Abstract
Key message
Four MaHDZs are possibly involved in banana fruit ripening by activating the transcription of genes related to ethylene biosynthesis and cell wall degradation, such as MaACO5, MaEXP2, MaEXPA10, MaPG4 and MaPL4.
Abstract
The homeodomain-leucine zipper (HD-ZIP) proteins represent plant-specific transcription factors, which contribute to various plant physiological processes. However, little information is available regarding the association of HD-ZIPs with banana fruit ripening. In this study, we identified a total of 96 HD-ZIP genes in banana genome, which were divided into four different groups consisting of 35, 31, 9 and 21 members in the I, II, III and IV subfamilies, respectively. The expression patterns of MaHDZ genes during fruit ripening showed that MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 were significantly up-regulated in the ripening stage and thus suggested to be potential regulators of banana fruit ripening. Furthermore, MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 were found to localize exclusively in the nucleus and exhibit transcriptional activation capacities. Importantly, MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 stimulated the transcription of several ripening-related genes including MaACO5 related to ethylene biosynthesis, MaEXP2, MaEXPA10, MaPG4 and MaPL4 were associated with cell wall degradation, through directly binding to their promoters. Taken together, our findings expand the functions of HD-ZIP transcription factors and identify four MaHDZs likely involved in regulating banana fruit ripening by activating the expression of genes related to ethylene biosynthesis and cell wall modification, which may have potential application in banana molecular breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Banana is the fourth major food crop in the world after rice, wheat and maize, and it is an important source of carbohydrates and valuable bioactive compounds for human beings (Ba et al. 2014; Han et al. 2016a, b; Kuang et al. 2017). Bananas are typical climacteric fruit, the ripening of which is accompanied with increased respiration, ethylene release, starch and cell wall degradation. These attributes cause rapid fruit softening, thereby constraining the long-term storage and transportation of the fruit (Kuang et al. 2013; Han et al. 2016a, b; Fan et al. 2018a; Guo et al. 2019). Therefore, a better understanding of the mechanism underlying banana fruit ripening is critical for improving the technology to maintain quality and extend fruit shelf life.
Control of transcription is an essential machinery in various plant physiological events, such as plant growth and development, abiotic and biotic stress responses and fruit ripening (Goel et al. 2016). Transcription is a highly complex process controlled in large part by transcription factors (TFs), a kind of proteins that determine when and how the genes are expressed on the transcriptional level. To date, a number of TFs have been found in plants, such as MYB, bHLH, bZIP, WRKY and HD-ZIP (Lehti-Shiu et al. 2017). The HD-ZIP family is a unique TF in plant. According to the evolutionary relationship and gene structure, HD-ZIP TFs are generally divided into four subfamilies, namely HD-ZIP I, HD-ZIP II, HD-ZIP III and HD-ZIP IV (Ariel et al. 2007). A number of studies have implicated that various subfamilies of HD-ZIP proteins play diverse roles in multiple cellular and developmental processes and stress responses (Schena et al. 1993; Sessa et al. 1998; Henriksson et al. 2005; Bang et al. 2018). For instance, in the late stage of seed germination, HD-ZIP I and HD-ZIP II proteins can be stimulated by far red light, thereby leading to shade avoidance response (Ciarbelli et al. 2008; Stamm and Kumar 2010). Moreover, HD-ZIP I and HD-ZIP II proteins are found to respond to various biotic and abiotic stresses, through controlling hormone signaling pathways and expression of related genes (Agalou et al. 2008; Manavella et al. 2008; Brinker et al. 2010; Harris et al. 2011). The HD-ZIP III subfamily is mainly responsible for regulating the differentiation of plant cells, apical meristems, embryonic development and formation of plant vascular system (Green et al. 2005; Prigge et al. 2005; Javelle et al. 2011). In addition, HD-ZIP IV proteins mediate the epidermal cell differentiation and root formation (Kamata et al. 2013; Yan et al. 2018). It is worth noting that some members of HD-ZIPs are involved in the regulation of fruit maturation and ripening. Tomato LeHB-1, a member of the HD-ZIP I subfamily, specifically recognizes the promoter region of ethylene biosynthetic gene LeACO1, and silencing LeHB-1 gene leads to a significant decreased expression of LeACO1 and the delay of fruit ripening (Lin et al. 2008). However, information concerning the possible roles of HD-ZIPs in fruit ripening has not been entirely understood.
The completed sequencing project of banana genome (D’Hont et al. 2012) provides a useful platform for investigating banana functional genomics and identifying potential candidate functional genes (Martin et al. 2016). Until now, several TF families have been characterized from banana, such as WRKYs (Goel et al. 2016), bZIPs (Hu et al. 2016), MADS-boxes (Liu et al. 2017) and IV subfamily of HD-ZIPs (Pandey et al. 2016). These findings provide detailed information on morphological and physiological diversity underlying banana fruit ripening. In this study, four HD-ZIP members, namely MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7, were identified to be markedly up-regulated during banana fruit ripening. The subcellular localization, transactivation capacities and potential target genes of these four MaHDZs were investigated. Our findings provide novel insight into deciphering the mechanism of HD-ZIPs involved in regulating banana fruit ripening.
Materials and methods
Plant materials and treatments
Pre-climacteric banana (Musa acuminata) fruits at 70–80% maturity were harvested from a local commercial plantation near Guangzhou, China. Harvested fruits were separated into single fruit, and fruits with uniform size, maturity and absence of mechanical injury, pests and diseases were selected. Fruits were sterilized with 500 mg/L prochloraz for 2 min, and then air-dried. Three different ripening treatments, including the control (natural ripening), ethylene-induced ripening (100 μL/L ethylene, 18 h), 1-MCP-delayed ripening (0.5 μL/L 1-MCP, 18 h), were performed according to our previous studies (Kuang et al. 2017; Xiao et al. 2018). After each treatment, fruits were stored at 22 °C and 90% relative humidity until completely ripe. For each treatment, samples were taken based on the rate of ethylene production and decline of fruit firmness during ripening. Samples of natural ripening were taken at 0, 1, 3, 5, 7, 15, 18, 20 and 23 days of storage. As for ethylene-induced ripening, samples were obtained at 0, 1, 3, 5, and 7 days of storage. In the case of 1-MCP-delayed ripening, samples were taken at 0, 1, 3, 5, 7, 15, 18, 20, 23, 26, 30, 35 and 40 days of storage. Fruit pulps of all samples were frozen in liquid nitrogen immediately after sampling and stored at − 80 °C for further use. All assessments were conducted using three biological replicates.
Identification and phylogenetic analysis of MaHDZs in banana
Whole banana (Musa accuminata) sequences were available from the banana genome database (https://banana-genome-hub.southgreen.fr/), and Arabidopsis thaliana HD-ZIP sequences was obtained from the TAIR (http://www.arabidopsis.org/) database. HD-ZIPs were identified from the banana genome through ‘blast comparing two seq’ in TBtools (Chen et al. 2018). A conserved domain search of the potential banana HD-ZIPs was further performed using CDD (http://www.ncbi.nlm.nih.gov/cdd/) database. Phylogenetic analysis of HD-ZIPs amino acid sequences from banana and Arabidopsis thaliana were carried out using MEGA X software with bootstrap values for 1000 replicates.
Protein properties analysis
Molecular weight (Mw) and isoelectric point (pI) of MaHDZ proteins were predicted using the ExPASy database (https://web.expasy.org/compute_pi/). Subcellular localization of MaHDZ proteins was predicted using the Wolfpsort database (https://wolfpsort.hgc.jp/).
Gene expression analysis and cloning
Total RNA was extracted using the hot borate method (Wan and Wilkins 1994). The extracted RNA was used to synthesize cDNA by PrimeScript™ RT reagent kit with gDNA Eraser (TaKaRa, Shiga, Japan). Quantitative real-time PCR (qRT-PCR) was carried out on a Bio-Rad CFX96 Real-Time PCR System (Bio-Rad, Hercules, CA, USA) using the Hieff® qPCR SYBR® Green Master Mix (Yeasen). The expression levels of target genes were normalized according to the cycle threshold (Ct) value using MaRPS4 (ribosomal protein 4) as the reference gene (Chen et al. 2011). Based on the sequence information of the NCBI (https://www.ncbi.nlm.nih.gov/) and the banana genome database (https://banana-genome-hub.southgreen.fr/), the coding regions of MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 were cloned using gene-specific primers.
Subcellular localization analysis
The coding regions of MaHDZs without the stop codon were amplified and cloned into the pEAQ vector (Sainsbury, et al. 2009) containing the gene in fusion with green fluorescent protein (GFP). The fusion constructs, GFP vector and NLS-mCherry were electroporated into Agrobacterium tumefaciens strain EHA105. GFP vector was used as a control, and NLS-mCherry pEAQ served as an indication of nucleus. Then, the above gene constructs or the control vector was transiently transformed into the leaves of 3-week-old tobacco (Nicotiana benthamiana) plants by Agrobacterium tumefaciens infiltration, respectively, as described previously (Tan et al. 2018; Xiao et al. 2018). After 36–48 h of infiltration, infected leaf tissues were analyzed for GFP signal using a fluorescence microscope (Zeiss Axioskop 2 Plus). All transient expression assays were repeated at least three times with similar results.
Transcriptional activation analysis in yeast cells
The coding regions of MaHDZs were cloned into the pGBKT7 vector (Clontech, Mountain View, CA, USA) containing a GAL4 DNA-binding domain (DBD) to create the pGBKT7-MaHD-ZIPs fusion constructs. Then, pGBKT7-MaHD-ZIPs, the positive control pGBKT7-53 + pGADT7-T-antigen and the negative control pGBKT7 were separately transformed into yeast strain GoldY2H using the lithium acetate method according to the protocols provided by the manufacturer (Clontech). The transformed yeast cells were streaked onto plates with minimal medium without tryptophan (SD/-Trp) or tryptophan, histidine and adenine (SD/-Trp-His-Ade), and the transactivation activity of each MaHD-ZIP protein was evaluated according to their growth status and the activity of x-α-galactosidase (X-α-Gal).
Dual-luciferase transient expression assay
The transcriptional activities of MaHDZs were assayed using the dual-luciferase transient expression system in tobacco leaves as described previously (Fan et al. 2018b; Tan et al. 2019). The reporter vector was modified from the pGreenII 0800-LUC vector. The firefly luciferase (LUC) was driven by the minimal TATA box of the CaMV35S promoter plus five copies of the GAL4 binding element (5 × GAL4), and the renilla luciferase (REN) driven by CaMV35S at the same vector was used as an internal control. The full-length coding sequences of MaHDZs were fused with the GAL4 DNA-binding domain (GAL4BD) under the control of CaMV35S promoter as effectors. To assess the transcriptional effects of MaHDZs on the promoters of MaACO5, MaEXP2, MaEXPA10, MaPG4 and MaPL4, these promoters were inserted into pGreenII 0800-LUC vector as reporters, while MaHDZs were cloned into the pGreenII 62-SK vector as effectors.
The constructed reporter and effector plasmids were transiently co-expressed in tobacco leaves as described above. After 48 h of infiltration, LUC and REN activities were measured by dual-luciferase assay kit (Promega, Madison, WI, USA). The transcriptional activities of MaHDZs were indicated by the ratio of LUC to REN. At least six transient assay measurements were included for each pair.
Protein expression and electrophoretic mobility shift assay (EMSA)
MaHDZs were cloned into pGEX-4T-1 to fuse in frame with GST and transformed into Escherichia coli strain BM Rosetta (DE3), respectively. The recombinant proteins were induced by 1 mM isopropyl thio-β-d-galactoside (IPTG) for 6 h at 28 °C, and purified with Glutathione-Superflow Resin (Clontech) according to manufacturer’s protocols. The fragments of ~ 50 bp covering the putative HD-ZIP binding region derived from the promoters of MaACO5, MaEXP2, MaEXPA10, MaPG4 and MaPL4 were biotin labeled using the Biotin 3′ End DNA Labeling Kit (Thermo Scientific, Rockford, IL, USA). Then the EMSA was performed using the LightShift Chemiluminescent EMSA kit (Thermo) following the manufacturer’s guidelines. As for competition, 800-fold molar excess of unlabeled DNA fragments with the same or mutant sequences were used as competitors, respectively. The GST protein alone was used as the negative control. After electrophoresis separation, protein–DNA complexes were transferred to the nylon membrane and detected using the chemiluminescence method and photos were taken on a ChemiDoc MP Imaging System (Bio-Rad).
Statistical analysis
Data are presented as mean ± standard errors (SE) of three or six independent biological replicates. Figures were made by TBtools, Weblogos and Origin 2017. Statistical differences between samples were analyzed by Student’s t test (P < 0.05 or 0.01).
Primers
The sequences of all primers used in this study are listed in Supplementary Table S1.
Results
Changes in ethylene production and firmness in fruit with three different ripening characteristics
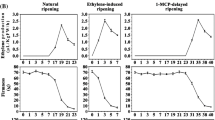
The ethylene production and fruit firmness during banana fruit ripening with three different characteristics including natural, ethylene-induced and 1-MCP-delayed ripening (Fig. 1a) were measured. As shown in Fig. 1b, ethylene production in natural ripening fruits increased significantly after 15 days of storage, reached a maximum on 18 days and decreased thereafter. The fruit firmness began to decrease during the 15 days of storage and reached a minimum on 23 days. Ethylene treatment accelerated the ripening of banana fruit, as evidenced by the ethylene peak and the sharp decrease in fruit firmness in 3 days. In contrast, 1-MCP treatment suppressed ethylene production and delayed fruit ripening, with ethylene peak and firmness loss appearing in 30 days (Fig. 1b).
Identification and classification of banana MaHDZs
A total of 96 MaHD-ZIP genes were identified as members of the HD-ZIP gene family in banana genome through the simultaneous consideration of the conservation of HD-ZIP domains from the Conserved Domain (CD) search. In accordance with the previous study that identified IV subfamily of HD-ZIPs in banana (Pandey et al. 2016), HD-ZIP genes of banana were termed as MaHDZs.
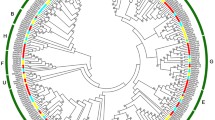
To explore the evolutionary and the classification characteristics of these banana MaHDZs, an unrooted phylogenetic tree was constructed in MEGA X based on the protein sequences of banana and Arabidopsis HD-ZIPs (Fig. 2). According to the phylogenetic analysis, 96 members of banana HD-ZIPs were divided into four subfamilies, namely MaHDZI (homologous to AtHB1/3/5/6/7/12/13/16, AtHB20-23 and AtHB40/51/52/53/54), MaHDZII (homologous to AtHB2/4/17 and HAT1/2/3/9/14/22), MaHDZIII (homologous to AtHB8/9/14/15 and IFL1), and MaHDZIV (homologous to ATHB10, ATML1, ANL2, PDF2 and HDG1/2/3/4/5/6/7/8/9/10/11/12), which contain 35, 31, 9 and 21 members, respectively. Since MaHDZIV have been reported previously (Pandey et al. 2016), we focused on the other 75 members of MaHDZI-III in this study. These 75 MaHDZI-III genes were named as MaHDZI.1-35, MaHDZII.1-31 and MaHDZIII.1-9 according to their orders on the chromosomes and typical characteristics (Supplementary Table S2).
Phylogenetic tree of HD-ZIP proteins. A phylogenetic tree was constructed based on a multiple alignment of predicted full-length banana (Musa accuminata) MaHDZs and Arabidopsis thaliana AtHD-ZIPs protein sequences. A total of 96 MaHDZ genes were identified as members of the HD-ZIP gene family in banana and divided into four subfamilies. MaHDZI, MaHDZII, MaHDZIII and MaHDZIV are represented by green hollow circles, purple solid circles, blue hollow triangles and orange solid triangles, respectively. Lighter colors represent AtHD-ZIPs
The 75 identified full-length MaHDZs sequences vary in length from 456 to 2577 bp, which encode proteins ranging from 154 to 858 aa, with molecular weight and isoelectric points ranging from 18.28 to 93.75 kDa and 4.59 to 9.22, respectively. In addition, most of the MaHDZI and MaHDZII proteins are predicted to locate in the nucleus, while MaHDZIII proteins are likely to locate in the nucleus, cytoplasm, chloroplast and mitochondrion (Supplementary Table S2). These results indicate that different MaHDZ proteins might function differentially based on their evolution relationship.
Expression profiles of MaHDZs under three different ripening behaviors in bananas
On the basis of our RNA-seq database of banana fruit ripening (Kuang et al. 2017), among 75 MaHDZI-III genes, 13 MaHDZs were up-regulated in the course of banana fruit ripening. Their expression profiles in banana fruit under three different ripening behaviors were further verified by qRT-PCR. As shown in Fig. 3, particularly, eight MaHDZs such as MaHDZI.19, MaHDZI.20, MaHDZI.26, MaHDZII.4, MaHDZII.7, MaHDZII.17, MaHDZII.22 and MaHDZIII.2 were most obviously elevated following the ripening, which parallel with the decline in fruit firmness and increase in ethylene production (Fig. 1). As the full lengths of only four MaHDZs (MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7) were cloned successfully, these four MaHDZs were focused on for further investigation.
Hierarchical clustering of the transcript accumulation profiles of 13 MaHDZ genes in banana fruit pulp with three different ripening characteristics, including natural (control), ethylene-induced and 1-MCP-delayed ripening. Three biological replicates were included in each sample. Differences in gene expression changes are indicated by the color scale bar
MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 are nucleus-localized transcription activators
To investigate the subcellular localizations of MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7, their full-length coding sequences were fused in frame with GFP and transiently expressed in tobacco leaves, respectively. As shown in Fig. 4a, the fluorescence emitted from these proteins was localized exclusively in the nucleus of tobacco leaf epidermal cells, while the control GFP fluorescence was observed throughout the cytoplasm and the nucleus.
Sub-cellular localization and transcriptional activation of MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7. a The fusion protein (MaHDZs-GFP) and control (GFP) were transiently expressed in Nicotiana benthamiana leaves by Agrobacterium tumefaciens strain EHA105, respectively. GFP fluorescence was observed with a fluorescence microscope. Images were taken in a fluorescence field for GFP and mCherry, while the outline of the cell and the merged were photographed in a bright field. NLS-mCherry was included in each transfection to serve as a control for nuclear localization. Bars, 25 μm. b Transcriptional activation of MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 in yeast cells. The coding region of MaHDZs was inserted into the pGBKT7 (GAL4DBD) to create the pGBKT7-MaHDZs construct. The yeast cells of strain Y2H Gold harboring the pGBKT7-MaHDZs plasmids were grown on SD plates without tryptophan (Trp−) or without tryptophan, histidine and adenine (Trp−His−Ade−) for 3 days at 28 °C, followed by the α-galactosidase assay (α-Gal staining). pGBKT7 and pGBKT7-53 + pGADT7-T were used as negative and positive control, respectively. c Diagrams of the reporter and effector vectors. d Transcriptional activation of MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 in Nicotiana benthamiana leaves. The transactivation ability of MaHDZs was demonstrated by the ratio of luciferase (LUC) to renilla luciferase (REN). The LUC/REN ratio of the empty BD-62SK vector was used as a calibrator (set as 1). BD-62SK-VP16 was used as a positive control. Data are mean ± SE of six independent biological replicates. Asterisks represent significant difference at 0.01 level by Student’s t test, compared to BD-62SK
Then we studied the transcriptional activation abilities of MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 using the GAL4BD system in yeast cells. The full-length coding regions of MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 were fused to the GAL4BD to generate BD-MaHDZI.19, BD-MaHDZI.26, BD-MaHDZII.4 and BD-MaHDZII.7 fusion plasmids, respectively. Then the plasmids were transformed into yeast cells GoldY2H to determine the abilities of the transformants to activate transcription from the upstream activation sequence of GAL4 and to promote yeast growth in medium lacking Trp, His and Ade (SD/− Trp − His − Ade). The transformants containing pGBKT7-53 + pGADT7-T and pGBKT7 vectors (empty) were used as positive and negative controls, respectively. As shown in Fig. 4b, the transformed yeast cells containing BD-MaHDZI.19, BD-MaHDZI.26, BD-MaHDZII.4 and BD-MaHDZII.7 and pGBKT753 + pGADT7-T (positive control) grew well in SD/− Trp − His − Ade and showed α-Gal activity, whereas cells containing pGBKT7 alone (negative control) did not, suggesting that these four MaHDZs behave as transcriptional activators in gene regulation.
The transcriptional activation activities of MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 in vivo were confirmed using the dual-luciferase reporter system (Fig. 4c). Compared with the negative control BD-62SK, expression of the positive control BD-62SK-VP16 as well as BD-62SK-MaHDZI.19, BD-62SK-MaHDZI.26, BD-62SK-MaHDZII.4 and BD-62SK-MaHDZII.7 resulted in higher values of LUC/REN ratio (Fig. 4d), which is consistent with the results found in yeast cells. Collectively, these data indicate that MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 are nucleus-localized transcriptional activators.
MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 stimulate the transcription of several ripening-associated genes by binding to their promoters
It has been reported previously that HD-ZIP I and HD-ZIP II bind preferentially to CAAT(A/T)ATTG and CAAT(C/G)ATTG elements in the promoters of target genes, respectively (Ariel et al. 2007). To investigate the potential targets of MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7, we searched CAAT(A/T)ATTG and CAAT(C/G)ATTG elements in the promoters of ripening-associated genes related to ethylene biosynthesis and cell wall degradation (Asif et al. 2014; Kuang et al. 2017). Five ripening-associated genes such as ethylene biosynthetic gene MaACO5, and cell wall degradation-related genes MaEXP2, MaEXPA10, MaPG4 and MaPL4, contain at least one CAAT(A/T)ATTG and CAAT(C/G)ATTG element in their promoters (Supplementary Text S1). The transcription levels of these selected five genes were increased at the ripening stage (Supplementary Fig. S1).
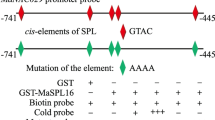
EMSAs were carried out to detect whether MaHDZI.19/26 and MaHDZII.4/7 can bind to the promoters of MaACO5, MaEXP2, MaEXPA10, MaPG4 and MaPL4. The recombinant GST-MaHDZI.19, -MaHDZI.26, -MaHDZII.4 and -MaHDZII.7 proteins were successfully purified (Supplementary Fig. S2). As shown in Fig. 5, the recombinant GST-MaHDZI.19/26 proteins could bind to CAAT(A/T)ATTG cis-acting elements present in the MaPL4 promoter, while GST-MaHDZII.4/I7 proteins could bind to CAAT(C/G)ATTG elements present in the promoters of MaACO5, MaEXP2, MaEXPA10, MaPG4 and MaPL4. When the corresponding unlabeled probes were added as cold competitors, the formation of the DNA–protein complex was effectively eliminated. However, it was not abolished when mutated probes were used in the assay (Fig. 5).
EMSA showing the binding of MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 to the promoter of ripening-associated genes. Purified GST-MaHDZ protein was incubated with the biotin-labeled wild-type probe containing CAAT (A/T)ATTG or CAAT (C/G)ATTG elements in MaACO5, MaEXP2, MaEXPA10, MaPG4 and MaPL4, respectively. The DNA–protein complexes were separated on native polyacrylamide gels. Sequences of both the wild-type and mutant probes are shown at the top of the image (wild-type and mutant CAAT (A/T)ATTG or CAAT (C/G)ATTG elements are marked with red letters). The mutant probe was used to test binding specificity. Shifted bands, suggesting the formation of DNA–protein complexes, are indicated by arrows. ‘‘−’’ represents absence, ‘‘+’’ represents presence. Competition experiments were carried out by adding 800-fold molar excess of unlabeled cold probe or mutant probe
To assess whether MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 act as transcriptional activators of MaACO5, MaEXP2, MaEXPA10, MaPG4 and MaPL4, transient dual-luciferase assays in tobacco leaves were performed. As shown in Fig. 6, transient overexpression of MaHDZI.19/26 stimulated the transcription of MaPL4, and MaHDZII.4/7 induced the transcription of MaACO5, MaEXP2, MaEXPA10, MaPG4 and MaPL4, as the higher value of the LUC/REN ratio was observed, compared to the empty control (pGreenII 62-SK). These data reveal that MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 act as positive regulators of MaACO5, MaEXP2, MaEXPA10, MaPG4 and MaPL4 through directly targeting CAAT(A/T)ATTG and CAAT(C/G)ATTG cis-elements on their promoters, respectively.
MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 activate the transcription of ripening-associated genes including MaACO5, MaEXP2, MaEXPA10, MaPG4 and MaPL4 by dual-luciferase transient expression assay in Nicotiana benthamiana leaves. a Diagrams of the reporter and effector vectors. b MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 activate the transcription of ripening-associated genes. The ratio of LUC/REN of the empty vector (62SK) plus promoter was used as a calibrator (set as 1). Data are mean ± SE of six independent biological replicates. Asterisks indicate significant differences by Student’s t test (**P < 0.01)
Discussion
The HD-ZIP family is one of the unique transcriptional regulators in plants, which are widely involved in diverse plant biological processes, such as plant growth and development, and stress responses (Ariel et al. 2010; Harris et al. 2011; Cabello et al. 2015; Bang et al. 2018; Zhu et al. 2018; Zhang, et al. 2019). In tomato, VAHOX1 (a member of HD-ZIP I subfamily) could be induced by low temperature treatment (Bartley and Ishida 2007). Sunlower HAHB4 positively regulates jasmomic acid and ethylene production, but negatively regulates ethylene sensitivity and salicylic acid accumulation during biotic stress responses and mechanical damage (Manavella et al. 2008). In Arabidopsis, overexpression of the AtHB8 gene leads to the early formation of xylem and cell lignification (Baima et al. 2001; Emery et al. 2003). In another case, Arabidopsis plants overexpressing the AtHB15 gene showed significant dwarfing of the plants, decreased lignification and uncurled leaves (Ohashiito and Fukuda 2003). Previously, the identification of HD-ZIP IV subfamily has been reported (Pandey et al. 2016), but the remaining subfamilies have not been unraveled yet. Due to the emergence of genome of banana (Musa acuminata) (Martin et al. 2016), identification of the whole HD-ZIP gene family in banana is feasible.
In this study, 96 MaHD-ZIP genes were found in banana genome. Phylogenetic analysis indicated that MaHD-ZIP proteins can be divided into four classes (Class I–IV) (Fig. 2). Different subfamilies and different species may indicate that banana HD-ZIPs may function diversely. Stated roughly, HD-ZIP I primarily responds to external signals, such as drought, extreme temperatures, and osmotic pressure in abiotic stresses, regulating plant growth to adapt to the environment (Cabello et al. 2015; Shao et al. 2018). HD-ZIP II is mainly involved in light stress response and auxin signal transduction (Turchi et al. 2013, 2015). HD-Zip III is involved in the regulation of plant growth and development, such as meristem formation, polar auxin transport and vascular system development (Hu et al. 2012; Franco et al. 2015; Turchi et al. 2015). HD-ZIP IV is mainly involved in root development, trichome formation and anthocyanin accumulation (Yan et al. 2018; Yang et al. 2018; Zhu et al. 2018).
Despite that HD-ZIP members have been widely studied in plant growth and development, little is known regarding their roles in banana fruit ripening. It is noteworthy that tomato LeHB-1 (HD-ZIP I subfamily) is involved in the control of fruit ripening (Lin et al. 2008). Interestingly, HD-ZIPs-mediated fruit ripening were different in peach and tomato (Gu et al. 2019). In peach, 11 HB genes in the HD-ZIP I subfamily were not differentially expressed between ripening and developing fruits, indicating that these genes were not associated with fruit ripening in peach. Instead, PpHB.G7 (HD-ZIP II subfamily) was found to mediate ethylene biosynthesis during fruit ripening (Gu et al. 2019). Recently litchi HD-ZIPs member LcHB2 and LcHB3 (HD-ZIP I subfamily) have been reported to be involved in fruitlet abscission by directly activating cell wall degradation-related genes LcCEL2/8 and ethylene biosynthetic genes LcACO2/3 and LcACS1/4/7, respectively (Li et al. 2019; Ma et al. 2019). In the present work, 13 MaHDZs were up-regulated in response to ethylene regulation in the course of banana fruit ripening (Fig. 3). We further demonstrated that four nucleus-localized transcriptional activators MaHDZs (MaHDZI.19/26 and MaHDZII.4/7) were directly bound to CAAT(A/T)ATTG and CAAT(C/G)ATTG elements on MaACO5, MaEXP2, MaEXPA10, MaPG4 and MaPL4 promoters, respectively, and activated their transcription (Figs. 4, 5 and 6). Our results, together with previous findings in tomato, peach and litchi, reveal that HD-ZIPs are involved in biological processes such as fruit ripening by modulating ethylene biosynthesis and cell wall degradation. It should be pointed out that further targeted transgenic research will contribute to the full clarification of MaHDZs involved in fruit ripening by regulating ethylene biosynthesis and cell wall modification.
In summary, four ripening-induced MaHDZ genes including MaHDZI.19, MaHDZI.26, MaHDZII.4 and MaHDZII.7 were identified. These four MaHDZs acted as nucleus-localized transcriptional activators of ripening-associated genes involved in ethylene biosynthesis and cell wall degradation. These findings expand the functions of HD-ZIP TFs and provide novel insight into deciphering the mechanism of HD-ZIPs involved in regulating banana fruit ripening.
Abbreviations
- 1-MCP:
-
1-Methylcyclopropene
- ACO:
-
1-Aminocyclopropane-1-carboxylate oxidase
- EMSA:
-
Electrophroretic mobility shift assay
- EXP:
-
Expansin
- GFP:
-
Green fluorescent protein
- HD-ZIP:
-
Homeodomain-leucine zipper
- IPTG:
-
Isopropyl thio-β-d-galactoside
- TFs:
-
Transcription factors
- M w :
-
Molecular weight
- PG:
-
Polygalacturonase
- pI:
-
Isoelectric point
- PL:
-
Pectinate lyases
- qRT-PCR:
-
Quantitative real-time polymerase chain reaction
- X-α-Gal:
-
X-α-galactosidase
- Y2H:
-
Yeast two-hybrid
References
Agalou A, Purwantomo S, Övernäs E et al (2008) A genome-wide survey of HD-Zip genes in rice and analysis of drought-responsive family members. Plant Mol Biol 66:87–103
Ariel FD, Manavella PA, Dezar CA, Chan RL (2007) The true story of the HD-Zip family. Trends Plant Sci 12:419–426
Ariel F, Diet A, Verdenaud M et al (2010) Environmental regulation of lateral root emergence in Medicago truncatula requires the HD-Zip I transcription factor HB1. Plant Cell 22:2171–2183
Asif MH, Lakhwani D, Pathak S et al (2014) Transcriptome analysis of ripe and unripe fruit tissue of banana identifies major metabolic networks involved in fruit ripening process. BMC Plant Biol 14:316
Ba LJ, Shan W, Xiao YY et al (2014) A ripening-induced transcription factor MaBSD1 interacts with promoters of MaEXP1/2 from banana fruit. Plant Cell Rep 33:1913–1920
Baima S, Possenti M, Matteucci A et al (2001) The arabidopsis ATHB-8 HD-zip protein acts as a differentiation-promoting transcription factor of the vascular meristems. Plant Physiol 126:643–655
Bang SW, Lee DK, Jung H et al (2018) Overexpression of OsTF1L, a rice HD-Zip transcription factor, promotes lignin biosynthesis and stomatal closure that improves drought tolerance. Plant Biotechnol J 17:118–131
Bartley GE, Ishida BK (2007) Ethylene-sensitive and insensitive regulation of transcription factor expression during in vitro tomato sepal ripening. J Exp Bot 58:2043–2051
Brinker M, Brosché M, Vinocur B et al (2010) Linking the salt transcriptome with physiological responses of a salt-resistant Populus species as a strategy to identify genes important for stress acclimation. Plant Physiol 154:1697–1709
Cabello JV, Arce AL, Chan RL (2015) The homologous HD-Zip I transcription factors HaHB1 and AtHB13 confer cold tolerance via the induction of pathogenesis-related and glucanase proteins. Plant J 69:141–153
Chen L, Zhong H, Kuang J et al (2011) Validation of reference genes for RT-qPCR studies of gene expression in banana fruit under different experimental conditions. Planta 234:377–390
Chen C, Chen H, He Y, Xia R (2018) TBtools, a Toolkit for Biologists integrating various HTS-data handling tools with a user-friendly interface. BioRxiv. https://doi.org/10.1101/289660
Ciarbelli AR, Ciolfi A, Salvucci S et al (2008) The Arabidopsis Homeodomain-leucine Zipper II gene family: diversity and redundancy. Plant Mol Biol 68:465–478
D’Hont A, Denoeud F, Aury JM et al (2012) The banana (Musa acuminata) genome and the evolution of monocotyledonous plants. Nature 488:213–217
Emery JF, Floyd SK, Alvarez J et al (2003) Radial patterning of arabidopsis shoots by class III HD-ZIP and KANADI genes. Curr Biol 13:1768–1774
Fan ZQ, Ba LJ, Shan W et al (2018a) A banana R2R3-MYB transcription factor MaMYB3 is involved in fruit ripening through modulation of starch degradation by repressing starch degradation-related genes and MabHLH6. Plant J 96:1191–1205
Fan ZQ, Tan XL, Shan W et al (2018b) Characterization of a transcriptional regulator, BrWRKY6, associated with gibberellin-suppressed leaf senescence of Chinese flowering cabbage. J Agric Food Chem 66:1791–1799
Franco DM, Silva EM, Saldanha LL et al (2015) Flavonoids modify root growth and modulate expression of SHORT-ROOT and HD-ZIP III. J Plant Physiol 188:89–95
Goel R, Pandey A, Trivedi PK, Asif MH (2016) Genome-wide analysis of the Musa WRKY gene family: evolution and differential expression during development and stress. Front Plant Sci 7:299
Green KA, Prigge MJ, Katzman RB, Clark SE (2005) CORONA, a member of the class III homeodomain leucine zipper gene family in arabidopsis, regulates stem cell specification and organogenesis. Plant Cell 17:691–704
Gu C, Guo ZH, Cheng HY et al (2019) A HD-ZIP II HOMEBOX transcription factor, PpHB. G7, mediates ethylene biosynthesis during fruit ripening in peach. Plant Sci 278:12–19
Guo Y, Shan W, Liang S et al (2019) MaBZR1/2 act as transcriptional repressors of ethylene biosynthetic genes in banana fruit. Physiol Plant 165:555–568
Han YC, Fu CC, Kuang JF et al (2016a) Two banana fruit ripening-related C2H2 zinc finger proteins are transcriptional repressors of ethylene biosynthetic genes. Postharvest Biol Technol 116:8–15
Han YC, Kuang JF, Chen JY et al (2016b) Banana transcription factor MaERF11 recruits histone deacetylase MaHDA1 and represses the expression of MaACO1 and expansins during fruit ripening. Plant Physiol 171:1070–1084
Harris JC, Hrmova M, Lopato S, Langridge P (2011) Modulation of plant growth by HD-Zip class I and II transcription factors in response to environmental stimuli. New Phytol 190:823–837
Henriksson E, Olsson AS, Johannesson H et al (2005) Homeodomain leucine zipper class I genes in Arabidopsis. Expression patterns and phylogenetic relationships. Plant Physiol 139:509–518
Hu R, Chi X, Chai G et al (2012) Genome-Wide identification, evolutionary expansion, and expression profile of homeodomain-leucine zipper gene family in poplar (Populus trichocarpa). PLoS One 7:e31149
Hu W, Wang L, Tie W et al (2016) Genome-wide analyses of the bZIP family reveal their involvement in the development, ripening and abiotic stress response in banana. Sci Rep 6:30203
Javelle M, Klein-Cosson C, Vernoud V et al (2011) Genome-wide characterization of the HD-ZIP IV transcription factor family in maize: preferential expression in the epidermis. Plant Physiol 157:790–803
Kamata N, Okada H, Komeda Y, Takahashi T (2013) Mutations in epidermis-specific HD-ZIP IV genes affect floral organ identity in Arabidopsis thaliana. Plant J 75:430–440
Kuang JF, Chen L, Shan W et al (2013) Molecular characterization of two banana ethylene signaling component MaEBFs during fruit ripening. Postharvest Biol Technol 85:94–101
Kuang JF, Chen JY, Liu XC et al (2017) The transcriptional regulatory network mediated by banana (Musa acuminata) dehydration-responsive element binding (MaDREB) transcription factors in fruit ripening. New Phytol 214:762–781
Lehti-Shiu MD, Panchy N, Wang P et al (2017) Diversity, expansion, and evolutionary novelty of plant DNA-binding transcription factor families. Biochim Biophys Acta Gene Regul Mech 1860:3–20
Li C, Zhao M, Ma X et al (2019) The HD-Zip transcription factor LcHB2 regulates litchi fruit abscission through the activation of two cellulase genes. J Exp Bot 70:5189–5203
Lin Z, Hong Y, Yin M et al (2008) A tomato HD-Zip homeobox protein, LeHB-1, plays an important role in floral organogenesis and ripening. Plant J 55:301–310
Liu J, Zhang J, Zhang J et al (2017) Genome-wide analysis of banana MADS-box family closely related to fruit development and ripening. Sci Rep 7:3467
Ma X, Li C, Huang X et al (2019) Involvement of HD-ZIP I transcription factors LcHB2 and LcHB3 in fruitlet abscission by promoting transcription of genes related to the biosynthesis of ethylene and ABA in litchi. Tree Physiol 39:1600–1613
Manavella PA, Dezar CA, Bonaventure G et al (2008) HAHB4, a sunflower HD-Zip protein, integrates signals from the jasmonic acid and ethylene pathways during wounding and biotic stress responses. Plant J 56:376–388
Martin G, Baurens FC, Droc G et al (2016) Improvement of the banana “Musa acuminata” reference sequence using NGS data and semi-automated bioinformatics methods. BMC Genom 17:243
Ohashiito K, Fukuda H (2003) HD-zip III homeobox genes that include a novel member, ZeHB-13 (Zinnia)/ATHB-15 (Arabidopsis), are involved in procambium and xylem cell differentiation. Plant Cell Physiol 44:1350–1358
Pandey A, Misra P, Alok A et al (2016) Genome-wide identification and expression analysis of homeodomain leucine zipper subfamily IV (HDZ IV) gene family from Musa accuminata. Front Plant Sci 7:20
Prigge MJ, Otsuga D, Alonso JM et al (2005) Class III homeodomain-leucine zipper gene family members have overlapping, antagonistic, and distinct roles in Arabidopsis development. Plant Cell 17:61–76
Sainsbury F, Thuenemann EC, Lomonossoff GP (2009) pEAQ: versatile expression vectors for easy and quick transient expression of heterologous proteins in plants. Plant Biotechnol J 7:682–693
Schena M, Lloyd AM, Davis RW (1993) The HAT4 gene of Arabidopsis encodes a developmental regulator. Genes Dev 7:367–379
Sessa G, Steindler C, Morelli G, Ruberti I (1998) The Arabidopsis Athb-8, -9 and genes are members of a small gene family coding for highly related HD-ZIP proteins. Plant Mol Biol 38:609–622
Shao J, Haider I, Xiong L et al (2018) Functional analysis of the HD-Zip transcription factor genes Oshox12 and Oshox14 in rice. PLoS One 13:e0199248
Stamm P, Kumar PP (2010) The phytohormone signal network regulating elongation growth during shade avoidance. J Exp Bot 61:2889–2903
Tan XL, Fan ZQ, Shan W et al (2018) Association of BrERF72 with methyl jasmonate-induced leaf senescence of Chinese flowering cabbage through activating JA biosynthesis-related genes. Hortic Res 5:22
Tan XL, Fan ZQ, Kuang JF et al (2019) Melatonin delays leaf senescence of Chinese flowering cabbage by suppressing ABFs-mediated abscisic acid biosynthesis and chlorophyll degradation. J Pineal Res 67:e12570
Turchi L, Carabelli M, Ruzza V et al (2013) Arabidopsis HD-Zip II transcription factors control apical embryo development and meristem function. Development 140:2118–2129
Turchi L, Baima S, Morelli G, Ruberti I (2015) Interplay of HD-Zip II and III transcription factors in auxin-regulated plant development. J Exp Bot 66:5043–5053
Wan CY, Wilkins TA (1994) A modified hot borate method significantly enhances the yield of high-quality RNA from cotton (Gossypium hirsutum L.). Anal Biochem 223:7–12
Xiao YY, Kuang JF, Qi XN et al (2018) A comprehensive investigation of starch degradation process and identification of a transcriptional activator MabHLH6 during banana fruit ripening. Plant Biotechnol J 16:151–164
Yan T, Li L, Xie L et al (2018) A novel HD-ZIP IV/MIXTA complex promotes glandular trichome initiation and cuticle development in Artemisia annua. New Phytol 218:567–578
Yang Q, Niu Q, Li J et al (2018) PpHB22, a member of HD-Zip proteins, activates PpDAM1 to regulate bud dormancy transition in ‘Suli’ pear (Pyrus pyrifolia White Pear Group). Plant physiol Bioch 127:355–365
Zhang L, Zhou D, Hu H et al (2019) Genome-wide characterization of a SRO gene family involved in response to biotic and abiotic stresses in banana (Musa spp.). BMC Plant Biol 19:211
Zhu H, Sun X, Zhang Q et al (2018) GLABROUS (CmGL) encodes a HD-ZIP IV transcription factor playing roles in multicellular trichome initiation in melon. Theor Appl Genet 131:569–579
Acknowledgements
We thank Dr. George P. Lomonossoff (John Innes Centre, Norwich Research Park) for the generous gift of pEAQ vectors. This work was financially supported by the grants from the Natural Science Foundation of China (Grant no. 31830071), and China Agriculture Research System (Grant no. CARS-31-11).
Author information
Authors and Affiliations
Contributions
YYY performed the experiments, interpreted the results and wrote the article; YYY and WS analyzed the data and performed statistical analyses; JFK and JYC contributed to experiment design and manuscript editing; WJL designed and coordinated the experiments, and revised and improved the manuscript; WJL is responsible for the manuscript as a whole.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Additional information
Communicated by Prakash Lakshmanan.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, YY., Shan, W., Kuang, JF. et al. Four HD-ZIPs are involved in banana fruit ripening by activating the transcription of ethylene biosynthetic and cell wall-modifying genes. Plant Cell Rep 39, 351–362 (2020). https://doi.org/10.1007/s00299-019-02495-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-019-02495-x