Abstract
To investigate the genetic basis of maize seedling response to waterlogging, we performed a genome-wide association study in 144 maize inbred lines, measuring length, fresh and dry weight of roots and shoots under normal and waterlogged conditions using 45,868 SNPs. This panel was divided into three subgroups based on the population structure results and the LD decay distance was 180 kb. A biparental advanced backcross (AB) population was also used to detect quantitative trait loci (QTL). In a comparison of 16 different models, principal components analysis (PCA/top PC3) + K was found to be best for reduction of false-positive associations for further analysis. A whole-genome scan detected four strong peak signals (P < 2.18 × 10−5) significantly associated with the waterlogging response on chromosomes 5, 6 and 9. SNP4784, SNP200, SNP298, and SNP6314 showed significant association with corresponding traits under waterlogging and explained 14.99–19.36 %, 15.75–17.64 %, 16.08 % and 15.44 % of the phenotypic variation, respectively. The identified SNPs were located in GRMZM2G012046, GRMZM2G009808, GRMZM2G137108 and GRMZM2G369629 (AGPV1). SNP4784 (GRMZM2G012046) was colocalized with the major QTL that was identified with the same traits in the AB population. Forty-seven SNPs significantly associated (P < 2.18 × 10−4) with six traits in association mapping were identified and, among these, 33 SNPs were already reported in literature as waterlogging-related traits. These results will help elucidate the genetic basis of differential responses and tolerance to waterlogging stress among maize inbred lines, and provide novel loci for improvement of waterlogging tolerance of maize inbred lines using marker-assisted selection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Waterlogging, which results in oxygen deprivation in the rhizosphere, is a serious abiotic stress in plants (Visser et al. 2003). In the life cycle of maize (Zea mays L.), the grain yield will be reduced in both natural and agricultural systems when seedlings encounter oxygen limitation frequently because of waterlogging. In general, the early seedling growth phase from the second leaf stage (V2) to the seventh leaf stage (V7) is sensitive to waterlogging (Zaidi et al. 2004; Liu et al. 2010), but substantial variation exists among maize genotypes in response to waterlogging stress (Mano et al. 2002; Zaidi et al. 2004; Liu et al. 2010). In recent years, genetic and molecular biological dissection has focused mainly on exploiting the genetic variability in waterlogging tolerance available in maize and its wild relative, Zea luxurians and Zea nicaraguensis. Mapping of quantitative trait loci (QTL) has revealed a number of chromosomal regions that affect important traits, such as root and shoot development-associated traits (Qiu et al. 2007), capacity for root aerenchyma formation (Mano et al. 2007, 2008, 2012; Mano and Omori 2008, 2009), adventitious root formation (Mano et al. 2005a, b, 2009), tolerance to toxins under reducing soil conditions and leaf injury (Mano et al. 2006) under waterlogged conditions in maize seedlings. QTL associated with waterlogging tolerance have also been studied in other crops, such as rice (Xu et al. 2006; Hattori et al. 2009), soybean (VanToai et al. 2001), wheat (Burgos et al. 2001), and barley (Li et al. 2008), and provide insights into mechanisms of crop survival under waterlogging stress that are potentially translatable to maize.
Successful map-based cloning revealed that major genes control tolerance of complete submergence (SUB1) and rapid outgrowth of adverse partial submergence (SK) in rice (Xu et al. 2006; Hattori et al. 2009). However, the waterlogging-tolerant mechanism in maize is potentially different from that in rice; a large number of minor QTL is considered responsible for waterlogging tolerance in maize, and only a few major QTLs (R 2 > 10 % often was adopted as major QTL) (Li et al. 2012) were associated with waterlogging tolerance, for example, Qarf7.04-5 (Mano et al. 2005c), Qaer1.06 (Mano and Omori 2009), Qaer1.06–1.07 (Mano et al. 2012) and sdw9-1, rdw9-1, tdw9-1, tdw9-2, tdw9-3, sdw9-4 (Qiu et al. 2007).
Association mapping is an approach to study the relationship between phenotypic variation and genetic polymorphisms on the basis of linkage disequilibrium (LD) (Zondervan and Cardon 2004), which is rapidly becoming the main strategy to dissect the genetic architecture of complex traits in plants with the advent of abundant single nucleotide polymorphisms (SNPs). Currently, most association mapping studies in plants are performed using sets of genes selected to be putative candidates for the trait of interest (Su et al. 2011; Sharma and Chauhan 2012), but rapid developments in genomics will allow for genome-wide association study (GWAS) in virtually any plant species in the near future (Rafalski 2010; Ingvarsson and Street 2011; Yan et al. 2011). The method has been applied in diverse plants, such as maize (Belo et al. 2008; Yan et al. 2010a; Yang et al. 2010; Kump et al. 2011; Tian et al. 2011), rice (Huang et al. 2010, 2012), wheat (Breseghello and Sorrells 2006) and Arabidopsis (Atwell et al. 2010). Excellent results have been achieved for complex traits, such as rice flowering time and grain yield traits (Huang et al. 2012), maize kernel composition (Cook et al. 2012), and resistance to northern (Wisser et al. 2011) and southern (Kump et al. 2011) leaf blight.
In the present study, the GWAS approach was used to detect QTL involved in the response to waterlogging during the maize seedling stage under normal and waterlogged conditions. The goals were (1) to identify the major putative alleles associated with root and shoot traits under normal and waterlogged conditions by GWAS and further predict the candidate genes for waterlogging tolerance; and (2) to compare SNP-trait associations with the results of advanced backcross-QTL (AB-QTL) mapping.
Materials and Methods
Maize Germplasm and Genotyping
A set of 144 maize inbred lines representing the genetic diversity among different heterotic groups in China were chosen for GWAS of the traits associated with waterlogging tolerance in the seedling stage (Table S1) (Wang et al. 2011). Total genomic DNA was bulked for extraction from the young leaves of six randomly selected 30-day-old seedlings per inbred line using a standard CTAB extraction method (Saghai-Maroof et al. 1984).
DNA of each line was quantified and genotyped using the Illumina MaizeSNP50 BeadChip following the manufacturer’s instructions (http://www.emtd.com.cn/index_en.asp). The MaizeSNP50 Genotyping BeadChip contains 56,110 SNPs spaced at an average distance of 40 kb. (http://www.illumina.com/products/maize_snp50_whole_genome_genotyping_kits.ilmn).
Illumina GenomeStudio genotyping software was used to assess gene clusters, which were rechecked manually, and heterozygous SNPs were attributed to missing data (Yan et al. 2010b). A total of 45,868 SNPs with a minor allele frequency (MAF) of more than 0.05 and missing data less than 20 % were confirmed for further association analysis. In total, 92 % (43,102/45,868) of oligomer sequences derived from the SNPs were mapped with a single megablast hit blasted against the maize sequence database AGPV1 (http://www.maizesequence.org), and were well distributed across the maize genome by in silico mapping (data not shown). The heterozygotes frequency of SNPs, gene diversity, polymorphism information content (PIC) were estimated using PowerMarker V3.25 (Liu and Muse 2005).
Development of the Backcross Population
Linkage analysis was conducted with 180 BC2F2 individuals developed from a cross between HZ32 (waterlogging tolerant) as the donor parent and K12 (waterlogging sensitive) as the recurrent parent.
Phenotypic Evaluation
Three pot experiments (EXP.1, EXP.2 and EXP.3) with the association panel were conducted in August, September and October 2009, respectively. A total of 180 BC2F2:3 families was phenotyped in a similar manner at Huazhong Agriculture University’s experimental station in 2010. The day/night temperatures were 36/11 °C and the photoperiod was 13/11 h (day/night). The growth substrate and waterlogging treatment followed the description by Qiu et al. (2007). Trials with the association panel and AB population were laid out in a randomized complete-block design with two replications and three replications, respectively. Six pots per genotype and five plants per pot were included in each replication, of which three pots were the control and three pots were subjected to waterlogging stress. The waterlogging treatment was applied at the seedling V2 stage and 15 seedlings per genotype were used for trait scoring under the control and waterlogged conditions after waterlogging treatment for 6 days. The treatment method, sampling, and measurement of root and shoot traits were performed in accordance with the methods of Qiu et al. (2007). Six traits, namely seedling height (SH), root length (RL), shoot fresh weight (SFW), root fresh weight (RFW), shoot dry weight (SDW) and root dry weight (RDW), were dissected under the control and waterlogged conditions.
Phenotypic Data Analysis
Statistical analysis for all seedling traits was performed with software SAS8.02 (SAS Institute, Cary, NC). Using the average value of 15 plants of each genotype per replicate across the three trials, the effects of genotype by experiment (G × E) and genotype by treatment (G × T) (i.e. , normal and waterlogged conditions) were evaluated by PROC GLM. The PROC MIXED procedure was used to calculate the adjusted mean values for each trait in different treatments (the normal and waterlogged conditions, respectively) of each inbred line using best linear unbiased predictors (BLUP), estimated from ANOVA of data sets for the three pot experiments with genotype, experiments, and G × E considered. Broad-sense heritabilities (h 2) of seedling traits under waterlogged and control conditions were calculated using the combined data from the three pot experiments with the following formula:
where \( \delta_{\mathrm{G}}^2 \) is the genotypic variance, \( \delta_{\mathrm{GE}}^2 \) is the genotype × experiments (GE) variance, \( \delta_{\mathrm{e}}^2 \) is the residual error variance, and n and r are the number of experiments and replications, respectively (Knapp et al. 1985). Phenotypic Pearson’s correlations among six seedling traits were estimated by PROC CORR, using the adjusted means of this maize panel across three experiments. Normality of distributions for each trait under different moisture regimes was evaluated with the Shapiro-Wilk test (Shapiro and Wilk 1965). To estimate genotypic performances, the adjusted means across the three experiments for the phenotypic trait of each inbred line under the control and waterlogged conditions were used as input values for GWAS.
Linkage Mapping
A linkage map consisting of 121 SSR markers was constructed by Kosambi’s mapping function with JoinMap 4.0 (Van Ooijen and Voorrips 2001). The AB-QTL analysis was conducted on arithmetic mean values of three replicates for each trait and each family by the inclusive composite interval mapping (ICIM) method described by Li et al. (2007). The logarithm of odds (LOD) score for declaring a QTL was 2.5 for each trait and the walking speed for all QTL was 1 cM.
Whole-Genome Scan Association Mapping
The software STRUCTURE (Pritchard et al. 2000), based on a Bayesian Markov chain Monte Carlo model, was used to estimate the population structure and assign inbred lines to subpopulations in accordance with the methods reported by Evanno et al. (2005), which took the parameter of 100,000 burn-in, 100,000 run length and three iterations for each K with 28,791 SNPs with MAF of more than 0.05 and missing data less than 10 %. Nei’s genetic distances among given subgroups and lines and a neighbor-joining tree were calculated with PowerMarker 3.25 (Liu and Muse 2005). To correct for population stratification, data from the 28,791 SNPs in the 144 inbred lines were subjected to principal component analysis (PCA), and distance matrices were used to obtain eigenvectors based on Nei’s genetic matrices, with the modules DCENTER and EIGEN implemented in NTSYSpc 2.1 (Rohlf 2000). To account for relatedness among individuals, 28,791 SNPs were used to assess the kinship matrix (K) between each pair of lines in this maize panel using the SPAGeDi software package (Hardy and Vekemans 2002).
The pairwise estimates LD (r 2) based on 28,791 SNPs were calculated using the software package Haploview 3.31 (Barrett et al. 2005). The physical distance of each SNP on a reference genome in the Maize B73 RefGen_v1 sequence, and the sequence and function prediction of candidate genes were obtained from MaizeGDB (http://www.maizegdb.org/). A schematic representation of LD decay was exhibited in accordance with the methods of Yan et al. (2009). To correct for false positives, 16 models were compared to identify the optimal model of association analysis for seedling traits under waterlogging, which referred to the population structure (Q, PCA) and kinship (K). The first six models were a simple model (GLM), Q model, PCA model, K model, Q + K model and PCA + K model, respectively (Yang et al. 2011; Zhang et al. 2010b). The other ten models were a PCA + K model that each incorporated one of the top ten components. Each SNP was fit as a fixed effect singly to test the association between the SNP and phenotype. The simple modes, Q and PCA models were applied using a general linear model (GLM) with TASSEL 3.0.50; the K, Q + K and PCA + K models were applied with a compressed mixed linear model (MLM) in TASSEL 3.0.50 (Yu et al. 2006; Zhang et al. 2010b). Correction for multiple comparisons followed Setter et al. (2011), i.e., a family-wise probability level of α < 0.10 was chosen, and a Bonferroni-corrected threshold probability based on individual tests (β) was calculated as β < 0.10/n, where n was the number of individual trait–SNP combinations tested. Because of SNP dependence, chromosome independence and empirical data (Lu et al. 2010; Setter et al. 2011), the cutoff value is important and an alternative cutoff 10/n and 1/n was adopted. Both the quantile-quantile plots and Manhattan plots were drawn with the gplot library using R version 2.12.0 (R Foundation for Statistical Computing, http://www.r-project.org/).
Results
Analysis of Phenotypes
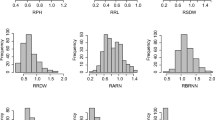
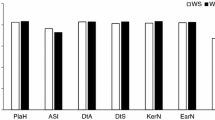
Phenotype data for the six traits of interest were collected from the association panel under normal and waterlogged conditions. The phenotypic frequency of the six traits fitted a normal distribution as determined by the Shapiro-Wilk test (Shapiro and Wilk 1965), which indicated that the traits were quantitative traits. A wide range of phenotypic variation was observed for all traits measured under normal and stress conditions. Most traits showed the lowest means in EXP.3 and the highest means in EXP.2 (Table 1). Generally, in all genotypes, the six traits were significantly reduced (5.38–41.67 % decrease) under waterlogged conditions relative to the control across the three experiments. Overall, RL, RFW and RDW showed a greater reduction than SH, SFW and SDW under waterlogging in the three experiments (Table 1).
ANOVA indicated that the mean values of all traits under waterlogged and control conditions were significantly different (P < 0.01) in all three experiments, and the G × E and G × T interactions were also significant (P < 0.001) for all traits (Table 1), which indicated that the genotypes responded differentially to experiments and treatments. The ANOVA indicated that most of the variation among the three experiments was explained by the waterlogging treatment, and the difference among the three experiments for all traits investigated was significant (P < 0.001); therefore, the maize association panel was tested across a broad range of growing conditions in the three experiments (data not shown). Broad-sense heritability (h 2) for the six traits was high under both moisture conditions, but differed between the two conditions. SH, RL, SFW, RFW, SDW and RDW had relatively higher h 2 (0.87, 0.85, 0.91, 0.92, 0.90 and 0.91, respectively) under the control condition and lower h 2 (0.85, 0.81, 0.90, 0.90, 0.87 and 0.88, respectively) under the waterlogged condition (Table 1). Compared with h 2 under the control condition, RL, RFW and RDW were indicated to be more prone to waterlogging stress than the other traits.
All seedling traits were significantly positively correlated (P < 0.001) under waterlogged and control conditions (Table 2). Furthermore, most correlation coefficients for pairs of measured traits were very similar between waterlogged and control conditions, but were usually lower under waterlogging stress than under the control condition. As expected, the phenotypic correlation coefficients were relatively higher between SDW and RDW, and between SFW and RFW than between one of these and other traits. However, RL showed relatively low correlation coefficients with the other five traits under both moisture conditions. For example, the correlation coefficients between RL and RDW were 0.52 and 0.58 under waterlogged and control conditions, respectively. This result indicated that RL expression showed greater independence of moisture regime than the other traits.
Genotypic Data
The frequency distribution of minor alleles from 56,110 SNPs was evaluated with 50 continuous classes from 0.01 to 0.50 with a similar number of SNPs in each MAF class (Fig. S1a). The mean value of MAFs from the 56,110 SNPs was 0.25, and about 50 % of the SNPs had a MAF exceeding 0.25. The average heterozygote frequency of 53,014 informative SNPs in the 144-strong maize panel is 0.037; 51,311 SNPs occurred in less than 0.1 % heterozygotes and 43,417 SNPs in less than 0.05 % heterozygotes. The average values of PIC and gene diversity were 0.271 (maximum 0.375) and 0.339 (maximum 0.500) of all informative SNPs, respectively. (Fig. S1b).
Linkage disequilibrium (LD) patterns and distributions in the panel of maize inbred lines determine the power and resolution of GWAS. To explore LD decay, the pairwise correlation coefficient (r 2) based on 28,791 SNPs with respect to the physical distance for each chromosome as well as all chromosomes is presented in Fig. S2. As expected, a rapid decline in the pairwise r 2 value was observed as the physical distance increased. For the maize panel, it was expected that LD would decay over relatively short distances because of its diversity and historical recombination. The rate of LD decay varied among different chromosomes. When the r 2 cut-off value was set to 0.1, the LD decay rate was estimated at 130–180 kb for every independent chromosome and 180 kb at the whole-genome level. This was a relatively long-range LD, which might be the result of the small population size and origin of the accessions. On the basis of the LD decay results, 45,868 SNPs were suitable for GWAS with the association panel.
Population Structure and Kinship
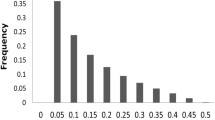
Population structure in the panel and a long-range LD represent major challenges in GWAS and could cause spurious associations. In other words, significant associations may not be the consequence of physical linkage between markers and causal genes and false positives could be avoided by accounting for population structure before GWAS. Therefore, first a panel of 28,791 SNP markers was selected to infer population substructure using STRUCTURE, and a Q matrix assigning individuals to subpopulations was created. The maize panel containing 144 inbred lines was divided into three subgroups based on the known pedigree, LnP(D) and delta k (Fig. S3a) (Pritchard et al. 2000; Evanno et al. 2005). Principal component analysis (PCA) is a method widely used to identify population substructure and to build the P matrix for population correction to remove stratification effects owing to adaptation among lines (Price et al. 2006), which is an alternative strategy to the STRUCTURE algorithm (Zhao et al. 2007). PCA separated the maize populations easily into three subpopulations by plotting the first two eigenvectors generated with SNPs (Fig. S3b). A neighbor-joining tree based on Nei’s genetic distance was constructed, and three subgroups similar to those obtained with STRUCTURE and PCA were revealed (Fig. S3c). The distribution of kinship coefficients between 0 and 0.45 represented 97.6 % of the data (Fig. S4). A total of 62.9 % of the pairwise kinship estimates were equal to 0, which indicated there was no relatedness between these pairs of lines, and the remaining estimates decreased continuously with the increase in number of categories. The kinship analysis indicated few lines showed strong similarities, which suggested that the simple relatedness among inbreds will decrease the frequency of spurious associations because of the effect of population structure (Yang et al. 2011). Thus the maize panel was suitable for GWAS.
Association Mapping
To reduce the frequency of spurious associations because of unrecognized population structure or kinship between pairs of lines in the association population, the GLM and compressed MLM procedures (Zhang et al. 2010b) with different parameters were applied in TASSEL, which required a population structure (Q matrix) and kinship matrix (K). To determine which of the simple, Q, PCA, K, Q + K and PCA + K models most efficiently reduced the number of spurious associations, quantile-quantile plots of estimated −log10(P) were calculated using all traits. The results indicated that the PCA + K model was optimal in terms of reduction of the false-positive frequency; because of the high correlation of different traits, and consistent trends in the P value of the six traits, the results for one trait (SFW) under the waterlogged condition for each model are shown in Fig. S5. GWAS using principal components may be sensitive to the number of principal components, while no significant differences in the reduction of the false-positive frequency for SFW under the waterlogged condition were observed when the number of principal components increased from one to ten with the PCA + K model (Fig. S6). Thus, the top three principal components with the lowest P value were selected to build the P matrix accounting for population structure correction. Overall, the compressed MLM approach, which took population structure (PCA) and kinship (K matrix) into account, was used to perform GWAS between 45,868 SNPs and the six physiological traits under the different moisture regimes in a panel of 144 inbred lines, as shown in Manhattan plots (Fig. 1).
Genome-wide association study of root and shoot traits under the control and waterlogged conditions. a Manhattan plots of the compressed mixed linear model (MLM) for all seedling traits under the two moisture regimes for the whole genome. The horizontal axis shows the single nucleotide polymorphism (SNP) physical location and the vertical axis shows negative log10-transformed P values from a genome-wide scan for each SNP tested. The blue horizontal line indicates a genome-wide significance threshold of 1/n (n = no. of SNP with association mapping). The different combinations of colors and forms show different traits as indicated. b Quantile-quantile plots of test statistics for SNP-trait associations using the compressed MLM for the whole genome. The horizontal axis shows the expected negative log10-transformed P values assuming no SNP-trait associations; the vertical axis is the observed negative log10-transformed P values indicating true marker associations
Out of 550,416 possible associations (45,868 SNP × six traits × two treatments), a total of 98 SNP-trait associations with P < 2.18 × 10−4 were identified, which comprised 47 unique SNPs (uni-SNPs: unique position) blasted against the maize sequence database AGPV1 (http://www.maizesequence.org), for which detailed information is shown in Table S2. Significant associated uni-SNPs were distributed on all chromosomes except chromosome 4. Of the uni-SNPs 70.12 % (33/47) were coincident with previously reported QTL associated with waterlogging-responsive traits in different biparental populations. We also identified 18 SNP-trait associations with P < 2.18 × 10−5, which comprised five uni-SNPs (Table 3). The SNP markers significantly associated with phenotype with P < 2.18 × 10−6 were SNP 200 (RFW, P = 2.18 × 10−6, R 2 = 18.28 %) and SNP 4784 (SFW, P = 1.39 × 10−6, R 2 = 19.36 %) under the control and waterlogged conditions, respectively (Table S2).
Because of the slow LD decay distance of 180 kb in the maize panel, the putative genes were predicted in a 400 kb region around each significant SNP with P < 2.18 × 10−6. SNP 200 was located within GRMZM2G009808, which encodes the aconitase/3-isopropylmalate dehydratase large subunit involved in energy metabolism on chromosome 9, in bin 9.07; the functional annotations of ten putative genes within a 400 kb region around SNP200 are shown in Table S3. SNP 4784 was located within GRMZM2G012046, which encoded the membrane attack complex component/perforin/complement C9 on chromosome 5, in bin 5.04; the function annotations of eight putative genes within a 400 kb window around SNP4784 are shown in Table S4.
Linkage Mapping
Linkage analysis of a biparental mapping population is extremely useful in identifying QTL that influence complex traits. In the present study, the linkage map had a total length of 1,286.9 cM with an average interval of 10.6 cM between adjacent markers; the details are shown in Table S5. AB-QTL mapping was used to test the results of association mapping. A list of the putative QTL flanked by SSR markers along with their LOD scores, phenotypic variance, and additive and dominance effects is presented in Table 4. A graphical presentation of QTL locations on the linkage map is shown in Fig. 2. QTL that control RDW, RFW, SDW and SFW were mapped to a similar region of chromosome 5; the marker umc1171 closest to the QTL peak was located at 168,868,089 bp against the maize sequence database AGPV1 (http://www.maizesequence.org), and was located extremely close to SNP 4784 by GWAS. Consistent results were obtained for both moisture regimes. Individual QTL accounted for 6.32–12.01 % of the phenotypic variation. The alleles from the QTL flanked by umc1171/umc1722 on chromosome 5, which contributed to an increase in the trait values, were from the waterlogging-tolerant parental genotype HZ32, which was in agreement with the genetic direction in association mapping. One QTL associated with RDW under the control condition was mapped between bnlg1525 and umc1714 on chromosome 9 in the BC2F2:3 families, which also partly overlapped with SNP 200 by GWAS. Individual QTLs accounted for 5.73 % of the phenotypic variation and alleles from the waterlogging-tolerant parental genotype HZ32 tended to increase the trait score, which was consistent with the genetic direction in association mapping. Two major peaks of association identified on chromosome 6 in bin 6.04 and bin 6.05 were not confirmed in the AB population.
Linkage map and chromosome locations of QTL for waterlogging response-related traits on chromosomes 5 and 9 in a BC2F2 population consisting of 180 BC2F2:3 families developed from a cross between two maize inbred lines (K12 and HZ32) using inclusive composite interval mapping (ICIM). LOD curves obtained by ICIM are shown in chromosome 5 and 9, where the significant SNP-trait associations with P < 2.18 × 10−5 were detected in association mapping. Bin numbers are in parentheses after marker names
Discussion
Maize seedlings prior to the sixth leaf stage (V6) are extremely susceptible to waterlogging because the shoot growing point is below the soil surface at that time (Nielsen 2004), and the most sensitive phase is V2 (Zaidi et al. 2004; Liu et al. 2010). The present results reveal substantial genetic variation in SH, RL, SFW, RFW, SDW and RDW existing in the maize inbred lines and BC2F2 population. As expected, in maize seedlings in response to waterlogging stress, the growth and development of root and shoot traits in most inbred lines and BC2F2:3 families were seriously inhibited, which is consistent with previous reports (Dennis et al. 2000; Qiu et al. 2007; Liu et al. 2010). However, shoots seemed to possess a different acclimation mechanism for more rapid growth under waterlogging stress than under control conditions for some inbred lines, which has also been reported in Arabidopsis (Ellis et al. 1999) and Zea nicaraguensis (Abiko et al. 2012). The significantly positive correlations between the waterlogged and control conditions for the traits investigated in the present study indicated that common genetic mechanisms exist for these traits.
The magnitude and distribution of LD determines the choice of association mapping methodology (Rafalski 2002). GWAS may help identify genomic regions that are associated with interesting traits, and the size of the genomic regions corresponds to LD decay distance. Rapid LD decay in a given association mapping panel implies that a higher density of markers is needed in GWAS to detect significant marker–trait associations. In the present study, the decrease in r 2 to 0.1 within 180 kb indicated it was possible to achieve resolution down to ∼180 kb in GWAS. On the basis of previous studies (Purcell et al. 2003; Belo et al. 2008; Yan et al. 2010a; Yang et al. 2010; Setter et al. 2011), the 2,300 Mb complete maize genome (Schnable et al. 2009) requires 23,000 or 11,500 markers with an average gap of 100 kb or 200 kb assuming that two selected SNPs can represent the diversity of one gene. The relatively high LD observed in the maize panel contributed to the feasibility of covering the whole genome with 45,868 SNPs. The number of markers ideally needed to cover the entire maize genome was only for reference because previous studies showed that LD levels varied both between the different loci Y1 and PSY2 in the same population (Palaisa et al. 2003) and within the same locus Y1 in different populations (Palaisa et al. 2003, 2004). For example, in previous studies of maize, the LD decay ranges from less than 1 kb (Tenaillon et al. 2001; Su et al. 2011) or roughly 2–5 kb (Remington et al. 2001; Yan et al. 2009) to as high as 100 kb (Ching et al. 2002). Comparing with the results of a simulation of QTL detection power (Yan et al. 2011), the panel in the present study provided 10–50 % detection power for QTL that contributed 0.5–5.0 % of the total phenotypic variation of the target traits. Marker density, however, is another determinant for increasing the detection power of association analysis (Mackay 2009), especially for GWAS. In addition, a high density of markers will lead to finer resolution QTL mapping. Thus, the population size and marker density of the maize panel in the present study were suitable for investigation of the genetic basis of traits controlled by waterlogging-tolerance genes (Yu et al. 2006; Yan et al. 2011).
The detection of false-positive associations caused by population substructure identified within a germplasm group and multiple testing across the entire genome is one of the main limitations to the use of GWAS (Aranzana et al. 2005). Several statistical methods have been proposed to account for population structure and familial relatedness among individuals in association panels, including genomic control (Devlin and Roeder 1999), structured association (Pritchard et al. 2000; Falush et al. 2003), the unified mixed model approach (Yu et al. 2006), principal component approach (Patterson et al. 2006; Price et al. 2006), and nonmetric multidimensional scaling (Zhu and Yu 2009). For maize, spurious associations can be controlled efficiently by the population structure (Q matrix) and kinship matrix (K) (Yu et al. 2006; Patterson et al. 2006; Price et al. 2006).
The MLM approach, which was used to account for both population structure (Q matrix) and pairwise kinship (K matrix), is one of the most common methods of association analysis used to eliminate possible false-positive associations and previously has been applied successfully in plants (Yu et al. 2006). Estimation of the Q matrix with STRUCTURE software is computationally intense, therefore the MLM with Q matrix calculated by PCA may be an ideal method for large data sets (Patterson et al. 2006; Price et al. 2006; Zhao et al. 2007; Zhu and Yu 2009). In the present study, the performance of six statistical models was evaluated to control for false positives in the maize panel; the results indicated that the MLM (PCA + K) model provided the best reduction in false-positive frequency (Fig. S5). In addition, false-positive correction was not sensitive to the number of principal components when the number of axes ranged from one to ten, as reported in previous studies (Patterson et al. 2006; Price et al. 2006). These results indicated PCA models that incorporate the top three principal components were appropriate to control for population structure and achieve the lowest frequency of false-positive associations in our association panel. If the distribution of functional alleles is highly correlated with population structure, statistically controlling for population structure might also result in false-negatives. Because GWAS can examine abundant SNPs throughout the entire genome simultaneously, an exceptionally high threshold was used in the present study to reduce the false-positive frequency but increase the frequency of false-negatives, especially if an individual SNP had a small effect on the six traits studied.
Four major peaks, which were identified by association mapping of root and shoot traits for the 45,868 SNPs, were located on chromosomes 5, 6 and 9, respectively (Fig. 1). Compared with reported QTL associated with waterlogging responses, a novel genomic region in bin 5.04 that controls waterlogging responses in RDW, RFW, SDW, SFW and SH was identified, and the SNP–trait associations found by GWAS were validated by AB-QTL mapping. Moreover, two recombinant inbred lines, A3237 (waterlogging tolerant) and A3239 (waterlogging susceptible), derived from the cross between Huangzao4 and Mo17 (both susceptible to waterlogging) were analyzed by bulk segregation analysis. Two SSR markers (umc1155 and umc0481) were tightly linked with waterlogging-tolerance gene(s) and were located close together in the same region in bin 5.04. A3237 and A3239 were also sequenced using the genotyping-by-sequencing method (Elshire et al. 2011), and the effective SNPs for waterlogging tolerance were located in the same QTL region in bin 5.04 (unpublished data). SNP4784 (bin5.04: 159998918) was located in GRMZM2G012046, which encodes the membrane attack complex component/perforin/complement C9 involved in Arabidopsis stress responses (Morita-Yamamuro et al. 2005; Noutoshi et al. 2006). Salavati et al. (2012) reported that the ratio of change in abundance of the membrane attack complex component/perforin/complement C9 (ABE79564.1) protein spots compared to that of the control was 2.22 in an analysis of proteomic changes in roots of soybean seedlings during recovery after flooding. All of these results indicated that the genomic region identified by GWAS was an important region for maize seedling waterlogging tolerance. However, it was unclear whether this major QTL on chromosome 5 had a pleiotropic effect on seedling root and shoot traits.
On the basis of the results of Qiu et al. (2007) and AB-QTL mapping, the major peak located on chromosome 9 also might be an important region affecting maize seedling tolerance to waterlogging. Citrate synthase (CS; EC 2.3.3.1) and alanine aminotransferase (AlaAT; EC 2.6.1.2) QTL were colocated on chromosome 9 (146,862,896–147,007,976) (Zhang et al. 2010a), which is extremely close to SNP200 (147,425,437). CS plays key roles in the tricarboxylic acid cycle. AlaAT catalyzes the reversible transfer of the amino group from the second position of glutamate to pyruvate, to yield 2-oxoglutarate and alanine. Recently, it was reported that transgenic Brassica napus and rice plants overexpressing a barley (Hordeum vulgare) AlaAT cDNA show increased biomass and seed yield (Good et al. 2007; Shrawat et al. 2008). It is also interesting to note that AlaAT is significantly induced in response to waterlogging stress (Zou et al. 2010). These authors concluded that the crosstalk between carbon and amino acid metabolism revealed that amino acid metabolism performs two main roles at the waterlogging late stage: the regulation of cytoplasmic pH and energy supply through breakdown of the carbon skeleton (Zou et al. 2010).
In addition, the major peak on chromosome 6 detected by GWAS was novel compared to the results of AB-QTL mapping, but Qiu et al. (2007) identified a QTL in bin 6.04 for RDW under waterlogged conditions in F2 populations (Table S2). Thus, classical QTL mapping based on a biparental cross can be used as a complementary method to GWAS. From the results of QTL mapping and GWAS, we concluded that four major peaks of association might be associated positively with waterlogging tolerance.
Of uni-SNPs with P < 2.18 × 10−4, 70.12 % (33/47) were coincident with previously reported QTL associated with waterlogging-responsive traits in different biparental populations. For example, Qiu et al. (2007) identified an important cluster of QTL for RL on chromosome 7 that overlapped with SNP43260 (bin 7.02: 21,185,816) associated with RL under waterlogging with P = 1.74 × 10−4 (Table S2). Mano et al. (2009) reported that Qaer1.05-1.06 overlapped with SNP30312 (bin 1.05: 154,122,968) associated with RL under waterlogging with P = 1.26 × 10−4 (Table S2). These results indicated that most putative QTL existed within the empirical significance threshold.
The present results prove that GWAS is invaluable for identification of waterlogging response-related genes and offered novel insights for the genetic study of other environmentally dependent and genetically complex traits. In order to fine map and clone the waterlogging-tolerance gene(s) in maize seedlings, it will be necessary for future research to validate the present findings, including development of markers based on associated SNPs, increasing the sample size and marker density of the association population, and validating the results in different genetic backgrounds and environments.
Abbreviations
- SNP:
-
Single nucleotide polymorphism
- AB:
-
Advanced backcross
- PCA:
-
Principal components analysis
- LD:
-
Linkage disequilibrium
- QTL:
-
Quantitative trait locus
- GWAS:
-
Genome-wide association study
- SH:
-
Seedling height
- RL:
-
Root length
- SFW:
-
Shoot fresh weight
- RFW:
-
Root fresh weight
- SDW:
-
Shoot dry weight
- RDW:
-
Root dry weight
References
Abiko T, Kotula L, Shiono K, Malik AI, Colmer TD, Nakazono M (2012) Enhanced formation of aerenchyma and induction of a barrier to radial oxygen loss in adventitious roots of Zea nicaraguensis contribute to its waterlogging tolerance as compared with maize (Zea mays ssp. mays). Plant Cell Environ 35:1618–1630. doi:10.1111/j.1365-3040.2012.02513.x
Aranzana MJ, Kim S, Zhao K, Bakker E, Horton M, Jakob K, Lister C, Molitor J, Shindo C, Tang C, Toomajian C, Traw B, Zheng H, Bergelson J, Dean C, Marjoram P, Nordborg M (2005) Genome-wide association mapping in Arabidopsis identifies previously known flowering time and pathogen resistance genes. PLoS Genet 1(5):e60. doi:10.1371/journal.pgen.0010060
Atwell S, Huang YS, Vilhjalmsson BJ, Willems G, Horton M, Li Y, Meng D, Platt A, Tarone AM, Hu TT, Jiang R, Muliyati NW, Zhang X, Amer MA, Baxter I, Brachi B, Chory J, Dean C, Debieu M, de Meaux J, Ecker JR, Faure N, Kniskern JM, Jones JD, Michael T, Nemri A, Roux F, Salt DE, Tang C, Todesco M, Traw MB, Weigel D, Marjoram P, Borevitz JO, Bergelson J, Nordborg M (2010) Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature 465(7298):627–631
Barrett JC, Fry B, Maller J, Daly MJ (2005) Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21(2):263–265
Belo A, Zheng P, Luck S, Shen B, Meyer DJ, Li B, Tingey S, Rafalski A (2008) Whole genome scan detects an allelic variant of fad2 associated with increased oleic acid levels in maize. Mol Genet Genomics 279(1):1–10
Breseghello F, Sorrells ME (2006) Association mapping of kernel size and milling quality in wheat (Triticum aestivum L.) cultivars. Genetics 172(2):1165–1177
Burgos M, Messmer M, Stamp P, Schmid J (2001) Flooding tolerance of spelt (Triticum spelta L.) compared to wheat (Triticum aestivum L.)—a physiological and genetic approach. Euphytica 122(2):287–295
Ching A, Caldwell KS, Jung M, Dolan M, Smith OS, Tingey S, Morgante M, Rafalski AJ (2002) SNP frequency, haplotype structure and linkage disequilibrium in elite maize inbred lines. BMC Genet 3:19
Cook JP, McMullen MD, Holland JB, Tian F, Bradbury P, Ross-Ibarra J, Buckler ES, Flint-Garcia SA (2012) Genetic architecture of maize kernel composition in the nested association mapping and inbred association panels. Plant Physiol 158(2):824–834
Dennis ES, Dolferus R, Ellis M, Rahman M, Wu Y, Hoeren FU, Grover A, Ismond KP, Good AG, Peacock WJ (2000) Molecular strategies for improving waterlogging tolerance in plants. J Exp Bot 51(342):89–97
Devlin B, Roeder K (1999) Genomic control for association studies. Biometrics 55(4):997–1004
Ellis MH, Dennis ES, Peacock WJ (1999) Arabidopsis roots and shoots have different mechanisms for hypoxic stress tolerance. Plant Physiol 119(1):57–64
Elshire RJ, Glaubitz JC, Sun Q, Poland JA, Kawamoto K, Buckler ES, Mitchell SE (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS One 6(5)
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14(8):2611–2620
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164(4):1567–1587
Good AG, Johnson SJ, DePauw M, Carroll RT, Savidov N (2007) Engineering nitrogen use efficiency with alanine aminotransferase. Can J Bot 85:252–262
Hardy OJ, Vekemans X (2002) SPAGEDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2(4):618–620
Hattori Y, Nagai K, Furukawa S, Song XJ, Kawano R, Sakakibara H, Wu J, Matsumoto T, Yoshimura A, Kitano H, Matsuoka M, Mori H, Ashikari M (2009) The ethylene response factors SNORKEL1 and SNORKEL2 allow rice to adapt to deep water. Nature 460(7258):1026–1030
Huang X, Wei X, Sang T, Zhao Q, Feng Q, Zhao Y, Li C, Zhu C, Lu T, Zhang Z, Li M, Fan D, Guo Y, Wang A, Wang L, Deng L, Li W, Lu Y, Weng Q, Liu K, Huang T, Zhou T, Jing Y, Lin Z, Buckler ES, Qian Q, Zhang QF, Li J, Han B (2010) Genome-wide association studies of 14 agronomic traits in rice landraces. Nat Genet 42(11):961–967
Huang X, Zhao Y, Wei X, Li C, Wang A, Zhao Q, Li W, Guo Y, Deng L, Zhu C, Fan D, Lu Y, Weng Q, Liu K, Zhou T, Jing Y, Si L, Dong G, Huang T, Lu T, Feng Q, Qian Q, Li J, Han B (2012) Genome-wide association study of flowering time and grain yield traits in a worldwide collection of rice germplasm. Nat Genet 44(1):32–39
Ingvarsson PK, Street NR (2011) Association genetics of complex traits in plants. New Phytol 189(4):909–922
Knapp SJ, Stroup WW, Ross WM (1985) Exact confidence intervals for heritability on a progeny mean basis. Crop Sci 25:192–195
Kump KL, Bradbury PJ, Wisser RJ, Buckler ES, Belcher AR, Oropeza-Rosas MA, Zwonitzer JC, Kresovich S, McMullen MD, Ware D, Balint-Kurti PJ, Holland JB (2011) Genome-wide association study of quantitative resistance to southern leaf blight in the maize nested association mapping population. Nat Genet 43(2):163–168
Li HH, Ye GY, Wang JK (2007) A modified algorithm for the improvement of composite interval mapping. Genetics 175(1):361–374
Li H, Vaillancourt R, Mendham N, Zhou M (2008) Comparative mapping of quantitative trait loci associated with waterlogging tolerance in barley (Hordeum vulgare L.). BMC Genomics 9:401
Li HM, Liang H, Tang ZX, Zhang HQ, Yan BJ, Ren ZL (2012) QTL Analysis for grain pentosans and hardness index in a Chinese 1RS.1BL × non-1RS.1BL wheat cross. Plant Mol Biol Report. doi:10.1007/s11105-012-0517-4
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21(9):2128–2129
Liu YZ, Tang B, Zheng YL, Ma KJ, Xu SZ, Qiu FZ (2010) Screening methods for waterlogging tolerance at Maize (Zea mays L.) seedling stage. Agric Sci China 9(3):362–369
Lu Y, Zhang S, Shah T, Xie C, Hao Z, Li X, Farkhari M, Ribaut JM, Cao M, Rong T, Xu Y (2010) Joint linkage-linkage disequilibrium mapping is a powerful approach to detecting quantitative trait loci underlying drought tolerance in maize. Proc Natl Acad Sci USA 107(45):19585–19590
Mackay TF (2009) Genetics. A-maize-ing diversity. Science 325(5941):688–689
Mano Y, Omori F (2008) Verification of QTL controlling root aerenchyma formation in a maize × teosinte “Zea nicaraguensis” advanced backcross population. Breeding Sci 58:217–223
Mano Y, Omori F (2009) High-density linkage map around the root aerenchyma locus Qaer1.06 in the backcross populations of maize Mi29 × teosinte “Zea nicaraguesis”. Breeding Sci 59:427–733
Mano Y, Muraki M, Komatsu T, Fujimori M, Akiyama F, Takamizo T (2002) Varietal difference in pre-germination flooding tolerance and waterlogging tolerance at the seedling stage in maize inbred lines. Jpn J Crop Sci 71(3):361–367
Mano Y, Muraki M, Fujimori M, Takamizo T, Kindiger B (2005a) AFLP–SSR maps of maize × teosinte and maize × maize: comparison of map length and segregation distortion. Plant Breeding 124:432–439
Mano Y, Muraki M, Fujimori M, Takamizo T, Kindiger B (2005b) Identification of QTL controlling adventitious root formation during flooding conditions in teosinte (Zea mays ssp. huehuetenangensis) seedlings. Euphytica 142:33–42
Mano Y, Omori F, Muraki M, Takamizo T (2005c) QTL mapping of adventitious root formation under flooding conditions in tropical maize (Zea mays L.) seedlings. Breeding Sci 55(3):343–347
Mano Y, Muraki M, Takamizo T (2006) Identification of QTL controlling flooding tolerance in reducing soil conditions in maize (Zea mays L.) seedlings. Plant Prod Sci 9(2):176–181
Mano Y, Omori F, Takamizo T, Kindiger B, Bird RM, Loaisiga CH, Takahashi H (2007) QTL mapping of root aerenchyma formation in seedlings of a maize × rare teosinte “Zea nicaraguensis” cross. Plant Soil 295:103–113
Mano Y, Omori F, Kindiger B, Takahashi H (2008) A linkage map of maize × teosinte Zea luxurians and identification of QTLs controlling root aerenchyma formation. Mol Breeding 21:327–337
Mano Y, Omori F, Loaisiga CH, Bird RM (2009) QTL mapping of above-ground adventitious roots during flooding in maize × teosinte “Zea nicaraguensis” backcross population. Plant Root 3:3–9
Mano Y, Omori F, Takeda K (2012) Construction of intraspecific linkage maps, detection of a chromosome inversion, and mapping of QTL for constitutive root aerenchyma formation in the teosinte “Zea nicaraguensis”. Mol Breeding 29:137–146
Morita-Yamamuro C, Tsutsui T, Sato M, Yoshioka H, Tamaoki M, Ogawa D, Matsuura H, Yoshihara T, Ikeda A, Uyeda I, Yamaguchi J (2005) The Arabidopsis gene CAD1 controls programmed cell death in the plant immune system and encodes a protein containing a MACPF domain. Plant Cell Physiol 46(6):902–912
Nielsen R (2004) Late season flooding of maize. Pioneer Technical Insights 342:1–5
Noutoshi Y, Kuromori T, Wada T, Hirayama T, Kamiya A, Imura Y, Yasuda M, Nakashita H, Shirasu K, Shinozaki K (2006) Loss of Necrotic Spotted Lesions 1 associates with cell death and defense responses in Arabidopsis thaliana. Plant Mol Biol 62(1–2):29–42
Palaisa KA, Morgante M, Williams M, Rafalski A (2003) Contrasting effects of selection on sequence diversity and linkage disequilibrium at two phytoene synthase loci. Plant Cell 15(8):1795–1806
Palaisa K, Morgante M, Tingey S, Rafalski A (2004) Long-range patterns of diversity and linkage disequilibrium surrounding the maize Y1 gene are indicative of an asymmetric selective sweep. Proc Natl Acad Sci USA 101(26):9885–9890
Patterson N, Price AL, Reich D (2006) Population structure and eigenanalysis. Plos Genetics 2(12):e190
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38(8):904–909
Pritchard JK, Stephens M, Rosenberg NA, Donnelly P (2000) Association mapping in structured populations. Am J Hum Genet 67(1):170–181
Purcell S, Cherny SS, Sham PC (2003) Genetic power calculator: design of linkage and association genetic mapping studies of complex traits. Bioinformatics 19(1):149–150
Qiu F, Zheng Y, Zhang Z, Xu S (2007) Mapping of QTL associated with waterlogging tolerance during the seedling stage in maize. Ann Bot 99(6):1067–1081
Rafalski A (2002) Applications of single nucleotide polymorphisms in crop genetics. Curr Opin Plant Biol 5(2):94–100
Rafalski JA (2010) Association genetics in crop improvement. Curr Opin Plant Biol 13(2):174–180
Remington DL, Thornsberry JM, Matsuoka Y, Wilson LM, Whitt SR, Doebley J, Kresovich S, Goodman MM, Buckler ES (2001) Structure of linkage disequilibrium and phenotypic associations in the maize genome. Proc Natl Acad Sci USA 98(20):11479–11484
Rohlf FJ (2000) NTSYS-pc numerical taxonomy and multivariate analysis system. version 21 Exeter Software: New York
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci USA 81(24):8014–8018
Salavati A, Khatoon A, Nanjo Y, Komatsu S (2012) Analysis of proteomic changes in roots of soybean seedlings during recovery after flooding. J Proteomics 75(3):878–893
Schnable PS, Ware D, Fulton RS, Stein JC, Wei F, Pasternak S, Liang C, Zhang J, Fulton L, Graves TA, Minx P, Reily AD, Courtney L, Kruchowski SS, Tomlinson C, Strong C, Delehaunty K, Fronick C, Courtney B, Rock SM, Belter E, Du F, Kim K, Abbott RM, Cotton M, Levy A, Marchetto P, Ochoa K, Jackson SM, Gillam B, Chen W, Yan L, Higginbotham J, Cardenas M, Waligorski J, Applebaum E, Phelps L, Falcone J, Kanchi K, Thane T, Scimone A, Thane N, Henke J, Wang T, Ruppert J, Shah N, Rotter K, Hodges J, Ingenthron E, Cordes M, Kohlberg S, Sgro J, Delgado B, Mead K, Chinwalla A, Leonard S, Crouse K, Collura K, Kudrna D, Currie J, He R, Angelova A, Rajasekar S, Mueller T, Lomeli R, Scara G, Ko A, Delaney K, Wissotski M, Lopez G, Campos D, Braidotti M, Ashley E, Golser W, Kim H, Lee S, Lin J, Dujmic Z, Kim W, Talag J, Zuccolo A, Fan C, Sebastian A, Kramer M, Spiegel L, Nascimento L, Zutavern T, Miller B, Ambroise C, Muller S, Spooner W, Narechania A, Ren L, Wei S, Kumari S, Faga B, Levy MJ, McMahan L, Van Buren P, Vaughn MW, Ying K, Yeh CT, Emrich SJ, Jia Y, Kalyanaraman A, Hsia AP, Barbazuk WB, Baucom RS, Brutnell TP, Carpita NC, Chaparro C, Chia JM, Deragon JM, Estill JC, Fu Y, Jeddeloh JA, Han Y, Lee H, Li P, Lisch DR, Liu S, Liu Z, Nagel DH, McCann MC, SanMiguel P, Myers AM, Nettleton D, Nguyen J, Penning BW, Ponnala L, Schneider KL, Schwartz DC, Sharma A, Soderlund C, Springer NM, Sun Q, Wang H, Waterman M, Westerman R, Wolfgruber TK, Yang L, Yu Y, Zhang L, Zhou S, Zhu Q, Bennetzen JL, Dawe RK, Jiang J, Jiang N, Presting GG, Wessler SR, Aluru S, Martienssen RA, Clifton SW, McCombie WR, Wing RA, Wilson RK (2009) The B73 maize genome: complexity, diversity, and dynamics. Science 326(5956):1112–1115
Setter TL, Yan J, Warburton M, Ribaut JM, Xu Y, Sawkins M, Buckler ES, Zhang Z, Gore MA (2011) Genetic association mapping identifies single nucleotide polymorphisms in genes that affect abscisic acid levels in maize floral tissues during drought. J Exp Bot 62(2):701–716
Shapiro SS, Wilk MB (1965) Analysis of variance test for normality (complete samples). Biometrika 52:591–611
Sharma A, Chauhan RS (2012) Identification and association analysis of castor bean orthologous candidate gene-based markers for high oil content in Jatropha curcas. Plant Mol Biol Rep 30(4):1025–1031. doi:10.1007/s11105-011-0408-0
Shrawat AK, Carroll RT, DePauw M, Taylor GJ, Good AG (2008) Genetic engineering of improved nitrogen use efficiency in rice by the tissue-specific expression of alanine aminotransferase. Plant Biotechnol J 6:722–732
Su Z, Li X, Hao Z, Xie C, Li M, Weng J, Zhang D, Liang X, Wang Z, Gao J, Zhang S (2011) Association analysis of the nced and rab28 genes with phenotypic traits under water stress in maize. Plant Mol Biol Rep 29(3):714–722. doi:10.1007/s11105-010-0279-9
Tenaillon MI, Sawkins MC, Long AD, Gaut RL, Doebley JF, Gaut BS (2001) Patterns of DNA sequence polymorphism along chromosome 1 of maize (Zea mays ssp. mays L.). Proc Natl Acad Sci USA 98(16):9161–9166
Tian F, Bradbury PJ, Brown PJ, Hung H, Sun Q, Flint-Garcia S, Rocheford TR, McMullen MD, Holland JB, Buckler ES (2011) Genome-wide association study of leaf architecture in the maize nested association mapping population. Nat Genet 43(2):159–162
Van Ooijen JW, Voorrips RE (2001) JoinMap 3.0, software for the calculation of genetic linkage maps. Plant Research International Wageningen, The Netherlands
VanToai T, St Martin S, Chase K, Boru G, Schnipke V, Schmitthenner A, Lark K (2001) Identification of a QTL associated with tolerance of soybean to soil waterlogging. Crop Sci 41(4):1247–1252
Visser EJW, Voesenek LACJ, Vartapetian BB, Jackson MB (2003) Flooding and plant growth. Ann Bot-London 91(2):107–109
Wang M, Zhang X, Zhao J, Song W, Zheng Y (2011) Evaluation of the genetic diversity and genome-wide linkage disequilibrium of Chinese maize inbred lines. Aust J Crop Sci 5:1790–1795
Wisser RJ, Kolkman JM, Patzoldt ME, Holland JB, Yu JM, Krakowsky M, Nelson RJ, Balint-Kurti PJ (2011) Multivariate analysis of maize disease resistances suggests a pleiotropic genetic basis and implicates a GST gene. Proc Natl Acad Sci USA 108(18):7339–7344
Xu K, Xu X, Fukao T, Canlas P, Maghirang-Rodriguez R, Heuer S, Ismail AM, Bailey-Serres J, Ronald PC, Mackill DJ (2006) Sub1A is an ethylene-response-factor-like gene that confers submergence tolerance to rice. Nature 442(7103):705–708
Yan J, Shah T, Warburton ML, Buckler ES, McMullen MD, Crouch J (2009) Genetic characterization and linkage disequilibrium estimation of a global maize collection using SNP markers. PLoS One 4(12):e8451. doi:10.1371/journal.pone.0008451
Yan J, Kandianis CB, Harjes CE, Bai L, Kim EH, Yang X, Skinner DJ, Fu Z, Mitchell S, Li Q, Fernandez MG, Zaharieva M, Babu R, Fu Y, Palacios N, Li J, Dellapenna D, Brutnell T, Buckler ES, Warburton ML, Rocheford T (2010a) Rare genetic variation at Zea mays crtRB1 increases beta-carotene in maize grain. Nat Genet 42(4):322–327
Yan J, Yang X, Shah T, Sanchez-Villeda H, Li J, Warburton M, Zhou Y, Crouch JH, Xu Y (2010b) High-throughput SNP genotyping with the GoldenGate assay in maize. Mol Breeding 25:441–451
Yan J, Warburton M, Crouch J (2011) Association mapping for enhancing maize (Zea mays L.) genetic improvement. Crop Sci 51. doi:10.2135/cropsci2010.04.0233
Yang X, Yan J, Shah T, Warburton ML, Li Q, Li L, Gao Y, Chai Y, Fu Z, Zhou Y, Xu S, Bai G, Meng Y, Zheng Y, Li J (2010) Genetic analysis and characterization of a new maize association mapping panel for quantitative trait loci dissection. Theor Appl Genet 121(3):417–431
Yang X, Gao S, Xu S, Zhang Z, Prasanna BM, Li L, Li J, Yan J (2011) Characterization of a global germplasm collection and its potential utilization for analysis of complex quantitative traits in maize. Mol Breeding 28(4):511–526. doi:10.1007/s11032-010-9500-7
Yu JM, Pressoir G, Briggs WH, Bi IV, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38(2):203–208
Zaidi PH, Rafiquea S, Rai PK, Singha NN, Srinivasanb G (2004) Tolerance to excess moisture in maize (Zea mays L.): susceptible crop stages and identification of tolerant genotypes. Field Crop Res 90(2–3):189–202
Zhang N, Gibon Y, Gur A, Chen C, Lepak N, Hohne M, Zhang Z, Kroon D, Tschoep H, Stitt M, Buckler E (2010a) Fine quantitative trait loci mapping of carbon and nitrogen metabolism enzyme activities and seedling biomass in the maize IBM mapping population. Plant Physiol 154(4):1753–1765
Zhang Z, Ersoz E, Lai CQ, Todhunter RJ, Tiwari HK, Gore MA, Bradbury PJ, Yu J, Arnett DK, Ordovas JM, Buckler ES (2010b) Mixed linear model approach adapted for genome-wide association studies. Nat Genet 42(4):355–360
Zhao KY, Aranzana MJ, Kim S, Lister C, Shindo C, Tang CL, Toomajian C, Zheng HG, Dean C, Marjoram P, Nordborg M (2007) An Arabidopsis example of association mapping in structured samples. Plos Genetics 3(1)
Zhu C, Yu J (2009) Nonmetric multidimensional scaling corrects for population structure in association mapping with different sample types. Genetics 182(3):875–888
Zondervan KT, Cardon LR (2004) The complex interplay among factors that influence allelic association. Nat Rev Genet 5(2):89–100
Zou X, Jiang Y, Liu L, Zhang Z, Zheng Y (2010) Identification of transcriptome induced in roots of maize seedlings at the late stage of waterlogging. BMC Plant Biol 10:189
Acknowledgments
This research was supported by the Fundamental Research Funds for the Central University (no. 2011PY132), the National Natural Science Foundation of China (31071428), and the National Basic Research Program of China (973 Program) (2009CB118402).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, X., Tang, B., Yu, F. et al. Identification of Major QTL for Waterlogging Tolerance Using Genome-Wide Association and Linkage Mapping of Maize Seedlings. Plant Mol Biol Rep 31, 594–606 (2013). https://doi.org/10.1007/s11105-012-0526-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-012-0526-3