Abstract
Millets are small seeded cereal crops predominantly cultivated and consumed by resource-poor farmers in the semi-arid tropics of Asia and Africa. Millets possess rich nutrients and a climate resilience property when compared to the other cereals such as rice and wheat. Millet improvement using modern genetic and genomic tools is falling behind other cereal crops due to their cultivation being restricted to less developed countries. Genome editing tools have been successfully applied to major cereal crops and, as a result, many key traits have been introduced into rice, wheat and maize. However, genome editing tools have not yet been used for most millets although they possess rich nutrients. The foxtail millet is the only millet utilised up to now for genome editing works. Limited genomic resources and lack of efficient transformation systems may slow down genome editing in millets. As millets possess many important traits of agricultural importance, high resolution studies with genome editing tools will help to understand the specific mechanism and transfer such traits to major cereals in the future. This review covers the current status of genome editing studies in millets and discusses the future prospects of genome editing in millets to understand key traits of nutrient fortification and develop climate resilient crops in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Millets are small-seeded cereal crops widely cultivated in the semi-arid tropics of Asia and Africa. Millets are key crops for strengthening food and nutrient security in the drought prone areas of these regions. The most frequently cultivated millets are finger millet (Eleusine coracana (L.) Gaertn.), foxtail millet (Setaria italica (L.) P. Beauvois), pearl millet (Cenchrus americanus (L.) Morrone.), kodo millet (Paspalum scrobiculatum L.), little millet (Panicum sumatrense Roth ex Roem. and Schult.), proso millet (Panicum miliaceum L.), and barnyard millet (Echinochloa crusgalli (L.) P. Beauvois). Millets have not been utilized frequently for modern genetic and genomic studies since they are cultivated in less developed countries of Asia and Africa. Genome sequences of millets also have only been released in recent years, a few decades after the release of genome sequences of stable cereals like rice [1]. However, millets have started receiving attention in recent years, especially from researchers in Europe and America, due to their dense nutrients and climate resilient properties [2].
Genome editing has emerged as a central tool for modern plant genetic studies. The recently developed and Nobel Prize winning genome editing tool, clustered regularly interspaced palindromic repeats (CRISPR) and CRISPR associated protein 9 (Cas9), has emerged as the most popular tool for genome editing in a diverse range of organisms [3,4,5]. The CRISPR/Cas system has been championed to accelerate crop improvement, with an option for precision breeding [6,7,8,9]. Many labs around the world have quickly adopted the CRISPR/Cas system and successfully produced mutant plants and scores of reports are available on crop genome editing (reviewed in [10, 11]).
Millets have been generally considered to be orphan crops due to the lower importance they have been awarded for crop improvement programs in the green revolution and modern genetic studies. For example, genetic engineering and genome sequencing works are lagging far behind in millets when compared with other major cereals [1, 2, 12]. Apart from foxtail millet, other millets do not have complete and annotated genome sequences and this hampers modern genetic studies, including the application of genome editing in these crops. Many traits associated with climate resilience and nutrient enrichment have not yet been studied in millets with high resolution molecular studies. Studying the molecular mechanism of abiotic stress tolerance and nutrient enrichment with the help of precise genome editing will be helpful to understand the mechanism and transfer such traits to other cereals. The availability of fully annotated genome sequence and efficient transformation system are the key factors to apply the genome editing in millets. In this review, nutrient benefits of millets, the scope of genome editing in millets, the need for genome editing in millets and future prospects for genome editing in millets are discussed.
Nutritional importance of millets
Nutrient contents of millets are superior to main cereals like rice and wheat [13]. Seeds of millets are the richest source of several nutrients like calcium, magnesium, phosphorus, iron, and proteins [1, 14, 15]. Millets also have high amounts of essential amino acids, dietary fibers, minerals, and vitamins [16]. For example, proso millet is a rich source of protein (12.5%), barnyard millet has highest crude fiber (13.6%) and iron (186 mg/kg dry matter) [17]. Finger millet is a rich source of calcium, magnesium, and potassium [14, 17]. It is noteworthy to mention that the calcium content of finger millet (344 mg) is the highest among cereals [1, 18]. Foxtail millet is also rich in protein (11%) and fat (4%) [19]. Pearl millet has high amounts of zinc, iron, and lysine [20]. Kodo millet is rich in magnesium (1.1 g/kg dry matter) and essential amino acids such as lysine, threonine, valine, and sulfur-containing amino acids [21]. Consumption of millets minimizes the risk of diseases like duodenal ulcers, anaemia, constipation, and atopic dermatitis [22, 23]. It has been predicted that consumption of millets reduced the incidence of cardiovascular complaints, diabetes, and certain cancer diseases [24, 25]. Consequently, millets are considered as nutra-cereals due to their nutrient dense seeds providing potential health benefits. Studying millets with genome editing tools will help to understand key roles of nutrient fortification.
Genome editing for agricultural improvement
Genome editing is a recent addition to the toolbox for plant breeders to develop new and improved varieties of crops. The commonly used genome editing tools include zinc finger nucleases (ZFNs), transcription activator like effector nucleases (TALENs), and the most popular, the CRISPR/Cas system [26]. Among these three systems, CRISPR/Cas system has been widely adopted by many labs for efficient genome editing due to its simple, user friendly, and cost effective construct designing [27,28,29]. Researchers can choose from the wide choices of constructs available for academic research at Addgene, a non-profit plasmid repository (https://www.addgene.org/). Basic plant biotech labs that have expertise with basic cloning and plant transformation experiments can successfully use CRISPR/Cas system-mediated genome editing due to its user friendly construct design option. This has opened a broad scope for the variety of genome editing works in many plant species.
Genome editing tools have been predicted to solve many issues of agriculture. In particular, the CRISPR-Cas9 system has been predicted to play a key role in imparting new traits in crop plants. Many excellent reviews are available on possible application of genome editing for crop improvement [11, 29,30,31,32]. Genome editing has been believed to benefit agriculture by imparting key traits including seed quality, drought tolerance, higher yield, disease resistance, improved nutrient uptake, and herbicide tolerance. For example, in rice, through knocking out SWEET genes that are responsible for disease susceptibility, researchers imparted resistance to bacterial blight [33, 34]. Improving yield is the primary objective of plant breeding, and genome editing has been considered to play a key role in this aim. Key genes responsible for yield improvement could help improve the yield. For example, editing genes responsible for tillering is believed to improve the yield of cereals [29, 31, 32, 35,36,37]. Genome editing could also be harnessed for enhancing the quality of seeds with lower gluten content, enriched carotenoid, and reduced phytic acid levels. Similarly, genome editing could help to overcome other stresses impacting crop production like drought, salinity, and cold, and could offer herbicide tolerance [11]. Imparting these types of novel traits by genome editing for enriching seed nutrient content with yield improvement will ensure both food and nutritional security.
Genome editing in major cereals
The CRISPR/Cas system has been applied to many cereal crops for crop improvement [7]. Many important traits have been imparted in cereals like rice, maize, and wheat through CRISPR/Cas9-mediated genome editing (Fig. 1). Genome editing tools have been successfully applied to many cereal crops and in particular scores of reports are available on the use of the CRISPR/Cas system (Reviewed in [7, 38, 39, 39]). Many studies were conducted on testing the genome editing tools with non-functional genes in major cereals like rice and wheat [7]. However, a few traits have also been imparted by precision genome editing in rice, barley, wheat, and maize (reviewed in [38]). These include development of fungal resistance in wheat [40,41,42], blast resistance in rice [43], grain nutrient quality improvement in rice [31, 44, 45] and maize [46], resistance to various biotic and abiotic stresses in wheat [47] and rice [48,49,50,51]. Major cereals like rice, maize, and wheat continue to get more attention for genome editing for improvement of various traits.
Need for millet genome editing
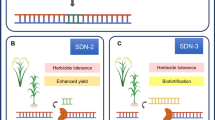
Application of genome editing tools in millets is helpful to improve millet research in the future. Genome editing may help in understanding the mechanism of nutrient fortification and climate resilience and to exchange such useful traits with other cereals. As discussed above, functional genomics studies of millets are lagging far behind when compared to those in main-stream cereals. However, millets possess superior traits to main cereals for agricultural performance and seed nutrient profile. Genes and QTL responsible for such traits need to be dissected through high resolution functional genomic studies with the help of genome editing tools. This will help to identify and transfer such traits to main-stream cereals. Millets show more plasticity in the field under adverse climate conditions [52]. Rice and wheat cannot produce good yield above 37 °C but millets like pearl millet can grow up to 46 °C, a very crucial trait to cope with drought induced by global warming [52]. When compared with main cereals like rice and wheat, millets are short duration crops demanding less maintenance in the field. Similarly, millets require minimum water and rainfall compared to major cereals. These traits have not yet been dissected by functional genomics studies. High resolution functional genomics studies aided by genome editing may help to unravel the candidate genes responsible for these traits and would help to transfer such traits to main cereals. Exceptionally higher accumulation of mineral nutrients like calcium (finger millet) and iron (barnyard millet) and B-vitamins in millet seeds need to be studied with genome editing tools to identify the transporters and signals responsible for the seed fortification. These traits could be transferred to within and outside the millet cereals to strengthen food security. An illustration on the application of various CRISPR/Cas systems for functional genomics studies and trait improvement in millets is included (Fig. 2).
Application of CRISPR/Cas system mediated genome editing in millet improvement. Use of various CRISPR/Cas systems to edit the genomes of various millets to understand functions of genes and improve the millet is illustrated. A Gene knock-out studies with the use of the CRISPR-Cas9 system is helpful to study the functions of key genes involved in climate resilience and the nutrient fortification process. B Gene knock-in studies with CRISPR-Cas9 aided by homologous recombination (HR) is useful to add novel genes and replace the defective portion of a gene to improve the performance. Base editing with the help of cytidine (C) and adenosine (D) base editors may help to change the ligand binding site residues of key nutrient transporters for improving nutrient transport in plants growing in low nutrient soils. E Expression of genes could be regulated at the transcriptional level using death Cas9 (dCas9) fused with either transcriptional activator or repressor to induce and down regulate respectively the specific genes in millets
Although millets possess climate resilient properties, their yield is affected by low nutrient soils since they are mostly cultivated under low input agriculture systems by resource poor farmers. For example, zinc deficiency severely affects the productive stage of pearl millet. It reduces the size of the panicle and delays the development and maturation of the grain [53]. Phosphate and nitrogen deficiencies significantly reduce the growth and yield of foxtail millet [54, 55]. Particularly, phosphate deficiency also significantly influences the size of the panicle and reduces the grain yield of all millets [56]. Precision genome editing like base editing could help to improve nutrient transport and improve the yield of millets under low nutrient soils in semi-arid regions of Asia and Africa.
Current status of genome editing in millets
Among the cereal crops, most reports are available for genome editing in rice. This is followed by wheat, maize, and barley based on a recent Pubmed search. But only 2 reports are available until now on genome editing in millets (Fig. 3). It may be due to lack of funding and the fact that millets are cultivated and consumed by less developed countries in Asia and Africa. It is also one of the reasons for delayed sequencing of millet genomes. Foxtail millet is the only millet which was utilized for genome editing studies. Foxtail millet is considered as a model crop due to its small genome (~ 450 mb), diploid in nature and has C4 photosynthesis chemistry [57]. Genome sequences of two different foxtail millet genotypes were released in 2012 and the genome is also completely annotated [58, 59]. Many genetic studies have been conducted in foxtail millet as it is a model C4 plant with a small genome. As a result, foxtail millet is the first millet utilized for genome editing studies. The first genome editing study using CRISPR/Cas9 was reported in foxtail millet during 2018 by Lin et al. [60]. They have used the protoplasts of several monocot and dicot plants including foxtail millet to assess CRISPR/Cas9-mediated genome editing in single cells (protoplast). The phytoene desaturase gene of foxtail millet was targeted by the plasmid pCAMBIA1300-35s-Cas9-OsU3-SiPDS. The targeted mutagenesis was detected by RFLP and sequencing [60]. Both deletion of bases up to 43 with insertion of single base were detected by sequencing results [60]. This is the first report on the successful application of the CRISPR/Cas9 system in foxtail millet.
Recently, CRISPR/Cas9-mediated genome editing has been applied in foxtail millet for the induction of double haploid (DH) lines by targeting the S. italica Matrilineal (SiMTL) gene [61]. The exonic region of SiMTL was targeted with two different gRNAs and was expressed under the control of the OsU3 promoter, with Cas9 kept under the control of the Uq promoter of maize. Foxtail millet genotype Ci846 was used for this study and the transformation was mediated by Agrobacterium. The authors reported a haploid induction frequency of 2.8% in the T2 progeny. It is encouraging to note that foxtail millet, being a model C4 crop, is utilized for genetic studies by genome editing. Apart from foxtail millet, other millets have not yet been used for high resolution genetic studies by genome editing.
Future prospects of millet genome editing
Considering the geographical area of cultivation, available resources, and the expertise of native researchers, millet genome editing is expected to progress at a slow pace. Due to limited expertise and infrastructure among millet research labs in developing countries of Asia and Africa, the reach of modern tools like genome editing is delayed. Among the available genome editing tools, the CRISPR/Cas system may play a key role for genome editing in millets due to it user friendly and cost effective construct design and tools like ZFN and TALEN may not be utilized for genome editing in the labs of less developed countries with millet research. Other key aspects influencing millet genome editing are discussed below.
Whole genome sequence of millet
Availability of a fully annotated genome sequence is the vital prerequisite for undertaking genome editing studies in any organism. A fully annotated whole genome sequence is required for the prediction of target regions and guide RNA (gRNA) in the CRISPR/Cas9-mediated genome editing system. For targeting individual genes, details on the structure of the whole gene, including intronic and exonic regions, are needed. Further, Cas9 from Streptococcus pyogenes requires “NGG” at the 3′ end of the target region [3]. So, mining such regions and designing gRNAs targeting the specific sequence demand a fully annotated whole genome sequence. Unfortunately, a fully annotated genome sequence is available only for foxtail millet as of now [2]. Only draft genome sequences were released for finger millet [62], pearl millet [63], and proso millet [64]. Fully annotated genome sequences are not yet available for these millets. Not even a draft genome sequence is reported for other millets (little millet, barnyard millet, and kodo millet) up to now. In genome editing studies, the editing efficiency and type of editing (insertion and/or deletion) are being analyzed by the sequencing of target regions and sometimes by whole genome re-sequencing of the plants especially to detect any off-target effects. Whole genome re-sequencing has been applied in rice [65,66,67], sorghum [68], soybean [69], and Arabidopsis [70] to find off-target effects of genome-edited plants. Such studies could not be conducted in millets except for foxtail millet since it is the only millet having a complete and annotated genome sequence. So, the availability of complete and annotated genomes of millets will help in the efficient application of genome editing systems in the future.
Efficient transformation and regeneration protocols
In addition to having the fully annotated genome, existence of an efficient system for the transformation and regeneration is another important prerequisite for the successful development of mutant plants by genome editing. Generally, monocots are considered as recalcitrant for transformation and regeneration [71] studies. Thanks to the efforts made by labs around the world during the last two decades for the development of an efficient transformation system especially based on Agrobacterium, several efficient transformation works were reported for cereals [72, 73]. However, millet crops are lagging behind when compared with mainstream cereals on the transformation studies too, and only a few reports are available on millet transformation (Table 1) [12, 74]. We have recently reviewed the status of millet transformation works [2].
Millets were predominantly transformed by the Agrobacterium-mediated system. Finger millet has several transformation reports and transgenic plants were regenerated by both direct [75] and indirect [76, 77] regeneration methods (reviewed in [2]). Following finger millet, foxtail millet has a few reports on transformation studies. The Agrobacterium-mediated system was used frequently and it was first report by Yinghui et al. [78]. Many labs in China used the Jigu11 genotype of foxtail millet for transformation studies (reviewed in [2]). We have also reported a transformation system for the Maxima (Acc. No: Bs 3875) genotype with direct regeneration of transformed explants [79]. Two recent studies of the same period also reported the optimization of conditions for Agrobacterium-mediated transformation of foxtail millet [80, 81]. A few reports are also available on genetic transformation of pearl millet and this millet was predominately transformed by the biolistic method (reviewed in [74]). Embryogenic calli were mostly used as the explants for the transformation of pearl millet. Only one old report is available on the transformation of barnyard millet by the biolistic method in which the efficiencies of various promoters were tested for the expression of the GUS reporter gene [82]. Agrobacterium-mediated transformation of kodo millet was reported recently by Bhatt et al. [83]. They have optimized various parameters for the successful transformation of kodo millet in this study. No report is available on the transformation of little millet and proso millet. This will hamper application of genome editing in these millets. Among the available transformation methods, the Agrobacterium-based system is expected to dominate millet transformation and the same could be extended to successful delivery of CRISPR/Cas constructs to millets. Biolistic system-based transformation may not be utilized as many millet research labs in developing countries may not afford this expensive system.
As a leading millet with completely annotated genome with application of genome editing tools, foxtail millet is expected to lead the genome editing studies among millets. To further supplement the genetic studies, a mini foxtail millet variety with a shorter life cycle has been developed recently [84]. This is named as Xiaomi, and has a point mutation in the Phytochrome C gene and shows a heading date of 39 days after sowing whereas for wild type plants it takes 82 days. The authors have also assembled the genome and transcriptome of the Xiaomi variety with establishment of an efficient transformation system [84]. This variety of foxtail millet could serve as a model system for functional genomics studies in other millets in future. The Xiaomi variety could be harnessed for several genome editing studies and genes of many other millets could be tested by comparative genomics approaches in this variety to accelerate the gene characterization.
Conclusion
Millets are nutri-rich cereal crops supplying energy and nutrients to the majority of the people in the semi-arid tropics of Asia and Africa. They also possess good climate resilient properties like tolerance to drought and salinity. Unfortunately, modern genetic research is still far from reaching these crops due to the fact that these are cultivated in less developed countries where the resource for modern molecular research is limited. Application of genome editing has been considered to accelerate crop breeding by helping to impart key traits precisely. Apart from foxtail millet, other millets have not yet been used for genome editing studies. Development of completely annotated genomes and an efficient transformation system for millets may aid high resolution studies using genome editing tools like CRISPR/Cas9 in the near future to help conserve food security in Asia and Africa.
References
Ceasar S, Maharajan T, Ajeesh Krishna TP et al (2018) Finger millet [Eleusine coracana (L.) Gaertn.] improvement: current status and future interventions of whole genome sequence. Front Plant Sci 9:1054
Vetriventhan M, Azevedo VCR, Upadhyaya HD et al (2020) Genetic and genomic resources, and breeding for accelerating improvement of small millets: current status and future interventions. Nucleus 63:217–239. https://doi.org/10.1007/s13237-020-00322-3
Ceasar SA, Rajan V, Prykhozhij SV (2016) Insert, remove or replace: A highly advanced genome editing system using CRISPR/Cas9. Biochim Biophys Acta 1863:2333–2344
Manghwar H, Lindsey K, Zhang X, Jin S (2019) CRISPR/Cas system: recent advances and future prospects for genome editing. Trends Plant Sci 24:1102–1125
Wang H, La Russa M, Qi LS (2016) CRISPR/Cas9 in genome editing and beyond. Annu Rev Biochem 85:227–264. https://doi.org/10.1146/annurev-biochem-060815-014607
Chen K, Wang Y, Zhang R et al (2019) CRISPR/Cas genome editing and precision plant breeding in agriculture. Annu Rev Plant Biol 70:667–697. https://doi.org/10.1146/annurev-arplant-050718-100049
Hillary VE, Ceasar SA (2019) Application of CRISPR/Cas9 genome editing system in cereal crops. Open Biotechnol J. https://doi.org/10.2174/1874070701913010173
Langner T, Kamoun S, Belhaj K (2018) CRISPR crops: plant genome editing toward disease resistance. Annu Rev Phytopathol 56:479–512. https://doi.org/10.1146/annurev-phyto-080417-050158
Song G, Jia M, Chen K et al (2016) CRISPR/Cas9: a powerful tool for crop genome editing. Crop J 4:75–82
Zaidi SS-A, Mahas A, Vanderschuren H, Mahfouz MM (2020) Engineering crops of the future: CRISPR approaches to develop climate-resilient and disease-resistant plants. Genome Biol 21:289. https://doi.org/10.1186/s13059-020-02204-y
Zhu H, Li C, Gao C (2020) Applications of CRISPR–Cas in agriculture and plant biotechnology. Nat Rev Mol Cell Biol 21:661–677. https://doi.org/10.1038/s41580-020-00288-9
Ceasar SA, Ignacimuthu S (2009) Genetic engineering of millets: current status and future prospects. Biotechnol Lett 31:779–788. https://doi.org/10.1007/s10529-009-9933-4
Shobana S, Krishnaswamy K, Sudha V, Malleshi NG, Anjana RM, Palaniappan L, Mohan V (2013) Finger millet (Ragi, Eleusine coracana L.): a review of its nutritional properties, processing, and plausible health benefits. Adv Food Nut Res 69:1–39. https://doi.org/10.1016/B978-0-12-410540-9.00001-6
Devi PB, Vijayabharathi R, Sathyabama S, Malleshi NG, Priyadarisini VB (2014) Health benefits of finger millet (Eleusine coracana L.) polyphenols and dietary fiber: a review. J Food Sci Techn 51:1021–1040. https://doi.org/10.1007/s13197-011-0584-9
Maharajan T, Ceasar SA, Krishna ATP, Ignacimuthu S (2021) Finger millet [Eleusine coracana (L.) Gaertn]: an orphan crop with a potential to alleviate the calcium deficiency in the semi-arid tropics of Asia and Africa. Front Sust Food Syst 5:258. https://doi.org/10.3389/fsufs.2021.684447
Saha D, Gowda MVC, Arya L, Verma M, Bansal KC (2016) Genetic and genomic resources of small millets. Crit Rev Plant Sci 35:56–79
Saleh ASM, Zhang Q, Chen J, Shen Q (2013) Millet grains: nutritional quality, processing, and potential health benefits. Compr Rev Food Sci Food Saf 12:281–295. https://doi.org/10.1111/1541-4337.12012
Puranik S, Kam J, Sahu PP et al (2017) Harnessing finger millet to combat calcium deficiency in humans: challenges and prospects. Front Plant Sci 8:1311
Zhang LZ, Liu RH (2015) Phenolic and carotenoid profiles and antiproliferative activity of foxtail millet. Food Chem 174:495–501. https://doi.org/10.1016/j.foodchem.2014.09.089
Hadimani N, Muralikrishna G, Tharanathan R, Malleshi N (2001) Nature of carbohydrates and proteins in three pearl millet varieties varying in processing characteristics and kernel texture. J Cereal Sci 33:17–25
Antony U, Sripriya G, Chandra TS (1996) Effect of fermentation on the primary nutrients in finger millet (Eleusine coracana). J Agric Food Chem 44:2616–2618. https://doi.org/10.1021/jf950787q
Nambiar V, Dhaduk J, Sareen N, Shahu T, Desai R (2011) Potential functional implications of Pearl millet (Pennisetum glaucum) in health and disease. J Appl Pharm Sci 1:62–67
Watanabe M (1999) Antioxidative phenolic compounds from Japanese barnyard millet (Echinochloa utilis) grains. J Agric Food Chem 47:4500–4505. https://doi.org/10.1021/jf990498s
Radhika G, Sathya RM, Ganesan A, Saroja R, Vijayalakshmi P, Sudha V, Mohan V (2011) Dietary profile of urban adult population in South India in the context of chronic disease epidemiology (CURES-68). Public Health Nutr 14:591–598. https://doi.org/10.1017/S136898001000203X
Singh P, Raghuvanshi RS (2012) Finger millet for food and nutritional security. Afr J Food Sci 6:77–84
Gaj T, Sirk SJ, Shui S-L, Liu J (2016) Genome-editing technologies: principles and applications. Cold Spring Harb Perspect Biol 8:a023754. https://doi.org/10.1101/cshperspect.a023754
Kamburova VS, Nikitina EV, Shermatov SE et al (2017) Genome editing in plants: an overview of tools and applications. Int J Agron 2017:7315351. https://doi.org/10.1155/2017/7315351
Nadakuduti SS, Buell CR, Voytas DF et al (2018) Genome editing for crop improvement-applications in clonally propagated polyploids with a focus on potato (Solanum tuberosum L.). Front Plant Sci 9:1607
Zhang J, Zhang H, Botella JR, Zhu J-K (2018) Generation of new glutinous rice by CRISPR/Cas9-targeted mutagenesis of the Waxy gene in elite rice varieties. J Integr Plant Biol 60:369–375. https://doi.org/10.1111/jipb.12620
Ricroch A (2019) Global developments of genome editing in agriculture. Trans Res 28:45–52. https://doi.org/10.1007/s11248-019-00133-6
Zhang Y, Li D, Zhang D, Zhao X, Cao X, Dong L, Liu J, Chen K, Zhang H, Gao C, Wang D (2018) Analysis of the functions of TaGW2 homoeologs in wheat grain weight and protein content traits. Plant J 94:857–866. https://doi.org/10.1111/tpj.13903
Zhang Y, Massel K, Godwin ID, Gao C (2018) Applications and potential of genome editing in crop improvement. Genome Biol 19:210. https://doi.org/10.1186/s13059-018-1586-y
Oliva R, Ji C, Atienza-Grande G, Huguet-Tapia JC, Perez-Quintero A, Li T, Eom JS, Li C, Nguyen H, Liu B, Auguy F, Sciallano C, Luu VT, Dossa GS, Cunnac S, Schmidt SM, Slamet-Loedin IH, Vera Cruz C, Szurek B, Yang B (2019) Broad-spectrum resistance to bacterial blight in rice using genome editing. Nat Biotech 37:1344–1350. https://doi.org/10.1038/s41587-019-0267-z
Xu Z, Xu X, Gong Q, Li Z, Li Y, Wang S, Yang Y, Ma W, Liu L, Zhu B, Zou L, Chen G (2019) Engineering broad-spectrum bacterial blight resistance by simultaneously disrupting variable TALE-binding elements of multiple susceptibility genes in rice. Mol Plant 12:434–1446. https://doi.org/10.1016/j.molp.2019.08.006
Liu J, Chen J, Zheng X, Wu F, Lin Q, Heng Y, Tian P, Cheng Z, Yu X, Zhou K, Zhang X, Guo X, Wang J, Wang H, Wan J (2017) GW5 acts in the brassinosteroid signalling pathway to regulate grain width and weight in rice. Nat Plants 3:17043. https://doi.org/10.1038/nplants.2017.43
Zeng Y, Wen J, Zhao W, Wang Q, Huang W (2020) rational improvement of rice yield and cold tolerance by editing the three genes OsPIN5b, GS3, and OsMYB30 with the CRISPR–Cas9 system. Fronti Plant Sci 10:1663. https://doi.org/10.3389/fpls.2019.01663
Zhou J, Xin X, He Y, Chen H, Li Q, Tang X, Zhong Z, Deng K, Zheng X, Akher SA, Cai G, Qi Y, Zhang Y (2019) Multiplex QTL editing of grain-related genes improves yield in elite rice varieties. Plant Cell Rep 38:475–485. https://doi.org/10.1007/s00299-018-2340-3
Ansari WA, Chandanshive SU, Bhatt V et al (2020) Genome editing in cereals: approaches, applications and challenges. Int J Mol Sci 21:4040. https://doi.org/10.3390/ijms21114040
Zhu C, Bortesi L, Baysal C et al (2017) Characteristics of genome editing mutations in cereal crops. Trends Plant Sci 22:38–52. https://doi.org/10.1016/j.tplants.2016.08.009
Shan Q, Wang Y, Li J, Gao C (2014) Genome editing in rice and wheat using the CRISPR/Cas system. Nat Protoc 9:2395–2410. https://doi.org/10.1038/nprot.2014.157
Wang Y, Cheng X, Shan Q et al (2014) Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat Biotechnol 32:947–951. https://doi.org/10.1038/nbt.2969
Zhang Y, Bai Y, Wu G et al (2017) Simultaneous modification of three homoeologs of TaEDR1 by genome editing enhances powdery mildew resistance in wheat. Plant J 91:714–724. https://doi.org/10.1111/tpj.13599
Wang F, Wang C, Liu P et al (2016) Enhanced rice blast resistance by CRISPR/Cas9-targeted mutagenesis of the ERF transcription factor gene OsERF922. PLoS ONE 11:e0154027
Jiang M, Liu Y, Liu Y et al (2019) Mutation of inositol 1,3,4-trisphosphate 5/6-kinase6 impairs plant growth and phytic acid synthesis in rice. Plants 8:114
Lu Y, Zhu J-K (2017) Precise editing of a target base in the rice genome using a modified CRISPR/Cas9 system. Mol Plant 10:523–525. https://doi.org/10.1016/j.molp.2016.11.013
Zhu J, Song N, Sun S et al (2016) Efficiency and inheritance of targeted mutagenesis in maize using CRISPR-Cas9. J Genet Genom 43:25–36. https://doi.org/10.1016/j.jgg.2015.10.006
Xie K, Yang Y (2013) RNA-guided genome editing in plants using a CRISPR–Cas system. Mol Plant 6:1975–1983. https://doi.org/10.1093/mp/sst119
Macovei A, Sevilla NR, Cantos C et al (2018) Novel alleles of rice eIF4G generated by CRISPR/Cas9-targeted mutagenesis confer resistance to Rice tungro spherical virus. Plant Biotechnol J 16:1918–1927. https://doi.org/10.1111/pbi.12927
Mao X, Zheng Y, Xiao K et al (2018) OsPRX2 contributes to stomatal closure and improves potassium deficiency tolerance in rice. Biochem Biophys Res Commun 495:461–467. https://doi.org/10.1016/j.bbrc.2017.11.045
Tang L, Mao B, Li Y et al (2017) Knockout of OsNramp5 using the CRISPR/Cas9 system produces low Cd-accumulating indica rice without compromising yield. Sci Rep 7:14438. https://doi.org/10.1038/s41598-017-14832-9
Zhang A, Liu Y, Wang F et al (2019) Enhanced rice salinity tolerance via CRISPR/Cas9-targeted mutagenesis of the OsRR22 gene. Mol Breed 39:47. https://doi.org/10.1007/s11032-019-0954-y
Kumar A, Tomer V, Kaur A, Kumar V, Gupta K (2018) Millets: a solution to agrarian and nutritional challenges. Agric Food Sec 7(1):31. https://doi.org/10.1186/s40066-018-0183-3
Krishna TPA, Ceasar SA, Maharajan T, Ramakrishnan M, Duraipandiyan V, Al-Dhabi N, Ignacimuthu S (2017) Improving the zinc-use efficiency in plants: A review. SABRAO J Breed Genet 49:221–230
Ceasar SA, Hodge A, Baker A, Baldwin SA (2014) Phosphate concentration and arbuscular mycorrhizal colonisation influence the growth, yield and expression of twelve PHT1 family phosphate transporters in foxtail millet (Setaria italica). PLoS ONE. https://doi.org/10.1371/journal.pone.0108459
Nadeem F, Ahmad Z, Wang R, Han J, Shen Q, Chang F, Diao X, Zhang F, Li X (2018) Foxtail millet [Setaria italica (L.) Beauv.] grown under low nitrogen shows a smaller root system, enhanced biomass accumulation, and nitrate transporter expression. Front Plant Sci 9:205. https://doi.org/10.3389/fpls.2018.00205
Maharajan T, Ceasar SA, Krishna TPA, Ignacimuthu S (2019) Phosphate supply influenced the growth, yield and expression of PHT1 family phosphate transporters in seven millets. Planta 250:1433–1448
Doust AN, Kellogg EA, Devos KM, Bennetzen JL (2009) Foxtail Millet: a sequence-driven grass model system. Plant Physiol 149:137–141. https://doi.org/10.1104/pp.108.129627
Bennetzen JL, Schmutz J, Wang H et al (2012) Reference genome sequence of the model plant Setaria. Nat Biotechnol 30:555–561. https://doi.org/10.1038/nbt.2196
Zhang G, Liu X, Quan Z et al (2012) Genome sequence of foxtail millet (Setaria italica) provides insights into grass evolution and biofuel potential. Nat Biotechnol 30:549
Lin C-S, Hsu C-T, Yang L-H et al (2018) Application of protoplast technology to CRISPR/Cas9 mutagenesis: from single-cell mutation detection to mutant plant regeneration. Plant Biotechnol J 16:1295–1310. https://doi.org/10.1111/pbi.12870
Cheng Z, Sun Y, Yang S et al (2021) Establishing in planta haploid inducer line by edited SiMTL in foxtail millet (Setaria italica). Plant Biotechnol J. https://doi.org/10.1111/pbi.13584
Hittalmani S, Mahesh HB, Shirke MD et al (2017) Genome and transcriptome sequence of finger millet (Eleusine coracana (L.) Gaertn.) provides insights into drought tolerance and nutraceutical properties. BMC Genomics 18:465. https://doi.org/10.1186/s12864-017-3850-z
Varshney RK, Shi C, Thudi M et al (2017) Pearl millet genome sequence provides a resource to improve agronomic traits in arid environments. Nat Biotechnol 35:969–976. https://doi.org/10.1038/nbt.3943
Zou C, Li L, Miki D et al (2019) The genome of broomcorn millet. Nat Commun 10:436. https://doi.org/10.1038/s41467-019-08409-5
Li G, Jain R, Chern M et al (2017) The sequences of 1504 mutants in the model rice variety kitaake facilitate rapid functional genomic studies. Plant Cell 29:1218–1231. https://doi.org/10.1105/tpc.17.00154
Li S, Zheng Y, Cui H et al (2016) Frequency and type of inheritable mutations induced by γ rays in rice as revealed by whole genome sequencing. J Zhejiang Univ B 17:905–915. https://doi.org/10.1631/jzus.B1600125
Tang X, Liu G, Zhou J et al (2018) A large-scale whole-genome sequencing analysis reveals highly specific genome editing by both Cas9 and Cpf1 (Cas12a) nucleases in rice. Genome Biol 19:84. https://doi.org/10.1186/s13059-018-1458-5
Jiao Y, Burke J, Chopra R et al (2016) A sorghum mutant resource as an efficient platform for gene discovery in grasses. Plant Cell 28:1551–1562. https://doi.org/10.1105/tpc.16.00373
Tsuda M, Kaga A, Anai T et al (2015) Construction of a high-density mutant library in soybean and development of a mutant retrieval method using amplicon sequencing. BMC Genomics 16:1014. https://doi.org/10.1186/s12864-015-2079-y
Belfield EJ, Gan X, Mithani A et al (2012) Genome-wide analysis of mutations in mutant lineages selected following fast-neutron irradiation mutagenesis of Arabidopsis thaliana. Genome Res 22:1306–1315. https://doi.org/10.1101/gr.131474.111
Hofmann NR (2016) A breakthrough in monocot transformation methods. Plant Cell 28:1989. https://doi.org/10.1105/tpc.16.00696
Hiei Y, Ishida Y, Komari T (2014) Progress of cereal transformation technology mediated by Agrobacterium tumefaciens. Front Plant Sci 5:628
Shrawat AK, Lörz H (2006) Agrobacterium-mediated transformation of cereals: a promising approach crossing barriers. Plant Biotechnol J 4:575–603. https://doi.org/10.1111/j.1467-7652.2006.00209.x
Sood P, Singh RK, Prasad M (2019) Millets genetic engineering: the progress made and prospects for the future. Plant Cell Tissue Organ Cult’ 137:421–439. https://doi.org/10.1007/s11240-019-01587-6
Satish L, Ceasar SA, Ramesh M (2017) Improved Agrobacterium-mediated transformation and direct plant regeneration in four cultivars of finger millet (Eleusine coracana (L.) Gaertn.). Plant Cell Tissue Organ Cult 131:547–565. https://doi.org/10.1007/s11240-017-1305-5
Ceasar S, Ignacimuthu S (2011) Agrobacterium-mediated transformation of finger millet (Eleusine coracana (L.) Gaertn.) using shoot apex explants. Plant Cell Rep 30:1759–1770. https://doi.org/10.1007/s00299-011-1084-0
Ignacimuthu S, Ceasar SA (2012) Development of transgenic finger millet (Eleusine coracana (L.) Gaertn.) resistant to leaf blast disease. J Biosci. https://doi.org/10.1007/s12038-011-9178-y
Yinghui L, Jingjuan Y, Qian Z et al (2005) Genetic transformation of millet (Tetaria italica) by Agrobacterium-mediated. Nong ye sheng wu ji shu xue bao =. J Agric Biotechnol 13:32–37
Ceasar SA, Baker A, Ignacimuthu S (2017) Functional characterization of the PHT1 family transporters of foxtail millet with development of a novel Agrobacterium-mediated transformation procedure. Sci Rep 7:14064. https://doi.org/10.1038/s41598-017-14447-0
Santos CM, Romeiro D, Silva JP et al (2020) An improved protocol for efficient transformation and regeneration of Setaria italica. Plant Cell Rep 39:501–510. https://doi.org/10.1007/s00299-019-02505-y
Sood P, Singh RK, Prasad M (2020) An efficient Agrobacterium-mediated genetic transformation method for foxtail millet (Setaria italica L.). Plant Cell Rep 39:511–525. https://doi.org/10.1007/s00299-019-02507-w
Gupta P, Raghuvanshi S, Tyagi A (2001) Assessment of the efficiency of various gene promoters via biolistics in leaf and regenerating seed callus of millets, Eleusine coracana and Echinochloa crusgalli. Plant Biotechnol 18:275–282. https://doi.org/10.5511/plantbiotechnology.18.275
Bhatt R, Asopa PP, Jain R, Kothari-Chajer A, Kothari SL, Kachhwaha S (2021) Optimization of Agrobacterium-mediated genetic transformation in Paspalum scrobiculatum L. (kodo millet). Agronomy 11:1104. https://doi.org/10.3390/agronomy11061104
Yang Z, Zhang H, Li X et al (2020) A mini foxtail millet with an Arabidopsis-like life cycle as a C4 model system. Nat Plants 6:1167–1178. https://doi.org/10.1038/s41477-020-0747-7
Fraiture MA, Roosens NHC, Taverniers I et al (2016) Biotech rice: current developments and future detection challenges in food and feed chain. Trends Food Sci Technol 52:66–79. https://doi.org/10.1016/j.tifs.2016.03.011
Acknowledgements
The author thanks Dr David Pilbeam, Visiting Research Fellow, University of Leeds for critically reading the manuscript.
Funding
The research works in our lab are funded by the Department of Biotechnology, Govt. of India under Grant No: BT/PR21321/GET/119/76/2016.
Author information
Authors and Affiliations
Contributions
SAC conceptualized, drafted, edited, and produced the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The author declares that no conflict of interest exists.
Consent to participate
This article does not involve any clinical studies that need consent of participants.
Consent to publish
This article does not involve any clinical studies that need consent of participants to publish.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ceasar, A. Genome-editing in millets: current knowledge and future perspectives. Mol Biol Rep 49, 773–781 (2022). https://doi.org/10.1007/s11033-021-06975-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06975-w