Abstract
Key message
This is the first time to dissect the mechanism of NACs-mediated disease resistance in plants using metabolomic approach and discover the involvement of ABA signaling pathway in NACs-mediated disease resistance.
Abstract
NAC transcription factors have been validated as important regulators in stress responses, but their molecular mechanisms in plant disease resistance are still largely unknown. Here we report that the NAC gene ONAC066 (LOC_Os01g09550) is significantly activated by rice blast infection. ONAC066 is ubiquitously expressed and this protein is localized in the nucleus. Overexpression of ONAC066 quantitatively enhances resistance to blast disease and bacterial blight in rice. The transcript levels of PR genes are also dramatically induced in ONAC066 overexpressing plants. Exogenous abscisic acid (ABA) strongly activates the transcription of ONAC066 in rice. Further analysis shows that overexpression of ONAC066 remarkably suppresses the expression of ABA-related genes, whereas there are no obvious differences for salicylic acid (SA) and jasmonic acid (JA)-related genes between wild-type and ONAC066 overexpressing plants. Consistently, lower endogenous ABA levels are identified in ONAC066 overexpressing plants compared with wild-type plants before and after blast inoculation, while no significant differences are observed for the SA and JA levels. Yeast one-hybrid assays demonstrate that ONAC066 directly binds to the promoters of LIP9 and NCED4 to modulate their expression. Moreover, the metabolomic study reveals that the ONAC066 overexpressing plants accumulated higher contents of soluble sugars and amino acids both before and after pathogen attack, when compared to wild-type plants. Taken together, our results suggest that ONAC066 positively regulates rice resistance to blast and bacterial blight, and ONAC066 exerts its functions on disease resistance by modulating of ABA signaling pathway, sugars and amino acids accumulation in rice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As sessile organisms, plants are frequently under the attack by various fungus, bacterial and viruses. To survive, sophisticated mechanisms have been evolved in plants to cope with these adverse stresses (Sharma et al. 2013). Upon exposure to the pathogen attack, numerous genes are induced at the transcriptional level. Some of these induced genes can protect plant cells from pathogen-induced damage by the generation of pivotal metabolic proteins, while the others function as regulatory proteins that modulate the genes for signal transduction in defense response pathways (Nakashima et al. 2007; Song et al. 2011). Typical examples of such regulatory proteins are transcription factors, including NACs, bZIPs, MYBs and WRKYs, which have been demonstrated to play pivotal roles in plant disease resistance by modulating the transcription of downstream defense-related genes (Nakashima et al. 2009; Song et al. 2011; Puranik et al. 2012; Jiang et al. 2017).
NAC (NAM, ATAF1/2, and CUC2) is one of the largest families of transcription factors that are specifically expressed in plants (Puranik et al. 2012). They have a highly conserved DNA-binding domain in the N-terminal but a usually diversified transcriptional regulating domain in the C-terminal, and this remarkable diversification across plants reflects their numerous functions (Yokotani et al. 2014; Fang et al. 2015). Thus far, NAC proteins have been validated to regulate a series of biological processes, including the development of embryos and flowers, leaf senescence, cell division, hormone signaling and the synthesis of cell wall (Takada et al. 2001; Olsen et al. 2005; Guo and Gan 2006; Zhong et al. 2006; Kim et al. 2016). In addition, they also act as master regulators in regulating plant response to both biotic and abiotic stresses (Olsen et al. 2005; Puranik et al. 2012; Nuruzzaman et al. 2013). For instance, ATAF1 negatively regulates defense against necrotrophic fungal and bacterial pathogens but positively regulates resistance against non-host pathogen Blumeria graminis f. sp. hordei in Arabidopsis (Jensen et al. 2008; Wang et al. 2009). Overexpressing of GsNAC019 exhibited enhanced tolerance to alkaline stress in Arabidopsis (Cao et al. 2017a) and overexpression of TaRNAC1 caused enhanced drought tolerance in wheat root (Chen et al. 2017).
The rice genome encodes 151 NAC transcription factors (Nuruzzaman et al. 2010). However, so far, only 6 NAC genes (OsNAC6, ONAC122, ONAC131, OsNAC111, OsNAC4 and RIMI) have been validated to be involved in defense responses against pathogen attack (Nakashima et al. 2007; Kaneda et al. 2009; Yoshii et al. 2009; Sun et al. 2013; Yokotani et al. 2014). ONAC122, ONAC131 and OsNAC111 positively regulate plant resistance to Magnaporthe oryzae (Sun et al. 2013; Yokotani et al. 2014), whereas silencing of RIMI confers resistance to Rice dwarf virus (Yoshii et al. 2009). The functions of other rice NAC genes in disease resistance are still unknown. Moreover, few have been reported about the detailed mechanisms of NAC genes in response to plant disease.
In this study, we show that the expression of ONAC066 (LOC_Os03g56580) is strongly induced by M. oryzae infection. Overexpression of ONAC066 causes enhanced resistance to rice blast and bacterial blight disease in rice. Further analysis reveals that ONAC066 regulates rice responses to pathogen infection through modulating the abscisic acid (ABA) signaling pathways. Moreover, we also explore the primary metabolomes in the wild-type Nipponbare and ONAC066 overexpressing plants using a GC–MS based metabolomic approach. The results show that ONAC066-mediated disease resistance is also associated with the regulation of sugar and amino acids metabolism in rice.
Materials and methods
Plant materials and pathogens
The japonica rice Nipponbare and the blast-resistant line BC10 (Liu et al. 2004) were used in this study. The blast isolate (M. oryzae) GD08-T13 and Chinese Xanthomonas oryzae pv. Oryzae (Xoo) race 4 isolate were used for rice blast and bacterial blight inoculation, respectively.
Hormone treatments
Phytohormone treatments were performed using the same method as described in Liu et al. (2016a). Briefly, rice seedlings at the three- to four-leaf stage were sprayed with different hormone solutions, each at a concentration of 100 µM. Plants sprayed with water were used as the controls. Seedling samples were collected at 3 h, 6 h, 12 h and 24 h after treatment. The experiments were repeated twice.
Pathogen inoculation
Spraying and punch methods were used for blast inoculation. M. oryzae isolate GD08-T13 inoculum was prepared as described by Beltenev et al. (2007). The revived isolate GD08-T13 was cultured on prune agar plates (three pieces prune, 5 g lactose, 1 g yeast extract, and 20 g agar bar in 1 L, pH of 6–6.5, autoclaved) for 7 days at room temperature, and mycelial growth was scraped with sterilized spatula and exposed to fluorescent light for 4–5 days to induce sporulation. Spores were collected and suspended in water to reach a concentration of 1 × 106 mL−1. For the spraying inoculation, rice seedlings at the three- to four-leaf stage were inoculated with this isolate by the spraying method (Liu et al. 2016a). The inoculated plants were firstly incubated at 25 °C in dark for 24 h, and then followed by a 12-h photoperiod. Sampling for RNA extraction was conducted at 6 h, 12 h, 24 h and 48 h after inoculation. Disease was assessed 6 days after inoculation by measuring the DLA. For punch inoculation, leaves of Nipponbare and ONAC066 overexpressing plants at the tillering stage were inoculated with GD08-T13 using the punch method described by Ding et al. (2012). Lesion size was determined 10 days after inoculation. For determination of in planta sporulation after punch inoculation, leaf strips containing a lesion spot were excised and submerged in 100 µL of distilled water in a 1.5 mL microcentrifuge tube. After the suspension was vigorously mixed, spores were counted with a microscope (Ding et al. 2012).
To evaluate bacterial blight resistance, Chinese Xoo race 4 isolate was grown on nutrient yeast sucrose broth for 72 h at 30 °C, and then re-suspended in sterile water at an optical density (OD600) of 0.5. Plants were inoculated with the isolate at the booting stage by the leaf-clipping method (Kauffman et al. 1973). Lesion length (centimeters) was measured after 14 days post inoculation (dpi).
Subcellular localization of ONAC066 protein
The CDS sequence of ONAC066 was amplified using primers ONAC066-GFP-F/R (Supplemental Table 1) and the fragment was inserted into the pGFP1 vector to generate the fusion protein pGFP1–ONAC066. The empty pGFP1 and pGFP1–ONAC066 plasmids (1 µg) were transformed into rice stem protoplasts as described by Zhang et al. (2011), or introduced into onion epidermal cells as described by Liu et al. (2016b), respectively. The GFP signals were detected by a laser confocal microscopy (Zeiss LSM710, Germany) after 24 h incubation at 25 °C.
Vector construction and plant transformation
We amplified the full-length CDS of ONAC066 from BC10 using primers ONAC066-OE-F/R (Supplemental Table 1) and subsequently the product was cloned into the pOX vector which has an ubiquitin promoter to generate the ONAC066 overexpressing plants. The promoter region of ONAC066 (1330 bp) was amplified from BC10 using primers ONAC066-GUS-F/R (Supplemental Table 1), and then the fragment was inserted into the pcambia1381Z vector. The constructed plasmids were transformed into the wild-type Nipponbare plants using the method as described by Toki et al. (2006).
Cis-elements analysis of the promoter and the promoter–GUS reporter assay
We downloaded the promoter sequence (about 1330 bp) of ONAC066 from MSU Rice Genome Annotation Project, and then the sequence was scanned by PlantCARE for cis-elements analysis (Lescot et al. 2002).
The GUS reporter assay was conducted using the same method according to our previous report (Liu et al. 2016a).
Real-time PCR analysis
Total RNA was extracted from rice seedlings using Eastep Super total RNA extraction kit (Promega Biotech Co., Ltd, USA). The methods for real-time PCR have been described previously (Liu et al. 2016a). Three plants were used for each replicate. The primers used in the present study are shown in Supplemental Table 1.
Metabolic profiling by GC–MS
The primary metabolites in rice leaves were extracted following the procedures described in detail in the previous study (Salem et al. 2016). Four biological replicates were performed for each treatment, and 12 plants were used for each biological replicate. The dried extract from the lower polar phase was derivatized with N-methyl-N-(trimethylsilyl)trifluoroacetamide (MSTFA), and further analyzed by GC–MS (7890A-5975C, Agilent) following the protocol described previously (Lisec et al. 2006). LECO-Fiehn Rtx5 database and the NIST library were used for peak identification after the pretreatment of GC–MS raw data by Chroma TOF 4.3X software (LECO Corporation) (Kind et al. 2009). The resulting three-dimensional data involving the peak number, sample name, and normalized peak area were fed to the SIMCA software package (V14, Umetrics AB, Umea, Sweden) for multivariate statistical analyses. To refine this statistical analysis for significantly changed metabolites, the metabolites with |P(corr)| ≥ 0.5 and a P-value < 0.05 were considered different between the two comparison groups. P-value was obtained by one way ANOVA (Kind et al. 2009).
Quantification of endogenous hormones and their precursors
The hormone was extracted and quantified using the same method as described previously (Liu et al. 2016b). The optimized MS/MS conditions that used for quantification of ABA, violaxanthin, SA and jasmonic acid (JA) were listed in Supplemental Table 2.
Seed germination assays
The seeds of wild-type Nipponbare and ONAC066 overexpressing plants were sterilized with 75% ethanol and 2.5% NaClO and pre-germinated on 1/2 MS medium for 2 days. Then, the identically sprouted seeds were transplanted on fresh 1/2 MS medium supplemented with ABA or without supplementation of ABA (control). 7 days later, the lengths of the shoot and primary roots were measured.
Yeast one-hybrid assays
The yeast one-hybrid assays were conducted using the same method as described in our previous study (Liu et al. 2016a). The coding sequence of ONAC066 was inserted into the pGADT7 vector while the 1.0 kb promoters of LEA3, LIP9, Rab16A, ABI2, NCED3 and NCED4 were inserted into the pAbAi vector, respectively. The experiments were performed according to the manufacturer’s protocol (Clontech Yeast Protocol Handbook; BD Biosciences Clontech). P53 is used as the positive control.
Results
Expression of ONAC066 is induced by blast inoculation
In our previous study to identify genes that were regulated by leaf blast infection using microarray in a blast resistant line BC10 (Liu et al. 2004), we identified that the transcription of ONAC066 was significantly induced during leaf blast inoculation (data not shown). To further confirm this result in the present study, we performed quantitative RT-PCR to analyze the expression of ONAC066 in Nipponbare seedlings after inoculation with blast isolate GD08-T13. We identified that the expression of ONAC066 was induced by M. oryzae infection and the level remained high until 48 h after inoculation (Fig. 1a), suggesting that ONAC066 might play an important role in rice against blast pathogen attack.
The expression patterns of ONAC066 to rice blast infection, and in different rice tissues. a The transcription of ONAC066 was induced by blast inoculation. Values are means ± standard error (SE) from three biological replicates. The asterisks represent significant differences relative to 0 h treatment (t test, **P < 0.01 and *P < 0.05). b Relative expression levels of ONAC066 in different tissues of Nipponbare plants by real-time PCR. Values are means ± SE from three biological replicates. c–k GUS staining analysis of ONAC66 in different tissues of Nipponbare. c 1-week old seedling; d root of 1-week old seedling; e the first node at the booting stage; f the second node at the booting stage; g the third node at the booting stage; h panicle at the booting stage; i panicle at the initial heading stage; j leaf at the booting stage; k mature seeds. c–j Bar = 1 cm. k Bar = 0.5 cm
ONAC066 is highly expressed in leaves and panicles
To investigate the spatial expression of ONAC066 in rice plants, we analyzed the expression pattern of ONAC066 in various tissues of rice by quantitative RT-PCR. As shown in Fig. 1b, ONAC066 was expressed in all rice tissues examined, but the expression was relatively higher in roots, leaves and panicles. To further confirm this result, transgenic rice plants were generated using the GUS reporter gene, which was driven by the ONAC066 promoter (approximately 1.4 kb upstream of the translation start site). GUS signal was detected in the root and shoot, node, panicles at the booting stage and heading stage, and leaves at the booting stage (Fig. 1c–k), agreeing well with the results from qRT-PCR.
Nuclear localization of ONAC066 protein
To determine the subcellular localization of ONAC066 protein, the coding region of ONAC066 was fused to green fluorescence protein (GFP) which was under the control of the CaMV 35S promoter. The transient expression analysis showed that the GFP signal was localized in the nucleus of rice protoplast cells transfected with GFP-ONAC066 protein, whereas the control cells (transformed with the empty vector) exhibited ubiquitous GFP signal (Fig. 2a). We also assayed the localization using onion epidermal cells and similar results were obtained, indicating that ONAC066 is a nucleus localized protein (Fig. 2b).
Overexpression of ONAC066 quantitatively enhanced resistance to blast and bacterial blight in rice
To explore the potential functions of ONAC066 in rice disease resistance, transgenic plants overexpressing ONAC066 (OX-ONAC066) were generated by using the maize ubiquitin promoter. The plant heights of OX-ONAC066 plants were remarkably higher than wild-type Nipponbare (NPB) plants. However, the tiller numbers of OX-ONAC066 plants were significantly reduced, and as a result the grain weights per plant were also significantly lower in the OX-ONAC066 plants compared with the weights in the Nipponbare plants (Supplemental Table 3). Two independent homozygous lines (OX-2 and OX-3) were selected for disease evaluation (Supplemental Fig. 1). Real-time PCR analysis showed that the transcript level of ONAC066 was significantly increased in these transgenic lines (Fig. 3a). We firstly inoculated the 2-week-old OX-ONAC066 plants with blast isolate GD08-T13, which showed strong virulence on wild-type Nipponbare plants, using the spray-inoculation method. The OX-ONAC066 plants showed significantly enhanced resistance to blast disease (P < 0.01), with the diseased areas ranging from 29.8 to 32.2%, compared with 61.3% for wild-type Nipponbare plants (Fig. 3b, c). To confirm this result, the OX-ONAC066 plants were inoculated with the same isolate by punch method and a smaller lesion size was identified in the OX-ONAC066 plants relative to the wild-type Nipponbare plants (Fig. 3d, e). The spores in the infected OX-ONAC066 leaves were also fewer than that in the wild-type Nipponbare leaves (Fig. 3f). Moreover, the OX-ONAC066 plants also exhibited remarkably increased resistance to bacterial blight (P < 0.05), with the lesion length ranging from 10.27 to 10.77 cm for the transgenic plants versus 14.21 cm for wild-type Nipponbare plants (Fig. 4a, b). The growth rate of Xoo on the transgenic plants was also much slower (P < 0.05) than that on the wild-type Nipponbare plants at 12 and 16 days after inoculation (Fig. 4c).
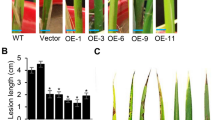
Overexpressing ONAC066 causes enhanced resistance to rice blast. Asterisks represent significant differences relative to wild-type Nipponbare (NPB) plants (Student’s t test, **P < 0.01). a Expression analysis of ONAC066 overexpressing (OX-ONAC066) plants by real-time PCR. Values are mean ± SE from three replicates. b The OX-ONAC066 plants produced less diseased lesions compared to the Nipponbare plants after inoculation with GD08-T13 using the spraying method. c Relative diseased area in the OX-ONAC066 and Nipponbare plants after blast inoculation. Values are means ± SE from 30 biological replicates. d The OX-ONAC066 plants produced smaller diseased lesions compared to the Nipponbare plants after inoculation with GD08-T13 using punch method. e Relative lesion size in the OX-ONAC066 and Nipponbare plants after blast infection. Values are means ± SE from at least ten biological replicates. f The infection ratio of the OX-ONAC066 and Nipponbare plants after punch inoculation. Values are means ± SE from at least ten biological replicates
The phenotypes of ONAC066 overexpressing (OX-ONAC066) plants to bacterial blight disease. The asterisks represent significant differences relative to wild-type Nipponbare (NPB) plants (t test, **P < 0.01 and *P < 0.05). a Overexpression of ONAC066 enhanced resistance to bacterial blight. b Lesion length in the OX-ONAC066 and Nipponbare plants after Xoo inoculation. Values are mean ± SE from 16 biological replicates. c The growth rates of Xoo race 4 in the leaves of OX-ONAC066 and Nipponbare plants. Three leaves 12 or 16 days after inoculation were collected for determination of bacterial populations by counting colony-forming units (cfu). Two independent biological experiments were performed, resulting in similar results. Values are means ± SE from three biological replicates
Since Yokotani et al. (2014) reported that OsNAC111 could enhance rice blast resistance through activation of PR2 (acidic pathogenesis-related protein 2) and PR8, we also investigated if ONAC066 enhanced blast resistance through activation of the PR genes. The expression patterns of nine PR genes were analyzed in the OX-ONAC066 and wild-type Nipponbare plants both before and blast infection. The results showed that pathogen inoculation strongly induced the transcription of PR1a, PR2, PR3, PR4, PR5, PR5-1, PR8 and PR10 both in the wild-type Nipponbare and OX-ONAC066 plants (Fig. 5). Nevertheless, the transcripts of the nine genes were significantly higher (P < 0.05) in OX-ONAC066 plants compared with Nipponbare plants both before and after M. oryzae inoculation. Taken together, these results suggest that ONAC066 positively regulates the responses of rice plants to both blast and bacterial blight diseases.
ONAC066 reduces the expression of genes involved in ABA signaling pathway
To dissect the potential mechanism of ONAC066 in regulation of disease resistance in rice, the promoter sequence of ONAC066 was subjected for cis-elements analysis firstly. Two TC-rich repeats involved in defense and stress responsiveness were identified (Supplemental Table 4), further supporting the regulatory effect of ONAC066 on disease response. Besides, we also observed three ABA-responsive elements (ABREs), two salicylic acid (SA)-responsive elements (TCA element) and two methyl jasmonic acid (MeJA)-responsive elements (TGACG-motif) (Supplemental Table 4). These hormone response elements in the promoter region of ONAC066 imply that ONAC066-mediated disease resistance may involve regulation of these hormone signaling pathways in rice. To confirm this inference, we treated the wild-type Nipponbare plants with exogenous ABA, SA and JA at the three- to four-leaf stage, respectively, and then analyzed the transcription of ONAC066 at different time points after hormone treatment using qRT-PCR. The expression levels of ONAC066 were remarkably increased at 6, 12 and 24 h after ABA treatment in Nipponbare plants, whereas its expression level showed no obvious change under SA or JA treatment (Supplemental Fig. 2).
Moreover, experiments were also conducted to analyze the expression patterns of several well-known stress-related genes that are involved in ABA, JA or SA signaling pathways. The ABA-related genes include three ABA responsive genes (LEA3, Rab16A and LIP9), an ABA signaling gene (ABI2) and two ABA biosynthesis genes (NCED3 and NCED4) (Liu et al. 2012; Chen et al. 2015). LOX and AOS2 are involved in the biosynthesis of JA, while PAL1, ICS1, NH1 and PAD4 are four genes that related to the SA signaling pathway (Deng et al. 2012). Under normal conditions, significantly lower transcription levels of the six ABA-related genes were observed in OX-ONAC066 plants compared with the wild-type Nipponbare plants (Fig. 6a). Pathogen inoculation significantly reduced the transcription of the six genes in both Nipponbare and transgenic plants, while these six genes also exhibited lower expression levels in OX-ONAC066 plants relative to Nipponbare plants after pathogen infection (Fig. 6a). However, the expression levels of the genes involved in JA or SA signaling pathways showed no obvious differences between Nipponbare plants and OX-ONAC066 plants before and after blast inoculation even though the transcripts of LOX, AOS2, ICS1, NHI and PAD4 were remarkably up-regulated by pathogen infection (Fig. 6b). These results together indicate that the disease resistance conferred by ONAC066 is associated with the ABA signaling pathway but not the JA or SA signaling pathways.
ONAC066 reduces the expression of genes involved in ABA signaling pathway. Values are mean ± SE from nine biological replicates. Asterisks represent significant differences relative to Nipponbare plants (t test, **P < 0.01 and *P < 0.05). a The expression patterns of genes involved in ABA signaling pathway in OX-ONAC066 and Nipponbare plants before (0 h) and after blast (24 h) inoculation. b The expression patterns of genes involved in SA or JA signaling pathway in OX-ONAC066 and Nipponbare plants before and after pathogen attack
ONAC066 functions dependent on the ABA signaling pathway
To further understand the relationship between ONAC066 and the ABA signaling pathway, we quantified the contents of the endogenous ABA in the seedlings of wild-type Nipponbare and OX-ONAC066 plants before and after M. oryzae infection. Blast inoculation significantly reduced the endogenous level of ABA in both Nipponbare and OX-ONAC066 plants (Fig. 7). Nevertheless, the endogenous ABA concentration was significantly lower (P < 0.05) in transgenic (OX-ONAC066) plants than that in Nipponbare plants both before and after pathogen inoculation (Fig. 7a). In consistent with the results of gene expression analysis, endogenous JA and SA levels showed no significant differences between Nipponbare and OX-ONAC066 plants though blast inoculation remarkably induced the accumulation of SA in both these plants (Fig. 7). These results further suggest that ONAC066-mediated defense response is at least partially dependent on the ABA signaling pathway in rice.
ONAC066 functions dependent on the ABA signaling pathway. a The endogenous levels of ABA, violaxanthin, SA and JA in the wild-type Nipponbare and OX-ONAC066 plants before (0 h) and after blast (24 h) inoculation. The values are the mean of 12 biological replicates. Asterisks represent significant differences relative to Nipponbare plants before and after blast inoculation (t test, **P < 0.01 and *P < 0.05). FW fresh weight. b Yeast one-hybrid assays indicate the direct interactions between ONAC066 and the promoters of LIP9 and NCED4. P53 is the positive control
To further confirm the interactions between ONAC066 and the ABA signaling pathway, we quantified the endogenous level of violaxanthin, the precursor of ABA (Neuman et al. 2014), in the seedlings of the same plants used for measuring the concentration of hormones before and after M. oryzae inoculation. Similar to ABA, the endogenous level of violaxanthin was significantly reduced in both Nipponbare and OX-ONAC066 plants, but the violaxanthin level was remarkably lower in transgenic plants than in Nipponbare plants before and after blast infection (Fig. 7a). A yeast one-hybrid assays were also performed to determine that whether ONAC066 can directly bind to the promoters of the six ABA-related genes. We identified the direct interactions between ONAC066 and the LIP9 and NCED4 promoters (Fig. 7b), whereas no direct binding were observed between ONAC066 and the promoters of LEA3, Rab16A, ABI2, and NCED3 (data not shown). Importantly, the seed germination assays showed that the shoot and root length of the OX-ONAC066 seeds were more sensitive to ABA treatments than the wild-type Nipponbare seeds (Supplemental Fig. 3). Taken together, these results indicate that ONAC066-mediated defense response is indeed involved in the ABA signaling pathway in rice.
Metabolic changes are different between wild-type and transgenic plants
To further compare the metabolic responses between wild-type Nipponbare and ONAC066 overexpressing plants before and after M. oryzae infection, metabolomic studies were conducted in the leaves of Nipponbare (NPB) and ONAC066 overexpressing plants (the transgenic line, OX-2) both before (0 h) and after 24-h (24 h) blast inoculation. Totally, 84 metabolite peaks were detected in the rice leaf by gas chromatography tandem mass spectrometry (GC–MS), and 51 peaks (representing 48 metabolites) were identified based on their authentic standards, 33 peaks were annotated by matching the MS spectra to those in the NIST11 library (Supplemental Table 5). Retention times and levels of these compounds are provided in Supplemental Table 5. Among the 48 identified metabolites, 11 were identified as sugar-related compounds, 17 were amino acid-related compounds, 5 were free fatty acids, 5 were tricarboxylic acid (TCA) cycle intermediates, and 10 were other compounds. Most were primary metabolites.
The relative intensities of the 84 metabolite peaks were normalized based on the intensity of an internal standard, and used to perform multivariate statistical analyses. Orthogonal partial least squares-discriminant analysis (OPLS-DA) with supervised pattern recognition was conducted to generate an overview of the metabolic pattern of the four samples. The results showed that separation was more evident between the NPB_0h and OX-2_0h groups, suggesting more metabolic differences between NPB and OX-2 rice leaves before infection (Fig. 8). Pairwise comparisons of the leaf metabolome using the OPLS-DA model were performed to find metabolites that differed significantly in the two comparison groups. The OPLS-DA score plots showed distinct differences among metabolomes (Supplemental Figs. 4–7). Parameters of OPLS-DA models were used to evaluate model fitting and the predictive ability of the model. All models were found to have a satisfactory fit with good predictive power values: all R2Y and Q2 values were higher than 0.9, and the P-values of all CV-ANOVA were < 0.05 (Supplemental Table 6).
ONAC066 induces the accumulation of soluble sugars and amino acids
P(corr)-plot analyses allowed us to identify metabolites that were up- or down-regulated between the pair-wise comparisons. For each pairwise comparison, 63 metabolites were remarkably different between NPB_0h and NPB_24h (Supplemental Fig. 8); 61 metabolites were remarkably different between OX-0h and OX-2_24h (Supplemental Fig. 9); 39 metabolites were remarkably different between NPB_0h and OX-0h (Supplemental Fig. 10); and 17 metabolites were remarkably different between NPB_24h and OX-2_24h (Fig. 9), using univariate statistical analyses based on the Student’s t test.
The p (corr)-plot analysis of all metabolites detected by GC–MS in the OX-ONAC066 plants (OX-2) and Nipponbare (NPB) plants after 24 h leaf blast infection. Red column represents up-regulation and green column represents down-regulation with the threshold |P(corr)| > 0.5. Asterisks represent significant differences relative to Nipponbare plants (t test, **P < 0.01, *P < 0.05)
We identified that the concentrations of many kinds of sugars were significantly down-regulated after pathogen inoculation in both the wild-type Nipponbare and OX-ONAC066 plants (Supplemental Figs. 8, 9). However, the contents of several sugars, such as sucrose, glucose, fructose, mannopyranose, and fucopyranose, were remarkably increased in OX-ONAC066 plants relative to Nipponbare plants either before or after blast infection (Fig. 9; Supplemental Fig. 10). Furthermore, the contents of amino acids were significantly up-regulated in both Nipponbare and OX-ONAC066 plants after pathogen infection (Supplemental Figs. 8, 9). The concentrations of many amino acids, including valine, glycine, leucine, proline, and alanine, were also much higher in OX-ONAC066 plants compared to Nipponbare plants either before or after pathogen inoculation (Fig. 9; Supplemental Fig. 10). Adversely, the level of oxalic acid was dramatically down-regulated in OX-ONAC066 plants compared with Nipponbare plants both before and after blast inoculation (Fig. 9; Supplemental Fig. 10). These results together suggest the important roles of sugars, amino acids and oxalic acid in ONAC066-mediated disease resistance.
Discussion
Although several NAC transcription factors have been demonstrated to play crucial roles in biotic stress response in rice (Nakashima et al. 2007; Kaneda et al. 2009; Yoshii et al. 2009; Sun et al. 2013; Yokotani et al. 2014), the regulatory roles of other family members in disease resistance are still largely unknown. Besides, the regulatory mechanisms of NAC genes in rice defense response were not well understood. In this study, we have shown that the transcription of ONAC066 was strongly induced by blast infection and ONAC066 overexpressing transgenic rice plants exhibited quantitatively enhanced resistance to blast and bacterial blight resistance. We also revealed that ONAC066 influenced the metabolisms of sugars and amino acids in normal condition or after pathogen inoculation in rice. To our knowledge, this is the first report to dissecting the mechanism of NACs-mediated disease resistance using metabolomic approach.
ONAC066 exerts its functions on disease resistance through modulating the transcription of stress-related genes
As transcription factors, previous reports have indicated that NAC genes can directly bind to the promoters of downstream target genes to regulate their expression in plants (Hu et al. 2006; Shen et al. 2017). In this study, we have shown that the transcription of nine defense-related PR genes were significantly induced in ONAC066 overexpression plants relative to Nipponbare plants both before and after pathogen inoculation whereas the expression of ABA pathway genes (LEA3, LIP9, Rab16A and so on) were remarkably reduced in ONAC066 overexpressing plants relative to Nipponbare plants before blast infection. These data strongly suggest that ONAC066 exerts its functions on disease resistance through modulating the transcription of well-known stress-related genes in rice. Since Yokotani et al. (2014) reported that OsNAC111 directly activates the promoters of PR2 and PR8 to induce their expression by M. oryzae infection, and Shen et al. (2017) demonstrated that LEA3 and Rab16A are the direct targets of OsNAC2 in the rice response to drought and salt stresses, we deduced that the stress-related genes in our study were also likely to be the direct targets of ONAC066. To verify this inference, a series of yeast one-hybrid assays were conducted. Six ABA-related genes (LEA3, LIP9, Rab16A, ABI2, NCED3 and NCED4) and seven PR genes (PR1a, PR10, PR4, PR5, PR2, PR3 and PR8) were chosen to perform this experiment. As expected, we identified the interactions between ONAC066 and the promoters of LIP9 and NCED4, indicating the direct regulation of LIP9 and NCED4 by ONAC066. The direct binding of ONAC066 to PR gene promoters was not found (data not shown).
ONAC066 confers disease resistance by regulating the ABA-dependent pathway
Generally, it is considered that the responses of plant to pathogens are modulated either by the SA signaling pathway or the JA signaling pathway (Bari and Jones 2009). However, in this study, we show that the disease resistance conferred by ONAC066 is dependent on the ABA signaling pathway but not the SA or JA signaling pathways. The transcripts of stress-related genes, LEA3, LIP9, Rab16A, ABI2, NCED3 and NCED4, which are involved in the ABA-dependent pathway, were remarkably reduced in OX-ONAC066 plants compared with wild-type Nipponbare plants either before or after blast infection. Yeast one-hybrid assays confirmed that ONAC066 could bind directly to the promoters of LIP9 and NCED4 to regulate their transcription. Besides, the expression level of ONAC066 was strongly induced by exogenous ABA, and the OX-ONAC066 plants were more sensitive to ABA treatments than wild-type Nipponbare plants. Moreover, we also identified that the endogenous ABA and violaxanthin levels were significantly lower in ONAC066 overexpressing plants compared with Nipponbare plants after pathogen infection. In contrast, the expression of ONAC066 was not influenced by exogenous SA and JA, and the endogenous levels of these two hormones also exhibited no obvious changes between Nipponbare and ONAC066 overexpressing plants before and after pathogen attack. These results together suggest that ONAC066 confers disease resistance by regulating the ABA-dependent pathway. This is consistent with the previous report that ABA plays a negative role in rice-M. oryzae interactions (Jiang et al. 2010; Ulferts et al. 2015). Recently, several studies suggested that the balance between ABA and SA signaling pathways is an important determinant for the outcome of plant–pathogen interactions, and ABA-mediated susceptibility to pathogen infection may partially depend on the suppression of the SA signaling pathway (Jiang et al. 2010; Xu et al. 2013; Gao et al. 2016). Our results here also indicate that ONAC066-mediated disease resistance involves regulation of the ABA signaling pathway and ONAC066 alleviates ABA-mediated disease susceptibility.
Roles of soluble sugars in ONAC066-mediated defense response
Recently, more and more evidence has shown the important roles of sugars in plant disease resistance (Gómez-Ariza et al. 2007; Qian et al. 2015; Machado et al. 2015; Moore et al. 2015). For instance, a higher level of sucrose was accumulated in the leaves of PRms overexpressing rice plants that exhibited enhanced blast resistance, and pretreatment of wild-type rice plants with exogenous sucrose causes enhanced resistance to M. oryzae (Gómez-Ariza et al. 2007). In the present study, higher concentrations of sucrose and fructose were also identified in resistance-enhanced ONAC066 overexpressing plants relative to Nipponbare plants both before and after M. oryzae inoculation. These results together suggest the positive roles of sugars in regulating plant response to blast disease in rice. However, we also found that the levels of most sugars (sucrose, fructose, and others) were dramatically reduced after 24 h blast infection in both Nipponbare and ONAC066 overexpressing plants. Similar results were also observed by Parker et al. (2009). In their experiments, the levels of sucrose, glucose and fructose were significantly increased upon the appearance of visible blast lesions in rice plants. Our result may reflect the status of sugar contents at the early stage during M. oryzae infection.
The potential roles of amino acids in plant disease resistance
Amino acids are well-known for their central roles in regulating plant growth and development, including protein synthesis, nitrogen metabolism and energy supply (Jespersen et al. 2017). In recent years, emerging evidences have shown that amino acids were also involved in the plant response to various stresses (Parker et al. 2009; Yun et al. 2013; Cao et al. 2017b; Jespersen et al. 2017). For example, increased accumulation of glycine, serine and other amino acids were associated with improved tolerance to heat or drought stresses in creeping bentgrass (Jespersen et al. 2017), and glycine treatment caused enhanced cold tolerance by improving leaf photosynthesis in rice (Cao et al. 2017b). In this study, the metabolic analyses revealed the significant accumulation of many amino acids in both the wild-type and ONAC066 overexpressing plants after blast inoculation, consistent with the previous reports that increases in amino acids were identified in biotrophic pathogen-challenged leaves (Solomon et al. 2003; Parker et al. 2009). Notably, higher levels of glycine, alanine and valine were identified in ONAC066 overexpressing plants compared with wild-type plants after pathogen inoculation, suggesting their important roles in ONAC066-mediated disease resistance. Just as expected, glycine is the necessary component for the synthesis of glutathione, which has been well-documented for its role in the plant defense response (Noctor et al. 1997). However, less information was available on how alanine and valine function in plant disease resistance. Further research will be needed to address these questions.
References
Bari R, Jones JD (2009) Role of plant hormones in plant defense responses. Plant Mol Biol 69:473–488
Beltenev V, Ivanov V, Rozhdestvenskaya I, Cherkashov G, Stepanova T, Shilov V, Pertsev A, Davydov M, Egorov I, Melekestseva I, Narkevsky E, Ignatov V (2007) Protocols on rice blast of IRRI. Internal Rice Research Institute, Manila
Cao L, Yu Y, Ding X, Zhu D, Yang F, Liu B, Sun X, Duan X, Yin K, Zhu Y (2017a) The Glycine soja NAC transcription factor GsNAC019 mediates the regulation of plant alkaline tolerance and ABA sensitivity. Plant Mol Biol 95:253–268
Cao XC, Zhong C, Zhu LF, Zhang JH, Ssjid H, Wu LH, Jin QY (2017b) Glycine increases cold tolerance in rice via the regulation of N uptake, physiological characteristics, and photosynthesis. Plant Physiol Biochem 112:251–260
Chen D, Chai S, McIntyre CL, Xue GP (2017) Overexpression of a predominantly root-expressed NAC transcription factor in wheat roots enhances root length, biomass and drought tolerance. Plant Cell Rep. https://doi.org/10.1007/s00299-017-2224-y
Chen M, Zhao Y, Zhuo C, Lu S, Guo Z (2015) Overexpression of a NF-YC transcription factor from bermudagrass confers tolerance to drought and salinity in transgenic rice. Plant Biotechnol J 13:482–491
Deng H, Liu H, Li X, Xiao J, Wang S (2012) A CCCH-type zinc finger nucleic acid-binding protein quantitatively confers resistance against rice bacterial blight disease. Plant Physiol 158:876–889
Ding B, Bellizzi Mdel R, Ning Y, Meyers BC, Wang GL (2012) HDT701, a histone H4 deacetylase, negatively regulates plant innate immunity by modulating histone H4 acetylation of defense-related genes in rice. Plant Cell 24:3783–3794
Fang Y, Liao K, Du H, Xu Y, Song H, Li X, Xiong L (2015) A stress-responsive NAC transcription factor SNAC3 confers heat and drought tolerance through modulation of reactive oxygen species in rice. J Exp Bot 66:6803–6817
Gao S, Guo W, Feng W, Liu L, Song X, Chen J, Hou W, Zhu H, Tang S, Hu J (2016) LTP3 contributes to disease susceptibility in Arabidopsis by enhancing abscisic acid (ABA) biosynthesis. Mol Plant Pathol 17:412–426
Gómez-Ariza J, Campo S, Rufat M, Estopà M, Messeguer J, San Segundo B, Coca M (2007) Sucrose-mediated priming of plant defense responses and broad-spectrum disease resistance by overexpression of the maize pathogenesis-related PRms protein in rice plants. Mol Plant Microbe Interact 20:832–842
Guo Y, Gan S (2006) AtNAP, a NAC family transcription factor, has an important role in leaf senescence. Plant J 46:601–612
Hu H, Dai M, Yao J, Xiao B, Li X, Zhang Q, Xiong L (2006) Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Natl Acad Sci USA 103:12987–12992
Jensen MK, Hagedorn PH, de Torres-Zabala M, Grant MR, Rung JH, Colling DB, Lyngkjaer MF (2008) Transcriptional regulation by an NAC (NAM–ATAF1, 2–CUC2) transcription factor attenuates ABA signaling for efficient basal defense towards Blumeria graminis f. sp. hordei in Arabidopsis. Plant J 56:867–880
Jespersen D, Yu J, Huang B (2017) Metabolic effects of acibenzolar-S-methyl for improving heat or drought stress in creeping bentgrass. Front Plant Sci 8:1224
Jiang CJ, Shimono M, Sugano S, Kojima M, Yazawa K, Yoshida R, Inoue H, Hayashi N, Sakakibara H, Takatsuji H (2010) Abscisic acid interacts antagonistically with salicylic acid signaling pathway in rice-Magnaporthe grisea interaction. Mol Plant Microbe Interact 23:791–798
Jiang J, Ma S, Ye N, Jiang M, Cao J, Zhang J (2017) WRKY transcription factors in plant responses to stresses. J Integr Plant Biol 59:86–101
Kaneda T, Taga Y, Takai R, Iwano M, Matsui H, Takayama S, Isogai A, Che FS (2009) The transcription factor OsNAC4 is a key positive regulator of plant hypersensitive cell death. EMBO J 28:926–936
Kauffman HE, Reddy APK, Hsieh SPY, Merca SD (1973) An improved technique for evaluating resistance of rice varieties to Xanthomonas oryzae. Plant Dis Rep 57:537–541
Kim HJ, Nam HG, Lim PO (2016) Regulatory network of NAC transcription factors in leaf senescence. Curr Opin Plant Biol 33:48–56
Kind T, Wohlgemuth G, Lee DY, Lu Y, Palazoglu M, Shahbaz S, Fiehn O (2009) FiehnLib: mass spectral and retention index libraries for metabolomics based on quadrupole and time-of-flight gas chromatography/mass spectrometry. Anal Chem 81:10038–10048
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327
Lisec J, Schauer N, Kopka J, Willmitzer L, Fernie AR (2006) Gas chromatography mass spectrometry-based metabolite profiling in plants. Nat Protoc 1:387–396
Liu D, Chen X, Liu J, Ye J, Guo Z (2012) The rice ERF transcription factor OsERF922 negatively regulates resistance to Magnaporthe oryzae and salt tolerance. J Exp Bot 63:3899–3911
Liu B, Zhang S, Zhu X, Yang Q, Wu S, Mei M, Mauleon R, Leach J, Mew T, Leung H (2004) Candidate defense genes as predictors of quantitative blast resistance in rice. Mol Plant Microbe Interact 17:1146–1152
Liu Q, Yang J, Zhang S, Zhao J, Feng A, Yang T, Wang X, Mao X, Dong J, Zhu X, Leung H, Leach JE, Liu B (2016a) OsGF14b positively regulates panicle blast resistance, but negatively regulates leaf blast resistance in rice. Mol Plant Microbe Interact 29:46–56
Liu Q, Yang J, Yan S, Zhang S, Zhao J, Wang W, Yang T, Wang X, Mao X, Dong J, Zhu X, Liu B (2016b) The germin-like protein OsGLP2-1 enhances resistance to fungal blast and bacterial blight in rice. Plant Mol Biol 92:411–423
Machado RA, Arce CC, Ferrieri AP, Baldwin IT, Erb M (2015) Jasmonate-dependent depletion of soluble sugars compromises plant resistance to Manduca sexta. New Phytol 207:91–105
Moore JW, Herrera-Foessel S, Lan C, Schnippenkoetter W, Ayliffe M, Huerta-Espino J, Lillemo M, Viccars L, Milne R, Periyannan S, Kong X, Spielmeyer W, Talbot M, Bariana H, Patrick JW, Dodds P, Singh R, Lagudah E (2015) A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nat Genet 47:1494–1498
Nakashima K, Tran LS, Van Nguyen D, Fujita M, Maruyama K, Todaka D, Ito Y, Hayashi N, Shinozaki K, Yamaguchi-Shinozaki K (2007) Functional analysis of a NAC-type transcription factor OsNAC6 involved in abiotic and biotic stress-responsive gene expression in rice. Plant J 51:617–630
Nakashima K, Ito Y, Yamaguchi-Shinozaki K (2009) Transcriptional regulatory networks in response to abiotic stresses in Arabidopsis and grasses. Plant Physiol 149:88–95
Neuman H, Galpaz N, Cunningham FX Jr, Zamir D, Hirschberg J (2014) The tomato mutation nxd1 reveals a gene necessary for neoxanthin biosynthesis and demonstrates that violaxanthin is a sufficient precursor for abscisic acid biosynthesis. Plant J 78:80–93
Noctor G, Arisic CM, Jouanin L, Valadier HM, Roux Y, Foyer CH (1997) The role of glycine in determining the rate of glutathione synthesis in poplar. Possible implications for glutathione production during stress. Physiol Plant 100:255–263
Nuruzzaman M, Manimekalai R, Sharoni AM, Satoh K, Kondoh H, Ooka H, Kikuchi S (2010) Genome-wide analysis of NAC transcription factor family in rice. Gene 465:30–44
Nuruzzaman M, Sharoni AM, Kikuchi S (2013) Roles of NAC transcription factors in the regulation of biotic and abiotic stress responses in plants. Front Microbiol 4:248
Olsen AN, Ernst HA, Leggio LL, Skriver K (2005) NAC transcription factors: structurally distinct, functionally diverse. Trends Plant Sci 10:79–87
Parker D, Beckmann M, Zubair H, Enot DP, Caracuel-Rios Z, Overy DP, Snowdon S, Talbot NJ, Draper J (2009) Metabolomic analysis reveals a common pattern of metabolic re-programming during invasion of three host plant species by Magnaporthe grisea. Plant J 59:723–737
Puranik S, Sahu PP, Srivastava PS, Prasad M (2012) NAC proteins: regulation and role in stress tolerance. Trends Plant Sci 17:369–381
Qian Y, Tan DX, Reiter RJ, Shi H (2015) Comparative metabolomics analysis highlights the involvement of sugars and glycerol in melatonin-mediated innate immunity against bacterial pathogen in Arabidopsis. Sci Rep 5:15815
Salem MA, Jüppner J, Bajdzienko K, Giavalisco P (2016) Protocol: a fast, comprehensive and reproducible one-step extraction method for the rapid preparation of polar and semi-polar metabolites, lipids, proteins, starch and cell wall polymers from a single sample. Plant Methods 12:45
Sharma R, De Vleesschauwer D, Sharma MK, Ronald PC (2013) Recent advances in dissecting stress-regulatory crosstalk in rice. Mol Plant 6:250–260
Shen J, Lv B, Luo L, He J, Mao C, Xi D, Ming F (2017) The NAC-type transcription factor OsNAC2 regulates ABA-dependent genes and abiotic stress tolerance in rice. Sci Rep 7:40641
Solomon PS, Tan KC, Oliver RP (2003) The nutrient supply of pathogenic fungi; a fertile field for study. Mol Plant Pathol 4:203–210
Song SY, Chen Y, Chen J, Dai XY, Zhang WH (2011) Physiological mechanisms underlying OsNAC5-dependent tolerance of rice plants to abiotic stress. Planta 234:331–345
Sun L, Zhang H, Li D, Huang L, Hong Y, Ding XS. Nelson RS, Zhou X, Song F (2013) Functions of rice NAC transcriptional factors, ONAC122 and ONAC131, in defense responses against Magnaporthe grisea. Plant Mol Biol 81:41–56
Takada S, Hibara K, Ishida T, Tasaka M (2001) The CUP-SHAPED COTYLEDON1 gene of Arabidopsis regulates shoot apical meristem formation. Development 128:1127–1135
Toki S, Hara N, Ono K, Onodera H, Tagiri A, Oka S, Tanaka H (2006) Early infection of scutellum tissue with Agrobacterium allows high-speed transformation of rice. Plant J 47:969–976
Ulferts S, Delventhal R, Splivallo R, Karlovsky P, Schaffrath U (2015) Abscisic acid negatively interferes with basal defence of barley against Magnaporthe oryzae. BMC Plant Biol 15:7
Wang X, Basnayake BM, Zhang H, Li G, Li W, Virk N, Menqiste T, Song F (2009) The Arabidopsis ATAF1, a NAC transcription actor, is a negative regulator of defense responses against necrotrophic fungal and bacterial pathogens. Mol Plant-Microbe Interact 22:1227–1238
Xu J, Audenaert K, Hofte M, De Vleesschauwer D (2013) Abscisic acid promotes susceptibility to the rice leaf blight pathogen Xanthomonas oryzae pvoryzae by suppressing salicylic acid-mediated defenses. PLoS ONE 8:e67413
Yokotani N, Tsuchida-Mayama T, Ichikawa H, Mitsuda N, Ohme-Takagi M, Kaku H, Minami E, Nishizawa Y (2014) OsNAC111, a blast disease-responsive transcription factor in rice, positively regulates the expression of defense-related genes. Mol Plant-Microbe Interact 27:1027–1034
Yoshii M, Shimizu T, Yamazaki M, Higashi T, Miyao A, Hirochika H, Omura T (2009) Disruption of a novel gene for a NAC-domain protein in rice confers resistance to Rice dwarf virus. Plant J 57:615–625
Yun Z, Gao H, Liu P, Liu S, Luo T, Jin S, Xu Q, Xu J, Cheng Y, Deng X (2013) Comparative proteomics and metabolomics profiling of citrus fruit with enhancement of disease resistance by postharvest heat treatment. BMC Plant Biol 13:44
Zhang Y, Su J, Duan S, Ao Y, Dai J, Liu J, Wang P, Li Y, Liu B, Feng D, Wang J, Wang H (2011) A highly efficient rice green tissue protoplast system for transient gene expression and studying light/chloroplast-related processes. Plant Methods 7:30
Zhong R, Demura T, Ye ZH (2006) SND1, a NAC domain transcription factor, is a key regulator of secondary wall synthesis in fibers of Arabidopsis. Plant Cell 18:3158–3170
Acknowledgements
This research was supported partially by the NSFC-IRRI Project (31461143019), Science and Technology Projects of Guangdong Province (2015B020231002, 2016A050502030 and 2014B070706013), the Natural Science Foundation of Guangdong Province (2014A030310489), and the Common Technical Innovation Team of Guangdong Province on Agricultural Seed Industry (2017LM2148).
Author information
Authors and Affiliations
Contributions
QL conducted the quantitative qRT-PCR assay, transgenic functional confirmation and chemical treatment experiments, drafting the manuscript and proposal writing. SY conducted the metabolic study, drafting the manuscript and proposal writing. WH performed the quantification of endogenous ABA. JY evaluated the blast and Xoo resistance. JD, SZ, JZ, TY and XM participated in RNA extraction and quantitative qRT-PCR assays. XZ and BL conceived of the study, drafted proposal and corrected manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, Q., Yan, S., Huang, W. et al. NAC transcription factor ONAC066 positively regulates disease resistance by suppressing the ABA signaling pathway in rice. Plant Mol Biol 98, 289–302 (2018). https://doi.org/10.1007/s11103-018-0768-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-018-0768-z