Abstract
Black rice is a rare type of rice germplasm with various health benefits that are largely attributed to anthocyanin pigment accumulation in the pericarps. The anthocyanin biosynthesis in plant tissues is activated mainly by the MBW complexes, consisting of three types of transcription factors R2R3-MYB, bHLH, and WDR. In black rice, the bHLH and WDR components regulating anthocyanin biosynthesis in pericarps have been characterized, while the R2R3-MYB factor remains unknown. By examining the expression correlation between all putative rice MYB genes and anthocyanin biosynthesis–related genes based on transcriptome data of pericarps in combination with further molecular and genetic analysis, we proved that OsMYB3 (LOC_Os03g29614) was the determinant R2R3-MYB gene for anthocyanin biosynthesis in rice pericarps. The expression level of OsMYB3 in pericarps of black rice was significantly higher than that of white rice. The knockout of OsMYB3 in a black rice variety caused significant downregulation of 19 anthocyanin metabolites and many other flavonoids in grains. Our research deepens the understanding of regulatory system for anthocyanin biosynthesis in rice pericarps and provides implications for breeding black rice varieties with high anthocyanin level.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As one of the most important staple foods in the world, rice (Oryza sativa L.) generally provides energy and a portion of protein for daily need (Al-Kanhal et al. 1999). However, black rice, a special type of rice germplasm with anthocyanin accumulation in the pericarps, have been proved to provide many additional health benefits compared with common white rice, such as preventing insulin resistance (Guo et al. 2007) and cholesterol absorption (Yao et al. 2013), inhibiting breast cancer cell growth (Hui et al. 2010) and D-galactose-induced senescence (Lu et al. 2014). Moreover, high anthocyanin intake has also shown to reduce the risk of cancer (Peiffer et al. 2016), inflammation (Miyake et al. 2011), and neurological diseases (Strathearn et al. 2014), cardiovascular diseases (Cassidy 2018), obesity (Li et al. 2013), and other chronic diseases. Therefore, the consumption of black rice has been becoming more and more popular (Kushwaha 2016). Recently, black rice has been advocated to be consumed as staple food to substitute white rice due to its outstanding health-promoting effects (Zhang 2021).
Anthocyanins are water-soluble flavonoid pigments that are broadly accumulated in plants. The anthocyanin biosynthesis pathway is catalyzed by a set of enzymes including chalcone synthase (CHS), chalcone isomerase (CHI), flavanone 3-hydroxylase (F3H), flavonoid 3′-hydroxylase (F3’H), dihydroflavonol 4-reductase (DFR), anthocyanidin synthase (ANS), and UDPG-flavonoid glucosyltransferase (UFGT). The activation of the anthocyanin biosynthetic genes largely relies on the MBW complexes, consisting of three types of transcription factors (TFs) R2R3-MYB, bHLH, and WDR, and this regulatory model is widely conserved in higher plants (Xu et al. 2015). According to the conservative amino acid motifs in the MYB repeat sequence, R2R3-MYBs in Arabidopsis thaliana can be further divided into 22 subgroups (subgroup 1 to 22, SG1-22) (Kranz et al. 1998). In MBW complexes, the R2R3-MYBs are generally responsible for the spatiotemporal patterns of anthocyanin production (Albert et al. 2014). For instance, R2R3-MYBs of SG6 (PAP1, PAP2, AtMYB113, and AtMYB114) regulate the anthocyanin biosynthetic genes by forming the MBW complex with bHLHs (GL3, EGL3, or TT8) and TTG1 in vegetative tissues (Gonzalez et al. 2008). Nevertheless, in the seed coat of Arabidopsis, the MBW complex composed of TT2 (R2R3-MYB, SG5), TT8 (bHLH), and TTG1 (WDR) activates the biosynthesis of proanthocyanidin, belonging to a branch closely related to anthocyanin in the flavonoid pathway (Baudry et al. 2004). Interestingly, AtMYB113 (At1g66370), AtMYB114 (At1g66380), and PAP2 (At1g66390) are clustered on chromosome 1, which indicated that these genes probably derived from the tandem repeats of PAP1 (At1g56650) during the evolution of Arabidopsis (Lin-Wang et al. 2010). In maize (Zea mays), ZmPl and ZmC1 of SG5 are responsible for anthocyanin biosynthesis via interacting with bHLHs (R/B1) and the WDR protein PAC1, i.e., ZmPl is responsible for the regulation in vegetative tissues and floral organs, and ZmC1 functions in caryopsis (Petroni et al. 2014). Different from the R2R3-MYBs of SG5 and SG6, those of SG7 could activate the flavonoid biosynthetic pathway in absence of bHLHs. In Arabidopsis, AtMYB11, AtMYB12, and AtMYB111 activate the expression of the early biosynthetic genes (EBGs) AtCHS, AtCHI, and AtF3H and the flavonol synthase gene AtFLS, and their activations on AtFLS are partial redundancy (Stracke et al. 2007). In maize, P1 regulates the expression of EBGs and tannin synthesis in floral organs (Grotewold et al. 1994). In apple (Malus × domestica), MdMYB1 and MdMYBA mainly regulate the biosynthesis of anthocyanin in fruit skins, and MdMYB10 participates in the regulation of anthocyanin in apple flesh and leaves (Takos et al. 2006; Ban et al. 2007; Espley et al. 2007).
Rice may accumulate anthocyanin pigments in various tissues including leaves, leaf sheathes, internodes, ligules, pericarps, apiculi, and stigmas. The anthocyanin biosynthetic genes of rice have been well characterized (Reddy et al. 1996, 2007; Druka et al. 2003; Furukawa et al. 2007; Kim et al. 2008; Shih et al. 2008; Tanaka et al. 2008). However, anthocyanin production in different rice tissues was mainly determined by R2R3-MYB, bHLH regulators, and the biosynthetic gene OsDFR (also known as Rd). For instance, OsC1 (R2R3-MYB), OsRb (bHLH), and OsDFR regulated anthocyanin biosynthesis in leaves (Zheng et al. 2019). OsC1, OsKala4 (bHLH, also known as OsB2), and OsDFR regulated anthocyanin biosynthesis in hulls (Sun et al. 2018). While OsC1-OsPa (bHLH)-OsDFR and OsC1-OsPs (bHLH)-OsDFR regulated anthocyanin accumulation in apiculi and stigmas, respectively (Meng et al. 2021). Moreover, OrC1, a novel allele of OsC1 from O. rufipongon, was reported to have different tissue-specificities in different rice subspecies, i.e., OrC1 promotes anthocyanin accumulation in apiculi, leaf sheathes, and stigmas in indica rice, but only in apiculi in japonica rice (Qiao et al. 2021). Oikawa et al. (2015) demonstrated that the gain-of-function mutation of a rice bHLH gene OsKala4 activated the expression of anthocyanin biosynthetic genes in pericarps and caused the anthocyanin accumulation in pericarps. Recently, OsPAC1, a WDR that participated in activating anthocyanin biosynthetic genes in rice leaves (Zheng et al. 2019), was proved to be crucial for anthocyanin biosynthesis in pericarps as well (Yang et al. 2021). However, the R2R3-MYB component regulating anthocyanin biosynthesis in rice pericarps remained uncharacterized. Although OsC1 widely regulates anthocyanin biosynthesis in multiple rice tissues as described above, it is not the R2R3-MYB regulator for anthocyanin biosynthesis in rice pericarps. It was because that the expression of OsC1 was essentially absent in rice pericarps (Zheng et al. 2019).
Rice contains approximately 230 MYB genes (Feller et al. 2011). To identify the R2R3-MYB regulator for anthocyanin biosynthesis in rice pericarps, in this study, we examined the expression correlation between all rice MYB genes and anthocyanin biosynthesis–related genes to select putative MYB candidates based on transcriptome data of pericarps from 27 black rice accessions. We finally determined the R2R3-MYB regulator for anthocyanin biosynthesis in rice pericarps through further molecular and genetic analysis of the selected MYB candidate genes.
Materials and methods
Plant materials
A total of 27 black rice accessions were collected in China and used for transcriptome analysis (Supplementary Table 1).
Rice pericarp sampling and RNA extraction
The black rice materials were grown in the field. The rice grains of 27 black rice accessions were harvested at 8 days after pollination (DAP) and placed into liquid nitrogen immediately. All grain samples of 8 DAP were transferred in the liquid nitrogen to the laboratory and were stored at – 80 °C before RNA extraction.
In the laboratory, rice grains of 8 DAP were first dulled using the forceps, and immature embryos were removed from the immature seeds by a scalpel. Then, the endosperms were subsequently squeezed out, and the remaining pericarps were place liquid nitrogen. The RNA extraction followed the procedure as described by Yang et al (2006).
Transcriptome analysis
Transcriptome data of leaves from 268 rice accessions, transcriptome data of young spikes from 265 rice accessions, transcriptome data of 8 DAP pericarps from 145 rice accessions, and transcriptome data of 8 DAP endosperms from 60 rice accessions, which were all sequenced previously by our laboratory, were used to profile the expression of OsMYB3.
RNA samples were sent to Novogene Corporation (Beijing, China) for RNA sequencing. Reference genome and gene model annotation files were obtained from MSU Rice Genome Annotation Project Release 7 (http://rice.plantbiology.msu.edu). HISAT2 v.2.1.0 (http://ccb.jhu.edu/software/hisat2/ index.shtml) was used for mapping clean reads to the reference genome, and fragments per kb of transcript per million fragments mapped (FPKMs) of known genes were calculated by CUFFLINKS v.2.2.1 (http://cole-trapnell-lab.github.io/cufflinks/) from the reference annotation file. The FPKM values were scaled to 0–1, a heatmap was illustrated using R/pheatmap, and the correlation coefficient matrices were generated and displayed using R/corrgram.
qRT-PCR
Two micrograms of total RNA for each sample was treated with RNA-free DNase I (Promega). Reverse transcription was performed using M-MLV Reverse Transcriptase (Invitrogen). Real-time PCR was conducted on a ViiA7 Real-time PCR system (Applied Biosystems, Foster City, CA, USA) using FastStart Universal SYBR Green Master (ROX) (Roche) as described by Zheng et al. (2019). The ubiquitin gene was used as the reference, and each sample was assessed in triplicate of technical replications. All primers used are listed in Supplementary Table 3.
Phylogenetic analysis
The amino acid sequences of known maize and Arabidopsis MYBs involved in anthocyanin or proanthocyanidin biosynthesis were obtained from Phytozome (https://phytozome.jgi.doe.gov/pz/portal.html#). The amino acid sequences of 233 putative MYB TFs of rice (Feller et al. 2011) were extracted from RGAP 7 (http://rice.plantbiology.msu.edu/index.shtml). The amino acid sequences were aligned using the website tool CLUSTALOMEGA (https://www.ebi.ac.uk/Tools/msa/clustalo/). Phylogenetic trees were constructed by a comprehensive molecular biology analysis tool suite, GENEIOUS (https://www.geneious.com/).
Anthocyanin content measurement
Anthocyanin extraction and content analysis followed the protocol described by Zhu et al. (2010). The anthocyanin content was determined by high performance liquid chromatography (HPLC) using an Agilent 1260 series system (Agilent Technologies, Palo Alto, CA, USA).
Overexpression and knockout of OsMYB3 in transgenic rice
The full-length cDNA of OsMYB3 (LOC_Os03g29614) was isolated from a black rice variety Zixiangnuo1 (Oryza sativa ssp. japonica) and inserted into pCAMBIA1300 under the control of the maize ubiquitin promoter and NOS terminator to form the overexpression vector. The CRISPR/Cas9-based genome editing method was used to generate OsMYB3 knockout lines. OsMYB3 was inserted in an sgRNA-Cas9 expression vector as described by Ma et al. (2015). The overexpression vector of OsMYB3 was introduced into Zixiangnuo1 and Chao 2–10 (a white rice variety), while the knockout vector was introduced into Zixiangnuo1. Agrobacterium-mediated rice transformation followed the protocol as described by Lin et al. (2002).
Yeast two-hybrid (Y2H)
Yeast AH109 cells were co-transformed with specific bait and prey constructs through the LiCl-PEG method according to the manufacturer’s manual (Clontech, Palo Alto, CA, USA). The transformants were selected on SD/-Leu/-Trp + X-α-gal medium. Interactions were tested on SD/-Leu/-Trp/-His/-Ade + X-α-gal medium.
Transcriptional activity assay using rice protoplasts
The cDNAs of OsMYB3, OsKala4, and OsPAC1 genes were isolated from Zixiangnuo1 and inserted into the “None” vector as effectors. Approximately 2-kb promoter regions of the OsCHS, OsCHI, OsF3′H, OsF3H, OsDFR, and OsANS1 genes were isolated from Zixiangnuo1 and inserted into the “190fLUC” vector to drive firefly luciferase (fLUC) as reporters. The internal control vector contains renilla luciferase (rLUC) driven by the ubiquitin promoter of Arabidopsis. Isolation of rice protoplasts and dual luciferase transcriptional activity assays were performed as described previously (Zong et al. 2016).
Luciferase activity was measured using the Dual-Luciferase® Reporter Assay System (Promega). Three independent transformations and measurements for each sample were performed.
Haplotype analysis
Nucleotide polymorphism information of OsMYB3 of 533 rice accessions was downloaded from RiceVarMap (http://ricevarmap.ncpgr.cn/) (Zhao et al. 2015). All SNPs and InDels with minor allele frequency (MAF) ≥ 0.05 in coding regions of OsMYB3 were used for haplotype analysis by DnaSP v.6.12.03 (http://www.ub.edu/dnasp/), and the haplotype network was drawn by haplotype viewer (http://www.cibiv.at/~greg/haploviewer).
Metabolomic profiling by LC–MS/MS
Whole-grain samples of OsMYB3 knockout lines and the original Zixiangnuo1, with three biological replicates for each, were sent to Wuhan MetWare Biotechnology Co., Ltd. (www.metware.cn) for a flavonoid metabolite analysis. In brief, the freeze-dried sample was ground in a mixer mill with a zirconia bead for 1.5 min at 30 Hz, and 100 mg of this powder was extracted with 1 mL of 70% aqueous methanol at 4 °C overnight. During this time, the extract was vortexed three times to increase extraction rate. After centrifugation at 10,000 × g for 10 min, the extracts were absorbed (CNWBOND Carbon-GCB SPE Cartridge, 250 mg, 3 mL; ANPEL, Shanghai, China) and filtered (SCAA-104, 0.22 μm pore size; ANPEL) before LC–MS/MS analysis.
The treated extracts were analyzed using an LC–ESI–MS/MS system (HPLC, Shim-pack UFLC CBM30A, Shimadzu, Kyoto, Japan; MS, 6500 QTRAP, Applied Biosystems, Norwalk, USA). The analytical conditions were as follows, HPLC: column, Waters ACQUITY UPLC HSS T3 C18 (1.8 μm, 2.1 mm * 100 mm); solvent system, water (0.04% acetic acid): acetonitrile (0.04% acetic acid); gradient program, 100:0 V/V at 0 min, 5:95 V/V at 11.0 min, 5:95 V/V at 12.0 min, 95:5 V/V at 12.1 min, 95:5 V/V at 15.0 min; flow rate, 0.40 mL/min; temperature, 40 °C; injection volume: 2 μL. The MS was operated in positive and negative ion modes. The ESI source operation parameters were as follows: ion source, turbo spray; source temperature 500 °C; ion spray voltage (IS) 5500 V; ion source gas I (GSI), gas II(GSII), curtain gas (CUR) were set at 55, 60, and 25.0 psi, respectively; the collision gas (CAD) was high. Metabolite quantification was performed using a multiple reaction monitoring method (MRM). De-clustering potential (DP) and collision energy (CE) for individual MRM transitions were done with further DP and CE optimization. A specific set of MRM transitions was monitored for each period according to the metabolites eluted within this period. Data were processed using Analyst 1.6.3 software.
Statistical analysis
ANOVA and Tukey’s honest significant difference test were performed using R/multcomp. For metabolomic profiling, all identified metabolites were subjected to orthogonal partial least squares discriminant analysis (OPLS-DA), and those with variable importance in project (VIP) ≥ 1 and fold change (FC) ≥ 2 or ≤ 0.5 were regarded as significantly differential metabolites.
Association study between the genomic polymorphisms of OsMYB3 and rice pericarp phenotype
For association study, we integrated the core collection of 533 rice accessions with 45 additional black rice accessions that we collected somewhere else and 25 black rice accessions from the 3 k genome project (Wang et al. 2018), because there are only six black rice accessions in the core collection. Only the SNP sites appearing on the panel are used to complete genotyping using the samtools (v1.8), and then use the beagle software to construct a reference panel and the imputation of the genotyping results. Nucleotide polymorphism information of OsMYB3 of 533 rice accessions was downloaded from RiceVarMap. Genomes of 45 black rice accessions were re-sequenced previously by our laboratory. Nucleotide polymorphism information of OsMYB3 of 25 black rice accessions from the 3 k genome project was downloaded from IRRI (http://iric.irri.org/resources/3000-genomes-project) (Wang et al. 2018). Association study was performed following the logistic regression method using PLINK v.1.90b3.40 (http://www.cog-genomics.org/plink/1.9/general_usa ge#cite). The significance thresholds were determined following a modified Bonferroni correction a* = a / Me.
Results
Transcriptome analysis revealed the putative MYB component for anthocyanin biosynthesis in the pericarps
We sequenced the transcriptomes of pericarps from 27 black rice accessions (Supplementary Table 1), which was classified by the phylogenetic analysis into two clades, corresponding to two main rice subspecies indica and japonica (Supplementary Fig. 1). The expression correlation between all 233 putative rice MYB genes and the flavonoid biosynthesis-related genes including OsCHS, OsCHI, OsF3’H, OsF3H, OsF3H2, OsDFR, OsANS1, OsANS2, OsUFGT, anthocyanidin reductase (OsANR), leucoanthocyanidin reductase (OsLAR), OsFLS, OsKala4, and OsPAC1 (Supplementary Fig. 2) was analyzed according to the transcriptome data (Supplementary Table 2). The result showed that four MYB genes (LOC_Os01g49160, LOC_Os01g63460, LOC_Os03g29614, and LOC_Os12g07640) out of all 233 putative rice MYB genes had a highly positive expression correlation with the anthocyanin biosynthetic genes OsCHS, OsCHI, OsF3’H, OsF3H, OsDFR, OsANS1, OsANS2, and OsUFGT, but low or no expression correlation with the biosynthetic genes of other flavonoid branches like OsANR, OsLAR, OsFLS, and OsF3H2 (Supplementary Fig. 2).
The phylogenetic analysis of the four MYB candidates (LOC_Os01g49160, LOC_Os01g63460, LOC_Os03g29614, and LOC_Os12g07640) and 28 known MYBs showed that LOC_Os03g29614 along with AtTT2 (Arabidopsis), ZmC1 (maize), ZmPl (maize), and OsC1 (rice) were grouped into the clade of SG5 that was related to anthocyanin or proanthocyanidin biosynthesis (Fig. 1a). In addition, the dual-luciferase transient transcriptional activity assay using rice protoplasts showed that LOC_Os03g29614 could activate the promoters of the two anthocyanin biosynthetic genes OsCHS and OsDFR effectively when co-expressed with OsKala4, the known bHLH component for anthocyanin biosynthesis in rice pericarps, but the other three MYB candidates (LOC_Os01g49160, LOC_Os01g63460, and LOC_Os12g07640) did not (Fig. 1b). Therefore, LOC_Os03g29614 was selected as the putative MYB component of the MBW complex activating the anthocyanin biosynthesis in the pericarps of black rice.
Characterization of the four MYB candidates for anthocyanin biosynthesis in the pericarp of black rice. a Phylogenetic analysis of the four MYB candidates and other known R2R3-MYBs. b The activation effects of the four MYBs on the anthocyanin biosynthetic genes OsCHS and OsDFR using a dual-luciferase transient transcriptional activity assay when co-expressed with OsKala4. Error bars in represent the SD
Functional characterization of LOC_Os03g29614
LOC_Os03g29614 was designated as to OsMYB3 as it was located in chromosome 3 of rice. OsMYB3 was predicted to contain a 966-bp open reading frame (ORF) encoding a protein product of 321 amino acids (aa). The aa sequence alignment of OsMYB3 and four known R2R3-MYBs involved in anthocyanin or proanthocyanidin biosynthesis showed that all of these R2R3-MYBs were highly conserved at the R2R3 domain located on N-terminus, but variable on the C-terminus (Supplementary Fig. 3a).
We further investigated the expression profiles of OsMYB3 based on the transcriptome data of four different tissues (leave, spikes, pericarps, and endosperms). The result showed that OsMYB3 predominantly expressed in the pericarps. OsMYB3 had no or extremely low expression level in the leaves, spikes, and endosperms, but comparatively high expression in pericarps (Supplementary Fig. 3b). The expression level of OsMYB3 in the pericarps differed significantly among black rice, white rice, and red rice, although it expressed in the pericarps of all the three rice types. The expression level of OsMYB3 in the pericarps of black rice was significantly higher than that of white rice, but was not significantly different from that of red rice (Supplementary Fig. 3c). Moreover, the expression level of OsMYB3 in the pericarps did not show significant difference among the different rice subspecies indica, japonica, and aus (Supplementary Fig. 3d).
As known, R2R3-MYBs of SG5 participate in activating anthocyanin or proanthocyanidin biosynthesis via interacting with bHLH and WDR proteins to form the MBW complexes. The Y2H assay proved that OsMYB3 interacted directly with any of the bHLHs OsRb, OsB1, and OsKala4 and the WDR OsPAC1 (Fig. 2a), indicating that OsMYB3 should function as a component of a MBW complex like other anthocyanin biosynthesis-activating R2R3-MYBs of SG5. A dual-luciferase transient transcriptional activity assay using rice protoplasts showed that none of OsMYB3, OsKala4, or OsPAC1 could activate the anthocyanin biosynthetic genes (OsCHS, OsCHI, OsF3′H, OsF3H, OsDFR, and OsANS1) alone (Fig. 2b). While co-transformation of OsMYB3 and OsKala4 (i.e., OsMYB3 + OsKala4) effectively activated these anthocyanin biosynthetic genes, and OsMYB3 + OsKala4 + OsPAC1 further improved the activation effect compared with OsMYB3 + OsKala4 (Fig. 2b). These results confirmed that the activation effect of OsMYB3 on anthocyanin biosynthesis relied on the formation of MBW complex.
Characterization of the regulatory role of OsMYB3 in anthocyanin biosynthesis. a Yeast two-hybrid assay of OsMYB3 and known bHLH or WDR partners in rice OsKala4, OsB1, OsRb, and OsPAC1. b The activation effects of OsMYB3 + OsKala4 + OsPAC1 complex on the anthocyanin biosynthetic genes OsCHS, OsCHI, OsF3’H, OsF3H, OsDFR, and OsANS1, respectively, using dual-luciferase transient transcriptional activity. Error bars represent the SD
OsMYB3 is the responsible R2R3-MYB for anthocyanin biosynthesis in rice pericarps
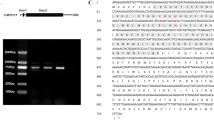
OsMYB3 was knocked out in a black rice cultivar Zixiangnuo1 by using the CRISPR/Cas9 system to validate its function. Three independent knockout lines of OsMYB3 (namely KO-5, KO-8, and KO-10) were generated with the target site located in the first exon encoding the R2 MYB domain (Fig. 3). KO-10 had a 2-bp deletion and KO-5 and KO-8 had a 1-bp insertion in the first exon of OsMYB3, respectively (Fig. 3a), all of which caused a frame-shift mutation. In contrast to the wild-type (WT) Zixiangnuo1 that showed dark black grains, the three OsMYB3-knockout lines appeared light brown grains (Fig. 3b). The HPLC analysis showed that Zixiangnuo1 accumulated 997.1 μg/g cyanidin 3O-glucosid (C3G) and 151.7 μg/g peonidin 3O-glucoside (P3G) in grains, while the anthocyanins in grains of the three knockout lines were all undetectable (Fig. 3c).
Knockout and overexpression of OsMYB3 in the black rice cultivar Zixiangnuo1. a Sequencing for the CRISPR/Cas9-targeted sites close to the 5′end of OsMYB3 knockout plantlet lines. b Grain color of the wild type Zixiangnuo1 and three knockout lines of OsMYB3. c Anthocyanin content in grains of Zixiangnuo1 and OsMYB3 knockout transgenic lines. KO-5, KO-8, and KO-10 were three independent knockout lines of OsMYB3. d Grain color of the wild type Zixiangnuo1 and three OsMYB3-overexpressed transgenic lines. e Anthocyanin content in grains of Zixiangnuo1 and OsMYB3-overexpressed transgenic lines. The asterisk (*) and double asterisk (**) indicate significant differences as compared to Zixiangnuo1 at P < 0.05 and P < 0.01, respectively. OE-Z3, OE-Z4, and OE-Z5 were three independent overexpression lines of OsMYB3. Error bars represent the standard deviation in c and e
OsMYB3 was then overexpressed in Zixiangnuo1 and a white rice cultivar Chao2-10 driven by the maize ubiquitin promoter. Three independent OsMYB3-overexpressing lines of Zixiangnuo1 (OE-Z3, OE-Z4, and OE-Z5) and Chao 2–10 (OE-C3, OE-C6, and OE-C9) were acquired. The anthocyanin content in grains of OE-Z3, OE-Z4, and OE-Z5 was 1274.2 μg/g (P = 0.0040), 1211.4 μg/g (P = 0.0023), and 1117.9 μg/g (P = 0.0205), which were all significantly higher than that of the WT Zixiangnuo1 (993.1 μg/g: Fig. 3d and e), while OE-C3, OE-C6, OE-C9, and the WT Chao2-10 did not show anthocyanin pigmentation in grains. Taken together, OsMYB3 is the responsible R2R3-MYB for anthocyanin biosynthesis in the pericarps of black rice.
Overexpression of OsMYB3 complemented the function of OsC1 in leaves
The previous study demonstrated that OsC1 was the determinant R2R3-MYB for anthocyanin biosynthesis in rice leaves, and non-functional alleles of OsC1 caused complete anthocyanin absence in rice leaves (Zheng et al. 2019). The leaves of Zixiangnuo1 and Chao2-10 were non-anthocyanin-pigmented because both cultivars contained the non-functional Osc1 allele with 10-bp deletion in exon 3 that was the most frequent null mutation of Osc1 alleles. However, leaves of all three OsMYB3-overexpressing lines of Zixiangnuo1 OE-Z3, OE-Z4, and OE-Z5 accumulated 325.6 μg/g, 118.4, and 15.8 μg/g anthocyanins respectively, in contrast to undetectable anthocyanin in leaves the WT Zixiangnuo1 (Fig. 4a and b). Similarly, leaves in OsMYB3-overexpressing lines of Chao2-10 OE-C3, OE-C6, and OE-C9 accumulated 58.9 μg/g, 247.3 μg/g, and 79.0 μg/g anthocyanins, respectively, but leaves of the WT Chao2-10 did not (Supplementary Fig. 4a and b). In addition, OE-C3, OE-C6, and OE-C9 exhibited purple apiculus with anthocyanin accumulation, but the WT Chao2-10 did not (Supplementary Fig. 4c).
Overexpression of OsMYB3 in black rice cultivar Zixiangnuo1. a Anthocyanin content in leaves of Zixiangnuo1 and OsMYB3-overexpressed transgenic lines. b Comparison of leaves between Zixiangnuo1 and OE-Z3. c–i Relative expression level of anthocyanin biosynthetic genes in leaves of Zixiangnuo1 and OsMYB3-overexpressed transgenic lines. j–l Relative expression level of transcriptional regulatory genes in leaves of Zixiangnuo1 and OsMYB3-overexpressed transgenic lines. Error bars represent the standard deviation
qRT-PCR analysis showed that in leaves of OE-Z3, OE-Z4, and OE-Z5, the expression level of OsMYB3 and LBGs (OsF3H, OsDFR, OsANS, and OsUFGT) were significantly upregulated compared with the WT control, whereas the expression level of EBGs (OsCHS, OsCHI, and OsF3′H) together with several regulator genes OsKala4 and OsPAC1 remained unaffected (Fig. 4c–l). In leaves of OE-C3, OE-C6, and OE-C9, OsMYB3 and six anthocyanin biosynthetic genes (OsCHS, OsF3’H, OsF3H, OsDFR, OsANS, and OsUFGT) were significantly upregulated, while the expression of OsKala4 and OsPAC1 remained unaffected (Supplementary Fig. 4d-m). Overexpression of OsMYB3 activated more anthocyanin biosynthetic genes in leaves of Chao2-10 compared with the case in Zixiangnuo1, indicating that the activation effect of OsMYB3 on anthocyanin biosynthetic genes in rice leaves might vary among different genetic backgrounds. These results demonstrated that overexpression of OsMYB3 was able to complement the function of OsC1 in rice leaves.
Haplotype analysis of OsMYB3
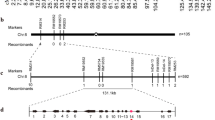
Li et al. (2020) demonstrated that OsMYB3 is also the responsible gene of a minor quantitative trait locus small grain 3 (SG3), which negatively regulates grain length in rice. Moreover, a 12-bp insertion in exon 3 of OsMYB3 was the functional mutation that was significantly associated with grain length in the indica subpopulation. OsMYB3 alleles with the 12-bp insertion were regarded as non-functional ones (i.e., sg3), because the insertion mutation caused a substitution of 7 continuous amino acid (aa) residues due to a frame shift followed by a loss of 20 aa residues at the C-terminus of the protein product due to a premature stop codon compared with the alleles without the insertion (Li et al. 2020).
To investigate whether the 12-bp insertion was also associated with the function of OsMYB3 in regulating anthocyanin biosynthesis in the pericarps, we analyzed the genomic sequences of OsMYB3 from 533 rice accessions. According to the variation in the coding sequence region, a total of 26 haplotypes (H1 to H26) of OsMYB3 were identified, and 4 haplotypes (H2, H3, H6, and H20) representing 312 rice accessions contained the 12-bp insertion (Fig. 5a and b). It was worth noticing that H3 that contains the 12-bp insertion presents in all the five rice subpopulations indica, japonica, intermidiate, aus, and aromatic. This indicated that H3 was likely the ancestral haplotype, and those ones without the 12-bp insertion should be the mutants.
We noticed that the expression level of OsMYB3 in the pericarps differs significantly between black rice and white rice (Supplementary Fig. 3c). A correlation analysis between the genomic polymorphisms of OsMYB3 (including promoter and terminator regions) and its expression level in pericarps was conducted. However, none of the genomic variations in OsMYB3 were found to be significantly correlated with its expression (Supplementary Fig. 5a). Furthermore, an association study between the genomic polymorphisms of OsMYB3 and rice pericarp phenotype also did not identify significantly correlated locus (Supplementary Fig. 5b).
Identification of differential metabolites associated with anthocyanin biosynthesis
To further investigate the affection of OsMYB3 expression on metabolites associated with anthocyanin biosynthesis, the LC–MS/MS analysis was used to profile the metabolites especially anthocyanins and flavonoids in grains of OsMYB3 knockout lines and the original Zixiangnuo1. The principal component analysis (PCA) of identified metabolites showed that three biological replicates of OsMYB3 knockout lines and Zixiangnuo1 were classified into two different groups (Supplementary Fig. 6a). A total of 205 metabolites were identified with 81 differentially accumulated metabolites (DAMs). Among the 81 DAMs, 75 were downregulated in OsMYB3 knockout lines compared with Zixiangnuo1, while 6 were upregulated (Supplementary Fig. 6b). KEGG enrichment showed that DAMs in anthocyanin biosynthesis pathway exhibited the highest rich factor and significance (Supplementary Fig. 6c).
In the anthocyanin biosynthesis pathway, a total of 20 DAMs were detected and normalized. Among them, 12 anthocyanin DAMs were completely undetected in OsMYB3 knockout lines, and 7 anthocyanin DAMs were markedly downregulated. Only one anthocyanin metabolite in OsMYB3-knockout lines was upregulated (Supplementary Fig. 7). These results indicated that the function of OsMYB3 has a wide impact on the accumulation of anthocyanin and flavonoid-related metabolites.
Discussion
By far, the bHLH (OsKala4) and WDR (OsPAC1 or OsTTG1) components regulating anthocyanin biosynthesis in rice pericarps have been characterized successively (Oikawa et al. 2015; Yang et al. 2021). In this study, we determined OsMYB3 as the R2R3-MYB regulator for anthocyanin biosynthesis in rice pericarps. OsMYB3 fell into the region of the rice genomic locus Kala3, which was previously demonstrated to control the black grain trait together with the loci Kala1 (OsDFR) and Kala4 according to the phenotypic observation in near isogenic lines (Maeda et al. 2014). However, the functions of OsMYB3 gene in anthocyanin biosynthesis had not been well characterized yet. The previous research showed that the mutated OsKala4, which caused the origination of black rice, expressed in both pericarps and leaves (Zheng et al. 2019), while OsPAC1 was also confirmed to participate in the activation of anthocyanin biosynthesis in both rice leaves and pericarps (Yang et al. 2021; Zheng et al. 2019). These results indicated that neither OsKala4 nor OsPAC1 is the pericarp-specific regulators for anthocyanin biosynthesis of black rice. Our results showed that among four different tissues leaves, spikes, pericarps, and endosperms, OsMYB3 expressed predominantly in pericarps. Therefore, OsMYB3 is the pericarp-specific regulator for anthocyanin biosynthesis in black rice. Because black rice was originated from the function acquired mutation of OsKala4, the original function of OsMYB3 in pericarps should not be associated with anthocyanin biosynthesis. A recent study demonstrated that OsMYB3 was also negatively regulating grain length (Li et al. 2020), which is consistent with our inference that OsMYB3 had other function in pericarps besides activating anthocyanin biosynthesis as a partner of OsKala4 and OsPAC1. Therefore, our study demonstrated an interesting paradigm how a pleiotropic gene evolves a novel function.
Our study showed that overexpression of OsMYB3 significantly enhanced anthocyanin accumulation in grains of black rice. This indicated that the expression level of OsMYB3 might be associated with anthocyanin content in grains of black rice. Actually, transcriptomic analysis showed significant differences in expression level between black rice and white rice. However, we did not identify DNA sequence variants associated with grain color or expression level of OsMYB3 in CDS or promoter region of OsMYB3. Probably, the difference in expression level of OsMYB3 between black rice and white rice might be associated with the expression level of certain upstream regulators. For instance, FaMYB10 was a key regulator of the anthocyanin synthesis pathway in strawberry, and the RAV transcription factor FaRAV1 activated the FaMYB10 to promote the synthesis of anthocyanin (Medina-Puche et al. 2014; Zhang et al. 2020). Moreover, knockout of OsMYB3 also caused significant downregulation of most flavonoid metabolites besides anthocyanins in black rice, indicating that OsMYB3 also plays roles in activating other branches of the flavonoid pathway. Taken together, OsMYB3 is an important regulator determining nutrients of rice, and characterization of OsMYB3 provides valuable implications to breed highly nutritious rice varieties.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Code availability.
Not applicable.
References
Albert NW, Davies KM, Lewis DH, Zhang H, Montefiori M, Brendolise C, Boase MR, Ngo H, Jameson PE, Schwinn KE (2014) A conserved network of transcriptional activators and repressors regulates anthocyanin pigmentation in eudicots. Plant Cell 26(3):962–980. https://doi.org/10.1105/tpc.113.122069
Al-Kanhal MA, Al-Mohizea IS, Al-Othaimeen AI, Khan MA (1999) Nutritive value of various rice based dishes in Saudi Arabia. Ecol Food Nutr 38(3):223–235. https://doi.org/10.1080/03670244.1999.9991579
Ban Y, Honda C, Hatsuyama Y, Igarashi M, Bessho H, Moriguchi T (2007) Isolation and functional analysis of a MYB transcription factor gene that is a key regulator for the development of red coloration in apple skin. Plant Cell Physiol 48(7):958–970. https://doi.org/10.1093/pcp/pcm066
Baudry A, Heim MA, Dubreucq B, Caboche M, Weisshaar B, Lepiniec L (2004) TT2, TT8, and TTG1 synergistically specify the expression of BANYULS and proanthocyanidin biosynthesis in Arabidopsis thaliana. Plant J 39(3):366–380. https://doi.org/10.1111/j.1365-313X.2004.02138.x
Cassidy A (2018) Berry anthocyanin intake and cardiovascular health. Mol Aspects Med 61:76–82. https://doi.org/10.1016/j.mam.2017.05.002
Druka A, Kudrna D, Rostoks N, Brueggeman R, von Wettstein D, Kleinhofs A (2003) Chalcone isomerase gene from rice (Oryza sativa) and barley (Hordeum vulgare): physical, genetic and mutation mapping. Gene 302(1–2):171–178. https://doi.org/10.1016/S0378-1119(02)01105-8
Espley RV, Hellens RP, Putterill J, Stevenson DE, Kutty-Amma S, Allan AC (2007) Red colouration in apple fruit is due to the activity of the MYB transcription factor, MdMYB10. Plant J 49(3):414–427. https://doi.org/10.1111/j.1365-313X.2006.02964.x
Feller A, Machemer K, Braun EL, Grotewold E (2011) Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. Plant J 66(1):94–116. https://doi.org/10.1111/j.1365-313X.2010.04459.x
Furukawa T, Maekawa M, Oki T, Suda I, Iida S, Shimada H, Takamure I, Kadowaki K (2007) The Rc and Rd genes are involved in proanthocyanidin synthesis in rice pericarp. Plant J 49(1):91–102. https://doi.org/10.1111/j.1365-313X.2006.02958.x
Gonzalez A, Zhao M, Leavitt JM, Lloyd AM (2008) Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J 53(5):814–827. https://doi.org/10.1111/j.1365-313X.2007.03373.x
Grotewold E, Drummond BJ, Bowen B, Peterson T (1994) The myb-homologous P gene controls phlobaphene pigmentation in maize floral organs by directly activating a flavonoid biosynthetic gene subset. Cell 76(3):543–553. https://doi.org/10.1016/0092-8674(94)90117-1
Guo H, Ling W, Wang Q, Liu C, Hu Y, Xia M, Feng X, Xia X (2007) Effect of anthocyanin-rich extract from black rice (Oryza sativa L. indica) on hyperlipidemia and insulin resistance in fructose-fed rats. Plant Foods Hum Nutr 62(1):1–6. https://doi.org/10.1007/s11130-006-0031-7
Hui C, Bin Y, Xiaoping Y, Long Y, Chunye C, Mantian M, Wenhua L (2010) Anticancer activities of an anthocyanin-rich extract from black rice against breast cancer cells in vitro and in vivo. Nutr Cancer 62(8):1128–1136. https://doi.org/10.1080/01635581.2010.494821
Kim JH, Lee YJ, Kim BG, Lim Y, Ahn JH (2008) Flavanone 3beta-hydroxylases from rice: key enzymes for favonol and anthocyanin biosynthesis. Mol Cells 25(2):312–316
Kranz HD, Denekamp M, Greco R, Jin H, Leyva A, Meissner RC, Petroni K, Urzainqui A, Bevan M, Martin C, Smeekens S, Tonelli C, Paz-Ares J, Weisshaar B (1998) Towards functional characterisation of the members of the R2R3-MYB gene family from Arabidopsis thaliana. Plant J 16(2):263–276. https://doi.org/10.1046/j.1365-313x.1998.00278.x
Kushwaha UKS (2016) Black rice: research, history and development. Springer International Publishing, Berlin
Li G, Zhu Y, Zhang Y, Lang J, Chen Y, Ling W (2013) Estimated daily flavonoid and stilbene intake from fruits, vegetables, and nuts and associations with lipid profiles in Chinese adults. J Acad Nutr Diet 113(6):786–794. https://doi.org/10.1016/j.jand.2013
Li Q, Lu L, Liu H, Bai X, Zhou X, Wu B, Yuan M, Yang L, Xing Y (2020) A minor QTL, SG3, encoding an R2R3-MYB protein, negatively controls grain length in rice. Theor Appl Genet 133(8):2387–2399. https://doi.org/10.1007/s00122-020-03606-z
Lin YJ, Chen H, Cao YL, Wu CY, Wen J, Li YF, Hua HX (2002) Establishment of high-efficiency Agrobacterium-mediated genetic transformation system of Mudanjiang 8. Acta Agron Sin 28(3):294–300
Lin-Wang K, Bolitho K, Grafton K, Kortstee A, Karunairetnam S, McGhie TK, Espley RV, Hellens RP, Allan AC (2010) An R2R3 MYB transcription factor associated with regulation of the anthocyanin biosynthetic pathway in Rosaceae. BMC Plant Biol 10:50–66. https://doi.org/10.1186/1471-2229-10-50
Lu X, Zhou Y, Wu T, Hao L (2014) Ameliorative effect of black rice anthocyanin on senescent mice induced by D-galactose. Food Funct 5(11):2892–2897. https://doi.org/10.1039/c4fo00391h
Ma X, Zhang Q, Zhu Q, Liu W, Chen Y, Qiu R, Wang B, Yang Z, Li H, Lin Y, Xie Y, Shen R, Chen S, Wang Z, Chen Y, Guo J, Chen L, Zhao X, Dong Z, Liu YG (2015) A robust CRISPR/Cas9 system for convenien, t high-efficiency multiplex genome editing in monocot and dicotplants. Mol Plant 8(8):1274–1284. https://doi.org/10.1016/j.molp.2015.04.007
Maeda H, Yamaguchi T, Omoteno M, Takarada T, Fujita K, Murata K, Iyama Y, Kojima Y, Morikawa M, Ozaki H, Mukaino N, Kidani Y, Ebitani T (2014) Genetic dissection of black grain rice by the development of a near isogenic line. Breed Sci 64(2):134–141. https://doi.org/10.1270/jsbbs.64.134
Medina-Puche L, Cumplido-Laso G, Amil-Ruiz F, Hoffmann T, Ring L, Rodriguez-Franco A, Caballero JL, Schwab W, Munoz-Blanco J, Blanco-Portales R (2014) MYB10 plays a major role in the regulation of flavonoid/phenylpropanoid metabolism during ripening of Fragaria × ananassa fruits. J Exp Bot 65(2):401–417. https://doi.org/10.1093/jxb/ert377
Meng L, Qi C, Wang C, Wang S, Zhou C, Ren Y, Cheng Z, Zhang X, Guo X, Zhao Z, Wang J, Lin Q, Zhu S, Wang H, Wang Z, Lei C, Wan J (2021) Determinant factors and regulatory systems for anthocyanin biosynthesis in rice apiculi and stigmas. Rice 14(1):1–18. https://doi.org/10.1186/s12284-021-00480-1
Miyake S, Takahashi N, Sasaki M, Kobayashi S, Tsubota K, Ozawa Y (2011) Vision preservation during retinal inflammation by anthocyanin-rich bilberry extract: cellular and molecular mechanism. Lab Invest 92(1):102–109. https://doi.org/10.1038/labinvest.2011.132
Oikawa T, Maeda H, Oguchi T, Yamaguchi T, Tanabe N, Ebana K, Yano M, Ebitani T, Izawa T (2015) The birth of a black rice gene and its local spread by introgression. Plant Cell 27(9):2401–2414. https://doi.org/10.1105/tpc.15.00310
Peiffer DS, Wang LS, Zimmerman NP, Ransom BW, Carmella SG, Kuo CT, Chen JH, Oshima K, Huang YW, Hecht SS, Stoner GD (2016) Dietary consumption of black raspberries or their anthocyanin constituents alters innate immune cell trafficking in esophageal cancer. Cancer Immunol Res 4(1):72–82. https://doi.org/10.1158/2326-6066.CIR-15-0091
Petroni K, Pilu R, Tonelli C (2014) Anthocyanins in corn: a wealth of genes for human health. Planta 240(5):901–911. https://doi.org/10.1007/s00425-014-2131-1
Qiao W, Wang Y, Xu R, Yang Z, Sun Y, Su L, Zhang L, Wang J, Huang J, Zheng X, Liu S, Tian Y, Chen L, Liu X, Lan J, Yang Q (2021) A functional chromogen gene C from wild rice is involved in a different anthocyanin biosynthesis pathway in indica and japonica. Theor Appl Genet 134:1531–1543. https://doi.org/10.1007/s00122-021-03787-1
Reddy AR, Scheffler B, Madhuri G, Srivastava MN, Kumar A, Sathyanarayanan PV, Nair S, Mohan M (1996) Chalcone synthase in rice (Oryza sativa L.): Detection of the CHS protein in seedlings and molecular mapping of the chs locus. Plant Mol Biol 32(4):735–743. https://doi.org/10.1007/BF00020214
Reddy AM, Reddy VS, Scheffler BE, Wienand U, Reddy AR (2007) Novel transgenic rice overexpressing anthocyanidin synthase accumulates a mixture of flavonoids leading to an increased antioxidant potential. Metab Eng 9(1):95–111. https://doi.org/10.1016/j.ymben.2006.09.003
Shih CH, Chu H, Tang LK, Sakamoto W, Maekawa M, Chu IK, Wang M, Lo C (2008) Functional characterization of key structural genes in rice flavonoid biosynthesis. Planta 228(6):1043–1054. https://doi.org/10.1007/s00425-008-0806-1
Stracke R, Ishihara H, Huep G, Barsch A, Mehrtens F, Niehaus K, Weisshaar B (2007) Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling. Plant J 50(4):660–677. https://doi.org/10.1111/j.1365-313X.2007.03078.x
Strathearn KE, Yousef GG, Grace MH, Roy SL, Tambe MA, Ferruzzi MG, Wu QL, Simon JE, Lila MA, Rochet JC (2014) Neuroprotective effects of anthocyanin- and proanthocyanidin-rich extracts in cellular models of Parkinson’s disease. Brain Res 1555:60–77. https://doi.org/10.1016/j.brainres.2014.01.047
Sun X, Zhang Z, Chen C, Wu W, Ren N, Jiang C, Yu J, Zhao Y, Zheng X, Yang Q, Zhang H, Li J, Li Z (2018) The C-S-A gene system regulates hull pigmentation and reveals evolution of anthocyanin biosynthesis pathway in rice. J Exp Bot 69(7):1485–1498. https://doi.org/10.1093/jxb/ery001
Takos AM, Jaffe FW, Jacob SR, Bogs J, Robinson SP, Walker AR (2006) Light-induced expression of a MYB gene regulates anthocyanin biosynthesis in red apples. Plant Physiol 142(3):1216–1232. https://doi.org/10.1104/pp.106.088104
Tanaka Y, Sasaki N, Ohmiya A (2008) Biosynthesis of plant pigments: anthocyanins, betalains and carotenoids. Plant J 54(4):733–749. https://doi.org/10.1111/j.1365-313X.2008.03447.x
Wang W, Mauleon R, Hu Z, Chebotarov D, Tai S, Wu Z, Li M, Zheng T, Fuentes RR, Zhang F et al (2018) Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 557:43–49. https://doi.org/10.1038/s41586-018-0063-9
Xu W, Dubos C, Lepiniec L (2015) Transcriptional control of flavonoid biosynthesis by MYB-bHLH-WDR complexes. Trends Plant Sci 20(3):176–185. https://doi.org/10.1016/j.tplants.2014.12.001
Yang J, Song B, Li Y, Liu J (2006) A simple and efficient method for rna extraction from potato tuber. J Agric Sci 14(2):297–298
Yang X, Wang J, Xia X, Zhang Z, He J, Nong B, Luo T, Feng R, Wu Y, Pan Y, Xiong F, Zeng Y, Chen C, Guo H, Xu Z, Li D, Deng G (2021) OsTTG1, a WD40 repeat gene, regulates anthocyanin biosynthesis in rice. Plant J. https://doi.org/10.1111/tpj.15285
Yao SL, Xu Y, Zhang YY, Lu YH (2013) Black rice and anthocyanins induce inhibition of cholesterol absorption in vitro. Food Funct 4(11):1602–1608. https://doi.org/10.1039/c3fo60196j
Zhang Q (2021) Purple tomatoes, black rice and food security. Nat Rev Genet 22(7):414. https://doi.org/10.1038/s41576-021-00359-3
Zhang Z, Shi Y, Ma Y, Yang X, Yin X, Zhang Y, Xiao Y, Liu W, Li Y, Li S, Liu X, Grierson D, Allan AC, Jiang G, Chen K (2020) The strawberry transcription factor FaRAV1 positively regulates anthocyanin accumulation by activation of FaMYB10 and anthocyanin pathway genes. Plant Biotechnol J 18(11):2267–2279. https://doi.org/10.1111/pbi.13382
Zhao H, Yao W, Ouyang Y, Yang W, Wang G, Lian X, Xing Y, Chen L, Xie W (2015) RiceVarMap: a comprehensive database of rice genomic variations. Nucleic Acids Res 43(D1):D1018–D1022. https://doi.org/10.1093/nar/gku894
Zheng J, Wu H, Zhu H, Huang C, Liu C, Chang Y, Kong Z, Zhou Z, Wang G, Lin Y, Chen H (2019) Determining factors, regulation system, and domestication of anthocyanin biosynthesis in rice leaves. New Phytol 223(2):705–721. https://doi.org/10.1111/nph.15807
Zhu F, Cai Y-Z, Bao J, Corke H (2010) Effect of γ-irradiation on phenolic compounds in rice grain. Food Chem 120(1):74–77. https://doi.org/10.1016/j.foodchem.2009.09.072
Zong W, Tang N, Yang J, Peng L, Ma S, Xu Y, Li G, Xiong L (2016) Feedback regulation of ABA signaling and biosynthesis by a bZIP transcription factor targets drought resistance related genes. Plant Physiol 171(4):2810–2825. https://doi.org/10.1104/pp.16.00469
Funding
This research was supported by National Natural Science Foundation of China (No. 32000233), Youth Innovation Talents Project of Department of Education of Guangdong Province (No. 2019KQNCX138), and the open funds of the National Key Laboratory of Crop Genetic Improvement (No. ZK201913).
Author information
Authors and Affiliations
Contributions
Jie Zheng performed all experiments, generated the figures and tables, wrote the original draft; Hao Wu performed all bioinformatics analyses and generated figures and tables; Mingchao Zhao performed the anthocyanin content measurement, PCR, and qRT-PCR; Zenan Yang performed rice pericarp sampling and RNA extraction; Zaihui Zhou performed rice transformation and grew the rice plants; Yongmei Guo collected rice germplasms; Yongjun Lin designed the experiments; and Hao Chen designed the experiments, reviewed, and edited the paper.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate.
Not applicable.
Consent for publication.
Not applicable.
Competing interests
The authors declare no competing interests.
Additional declarations for articles in life science journals that report the results of studies involving humans and/or animals.
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zheng, J., Wu, H., Zhao, M. et al. OsMYB3 is a R2R3-MYB gene responsible for anthocyanin biosynthesis in black rice. Mol Breeding 41, 51 (2021). https://doi.org/10.1007/s11032-021-01244-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-021-01244-x