Abstract
The most devastating diseases in rice (Oryza sativa) are sheath blight caused by the fungal necrotroph Rhizoctonia solani, rice blast by hemibiotrophic fungus Magnaporthe oryzae, and leaf blight by bacterial biotroph Xanthomonas oryzae (Xoo). It has been reported that the Class III acyl-CoA-binding proteins (ACBPs) such as those from dicots (Arabidopsis and grapevine) play a role in defence against biotrophic pathogens. Of the six Arabidopsis (Arabidopsis thaliana) ACBPs, AtACBP3 conferred protection in transgenic Arabidopsis against Pseudomonas syringae, but not the necrotrophic fungus, Botrytis cinerea. Similar to Arabidopsis, rice possesses six ACBPs, designated OsACBPs. The aims of this study were to test whether OsACBP5, the homologue of AtACBP3, can confer resistance against representative necrotrophic, hemibiotrophic and biotrophic phytopathogens and to understand the mechanisms in protection. Herein, when OsACBP5 was overexpressed in rice, the OsACBP5-overexpressing (OsACBP5-OE) lines exhibited enhanced disease resistance against representative necrotrophic (R. solani & Cercospora oryzae), hemibiotrophic (M. oryzae & Fusarium graminearum) and biotrophic (Xoo) phytopathogens. Progeny from a cross between OsACBP5-OE9 and the jasmonate (JA)-signalling deficient mutant were more susceptible than the wild type to infection by the necrotroph R. solani. In contrast, progeny from a cross between OsACBP5-OE9 and the salicylic acid (SA)-signalling deficient mutant was more susceptible to infection by the hemibiotroph M. oryzae and biotroph Xoo. Hence, enhanced resistance of OsACBP5-OEs against representative necrotrophs appears to be JA-dependent whilst that to (hemi)biotrophs is SA-mediated.
Similar content being viewed by others
Introduction
Despite the existence of natural defence mechanisms in plants, crops can become susceptible to bacterial and fungal pathogens which cause negative impacts on food security and food safety worldwide1. This happens when the virulence factors of a pathogen overwhelm the defence responses of a specific host plant2. Phytopathogens are classified into necrotrophs, hemibiotrophs and biotrophs based on their interaction with the host. Necrotrophic pathogens rapidly kill host tissues by secreting toxins and survive on the dead remains, whereas biotrophs derive nutrients from living cells and therefore sustain host viability3,4. Hemibiotrophic pathogens exhibit both forms of nutrient acquisition, starting from the suppression of host immune system in a biotrophic phase, followed by a later necrotrophic phase during which the pathogen induces host cell death by its toxin production5. Thus, necrotrophic pathogens tend to exert a more damaging effect on host plants than biotrophic pathogens and are deemed more detrimental to crop productivity4.
Necrotrophic soil-borne phytopathogenic fungi such as Fusarium and Rhizoctonia infect prominent economically important monocot crops such as rice (Oryza sativa), wheat and maize1,6,7. Fusarium solani infection resulted in a 10–30% drop in yield for pulse crops following severe root-rot8. In Japan, soybean foliar blight arising from the necrotroph Rhizoctonia solani AG-1 culminated to around 70% decrease in yield9. It has been reported that necrotrophic pathogens have a higher economic impact on agriculture over biotrophic pathogens10. Reduction in Australian wheat and barley production from necrotrophic pathogen diseases such as tan spot, and Stagonospora nodorum blotch, respectively, significantly exceeded that from wheat rusts and mildews caused by biotrophic pathogens10. In addition, the necrotroph, Botrytis cinerea, infects nearly all crop plants and creates global annual economic loss from $10 to $100 billion11. To safeguard a stable food supply to an escalating global population, there is an urgent need to control fungal diseases. To this end, proteins that can defend plants from plant pathogens need to be identified and tested for efficacy in transgenic monocots.
The acyl-CoA-binding proteins (ACBPs) represent a major group of proteins associated with acyl-CoA ester transfer in eukaryotes12. The Arabidopsis thaliana ACBPs bind to long-chain acyl-CoA esters and function in plant growth, development and stress responses13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28. Even though the expression of AtACBP3 was upregulated by both bacterial biotroph, Pseudomonas syringae pv tomato DC3000, and fungal necrotroph, B. cinerea, transgenic Arabidopsis AtACBP3-overexpressing (OE) lines were conferred protection only against P. syringae17. Moreover, atacbp3 was susceptible to P. syringae, hemibiotrophic fungal pathogen Colletotrichum higginsianum and the fungal necrotroph B. cinerea in comparison to the wild type (WT)29. It appears that both the overexpression and loss of AtACBP3 affected basal defence against the fungal necrotroph B. cinerea17,29, suggesting that susceptibility of mutant lines to a pathogen does not always correspond to resistance in OE lines.
A homologous Class III ACBP from Vitis vinifera (grape) had displayed enhanced resistance to P. syringae when overexpressed in Arabidopsis30. The same study revealed that transgenic Arabidopsis VvACBP-OEs were more tolerant against hemibiotrophic fungal pathogen C. higginsianum infection in detached leaves, but the response of Arabidopsis VvACBP-OEs against necrotrophic pathogens was not reported30. When compared to ACBPs from dicotyledonous plants17,29,30, ACBPs from monocotyledonous plants such as rice are less well understood. Rice ACBP5 (OsACBP5) resembles AtACBP3 in being classified as the sole rice Class III ACBP, with the acyl-CoA-binding (ACB) domain located at the C-terminus unlike the other three classes31. Previous studies revealed that the ACB domain of OsACBP5 binds to 18:3-acyl-CoA ester31, and interestingly the 18:3 fatty acid (FA) is associated with resistance to the biotroph P. syringae in tomato by activating NADPH oxidase in reactive oxygen species (ROS) production32. The rise in ROS initiates the biosynthesis of the plant hormone salicylic acid (SA) leading to hypersensitive response against the pathogen32,33.
Phytohormones such as SA and jasmonic acid (JA) are well known in regulating plant defence34. SA is synthesized from chorismate, a primary metabolite, through two enzymatic pathways, one involving PHENYLALANINE AMMONIA LYASE and the other ISOCHORISMATE SYNTHASE35,36,37,38. Pattern- and effector-triggered immunity activates SA biosynthesis upon recognition of pathogen-associated molecular patterns (PAMPs) and effectors of pathogens, respectively36,39. The regulatory protein NONEXPRESSOR OF PATHOGENESIS-RELATED GENES1 (NPR1) mostly controls the SA-signalling pathway36. NPR1, triggered by SA, functions as a transcriptional coactivator of the PATHOGENESIS-RELATED (PR) genes36,40,41. PR proteins demonstrate antimicrobial activity and PR-1 represents an important marker for SA-responsive gene expression36,42, while the activation of PR genes leads to SA-mediated defence36.
JA and its structurally-related compounds are synthesized by the oxylipin biosynthesis pathway following pathogen invasion43. JA carboxyl methyl transferase catalyses the methylation of JA to methyl jasmonate (MeJA)44. JA-amidosynthetase (JAR1) catalyses the production of JASMONYL-ISOLEUCINE (JA-Ile) by conjugation of JA to the amino acid isoleucine45. CORONATINE INSENSITIVE1 (COI1), JAR1 and JASMONATE INSENSITIVE1/MYC2 (JIN1/MYC2) are the key components involved in JA-dependent defence responses in plants. COI1 interacts with SKP1/Cullin counterparts and forms the SKP1–Cullin–F-box (SCF)COI1ubiquitin E3 ligase complex. The SCF complex mediates protein degradation which is essential for JA-dependent defence responses46. The expression of JA-related genes is transcriptionally regulated by the JIN1/MYC2 transcription factor47. Jasmonate zim-domain (JAZ) repressor proteins interact with the JIN1/MYC2 transcription factor to suppress the expression of JA-associated genes47. COI1 interacts with JAZ proteins to form the COI1-JAZ complex48. COI1-JAZ functions as a JA-Ile receptor in the SCF complex49. Degradation of the JAZ repressor protein is mediated by the interaction of JA-Ile with COI150. JAZ protein degradation results in the stimulation of JA-related genes by JIN1/MYC2 transcription factors leading to plant defence50.
While the JA pathway generally provides protection against necrotrophic pathogens, the SA pathway is normally associated with resistance to biotrophic pathogens47,51,52. Several studies have demonstrated that the SA pathway functions antagonistically to the JA-signalling pathway51,53. Hence, enhanced tolerance to biotrophic infection is often associated with elevated susceptibility to necrotrophs and vice versa52,54 as demonstrated in AtACBP3-OEs17. The cross talk between SA and JA has been clarified in dicots such as Arabidopsis whereby suppression of the JA pathway by SA functions downstream of the E3 ubiquitin-ligase SCF complex, which targets JAZs for proteasome-mediated degradation55. In comparison, the antagonistic interaction between the SA- and JA-signalling pathways is less well understood in the monocots56,57. Therefore, it would be pertinent to decipher the role of OsACBP5 in plant defence, given its ability to bind 18:3-acyl-CoA ester possibly associated with phytohormone signalling.
Meng et al. had earlier shown in quantitative real-time PCR (qRT-PCR) analysis that of the six OsACBPs, only OsACBP5 mRNA expression was upregulated following infection with the hemibiotrophic rice blast fungal pathogen, Magnaporthe oryzae31. Rice blast is a destructive fungal disease causing a 30% decline in rice production globally58. Our recent study demonstrated that overexpression of OsACBP5 in transgenic Arabidopsis conferred resistance to representative necrotrophic (R. solani, B. cinerea, Alternaria brassicicola), hemibiotrophic (Colletotrichum siamense) and biotrophic (P. syringae) phytopathogens through cell wall-mediated defence as well as SA- and JA-mediated defence pathways59,60. Given the need for crops to be protected against necrotrophic fungal pathogens, the present study follows up on our initial investigations on OsACBP531. To this end, transgenic rice lines overexpressing OsACBP5 were generated and tested against representatives derived from three groups of phytopathogens (the necrotrophs, hemibiotrophs and biotrophs). Transgenic rice harbouring OsACBP5pro::GUS were also produced to examine OsACBP5 regulation.
Results
Transgenic rice OsACBP5-OEs showed enhanced tolerance against two necrotrophic fungal pathogens, R. solani and C. oryzae
Transgenic rice lines overexpressing OsACBP5 (OsACBP5-OEs) were generated and verified by western blot analysis (Supplemental Fig. S1). To explore the function of OsACBP5 in defence, the resistance level of transgenic rice OsACBP5-OEs (OE-1, OE-3, OE-6, OE-9 and OE-11) was first evaluated against sheath blight disease, a severe fungal disease in rice caused by R. solani, using sheath inoculation assays. While typical disease lesions on the WT and vector (pCAMBIA1304)-transformed plants were observed, fewer disease lesions appeared on transgenic rice OsACBP5-OEs (Fig. 1A). At 14 days-post-inoculation (DPI), the lesion lengths on the sheaths of WT and vector-transformed plants were 4 cm and 4.5 cm, respectively, while the average lesion length on sheaths of the OsACBP5-OEs was 1.6 cm, representing an approximately twofold reduction in lesion length in comparison to the controls (Fig. 1B). Consistent results were observed when transgenic rice OsACBP5-OEs were infected with the necrotroph C. oryzae causing narrow brown leaf spot in rice plants (Fig. 1C). The average disease scores in WT and vector-transformed plants at 9 DPI were 5.5 and 5, respectively, while the average disease score on OsACBP5-OEs was 2.3, resulting in an approximately 2.5-fold reduction in comparison to the controls (Fig. 1D). Taken together, these results suggest that the overexpression of OsACBP5 in transgenic rice conferred enhanced resistance to necrotrophs, R. solani and C. oryzae, in comparison to WT and vector-transformed plants.
OsACBP5-OE transgenic rice plants displayed improved resistance against fungal necrotrophs. (A) Sheath blight symptoms on five-week-old WT, vector-transformed control and OsACBP5-OE (OE-1, OE-3, OE-6, OE-9, OE-11) rice inoculated with R. solani. Blue bars = 1 cm. (B) Lesion length following R. solani infection in WT, vector-transformed control and OsACBP5-OEs 14 days-post-inoculation (dpi). (C) Disease phenotype following C. oryzae infection on three-week-old WT, vector-transformed control and OsACBP5-OEs at 9 dpi. Blue bars = 7 mm. (D) Disease scores from C. oryzae-infected leaves. Data points represent means ± SD from three independent experiments. Asterisks indicate significant difference (P < 0.05) in comparison to the controls by Student’s t-test.
Transgenic rice OsACBP5-OEs were more resistant to two hemibiotrophic fungal pathogens, M. oryzae and Fusarium graminearum
The resistance level of transgenic rice OsACBP5-OEs was next investigated against blast disease caused by the hemibiotroph M. oryzae. When spray inoculation (Fig. 2A) was performed to explore the resistance of OsACBP5-OEs against M. oryzae, the average lesion areas in WT and vector-transformed plants at 7 dpi were 5.3 mm2 and 5.2 mm2, respectively, while the average lesion area on OsACBP5-OEs was 1.7 mm2, resulting in approximately 50% reduction in lesion area (Fig. 2B). When the seeds of WT, vector-transformed control and OsACBP5-OEs were imbibed with conidial suspensions of seed-infecting hemibiotroph, F. graminearum, the OsACBP5-OEs exhibited 80–100% of seed germination rates, whereas those of WT and vector-transformed controls were 20% and 40%, respectively, suggesting that OsACBP5-OEs were more resistant to F. graminearum infection (Fig. 2C,D).
Increased resistance of OsACBP5-OE transgenic rice plants to hemibiotrophic fungal infection. (A) Rice blast symptoms after inoculation with M. oryzae on 3-week-old WT, vector-transformed control and OsACBP5-OE (OE-1, OE-3, OE-6, OE-9, OE-11) at 7 days-post-inoculation (dpi). Blue bars = 5 mm. (B) Lesion area on WT, vector-transformed control and OsACBP5-OEs inoculated with M. oryzae. (C) Germination rate of WT, vector-transformed control and OsACBP5-OE rice plants grown from the seeds inoculated with F. graminearum at 7 dpi. (D) WT, vector-transformed control, and OsACBP5-OE rice plants germinated from F. graminearum-infected seeds. Blue bars = 1 cm. Data points represent means ± SD from three independent experiments. Asterisks indicate significant difference (P < 0.05) in comparison to the controls by Student’s t-test.
Transgenic rice OsACBP5-OEs conferred protection against biotrophic bacterial pathogen Xanthomonas oryzae
When transgenic rice OsACBP5-OEs were evaluated against the bacterial leaf blight disease caused by Xanthomonas oryzae pv. oryzae (Xoo) using the leaf-clipping method (Fig. 3A), the average lesion length on OsACBP5-OEs at 14 dpi was 0.7 cm, while the average lesion lengths in WT and vector-transformed plants were 5.7 cm and 5.5 cm, respectively, representing a fivefold reduction in disease development (Fig. 3B). These results indicate that OsACBP5-OEs displayed enhanced tolerance to Xoo.
OsACBP5-OE transgenic rice plants are more resistant to the biotrophic bacterial pathogen Xoo infection. (A) Disease symptoms on three-week-old WT, vector-transformed control and OsACBP5-OE (OE-1, OE-3, OE-6, OE-9, OE-11) plants inoculated with Xoo. Leaves were photographed 14 days-post-inoculation. Blue bars = 5 mm. (B) Lesion length after inoculation with Xoo in WT, vector-transformed control and OsACBP5-OEs. Data points represent means ± SD from three independent experiments. Asterisks indicate significant difference (P < 0.05) in comparison to the controls by Student’s t-test.
SA and JA levels in transgenic rice OsACBP5-OEs were elevated
As SA and JA play important roles in regulating immune responses in rice61, possible relationship between SA and JA in enhanced pathogen-resistance in OsACBP5-OEs, was investigated by SA and JA content measurements using gas chromatography-mass spectrometry (GC-MS) on uninfected and R. solani-infected plant samples. A two-fold increase in endogenous SA was shown in OsACBP5-OEs in comparison to the controls (Fig. 4A,B). Similarly, the endogenous JA content in OsACBP5-OEs were two-fold higher than the controls (Fig. 4A, B). When qRT-PCR was performed, the expression of OsNPR1, an SA-signalling regulatory gene, and ALLENE OXIDE SYNTHASE1 (OsAOS1) encoding allene oxide synthase in JA biosynthesis, were upregulated in uninfected and R. solani-infected plant samples of OsACBP5-OEs in comparison to that of controls (Fig. 4C,D). These results indicate that both SA- and JA-signalling pathways are activated in OsACBP5-OEs.
OsACBP5-OE transgenic rice plants displayed elevated SA and JA level. Total SA and JA were extracted from uninfected (A) and R. solani-infected (B) WT, vector-transformed control and OsACBP5-OEs (OE-1, OE-3, OE-6, OE-9, OE-11) and analyzed by GC-MS. qRT-PCR analyses showing the expression of OsNPR1 and OsAOS1 in uninfected (C) and R. solani-infected (D) WT, vector-transformed control and OsACBP5-OEs. Data points represent means ± SD from three independent experiments. FW, fresh weight. Asterisks indicate significant difference (P < 0.05) in comparison to the controls by Student’s t-test.
Protection in rice OsACBP5-OEs are dependent on both SA- and JA-signalling pathways
To determine if SA- and JA-signalling pathways are involved in the enhanced resistance of rice OsACBP5-OEs against various plant pathogens, WT, OsACBP5-OE9 (OE-9), osnpr1 (SA-signalling-deficient mutant), oscoi1 (JA-signalling-deficient mutant) and OE-9 in the osnpr1 or oscoi1 backgrounds were infected with the necrotroph R. solani (Fig. 5A), the hemibiotroph M. oryzae (Fig. 5B) and the biotroph Xoo (Fig. 5C). The measurement of lesion length in R. solani-infected plants showed significantly higher susceptibility in OE-9oscoi1 and oscoi1 plants in comparison to the WT (Fig. 5D). No significant difference was observed between osnpr1 and the WT (Fig. 5D). The OE-9osnpr1 mimicked the response of OE-9 (Fig. 5D). These results indicated that the improved resistance of OsACBP5-OEs to the fungal necrotroph R. solani is JA-dependent. When the plants (WT, OE-9, osnpr1, oscoi1, OE-9osnpr1 and OE-9oscoi1) were infected with the hemibiotrophic fungal pathogen M. oryzae and the biotroph Xoo, OE-9osnpr1 plants were no longer resistant to the pathogen similar to osnpr1 (Fig. 5, E and F). No significant difference was observed in M. oryzae- or Xoo-infected WT and oscoi1 mutant (Fig. 5E,F). OE-9 and OE-9oscoi1 showed similar responses to representative hemibiotrophic (M. oryzae) and biotrophic (Xoo) pathogen infection (Fig. 5E,F). These findings illustrate that the SA-signalling pathway is responsible for the enhanced resistance of OsACBP5-OEs to infection caused by the representative hemibiotroph and biotroph.
OsACBP5-OE transgenic rice plants showed protection against various plant pathogens via SA- and JA-induced defence responses. (A) Sheath blight symptoms after inoculation with the necrotroph R. solani on five-week-old WT, OsACBP5-OE line 9 (OE-9), JA-signalling deficient mutant oscoi1, oscoi1 in the OE-9 background (OE-9oscoi1), SA-signalling deficient mutant osnpr1 and osnpr1 in the OE-9 background (OE-9osnpr1) at 14 days post-inoculation (dpi). Blue bars = 1 cm. (B) Rice blast symptoms on 3-week-old WT, oscoi1, OE-9oscoi1, osnpr1 and OE-9osnpr1 inoculated with the hemibiotrophic fungal pathogen, M. oryzae 7 dpi. Blue bars = 5 mm. (C) Disease phenotype in 3-week-old WT, oscoi1, OE-9oscoi1, osnpr1 and OE-9osnpr1 inoculated with the bacterial biotroph Xoo 14 dpi. Blue bars = 5 mm. (D) Lesion length following R. solani infection; (E) lesion area after inoculation with M. oryzae; (F) lesion length following Xoo infection in the WT, OE-9, oscoi1, OE-9oscoi1, osnpr1 and OE-9osnpr1. Data points represent means ± SD from three independent experiments. Asterisks indicate significant difference (P < 0.05) in comparison to the controls by Student’s t-test.
Furthermore, GC-MS was performed to measure SA and JA content in R. solani-, M. grisea- and Xoo-infected WT, OE-9, osnpr1, oscoi1, OE-9osnpr1 and OE-9oscoi1. R. solani-infected oscoi1 and OE-9oscoi1 respectively showed 40- and three-fold lower JA content compared to the WT (Supplemental Fig. S2A). When qRT-PCR was performed, oscoi1 and OE-9oscoi1 respectively showed ~ ten- and three-fold lower expression of OsAOS1 in comparison to that of WT (Supplemental Fig. S3A). Similarly, M. grisea- and Xoo-infected osnpr1 and OE-9osnpr1 respectively showed ~ 20- and 2.5-fold lower SA content compared to the WT (Supplemental Fig. S2B,C). When qRT-PCR was performed, osnpr1 and OE-9osnpr1 respectively showed ~ ten- and three-fold lower expression of OsNPR1 compared to the WT (Supplemental Fig. S3B,C). These results further confirm that the improved resistance of OsACBP5-OEs to the necrotroph R. solani is JA-dependent and enhanced resistance of OsACBP5-OEs to hemibiotroph (M. grisea) and biotroph (Xoo) is SA-dependent.
Two W-boxes in the OsACBP5 5′-flanking region bind infected rice nuclear proteins
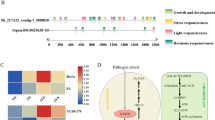
Given that OsACBP5 functions in plant defence, the 5′-flanking region of OsACBP5 was investigated using the PlantCARE62 (https://bioinformatics.psb.ugent.be/webtools/plantcare/html/) and PLACE63 (https://www.dna.affrc.go.jp/PLACE/) databases. Potential cis-elements identified in the OsACBP5 5′-flanking region include pathogen-responsive cis-elements such as the W-box64 (− 1713/− 1708, − 1,560/− 1555, − 413/− 408 and − 157/− 152), MeJA-responsive element CGTCA65 (− 1,620/− 1616, − 1,540/− 1536 and − 751/− 747) and seed-specific motifs such as Skn-166 (− 1,790/− 1786 and − 371/− 367) (Fig. 6A). EMSAs using crude nuclear extracts from R. solani-infected 5-week-old WT rice, C. oryzae-infected three-week-old WT rice, M. oryzae-infected three-week-old WT rice, Xoo-infected three-week-old WT rice and C. oryzae-infected three-week-old WT rice showed strong DNA–protein binding complexes with the W-boxes at − 1713/− 1708 and − 157/− 152 (Fig. 6B), indicating that two of the four putative W-boxes are essential in regulating OsACBP5 expression. In contrast, when the CGTCA and Skn-1 boxes were tested, they did not bind to nuclear extracts in EMSA (Supplemental Fig. S4).
Analysis of the OsACBP5 5′-flanking region. (A) A schematic diagram of constructs (pOS820, pOS891 and pOS895) developed by 5′-end deletion of the OsACBP5 5′-flanking region (− 1926/ + 304). Promoter fragments of different lengths were introduced into the binary vector DX2181 comprising the GUS reporter gene. Black bars (not to scale) represent each truncated fragment and the end position of each deletion is denoted on the left. Putative cis-elements (in black, blue and red symbols are labelled) on the OsACBP5pro::GUS construct pOS820. The cis-elements labelled in green have been verified experimentally. Forward and reverse arrows indicate the PCR primers used for generating constructs. (B) Interaction of the infected WT rice leaf nuclear extract (I) with the W-box (− 1713/− 1708 and − 157/− 152) probe. Nucleotide sequences of double-stranded oligonucleotides used in EMSAs are shown in bold. The protein-DNA complexes are indicated by red arrowheads. Lane 1, free probe without the addition of leaf nuclear extracts. Crude nuclear extracts from infected leaves were incubated with biotin end-labelled probes (lane 2) in the presence of a 500-fold molar excess of an unlabelled competitor (lane 3). Lane 4, a negative control with labelled probe and untreated leaf nuclear extract (U). I, R. solani-infected nuclear extract in panels i and v; M. oryzae-infected nuclear extract in panels ii and vi; Xoo-infected nuclear extract in panels iii and vii and C. oryzae-infected nuclear extract in panels iv and viii. Quantitative fluorometric measurement of GUS activity in (C) SA-treated, (D) MeJA-treated and (E) R. solani-infected OsACBP5pro::GUS constructs pOS820, pOS891 and pOS895 0 h, 5 h, 12 h and 24 h post-treatment. Five independent lines were used per construct. Data points represent means ± SD from three independent experiments.
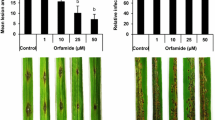
OsACBP5pro::GUS expression is induced by SA, MeJA and R. solani infection
To identify the cis-elements of the OsACBP5 5′-flanking region in SA-, methyl jasmonate (MeJA)- and pathogen-induced regulation, quantitative GUS assays were performed on three-week-old (T3-generation) transgenic rice pOS820 (2.2-kb OsACBP5pro::GUS), pOS891 (1.3-kb OsACBP5pro::GUS) and pOS895 (0.6-kb OsACBP5pro::GUS) transformants. When SA- (100 µM) and MeJA- (100 µM) treated three-week-old rice seedlings were analysed 0 h, 5 h, 12 h and 24 h post-treatment, pOS820, pOS891 and pOS895 transformants showed higher GUS activity 5 h post-treatment (Fig. 6C,D). However, the pOS820 transformants showed 1.7-fold and 2.4-fold increased GUS activity 5 h post-SA treatment over the pOS891 and pOS895 transformants, respectively (Fig. 6C). Similarly, the pOS820 transformants showed 1.6-fold and 2.5-fold increased GUS activity 5 h post-MeJA treatment over the pOS891 and pOS895 transformants, respectively (Fig. 6D). Likewise, when R. solani-infected rice seedlings were analysed, the pOS820 transformants showed twofold and threefold increased GUS activity over the pOS891 and pOS895 transformants, respectively (Fig. 6E), demonstrating that OsACBP5pro::GUS expression in seedlings was induced by SA, MeJA and R. solani treatment. Reduction in GUS activity in the pOS891 and pOS895 transformants suggested that the W-boxes (− 1713/− 1708 and − 157/− 152) play an important role in the regulation of OsACBP5.
Recombinant OsACBP5 binds 18:3-acyl-CoA ester
Lipidex assays by Meng et al. have shown the binding of (His)6-tagged OsACBP5 to 18:3-acyl-CoA esters28. As 18:3-FA is important for basal defence against fungal pathogens and is a precursor for JA biosynthesis67, the binding affinity of (His)6-OsACBP5 to 18:3-acyl-CoA ester was investigated by isothermal titration calorimetry (ITC) which provides a more precise method to measure protein-ligand binding than Lipidex assays20. Consistent with Lipidex assays, recombinant OsACBP5 (rOsACBP5) was shown to bind to 18:3-acyl-CoA with high affinities (Supplemental Fig. S5). ITC results (Supplemental Table S1) indicated that rOsACBP5 has a strong binding affinity to 18:3-acyl-CoA ester with a dissociation constant (Kd) value of 59.5 nM.
When OsACBP5-OE leaves were further examined using GC-MS to test the level of the six major FA species (14:0-, 16:0-, 18:1-, 18:2-, 18:3- and 20:0-FAs), the three most abundant species were 16:0-, 18:2- and 18:3-FAs (Supplemental Fig. S6). OsACBP5-OEs (OE-1, OE-3, OE-6, OE-9 and OE-11) showed twofold higher 18:3-FA content in leaves than the wild-type and vector-transformed controls (Fig. S6). However, no significant differences were detected for 14:0-, 16:0-, 18:1-, 18:2-, and 20:0-FAs between the OsACBP5-OEs and the controls (Fig. S6).
Rice genes are differentially expressed between OsACBP5-OEs and the wild type in response to R. solani infection
When transcriptomic analysis was performed on R. solani-infected transgenic rice OsACBP5-OEs, a total of 22,063 (15,253 up-regulated and 6,810 down-regulated) differentially expressed genes (DEGs) were identified between OsACBP5-OEs and the wild type control. Sixteen genes upregulated in the plant-pathogen interaction pathway (Kyoto Encyclopedia of Genes and Genomes (KEGG) map 04626) in OsACBP5-OEs following R. solani infection were CYCLIC NUCLEOTIDE GATED CHANNELS (CNGCS), CALCIUM-DEPENDENT PROTEIN KINASE (CDPK), CALMODULIN/CALMODULIN-LIKE PROTEINS (CAM/CML), RESPIRATORY BURST OXIDASE HOMOLOG (RBOH), NITRIC OXIDE SYNTHASE (NOS), FLAGELLIN-SENSING2 (FLS2), MITOGEN-ACTIVATED PROTEIN KINASE KINASE1/2 (MKK1/2), MITOGEN-ACTIVATED PROTEIN KINASE KINASE4/5 (MKK4/5), WRKY TRANSCRIPTION FACTOR22 (WRKY22), WRKY TRANSCRIPTION FACTOR33 (WRKY33), DISEASE RESISTANT PROTEINS (RPM1, RPS2, RAR1), RPM1-INTERACTING PROTEIN4 (RIN4), SUPPRESSOR OF G2 ALLELE OF SKP1 (SGT1) and HEAT SHOCK PROTEIN90 (HSP90) (Fig. 7A). Ten DEGs related to the PAMP-triggered immunity (PTI) signalling pathway include those encoding CNGCs, CDPK, CaM/CML, RbOH, NOS, FLS2, MKK1/2, MKK4/5, WRKY22 and WRKY33 (Fig. 7A). The six DEGs upregulated in the effector-triggered immunity (ETI) signalling pathway were RPM1, RPS2, RAR1, RIN4, SGT1 and HSP90 (Fig. 7A).
DEGs associated with the plant-pathogen interaction pathway as well as SA- and JA-signalling pathways in R. solani-infected OsACBP5-OEs. The KEGG database148,149,150 was used for pathway analysis. (A) Increase in the cytosolic Ca2+ concentration by the activation of CNGCs, CDPK and CaM/CML in PTI, is a regulator for production of ROS and NOS which results in the hypersensitive response121,122. Activation of FLS2 in PTI triggers the MAPK signalling pathway that induces known defence genes for the generation of antimicrobial compounds such as phytoalexins, camalexin and lignin123,124. Pathogen infection induces RIN4 in ETI, which activates the disease resistant proteins RPM1 and RPS292. RPM1 and RPS2 then trigger a complex formed by HSP90, RAR1 and SGT1 leading to the hypersensitive response96,97,98. SGT1 also regulates early R gene-mediated plant defences upon pathogen infection125. Up-regulated genes are boxed in red, genes those are not affected are boxed in grey. Those marked with asterisks in this figure have been previously discussed126. (B) SA triggers the accumulation NPR1 which activates the TGA transcription factor and PR1 resulting in plant defence127,128. Activation of JAR1 following pathogen infection catalyses the production of JA-Ile from JA. Production of JA-Ile is crucial for the JA-signalling pathway involving COI1, JAZ and MYC2 leading to plant defence47,129. Up-regulated genes are boxed in red.
Furthermore, seven DEGs related to the SA- and JA-signalling pathways were up-regulated in R. solani infected OsACBP5-OEs (Fig. 7B). In the JA signalling pathway, four DEGs encoding JASMONOYL ISOLEUCINE CONJUGATE SYNTHASE1 (JAR1), CORONATINE INSENSITIVE PROTEIN1 (COI1), JASMONATE ZIM-DOMAIN CONTAINING PROTEIN (JAZ) and TRANSCRIPTION FACTOR MYC2 were induced upon fungal infection (Fig. 7B). Similarly, three DEGs in the SA-signalling pathway such as NON-EXPRESSOR OF PATHOGENESIS-RELATED1 (NPR1), TRANSCRIPTION FACTOR TGA and PATHOGENESIS-RELATED PROTEIN1 (PR1) were up-regulated following pathogen invasion of OsACBP5-OEs leading to disease resistance (Fig. 7B). Table S2 shows fold changes of DEGs associated with the plant-pathogen interaction pathway as well as SA- and JA-signalling pathways in R. solani-infected OsACBP5-OEs.
Biotic stress-related proteins were induced in OsACBP5-OEs by R. solani infection
When SWATH-MS quantitative proteomic analysis was carried out to explore the effect of OsACBP5 action on R. solani infection, ProteinPilot software identified 1,365 proteins, 2,754 peptides, and 12,390 spectra with 99% confidence and 1% global false discovery rate (FDR). Of 1,365 identified proteins, 419 were significantly upregulated in rice OsACBP5-OEs versus WT and vector-transformed plants (P < 0.05). Consistent with transcriptomics data, proteins involved in the plant-pathogen interaction pathway (CDPK, FLS2, RPM1, RPS2 and HSP90), JA-signalling pathway (JAZ and MYC2) and SA-signalling pathway (PR1) were upregulated following R. solani infection in OsACBP5-OEs (Table 1).
When qRT-PCR was performed on OsACBP5-OEs to validate the results from transcriptomic and proteomic analyses, increased expression of genes involved in the plant-pathogen interaction pathway (CNGCs, CDPK, CaM/CML, RbOH, NOS, FLS2, MKK1/2, MKK4/5, WRKY22, WRKY33, RPM1, RPS2, RAR1, RIN4, SGT1 and HSP90), JA-signalling pathway (JAR1, COI1, JAZ and MYC2) and SA-signalling pathway (NPR1, TGA and PR1) in OsACBP5-OEs (Supplemental Figs. S7–S9) supported the transcriptomic and proteomic data. The expression level of ACTIN in R. solani-infected WT, VC and OsACBP5-OEs is shown in Supplemental Figure S10. Taken together, findings from transcriptomics and proteomics suggest that OsACBP5 plays a crucial role in the protection of plants against R. solani through activating plant-pathogen interaction pathway as well as JA- and SA-signalling pathways.
Discussion
OsACBP5 conferred broad-spectrum defence against phytopathogens
In this study, the function of OsACBP5 in plant defence was established by phenotypic analyses of five independent rice OsACBP5-OE lines in response to representative necrotrophic, hemibiotrophic and biotrophic pathogens. Previous work on transgenic Arabidopsis overexpressing its homologue, AtACBP3, had shown that AtACBP3 could confer NONEXPRESSOR OF PR GENES1 (NPR1)-dependent resistance to bacterial biotroph P. syringae, with increased susceptibility to the fungal necrotroph B. cinerea17. In contrast, this study revealed that transgenic rice OsACBP5-OEs displayed enhanced tolerance to necrotrophic fungal pathogens such as R. solani and C. oryzae (Fig. 1), hemibiotrophic fungal pathogens, M. oryzae and F. graminearum (Fig. 2) and a biotrophic bacterial pathogen, Xoo (Fig. 3). These findings demonstrated that OsACBP5 is more versatile against pathogens in transgenic rice. As Takato et al. had earlier illustrated that the overexpression of a Class III ACBP from grape (Vitis vinifera) could protect transgenic Arabidopsis against the biotroph P. syringae and a hemibiotroph C. higginsianum30, it appears that the Class III ACBPs are promising targets for disease prevention in both transgenic dicots and monocots. Similar to OsACBP5 in exhibiting broad-spectrum properties in defence, wide-range protection against R. solani, M. oryzae and Xoo have been reported in transgenic rice overexpressing a cysteine-rich antimicrobial defensin from Allium cepa (Ace-AMP), but the molecular regulation on its action is less understood68. In comparison, defence-related proteins such as the rice wall-associated kinase (OsWAK25) and MoSM1, encoding a cerato-platanin protein from M. oryzae, when overexpressed in rice, conferred protection only against the hemibiotroph M. oryzae and the bacterial biotroph Xoo, but displayed increased susceptibility to the fungal necrotroph R. solani57,69. Correspondingly, the expression of OsWRKY13, encoding transcription factor WRKY, was induced by M. oryzae and Xoo infection58,86. The constitutive expression of OsWRKY13 displayed protection to M. oryzae and Xoo via the SA-signalling pathway58,89. Likewise, M. oryzae and Xoo infection induced the expression of rice DEFENCE-RESPONSE PROTEIN8 (OsDR8), encoding a protein involved in thiamine biosynthesis58,86. OsDR8 accumulates thiamine and confers systemic acquired resistance (SAR) against M. oryzae and Xoo58,140. Similarly, the constitutive expression of rice INDOLE-3-ACETIC ACID (IAA) AMIDO SYNTHETASE (GH3-8), whose expression is induced by auxin, enhanced resistance to M. oryzae and Xoo infection in rice by suppressing pathogen-induced IAA accumulation58,141. Other rice genes that promote similar pathogen resistance are summarised in Table 2.
Rice OsACBP5-OEs showed JA-mediated response against necrotrophs and SA-mediated response against (hemi)biotrophs
SA and JA, the two critical defence signalling hormones that play vital roles against necrotrophic, hemibiotrophic and biotrophic pathogens in rice61,70, were observed to accumulate in rice OsACBP5-OEs (Fig. 4A,B). The upregulated expression of OsNPR1, an SA-signalling regulatory gene, and OsAOS2 encoding allene oxide synthase in JA biosynthesis, in OsACBP5-OEs (Fig. 4C,D) likely stimulates SA- and JA-mediated defence responses. Furthermore, results from bioassays on transgenic rice OsACBP5-OE9 in oscoi1 and osnpr1 backgrounds suggest that necrotrophic resistance in rice OsACBP5-OEs is JA-dependent and (hemi)biotrophic resistance is SA-dependent (Fig. 5). A recent study has shown that transgenic Arabidopsis overexpressing OsACBP5 were conferred resistance to necrotrophic (R. solani, B. cinerea, A. brassicicola), hemibiotrophic (C. siamense) and biotrophic (P. syringae) phytopathogens59,60. Proteomic analysis on the R. solani-infected transgenic Arabidopsis OsACBP-OEs showed upregulation of biotic stress-related proteins including cell wall-related proteins such as FASCILIN-LIKE ARABINOGALACTAN-PROTEIN10, LEUCINE-RICH REPEAT EXTENSIN-LIKE PROTEINS, XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE PROTEIN4 and PECTINESTERASE INHIBITOR18; proteins associated with glucosinolate degradation including GDSL-LIKE LIPASE23, EPITHIOSPECIFIER MODIFIER1, MYROSINASE1, MYROSINASE2 and NITRILASE1; as well as a protein involved in jasmonate biosynthesis, ALLENE OXIDE CYCLASE259,60. These results from proteomic analysis indicated that the defence responses arising from OsACBP5 overexpression in transgenic Arabidopsis involved cell wall-mediated defence as well as salicylic acid (SA)- and jasmonic acid (JA)-mediated defence pathways59,60. Similarly, transgenic Arabidopsis overexpressing AtACBP3 displayed enhanced SA-mediated resistance to the biotrophic pathogen P. syringae17.
The current results on transgenic rice OsACBP5-OEs also demonstrated the cooperation between the SA and JA pathways in defence against representative necrotrophs (R. solani, C. oryzae), hemibiotrophs (M. oryzae, F. graminearum) and biotrophs (Xoo). In contrast, the SA and JA defence signalling pathways generally interact antagonistically in dicots51,53,55,71,72,73,74,75,76,77,78, while such interaction is not well investigated in monocots such as rice56,79. Nonetheless, Tamaoki et al. reported that SA and JA can collaboratively stimulate a common defence signalling system in rice against pathogens80. Similar to the present study, transgenic MoSM1-OE rice displayed improved resistance to the hemibiotroph M. oryzae and the biotroph Xoo accompanied by elevated SA and JA content and upregulated expression of SA- and JA-signalling genes57. SA and JA accumulation in rice OsACBP5-OEs may have arisen from the ability of OsACBP5 in binding to 18:3-acyl-CoA ester because ITC data supported rOsACBP5 binding to 18:3-acyl-CoA ester with a Kd value in the nanomolar range (Fig. S5). Interestingly, OsACBP5-OEs showed higher linolenic acid (18:3) content than the controls (Fig. S6) and 18:3-FA is a precursor for JA biosynthesis67. Previous reports have demonstrated that the Arabidopsis ssi2 mutant contains lower 18:3-FA content than the WT and was more susceptible to necrotroph B. cinerea infection82. These results resonate well with the current study which revealed that decreased susceptibility of OsACBP5-OEs to various pathogens, in comparison to the WT, is associated with increase in 18:3-FA content.
The physiological significance in the role of AtACBP3 in trafficking lipids such as acyl-CoA esters was evident in transgenic Arabidopsis overexpressing AtACBP3 which displayed accelerated leaf senescence, in contrast to an atacbp3 T-DNA insertional mutant and AtACBP3 RNA interference (RNAi) transgenic Arabidopsis lines which were delayed in dark-induced leaf senescence15. Subsequent acyl-CoA and lipid profiling revealed that AtACBP3 overexpression culminated in the accumulation of acyl-CoA and phosphatidylethanolamine (PE), while the downregulation of AtACBP3 reduced PE15. In dark-treated and premature senescing AtACBP3-OE plants, PC and phosphatidylinositol levels declined accompanied by increases in PA, lysophospholipids, and oxylipin-containing galactolipids (arabidopsides). It was concluded that the accumulation of PA and arabidopsides (A, B, D, E, and G) resulting from lipid peroxidation in AtACBP3-OEs likely caused leaf senescence15. In another study, it was reported that oxylipin-related FA (18:2-FA, 18:3-FA and MeJA) content was lower in atacbp3 and AtACBP3-RNAi than wild-type phloem exudates upon GC-MS analysis28. On ITC analysis, recombinant AtACBP3 was shown to bind medium- and long-chain acyl-CoA esters with KD values in the micromolar range28. Hu et al. concluded that the phloem-mobile AtACBP3 likely affected the FA pool and JA content in the phloem by its binding to acyl-CoA esters, ultimately influencing the level of oxylipins, which are crucial components of the plant wound responses mobilized via the vasculature28.
Significance of W-boxes in regulating pathogen-inducible OsACBP5 expression
The WRKY family of TFs that regulate the transcription of plant defence genes through the W-box84, are crucial in protection against necrotrophic, hemibiotrophic and biotrophic pathogens84,85,86,87,88,89,90,91,92,93,94. For example, OsWRKY4 binds to the W-boxes in the 5′-flanking region in each of pathogenesis-related PR1b and PR5, and OsWRKY4 and OsWRKY80 were reported to be highly induced by R. solani infection91,95. Wang et al. also showed that transgenic rice overexpressing OsWRKY4 were protected against R. solani infection95. In this study, OsACBP5-OEs were proven tolerant to representative necrotrophs, hemibiotrophs and biotrophic phytopathogens and EMSAs revealed that only two of the four W-boxes (-1713/-1708 and -157/-152) in the 5′-flanking region of OsACBP5 regulate OsACBP5 expression during representative necrotrophic (Fig. 6B panels i, iv, v and viii), hemibiotrophic (Fig. 6B panels ii and vi) and biotrophic (Fig. 6B panels iii and vii) infection. These results correspond well with quantitative GUS assays on the pOS820 (2.2-kb OsACBP5pro::GUS; -1926/ + 304) transformants which displayed induced GUS expression after treatment with the pathogen-related phytohormones, SA and MeJA, in comparison to transformants from constructs that lacked either of these W-boxes, pOS891 (1.3-kb OsACBP5pro::GUS; − 1,281/ + 304) and pOS895 (0.6-kb OsACBP5pro::GUS; − 46/ + 304) (Fig. 6C,D). These results verified that the two W-boxes (− 1713/− 1708 and − 157/− 152) play a role in regulating OsACBP5 expression in response to necrotrophic, hemibiotrophic and biotrophic pathogens, as well as to the pathogen-related phytohormones, SA and MeJA.
Previous results suggest that increased protection to R. solani and M. oryzae in transgenic rice overexpressing OsWRKY30 was associated with elevated levels of JA, as well as the stimulated expression of JA synthesis-related genes (LOX and AOS2) and pathogenesis-related PR3 and PR10, following fungal pathogen infection90. Furthermore, Hiroyuki and Terauchi revealed that the W-boxes in the RICE THAUMATIN-LIKE PROTEIN1 (RTLP1) promoter function in response to M. oryzae infection96. Also, past investigations on the development of resistance in rice against the rice blast pathogen M. oryzae unveiled a critical role for WRKY TFs (OsWRKY45, OsWRKY13 and OsWRKY42) in plant defence92. OsWRKY45 has been assigned a vital role in SA-mediated signalling in rice against the hemibiotrophic pathogen M. oryzae84. Enhanced resistance of transgenic rice to the biotrophic pathogen Xoo and the hemibiotroph M. oryzae was achieved by OsWRKY13 overexpression that was related to activation of SA-signalling and suppression of the JA-dependent pathway89. Similar to OsACBP5, where W-boxes were observed to bind nuclear extracts from Xoo-infected rice, W-boxes in rice STRESS RESPONSIVE NAC1 (SNAC1) interacted with Xoo-treated nuclear proteins and OsWRKY13 was subsequently identified to regulate SNAC1 expression during biotic stress97. Furthermore, the overexpression of OsWRKY13 or OsWRKY71 culminated in better tolerance to Xoo in transgenic rice87,98. Taken together these studies support a role for OsWRKY TFs in biotrophic, hemibiotrophic and necrotrophic fungal tolerance in rice via the SA- and JA-defence signalling pathways.
Several defence-related genes were upregulated in R. solani-infected OsACBP5-OEs
The role of SA and JA in hemi(bio)trophic and necrotrophic pathogen defence in transgenic rice OsACBP5-OEs was partially confirmed from pathogen assays, GC-MS and qRT-PCR. Transcriptomic and proteomic analyses further confirmed the mechanism of defence in transgenic rice OsACBP5-OEs. Although transcriptomic and proteomic assays were performed only on necrotrophic pathogen R. solani-infected transgenic rice OsACBP5-OEs, the upregulated genes and proteins from these assays were reported to be involved in defence against necrotrophic, hemibiotrophic and biotrophic phytopathogens, which are discussed in this section.
Transcriptomics and proteomics data provided an insight into the defence responses of transgenic rice OsACBP5-OEs to the necrotrophic pathogen R. solani infection. The innate immunity in plants appeared to be triggered through PTI followed by ETI, providing the first line of defence upon pathogen challenge99,100. Ten genes involved in PTI were up-regulated in OsACBP5-OEs upon R. solani infection (Fig. 7A). Cytoplasmic Ca2+ concentration increases during PTI leading to the activation of CDPK in plant cells101. In this study, three genes involved in Ca2+ signalling were up-regulated in R. solani-infected OsACBP5-OEs including CNGCs, CDPK, CaM/CML. Calcium signalling was reportedly accompanied by an increase of both ROS and NO leading to SA-mediated defence30,101. Transcription factors WRKY22 and WRKY33 were activated by components of the MAPK cascade such as MEKK1, MKK1/2 and MKK4/5, resulting in induced expression of defence-related genes in R. solani-infected OsACBP5-OEs (Fig. 7A). Similar results were observed in Xoo-infected rice plants in which FLS2 perceived bacterial flagellin and activated the MAPK cascade which in turn activated WRKY22 and WRKY33 resulting in induced expression of defence-related genes92,99,100. Taken together, ROS production and activation of MAPKs and CDPKs cause an array of defences restricting pathogen progression.
In plants, a secondary immune response ETI is the basis for a second layer of defence99,100. The second signalling pathway consists of five genes encoding receptor proteins (RIN4, PBS1, RPM1, SGT1 and RAR1) to perceive pathogen infection. In this study, four such genes encoding receptor proteins including RIN4, RPM1, SGT1 and RAR1 displayed up-regulation in OsACBP5-OEs following R. solani infection (Fig. 7A). RPM1 recognizes modifications of RIN4 followed by P. syringae infection in Arabidopsis and RPM1 interacts with RIN4 triggering RPM1-mediated immunity102. RAR1 and SGT1 conferred resistance against Xoo and M. oryzae when overexpressed in rice103,104. RAR1 forms a complex with the molecular chaperones HSP90 and SGT1 to initiate a signalling cascade in diverse plant immune responses105,106,107,108. In this study, the upregulated expression of various components (HSP90, RAR1 and SGT1) of the complex likely caused a hypersensitive response in OsACBP5-OEs following R. solani infection (Fig. 7A). Previous studies have reported that hypersensitive responses are mostly accompanied by an increase in SA biosynthesis109,110.
Furthermore, several DEGs (NPR1, TGA, PR1, JAR1, COI1, JAZ and MYC2) involved in the SA- and JA-signalling pathways were enriched in R. solani-infected OsACBP5-OEs (Fig. 7B), suggesting that both pathways are involved (Fig. 7B), supporting the role of SA and JA against R. solani.
Conclusions
The present study demonstrates that OsACBP5 is effective and activates defence responses in transgenic rice against various representative necrotrophic, hemibiotrophic and biotrophic pathogens. Transgenic rice OsACBP5-OEs showed higher SA and JA levels when compared to the WT and vector-transformed control, suggesting that both SA- and JA-mediated signalling pathways are activated in OsACBP5-OEs. These results demonstrate that OsACBP5 overexpression in rice effectively conferred broad-spectrum resistance against several phytopathogens (R. solani, C. oryzae, M. oryzae, F. graminearum and Xoo), providing a potential for OsACBP5 in enhancing disease resistance in crop plants.
Methods
Plant materials and growth conditions
T-DNA insertion mutants osnpr1 and oscoi1 were purchased from Rice T-DNA Insertion Sequence Database (RISD DB; cbi.khu.ac.kr/RISD_DB.html). Plasmid vectors, pCAMBIA1304 and DX2181, were obtained from Shanghai Normal University and Huazhong Agricultural University, respectively. T3-generation seeds of transgenic rice derived from this study including vector-transformed controls (pCAMBIA1304 and DX2181), OsACBP5-OEs, OsACBP5pro::GUS, OsACBP5-OE9osnpr1 and OsACBP5-OE9oscoi1 as well as osnpr1, oscoi1, and Oryza sativa cv Zhonghua11 wild-type (WT) seeds (ten seeds were used for each line per experiment) were surface-sterilized with 70% ethanol for 5 min followed by 3% sodium hypochlorite solution for 40 min. The seeds were then washed with distilled water 5 times and germinated on half-strength MS115 medium containing 3% sucrose for 1 week at 28 °C. One-week-old seedlings (five seedlings for each line per experiment) were transferred to clay soil in separate pots in a growth chamber under a 12 h light (28 °C)/12 h dark (25 °C) photoperiod116. Supporting Information provides details on the generation of transgenic rice (OsACBP5-OEs, OsACBP5-OE9osnpr1, OsACBP5-OE9oscoi1, OsACBP5pro::GUS fusion and its deletion derivatives), pathogen assays, phytohormone treatments, fluorometric assays of GUS activity, electrophoretic mobility shift assays (EMSAs), isothermal titration calorimetry (ITC) experiments and Quantitative Real Time-Polymerase Chain Reaction (qRT-PCR).
Quantification of SA and JA
SA and JA quantification was performed following Fina et al.116. Leaf tissue (300 mg) was homogenized and SA extracted in 80% methanol by shaking for 16 h at − 20 °C. The samples were then purified on a C18 cartridge (Bond Elut C18 6 cc, 500 mg, Agilent, CA, USA) in 80% methanol. Formic acid was added for the binding of SA to the cartridge. The SA was eluted with diethyl ether. The eluent was evaporated under nitrogen gas after removing the residual water. The sample was further methylated using diazomethane and dried under nitrogen gas. The sample was subsequently dissolved in 100% hexane for GC-MS analysis. The same protocol was followed for JA quantification. SA (10 µM) and JA (10 µM) were used as internal standards.
Expression and purification of OsACBP5
The (His)6-OsACBP5 recombinant protein was expressed in the soluble fraction of Escherichia coli BL21(DE3) Star pLysS (Invitrogen) cells transformed with plasmid pOS543, derived from vector pRSETA (Life Technologies) following Meng et al. (2011). (His)6-OsACBP5 was purified by using a HisTrap HP column (GE Healthcare) charged with 0.1 M NiCl2 according to Guo et al.117.
Transcriptome analysis
Total RNA was extracted from R. solani-infected WT, vector (pCAMBIA1304)-transformed control and transgenic rice OsACBP5-OEs using the RNeasy Plant Mini Kit (Qiagen). RNAs samples were sequenced using BGISEQ-500 sequencer at Beijing Genomics Institute (BGI, Hong Kong). RNA concentration and quality were measured using Agilent 2100 Bio analyser (Agilent RNA 6000 Nano Kit). The BGISEQ-500 platform was used to sequence the cDNA libraries. SOAPnuke was used to filter reads and after filtering, the clean reads were stored in FASTQ format. The clean reads were mapped using Bowtie2 (https://bowtie-bio.sourceforge.net/Bowtie2/index.shtml) and the gene expression level was calculated using RSEM (https://deweylab.biostat.wisc.edu/RSEM). Differentially expressed genes (DEGs) were detected using DEGseq software based on the Poisson distribution. The KEGG database148,149,150 was used for pathway analysis. The open reading frame (ORF) of each DEG was identified using GETORF database. To predict the transcription factor of each DEG, the ORF was aligned to transcription factor domains using the PLNTFdb database. DEGs were mapped to the PRGdb database using BLAST to detect plant disease resistance genes.
Sequential window acquisition of all theoretical mass spectra quantitative proteomic analysis
The trichloroacetic acid/acetone method was used for proteins extraction following Wu et al.118. The protein pellet was resuspended in 2 mL urea buffer (6 M urea and 4 mM calcium chloride in 200 mM 3-(N-morpholino) propanesulfonic acid (MOPS), pH 8.0)120. An equivalent amount of protein (100 μg) was reduced using 10 mM dithiothreitol (DTT) and alkylated in 40 mM iodoacetamide (IAA) in the dark. After alkylation, the concentration of urea in the mixture was reduced to less than 2 M by diluting with 4 mM CaCl2. The protein was digested with trypsin (1:20) followed by incubation at 37 °C overnight. Subsequently, the peptides were desalted utilising C18 SepPak reverse-phase cartridges and SWATH-MS analysis was performed121. The data was analysed from five biological repeats.
Statistical analysis
Significant differences in data between different samples were analyzed by the Student’s t-test.
References
Savary, S., Ficke, A., Aubertot, J. N. & Hollier, C. Crop losses due to diseases and their implications for global food production losses and food security. Food Secur. 4, 519–537 (2012).
Casadevall, A. & Pirofski, L. A. Host-pathogen interactions: redefining the basic concepts of virulence and pathogenicity. Infect. Immun. 67, 3703–3713 (1999).
Lewis, D. H. Concepts in fungal nutrition and the origin of biotrophy. Biol. Rev. 48, 261–277 (1973).
Laluk, K. & Mengiste, T. Necrotroph attacks on plants: wanton destruction or covert extortion?. Arabidopsis Book 8, e0136 (2010).
Perfect, S. E. & Green, J. R. Infection structures of biotrophic and hemibiotrophic fungal plant pathogens. Mol. Plant Pathol. 2, 101–108 (2001).
Van Bruggen, A. H. C. & Finckh, M. R. Plant diseases and management approaches in organic farming systems. Annu. Rev. Phytopathol. 54, 25–54 (2016).
Singh, R. P. et al. Disease impact on wheat yield potential and prospects of genetic control. Annu. Rev. Phytopathol. 54, 303–322 (2016).
Cichy, K. A., Snapp, S. S. & Kirk, W. W. Fusarium root rot incidence and root system architecture in grafted common bean lines. Plant Soil 300, 233–244 (2007).
Narayanasamy, P. Biological Management of Diseases of Crops (Springer, Netherlands, 2013).
Murray, G. M. & Brennan, J. P. Estimating disease losses to the Australian wheat industry. Australas. Plant Pathol. 38, 558–570 (2009).
Wang, X., Jiang, N., Liu, J., Liu, W. & Wang, G. L. The role of effectors and host immunity in plant–necrotrophic fungal interactions. Virulence 5, 722–732 (2014).
Xiao, S. & Chye, M. L. New roles for acyl-CoA-binding proteins (ACBPs) in plant development, stress responses and lipid metabolism. Prog. Lipid Res. 50, 141–151 (2011).
Chen, Q. F., Xiao, S. & Chye, M. L. Overexpression of the Arabidopsis 10-kilodalton acyl-coenzyme A-binding protein ACBP6 enhances freezing tolerance. Plant Physiol. 148, 304–315 (2008).
Xiao, S., Gao, W., Chen, Q. F., Ramalingam, S. & Chye, M. L. Overexpression of membrane-associated acyl-CoA-binding protein ACBP1 enhances lead tolerance in Arabidopsis. Plant J. 54, 141–151 (2008).
Xiao, S. et al. Overexpression of Arabidopsis acyl-CoA-binding protein ACBP3 promotes starvation-induced and age-dependent leaf senescence. Plant Cell 22, 1463–1482 (2010).
Gao, W., Xiao, S., Li, H. Y., Tsao, S. W. & Chye, M. L. Arabidopsis thaliana acyl-CoA-binding protein ACBP2 interacts with heavy-metal-binding farnesylated protein AtFP6. New Phytol. 181, 89–102 (2009).
Xiao, S. & Chye, M. L. Overexpression of Arabidopsis ACBP3 enhances NPR1-dependent plant resistance to Pseudomonas syringae pv tomato DC3000. Plant Physiol. 156, 2069–2081 (2011).
Du, Z. Y., Chen, M. X., Chen, Q. F., Xiao, S. & Chye, M. L. Arabidopsis acyl-CoA-binding protein ACBP1 participates in the regulation of seed germination and seedling development. Plant J. 74, 294–309 (2013).
Du, Z. Y., Chen, M. X., Chen, Q. F., Xiao, S. & Chye, M. L. Overexpression of Arabidopsis acyl-CoA-binding protein ACBP2 enhances drought tolerance. Plant Cell Environ. 36, 300–314 (2013).
Du, Z. Y., Chen, M. X., Chen, Q. F., Gu, J. D. & Chye, M. L. Expression of Arabidopsis acyl-CoA-binding proteins AtACBP1 and AtACBP4 confers Pb (II) accumulation in Brassica juncea roots. Plant Cell Environ. 38, 101–117 (2015).
Du, Z. Y., Arias, T., Meng, W. & Chye, M. L. Plant acyl-CoA-binding proteins: an emerging family involved in plant development and stress responses. Prog. Lipid Res. 63, 165–181 (2016).
Liao, P., Chen, Q. F. & Chye, M. L. Transgenic Arabidopsis flowers overexpressing acyl-CoA-binding protein ACBP6 are freezing tolerant. Plant Cell Physiol. 55, 1055–1071 (2014).
Hsiao, A. S. et al. Arabidopsis cytosolic acyl-CoA-binding proteins ACBP4, ACBP5 and ACBP6 have overlapping but distinct roles in seed development. Biosci. Rep. 34, 865–877 (2014).
Hsiao, A. S., Yeung, E. C., Ye, Z. W. & Chye, M. L. The Arabidopsis cytosolic acyl-CoA-binding proteins play combinatory roles in pollen development. Plant Cell Physiol. 56, 322–333 (2015).
Ye, Z. W., Xu, J., Shi, J., Zhang, D. & Chye, M. L. Kelch-motif containing acyl-CoA-binding proteins AtACBP4 and AtACBP5 are differentially expressed and function in floral lipid metabolism. Plant Mol. Biol. 93, 209–225 (2017).
Lung, S. C. et al. Acyl-CoA-binding protein ACBP1 modulates sterol synthesis during embryogenesis. Plant Physiol. 174, 1420–1435 (2017).
Lung, S. C. et al. Arabidopsis ACYL-COA-BINDING PROTEIN1 interacts with STEROL C4-METHYL OXIDASE1-2 to modulate gene expression of homeodomain-leucine zipper IV transcription factors. New Phytol. 218, 183–200 (2018).
Hu, T. H., Lung, S. C., Ye, Z. W. & Chye, M. L. Depletion of Arabidopsis ACYL-COA-BINDING PROTEIN3 affects fatty acid composition in the phloem. Front. Plant Sci. 9, 2 (2018).
Xia, Y. et al. Acyl-CoA-binding proteins are required for cuticle formation and plant responses to microbes. Front. Plant Sci. 3, 224–242 (2012).
Takato, H., Shimidzu, M., Ashizawa, Y., Takei, H. & Suzuki, S. An acyl-CoA-binding protein from grape that is induced through ER stress confers morphological changes and disease resistance in Arabidopsis. J. Plant Physiol. 170, 591–600 (2013).
Meng, W., Su, Y. C., Saunders, R. M. & Chye, M. L. The rice acyl-CoA-binding protein gene family: phylogeny, expression and functional analysis. New Phytol. 189, 1170–1184 (2011).
Yaeno, T., Matsuda, O. & Iba, K. Role of chloroplast trienoic fatty acids in plant disease defence responses. Plant J. 40, 931–941 (2004).
Mur, L. A. et al. Biphasic ethylene production during the hypersensitive response in Arabidopsis: a window into defence priming mechanisms?. Plant Signal. Behav. 4, 610–613 (2009).
Kachroo, A. & Kachroo, P. Fatty acid-derived signals in plant defence. Annu. Rev. Phytopathol. 47, 153–176 (2009).
Garcion, C. & M´etraux, J. P. Salicylic acid in Plant Hormone Signaling: Annual Plant Reviews, (ed. PHedden, SG Thomas), 24:229–55 (Oxford: Blackwell, 2006)
Pieterse, C. M., Van der Does, D., Zamioudis, C., Leon-Reyes, A. & Van Wees, S. C. Hormonal modulation of plant immunity. Annu. Rev. Cell Dev. Biol. 28, 489–521 (2012).
Dempsey, D. A. & Klessig, D. F. How does the multifaceted plant hormone salicylic acid combat disease in plants and are similar mechanisms utilized in humans?. BMC Biol. 15, 23 (2017).
Rekhter, D. et al. Isochorismate-derived biosynthesis of the plant stress hormone salicylic acid. Science 365, 498–502 (2019).
Mishina, T. E. & Zeier, J. Pathogen-associated molecular pattern recognition rather than development of tissue necrosis contributes to bacterial induction of systemic acquired resistance in Arabidopsis. Plant J. 50, 500–513 (2007).
Dong, X. NPR1, all things considered. Curr. Opin. Plant Biol. 7, 547–552 (2004).
Moore, J. W., Loake, G. J. & Spoel, S. H. Transcription dynamics in plant immunity. Plant Cell 23, 2809–2820 (2011).
Van Loon, L. C., Rep, M. & Pieterse, C. M. J. Significance of inducible defense-related proteins in infected plants. Annu. Rev. Phytopathol. 44, 135–162 (2006).
Gfeller, A., Dubugnon, L., Liechti, R. & Farmer, E. E. Jasmonate biochemical pathway. Sci. Signal. 3, cm3 (2010).
Seo, H. S. et al. Jasmonic acid carboxyl methyltransferase: a key enzyme for jasmonate-regulated plant responses. Proc. Natl. Acad. Sci. 98, 4788–4793 (2001).
Staswick, P. E. & Tiryaki, I. The oxylipin signal jasmonic acid is activated by an enzyme that conjugates it to isoleucine in Arabidopsis. Plant Cell 16, 2117–2127 (2004).
Xie, D. X., Feys, B. F., James, S., Nieto-Rostro, M. & Turner, J. G. COI1: an Arabidopsis gene required for jasmonate-regulated defense and fertility. Science 280, 1091–1094 (1998).
Bari, R. & Jones. J. D. Role of plant hormones in plant defence responses. Plant Mol. Biol. 69, 473–488 (2009).
Katsir, L., Schilmiller, A. L., Staswick, P. E., He, S. Y. & Howe, G. A. COI1 is a critical component of a receptor for jasmonate and the bacterial virulence factor coronatine. Proc. Natl. Acad. Sci. 105, 7100–7105 (2008).
Sheard, L. B. et al. Jasmonate perception by inositol-phosphate-potentiated COI1–JAZ co-receptor. Nature 468, 400–405 (2010).
Thines, B. et al. JAZ repressor proteins are targets of the CF(COI1) complex during jasmonate signalling. Nature 448, 661–665 (2007).
Glazebrook, J. Contrasting mechanisms of defence against biotrophic and necrotrophic pathogens. Annu. Rev. Phytopathol. 43, 205–227 (2005).
Ali, S. et al. Overexpression of NPR1 in Brassica juncea confers broad-spectrum resistance to fungal pathogens. Front. Plant Sci. 8, 1693 (2017).
Vlot, A. C., Dempsey, D. M. A. & Klessig, D. F. Salicylic acid, a multifaceted hormone to combat disease. Annu. Rev. Phytopathol. 47, 177–206 (2009).
Grant, M. & Lamb, C. Systemic immunity. Curr. Opin. Plant Biol. 9, 414–420 (2006).
Van der Does, D. et al. Salicylic acid suppresses jasmonic acid signaling downstream of SCFCOI1-JAZ by targeting GCC promoter motifs via transcription factor ORA59. Plant Cell 25, 744–761 (2013).
Yamada, S. et al. Involvement of OsJAZ8 in jasmonate-induced resistance to bacterial blight in rice. Plant Cell Physiol. 53, 2060–2072 (2012).
Hong, Y. et al. Overexpression of MoSM1, encoding for an immunity-inducing protein from Magnaporthe oryzae, in rice confers broad-spectrum resistance against fungal and bacterial diseases. Sci. Rep. 7, 41037 (2017).
Ke, Y., Deng, H. & Wang, S. Advances in understanding broad-spectrum resistance to pathogens in rice. Plant J. 90, 738–748 (2017).
Panthapulakkal Narayanan, S. Overexpression of rice gene acyl-CoA-binding protein 5 leads to enhanced broad-spectrum pathogen defence in rice and Arabidopsis. HKU Theses Online (HKUTO, 2018).
Panthapulakkal Narayanan, S., Liao, P., Taylor, P. W., Lo, C. & Chye, M. L. Overexpression of a monocot acyl-CoA-binding protein confers broad-spectrum pathogen protection in a dicot. Proteomics 19, 1800368 (2019).
Yang, D. L., Yang, Y. & He, Z. Roles of plant hormones and their interplay in rice immunity. Mol. Plant 6, 675–685 (2013).
Lescot, M. et al. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 30, 325–327 (2002).
Higo, K., Ugawa, Y., Iwamoto, M. & Korenaga, T. Plant cis-acting regulatory DNA elements (PLACE) database. Nucleic Acids Res. 27, 297–300 (1999).
Berri, S. et al. Characterization of WRKY co-regulatory networks in rice and Arabidopsis. BMC Plant Biol. 9, 120 (2009).
Wang, Y., Liu, G. J., Yan, X. F., Wei, Z. G. & Xu, Z. R. MeJA-inducible expression of the heterologous JAZ2 promoter from Arabidopsis in Populus trichocarpa protoplasts. J. Plant Dis. Prot. 118, 69–74 (2011).
Washida, H. et al. Identification of cis-regulatory elements required for endosperm expression of the rice storage protein glutelin gene GluB-1. Plant Mol. Biol. 40, 1–12 (1999).
Zimmerman, D. C. & Feng, P. Characterization of a prostaglandin-like metabolite of linolenic acid produced by a flaxseed extract. Lipids 13, 313–316 (1978).
Patkar, R. N. & Chattoo, B. B. Transgenic indica rice expressing ns-LTP-like protein shows enhanced resistance to both fungal and bacterial pathogens. Mol. Breed. 17, 159–171 (2006).
Harkenrider, M. et al. Overexpression of rice wall-associated kinase 25 (OsWAK25) alters resistance to bacterial and fungal pathogens. PLoS ONE 11, e0147310 (2016).
De Vleesschauwer, D., Gheysen, G. & Hofte, M. Hormone defence networking in rice: tales from a different world. Trends Plant Sci. 18, 555–565 (2013).
Niki, T., Mitsuhara, I., Seo, S., Ohtsubo, N. & Ohashi, Y. Antagonistic effect of salicylic acid and jasmonic acid on the expression of pathogenesis-related (PR) protein genes in wounded mature tobacco leaves. Plant Cell Physiol. 39, 500–507 (1998).
Spoel, S. H. et al. NPR1 modulates cross-talk between salicylate-and jasmonate-dependent defence pathways through a novel function in the cytosol. Plant Cell 15, 760–770 (2003).
Koornneef, A. et al. Kinetics of salicylate-mediated suppression of jasmonate signalling reveal a role for redox modulation. Plant Physiol. 147, 1358–1368 (2008).
Koornneef, A. & Pieterse, C. M. Cross talk in defence signalling. Plant Physiol. 146, 839–844 (2008).
Leon-Reyes, A. et al. Salicylate-mediated suppression of jasmonate-responsive gene expression in Arabidopsis is targeted downstream of the jasmonate biosynthesis pathway. Planta 232, 1423–1432 (2010).
Caarls, L., Pieterse, C. M. & Van Wees, S. How salicylic acid takes transcriptional control over jasmonic acid signalling. Front Plant Sci. 6, 170 (2015).
Caarls, L. et al. Assessing the role of ETHYLENE RESPONSE FACTOR transcriptional repressors in salicylic acid-mediated suppression of jasmonic acid-responsive genes. Plant Cell Physiol. 58, 266–278 (2017).
Zhang, L., Zhang, F., Melotto, M., Yao, J. & He, S. Y. Jasmonate signaling and manipulation by pathogens and insects. J. Exp. Bot. 68, 1371–1385 (2017).
Mei, C., Qi, M., Sheng, G. & Yang, Y. Inducible overexpression of a rice allene oxide synthase gene increases the endogenous jasmonic acid level, PR gene expression, and host resistance to fungal infection. Mol. Plant-Microbe Interact. 19, 1127–1137 (2006).
Tamaoki, D. et al. Jasmonic acid and salicylic acid activate a common defence system in rice. Plant Signal. Behav. 8, e24260 (2013).
Truitt, C. L., Wei, H. X. & Paré, P. W. A plasma membrane protein from Zea mays binds with the herbivore elicitor volicitin. Plant Cell 16, 523–532 (2004).
Kachroo, P., Shanklin, J., Shah, J., Whittle, E. J. & Klessig, D. F. A fatty acid desaturase modulates the activation of defense signaling pathways in plants. Proc. Natl. Acad. Sci. 98, 9448–9453 (2001).
Meng, W. & Chye, M. L. Rice acyl-CoA-binding proteins OsACBP4 and OsACBP5 are differentially localised in the endoplasmic reticulum of transgenic Arabidopsis. Plant Signal. Behav. 9, e29544 (2014).
Shimono, M. et al. Rice WRKY45 plays a crucial role in benzothiadiazole-inducible blast resistance. Plant Cell 19, 2064–2076 (2007).
Kim, C. Y. et al. Identification of rice blast fungal elicitor-responsive genes by differential display analysis. Mol. Plant Microbe Interact. 13, 470–474 (2000).
Wen, N., Chu, Z. & Wang, S. Three types of defence responsive genes are involved in resistance to bacterial blight and fungal blast diseases in rice. Mol. Genet. Genomics 269, 331–339 (2003).
Liu, X. Q. et al. OsWRKY03, a rice transcriptional activator that functions in defence signalling pathway upstream of OsNPR1. Cell Res. 15, 593–603 (2005).
Ryu, H. S. et al. A comprehensive expression analysis of the WRKY gene superfamily in rice plants during defence response. Plant Cell Rep. 25, 836–847 (2006).
Qiu, D. et al. OsWRKY13 mediates rice disease resistance by regulating defence-related genes in salicylate-and jasmonate-dependent signalling. Mol. Plant-Microbe Interact. 20, 492–499 (2007).
Peng, X. et al. Constitutive expression of rice WRKY30 gene increases the endogenous jasmonic acid accumulation, PR gene expression and resistance to fungal pathogens in rice. Planta 236, 1485–1498 (2012).
Peng, X. et al. OsWRKY80-OsWRKY4 module as a positive regulatory circuit in rice resistance against Rhizoctonia solani. Rice 9, 63 (2016).
Cheng, H. et al. The WRKY45-2 WRKY13 WRKY42 transcriptional regulatory cascade is required for rice resistance to fungal pathogen. Plant Physiol. 167, 1087–1099 (2015).
Liu, J. et al. Alternative splicing of rice WRKY62 and WRKY76 transcription factor genes in pathogen defense. Plant Physiol. 171, 1427–1442 (2016).
Sheikh, A. H. et al. Regulation of WRKY46 transcription factor function by mitogen-activated protein kinases in Arabidopsis thaliana. Front. Plant Sci. 7, 61 (2016).
Wang, H. et al. Rice WRKY4 acts as a transcriptional activator mediating defense responses toward Rhizoctonia solani, the causing agent of rice sheath blight. Plant Mol. Biol. 89, 157–171 (2015).
Hiroyuki, K. & Terauchi, R. Regulation of expression of rice thaumatin-like protein: inducibility by elicitor requires promoter W-box elements. Plant Cell Rep. 27, 1521–1528 (2008).
Xiao, J. et al. Rice WRKY13 regulates cross talk between abiotic and biotic stress signaling pathways by selective binding to different cis-elements. Plant Physiol. 163, 1868–1882 (2013).
Liu, X., Bai, X., Wang, X. & Chu, C. OsWRKY71, a rice transcription factor, is involved in rice defence response. J. Plant Physiol. 164, 969–979 (2007).
Jones, J, D. & Dangl, J. L. The plant immune system. Nature, 444, 323–329 (2006).
Dodds, P. N. & Rathjen, J. P. Plant immunity: towards an integrated view of plant–pathogen interactions. Nat. Rev. Genet. 11, 539 (2010).
Cheval, C., Aldon, D., Galaud, J. P. & Ranty, B. Calcium/calmodulin-mediated regulation of plant immunity. Biochim. Biophys. Acta Mol. Cell Res. 1833, 1766–1771 (2013).
Liu, J., Elmore, J. M., Lin, Z. J. & Coaker, G. A receptor-like cytoplasmic kinase phosphorylates the host target RIN4, leading to the activation of a plant innate immune receptor. Cell Host Microbe 9, 137–146 (2011).
Wang, Y. et al. OsRAR1 and OsSGT1 physically interact and function in rice basal disease resistance. Mol. Plant-Microbe Interact. 21, 294–303 (2008).
Song, M. Y. et al. Differential requirement of Oryza sativa RAR1 in immune receptor-mediated resistance of rice to Magnaporthe oryzae. Mol. Cells 35, 327–334 (2013).
Shirasu, K. & Schulze-Lefert, P. Complex formation, promiscuity and multi-functionality: protein interactions in disease-resistance pathways. Trends Plant Sci. 8, 252–258 (2003).
Seo, P. J., Kim, S. G. & Park, C. M. Membrane-bound transcription factors in plants. Trends Plant Sci. 13, 550–556 (2008).
Shirasu, K. The HSP90-SGT1 chaperone complex for NLR immune sensors. Annu. Rev. Plant Biol. 60, 139–164 (2009).
Kadota, Y., Shirasu, K. & Guerois, R. NLR sensors meet at the SGT1–HSP90 crossroad. Trends Biochem Sci. 35, 199–207 (2010).
Hammond-Kosack, K. E. & Jones, J. D. Resistance gene-dependent plant defense responses. Plant Cell 8, 1773–1791 (1996).
Ryals, J. A. et al. Systemic acquired resistance. Plant Cell 8, 1809–1819 (1996).
Zhou, J. M. et al. NPR1 differentially interacts with members of the TGA/OBF family of transcription factors that bind an element of the PR-1 gene required for induction by salicylic acid. Mol. Plant-Microbe Interact. 13, 191–202 (2000).
Westfall, C. S. et al. Structural basis for prereceptor modulation of plant hormones by GH3 proteins. Science 336, 1708–1711 (2012).
Kazan, K. & Manners, J. M. MYC2: the master in action. Mol. Plant 6, 686–703 (2013).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 15, 473–497 (1962).
Yi, J. & An, G. Utilization of T-DNA tagging lines in rice. J. Plant Biol. 56, 85–90 (2013).
Fina, J. et al. UV-B inhibits leaf growth through changes in Growth-Regulating Factors and gibberellin levels. Plant Physiol. 174, 1110–1126 (2017).
Guo, Z. H., Chan, W. H., Kong, G. K., Hao, Q. & Chye, M. L. The first plant acyl-CoA-binding protein structures: the close homologues OsACBP1 and OsACBP2 from rice. Acta Crystallogr. D Struct. Biol. 73, 438–448 (2017).
Wu, X., Xiong, E., Wang, W., Scali, M. & Cresti, M. Universal sample preparation method integrating trichloroacetic acid/acetone precipitation with phenol extraction for crop proteomic analysis. Nat. Protoc. 9, 362 (2014).
Ross, P. L. et al. Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol. Cell. Proteomics 3, 1154–1169 (2004).
Zhu, F. Y. et al. SWATH-MS quantitative proteomic investigation reveals a role of jasmonic acid during lead response in Arabidopsis. J. Proteome Res. 15, 3528–3539 (2016).
Ma, W. & Berkowitz, G. A. The grateful dead: calcium and cell death in plant innate immunity. Cell. Microbiol. 9, 2571–2585 (2007).
Ma, W., Smigel, A., Verma, R. & Berkowitz, G. A. Cyclic nucleotide gated channels and related signaling components in plant innate immunity. Plant Signal. Behav. 4, 277–282 (2009).
Meng, X. & Zhang, S. MAPK cascades in plant disease resistance signaling. Annu. Rev. Phytopathol. 51, 12.1–12.22 (2013).
Mithoe, S. C. et al. Attenuation of pattern recognition receptor signaling is mediated by a MAP kinase kinase kinase. EMBO Rep. 17, 441–454 (2016).
Austin, M. J. et al. Regulatory role of SGT1 in early R gene-mediated plant defenses. Science 295, 2077–2080 (2002).
Liang, X. & Zhou, J. M. Receptor-like cytoplasmic kinases: central players in plant receptor kinase-mediated signaling. Annu. Rev. Plant Biol. 69, 267–299 (2018).
Mou, Z., Fan, W. & Dong, X. Inducers of plant systemic acquired resistance regulate NPR1 function through redox changes. Cell 113, 935–944 (2003).
Seyfferth, C. & Tsuda, K. Salicylic acid signal transduction: the initiation of biosynthesis, perception and transcriptional reprogramming. Front. Plant Sci. 5, 697 (2014).
Suza, W. P. & Staswick, P. E. The role of JAR1 in jasmonoyl-L-isoleucine production in Arabidopsis wound response. Planta 227, 1221–1232 (2008).
Sah, D. N. & Rush, M. C. Physiological races of Cercospora oryzae in the Southern United States. Plant Dis. 72, 263 (1988).
Mani, K. K., Hollier, C. A. & Groth, D. E. Effect of cultivar susceptibility and planting date on narrow brown leaf spot progression in rice. Crop Prot. 102, 88–93 (2017).
Park, C. H. et al. The Magnaporthe oryzae effector AvrPiz-t targets the RING E3 Ubiquitin Ligase APIP6 to suppress pathogen-associated molecular pattern-triggered immunity in rice. Plant Cell 24, 4748–4762 (2012).
Leslie, J. F. & Summerell, B. A. The Fusarium Laboratory Manual (Blackwell Publishing (Ames, IA, USA, 2006).
Shin, S. et al. A simple method for the assessment of Fusarium head blight resistance in Korean wheat seedlings inoculated with Fusarium graminearum. Plant Pathol. J. 30, 25–32 (2014).
Sun, X. et al. Xa26, a gene conferring resistance to Xanthomonas oryzae pv. oryzae in rice, encodes an LRR receptor kinase-like protein. Plant J. 37, 517–527 (2004).
Nakashima, K. et al. Functional analysis of a NAC-type transcription factor OsNAC6 involved in abiotic and biotic stress-responsive gene expression in rice. Plant J. 51, 617–630 (2007).
Jefferson, R. A., Kavanagh, T. A. & Bevan, M. W. GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 6, 3901–3907 (1987).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 3, 1101–1108 (2008).
Wang, G. et al. Dual function of rice OsDR8 gene in disease resistance and thiamine accumulation. Plant Mol. Biol. 60, 437–449 (2006).
Fu, J. et al. Manipulating broad-spectrum disease resistance by suppressing pathogen-induced auxin accumulation in rice. Plant Physiol. 155, 589–602 (2011).
Chen, X. et al. An XA21-associated kinase (OsSERK2) regulates immunity mediated by the XA21 and XA3 immune receptors. Mol. Plant 7, 874–892 (2014).
Hu, H., Xiong, L. & Yang, Y. Rice SERK1 gene positively regulates somatic embryogenesis of cultured cell and host defense response against fungal infection. Planta 222, 107–117 (2005).
Seo, Y.S. et al. Towards establishment of a rice stress response interactome. PLoS Genet. 7, (2011).
Liu, Q. et al. The germin-like protein OsGLP2-1 enhances resistance to fungal blast and bacterial blight in rice. Plant Mol. Biol. 92, 411–423 (2016).
Liu, B. et al. Lysin motif–containing proteins LYP4 and LYP6 play dual roles in peptidoglycan and chitin perception in rice innate immunity. Plant Cell 24, 3406–3419 (2012).
Yuan, Y. et al. Functional analysis of rice NPR1-like genes reveals that OsNPR1/NH1 is the rice orthologue conferring disease resistance with enhanced herbivore susceptibility. Plant Biotechnol. J. 5, 313–324 (2007).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M., Sato, Y., Furumichi, M., Morishima, K. & Tanabe, M. New approach for understanding genome variations in KEGG. Nucleic Acids Res. 47, D590–D595 (2019).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951 (2019).
Acknowledgements
We are grateful to Prof. Guoyin Kai (Shanghai Normal University) for providing pCAMBIA1304 vector, Prof. Yongjun Lin (Huazhong Agricultural University) for DX2181 vector, Prof. Gynheung An (Kyung Hee University) for the T-DNA insertion mutants, osnpr1 and oscoi1. We thank Dr Rumiana Ray (University of Nottingham) for the fungal strain Rhizoctonia solani AG-1-1 (ATCC 66157) and Dr Meng Wei (Northeast Forestry University) for the fungal strain Magnaporthe oryzae (RB22).
Funding
This work was supported by the Wilson and Amelia Wong Endowment Fund and the Research Grants Council of Hong Kong Special Administrative Region, China [HKU17109917, AoE/M-05/12, AoE/M-403/16, and Innovation Technology Fund of the Innovation Technology Commission (Funding Support to State Key Laboratory of Agrobiotechnology in Hong Kong)] to M.L.C. S.P.N. was supported by an HKU Postgraduate Studentship.
Author information
Authors and Affiliations
Contributions
S.P.N. performed most of the experiments. S.C.L made the constructs and generated transgenic rice lines. S.P.N and P.L. analysed GC-MS data. S.P.N., S.C.L., P.L., C.L., and M.L.C. analysed data. S.P.N. and M.L.C. designed the experiments and wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Panthapulakkal Narayanan, S., Lung, SC., Liao, P. et al. The overexpression of OsACBP5 protects transgenic rice against necrotrophic, hemibiotrophic and biotrophic pathogens. Sci Rep 10, 14918 (2020). https://doi.org/10.1038/s41598-020-71851-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-71851-9
- Springer Nature Limited
This article is cited by
-
Acyl-CoA-binding protein (ACBP) genes involvement in response to abiotic stress and exogenous hormone application in barley (Hordeum vulgare L.)
BMC Plant Biology (2024)
-
Advances in breeding, biotechnology, and nanotechnological approaches to combat sheath blight disease in rice
Molecular Biology Reports (2024)
-
Overexpression of cucumber CYP82D47 enhances resistance to powdery mildew and Fusarium oxysporum f. sp. cucumerinum
Functional & Integrative Genomics (2024)
-
Genome-wide identification and characterization of ACBP gene family in Populus reveal salinity alkali-responsive profiles
Journal of Forestry Research (2023)
-
A review of approaches to control bacterial leaf blight in rice
World Journal of Microbiology and Biotechnology (2022)