Abstract
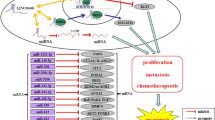
Long non-coding RNAs (lncRNAs) are a family of non-protein-coding RNAs with length more than 200 nucleotides. LncRNAs played important roles in many biological processes such as cell development, proliferation, invasion and migration. Deregulation of LncRNAs was found in multiple tumors where they can act as a tumor suppressor gene or oncogene. LncRNA ANRIL was identified as an oncogene involved in a number of tumors such as gastric cancer, lung cancer, hepatocellular carcinoma, and esophageal squamous cell carcinoma. Inhibition of ANRIL suppressed the cancer cell proliferation, migration and invasion. Increasing data has showed that ANRIL may act as a diagnostic and prognostic biomarker for some tumors. In our review, we summarize an overview of current knowledge concerning the expression and role of ANRIL in various cancers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Non-coding DNA accounts for approximately 98.5 % of the total human genomic DNA [1, 2]. Increasing evidences have demonstrated that transcription is also occurs in the non-coding regions [3–5]. Recent studies have showed that non-coding RNAs such as long non-coding RNA (lncRNA) and microRNA play a pivotal role in tumor development and progression [6–12]. LncRNAs are a group of non-protein-coding RNAs with more than 200 nucleotides in length [13, 14]. Accumulating studies have revealed that lncRNAs are involved in various biological and pathological processes including development, cell proliferation, metastasis, fate decision, invasion and migration [15–19]. Additionally, the deregulation of lncRNAs is identified in many cancers such as gastric cancer, hepatocellular carcinoma, lung cancer, renal cell carcinoma, and breast cancer [17, 20–23]. These lncRNAs act as tumor suppressors or oncogenes in the development of cancers [22, 24, 25].
ANRIL (CDKN2B antisense RNA 1) was originally identified in the familial melanoma patients [26]. Since its identification, accumulating studies have showed that ANRIL is deregulated in a number of malignancies such as gastric, breast, lung and bladder cancer [27–30]. In our review, we comment the current studies about the expression and role of lncRNA ANRIL in the development and progression of cancers.
Structural characterization of ANRIL
ANRIL was originally identified from familial melanoma patients with a large of germline deletion in the INK4B-ARF-INK4A gene cluster, which was located a 42-kb stretch on the chromosome 9p21 [26, 31–33]. This gene locus is transcriptionally silenced or homozygously deleted in a lot of tumors with a frequency of about 40 % exhibiting one of the most diversify genes in human tumors [31, 32]. The INK4B-ARF-INK4A locus is governed by PRCs and ANRIL is involved in suppressing this locus [34, 35].
ANRIL spans a region of 126.3 kb and is transcribed as a 3.8-kb lncRNA in the opposite direction from the INK4B-ARF-INK4A gene cluster [26]. ANRIL contains 19 exons and two exons of INK4B are overlapped [36]. The first exon of ANRIL is localized in the −300 bp upstream of ARF in the transcription start site [26, 37]. There is a positive correlation between the expression of ARF and ANRIL in cancers and normal tissue [38–40].
ANRIL in human cancers
Gastric cancer
Zhang et al. [27] demonstrated that ANRIL was upregulated in gastric cancer tissues. In addition, the expression level of ANRIL was positively correlated with tumor size and a higher TNM stage in a cohort of 120 GC patients. Multivariate analyses showed that the expression of ANRIL acted as an independent predictor for OS (overall survival). Moreover, inhibition of ANRIL suppressed GC cell proliferation both in vivo and in vitro. Overexpression of E2F1 promoted ANRIL and ANRIL-induce growth and was in part owing to epigenetic suppression of miR-449a/miR-99a through binding to PRC2. These data suggested that ANRIL acted as an oncogene in the development of gastric cancer.
Lung cancer
Nie et al. [41] demonstrated that ANRIL was upregulated in non-small cell lung cancer (NSCLC), and its expression status was positively correlated with tumor size and tumor node metastasis stages. In addition, patients with higher expression level of ANRIL had a poor prognosis. Moreover, knockdown of ANRIL suppressed NSCLC cell proliferation and promoted cell apoptosis both in vivo and in vitro. They also showed that overexpression of ANRIL could not suppress p15 expression in PC9 cells, but inhibited P21 and KLF2 transcription. Their data suggested that ANRIL played as an oncogene role in development of NSCLC.
Further study also demonstrated that ANRIL played an oncogenic role in NSCLC progression [42]. ANRIL was upregulated in NSCLC tissues and cells compared to the adjacent non-tumor tissues and normal bronchial epithelial cells. The expression level of ANRIL was positively associated with advanced lymph node metastasis and higher TNM stage. Moreover, patients with high ANRIL expression had shorter OS compared to low ANRIL patients. They also proved that high expression of ANRIL was an independent poor prognostic maker for patients with NSCLC. In addition, inhibition of ANRIL suppressed ANRIL cell migration, proliferation and invasion.
Another study demonstrated that ANRIL was upregulated after phospholipase D (PLD) inhibition in lung cancer cells [43]. Inhibition of ANRIL significantly inhibited PLD suppression-induced apoptosis. These findings demonstrated that ANRIL was responsible in anti-tumorigenesis induced by PLD inhibition. Recent research also found that inhibition of ANRIL could inhibit NSCLC proliferation [29].
Hepatocellular carcinoma
Huang et al. [44] demonstrated that ANRIL was upregulated in hepatocellular carcinoma (HCC) tissues and the expression level of ANRIL was positively associated with stage of Barcelona Clinic Liver Cancer (BCLC) and tumor size. They also found that knockdown of ANRIL suppressed HCC cell invasion and proliferation and induced cell apoptosis both in vivo and vitro. ANRIL inhibited the KLF2 (Kruppel-like factor 2) expression through binding with PRC2 and recruiting PRC2 to the promoter region of KLF2. They also showed that SP1 could regulate ANRIL expression.
Hua et al. [45] also found that the expression of ANRIL was higher in HCC tissues than in the adjacent non-tumor tissues. The expression of ANRIL was correlated with the TNM stage and histologic grade of HCC patients. Moreover, HCC patients with higher ANRIL expression had shorter OS. Multivariate analysis demonstrated that high ANRIL expression acted as an independent predictor for poor prognosis. Furthermore, they demonstrated that the downregulation of ANRIL could inhibit HCC cell proliferation, invasion and migration.
Esophageal squamous cell carcinoma
Chen et al. [46] found that ANRIL was upregulated in human esophageal squamous cell carcinoma (ESCC) tissues compared to adjacent non-tumor tissues. Moreover, downregulation of ANRIL could suppress cell proliferation and promote the expression of transforming growth factor β1 (TGFβ1) and p15 (INK4b) in ESCC cell. Another study genotyped the ANRIL rs2151280 T/C in 380 ESCC patients and 380 controls. They found that ANRIL rs2151280 T/C SNP was not associated with risk of ESCC [47].
Cervical cancer
Naemura et al. [29] demonstrated that ANRIL was upregulated in cervical cancer cells. Depletion of ANRIL promoted p15 expression, with no effect on ARF (alternative reading frame) or p16 expression. Depletion of ANRIL also induced cell-cycle arrest at the G2/M phase, suppressing the cell proliferation of H1299 and cervical cancer cells.
Melanoma
Pasmant et al. [26] demonstrated that there was deletion the region that contained the entire p16/CDKN2A, p15/CDKN2B and p14/ARF genes. Moreover, ANRIL was proved to locate within the 403-kb germline deletion, with a first exon located in the promoter of the p14/ARF gene. ANRIL expression was mainly co-clustered with p14/ARF both in physiologic and in pathologic conditions.
Bladder cancer
Zhu et al. [30] showed that ANRIL was upregulated in the bladder cancer tissues compared to adjacent non-tumor tissues. Moreover, inhibition of ANRIL suppressed bladder cancer cell proliferation and promoted cell apoptosis. It also suppressed Bcl-2 expression and induced expression of the cytoplasmic cytochrome c, Bax and Smac and caspase-3, cleaved caspase-9, and PARP in bladder cancer cell. Furthermore, they demonstrated that knockdown of ANRIL suppressed tumorigenic ability of bladder cancer cell line EJ cell in nude mice. This data proved ANRIL acted as an oncogene in bladder cancer. Another study showed that ANRIL had no determining role in high-graded and recurrent tumors associated with bad prognosis [48].
Ovarian cancer
Qiu et al. [49] demonstrated that ANRIL was upregulated in serous ovarian cancer (SOC) tissues compared to normal tissues. Moreover, the expression level of ANRIL was positively associated with high histological grade, advanced FIGO stage, poor prognosis and lymph node metastasis. More importantly, multivariate analysis showed that the expression ANRIL was an independent prognostic maker for predicting overall survival of SOC. They also showed that ANRIL was upregulated in the highly metastatic sublines compared with parental cell lines. Furthermore, inhibition of ANRIL suppressed SOC cell invasion and migration. MMP3 and MET were identified as the downstream genes of ANRIL.
Lymphoblastic leukemia
The rs564398 was mapped to the CDKN2BAS locus that encoded ANRIL antisense non-coding RNA. Lacobucci et al. [50] demonstrated that rs564398 showed a statistically correlation with the ALL (acute lymphoblastic leukemia) phenotype.
Neurofibromatosis
Pasmant et al. [51] showed that 9p21.3 deletions (CDKN2A/B-ANRIL locus) were the only recurrent somatic alterations in the Neurofibromatosis type 1 (NF1)-associated plexiform neurofibromas (PNFs). SNP rs2151280 was coated in ANRIL and was significantly correlated with the number of PNFs in NF1 patients. In addition, allele T of rs2151280 was associated with reduced ANRIL transcript levels. Another study also found that neither the PNF number nor PNF volume was found to be correlated with the T-allele of rs2151280 in 29 microdeletion patients [52] (Table 1).
Concluding remarks and future perspectives
ANRIL, upregulated in a lot of human tumors, was relatively a well-characterized oncogene. ANRIL can suppress cancer cell proliferation, invasion and migration and promoted cancer cell apoptosis. However, the detail molecular mechanism of ANRIL remains to be studied. It is hopeful that ANRIL will achieve clinical utility at last with more study.
References
Gibb EA, Brown CJ, Lam WL. The functional role of long non-coding RNA in human carcinomas. Mol Cancer. 2011;10:38.
Wan X, Ding X, Chen S, Song H, Jiang H, Fang Y, et al. The functional sites of miRNAs and lncRNAs in gastric carcinogenesis. Tumour Biol. 2015;36:521–32.
Tano K, Akimitsu N. Long non-coding RNAs in cancer progression. Front Genet. 2012;3:219.
Li CH, Chen Y. Targeting long non-coding RNAs in cancers: progress and prospects. Int J Biochem Cell Biol. 2013;45:1895–910.
Kitagawa M, Kotake Y, Ohhata T. Long non-coding RNAs involved in cancer development and cell fate determination. Curr Drug Targets. 2012;13:1616–21.
Gutschner T, Diederichs S. The hallmarks of cancer: a long non-coding RNA point of view. RNA Biol. 2012;9:703–19.
Taylor DH, Chu ET, Spektor R, Soloway PD. Long non-coding RNA regulation of reproduction and development. Mol Reprod Dev. 2015. doi:10.1002/mrd.22581.
Yang Y, Wen L, Zhu H. Unveiling the hidden function of long non-coding RNA by identifying its major partner-protein. Cell Biosci. 2015;5:59.
Yu X, Li Z. The role of miRNAs in cutaneous squamous cell carcinoma. J Cell Mol Med. 2015. doi:10.1111/jcmm.12649.
Li Z, Yu X, Shen J. The role of miRNAs in the pheochromocytomas. Tumour Biol. 2015. doi:10.1007/s13277-015-4199-z.
Yu X, Li Z. MicroRNA expression and its implications for diagnosis and therapy of tongue squamous cell carcinoma. J Cell Mol Med. 2015. doi:10.1111/jcmm.12650.
Li Z, Yu X, Shen J, Jiang Y. MicroRNA dysregulation in uveal melanoma: a new player enters the game. Oncotarget. 2015;6:4562–8.
Yu X, Li Z. Long non-coding RNA growth arrest-specific transcript 5 in tumor biology. Oncol Lett. 2015;10:1953–8.
Yu X, Li Z. Long non-coding RNA HOTAIR: a novel oncogene (review). Mol Med Rep. 2015;12:5611–8.
Silva A, Bullock M, Calin G. The clinical relevance of long non-coding RNAs in cancer. Cancers. 2015;7:2169–82.
Wang G, Liu C, Deng S, Zhao Q, Li T, Qiao S, et al. Long noncoding rnas in regulation of human breast cancer. Brief Funct Genomics. 2015. doi:10.1093/bfgp/elv049.
Tao H, Yang JJ, Zhou X, Deng ZY, Shi KH, Li J. Emerging role of long non-coding RNAs in lung cancer: current status and future prospects. Respir Med. 2015;110:12–9.
Meseure D, Drak Alsibai K, Nicolas A, Bieche I, Morillon A. Long non-coding RNAs as new architects in cancer epigenetics, prognostic biomarkers, and potential therapeutic targets. Biomed Res Int. 2015;2015:320214.
Shahryari A, Jazi MS, Samaei NM, Mowla SJ. Long non-coding RNA SOX2OT: expression signature, splicing patterns, and emerging roles in pluripotency and tumorigenesis. Front Genet. 2015;6:196.
Ding X, Wan X, Jiang H, Song H, Fang Y, Chen S, et al. The clinical value of ncRNAs in gastric cancer: a systematic review and meta-analyses. Tumour Biol. 2015;36:4017–25.
Hajjari M, Salavaty A. HOTAIR: an oncogenic long non-coding RNA in different cancers. Cancer Biol Med. 2015;12:1–9.
Yang X, Xie X, Xiao YF, Xie R, Hu CJ, Tang B, et al. The emergence of long non-coding RNAs in the tumorigenesis of hepatocellular carcinoma. Cancer Lett. 2015;360:119–24.
Martens-Uzunova ES, Bottcher R, Croce CM, Jenster G, Visakorpi T, Calin GA. Long non-coding RNA in prostate, bladder, and kidney cancer. Eur Urol. 2014;65:1140–51.
Yarmishyn AA, Kurochkin IV. Long non-coding RNAs: a potential novel class of cancer biomarkers. Front Genet. 2015;6:145.
Di Gesualdo F, Capaccioli S, Lulli M. A pathophysiological view of the long non-coding RNA world. Oncotarget. 2014;5:10976–96.
Pasmant E, Laurendeau I, Heron D, Vidaud M, Vidaud D, Bieche I. Characterization of a germ-line deletion, including the entire INK4/ARF locus, in a melanoma-neural system tumor family: identification of ANRIL, an antisense non-coding RNA whose expression coclusters with ARF. Cancer Res. 2007;67:3963–9.
Zhang EB, Kong R, Yin DD, You LH, Sun M, Han L, et al. Long non-coding RNA ANRIL indicates a poor prognosis of gastric cancer and promotes tumor growth by epigenetically silencing of mir-99a/mir-449a. Oncotarget. 2014;5:2276–92.
Iranpour M, Soudyab M, Geranpayeh L, Mirfakhraie R, Azargashb E, Movafagh A, et al. Expression analysis of four long non-coding RNAs in breast cancer. Tumour Biol. 2015. doi:10.1007/s13277-015-4135-2.
Naemura M, Murasaki C, Inoue Y, Okamoto H, Kotake Y. Long non-coding RNA ANRIL regulates proliferation of non-small-cell lung cancer and cervical cancer cells. Anticancer Res. 2015;35:5377–82.
Zhu H, Li X, Song Y, Zhang P, Xiao Y, Xing Y. Long non-coding RNA ANRIL is upregulated in bladder cancer and regulates bladder cancer cell proliferation and apoptosis through the intrinsic pathway. Biochem Biophys Res Commun. 2015;467:223–8.
Kia SK, Gorski MM, Giannakopoulos S, Verrijzer CP. SWI/SNF mediates polycomb eviction and epigenetic reprogramming of the INK4b-ARF-INK4a locus. Mol Cell Biol. 2008;28:3457–64.
Gil J, Peters G. Regulation of the INK4b-ARF-INK4a tumour suppressor locus: all for one or one for all. Nat Rev Mol Cell Biol. 2006;7:667–77.
Ozenne P, Eymin B, Brambilla E, Gazzeri S. The ARF tumor suppressor: structure, functions and status in cancer. Int J Cancer. 2010;127:2239–47.
Kheradmand Kia S, Solaimani Kartalaei P, Farahbakhshian E, Pourfarzad F, von Lindern M, Verrijzer CP. EZH2-dependent chromatin looping controls INK4a and INK4b, but not ARF, during human progenitor cell differentiation and cellular senescence. Epigenetics Chromatin. 2009;2:16.
Yap KL, Li S, Munoz-Cabello AM, Raguz S, Zeng L, Mujtaba S, et al. Molecular interplay of the non-coding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol Cell. 2010;38:662–74.
Aguilo F, Di Cecilia S, Walsh MJ. Long non-coding RNA ANRIL and polycomb in human cancers and cardiovascular disease. Curr Top Microbiol Immunol. 2015. doi:10.1007/82_2015_455.
He S, Gu W, Li Y, Zhu H. ANRIL/CDKN2B-AS shows two-stage clade-specific evolution and becomes conserved after transposon insertions in simians. BMC Evol Biol. 2013;13:247.
Aguilo F, Zhou MM, Walsh MJ. Long non-coding RNA, polycomb, and the ghosts haunting INK4b-ARF-INK4a expression. Cancer Res. 2011;71:5365–9.
Kotake Y, Nakagawa T, Kitagawa K, Suzuki S, Liu N, Kitagawa M, et al. Long non-coding RNA ANRIL is required for the PRC2 recruitment to and silencing of p15(INK4b) tumor suppressor gene. Oncogene. 2011;30:1956–62.
Rodriguez C, Borgel J, Court F, Cathala G, Forne T, Piette J. CTCF is a DNA methylation-sensitive positive regulator of the INK/ARF locus. Biochem Biophys Res Commun. 2010;392:129–34.
Nie FQ, Sun M, Yang JS, Xie M, Xu TP, Xia R, et al. Long non-coding RNA ANRIL promotes non-small cell lung cancer cell proliferation and inhibits apoptosis by silencing KLF2 and p21 expression. Mol Cancer Ther. 2015;14:268–77.
Lin L, Gu ZT, Chen WH, Cao KJ. Increased expression of the long non-coding RNA ANRIL promotes lung cancer cell metastasis and correlates with poor prognosis. Diagn Pathol. 2015;10:14.
Kang YH, Kim D, Jin EJ. Down-regulation of phospholipase d stimulates death of lung cancer cells involving upregulation of the long ncRNA anril. Anticancer Res. 2015;35:2795–803.
Huang MD, Chen WM, Qi FZ, Xia R, Sun M, Xu TP, et al. Long non-coding RNA ANRIL is upregulated in hepatocellular carcinoma and regulates cell apoptosis by epigenetic silencing of klf2. J Hematol Oncol. 2015;8:50.
Hua L, Wang CY, Yao KH, Chen JT, Zhang JJ, Ma WL. High expression of long non-coding RNA ANRIL is associated with poor prognosis in hepatocellular carcinoma. Int J Clin Exp Pathol. 2015;8:3076–82.
Chen D, Zhang Z, Mao C, Zhou Y, Yu L, Yin Y, et al. ANRIL inhibits p15(INK4b) through the TGF beta 1 signaling pathway in human esophageal squamous cell carcinoma. Cell Immunol. 2014;289:91–6.
Kang M, Sang Y, Gu H, Zheng L, Wang L, Liu C, et al. Long non-coding RNAs POLR2Ee rs3787016 c/t and HULC rs7763881 A/C polymorphisms are associated with decreased risk of esophageal cancer. Tumour Biol. 2015;36:6401–8.
Martinez-Fernandez M, Feber A, Duenas M, Segovia C, Rubio C, Fernandez M, et al. Analysis of the polycomb-related lncRNAs HOTAIR and ANRIL in bladder cancer. Clin Epigenetics. 2015;7:109.
Qiu JJ, Lin YY, Ding JX, Feng WW, Jin HY, Hua KQ. Long non-coding RNA ANRIL predicts poor prognosis and promotes invasion/metastasis in serous ovarian cancer. Int J Oncol. 2015;46:2497–505.
Iacobucci I, Sazzini M, Garagnani P, Ferrari A, Boattini A, Lonetti A, et al. A polymorphism in the chromosome 9p21 ANRIL locus is associated to Philadelphia positive acute lymphoblastic leukemia. Leuk Res. 2011;35:1052–9.
Pasmant E, Sabbagh A, Masliah-Planchon J, Ortonne N, Laurendeau I, Melin L, et al. Role of non-coding RNA ANRIL in genesis of plexiform neurofibromas in neurofibromatosis type 1. J Natl Cancer Inst. 2011;103:1713–22.
Mussotter T, Kluwe L, Hogel J, Nguyen R, Cooper DN, Mautner VF, et al. Non-coding RNA ANRIL and the number of plexiform neurofibromas in patients with NF1 microdeletions. BMC Med Genet. 2012;13:98.
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (NSFC) (Grant Numbers: 81401847, 81272053 and 81330044).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
None
Additional information
Zheng Li and Xin Yu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Li, Z., Yu, X. & Shen, J. ANRIL: a pivotal tumor suppressor long non-coding RNA in human cancers. Tumor Biol. 37, 5657–5661 (2016). https://doi.org/10.1007/s13277-016-4808-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-016-4808-5