stop codon was amplified by PCR (primers are listed in Supplementary Table S1) and subcloned into the pUC-GFP vector, in frame with the green fluorescent protein (GFP) sequence, resulting in 35S::gene-GFP vectors under the control of the cauliflower mosaic virus (CaMV) 35S promoter. The fusion constructs and the control GFP vector were used for transient assays in tobacco BY-2 suspension culture cell protoplasts as described above. GFP fluorescence was observed with a fluorescence microscope (Zeiss Axioskop 2 Plus). All transient expression assays were repeated at least three times.
Transient Assay in Protoplast
A dual luciferase assay was performed in the transient assay. For transactivation analysis of MaLBDs, the coding sequence of MaLBDs without the stop codon was cloned into the reconstructed GAL4DBD vector as effector. The double reporter vector includes a GAL4-LUC and a internal control REN driven by the 35S promoter. GAL4-LUC contains five copies of GAL4-binding element and minimal TATA region of the 35S promoter of CaMV, and these sequences are located upstream of the LUC.
For the assay of the binding activity of MaLBDs to the MaEXPs promoter, two pGreenII vectors, pGreenII 0800-LUC reporter vector and pGreenII 0029 62-SK effector vector, were used (Hellens et al. 2005). The MaEXPs promoter was cloned into pGreen 0800 at the HindIII and BamHI sites to fuse it with the LUC reporter gene (MaEXPs pro-LUC). MaLBDs were cloned into the pGreenII 62-SK vector. The pGreenII 0800-LUC vector carried a REN under the control of the 35S promoter as a positive control.
The constructed effector and reporter plasmids were co-transfected into BY-2 protoplasts as described above. The resulted protoplasts were incubated at 23 °C for 16 h in the dark and then were harvested to determine the LUC and REN luciferase activities as described above. The transactivation ability of MaLBDs and the binding activity of MaLBDs to the MaEXPs promoter are indicated by the ratio of LUC to REN. At least six transient assay measurements were included for each pair. The primers used in the construction of the effector and reporter vectors are listed in Supplementary Table S1.
Statistical Analysis
The experiment was performed using a completely randomized design. Data were presented as mean ± standard error (S.E.). Least significant difference (LSD) at the 5 % level was analyzed by DPS software (version 3.01; Zhejiang University, Hangzhou, China).
Results
Sequence Analysis of Banana Fruit MaLBDs Genes
Based on the banana whole-genome sequence (D’Hont et al. 2012), four LBD full-length cDNAs, designated as MaLBD1 to MaLBD4, were cloned from banana fruit pulp. MaLBD1–MaLBD4 were predicted to encode proteins of 236, 211, 287, and 250 aa, with calculated molecular weights of 24.4, 23.0, 30.7, and 26.7 kDa, and pI values of 7.46, 9.22, 6.20, and 6.14, respectively. Alignment of the deduced amino acid sequences of MaLBDs clearly showed that the conserved DNA-binding domain, designated as the LOB domain, existed in the N-terminal regions of the MaLBDs proteins (Fig. 1), which is a defining character of the LBD TF family (Majer and Hochholdinger 2011). The LOB domain comprises a C-domain presumably required for DNA binding which contains four perfectly conserved cysteine (C) residues in a CX2CX6CX3C zinc finger-like motif (Fig. 1). Moreover, an invariant glycine residue and a C-terminal leucine-zipper-like motif responsible for probable protein-protein interactions, which includes five hydrophobic amino acids (valine, isoleucine, leucine) separated by six variable amino acids, are observed within the LOB domain (Fig. 1) (Shuai et al. 2002; Matsumura et al. 2009; Majer and Hochholdinger 2011).
Alignment of the deduced amino acid sequences of four banana MaLBDs and their respective homologs from grape (VvLBD1, XP_002279521), cacao (TcLBD1, EOY20040), and soybean (GmLBD1, XP_003541285). Identical and similar amino acids were presented by black and gray shading, respectively. Gaps were introduced to optimize alignment. The LOB domain was underlined. The C-motif (CX2CX6CX3C) was included in the rectangle, and the invariant glycine residue was indicated by the asterisk
To determine the evolutionary relationship among MaLBDs and those of Arabidopsis and apple and then group them into established subfamilies, a phylogenetic tree was generated with full-length protein sequences (Fig. 2). The whole LBD proteins of Arabidopsis and apple could be classified into two major groups, I and II. Class I and class II families were further divided into 8 and 2 groups, as reported by Wang et al. (2013). In class I, the subgroups were named from class I a to class I i, while class II a and class II b comprised the class II group. Based on the tree, the four banana MaLBD proteins were clustered into four subgroups. MaLBD1 to MaLBD4 belonged to class I a, class I b, class II a, and class I f, respectively (Fig. 2). Together, these data indicate that MaLBD1 to MaLBD4 may exhibit diverse functions.
Phylogenetic tree of LBDs. Four banana MaLBDs (black circles) were aligned with the Arabidopsis and apple LBD family as designated by Wang et al. (2013). The multiple alignment was made using ClustalW, and the phylogenetic tree was constructed with MEGA 5.0 using a bootstrap test of phylogeny with minimum evolution test and default parameters
Expression of MaLBDs Genes during Fruit Ripening
The data related with fruit ripening and softening, including changes in fruit firmness and ethylene production in banana fruit in three different ripening treatments caused by natural, ethylene-induced, and 1-MCP-delayed treatments, have been described in the previous publication (Shan et al. 2012). Control natural fruit showed the ethylene climacteric at 15 days and reached a peak at 21 days. Ethylene treatment accelerated fruit ripening, with a climacteric peak appearance at 3 days. In contrast, 1-MCP treatment delayed ethylene production, with a peak appearance at day 30 (Shan et al. 2012).
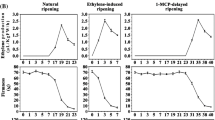
To investigate the possible role of MaLBD genes during banana fruit ripening, RT-qPCR was carried out to determine their mRNA accumulations in banana fruit pulp with those three different ripening characteristics. As shown in Fig. 3, among the four MaLBD genes, MaLBD1, MaLBD2, and MaLBD3 were strongly induced after ethylene treatment, and their transcript levels in fruit pulp of natural, ethylene-induced, or 1-MCP-delayed ripening clearly increased with ethylene evolution during ripening. However, MaLBD4 was expressed constitutively and its transcripts changed slightly during ripening (Fig. 3). These results indicate that MaLBD1, MaLBD2, and MaLBD3 may be more associated with fruit ripening. Thus, these three MaLBD genes were selected for the following analysis.
Expressions of MaLBDs in pulp during three ripening characteristics: natural (control), ethylene-induced, and 1-MCP-delayed ripening. The expression levels of each gene are expressed as a ratio relative to the harvest time (0 days of control), which was set at 1. Each value represents the mean ± S.E. of three biological replicates. The physiology data related with fruit ripening and softening, including changes in fruit firmness and ethylene production in banana fruit with these three different ripening characteristics, has been described in the work of Shan et al. (2012)
Promoter Activities of MaLBD1/2/3 are Induced by Ethylene
To further understand the regulation of MaLBD1/2/3 expression by ethylene, the upstream promoter region sequences from the transcription start site of MaLBD1/2/3 were isolated from the genome of M. acuminata using genome walking PCR methods. Amplification using specific primers for MaLBD1/2/3 upstream sequences resulted in 1,585-, 491-, and 1,067-bp amplicon, subsequently named MaLBD1, MaLBD2, and MaLBD3 promoter, respectively. Analysis of these promoters using the PLACE and Plant-CARE databases was illustrated in Supplementary Tables S2, S3, and S4. In addition to the core cis-acting elements like the TATA box and CAAT box, several regulatory motifs that are potentially related to light response and phytohormones such as salicylic acid (SA), methyl jasmonate (MeJA), auxin, abscisic acid (ABA), and gibberellin were also found. In particular, PERE motif, which has also been shown to function as an ethylene-responsive element (Solano et al. 1998; Yin et al. 2010), was observed in MaLBD1/2/3 promoters (Supplementary Tables S2, S3, and S4). These results indicate that the activities of MaLBD1/2/3 promoters may be modulated by ethylene.
To analyze MaLBD1/2/3 promoter activities in response to ethylene, a transient protoplast assay was conducted using a dual luciferase reporter vector containing the Firefly LUC driven by the MaLBD1/2/3 promoter and the REN luciferase driven by the CaMV35S promoter (CaMV35S-REN/MaLBDs pro-LUC, Fig. 4a). As shown in Fig. 4b–d, in BY-2 protoplast transfected with CaMV35S-REN/MaLBD1/2/3 pro-LUC, MaLBD1/2/3 pro-LUC/REN ratio apparently increased after ethylene treatment (Fig. 4b–d), suggesting that MaLBD1/2/3 promoter activities were induced by ethylene. These observations, together with the accumulation of MaLBD1/2/3 mRNA during ripening or following ethylene treatment, further strengthen the notion that MaLBD1/2/3 are likely to be involved in banana fruit ripening.
MaLBD promoter activities in response to ethylene. The top panel (a) shows schematics of the dual luciferase reporter vectors containing MaLBD promoters (CaMV35S-REN/MaLBDs pro-LUC). The bottom panel (b–d) shows promoter activities of MaLBD1/2/3 in response to ethylene. The reporter constructs containing MaLBD promoters were transiently transformed into tobacco BY-2 protoplasts using a modified PEG method and test for ethylene induction. After incubation for 14 h, LUC and REN luciferase activities were assayed, and the promoter activity is indicated by the ratio of LUC to REN. The asterisk indicates a significant difference at the 5 % level compared to the control. Each value represents the means of six biological replicates, and vertical bars represent the S.E
MaLBD1/2/3 are Nucleus-localized Proteins
LBD proteins are TFs that have previously been shown to localize to the nucleus (Husbands et al. 2007; Okushima et al. 2007; Rubin et al. 2009). To examine the subcellular localization of MaLBD1/2/3 in vivo, the full-length coding sequences of MaLBD1/2/3 were fused in-frame with the GFP gene and transient-expressed in tobacco BY-2 protoplasts. As shown in Fig. 5, the fluorescence of MaLBD1/2/3 proteins was localized exclusively in the nucleus, whereas the fluorescence from the GFP control was distributed throughout the entire cell (Fig. 5). These results indicate nuclear localization of MaLBD1/2/3, consistent with their predicted role as TFs.
Subcellular localization of MaLBDs in tobacco BY-2 protoplasts. Protoplasts were transiently transformed with 35S:MaLBDs–GFP constructs or 35S:GFP vector using a modified PEG method. GFP fluorescence was observed with a fluorescence microscope. VirD2NLS-mCherry was included in each transfection to serve as a control for successful transfection, as well as for nuclear localization. Images were taken in a dark field for green fluorescence, while the outline of the cell and the merged were photographed in a bright field. Bars, 25 μm
MaLBD1/2/3 Exhibit Transactivation Activity in Protoplasts
To investigate the abilities of MaLBD1/2/3 to activate transcription, the transactivation activities of MaLBD1/2/3 were examined using transient expression assay in BY-2 protoplasts. We employed a dual luciferase reporter plasmid harboring five copies of the Gal4-DNA binding element and minimal TATA region of 35S promoter fused to the LUC reporter, and a REN reporter under the control of the 35S promoter at the same vector was used as an internal control for successful transfection, while an effector plasmid encoding full-length MaLBD1/2/3 fused to the Gal4 DNA-binding domain (Gal4BD) (Fig. 6a). Compared with the GAL4-BD (pBD) negative control, MaLBD1/2/3 strongly activated the LUC reporter gene, and the LUC/REN ratio of MaLBD1/2/3 was 4.5-, 5.8-, and 0.9-fold higher than that of the negative control, respectively (Fig. 6b). These results indicate that MaLBD1/2/3 may function as transcriptional activators.
Transcriptional activation of MaLBDs in tobacco BY-2 protoplasts. a Reporter and effector constructs. The dual luciferase reporter construct contained the LUC reporter gene driven by the mini-35S (TATA box) plus five GAL4-binding elements. Each of the effectors contained a GAL4 DNA-binding domain (GAL4-BD), and pBD was used as a negative control. MaLBDs were fused with the GAL4-BD and driven by the 35S promoter plus the translation enhancer Ω sequence. b Transactivation ability of MaLBDs. Plasmid combinations of dual REN/LUC reporter, and effectors were cotransformed into BY-2 protoplasts. The protoplasts were incubated for 16 h, and the transactivation ability of MaLBDs is indicated by the ratio of LUC to REN. Each value represents the means of six biological replicates, and vertical bars represent the S.E. The asterisk indicates a significant difference at the 5 % level compared to the negative control pBD
MaLBD1/2/3 are Transcriptional Activators That Regulate MaEXP1/2 Genes
Recently, LBD18 is reported to function as a transcriptional activator that regulates the expression of EXP14/17 by directly binding to EXP14/17 promoters, facilitating the emergence of lateral root (Lee et al. 2013; Lee and Kim 2013). Five EXP genes have been isolated from banana fruit, and cell wall loosening mediated mainly by MaEXP1/2 is related to fruit ripening (Mbéguié-A-Mbéguié et al. 2009). We also investigated expressions of MaEXP1/2 in banana fruit pulp during ripening. As shown in Supplementary Fig. S1, similar with the expression patterns of MaLBD1/2/3, MaEXP1/2 were strongly induced after ethylene treatment, and their transcript levels in fruit pulp of natural, ethylene-induced, or 1-MCP-delayed ripening obviously increased with ethylene evolution during ripening (Supplementary Fig. S1), further confirming that MaEXP1/2 are associated with fruit ripening. Thus, it is reasonable to speculate that ripening-related MaLBDs might function as transcriptional activators to regulate MaEXP1/2 expression in banana fruit.
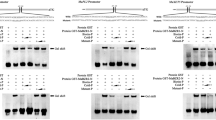
To confirm the speculation, transient expression assay in BY-2 protoplasts was performed with a dual luciferase reporter plasmid harboring the MaEXP1/2 promoter fused to LUC, and the REN driven by the CaMV35S promoter (CaMV35S-REN/MaLBDs pro-LUC, Fig. 7a), while an effector plasmid harboring MaLBD1/2/3 was expressed under the control of the CaMV 35S promoter (Fig. 7a). The results showed that MaLBD1/2/3 obviously activated the MaEXP1 promoter activity, with 10.1-, 9.8-, and 4.9-fold enhancement of LUC/REN ratio than that of the negative empty control vector (Fig. 7b). Similarly, MaLBD1/2/3 also activated the MaEXP2 promoter activity (Fig. 7b). These results clearly demonstrate that MaLBD1/2/3 are transcriptional activators that directly regulate MaEXP1/2 gene expressions.
MaLBDs activate the MaEXP1/2 promoter in a dual luciferase assay. a Schematic representation of the double reporters and effector plasmids used in the assay. The double-reporter plasmid contained the MaEXP1/2 promoter fused to LUC luciferase and REN luciferase drove by CaMV35S. The effector plasmid contained the MaLBDs drove by the CaMV35S. b MaLBDs activate the MaEXP1/2 promoter. The reporter and effector vectors, as indicated, were co-transformed into tobacco BY-2 protoplasts. The protoplasts were incubated for 16 h, and the activation of MaEXP1/2 promoter by MaLBDs was indicated by the ratio of LUC to REN. Each value represents the means of six biological replicates, and vertical bars represent the S.E. The asterisk indicates a significant difference at the 5 % level compared to the empty effector
Discussion
LBD proteins are a plant-specific TFs family with 42 members in Arabidopsis (Shuai et al. 2002), 35 in rice (Yang et al. 2006), 57 in poplar (Zhu et al. 2007), 43 in maize (Schnable et al. 2009), and 58 in apple (Wang et al. 2013). LBD proteins are characterized by the presence of a highly conserved N-terminal DNA-binding domain termed the LOB domain, whereas little conservation is found in their C-termini (Majer and Hochholdinger 2011). In this study, based on the banana whole-genome sequence, four banana fruit LBD genes, designated as MaLBD1 to MaLBD4, were cloned. Similar to the previous reports, alignment of the four MaLBD proteins showed that they shared a highly conserved LOB domain (Fig. 1). Phylogenetic analysis and evolutionary relationship of the LBD proteins have been intensively investigated in Arabidopsis and rice (Shuai et al. 2002; Yang et al. 2006; Matsumura et al. 2009), which divides the LBD gene family into class I and class II, characterizing by the presence or absence of functional leucine-zipper-like domains, respectively, and the majority of LBD proteins belong to class I (Majer and Hochholdinger 2011). Similarly, the four MaLBDs fell into class I and class II of the previously characterized LBD proteins, in which MaLBD1/2/4 belong to class I, while MaLBD3 belongs to class II (Fig. 2).
Most LBD proteins so far identified have been functionally characterized in Arabidopsis and crop plants. These LBD proteins are involved in regulating plant lateral organ such as embryo, root, leaf, inflorescence and flower development, anthocyanin or nitrogen metabolism, or responses to phytohormones such as cytokinin, auxin, gibberellins, and JA (Majer and Hochholdinger 2011). Recently, LBD proteins involved in biotic stress response are reported (Thatcher et al. 2012a, b). In the present work, among the four banana fruit MaLBD genes, MaLBD1/2/3 were ripening-inducible; moreover, their expressions also apparently upregulated after ethylene treatment (Fig. 3). Further, when MaLBD1/2/3 promoters transiently expressed in protoplasts, they were all activated by ethylene applications (Fig. 4). These data indicate that MaLBD1/2/3 might be associated with banana fruit ripening. This is the first time, to our knowledge, that a possible role for LBD proteins has been demonstrated in the involvement of fruit ripening. LBD proteins are suggested to act as TFs based on their nuclear localization and DNA-binding activity (Shuai et al. 2002; Husbands et al. 2007; Naito et al. 2007; Rubin et al. 2009). It has been demonstrated that Arabidopsis ASL4 (AtLOB) specifically binds to a DNA motif 5′-GCGGCG-3′ designated as the LBD motif (Husbands et al. 2007). Our present work also showed that MaLBD1/2/3 were localized in the nucleus (Fig. 5). Moreover, MaLBD1/2/3 possessed transcription-activating abilities in BY-2 protoplasts (Fig. 6). These results suggest that MaLBD1/2/3 may function as transcriptional activators. Accordingly, it is interesting to explore their downstream targets that related to banana fruit ripening.
Following the discovery of several important ripening-related TFs in tomato fruit, including MADS-RIN, NOR, and CNR, the identification of their downstream targets has been extenstively investigated. For example, MADS-RIN directly controls the expression of a wide range of other ripening-related genes, targeting the promoters of genes involved in ethylene biosynthesis and perception, cell wall metabolism, carotenoid formation, aroma biosynthesis, and ATP generation (Seymour et al. 2013). As to LBD TFs, several downstream targets of LBD TFs have been identified (Majer and Hochholdinger 2011), and interestingly, Arabidopsis LBD18/ASL20 activates EXPANSINA14/17 (EXP14/17) expression by directly binding to EXP14/17 promoters to enhance lateral root formation during the auxin response (Lee et al. 2013; Lee and Kim 2013). Expansins are primary wall-loosening factors that directly induce turgor-driven wall extension (Cosgrove 2000). Secondary wall-loosening factors such as xyloglucan endotransglicoylase/hydrolases (XET/XTH), glucanases, and polygalacturonase (PG), which enzymatically modify the structures of the cell wall, render it more responsive to wall-loosening events primarily mediated by expansins (Cosgrove 2000; Péret et al. 2009). Fruit softening is a typical textural change associated with ripening, and it has been suggested that cell wall modification enzymes and expansions contribute to fruit softening significantly (Cruz-Hernández and Paredes-lópez 2012). Previously, five EXP genes have been isolated from banana fruit, and MaEXP1/2 expressions are related to fruit ripening (Mbéguié-A-Mbéguié et al. 2009). During fruit ripening, MaEXP1/2 transcript levels obviously increased, as well as strongly induced after ethylene treatment (Supplementary Fig. S1). Using a transient expression assay in BY-2 protoplasts, we showed that ripening-related MaLBD1/2/3 are transcriptional activators that regulate MaEXP1/2 gene expressions by directly binding to MaEXP1/2 promoters (Fig. 7). In addition to EXP, other cell wall modification factors, such as XET/XTH, PG, and PME, are also reported to be associated with banana fruit ripeing (Mbéguié-A-Mbéguié et al. 2009); however, whether XET, PG, and PME are direct targets of MaLBD1/2/3 needs to be further investigated. In addition, it has been shown that several LBD proteins interact with MYB and bHLH TFs to co-regulate hormone-mediated organogenesis (Husbands et al. 2007; Guo et al. 2008; Wang et al. 2013); it is thus interesting to study whether MaLBD1/2/3 could interact with the reported ripening-related TFs of banana fruit, including MADSs (Choudhury et al. 2012), NACs (Shan et al. 2012), and ERFs (Xiao et al. 2013) to be involved in fruit ripening.
Overall, four banana fruit MaLBD genes were identified and characterized. Gene expression and promoter analysis demonstrated the possible involvement of MaLBD1/2/3 in fruit ripening. MaLBD1/2/3 were nucleus-localized proteins and had transactivation activity in protoplasts. More importantly, MaLBD1/2/3 were transcriptional activators that regulated MaEXP1/2 gene expressions by directly binding to MaEXP1/2 promoters. Taken together, our results elucidate the molecular mechanism by which MaLBDs involved in regulating fruit ripening, in part by transcriptional activation of the EXPANSIN expression related to cell wall modification. Our findings provide some new information about the functions of LBD TFs involved in the regulation of fruit ripening.
References
Abel S, Theologis A (1994) Transient transformation of Arabidopsis leaf protoplasts: a versatile experimental system to study gene expression. Plant J 5:421–427
Bapat VA, Trivedi PK, Ghosh A, Sane VA (2010) Ripening of fleshy fruit: molecular insight and the role of ethylene. Biotechnol Adv 28:94–107
Chen L, Zhong HY, Kuang JF, Li JG, Lu WJ, Chen JY (2011) Validation of reference genes for RT-qPCR studies of gene expression in banana fruit under different experimental conditions. Planta 234:377–390
Choudhury SR, Roy S, Nag A, Singh SK, Sengupta DN (2012) Characterization of an AGAMOUS-like MADS box protein, a probable constituent of flowering and fruit ripening regulatory system in banana. PLoS One 7:e44361
Cosgrove DJ (2000) Loosening of plant cell walls by expansins. Nature 407:321–326
Cruz-Hernández A, Paredes-López O (2012) Fruit quality: new insights for biotechnology. Crit Rev Food Sci Nutr 52:272–289
D’Hont A, Denoeud F, Aury JM, Baurens FC, Carreel F, Garsmeur O et al (2012) The banana (Musa acuminata) genome and the evolution of monocotyledonous plants. Nature 488:213–217
Elitzur T, Vrebalov J, Giovannoni JJ, Goldschmidt EE, Friedman H (2010) The regulation of MADS-box gene expression during ripening of banana and their regulatory interaction with ethylene. J Exp Bot 61:1523–1535
El-Sharkawy I, Sherif S, Mila I, Bouzayen M, Jayasankar S (2009) Molecular characterization of seven genes encoding ethylene responsive transcriptional factors during plum fruit development and ripening. J Exp Bot 60:907–922
Gapper NE, McQuinn RP, Giovannoni JJ (2013) Molecular and genetic regulation of fruit ripening. Plant Mol Biol 82:575–591
Guo M, Thomas J, Collins G, Timmermans MC (2008) Direct repression of KNOX loci by the ASYMMETRIC LEAVES1 complex of Arabidopsis. Plant Cell 20:48–58
Hellens R, Allan A, Friel E, Bolitho K, Grafton K, Templeton M, Karunairetnam S, Gleave A, Laing W (2005) Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods 1:13
Husbands A, Bell EM, Shuai B, Smith HMS, Springer PS (2007) LATERAL ORGAN BOUNDARIES defines a new family of DNA-binding transcription factors and can interact with specific bHLH proteins. Nucleic Acids Res 35:6663–6671
Klee HJ, Giovannoni JJ (2011) Genetics and control of tomato fruit ripening and quality attributes. Annu Rev Genet 45:41–59
Kuang JF, Chen JY, Luo M, Wu KQ, Sun W, Jiang YM, Lu WJ (2012) Histone deacetylase HD2 interacts with ERF1 and is involved in longan fruit senescence. J Exp Bot 63:441–454
Lee HW, Kim J (2013) EXPANSINA17 up-regulated by LBD18/ASL20 promotes lateral root formation during the auxin response. Plant Cell Physiol 54:1600–1611
Lee HW, Kim MJ, Kim NY, Lee SH, Kim J (2013) LBD18 acts as a transcriptional activator that directly binds to the EXPANSIN14 promoter in promoting lateral root emergence of Arabidopsis. Plant J 73:212–224
Lee HW, Kim NY, Lee DJ, Kim J (2009) LBD18/ASL20 regulates lateral root formation in combination with LBD16/ASL18 downstream of ARF7 and ARF19 in Arabidopsis. Plant Physiol 151:1377–1389
Liu H, Wang S, Yu X, Yu J, He X, Zhang S, Shou H, Wu P (2005) ARL1, a LOB-domain protein required for adventitious root formation in rice. Plant J 43:47–56
Majer C, Hochholdinger F (2011) Defining the boundaries: structure and function of LOB domain proteins. Trends Plant Sci 16:47–52
Martel C, Vrebalov J, Tafelmeyer P, Giovannoni JJ (2011) The tomato MADS-box transcription factor RIPENING INHIBITOR interacts with promoters involved in numerous ripening processes in a COLORLESS NONRIPENING-dependent manner. Plant Physiol 157:1568–1579
Matsumura Y, Iwakawa H, Machida Y, Machida C (2009) Characterization of genes in the ASYMMETRIC LEAVES2/LATERAL ORGAN BOUNDARIES (AS2/LOB) family in Arabidopsis thaliana, and functional and molecular comparisons between AS2 and other family members. Plant J 58:525–537
Mbéguié-A-Mbéguié D, Hubert O, Baurens FC, Matsumoto T, Chillet M, Fils-Lycaon B, Sidibé-Bocs S (2009) Expression patterns of cell wall-modifying genes from banana during fruit ripening and in relationship with finger drop. J Exp Bot 60:2021–2034
Naito T, Yamashino T, Kiba T, Koizumi N, Kojima M et al (2007) A link between cytokinin and ASL9 (ASYMMETRIC LEAVES 2 LIKE 9) that belongs to the AS2/LOB (LATERAL ORGAN BOUNDARIES) family genes in Arabidopsis thaliana. Biosci Biotechnol Biochem 71:1269–1278
Okushima Y, Fukaki H, Onoda M, Theologis A, Tasaka M (2007) ARF7 and ARF19 regulate lateral root formation via direct activation of LBD/ASL genes in Arabidopsis. Plant Cell 19:118–130
Péret B, De Rybel B, Casimiro I, Benková E, Swarup R, Laplaze L, Beeckman T, Bennett MJ (2009) Arabidopsis lateral root development: an emerging story. Trends Plant Sci 14:399–408
Rast MI, Simon R (2012) Arabidopsis JAGGED LATERAL ORGANS acts with ASYMMETRIC LEAVES2 to coordinate KNOX and PIN expression in shoot and root meristems. Plant Cell 24:2917–2933
Rubin G, Tohge T, Matsuda F, Saito K, Scheible WR (2009) Members of the LBD family of transcription factors repress anthocyanin synthesis and affect additional nitrogen responses in Arabidopsis. Plant Cell 21:3567–3584
Schnable PS, Ware D, Fulton RS, Stein JC, Wei F, Pasternak S et al (2009) The B73 maize genome: complexity, diversity, and dynamics. Science 326:1112–1115
Seymour GB, Østergaard L, Chapman NH, Knapp S, Martin C (2013) Fruit development and ripening. Annu Rev Plant Biol 64:219–241
Shan W, Kuang JF, Chen L, Xie H, Peng HH, Xiao YY, Li XP, Chen WX, He QG, Chen JY, Lu WJ (2012) Molecular characterization of banana NAC transcription factors and their interactions with ethylene signalling component EIL during fruit ripening. J Exp Bot 63:5171–5187
Shuai B, Reynaga-Peña CG, Springer PS (2002) The lateral organ boundaries gene defines a novel, plant-specific gene family. Plant Physiol 129:747–761
Solano R, Stepanova A, Chao QM, Ecker JR (1998) Nuclear events in ethylene signaling: a transcriptional cascade mediated by ETHYLENE-INSENSITIVE3 and ETHYLENE-RESPONSE-FACTOR1. Genes Dev 12:3703–3714
Soyano T, Thitamadee S, Machida Y, Chua NH (2008) ASYMMETRIC LEAVES2-LIKE19/LATERAL ORGAN BOUNDARIES DOMAIN30 and ASL20/LBD18 regulate tracheary element differentiation in Arabidopsis. Plant Cell 20:3359–3373
Thatcher LF, Powell JJ, Aitken EA, Kazan K, Manners JM (2012a) The lateral organ boundaries domain transcription factor LBD20 functions in Fusarium wilt susceptibility and jasmonate signaling in Arabidopsis. Plant Physiol 160:407–418
Thatcher LT, Kazan K, Manners JM (2012b) Lateral organ boundaries domain transcription factors: new roles in plant defense. Plant Signal Behav 7:1702–1704
Wan CY, Wilkins TA (1994) A modified hot borate method significantly enhances the yield of high-quality RNA from cotton (Gossypium hirsutum L.). Anal Biochem 223:7–12
Wang A, Tan DM, Takahashi A, Li TZ, Harada T (2007) MdERFs, two ethylene-response factors involved in apple fruit ripening. J Exp Bot 58:3743–3748
Wang XF, Zhang SZ, Su L, Liu X, Hao YJ (2013) A genome-wide analysis of the LBD (LATERAL ORGAN BOUNDARIES domain) gene family in Malus domestica with a functional characterization of MdLBD11. PLoS One 8:e57044
Xiao YY, Chen JY, Kuang JF, Shan W, Xie H, Jiang YM, Lu WJ (2013) Banana ethylene response factors are involved in fruit ripening through their interactions with ethylene biosynthesis genes. J Exp Bot 64:2499–2510
Yang Y, Yu X, Wu P (2006) Comparison and evolution analysis of two rice subspecies LATERAL ORGAN BOUNDARIES domain gene family and their evolutionary characterization from Arabidopsis. Mol Phylogenet Evol 39:248–262
Yin XR, Allan AC, Chen KS, Ferguson IB (2010) Kiwifruit EIL and ERF genes involved in regulating fruit ripening. Plant Physiol 153:1280–1292
Yordanov YS, Regan S, Busov V (2010) Members of the LATERAL ORGAN BOUNDARIES DOMAIN transcription factor family are involved in the regulation of secondary growth in Populus. Plant Cell 22:3662–3677
Zhu QH, Guo AY, Gao G, Zhong YF, Xu M, Huang M, Luo J (2007) DPTF: a database of poplar transcription factors. Bioinformatics 23:1307–1308
Acknowledgments
We thank Professor Seiichiro Hasezawa (Department of Integrated Biosciences, the University of Tokyo), Professor Shouyi Chen (Institute of Genetics and Developmental Biology, Chinese Academy of Sciences), and Professor Junping Gao (Department of Ornamental Horticulture, China Agricultural University) for the generous gift of tobacco BY-2 suspension cells and the transient expression vectors, respectively. This work was supported in part by the National Natural Science Foundation of China (grant no. 31372111), the China Agriculture Research System (grant no. CARS-32-09) and Guangdong Modern Agricultural Industry Technology System (grant no. LNSG2011-12).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ba, Lj., Shan, W., Kuang, Jf. et al. The Banana MaLBD (LATERAL ORGAN BOUNDARIES DOMAIN) Transcription Factors Regulate EXPANSIN Expression and Are Involved in Fruit Ripening. Plant Mol Biol Rep 32, 1103–1113 (2014). https://doi.org/10.1007/s11105-014-0720-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-014-0720-6