Abstract
Tea plant (Camellia sinensis) has very long history of cultivation and abundant germplasm resources in China. Purple bud is a characteristic variety, which has attracted the attention of breeding researchers because it accumulated a large number of anthocyanins naturally. In many species, R2R3-MYB transcription factors (TFs) were proved to be involved in the regulation of anthocyanin biosynthesis. Research on anthocyanin metabolism has been relatively clear in some species, but that needs to be further elucidated in tea plants. In this research, an R2R3-MYB transcription factor CsMYB113 related to the anthocyanin accumulation regulation was identified from tea plants. Spatial and temporal expression analysis revealed differential expression of CsMYB113 among different tissues and organs, with highest expression occurring in the roots. Subcellular localization assays showed that CsMYB113 localized in the nucleus. Ectopic expression of CsMYB113 increased pigmentation and anthocyanin contents by the upregulation of the expression levels of genes in anthocyanin biosynthesis pathway among different tissues of Arabidopsis. Moreover, transient overexpression of 35S::CsMYB113 in tea plant increased the anthocyanin contents in the leaves. Our results indicated that CsMYB113 plays important role in the anthocyanin biosynthesis regulation in tea plants. It will also provide useful candidate gene for the modification of anthocyanin metabolism by genetic engineering in plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Anthocyanin, which is classified to the sub-class of flavonoids, is one of the most important metabolites existing in horticultural crops (Sun et al. 2016). Anthocyanin has been proved to participate in plant multiple biological and physiological processes including pigmentation, pollen transmission, seed dispersal, UV radiation protection, cold temperatures resistance, drought stress response, and pathogen defense (Karageorgou and Manetas 2006; Liu et al. 2018a; Stuurman et al. 2004; Castellarin et al. 2007; Christie et al. 1994). Moreover, anthocyanin also exhibits biological activities in humans, such as anticancer, antioxidant, and cardiovascular diseases protection (He and Giusti 2010; Clifford et al. 2015). Due to these benefits, the high anthocyanin content (purple pigmentation) has become one of important traits for the breeders in tea plant (Maritim et al. 2021).
Almost all the pathway genes related to anthocyanin metabolism were identified to date, and these genes are showing higher similarity between many species including tea (Xi et al. 2019; Matsui et al. 2008; Wei et al. 2019). The pathway is catalyzed stepwise by a series of biosynthetic enzymes, such as cinnamate 4-hydroxylase (C4H), chalcone synthase (CHS), chalcone isomerase (CHI), flavanone 3-hydroxylase (F3H), flavonoid 3′-hydroxylase (F3′H), flavonoid 3′,5′-hydroxylase (F3′5′H), dihydroflavonol 4-reductase (DFR), anthocyanidin synthase (ANS), anthocyanin O-methyltransferase gene (AOMT), UDP glucose: flavonoid 3-glucosyltransferase (UFGT), anthocyanidin-3-glucoside rhamnosyltransferase (3RT), and methyltransferase (MT) (Jaakola 2013; Perez-Diaz et al. 2016). It is generally demonstrated that the structural genes directly involved in anthocyanin metabolism are activated by numerous regulators, comprising MYB, basic helix-loop-helix (bHLH), and WD-repeat proteins (WDR) (Xu et al. 2015; Peng et al. 2019; Qi et al. 2020; Deng et al. 2021). These transcription factors (TFs) could independently or combine with cofactors to function as regulators in anthocyanin metabolism (Baudry et al. 2004; Quattrocchio et al. 2006). In most species, the MYB TFs superfamily is known as the one of largest families. Based on the number of MYB domain repeats, the MYB family can be divided into four classes, including single repeat (1R-MYB), two repeats (R2R3-MYB), three repeats (3R-MYB), and four repeats (4R-MYB) (Dubos et al. 2010). So far, a lot of R2R3-MYB TFs related to the anthocyanin biosynthesis regulation have been identified in many plants, including AtMYB75/PAP1 in Arabidopsis (Baudry et al. 2004), MdMYB10 in apple (Espley et al. 2007), IbMYB1a in sweet potato (Chu et al. 2013), SmMYB1 in eggplant (Docimo et al. 2016), FvMYB10 in strawberry (Zhang et al. 2017), PpMYB15 in peach (Cao et al. 2019), and PpMYB140 in pear (Ni et al. 2021).
Tea (Camellia sinensis) as one of the oldest (since 3000 BC) commercial crops and most popular nonalcoholic beverage is widely cultivated in over 50 countries and regions (Fang et al. 2012; Mondal et al. 2004). The popularity of tea is not only attributed to its specific aroma and taste, but also owing to the health benefits for human body. These medicinal properties derived from the various secondary metabolites in tea plants, such as catechins, anthocyanins, and theanine (Shi et al. 2011). However, anthocyanins were trace amount detected in most of the tea varieties (He et al. 2018). In recent years, purple foliage has attracted a lot of attention by the tea plant breeding programmers. Many purple strains have been reported in different tea growing countries (Hsu et al. 2012; Kerio et al. 2012; Kilel et al. 2013; Jiang et al. 2013). With the great efforts of many researchers, some R2R3-MYB TFs related to the anthocyanin pathway regulation have been identified from tea plants. The R2R3-MYB TFs CsAN1 could combine with CsGL3 and CsTTG1 and activate the expression of genes involve in anthocyanin biosynthesis (Sun et al. 2016). In ectopic transgenic tobacco plant leaves, CHS and 3GT were activated by the CsMYB6A which result in the significantly increment of anthocyanins (He et al. 2018). In transgenic tobacco lines, CsMYB5a and CsMYB5e were reported to play important role in the regulation of anthocyanins and proanthocyanidins (Jiang et al. 2018). The PAP1-like MYB gene was proposed as a key regulator in controlling anthocyanin metabolism (He et al. 2018; Wei et al. 2016). CsMYB75 promoted the biosynthesis of catechins and anthocyanins by upregulating the expression of CsGSTF1 in transgenic tobacco (Wei et al. 2019). Recently, a collection of 122 R2R3-MYB TFs have been identified in the chromosome level genome from Camellia sinensis (Chen et al. 2021). Therefore, R2R3-MYB TFs related to anthocyanin metabolism regulation still need to be explored fatherly in tea plants.
In this research, the biological function of the MYB TF CsMYB113 which related to anthocyanin metabolism was studied. The phylogenetic and localization study indicated CsMYB113 belonging to R2R3-MYB TF family. Ectopic expression of CsMYB113 in Arabidopsis led to significantly increased pigmentation and production of anthocyanins in roots, seeds, stems, and leaves. The real-time quantitative PCR (qRT-PCR) analyses revealed that CsMYB113 activates the expression of anthocyanin-related structural genes in 35S::CsMYB113 Arabidopsis transgenic plants. Moreover, the transient expression assays were carried out in the leaves of tea plant for functional verification. This study advances our knowledge related to the anthocyanin metabolism regulation for tea plant.
Materials and Methods
Plant Materials
Five-year-old tea plant cultivars [Camellia sinensis (L.) O. Kuntze cv. “Fudingdabai,” “Yingshuang,” and “Wuniuzao”] were planted in the Tea Germplasm Resources Nursery, Huazhong Agricultural University (HAZU, Wuhan, China). Nicotiana benthamiana was used for subcellular localization analysis and Arabidopsis thaliana ecotype Columbia was used for gene overexpression experiments. These two plant materials were planted in growth chamber with a 16-h light/8-h dark photoperiod under illumination of 10,000 lx (normal intensity). The growth temperature was set to 22/19 °C (light/dark). All the collected samples were rapidly snap-frozen in liquid nitrogen, and then transferred to − 80 °C for the further processing.
Total RNA Extraction and cDNA Synthesis
The RN09-EASYspin plant RNA kit (Aidlab, Beijing, China) was used to isolate total RNA for all the sampled materials. One percent agarose gel electrophoresis was employed to assess the quality for all the extracted RNA. RNA concentrations and integrity were checked by the Qubit RNA Assay Kit in a Qubit 2.0 Fluorometer (Life Technologies, USA). The single strand cDNA was synthesized from 1 μg of each total RNA using the TRUEscript RT Kit with gDNA Eraser (Aidlab, Beijing, China).
Isolation of CsMYB113 Gene and the Sequence Analysis
The predicted nucleic acid sequences of CsMYB113 genes have been screened out from C. sinensis genome database and our transcriptomic data (Guo et al. 2017; Wei et al. 2018). The open reading frame (ORF) of CsMYB113 was cloned by using 2 × Ultra-Pfu Master Mix (Dye Plus) (Aidlab, Beijing, China) with gene-specific primers (forward primer: 5′-ATGGAAGGTGTTCCTTTAGGAG-3′; reverse primer: 5′-TCAAAGATCCCAAAGGTCCAT-3′). PCR products was ligated into pTOPO-Blunt Simple vector (Aidlab, Beijing, China) and checked through sequencing (TSINGKE, Beijing, China). ProtComp 9.0 of softberry (http://linux1.softberry.com) was used to predict the subcellular localization signal of CsMYB113. ProtParam tool (http://web.expasy.org/protparam/) was used to calculate the theoretical molecular weight and isoelectronic point (pI). The ScanProsite (Expasy; SIB Swiss Institute of Bioinformatics, Switzerland) was used to analyze conserved motifs of CsMYB113 proteins. DNAMAN version 6.0 was utilized to multiple sequence alignment analysis.
Based on the neighbor-joining (NJ) method with 1000 bootstrap replications, molecular Evolutionary Genetics Analysis (MEGA) version 7.0 was used to construct phylogenetic tree. The sequences of R2R3-MYB family from ten plants including tea were downloaded from the BLAST of NCBI (https://blast.ncbi.nlm.nih.gov/Blast.cgi). These above gene accession IDs are as follows: CsAN1 (KU745295), AcMYB110a (AHY00342), MdMYB1 (ADQ27443.1), MdMYB3 (AEX08668), MdMYB10 (ACQ45201.1), MdMYB110a (BAM84362.1), AtMYB90/PAP2 (75,338,996), AtMYB75/PAP1 (75,333,682), AtMYB123/TT2 (27,151,707), AtMYB113 (Q9FNV9), AtMYB114(Q9FNV8.1), VvMYBA1 (BAD18977), PcMYB10.6 (AKV89252.1), PcMYB10.1 (AKV89247.1), MrMYB1 (ADG21957), MrMYB2 (ADG21958), TaMYB14 (AFJ53059), FaMYB11 (AFL02461), VvMYBPA1 (NP_001268160), VvMYBPA2 (ACK56131), ZmMYBP (NP_001278607), CsMYB5a (ATC41981.1), CsMYB5e (ATC41985.1), and CsMYB6A (AQW35194.1).
Gene Expression Analysis
The gene expression was detected by the real-time quantitative PCR (qRT-PCR) with the Applied Biosystems StepOne Plus™ Real-Time PCR System (ABI, Foster City, USA). According to the manufacturer’s instructions of 2 × SYBR Green qPCR Mix (Aidlab, Beijing, China), the amplification program consisted of one cycle at 94 °C for 3 min, followed by forty cycles at 94 °C for 10 s, 60 °C for 34 s, and then finally 60 °C for 1 h. The expression level of genes was calculated using the 2−∆∆Ct method (Livak and Schmittgen 2001); CsGAPDH (Camellia sinensis) and AtACTIN2 (Arabidopsis) were employed as the internal reference genes to monitor gene expression. Each biological sample was examined with three technical replicates. The gene-specific oligonucleotide primers utilized for qRT-PCR are shown in Supplementary Table S1.
Subcellular Localization Analysis
To determine the subcellular localization, the ORF of CsMYB113 (without the stop codon) was sub-cloned into the BamHI and XbaI site of pCAMBIA2300-GFP vector (Wang et al. 2014) to fuse with GFP by using One Step Cloning Kit (Vazyme, Nanjing, China). Transient expression in tobacco (Nicotiana benthamiana) leaves was performed as described previously (Sparkes et al. 2006). With the empty vector as a control, Agrobacterium tumefaciens (GV3101) harboring 35S::CsMYB113-GFP and nucleus co-localization marker CBLn-RFP (red) were infiltrated into 4-week-old tobacco epidermal cells together. The GFP fluorescence in transformed cells was detected under a confocal microscope (SP8, Leica, Wetzlar, Germany) within 36–48 h. All transient expression experiments were repeated independently more than three times.
Transformation of Arabidopsis with CsMYB113
The plant expression vectors were constructed by using the Gateway Cloning System (Invitrogen, New York, USA). The completed ORF of CsMYB113 was cloned into pDONR221 by BP reaction, and then inserted into a pH2GW7 vector by LR reaction. The expression vector (35S::CsMYB113-pH2GW7) was transferred into Agrobacterium tumefaciens (strain GV3101). The Arabidopsis transgene lines were subsequently conducted using the floral-dip method described previously (Clough and Bent 1998). Transgenic plants were chosen according to their resistance of hygromycin (Hyg). Putative transgenic Arabidopsis plants were selected on the Murashige and Skoog (MS) solid medium adding Hyg (50 mg/L). The positive transgenic plants were further checked by genomic PCR and qRT-PCR analysis. The homozygous T3 transgene lines were collected for anthocyanin content and qRT-PCR analysis.
Transient Overexpression of CsMYB113 in the Leaves of Tea Plant
Transient expression assays in the leaves of tea plant were conducted as reported by Mo et al. (Mo et al. 2015), with some modification. Agrobacterium tumefaciens (strain GV3101) harboring empty vector and 35S::CsMYB113 was cultured overnight in 50 ml LB liquid medium with rifampicin (50 mg/L) and spectinomycin (100 mg/L), respectively. After being centrifuged, re-suspended, and incubated, the suspensions of Agrobacterium tumefaciens harboring empty vector and 35S::CsMYB113 were injected into the different sides of the same leaf using a 1-ml plastic syringe, respectively. Ten days after infiltration, different leaves were collected for the anthocyanin biosynthesis analysis.
Anthocyanin Extraction and Measurement
Anthocyanin in leaves of tea plant was extracted as described previously (Sun et al. 2016). After extracted, the absorbance level of supernatant was detected in two absorbencies (535 nm and 650 nm), and the contents of anthocyanin were described as A535–A650 g−1 fresh weight (FW). Each sample was calculated from three repeats. Anthocyanin in different tissues of A. thaliana was measured according to (Chen et al. 2018). The supernatant was measured in two absorbencies (530 nm and 657 nm), and the contents of anthocyanin were described as A530 − 0.25 A657. Each sample was calculated from three repeats.
Statistical Analysis
All data were expressed as a mean value from three technical replicates with error bars indicating ± SD. The one-way ANOVA analysis of variance was used for identification of significant difference. The results were considered statistically significant and indicated with asterisks when P < 0.01. Different letters indicate significant differences when α = 0.05 by using the multiple comparisons.
Results
Cloning and Sequence Analysis of CsMYB113
The CsMYB113 gene was cloned from the tea plant cultivar “Fudingdabai” by RT-PCR. The complete ORF sequence is 726 bp (Fig. 1) which encodes for a polypeptide of 241 amino acids. The isoelectric point (pI) and molecular weight (MW) were 9.88 and 27.96 kDa, respectively. The gene sequence BLASTx against the Arabidopsis showed the highest similar with AtMYB113, and then, this gene was named as CsMYB113. Sequence analysis via the ScanProsite program revealed that the CsMYB113 protein contained two myb-type HTH (helix-turn-helix) DNA-binding domain profiles at N-terminus. Furthermore, the amino acid sequences similar with CsMYB113 were selected in other species, and multiple sequence alignment was performed. The result suggested that the R2 and R3 domains were highly conserved in these species. Moreover, 13th, 33rd, and 53rd positions of the R2 domain encoded the same tryptophan residues, which were conducive to keep the stability of the “helix-turn-helix” configuration of the MYB protein domain (Figs. 1 and 2). Phylogenetic analysis indicated that CsMYB113 was divided into subgroup 6 (S6) and similar with MYBs involved in anthocyanin biosynthesis regulation, suggesting that CsMYB113 may play important role in the anthocyanin metabolism regulation. CsMYB113 shared the highest amino acid sequence identity with tea plant CsAN1 (62%) and kiwifruit AcMYB110 (57.5%), respectively.
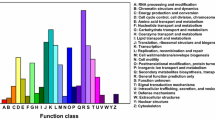
Conserved motif and phylogenetic analysis of amino acid sequences of CsMYB113 and anthocyanin-associated R2R3-MYB transcription factors in other species. A Conserved motifs analysis of CsMYB113. B Multiple sequence alignment of CsMYB113 and reported anthocyanin associated R2R3-MYB transcription factors in other species. C Phylogenetic tree of CsMYB113 and reported anthocyanin-associated R2R3-MYB transcription factors in tea and other plants. The green horizontal lines represent R2 domain and R3 domain; the red arrows represent three tryptophan (w) residues
Relative Gene Expression Analysis of CsMYB113 and the Subcellular Localization of Its Protein
To investigate the expression patterns of CsMYB113 gene in various tissues, qRT-PCR analysis was conducted to evaluate mRNA expression levels separated from ten tissues of “Fudingdabai.” As shown in Fig. 3A, CsMYB113 transcripts accumulated in all analyzed tissues. Overall, the transcript levels of CsMYB113 were the highest in the root, and decreased in the first leaf (FL). Second leaf (SL) showed the lowest expression level, which had no significant difference with other tissues, including the seed, bud, third leaf (TL), mature leaf (ML), old leaf (OL), stem, and flower.
Expression profile analysis of CsMYB113. A The expression profile of CsMYB113 genes in different tissues and organs of “Fuding dabai.” Data (error bars) are means (± SD) obtained from three technical replicates. Different letters indicate significant differences (α = 0.05). B Subcellular localization analysis of CsMYB113 in tobacco epidermal cells. Scale bar is 25 μm in the first row, 10 μm in the second row
To determine the subcellular localization of CsMYB113, the full-length ORF was fused in pCAMBIA2300 vector with the GFP reporter gene driven by the CaMV 35S promoter. Then, the construct was transformed into Agrobacterium and then infiltrated into tobacco epidermal cells. The results showed that GFP fluorescence in control (pCAMBIA2300-GFP) was ubiquitous distribution throughout the cell, while the GFP signal of CsMYB113-GFP overlaps with the nucleus co-localization fluorescence signal (as shown in pseudo colors green and red, respectively) (Fig. 3B). It is clearly indicated that CsMYB113 is exclusively localized in the nucleus of plant cell, which is similar with the CsAN1 (Sun et al. 2016). These results indicated that CsMYB113 is a transcription activator.
Overexpression of CsMYB113 Increased the Anthocyanin Contents in Transgenic Arabidopsis
To determine the potential role of CsMYB113 in the regulation of anthocyanin biosynthetic pathway, the 35S::CsMYB113 construct was employed to ectopically activate CsMYB113 expression into Arabidopsis (Col-0). There was visible phenotypic difference between Col-0 (wide type) plants and the transgenic lines after growing in a 16/8-h (light/dark) photoperiod under illumination of 10,000 lx in growth chamber. The overexpression of CsMYB113 resulted in the accumulation of anthocyanins in hypocotyl, veins, stems, seeds, and roots (Fig. 4A–G). And the anthocyanin contents in four tissues were significantly increased in transgenic Arabidopsis compared with wild type (Figs. 4H and 5A).
Phenotypes and anthocyanin contents of transgenic lines and wild type in Arabidopsis. A Phenotypes at hypocotyls of Arabidopsis seedlings. B Phenotypes of growing period. C Phenotypes of veins in the adult plants. D Phenotypes of stems in the adult plants. E Phenotypes of seeds. G Colors of tube during extracting anthocyanins from different tissues. F Phenotypes of roots
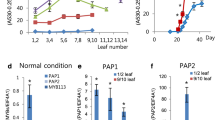
Relative anthocyanin contents and expression level of CsMYB113 in different tissues of Arabidopsis. A Increase of anthocyanin contents in different tissues of Arabidopsis. B Relative expression of CsMYB113 in different tissues of Arabidopsis. Data (error bars) are means (± SD) obtained from three technical replicates. With wild type as control, asterisks indicate significant differences (P < 0.01)
Further research showed that in transgenic lines, the anthocyanin contents in leaf, stem, root, and seed increased by 6.7-, 41.7-, 29.0-, and 4.5-fold compared with the wild type (average of the three lines), respectively. It indicated that the relative anthocyanin contents had significant differences in four tissues (Fig. 5). In order to verify whether the phenomenon is caused by the differential expression of CsMYB113, we detected the expression levels of CsMYB113 gene in four tissues of wild type (WT) and three overexpressing homozygous lines. The results showed that the successive decreasing order of the expression levels was tender stem, root, leaf, and seed (average of the three lines), which had the same trend as the increase folds of anthocyanin contents (Fig. 5). Therefore, we supposed that the differential expression of the CsMYB113 gene in four tissues leads to the differences in the anthocyanin contents. At the same time, combined with the results of CsMYB113 is root-specific expression in C. sinensis, which could further prove that the expression of CsMYB113 gene has obvious organizational differences.
Overexpression of CsMYB113 Increased the Expression Levels of Anthocyanin Biosynthetic Genes
R2R3-MYB TFs play important roles in activating structural genes involved in the anthocyanin biosynthesis. To further study the regulation of CsMYB113 gene, the expression levels of eight structural genes (AtPAL, AtCHS, AtCHI, AtF3H, AtF3’H, AtDFR, AtLDOX, AtUF3G) were detected between four tissues of wild-type and transgenic lines, respectively. In general, compared with the wild type, overexpressing of CsMYB113 could strongly increase the expression levels of AtCHI, AtF3H, AtDFR, AtLDOX, and AtUF3G (Fig. 6). In four tissues, the expression level of F3H gene was the most significantly upregulated by 5.5-fold in leaves (Fig. 6A), whereas the expression levels of CHI, F3H, and UF3G in stems were increased by 13-, 39-, and 114-fold, respectively (Fig. 6B). F3H, DFR, and UF3G genes in roots were upregulated by 42-, 20-, and 40-fold, respectively (Fig. 6C). The expression of each gene in seeds is less than fourfold (Fig. 6D). These results indicate that CsMYB113 can promote the expression levels of anthocyanin biosynthetic genes, thereby regulating the synthesis and accumulation of anthocyanin. However, the regulation profile has a certain difference, as CsMYB113 gene mainly upregulated different structural genes in the four tissues.
Transient Overexpression of CsMYB113 Stimulated Anthocyanin Accumulations in the Leaves of Tea Plant
With the development of research, transient transformation system has been established in many plants. It has the characteristics of high efficiency, short cycle, and fast realization of gene function verification, and has been widely used in herbs (tobacco, tomato, Arabidopsis, rice) and woody plants (citrus, poplar). Therefore, we further analyzed the function of CsMYB113 gene by using the transient expression system. We determine the content of anthocyanin and the expression level of CsMYB113 gene in leaves (Fig. 7). Leaves transformed with empty vector pK7WG2D (a) and target gene CsMYB113 (b) were collected, respectively. Control was set as the non-transformed leaves (ck). The results showed that the whole leaves grew well and were only slightly damaged near the injection hole. Moreover, phenotypic differences were observed among different treatments. Transformed with target gene CsMYB113 (b) appeared slight purple spots in leaves of “Fudingdabai” and “Wuniuzao” (Fig. 7A–C). The anthocyanin contents were significantly increased in leaves transformed with CsMYB113 gene (Fig. 7D). There was an almost twofold increase in leaves of three cultivars (P < 0.01).
Relative anthocyanin contents and expression level of CsMYB113 in the transient transfection of tea plant leaves. A–C Phenotypes and different colors during extracting anthocyanins with different treatments of “FD,” “YS,” and “WNZ.” D Anthocyanin contents of “FD,” “WNZ,” and “YS” with different treatments; E: Relative expression of CsMYB113 of “FD,” “WNZ,” and “YS” with different treatments. ck: Non-transformed leaves; a: Transformed leaves with pK7WG2D; b: Transformed leaves with 35S::CsMYB113-GFP. Data (error bars) are means (± SD) obtained from three technical replicates. With ck as control, asterisks indicate significant differences (P < 0.01). “Fudingdabai,” FD; “Yingshuang,” YS; “Wuniuzao,” WNZ
Meanwhile, qRT-PCR analysis showed that there was a significant increased in the expression level of CsMYB113 gene in transformed leaves (Fig. 7E). Compared with the non-transformed leaves (ck), the expression level increased almost 4.5 times in “Fudingdabai,” and almost 2 times in “Wuniuzao” and “Yingshuang.” The expression effect in “Fudingdabai” was better than that in the other two varieties. These results above further evidence that the CsMYB113 gene could transient expression in leaves of tea plant and the existence of CsMYB113 could accelerate the synthesis of anthocyanin in tea leaves to a certain extent.
Discussion
As a large subclass of MYB family, the R2R3-MYB transcription factor genes play important roles in anthocyanin metabolism regulation. In this research, an R2R3-MYB TF named CsMYB113 was successfully cloned from tea plant leaves. According to domain organization and phylogenetic analysis, CsMYB113 belongs to the S6 subgroup, which is important for the anthocyanin metabolism regulation (Liu et al. 2015). The protein sequence of CsMYB113 showed highest similarity with the CsAN1 protein (62%) from Camellia sinensis. In the “Zijuan” tea, the activation CsAN1 has been proved specifically unregulated anthocyanin biosynthesis genes to cause abnormal anthocyanin accumulation (Sun et al. 2016). A subcellular localization study showed that CsMYB113 is located in the nucleus (Fig. 3B), and indicated that CsMYB113 may act as a transcription factor. Taken as a whole, it indicates that CsMYB113 may function as transcription activating factor involved in the anthocyanin metabolism regulation in tea plants.
Further ectopic transgenic studies were implemented by overexpression of CsMYB113 in Arabidopsis plants. As compared to the wild-type Arabidopsis, the overexpressed lines were obviously turned to purple in the T3 homozygous plants, which was in accordance with obviously increased anthocyanin contents (Figs. 4 and 5). This finding is consistent with the exogenous gene expression patterns seen in Arabidopsis of other R2R3-MYB anthocyanin activator genes, such as MdMYB1 from apple (Takos et al. 2006), PUPRLE from cauliflower (Chiu et al. 2010), PyMYB10 from pear (Feng et al. 2010), EsMYBA1 from Herba epimedii (Huang et al. 2013a), MrMYB1 from Chinese bayberry (Huang et al. 2013b), BrMYB2 from Chinese cabbage (He et al. 2020), and FhPAP1 from Freesia hybrid (Li et al. 2020). Through tissue-specific analysis, it is found that the expression level of CsMYB113 in roots is the highest in all tissues. While there were little anthocyanins synthesized in the roots of “Fudingdabai.” Among the 124 R2R3-MYB transcription factors that have been discovered in Arabidopsis, AtMYB75, AtMYB90, AtMYB113, and AtMYB114 belong to the sixth subgroup. They usually form MBW complexes with bHLH and WD40, which positively regulate the structure of the late anthocyanin synthesis genes. Four belong to the fourth subfamily (AtMYB3, AtMYB4, AtMYB7, and AtMYB32) which acts as transcriptional repressors for upstream structural genes in the phenylpropane pathway of anthocyanin metabolism (Matsui et al. 2008). At present, studies have shown that CsMYB4-5 and CsMYB4-6 belong to such transcriptional repressors in tea plants, and they also have high expression levels in roots. They inhibit anthocyanin synthesis by downregulating the expression of C4H and 4CL (Gong et al. 2014). Therefore, we speculate that there are two opposite mechanisms of action in tea roots, positive regulators of CsMYB113 and negative regulators of CsMYB4-5 and CsMYB4-6, which work together to regulate anthocyanin content in tea roots.
The R2R3-MYB TFs, which regulate the synthesis of anthocyanin, are playing different roles in the various tissues of plants. In corn, the C1 gene regulates the biosynthesis of anthocyanin in aleurone (Cone et al. 1986). In GMYB10 overexpression transgenic tobacco plants (Nicotiana tabacum), the leaves, stems, and reproductive tissues turned to purple while no significantly anthocyanin accumulation in petal (Elomaa et al. 2003). Compared with the wild type, the MdMYBA overexpressing transgenic tobacco plants showed obviously increased anthocyanin in the reproductive tissues (Ban et al. 2007). In the MdMYB3 overexpression tobacco lines, the significantly increased anthocyanin pigmentation was observed in various tissues (Vimolmangkang et al. 2013). When ectopic expressed PyMYB10 in Arabidopsis, the anthocyanin content was significantly increased in immature seeds (Feng et al. 2010). In tea plants, the R2R3-MYB TFs showed abundant expression patterns, such as CsMYB4a expression was significantly higher in mature leaves, CsMYB42 is specifically expressed in pollen tubes, and CsMYB47 and CsMYB17 have the highest expression levels in leaves and buds (Li et al. 2017b; Wang et al. 2019; Chen et al. 2021). In this research, the contents of anthocyanin were determined in various tissues of transgenic Arabidopsis. It is worth noting that there were obvious differences in the level of accumulation of the anthocyanin in the different tissues (leaves, stem, roots, and seeds). The increments of anthocyanin in stem and roots were much higher than that in leaves and seeds (Fig. 5). The leave veins were obviously turned to purple while the mesophyll cells showed green color. We conclude that the CsMYB113 may play different roles in regulating the synthesis of anthocyanin among various tissues.
It is well known that MYB can promote gene expression levels of anthocyanin biosynthesis to active the anthocyanin accumulation. In transgenic cauliflower plants, upregulation of Purple (Pr) gene specifically activated three genes involved in anthocyanin biosynthesis which encodes F3’H, DFR, and LDOX (Chiu et al. 2010). In 35S::LfMYB113 transgenic Nicotiana tabacum plants, the expression levels of anthocyanin biosynthetic pathway genes were significantly increased including CHS, CHI, F3H, F3’H, DFR, ANS, and UFGT (Wen and Chu 2017). When ectopic expressed GhMYB1a in tobacco, the expression levels of CHS and F3H were significantly up-regulated than other genes related to the anthocyanin biosynthesis (Zhong et al. 2020). In the present study, the genes (AtPAL, AtCHS, AtCHI, AtF3H, AtF3’H, AtDFR, AtLDOX, AtUF3G) related to anthocyanin biosynthesis were significantly increased in transgenic Arabidopsis (stem, root, and seed) overexpressing CsMYB113. The relative expression levels increased in stem and root was much higher than that in the seed. There are only five genes (CHI, F3H, DFR, LDOX, UF3G) that were significantly increased in the leaves of transgenic Arabidopsis lines. These changes are highly consistent with the anthocyanin contents in the different tissues of transgenic lines. MYB TFs have been proved to regulate anthocyanin metabolism by combining with bHLH TFs in many plants (Gonzalez et al. 2008; Liu et al. 2018b; Li et al. 2017a). We conclude that CsMYB113 can integrate with tissue-specific bHLH to increase the transcription levels of anthocyanin biosynthesis, and lead to the anthocyanin content increment in different tissues.
In tea plants, the application of stable genetic transformation was limited by the problems of low transformation efficiency and difficulty in vitro regeneration (Mondal et al. 2004). Transient transformation system has many advantages compared to stable transformation, such as short period (the expression levels of genes could be analysis less than 12 h after transformation), high efficiency, easy to operation, and wide range of acceptor materials. Therefore, it has been frequently used for the gene function study in strawberry (Hoffmann et al. 2006), grape (Urso et al. 2013), orange (Jia and Wang 2014), persimmon (Mo et al. 2015), and other woody plants. In order to identify the possible function of CsMYB113 gene in tea plants, the homologous transient expression system was applied in this study. Transient transfection of tea plant leaves with the CsMYB113 overexpression caused the abnormal anthocyanin increment in three cultivars (“Fudingdabai,” “YingShuang,” and “Wuniuzao”) (Fig. 7), which is consistent with the result in Arabidopsis. It proved that CsMYB113 plays a vital role in the anthocyanin regulation in tea plants.
Conclusions
In the research, a R2R3-MYB TF CsMYB113 related to the regulation of anthocyanin biosynthesis was evaluated from tea plants. CsMYB113 was proved to localize in nucleus. Compared with wild type, some tissues (leaves, stems, roots, and seeds) were observed increased anthocyanin pigmentation inconsistent with the higher anthocyanin content in the CsMYB113 overexpression Arabidopsis plants. The ectopic expressed CsMYB113 in different tissues of transgenic Arabidopsis showed that the expression levels of genes related to anthocyanin biosynthesis were significantly enhanced. A distinguished anthocyanin content increment was detected in tea plant transient overexpression leaves. These results indicated that CsMYB113 plays a vital role in the regulation of anthocyanin metabolism.
References
Ban Y, Honda C, Hatsuyama Y, Igarashi M, Bessho H, Moriguchi T (2007) Isolation and functional analysis of a MYB transcription factor gene that is a key regulator for the development of red coloration in apple skin. Plant Cell Physiol 48(7):958–970
Baudry A, Heim MA, Dubreucq B, Caboche M, Weisshaar B, Lepiniec L (2004) TT2, TT8, and TTG1 synergistically specify the expression of BANYULS and proanthocyanidin biosynthesis in Arabidopsis thaliana. Plant J 39(3):366–380
Cao YL, Xie LF, Ma YY, Ren CH, Xing MY, Fu ZS, Wu XY, Yin XR, Xu CJ, Li X (2019) PpMYB15 and PpMYBF1 transcription factors are involved in regulating flavonol biosynthesis in peach fruit. J Agr Food Chem 67(2):644–652
Castellarin SD, Pfeiffer A, Sivilotti P, Degan M, Peterlunger E, Di Gaspero G (2007) Transcriptional regulation of anthocyanin biosynthesis in ripening fruits of grapevine under seasonal water deficit. Plant Cell Environ 30(11):1381–1399
Chen X, Wang P, Gu M, Lin X, Hou B, Zheng Y, Sun Y, Jin S, Ye N (2021) R2R3-MYB transcription factor family in tea plant (Camellia sinensis): genome-wide characterization, phylogeny, chromosome location, structure and expression patterns. Genomics 113(3):1565–1578
Chen XR, Wang Y, Zhao HH, Zhang XY, Wang XB, Li DW, Yu JL, Han CG (2018) Brassica yellows virus' movement protein upregulates anthocyanin accumulation, leading to the development of purple leaf symptoms on Arabidopsis thaliana. Sci Rep-Uk 8
Chiu LW, Zhou XJ, Burke S, Wu XL, Prior RL, Li L (2010) The purple cauliflower arises from activation of a MYB transcription factor. Plant Physiol 154(3):1470–1480
Christie PJ, Alfenito MR, Walbot V (1994) Impact of low-temperature stress on general phenylpropanoid and anthocyanin pathways - enhancement of transcript abundance and anthocyanin pigmentation in maize seedlings. Planta 194(4):541–549
Chu H, Jeong JC, Kim WJ, Chung DM, Jeon HK, Ahn YO, Kim SH, Lee HS, Kwak SS, Kim CY (2013) Expression of the sweetpotato R2R3-type IbMYB1a gene induces anthocyanin accumulation in Arabidopsis. Physiol Plantarum 148(2):189–199
Clifford T, Howatson G, West DJ, Stevenson EJ (2015) The potential benefits of red beetroot supplementation in health and disease. Nutrients 7(4):2801–2822
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16(6):735–743
Cone KC, Burr FA, Burr B (1986) Molecular analysis of the maize anthocyanin regulatory locus C1. P Natl Acad Sci USA 83(24):9631–9635
Deng J, Li JJ, Su MY, Lin ZY, Chen L, Yang PF (2021) A bHLH gene NnTT8 of Nelumbo nucifera regulates anthocyanin biosynthesis. Plant Physiol Bioch 158:518–523. https://doi.org/10.1016/j.plaphy.2020.11.038
Docimo T, Francese G, Ruggiero A, Batelli G, De Palma M, Bassolino L, Toppino L, Rotino GL, Mennella G, Tucci M (2016) Phenylpropanoids Accumulation in eggplant fruit: characterization of biosynthetic genes and regulation by a MYB transcription factor. Front Plant Sci 6
Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, Lepiniec L (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15(10):573–581
Elomaa P, Uimari A, Mehto M, Albert VA, Laitinen RAE, Teeri TH (2003) Activation of anthocyanin biosynthesis in Gerbera hybrida (Asteraceae) suggests conserved protein-protein and protein-promoter interactions between the anciently diverged monocots and eudicots. Plant Physiol 133(4):1831–1842
Espley RV, Hellens RP, Putterill J, Stevenson DE, Kutty-Amma S, Allan AC (2007) Red colouration in apple fruit is due to the activity of the MYB transcription factor, MdMYB10. Plant J 49(3):414–427
Fang W, Cheng H, Duan Y, Jiang X, Li X (2012) Genetic diversity and relationship of clonal tea (Camellia sinensis) cultivars in China as revealed by SSR markers. Plant Syst Evol 298(2):469–483
Feng SQ, Wang YL, Yang S, Xu YT, Chen XS (2010) Anthocyanin biosynthesis in pears is regulated by a R2R3-MYB transcription factor PyMYB10. Planta 232(1):245–255
Gong ND, Guo LL, Wang HX, Zhao L, Wang J, Wang WZ, Liu YJ, Wang YS, Gao LP, Xia T (2014) Cloning and functional verification of two MYB transcription factors in tea plant [Camellia sinensis (L.) ]. J Tea Sci 34(1):36–44
Gonzalez A, Zhao M, Leavitt JM, Lloyd AM (2008) Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J 53(5):814–827
Guo F, Guo YF, Wang P, Wang Y, Ni DJ (2017) Transcriptional profiling of catechins biosynthesis genes during tea plant leaf development. Planta 246(6):1139–1152
He JA, Giusti MM (2010) Anthocyanins: natural colorants with health-promoting properties. Annu Rev Food Sci T 1:163–187
He Q, Lu QQ, He YT, Wang YX, Zhang NN, Zhao WB, Zhang LG (2020) Dynamic changes of the anthocyanin biosynthesis mechanism during the development of heading Chinese cabbage (Brassica rapa L.) and Arabidopsis under the control of BrMYB2. Front Plant Sci 11
He XJ, Zhao XC, Gao LP, Shi XX, Dai XL, Liu YJ, Xia T, Wang YS (2018) Isolation and characterization of key genes that promote flavonoid accumulation in purple-leaf tea (Camellia sinensis L.). Sci Rep-Uk 8
Hoffmann T, Kalinowski G, Schwab W (2006) RNAi-induced silencing of gene expression in strawberry fruit (Fragaria x ananassa) by agroinfiltration: a rapid assay for gene function analysis. Plant J 48(5):818–826
Hsu CP, Shih YT, Lin BR, Chiu CF, Lin CC (2012) Inhibitory effect and mechanisms of an anthocyanins- and anthocyanidins-rich extract from purple-shoot tea on colorectal carcinoma cell proliferation. J Agr Food Chem 60(14):3686–3692
Huang WJ, Sun W, Lv HY, Luo M, Zeng SH, Pattanaik S, Yuan L, Wang Y (2013a) A R2R3-MYB transcription factor from Epimedium sagittatum regulates the flavonoid biosynthetic pathway. Plos One 8(8)
Huang YJ, Song S, Allan AC, Liu XF, Yin XR, Xu CJ, Chen KS (2013b) Differential activation of anthocyanin biosynthesis in Arabidopsis and tobacco over-expressing an R2R3 MYB from Chinese bayberry. Plant Cell Tiss Org 113(3):491–499
Jaakola L (2013) New insights into the regulation of anthocyanin biosynthesis in fruits. Trends Plant Sci 18(9):477–483
Jia HG, Wang N (2014) Targeted Genome Editing of Sweet Orange Using Cas9/sgRNA. Plos One 9(4)
Jiang LH, Shen XJ, Shoji T, Kanda T, Zhou JC, Zhao LM (2013) Characterization and activity of anthocyanins in Zijuan tea (Camellia sinensis var. kitamura). J Agr Food Chem 61(13):3306–3310
Jiang XL, Huang KY, Zheng GS, Hou H, Wang PQ, Jiang H, Zhao XC, Li MZ, Zhang SX, Liu YJ, Gao LP, Zhao L, Xia T (2018) CsMYB5a and CsMYB5e from Camellia sinensis differentially regulate anthocyanin and proanthocyanidin biosynthesis. Plant Sci 270:209–220
Karageorgou P, Manetas Y (2006) The importance of being red when young: anthocyanins and the protection of young leaves of Quercus coccifera from insect herbivory and excess light. Tree Physiol 26(5):613–621
Kerio LC, Wachira FN, Wanyoko JK, Rotich MK (2012) Characterization of anthocyanins in Kenyan teas: Extraction and identification. Food Chem 131(1):31–38. https://doi.org/10.1016/j.foodchem.2011.08.005
Kilel EC, Faraj AK, Wanyoko JK, Wachira FN, Mwingirwa V (2013) Green tea from purple leaf coloured tea clones in Kenya- their quality characteristics. Food Chem 141(2):769–775
Li CH, Qiu F, Ding L, Huang MZ, Huang SR, Yang GS, Yin JM (2017a) Anthocyanin biosynthesis regulation of DhMYB2 and DhbHLH1 in Dendrobium hybrids petals. Plant Physiol Bioch 112:335–345
Li MZ, Li YZ, Guo LL, Gong ND, Pang YZ, Jiang WB, Liu YJ, Jiang XL, Zhao L, Wang YS, Xie DY, Gao LP, Xia T (2017b) Functional characterization of tea (Camellia sinensis) MYB4a transcription factor using an integrative approach. Front Plant Sci 8
Li YQ, Shan XT, Tong LN, Wei C, Lu KY, Li SY, Kimani S, Wang SC, Wang L, Gao X (2020) The conserved and particular roles of the R2R3-MYB regulator FhPAP1 from Freesia hybrida in flower anthocyanin biosynthesis. Plant Cell Physiol 61(7):1365–1380
Liu JY, Osbourn A, Ma PD (2015) MYB transcription factors as regulators of phenylpropanoid metabolism in plants. Mol Plant 8(5):689–708
Liu Y, Tikunov Y, Schouten RE, Marcelis LFM, Visser RGF, Bovy A (2018a) Anthocyanin biosynthesis and degradation mechanisms in solanaceous vegetables: a review. Front Chem 6:52
Liu YJ, Hou H, Jiang XL, Wang PQ, Dai XL, Chen W, Gao LP, Xia T (2018b) A WD40 Repeat Protein from Camellia sinensis Regulates Anthocyanin and Proanthocyanidin Accumulation through the Formation of MYB-bHLH-WD40 Ternary Complexes. Int J Mol Sci 19(6)
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(T)(-Delta Delta C) method. Methods 25(4):402–408
Maritim TK, Masand M, Seth R, Sharma RK (2021) Transcriptional analysis reveals key insights into seasonal induced anthocyanin degradation and leaf color transition in purple tea (Camellia sinensis (L.) O. Kuntze). Sci Rep-Uk 11(1)
Matsui K, Umemura Y, Ohme-Takagi M (2008) AtMYBL2, a protein with a single MYB domain, acts as a negative regulator of anthocyanin biosynthesis in Arabidopsis. Plant J 55(6):954–967
Mo RL, Huang YM, Yang SC, Zhang QL, Luo ZR (2015) Development of Agrobacterium-mediated transient transformation in persimmon (Diospyros kaki Thunb.). Sci Hortic-Amsterdam 192:29–37
Mondal TK, Bhattacharya A, Laxmikumaran M, Ahuja PS (2004) Recent advances of tea (Camellia sinensis) biotechnology. Plant Cell Tiss Org 76(3):195–254
Ni JB, Premathilake AT, Gao YH, Yu WJ, Tao RY, Teng YW, Bai SL (2021) Ethylene-activated PpERF105 induces the expression of the repressor-type R2R3-MYB gene PpMYB140 to inhibit anthocyanin biosynthesis in red pear fruit. Plant J 105(1):167–181
Peng YY, Kui LW, Cooney JM, Wang T, Espley RV, Allan AC (2019) Differential regulation of the anthocyanin profile in purple kiwifruit (Actinidia species). Hortic Res-England 6
Perez-Diaz JR, Perez-Diaz J, Madrid-Espinoza J, Gonzalez-Villanueva E, Moreno Y, Ruiz-Lara S (2016) New member of the R2R3-MYB transcription factors family in grapevine suppresses the anthocyanin accumulation in the flowers of transgenic tobacco. Plant Mol Biol 90(1–2):63–76
Qi Y, Zhou L, Han LL, Zou HZ, Miao K, Wang Y (2020) PsbHLH1, a novel transcription factor involved in regulating anthocyanin biosynthesis in tree peony (Paeonia suffruticosa). Plant Physiol Bioch 154:396–408
Quattrocchio F, Verweij W, Kroon A, Spelt C, Mol J, Koes R (2006) PH4 of petunia is an R2R3 MYB protein that activates vacuolar acidification through interactions with basic-helix-loop-helix transcription factors of the anthocyanin pathway. Plant Cell 18(5):1274–1291
Shi CY, Yang H, Wei CL, Yu O, Zhang ZZ, Jiang CJ, Sun J, Li YY, Chen Q, Xia T, Wan XC (2011) Deep sequencing of the Camellia sinensis transcriptome revealed candidate genes for major metabolic pathways of tea-specific compounds. Bmc Genomics 12
Sparkes IA, Runions J, Kearns A, Hawes C (2006) Rapid, transient expression of fluorescent fusion proteins in tobacco plants and generation of stably transformed plants. Nat Protoc 1(4):2019–2025
Stuurman J, Hoballah ME, Broger L, Moore J, Basten C, Kuhlemeier C (2004) Dissection of floral pollination syndromes in petunia. Genetics 168(3):1585–1599
Sun BM, Zhu ZS, Cao PR, Chen H, Chen CM, Zhou X, Mao YH, Lei JJ, Jiang YP, Meng W, Wang YX, Liu SQ (2016) Purple foliage coloration in tea (Camellia sinensis L.) arises from activation of the R2R3-MYB transcription factor CsAN1. Sci Rep-Uk 6
Takos AM, Jaffe FW, Jacob SR, Bogs J, Robinson SP, Walker AR (2006) Light-induced expression of a MYB gene regulates anthocyanin biosynthesis in red apples. Plant Physiol 142(3):1216–1232
Urso S, Zottini M, Ruberti C, Lo Schiavo F, Stanca AM, Cattivelli L, Vale G (2013) An Agrobacterium tumefaciens-mediated gene silencing system for functional analysis in grapevine. Plant Cell Tiss Org 114(1):49–60
Vimolmangkang S, Han YP, Wei GC, Korban SS (2013) An apple MYB transcription factor, MdMYB3, is involved in regulation of anthocyanin biosynthesis and flower development. Bmc Plant Biol 13
Wang WD, Wang YH, Du YL, Zhao Z, Zhu XJ, Jiang X, Shu ZF, Yin Y, Li XH (2014) Overexpression of Camellia sinensis H1 histone gene confers abiotic stress tolerance in transgenic tobacco. Plant Cell Rep 33(11):1829–1841
Wang Y, Chang P, Pan J, Zhu J, Cui C, Ye X, Ma Y, Zhu X, Fang W, Jiang C (2019) Effect of aluminum and fluoride on R2R3-MYB transcription factor characterization and expression in Camellia sinensis. Biol Plantarum 63:298–307
Wei CL, Yang H, Wang SB, Zhao J, Liu C, Gao LP, Xia EH, Lu Y, Tai YL, She GB, Sun J, Cao HS, Tong W, Gao Q, Li YY, Deng WW, Jiang XL, Wang WZ, Chen Q, Zhang SH, Li HJ, Wu JL, Wang P, Li PH, Shi CY, Zheng FY, Jian JB, Huang B, Shan D, Shi MM, Fang CB, Yue Y, Li FD, Li DX, Wei S, Han B, Jiang CJ, Yin Y, Xia T, Zhang ZZ, Bennetzen JL, Zhao SC, Wan XC (2018) Draft genome sequence of Camellia sinensis var. sinensis provides insights into the evolution of the tea genome and tea quality. P Natl Acad Sci USA 115(18):E4151-E4158
Wei K, Wang LY, Zhang YZ, Ruan L, Li HL, Wu LY, Xu LY, Zhang CC, Zhou XG, Cheng H, Edwards R (2019) A coupled role for CsMYB75 and CsGSTF1 in anthocyanin hyperaccumulation in purple tea. Plant J 97(5):825–840
Wei K, Zhang YZ, Wu LY, Li HL, Ruan L, Bai PX, Zhang CC, Zhang F, Xu LY, Wang LY, Cheng H (2016) Gene expression analysis of bud and leaf color in tea. Plant Physiol Bioch 107:310–318
Wen CH, Chu FH (2017) A R2R3-MYB gene LfMYB113 is responsible for autumn leaf coloration in Formosan sweet gum (Liquidambar formosana Hance). Plant Cell Physiol 58(3):508–521
Xi WP, Feng J, Liu Y, Zhang SK, Zhao GH (2019) The R2R3-MYB transcription factor PaMYB10 is involved in anthocyanin biosynthesis in apricots and determines red blushed skin. Bmc Plant Biol 19
Xu WJ, Dubos C, Lepiniec L (2015) Transcriptional control of flavonoid biosynthesis by MYB-bHLH-WDR complexes. Trends Plant Sci 20(3):176–185
Zhang JX, Zhang YC, Dou YJ, Li WJ, Wang SM, Shi WJ, Sun YP, Zhang ZH (2017) Single nucleotide mutation in FvMYB10 may lead to the yellow fruit in Fragaria vesca. Mol Breeding 37 (3).
Zhong CM, Tang Y, Pang B, Li XK, Yang YP, Deng J, Feng CY, Li LF, Ren GP, Wang YQ, Peng JZ, Sun SL, Liang S, Wang XJ (2020) The R2R3-MYB transcription factor GhMYB1a regulates flavonol and anthocyanin accumulation in Gerbera hybrida. Hortic Res-England 7(1)
Funding
This work was supported by the National Natural Science Foundation of China (32072622, 31600556) and the Fundamental Research Funds for the Central Universities, Huazhong Agricultural University (2662019PY045).
Author information
Authors and Affiliations
Contributions
LS and FG designed the experiments. LS, MY, WL, and HL performed the experiments. LS and FG wrote the manuscript. WL and FG revise the manuscript. All authors provided helpful discussions and approved its final version.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key Message
• A R2R3-MYB transcription factor CsMYB113 clustered into S6 subgroup was identified in tea plant.
• CsMYB113 act as transcription factor and localize in the nucleus.
• Overexpression CsMYB113 in Arabidopsis enhanced pigmentation and anthocyanin accumulation in various organs.
• The anthocyanin contents were significantly increased in leaves transient transformed with CsMYB113 gene.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shui, L., Li, W., Yan, M. et al. Characterization of the R2R3-MYB Transcription Factor CsMYB113 Regulates Anthocyanin Biosynthesis in Tea Plants (Camellia sinensis). Plant Mol Biol Rep 41, 46–58 (2023). https://doi.org/10.1007/s11105-022-01348-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-022-01348-4