Abstract
Cancer widely affects the world’s health population and ranks second leading cause of death globally. Because of poor prognosis of various types of cancer such as sarcoma, lymphoma, adenomas etc., their high recurrence and metastasis rate and low early diagnosis rate have become concern lately. Role of autophagy in cancer progression is being studied since long. Autophagy is cell’s self-degradative mechanism towards stress and has role in degradation of the cytoplasmic macromolecules which has potential to damage other cytosolic molecules. Autophagy can promote as well as inhibit tumorigenesis depending upon the associated protein combinations in cancer cells. Recent studies have shown that non-coding RNAs (ncRNAs) do not code for protein but play essential role in modulation of gene expression. At transcriptional level, different ncRNAs like lncRNAs, miRNAs and circRNAs directly or indirectly affect different stages of autophagy like autophagy-dependent and non-apoptotic cell death in cancer cells. This review focuses on the involvement of ncRNAs in autophagy and the modulation of several cancer signal transduction pathways in cancers such as lung, breast, prostate, pancreatic, thyroid, and kidney cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer remains one of the major mortalities causing disease globally [1]. Cancer cells alter many cellular mechanisms which include uncontrolled nutrient supply, anti-senescence signals, uncontrolled blood supply and thus, acquire infinite growth and development [2]. According to statistics, there were 1,898,160 new cases of cancer and 608,570 cancer-related deaths in 2021.In male, the incidence and mortality rate is high for lung and prostate cancer. In contrary to males, breast cancer is of great concern for female, while the rate of pancreatic, kidney and thyroid cancers has been found to be increased in both [1].
Non-coding RNAs (ncRNAs) do not code for any protein, rather they protect the genome and act as regulators of autophagy and may aid in disease progression or inhibition [5]. Based on the number of nucleotides, the ncRNAs are classified into long non-coding RNAs (lncRNA) (more than 200 bp nucleotides), small ncRNA (less than 200 bp nucleotides) and circular RNAs (circRNA) [5, 10]. Small ncRNAs are further classified into microRNA (miRNA) (18–25 nucleotides), piwi interacting RNA (piRNA), small interfering RNA (siRNA) and small nucleolar RNA (snoRNA) [5]. ncRNAs do not code for any protein [11]. The epigenetic regulatory role of ncRNAs has been studied thoroughly with lncRNAs and miRNAs [5, 12]. circRNAs are a long chain ncRNAs, which has a closed loop structure. Research has shown that lncRNAs epigenetically control cancer cells by regulating invasion, metastasis, apoptosis and autophagy [5] and also regulates expression of autophagy related mRNAs and sponges miRNAs [13]. Like lncRNAs, circRNAs also sponge miRNAs and regulate autophagy associated cancer metastasis, progression [10].
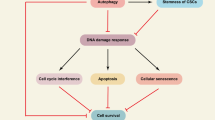
Autophagy is an evolutionary conserved cytoplasmic catabolic mechanism of cells, which is regulated by autophagy related genes (ATG) [3,4,5]. It aids in the removal of damaged cellular organelles, misfolded proteins, pathogens, and bulk cytoplasm from the cell, as well as the recycling of nutrients for cells. [2, 4]. Autophagy plays a crucial role in cancer metastasis and also helps in escaping tumor formation [4]. Autophagy can both kill cancer cells by degrading essential biological components, leading to apoptosis, and protect cancer cells by catabolic processes during conditions like administration of drugs administered and nutrient deficiency [6, 7]. Studies have shown the regulatory potentials of autophagy on anticancer drug discovery too [8]. ATG also regulate ncRNA, that modulate DNA methylation and histone modification, which may often induce autophagy too. The core autophagy process can be divided into 4 major steps. The first step, induction is initiated by the mTOR pathway where autophagic proteins such as ULK1, ATG13, ATG101 and FIP200 are involved. The following second step is phagophore nucleation, which is formed by BECLIN1 associated PIP3 kinase III. Autophagosomal membrane elongation is the third step which is modulated by two ubiquitin-like conjugation systems and involve autophagic factors like ATG3, ATG4, ATG5, ATG7 and ATG12. Until the maturation stage, the cellular machinery is recycled except a portion of LC3II, which gets attached to the membrane. The fourth step, also known as the maturation step, is marked by fusion of the autophagosome with lysosome [9]. Early autophagosome formation is regulated by ATG6 or BECLIN 1 [3]. BECLIN1 is found to be overexpressed in lung, prostate, breast, pancreatic cancers and are found to be regulated by different miRNAs and lncRNAs [3, 4, 14, 15]. Drug resistance poses as serious problem in cancer therapeutics, and it is primarily related to many signalling pathways [16]. Drug resistance is also modulated by autophagy related genes and associated miRNAs [17] and have thus been discussed in the study. Recent research on ncRNA-targeted medicines has significant implications for cancer therapy. The present review focuses on autophagy related ncRNAs (lncRNAs, miRNAs and circRNAs) in detailed manner, which can be used in lung, breast, prostate, pancreatic, kidney and thyroid cancers in future.
Autophagy related ncRNAs in lung cancer
Autophagy related miRNAs in lung cancer
Autophagy can act as an inhibitor or promotor of tumorigenesis and cancer progression. In Src tyrosine kinase inhibitor (TKI) induced autophagy, it is found out that upregulation of ULK1 and downregulation of miR-106a resulted into significant cell death in non-small cell lung cancer (NSCLC) cell line [18]. Overexpressed miR-21 was potentially found to be an oncogene, which act by inducing autophagy by up-regulating ULK1/LC3BII protein expressions, increasing autophagosome levels and downregulating p62, suggesting lymphatic metastasis in A549 cells [19]. In A549 and H460 lung cancer cell lines, miR-144 overexpression increased autophagy and apoptosis via LC3-II as well as BECLIN1 expression by targeting TIGAR protein, that functions to eliminate ROS from tumour cells, aiding suppression of tumour proliferation and metastasis inhibition [20]. miRNA also provides drug sensitivity towards NSCLC by downregulating autophagy related protein. miR-153-3p provide gefitinib-sensitivity in NSCLC cells by inhibiting ATG5 and upregulation of miR-1 is associated with DDP sensitivity by preventing ATG3-induced autophagy [21, 22]. Genetic inhibition of c-myc/miR-150 significantly repressed lung cancer by upregulating EPG2 concentration, which is autophagosome maturation-related protein [23].
Autophagy related lncRNAs in lung cancer
Metastasis-Associated Lung Adenocarcinoma Transcript 1 (MALAT1) is a lncRNA usually overexpressed in NSCLC. To activate autophagy, MALAT1 binds to RNA binding protein HuR [24]. Increased expression of lncRNA KTN1-AS1 in NSCLS acts as a prognostic marker for tumour progression and its knockdown results in increased apoptosis. miR-130a-5p acts as a negative regulator of KTN-AS1 and modulates autophagy targeting 3-phosphoinositide-dependent protein kinase 1 (PDPK1) [25]. Some lncRNAs like LCPAT1 can control the regulation of tumour suppressor miRNAs like miR-186-5p, miR-206, and miR-218-5p, which act on PI3K/AKT/mTOR pathway. This finding suggests that lncRNA can compete with miRNAs for activation of autophagy related proteins such as LC3, ATG and BECLIN2, resulting in tumour proliferation [26]. lncRNA loc146880 is related to biogenesis of lung cancer, and ROS induction by particulate matter enhances the expression of this lnRNA. This leads to BECLIN1 regulated autophagy activation and subsequent tumour cell migration and invasion [27]. A lncRNA, GAS5 is found to repress autophagy in NSCLC which is known to enhance the cisplatin sensitivity via downregulating BECLIN1, ATG3, ATG5, ATG7, ATG12 and LC3B expression [28]. BLACAT1 induced cisplatin chemoresistance in NSCLC was found to be modulated by upregulation of ATG7, MRP1 LC3-II/LC3-I and BECLIN1. miR-17 was found out to be the potential target for BLACAT1 lncRNA and may induce cisplatin sensitivity in NSCLC via miR-17/ATG7 [29]. lncRNA-HOTAIR supress ULK1, which is vital in initiation of autophagosome formation and upon silencing induces Crizotinib drug sensitivity and apoptosis in NSCLC [30]. lncRNA PANDAR acts as tumour suppressor by upregulating expression of BECLIN1, hence inducing autophagic response and BECLIN1 activates pro-apoptotic protein BAX in mitochondria, and eventually triggers apoptotic signals, leading to apoptosis of NSCLC cells [31]. Tumor-suppressor lncRNA on chromosome 8p12 (TSLNC8) inhibits tumour growth and supresses proliferation, metastasis in lung cancer via IL-6/STAT3/HIF-1α signaling pathways and eventually hypoxia induced autophagy is initiated [32]. Recent studies suggest that lncRNA ADAMTS9-AS2 act on AKT, PIK3CB, mTOR and autophagy-related proteins light chain 3-I/II (LC3-I/II), BECLIN1, sequestosome 1 (SQSTM1)] and apoptosis-related proteins (Bax and Bcl-2), contributing essential role in inhibition of tumour via modulating PI3K/AKT/mTOR signalling pathway [33].
Autophagy related circRNAs in lung cancer
Lately, circular RNAs are emerging as important controller of various cellular mechanisms and regulator of gene expression by acting as sponges of miRNA and lncRNA. One of the few circRNAs that play important role in cancer, circHIPK3 act as an oncogene in NSCLC. Silencing of circHIPK3 increases autophagy in tumour via activation of miR-124-3p, which is an activator of PRKAA/AMPK signalling pathway [34]. hsa_circ_0085131 circRNA activates autophagy related protein ATG7 by competing with miR-654-5p hence enhancing chemoresistance towards cisplatin in lung cancer cells [35]. In recent study by Fan and colleagues, it was found that six circRNAs are potentially intersecting miRNA-mRNA axes that regulate Chaperone-mediated autophagy, macroautophagy and regulatory protein of autophagy and act as potential modulatory mechanisms. This suggests that, in NSCLC, autophagy can be regulated by hsa_circ_0019709, hsa_circ_0040569, hsa_circ_0081983, hsa_circ_0098996, hsa_circ_0112354, hsa_circ_0135500 and these circRNAs can induce resistance to radiation therapy in NSCLC [36]. High expression of hsa_circ_0010235 in NSCLC downregulate expression of miR-433-3p. Downregulation of this circRNA and upregulation of miR-433-3p inhibited autophagy and initiated apoptosis in NSCLC via targeting mTOR signaling pathway regulator-like TIPRL protein that inhibit mTOR signaling [37].
Autophagy related ncRNAs in breast cancer
Autophagy related miRNAs in breast cancer
Breast cancer is one of the major cancers found in women. Many miRNAs provide drug resistance to cancer which is the main reason for invasion, metastasis and recurrence of breast cancer [38]. Oncogenic miR-638 acts on DACT3, the main modulator of Wnt/ β-catenin pathway and inhibits its activity. It was found that miR-638 activates mTOR induced autophagy and suppresses the expression of tumour suppressors such as BRCA1, PTEN and P53 [39], whereas tumour suppressive miR-135a inhibits proliferation of breast cancer MCF-7 and T47D cell lines via attenuating ELK1 and ELK3 oncogenes and decrease autophagy [40]. In a study, it was found that miR-224-5p inhibits autophagy via targeting SMAD4, which is a central mediator of transforming growth factor beta (TGFβ) and upon induction, it promotes proliferation and metastasis in MDA-MB-231 cells, a triple negative breast cancer cell line [41]. miRNA cluster analysis in breast cancer cell lines reveals that miR-106b, miR-93, and miR-25 modulate ULK1 (Unc-51 Like Autophagy Activating Kinase 1), TGFβ and promote invasiveness in breast cancer cells via AKT/MAPK signalling pathway [42]. In a study by Chen and colleagues, it was demonstrated that miR-23a increases breast cancer autophagic activity by XIAP and the mediated autophagic activation of LC3/p62 promotes metastasis and invasion of cancer cells [43]. In some breast cancer cases, it is reported that miRNA mediated loss of autophagy enhances the tumorigenesis via increasing intracellular reactive oxygen species (ROS) levels and DNA damage response (DDR) by targeting several key regulators of autophagy, including BECLIN 1, ATG16L1 and SQSTM1. A triple-negative breast cancer cell line and xenograft study demonstrates that miR-20a inhibits starvation induced autophagy and lysosomal activity through downregulating the expression of ATG16L1, BECLIN 1 and SQSTM1 [44]. In breast cancer, miR-489 and miR-567 reverse drug resistance of trastuzumab and doxorubicin by silencing autophagic flux [45, 46]. Taxol resistant breast cancer cell lines has been sensitized by upregulating miR-129-5p via supressing autophagy and enhancing apoptosis by inhibiting HMGB1, an autophagy-regulating protein that functions to displace Bcl-2 to bind to BECLIN1, in order to activate BECLIN1 for induction of autophagy [47].
Autophagy related lncRNAs in breast cancer
When lncRNAs were considered, DANCR-miR-758-3p-PAX6 axis was found to regulate autophagy and apoptosis as DANCR negatively regulates miR-758. Downregulation of DANCR was found to be beneficial in inhibiting malignant breast cancer [48]. HOTAIR, another lncRNA play important role in epigenetic gene silencing. HOTAIR cross talk with various ATGs and matrix metalloproteinases (MMPs) via various miRNAs, hence it has a pivotal role in breast cancer progression. When HOTAIR is downregulated, a decrease in migration and invasiveness in MCF-7 cells was observed with associated downregulation of MMP2, MMP9 and altered regulation of p53/Akt/JNK pathway, leading to arrest of cell cycle at G1 phase [49]. GAS5 as a lncRNA, was found to be overexpressed in MCF-7 cell line and BECLIN 1, ATG7, ATG12 and LC3B was found to be downregulated also, which affects ULK1 and ULK2 expression, resulting in upregulated apoptosis and supressed autophagy [50]. Upregulation of MAPT‐AS1 was also correlated with better patient survival in breast cancer [51].
Autophagy related circRNAs in breast cancer
Autophagy associated circRNAs such as circCDYL act as sponge on miR-1275 and modulate expression of autophagy associated downstream genes ATG7 and ULK1[52]. Increased expression of circular RNA circ-DNMT11 in breast cancer was positively correlated with increased proliferation and survival of breast cancer cells and it also stimulated autophagic flux downregulating p53 expression [53]. Autophagy related ncRNAs which are found to be involved in breast cancer and their targets are given in Table 1 below.
Autophagy related ncRNAs in prostate cancer
Autophagy related miRNAs in prostate cancer
Prostate cancer (PTC) is one of the most common cancers in men. In a study by Liao and colleagues, miR-381 has been found to downregulate the reclin (RELN) and promote the autophagy of PTC cells [61]. In another study, it was revealed that upregulation of LC3II and BECLIN 1 by miR-139 and miR-381 was led to increase in autophagy [15, 18]. There was inhibition of the PI3K/AKT/mTOR signalling pathway due to reduced expression of RELN, which was found to be modulated by miR-139 [15, 18]. The formation of autophagic flux blockade by miR-139 resulted in apoptosis of PTC [15]. Usually, ULK2 is overexpressed in PTC cells by regulating mTOR signalling pathway and ATG protein upregulation. Another study found out that when miR-26b has been upregulated, it led to downregulation of ULK2 and resulted in inhibition of the autophagy process in PTC [62].
Autophagy related lncRNAs in prostate cancer
TUG1, a lncRNA, has been found to balance oxidative stress and regulate autophagy by regulating antioxidant signalling pathways and NRF2 expression and enhances the invasion and proliferation of PTC. Knockdown of TUG1 resulted in decrease in NRF2 level [63,64,65]. TUG1 sponges miR-139-5p and positively regulates SMC1a. Knockdown of TUG1 results in radiosensitization of PTC cell lines [64]. Upregulation of a lncRNA TINCR decreases the proliferation, invasion and migration of PTC cells via suppressing TRIP13 expression [66]. LINC01116 knockdown has shown the increased expression of GAPDH, MAP1LC3B2 or LC3II and H2AFY, which resulted in inhibition of PTC cell proliferation. The expression of LINC01116 has been altered by sulforaphane following a novel mechanism and can be used as therapeutic strategies [67]. Radiosensitivity of PTC cells has been increased when GAS5 was upregulated by sponging miR-18a [68]. HULC has been found to suppress the phosphorylation of BECLIN 1 and reduced autophagy via mTOR signalling pathway. Knockdown of HULC, a lncRNA, has been found to enhance the radiosensitivity of PTC cells [69].
Autophagy related circRNAs in prostate cancer
circCSPP1 is modulated by HnRNP-1 and sponges miR-520 h, which in turn upregulate EGRI. As a result, circCSPP1 induces autophagy and accelerates the proliferation of PTC cells through circCSPP1-miR-520 h-EGR1 axis [70]. circ_0001747 promotes autophagy and increases the proliferation of PTC cells [71]. circRNA_CCNB2 knockdown enhances expression of miR-30b-5p and inhibits autophagy, which increases the sensitivity of irradiation resistance PTC cells towards irradiation by suppressing the expression of KIF18A [72].
Autophagy related ncRNAs in pancreatic cancer
Autophagy related miRNAs in pancreatic cancer
Pancreatic cancer (PC) is a common and important malignancy in the western world [73, 74]. ncRNAs are found to play a crucial role in metastasis, invasion, autophagy and apoptosis in PC [74]. Borchardt and colleagues have shown that expression of MYC is modulated by miR-24-3p, which induce apoptosis, modulate ROS generation and initiate autophagy in PC. The result showed the powerful effect of tumor inhibitory action of miR-24-3p in PC [73]. A study by Gu and colleagues depicted that miR-7 inhibits cell proliferation and metastasis of PC cells as miR-7 modulates the LKB1-AMPK-mTOR signaling pathway and decreases the autophagy. This process interferes with glycolysis to reduce the glucose levels in PC cells in in-vivo and in-vitro conditions [74]. Another study showed that miR-7 expression was found to be reduced in PC cells. Overexpression of miR-7 induced autophagy and reduced the glycolytic pathway, and also decreased the resistance of PC cells to gemcitabine [75]. Highly expressed miR-29c has inversely correlated with autophagy in PC [76]. Tetraspanin 1 (TSPAN1) is found to be actively participating in cell proliferation and enhances autophagy. FAM83A which is a positive regulator of TSPAN1 offers chemo-resistant to pancreatic cells. miR-454 targets both FAM83A and TSPAN1[77]. Overexpressed miR-137 directly targeted the ATG5 gene and downregulated autophagy in PC cells, and also increased the sensitivity to doxorubicin [78]. miR-9 downregulated eIF5A2 by directly acting upon 3’UTR of eIF5A2 and inhibited autophagy. PC cells with overexpressed miR-9 have shown more sensitivity to doxorubicin by repressing autophagy in PC cells [79].
Autophagy related lncRNAs in pancreatic cancer
In a study by Liu and colleagues, LINC01207 has been shown to interact with miR-143-5p which interacts with AGR2. Silencing of LINC01207 led to upregulation of miR-143-5p and downregulation of AGR2, which induced apoptosis and autophagy in PC cells [14]. PVT1, a lncRNA sponges miR-20a-5p in PC cells and modulate ULK1 expression [80].
Autophagy related circRNAs in pancreatic cancer
circRNA such as hsa_circ_0071922 act as miRNA sponge for miR-663a-5p by targeting the mTOR pathway in PC cells. This led to autophagy and inhibit cell proliferation in PC [10]. circRHOBTB3 acts as direct sponge to miR-600 and it enhances autophagy and proliferation of pancreatic ductal adenocarcinoma cells through inhibition of AKT/mTOR pathway [81]. Addition of chloroquine diphosphate causes autophagy suppression in pancreatic cancer and both miR-663a-5p and miR-154-3p were found to be under-expressed in chloroquine diphosphate treated cells [82]. These circRNAs can be exploited in pancreatic cancer therapeutics.
Autophagy related ncRNAs in thyroid cancer
Autophagy related miRNAs in thyroid cancer
miRNAs have been reported as prominent biomarkers and therapeutic targets in thyroid cancer (TC) [83,84,85]. Elevated expression of miR-183 and miR-375 were found in medullary thyroid cancer cells. Further study reported that knockdown of the elevated miR-183 resulted in increased expression of LC3B and autophagy in TC [84]. In another in-vitro study of medullary thyroid cancer cells, miR-9-3p transfection inhibited autophagy by targeting ATG5 and BECLIN 1 and resulted in decreased cell viability and increased apoptosis [85]. Binding of miR-30d to BECLIN 1 resulted to reduction of autophagy in anaplastic thyroid cancer cells and sensitivity of TC to cisplatin drug was increased [86]. In cisplatin treated anaplastic thyroid cells, miR-144 targets TGF-α and downregulates autophagy preventing TC progression [87]. Also, miR-30d and miR-144 enhance the sensitivity of anaplastic TC cells towards cisplatin, however, miR-30d reduces autophagy while miR-144 induces autophagy. Furthermore, overexpression of miR-125b negatively regulate FOXP3 expression which eventually modulate the expression of ATG7, BECLIN 1 and LC3II, and stimulate autophagy in thyroid cancer cells [88]. miR-524-5p directly downregulates ITGA3 and FOXE1 which leads to increase in the rate of apoptosis and autophagy and decline in proliferation and invasion of papillary TC cells. [89]. Therefore, expression of miR-183, miR-9-3p, miR-375, miR-524-5p, miR-125b, miR-144 and miR-30d have shown reduce TC cells proliferation by either downregulation or upregulation of autophagy associated gene expression.
Autophagy related lncRNAs in thyroid cancer
Altered lncRNAs expression have been reported in TC [90]. Yang and colleagues and Peng and colleagues have revealed that overexpression of a lncRNA TNRC6C-AS1 is associated with increased methylation of STK4 promoter gene which in turn induce TC proliferation. Silencing of TNRC6C-AS1 led to expression of STK4 which have been found to promote autophagy, apoptosis and proliferation of TC cells by activating Hippo signalling pathway. Suppression of TNRC6C-AS1 was also found to serve as protection against TC by inducing autophagy and apoptosis. [90, 91]. An in-vivo study has shown that lncRNA MALAT1 regulate miR-200a-3p/FOXA1 axis by acting as ceRNA and modulate the proliferation of anaplastic TC cells. Suppression of MALAT1 led to upregulation of miR-220a-3p and inhibition of miR-220a-3p resulted in overexpression of FOXA1. It was reported that inhibition of MALAT1 expression and upregulation of miR-220a-3p led to increase in BECLIN1 expression and LC-3II/I ratio, leading to increased apoptosis and autophagy, and eventually suppressed anaplastic TC cells proliferation [83]. In papillary TC, a lncRNA namely Linc00941 was found to be involved in the regulation of CHD6 expression and TGFβ signalling in papillary TC and was associated in inhibiting autophagy [92]. Overexpressed lncRNA GAS8-AS1 was also found to upregulate ATG5, leading to induction of autophagy [6]. In another study, lncRNA RP11-476D10.1 was found to downregulate miR-138-5p and upregulate LRRK2 expression. Silencing of lncRNA RP11-476D10.1 reverse its regulation on miR-138-5p and LRRK2, and resulted in increased BECLIN1, LC3B, Bax expression and decreased BECLIN 2 expression, leading to induction of autophagy and apoptosis of papillary thyroid cancer cells [93].
Autophagy related circRNAs in thyroid cancer
circ_0067934 sponges and inhibits miR-545-3p to repress ferroptosis of TC cells by upregulation of negative regulator of ferroptosis, SLC7A11. Upon silencing of circ_0067934, ferroptosis is induced and its knockdown eventually increases ferroptosis, autophagy and apoptosis in TC cells [94]. Another study reveals that knockdown of circEIF6 along with cisplatin treatment reduces LC3 II/LC3 I ratio, autophagy and cell proliferation of TC cells in vivo by enhancing miR-144-3p expression and inhibiting TGF-α expression [95].
Some of the ncRNAs along with their target molecules have been listed in Table 2. The role of different miRNAs, lncRNAs and circRNAs in kidney cancer and their associated process of modulation of autophagy have been provided in Table 3. Modulation of PI3K/AKT/mTOR pathway in autophagy by various ncRNAs in different cancers have been explained in Fig. 1.
Modulation of PI3K/AKT/mTOR pathway in Autophagy by various ncRNAs in lung, thyroid, prostate, breast, kidney and pancreatic cancers. ncRNAs shown in red inhibit the PI3K/AKT/mTOR pathway via sponging miRNA or directly inhibiting associated protein. ncRNA shown in green modulate PI3K/AKT/mTOR pathway promoting autophagy
Role of autophagy-based ncRNAs in drug resistance and therapeutics
Drug resistance
Drug resistance is an inevitable impediment for an effective and lasting cancer treatment [16]. Cancer cells are found to be resistant to many chemotherapeutic drugs such as cisplatin, crizotinib, trastuzumab, gemcitabine, gefitinib, doxorubicin, temozolomide and capecitabine [21, 28,29,30, 35, 45, 46, 75, 78, 79, 86,87,88, 92, 95, 113]. Trastuzumab and doxorubicin are commonly used drugs but they fail to exert bioactivity due to chemoresistance. Studies have shown that regulating autophagic flux can retrieve cancer cells sensitivity towards trastuzumab and doxorubicin. ncRNAs have been found to play pivotal roles in breast cancer, TC, PTC, LC and PC drug resistance. In breast cancer, miR-567 reverses drug resistance of trastuzumab by inactivating autophagic flux [45]. Furthermore, in TC, ncRNAs have been found to increase the drug sensitivity by modulating autophagy associated genes. Introduction of miR-30d in the system increases the sensitivity of TC towards cisplatin by inhibiting BECLIN 1 mediated autophagy [86]. miR-144 is found to enhance the effects of cisplatin by inhibiting cisplatin induced autophagy while miR-125b enhances cisplatin induced autophagy [87]. Moreover, overexpressed miR-125b improves the sensitivity of TC cells towards sorafenib and cisplatin [88]. circEIF6 improves cisplatin induced autophagy and increases its resistance in anaplastic and papillary TC cells by targeting miR-144-3p [95].
Therapeutic roles of autophagy associated ncRNAs in cancers
Autophagy related ncRNAs may be used in therapeutics to cure or get relieve from cancers [84]. Autophagy serves as either survival or destruction mechanism in cancers. Addition of some ncRNAs subsequently lead to significant decline in autophagy and some ncRNAs lead to overexpression of autophagy flux. Therefore, these ncRNAs can be therapeutically targeted to diminish cancer [85]. Several studies have been reported where ncRNAs acts as tumour suppressor. In a recent study, Qin and colleagues found that overexpression of lncRNA GAS8-AS1 lead to suppression of papillary TC through ATG5 regulated autophagy [6]. Another study by Zhang and colleagues suggests that miR-30d acts as critical regulator of cisplatin drug, so it can be exploited as therapeutic target of TC [86]. Cisplatin in combination with overexpressed miR-125b in TC cells have shown potential to reduce TC cells proliferation [88]. In another study Lu and colleagues have reported high expression of a circRNA, circCSP1, which significantly increases proliferation, migration and invasion of PTC cells via autophagy upregulation [70]. Yang and colleagues have found that a lncRNA, TNRC6C-AS1 enhance the proliferation of TC cells by promoting autophagy and apoptosis [90]. Therefore, these ncRNAs can be targeted accordingly in future for cancer therapeutics. Autophagy associated ncRNAs which can be targeted in different types of cancers have been depicted in Fig. 2.
Conclusion
In this review, the role of autophagy-related ncRNAs is highlighted which can be useful to devise newer therapeutic strategies for cancer in future. The ncRNAs either enhance or suppress the autophagy by modulating ATG genes, BECLIN1, PTEN, ULK, LC3-I/II etc. or pathways such as PIK3/AKT/ATG, MAPK/AKT, PI3K/ERK etc. Potential diagnostic and prognostic ncRNAs associated with six most lethal cancers (lung, breast, prostate, kidney, thyroid and pancreatic cancers) have been explained here. miRNA and lncRNAs interact with a mRNA which is directly or indirectly affects autophagy and cell proliferation of cancer cells. siRNA, circRNA, lncRNA and shRNA act as sponges for miRNA and these ncRNA either upregulate or downregulate the miRNA or related mRNA or other protein which either increases or decreases the incidence of autophagy of cancer cells. From various studies, it has been understood that not only upregulation of autophagy but downregulation of autophagy can also lead to inhibition of cancer cell proliferation, invasion and metastasis. For developing any new therapeutic strategies, it is very essential to understand of the mechanism of action of the therapeutic targets and its associated pathways in the disease. The mechanisms of action of all ncRNAs as autophagy inducers or inhibitors have been studied here and still demand more experiments to be conducted in this area. The ncRNAs mentioned in the study play important role in autophagy and can therefore can be explored as potential as well as promising therapeutic strategies for lung, breast, prostate, kidney, thyroid and pancreatic cancer treatment.
Abbreviations
- Non-coding RNA:
-
NcRNA
- lncRNA:
-
Long non coding RNA
- miRNA:
-
MicroRNA
- circRNA:
-
Circular RNA
- ATG:
-
Autophagy related gene
- ATG16L1:
-
Autophagy related 16 like 1 protein
- mTOR:
-
Mammalian target of rapamycin
- ULK1:
-
Unc-51 like autophagy activating kinase 1
- FIP200:
-
Focal adhesion kinase family-interacting protein of 200 kDa
- PIP3 kinase III:
-
Phosphatidylinositol 3 kinase III
- LC3II:
-
Microtubule-associated protein 1A/1B-light chain 3
- piRNA:
-
Piwi interacting RNA
- siRNA:
-
Small interfering RNA
- snoRNA:
-
Small nucleolar RNA
- ceRNA:
-
Competing endogenous RNA
- TKI:
-
Tyrosine kinase inhibitor
- NSCLC:
-
Non-small cell lung cancer
- TIGAR:
-
TP53-inducible glycolysis and apoptosis regulator
- ROS:
-
Reactive oxygen species
- DDP:
-
Dipeptidyl peptidase
- EPG2:
-
Ectopic P granules protein 2
- MALAT1:
-
Metastasis-Associated Lung Adenocarcinoma Transcript 1
- HuR:
-
Human antigen R
- PI3K:
-
Phosphoinositide 3-kinase
- GAS5:
-
Growth arrest specific 5 gene
- BLACAT 1:
-
Bladder cancer associated transcript 1
- MRP1:
-
Multidrug resistance associated protein 1
- TSLNC8:
-
Tumor-suppressive lncRNA on chromosome 8p12
- STAT3:
-
Signal transducer and activator of transcription 3
- HIF-1α:
-
Hypoxia inducible factor 1α
- PIK3CB:
-
Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit beta
- SQSTM1:
-
Sequestosome 1
- PRKAA/AMPK:
-
Protein kinase AMP activated catalytic subunit alpha
- TOR:
-
Target of rapamycin
- Wnt:
-
Wingless-related integration site
- BRCA1:
-
Breast cancer gene 1
- PTEN:
-
Phosphatase and Tensin Homolog deleted on chromosome 10
- ELK1:
-
ETS like-1 protien
- ELK3:
-
ETS like-2 protien
- SMAD4:
-
SMAD family member 4
- TGFβ:
-
Transforming growth factor beta
- MAPK:
-
Mitogen activated protein kinase
- XIAP:
-
X-linked inhibitor of apoptosis protein
- HMGB1:
-
High mobility group box 1
- PAX6:
-
Paired box protein 6
- MMPs:
-
Matrix metalloproteinases
- JNK:
-
C-Jun N-terminal kinase 1
- MAPT-AS1:
-
MAPT Antisense RNA 1
- ERK1/2:
-
Extracellular signal regulated kinase 1/2
- LAMP1:
-
Lysosomal-associated membrane protein 1
- RELN:
-
Reclin
- Nrf2:
-
Nuclear factor erythroid related factor 2
- SMC1A:
-
Structural maintenance of chromosome protein 1A
- TRIP13:
-
Thyroid receptor-interacting protein 13
- GAPDH:
-
Glyceraldehyde-3-phosphate dehydrogenase
- MAP1LC3B2:
-
Microtubule associated protein 1 light chain 3 beta 2
- H2AFY:
-
H2A histone family, member Y
- HnRNP-1:
-
Heterogeneous nuclear ribonucleoprotein A1
- EGR1:
-
Early growth response 1 gene
- KIF18A:
-
Kinesin family member 18A
- TUG1:
-
Taurine up-regulated 1
- LKB1:
-
Liver kinase B1
- AMPK:
-
AMP activated protein kinase
- TSPAN1:
-
Tetraspanin 1
- FAM83A:
-
Family member with sequence similarity 83A
- eIF5A2:
-
Eukaryotic translation initiation factor 5A2
- AGR2:
-
Anterior gradient protein 2 homolog
- STK4:
-
Serine/threonine-protein kinase 4
- FOXA1:
-
Forkhead box protein A1
- CHD6:
-
Chromodomain helicase DNA binding protein 6
- GAS8-AS1:
-
GAS8-antisense RNA 1
- LRRK2:
-
Leucine rich repeat kinase 2
- SLC7A11:
-
Solute carrier family 7 member 11
- FOXP3:
-
Foxhead box P3
- FOXE1:
-
Foxhead box E1
- ITGA3:
-
Integrin subunit alpha 3
- ITPR1:
-
Inositol 1,4,5-trisphosphate receptor type 1
- NOX4:
-
NADPH oxidase 4
- CCNB1:
-
Cyclin B1
References
Siegel RL, Miller KD, Fuchs HE, Jemal A (2021) Cancer statistics, 2021. Cancer J Clin 71(1):7–33. https://doi.org/10.3322/caac.21654
Cho YY, Kim DJ, Lee HS, Jeong CH, Cho EJ, Kim MO et al (2013) Autophagy and cellular senescence mediated by Sox2 suppress malignancy of cancer cells. PLoS ONE 8(2):e57172. https://doi.org/10.1371/journal.pone.0057172
Dragowska WH, Weppler SA, Wang JC, Wong LY, Kapanen AI, Rawji JS et al (2013) Induction of autophagy is an early response to gefitinib and a potential therapeutic target in breast cancer. PLoS ONE 8(10):e76503. https://doi.org/10.1371/journal.pone.0076503
Rezaei S, Mahjoubin-Tehran M, Aghaee-Bakhtiari SH, Jalili A, Movahedpour A, Khan H et al (2020) Autophagy-related MicroRNAs in chronic lung diseases and lung cancer. Crit Rev Oncol Hematol. https://doi.org/10.1016/j.critrevonc.2020.103063
Talebian S, Daghagh H, Yousefi B, Ȍzkul Y, Ilkhani K, Seif F, Alivand MR (2020) The role of epigenetics and non-coding RNAs in autophagy: a new perspective for thorough understanding. Mech Ageing Dev. https://doi.org/10.1016/j.mad.2020.111309
Qin Y, Sun W, Zhang H, Zhang P, Wang Z, Dong W, He L, Zhang T, Shao L, Zhang W, Wu C (2018) LncRNA GAS8-AS1 inhibits cell proliferation through ATG5-mediated autophagy in papillary thyroid cancer. Endocrine 59(3):555–564. https://doi.org/10.1007/s12020-017-1520-1
Qin Y, Sun W, Wang Z, Dong W, He L, Zhang T, Shao L, Zhang H (2020) ATF2-induced lncRNA GAS8-AS1 promotes autophagy of thyroid cancer cells by targeting the miR-187-3p/ATG5 and miR-1343-3p/ATG7 axes. Mol Ther Nucl Acids 22:584–600. https://doi.org/10.1016/j.omtn.2020.09.022
Zhang X, He Z, Xiang L, Li L, Zhang H, Lin F, Cao H (2019) Codelivery of GRP78 siRNA and docetaxel via RGD-PEG-DSPE/DOPA/CaP nanoparticles for the treatment of castration-resistant prostate cancer. Drug Des Dev Ther 13:1357. https://doi.org/10.2147/DDDT.S198400
Yang Y, Liang C (2015) MicroRNAs: an emerging player in autophagy. ScienceOpen Res. https://doi.org/10.14293/S2199-1006.1.SOR-LIFE.A181CU.v1
Jiang PC, Bu SR (2019) Clinical value of circular RNAs and autophagy-related miRNAs in the diagnosis and treatment of pancreatic cancer. Hepatobiliary Pancreat Dis Int 18(6):511–516. https://doi.org/10.1016/j.hbpd.2019.09.009
Islam Khan MZ, Tam SY, Law HKW (2019) Autophagy-modulating long non-coding RNAs (LncRNAs) and their molecular events in cancer. Front Genet 9:750. https://doi.org/10.3389/fgene.2018.00750
Doldi V, El Bezawy R, Zaffaroni N (2021) MicroRNAs as epigenetic determinants of treatment response and potential therapeutic targets in prostate cancer. Cancers 13(10):2380. https://doi.org/10.3390/cancers13102380
Wei J, Ge X, Tang Y, Qian Y, Lu W, Jiang K et al (2020) An autophagy-related long noncoding RNA signature contributes to poor prognosis in colorectal cancer. Journal of oncology. https://doi.org/10.1155/2020/4728947
Liu C, Wang JO, Zhou WY, Chang XY, Zhang MM, Zhang Y, Yang XH (2019) Long non-coding RNA LINC01207 silencing suppresses AGR2 expression to facilitate autophagy and apoptosis of pancreatic cancer cells by sponging miR-143-5p. Mol Cell Endocrinol 493:110424. https://doi.org/10.1016/j.mce.2019.04.004
Nam RK, Benatar T, Amemiya Y, Sherman C, Seth A (2020) Mir-139 regulates autophagy in prostate cancer cells through BECLIN-1 and mTOR signaling proteins. Anticancer Res 40(12):6649–6663. https://doi.org/10.21873/anticanres.14689
Son SW, Song MG, Yun BD, Park JK (2021) Noncoding RNAs associated with therapeutic resistance in pancreatic cancer. Biomedicines 9(3):263. https://doi.org/10.3390/biomedicines9030263
Xu P, Wu Q, Yu J, Rao Y, Kou Z, Fang G et al (2020) A systematic way to infer the regulation relations of miRNAs on target genes and critical miRNAs in cancers. Front Genet 11:278. https://doi.org/10.3389/fgene.2020.00278
Rothschild SI, Gautschi O, Batliner J, Gugger M, Fey MF, Tschan MP (2017) MicroRNA-106a targets autophagy and enhances sensitivity of lung cancer cells to Src inhibitors. Lung Cancer 107:73–83. https://doi.org/10.1016/j.lungcan.2016.06.004
Li S, Zeng X, Ma R, Wang L (2018) MicroRNA-21 promotes the proliferation, migration and invasion of non-small cell lung cancer A549 cells by regulating autophagy activity via AMPK/ULK1 signaling pathway. Exp Ther Med 16(3):2038–2045. https://doi.org/10.3892/etm.2018.6370
Chen S, Li P, Li J, Wang Y, Du Y, Chen X et al (2015) MiR-144 inhibits proliferation and induces apoptosis and autophagy in lung cancer cells by targeting TIGAR. Cell Physiol Biochem 35(3):997–1007. https://doi.org/10.1159/000369755
Zhang W, Dong YZ, Du X, Peng XN, Shen QM (2019) MiRNA-153–3p promotes gefitinib-sensitivity in non-small cell lung cancer by inhibiting ATG5 expression and autophagy. Eur Rev Med Pharmacol Sci 23:2444–2452. https://doi.org/10.26355/eurrev_201903_17391
Hua L, Zhu G, Wei J (2018) MicroRNA-1 overexpression increases chemosensitivity of non-small cell lung cancer cells by inhibiting autophagy related 3-mediated autophagy. Cell Biol Int 42(9):1240–1249. https://doi.org/10.1002/cbin.10995
Li H, Liu J, Cao W, Xiao X, Liang L, Liu-Smith F et al (2019) C-myc/miR-150/EPG5 axis mediated dysfunction of autophagy promotes development of non-small cell lung cancer. Theranostics 9(18):5134. https://doi.org/10.7150/thno.34887
Li S, Mei Z, Hu HB, Zhang X (2018) The lncRNA MALAT1 contributes to non-small cell lung cancer development via modulating miR-124/STAT3 axis. J Cell Physiol 233(9):6679–6688. https://doi.org/10.1002/jcp.26325
Li C, Zhao W, Pan X, Li X, Yan F, Liu S et al (2020) LncRNA KTN1-AS1 promotes the progression of non-small cell lung cancer via sponging of miR-130a-5p and activation of PDPK1. Oncogene 39(39):6157–6171. https://doi.org/10.1038/s41388-020-01427-4
Yu X, Ye X, Lin H, Feng N, Gao S, Zhang X, Wang Y, Yu H, Deng X, Qian B (2018) Knockdown of long non-coding RNA LCPAT1 inhibits autophagy in lung cancer. Cancer Biol Med 15(3):228–237. https://doi.org/10.20892/j.issn.2095-3941.2017.0150
Deng X, Feng N, Zheng M, Ye X, Lin H, Yu X et al (2017) PM2 5 exposure-induced autophagy is mediated by lncRNA loc146880 which also promotes the migration and invasion of lung cancer cells. Biochem Biophys Acta 1861(2):112–125. https://doi.org/10.1016/j.bbagen.2016.11.009
Zhang N, Yang GQ, Shao XM, Wei L (2016) GAS5 modulated autophagy is a mechanism modulating cisplatin sensitivity in NSCLC cells. Eur Rev Med Pharmacol Sci 20(11):2271–2277
Huang FX, Chen HJ, Zheng FX, Gao ZY, Sun PF, Peng Q et al (2019) LncRNA BLACAT1 is involved in chemoresistance of non-small cell lung cancer cells by regulating autophagy. Int J Oncol 54(1):339–347. https://doi.org/10.3892/ijo.2018.4614
Yang Y, Jiang C, Yang Y, Guo L, Huang J, Liu X et al (2018) Silencing of LncRNA-HOTAIR decreases drug resistance of non-small cell lung cancer cells by inactivating autophagy via suppressing the phosphorylation of ULK1. Biochem Biophys Res Commun 497(4):1003–1010. https://doi.org/10.1016/j.bbrc.2018.02.141
Zhang L, Wang Y, Xia S, Yang L, Wu D, Zhou Y, Lu J (2020) Long noncoding RNA PANDAR inhibits the development of lung cancer by regulating autophagy and apoptosis pathways. J Cancer 11(16):4783. https://doi.org/10.7150/jca.45291
Fan H, Li J, Wang J, Hu Z (2019) Long non-coding RNAs (lncRNAs) tumor-suppressive role of lncRNA on chromosome 8p12 (TSLNC8) inhibits tumor metastasis and promotes apoptosis by regulating interleukin 6 (IL-6)/signal transducer and activator of transcription 3 (STAT3)/hypoxia-inducible factor 1-alpha (HIF-1α) signaling pathway in non-small cell lung cancer. Med Sci Monitor 25:7624. https://doi.org/10.12659/MSM.917565
Li H, Huang H, Li S, Mei H, Cao T, Lu Q (2021) Long non-coding RNA ADAMTS9-AS2 inhibits liver cancer cell proliferation, migration and invasion. Exp Ther Med 21(6):1–9. https://doi.org/10.3892/etm.2021.9991
Shackelford DB, Shaw RJ (2009) The LKB1–AMPK pathway: metabolism and growth control in tumour suppression. Nat Rev Cancer 9(8):563–575. https://doi.org/10.1038/nrc2676
Kong R (2020) Circular RNA hsa_circ_0085131 is involved in cisplatin-resistance of non-small-cell lung cancer cells by regulating autophagy. Cell Biol Int 44(9):1945–1956. https://doi.org/10.1002/cbin.11401
Fan L, Li B, Li Z, Sun L (2021) Identification of autophagy related circRNA-miRNA-mRNA-subtypes network with radiotherapy responses and tumor immune microenvironment in non-small cell lung cancer. Front Genet. https://doi.org/10.3389/fgene.2021.730003
Zhang F, Cheng R, Li P, Lu C, Zhang G (2021) Hsa_circ_0010235 functions as an oncogenic drive in non-small cell lung cancer by modulating miR-433-3p/TIPRL axis. Cancer Cell Int 21(1):1–14. https://doi.org/10.1186/s12935-021-01764-8
Rahmani F, Tabrizi AT, Hashemian P, Alijannejad S, Rahdar HA, Ferns GA et al (2020) Role of regulatory miRNAs of the Wnt/β-catenin signaling pathway in tumorigenesis of breast cancer. Gene 754:144892. https://doi.org/10.1016/j.gene.2020.144892
Ren Y, Chen Y, Liang X, Lu Y, Pan W, Yang M (2017) MiRNA-638 promotes autophagy and malignant phenotypes of cancer cells via directly suppressing DACT3. Cancer Lett 390:126–136. https://doi.org/10.1016/j.canlet.2017.01.009
Ahmad A, Zhang W, Wu M, Tan S, Zhu T (2018) Tumor-suppressive miRNA-135a inhibits breast cancer cell proliferation by targeting ELK1 and ELK3 oncogenes. Genes Genom 40(3):243–251. https://doi.org/10.1007/s13258-017-0624-6
Cheng Y, Li Z, Xie J, Wang P, Zhu J, Li Y, Wang Y (2018) MiRNA-224-5p inhibits autophagy in breast cancer cells via targeting Smad4. Biochem Biophys Res Commun 506(4):793–798. https://doi.org/10.1016/j.bbrc.2018.10.150
Kandettu A, Radhakrishnan R, Chakrabarty S, Sriharikrishnaa S, Kabekkodu SP (2020) The emerging role of miRNA clusters in breast cancer progression. Biochem Biophys Acta. https://doi.org/10.1016/j.bbcan.2020.188413
Chen P, He YH, Huang X, Tao SQ, Wang XN, Yan H, Wu ZS (2017) MiR-23a modulates X-linked inhibitor of apoptosis-mediated autophagy in human luminal breast cancer cell lines. Oncotarget 8(46):80709. https://doi.org/10.18632/oncotarget.21080
Liu L, He J, Wei X, Wan G, Lao Y, Xu W et al (2017) MicroRNA-20a-mediated loss of autophagy contributes to breast tumorigenesis by promoting genomic damage and instability. Oncogene 36(42):5874–5884. https://doi.org/10.1038/onc.2017.193
Han M, Hu J, Lu P, Cao H, Yu C, Li X et al (2020) Exosome-transmitted miR-567 reverses trastuzumab resistance by inhibiting ATG5 in breast cancer. Cell Death Dis 11(1):1–15. https://doi.org/10.1038/s41419-020-2250-5
Soni M, Patel Y, Markoutsa E, Jie C, Liu S, Xu P, Chen H (2018) Autophagy, cell viability, and chemoresistance are regulated by miR-489 in breast cancer. Mol Cancer Res 16(9):1348–1360. https://doi.org/10.1158/1541-7786.MCR-17-0634
Shi Y, Gong W, Lu L, Wang Y, Ren J (2019) Upregulation of miR-129-5p increases the sensitivity to Taxol through inhibiting HMGB1-mediated cell autophagy in breast cancer MCF-7 cells. Braz J Med Biol Res. https://doi.org/10.1590/1414-431X20198657
Zhang XH, Li BF, Ding J, Shi L, Ren HM, Liu K et al (2020) LncRNA DANCR-miR-758-3p-PAX6 molecular network regulates apoptosis and autophagy of breast cancer cells. Cancer Manage Res 12:4073. https://doi.org/10.2147/CMAR.S254069
Yu Y, Lv F, Liang D, Yang Q, Zhang B, Lin H et al (2017) HOTAIR may regulate proliferation, apoptosis, migration and invasion of MCF-7 cells through regulating the P53/Akt/JNK signaling pathway. Biomed Pharmacother 90:555–561. https://doi.org/10.1016/j.biopha.2017.03.054
Li G, Qian L, Tang X, Chen Y, Zhao Z, Zhang C (2020) Long non-coding RNA growth arrest-specific 5 (GAS5) acts as a tumor suppressor by promoting autophagy in breast cancer. Mol Med Rep 22(3):2460–2468. https://doi.org/10.3892/mmr.2020.11334
Wang D, Li J, Cai F, Xu Z, Li L, Zhu H et al (2019) Overexpression of MAPT-AS1 is associated with better patient survival in breast cancer. Biochem Cell Biol 97(2):158–164. https://doi.org/10.1139/bcb-2018-0039
Liang G, Ling Y, Mehrpour M, Saw PE, Liu Z, Tan W et al (2020) Autophagy-associated circRNA circCDYL augments autophagy and promotes breast cancer progression. Mol Cancer 19(1):1–16. https://doi.org/10.1186/s12943-020-01152-2
Du WW, Yang W, Li X, Awan FM, Yang Z, Fang L et al (2018) A circular RNA circ-DNMT1 enhances breast cancer progression by activating autophagy. Oncogene 37(44):5829–5842. https://doi.org/10.1038/s41388-018-0369-y
Yu X, Li R, Shi W, Jiang T, Wang Y, Li C et al (2016) Silencing of MicroRNA-21 confers the sensitivity to tamoxifen and fulvestrant by enhancing autophagic cell death through inhibition of the PI3K-AKT mTOR pathway in breast cancer cells. Biomed Pharmacother 77:37–44. https://doi.org/10.1016/j.biopha.2015.11.005
Liu LM, Shen WF, Zhu ZH et al (2018) Combined inhibition of EGFR and c-ABL suppresses the growth of fulvestrant resistant breast cancer cells through miR-375-autophagy axis. Biochem Biophys Res Commun 498(3):559–565. https://doi.org/10.1016/j.bbrc.2018.03.019
Zhang FF, Wang BB, Long HL et al (2016) Decreased miR- 124–3p expression prompted breast cancer cell progression mainly by targeting BECLIN-1. Clin Lab 62(6):1139–1145. https://doi.org/10.7754/clin.lab.2015.151111
Pradhan AK, Talukdar S, Bhoopathi P, Shen X-N, Emdad L, Das SK et al (2017) mda-7/IL-24 mediates cancer cell-specific death via regulation of miR- 221 and the BECLIN-1 axis. Cancer Res 77:949–959
Cheng Y, Li Z, Xie J, Wang P, Zhu J, Li Y et al (2018) MiRNA-224-5p inhibits autophagy in breast cancer cells via targeting smad4. Biochem Biophys Res Commun 506:793–798
Chong ZX, Yeap SK, Ho WY (2021) Regulation of autophagy by microRNAs in human breast cancer. J Biomed Sci 28(1):1–18
Lu D, Luo P, Wang Q, Ye Y, Wang B (2017) lncRNA PVT1 in cancer: a review and meta-analysis. Int J Clin Chem 474:1–7. https://doi.org/10.1016/j.cca.2017.08.038
Liao W, Zhang Y (2020) MicroRNA-381 facilitates autophagy and apoptosis in prostate cancer cells via inhibiting the RELN-mediated PI3K/AKT/mTOR signaling pathway. Life Sci 254:117672. https://doi.org/10.1016/j.lfs.2020.117672
Clotaire DZJ, Zhang B, Wei N, Gao R, Zhao F, Wang Y et al (2016) MiR-26b inhibits autophagy by targeting ULK2 in prostate cancer cells. Biochem Biophys Res Commun 472(1):194–200. https://doi.org/10.1016/j.bbrc.2016.02.093
Liu, P. F., Farooqi, A. A., Peng, S. Y., Yu, T. J., Dahms, H. U., Lee, C. H., et al. (2020, October). Regulatory effects of noncoding RNAs on the interplay of oxidative stress and autophagy in cancer malignancy and therapy. In Seminars in Cancer Biology. Academic Press. https://doi.org/10.1016/j.semcancer.2020.10.009
Xiu D, Liu L, Cheng M, Sun X, Ma X (2020) Knockdown of lncRNA TUG1 enhances radiosensitivity of prostate cancer via the TUG1/miR-139-5p/SMC1A axis. Onco Targets Ther 13:2319. https://doi.org/10.2147/OTT.S236860
Yang G, Yin H, Lin F, Gao S, Zhan K, Tong H et al (2020) Long noncoding RNA TUG1 regulates prostate cancer cell proliferation, invasion and migration via the Nrf2 signaling axis. Pathol-Res Practice 216(4):152851. https://doi.org/10.1016/j.prp.2020.152851
Dong L, Ding H, Li Y, Xue D, Liu Y (2018) LncRNA TINCR is associated with clinical progression and serves as tumor suppressive role in prostate cancer. Cancer Manage Res 10:2799. https://doi.org/10.2147/CMAR.S170526
Beaver LM, Kuintzle R, Buchanan A, Wiley MW, Glasser ST, Wong CP et al (2017) Long noncoding RNAs and sulforaphane: a target for chemoprevention and suppression of prostate cancer. J Nutr Biochem 42:72–83. https://doi.org/10.1016/j.jnutbio.2017.01.001
Yang J, Hao T, Sun J, Wei P, Zhang H (2019) Long noncoding RNA GAS5 modulates α-Solanine-induced radiosensitivity by negatively regulating miR-18a in human prostate cancer cells. Biomed Pharmacother 112:108656. https://doi.org/10.1016/j.biopha.2019.108656
Chen C, Wang K, Wang Q, Wang X (2018) LncRNA HULC mediates radioresistance via autophagy in prostate cancer cells. Braz J Med Biol Res. https://doi.org/10.1590/1414-431X20187080
Lu J, Zhong C, Luo J, Shu F, Lv D, Liu Z et al (2021) HnRNP-L-regulated circCSPP1/miR-520h/EGR1 axis modulates autophagy and promotes progression in prostate cancer. Mol Ther-Nucl Acids 26:927–944. https://doi.org/10.1016/j.omtn.2021.10.006
Zhong C, Wu K, Wang S, Long Z, Yang T, Zhong W et al (2021) Autophagy-related circRNA evaluation reveals hsa_circ_0001747 as a potential favorable prognostic factor for biochemical recurrence in patients with prostate cancer. Cell Death Dis 12(8):1–12. https://doi.org/10.1038/s41419-021-04015-w
Cai F, Li J, Zhang J, Huang S (2020) Knockdown of Circ_CCNB2 sensitizes prostate cancer to radiation through repressing autophagy by the miR-30b-5p/KIF18A axis. Cancer Biother Radiopharm. https://doi.org/10.1089/cbr.2019.3538
Borchardt H, Ewe A, Morawski M, Weirauch U, Aigner A (2021) miR24–3p activity after delivery into pancreatic carcinoma cell lines exerts profound tumor-inhibitory effects through distinct pathways of apoptosis and autophagy induction. Cancer Lett 503:174–184. https://doi.org/10.1016/j.canlet.2021.01.018
Gu DN, Jiang MJ, Mei Z, Dai JJ, Dai CY, Fang C et al (2017) microRNA-7 impairs autophagy-derived pools of glucose to suppress pancreatic cancer progression. Cancer Lett 400:69–78. https://doi.org/10.1016/j.canlet.2017.04.020
Ye ZQ, Zou CL, Chen HB, Jiang MJ, Mei Z, Gu DN (2020) MicroRNA-7 as a potential biomarker for prognosis in pancreatic cancer. Dis Markers. https://doi.org/10.1155/2020/2782101
Huang L, Hu C, Cao H, Wu X, Wang R, Lu H et al (2018) MicroRNA-29c increases the chemosensitivity of pancreatic cancer cells by inhibiting USP22 mediated autophagy. Cell Physiol Biochem 47(2):747–758. https://doi.org/10.1159/000490027
Zhou C, Liang Y, Zhou L, Yan Y, Liu N, Zhang R et al (2021) TSPAN1 promotes autophagy flux and mediates cooperation between WNT-CTNNB1 signaling and autophagy via the MIR454-FAM83A-TSPAN1 axis in pancreatic cancer. Autophagy 17(10):3175–3195. https://doi.org/10.1080/15548627.2020.1826689
Wang ZC, Huang FZ, Xu HB, Sun JC, Wang CF (2019) MicroRNA-137 inhibits autophagy and chemosensitizes pancreatic cancer cells by targeting ATG5. Int J Biochem Cell Biol 111:63–71. https://doi.org/10.1016/j.biocel.2019.01.020
Wu Y, Tang Y, Xie S, Zheng X, Zhang S, Mao J et al (2020) Chimeric peptide supramolecular nanoparticles for plectin-1 targeted miRNA-9 delivery in pancreatic cancer. Theranostics 10(3):1151
Huang F, Chen W, Peng J, Li Y, Zhuang Y, Zhu Z et al (2018) LncRNA PVT1 triggers Cyto-protective autophagy and promotes pancreatic ductal adenocarcinoma development via the miR-20a-5p/ULK1 Axis. Mol Cancer 17(1):1–16. https://doi.org/10.1186/s12943-018-0845-6
Yang T, Shen P, Chen Q, Wu P, Yuan H, Ge W et al (2021) FUS-induced circRHOBTB3 facilitates cell proliferation via miR-600/NACC1 mediated autophagy response in pancreatic ductal adenocarcinoma. J Exp Clin Cancer Res 40(1):1–25. https://doi.org/10.1186/s13046-021-02063-w
Wei DM, Jiang MT, Lin P, Yang H, Dang YW, Yu Q et al (2019) Potential ceRNA networks involved in autophagy suppression of pancreatic cancer caused by chloroquine diphosphate: a study based on differentially-expressed circRNAs, lncRNAs, miRNAs and mRNAs. Int J Oncol 54(2):600–626. https://doi.org/10.3892/ijo.2018.4660
Gou L, Zou H, Li B (2019) Long noncoding RNA MALAT1 knockdown inhibits progression of anaplastic thyroid carcinoma by regulating miR-200a-3p/FOXA1. Cancer Biol Ther 20(11):1355–1365. https://doi.org/10.1080/15384047.2019.1617567
Abraham D, Jackson N, Gundara JS, Zhao J, Gill AJ, Delbridge L et al (2011) MicroRNA profiling of sporadic and hereditary medullary thyroid cancer identifies predictors of nodal metastasis, prognosis, and potential therapeutic targets. Clin Cancer Res 17(14):4772–4781. https://doi.org/10.1158/1078-0432.CCR-11-0242
Gundara JS, Zhao J, Gill AJ, Lee JC, Delbridge L, Robinson BG et al (2015) Noncoding RNA blockade of autophagy is therapeutic in medullary thyroid cancer. Cancer Med 4(2):174–182. https://doi.org/10.1002/cam4.355
Zhang Y, Yang WQ, Zhu H, Qian YY, Zhou L, Ren YJ et al (2014) Regulation of autophagy by miR-30d impacts sensitivity of anaplastic thyroid carcinoma to cisplatin. Biochem Pharmacol 87(4):562–570. https://doi.org/10.1016/j.bcp.2013.12.004
Liu J, Feng L, Zhang H, Zhang J, Zhang Y, Li S et al (2018) Effects of miR-144 on the sensitivity of human anaplastic thyroid carcinoma cells to cisplatin by autophagy regulation. Cancer Biol Ther 19(6):484–496. https://doi.org/10.1080/15384047.2018.1433502
Wang S, Wu J, Ren J, Vlantis AC, Li MY, Liu S, Ng E, Chan A, Luo DC, Liu Z, Guo W, Xue L, Ng SK, van Hasselt CA, Tong M, Chen GG (2018) MicroRNA-125b interacts with Foxp3 to induce autophagy in thyroid cancer. Mol Ther 26(9):2295–2303. https://doi.org/10.1016/j.ymthe.2018.06.015
Liu H, Chen X, Lin T, Chen X, Yan J, Jiang S (2019) MicroRNA-524-5p suppresses the progression of papillary thyroid carcinoma cells via targeting on FOXE1 and ITGA3 in cell autophagy and cycling pathways. J Cell Physiol 234(10):18382–18391. https://doi.org/10.1002/jcp.28472
Yang LX, Wu J, Guo ML, Zhang Y, Ma SG (2019) Suppression of long non-coding RNA TNRC6C-AS1 protects against thyroid carcinoma through DNA demethylation of STK4 via the Hippo signalling pathway. Cell Prolif 52(3):e12564. https://doi.org/10.1111/cpr.12564
Peng X, Ji C, Tan L, Lin S, Zhu Y, Long M, Luo D, Li H (2020) Long non-coding RNA TNRC6C-AS1 promotes methylation of STK4 to inhibit thyroid carcinoma cell apoptosis and autophagy via Hippo signalling pathway. J Cell Mol Med 24(1):304–316. https://doi.org/10.1111/jcmm.14728
Mila G, Manicardi V, Federica T, Elisabetta S, Riccardo B, Gloria M et al (2021) Linc00941 is a novel TGFβ target that primes papillary thyroid cancer metastatic behavior by regulating the expression of Cadherin. Thyroid. https://doi.org/10.1089/thy.2020.0001
Zhao Y, Zhao L, Li J, Zhong L (2019) Silencing of long noncoding RNA RP11-476D10. 1 enhances apoptosis and autophagy while inhibiting proliferation of papillary thyroid carcinoma cells via microRNA-138-5p-dependent inhibition of LRRK2. J Cell Physiol 234(11):20980–20991. https://doi.org/10.1002/jcp.28702
Wang HH, Ma JN, Zhan XR (2021) Circular RNA Circ_0067934 attenuates ferroptosis of thyroid cancer cells by miR-545–3p/SLC7A11 signaling. Front Endocrinol. https://doi.org/10.3389/fendo.2021.670031
Liu F, Zhang J, Qin L, Yang Z, Xiong J, Zhang Y et al (2018) Circular RNA EIF6 (Hsa_circ_0060060) sponges miR-144–3p to promote the cisplatin-resistance of human thyroid carcinoma cells by autophagy regulation. Aging 10(12):3806. https://doi.org/10.18632/aging.101674
Shi YP, Liu GL, Li S, Liu XL (2020) miR-17-5p knockdown inhibits proliferation, autophagy and promotes apoptosis in thyroid cancer via targeting PTEN. Neoplasma 67(2):249–258. https://doi.org/10.4149/neo_2019_190110N29
Gundara JS, Zhao J, Gill AJ, Lee JC, Delbridge L, Robinson BG, McLean C, Serpell J, Sidhu SB (2015) Noncoding RNA blockade of autophagy is therapeutic in medullary thyroid cancer. Cancer Med 4(2):174–182. https://doi.org/10.1002/cam4.355
Zhao Y, Zhao L, Li J, Zhong L (2019) Silencing of long noncoding RNA RP11–476D10.1 enhances apoptosis and autophagy while inhibiting proliferation of papillary thyroid carcinoma cells via microRNA-138–5p-dependent inhibition of LRRK2. J Cell Physiol 234(11):20980–20991. https://doi.org/10.1002/jcp.28702
Liu J, Feng L, Zhang H, Zhang J, Zhang Y, Li S, Qin L, Yang Z, Xiong J (2018) Effects of miR-144 on the sensitivity of human anaplastic thyroid carcinoma cells to cisplatin by autophagy regulation. Cancer Biol Ther 19(6):484–496. https://doi.org/10.1080/15384047.2018.1433502
Wen D, Liu WL, Lu ZW, Cao YM, Ji QH, Wei WJ (2021) SNHG9, a papillary thyroid cancer cell exosome-enriched lncRNA, inhibits cell autophagy and promotes cell apoptosis of normal thyroid epithelial cell Nthy-ori-3 through YBOX3/P21 pathway. Front Oncol 11:647034. https://doi.org/10.3389/fonc.2021.647034
Peng D, Li W, Zhang B, Liu X (2021) Overexpression of lncRNA SLC26A4-AS1 inhibits papillary thyroid carcinoma progression through recruiting ETS1 to promote ITPR1-mediated autophagy. J Cell Mol Med 25(17):8148–8158. https://doi.org/10.1111/jcmm.16545
Chen Y, Zhou J, Wu X, Huang J, Chen W, Liu D, Zhang J, Huang Y, Xue W (2020) miR-30a-3p inhibits renal cancer cell invasion and metastasis through targeting ATG12. Transl Androl Urol 9(2):646–653. https://doi.org/10.21037/tau.2019.12.10
Liu X, Zhong L, Li P, Zhao P (2020) MicroRNA-100 enhances autophagy and suppresses migration and invasion of renal cell carcinoma cells via disruption of NOX4-dependent mTOR pathway. Clin Transl Sci. https://doi.org/10.1111/cts.12798
Zheng B, Zhu H, Gu D, Pan X, Qian L, Xue B, Yang D, Zhou J, Shan Y (2015) MiRNA-30a-mediated autophagy inhibition sensitizes renal cell carcinoma cells to sorafenib. Biochem Biophys Res Commun 459(2):234–239. https://doi.org/10.1016/j.bbrc.2015.02.084
Takai T, Tsujino T, Yoshikawa Y, Inamoto T, Sugito N, Kuranaga Y, Heishima K, Soga T, Hayashi K, Miyata K, Kataoka K, Azuma H, Akao Y (2019) Synthetic miR-143 exhibited an anti-cancer effect via the downregulation of K-RAS networks of renal cell cancer cells in vitro and in vivo. Mol Ther 27(5):1017–1027. https://doi.org/10.1016/j.ymthe.2019.03.004
Mikhaylova O, Stratton Y, Hall D, Kellner E, Ehmer B, Drew AF, Gallo CA, Plas DR, Biesiada J, Meller J, Czyzyk-Krzeska MF (2012) VHL-regulated MiR-204 suppresses tumor growth through inhibition of LC3B-mediated autophagy in renal clear cell carcinoma. Cancer Cell 21(4):532–546. https://doi.org/10.1016/j.ccr.2012.02.019
Yan L, Liu G, Cao H, Zhang H, Shao F (2019) Hsa_circ_0035483 sponges hsa-miR-335 to promote the gemcitabine-resistance of human renal cancer cells by autophagy regulation. Biochem Biophys Res Commun 519(1):172–178. https://doi.org/10.1016/j.bbrc.2019.08.093
Jin Y, Huang R, Xia Y, Huang C, Qiu F, Pu J, He X, Zhao X (2020) Long noncoding RNA KIF9-AS1 regulates transforming growth factor-β and autophagy signaling to enhance renal cell carcinoma chemoresistance via microRNA-497-5p. DNA Cell Biol 39(7):1096–1103. https://doi.org/10.1089/dna.2020.5453
Shao Q, Wang Q, Wang J (2019) LncRNA SCAMP1 regulates ZEB1/JUN and autophagy to promote pediatric renal cell carcinoma under oxidative stress via miR-429. Biomed Pharmacother 120:109460. https://doi.org/10.1016/j.biopha.2019.109460
Li D, Li C, Chen Y, Teng L, Cao Y, Wang W, Pan H, Xu Y, Yang D (2020) LncRNA HOTAIR induces sunitinib resistance in renal cancer by acting as a competing endogenous RNA to regulate autophagy of renal cells. Cancer Cell Int 20:338. https://doi.org/10.1186/s12935-020-01419-0
Su Y, Lu J, Chen X, Liang C, Luo P, Qin C, Zhang J (2019) Long non-coding RNA HOTTIP affects renal cell carcinoma progression by regulating autophagy via the PI3K/Akt/Atg13 signaling pathway. J Cancer Res Clin Oncol 145(3):573–588. https://doi.org/10.1007/s00432-018-2808-0
Gui CP, Cao JZ, Tan L, Huang Y, Tang YM, Li PJ, Chen YH, Lu J, Yao HH, Chen ZH, Pan YH, Ye YL, Qin ZK, Chen W, Wei JH, Luo JH (2021) A panel of eight autophagy-related long non-coding RNAs is a good predictive parameter for clear cell renal cell carcinoma. Genomics 113(2):740–754. https://doi.org/10.1016/j.ygeno.2021.01.016
Sahin Lacin EE, Karakas Y, Yalcin S (2015) Metastatic medullary thyroid cancer: a dramatic response to a systemic chemotherapy (temozolomide and capecitabine) regimen. OncoTargets Ther 8:1039. https://doi.org/10.2147/OTT.S82906
Funding
This study was not funded by any authority.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Barnwal, S.K., Bendale, H. & Banerjee, S. Non-coding RNAs associated with autophagy and their regulatory role in cancer therapeutics. Mol Biol Rep 49, 7025–7037 (2022). https://doi.org/10.1007/s11033-022-07517-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07517-8