Abstract
Northern corn leaf blight (NCLB), which is caused by the hemibiotrophic fungal pathogen Setosphaeria turcica, is a devastating foliar disease that results in considerable maize yield losses. In the present study, quantitative trait locus (QTL) analysis was conducted across two environments using an ultra-high-density bin map constructed using recombinant inbred lines (RILs) derived from a cross between Ye478 and Qi319. A total of 11 QTLs, located on chromosomes 1, 4, 5, 6, 7, 8, 9, and 10, were detected that confer resistance to physiological race 0 of NCLB. Each QTL could explain 3.53–15.29% of the total phenotypic variation in disease resistance after artificial inoculation in two environments. Among these QTL, qNCLB7.02, which is located on chromosome 7, had the largest effect, accounting for 10.11 and 15.29% of the phenotypic variation in resistance in two field trials and BLUP. The common confidence interval (CI) for qNCLB7.02 was 1.4 Mb, according to the B73 RefGen_v3 sequence. The resistance effect of qNCLB7.02 was validated in 2016 by using chromosome segment substitution lines (CSSLs) derived from Qi319 as the donor in the genetic background of Ye478. The type 6 CSSL, which harbors introgressed qNCLB7.02, was found to be significantly associated with resistance to NCLB by linked marker bnlg1808 and exhibited greater resistance than the other CSSLs that did not carry this QTL (P = 0.0008). The combination of linkage mapping in RILs and validation in CSSLs is a powerful approach for the dissection of QTL for disease resistance in maize.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Northern corn leaf blight (NCLB), caused by the hemibiotrophic fungal pathogen Setosphaeria turcica, is one of the most devastating foliar diseases of maize. Typical symptoms include local lesions on leaves that progress until necrosis occurs (Pratt and Gordon 2005) and the accompanying yield loss can ranges from 20 to 50% and as high as 100% under severe epiphytotics (Perkins and Pedersen 1987; Tefferi et al. 1996). At present, physiological races 0, 1, 2, 3, N, and a few others have been identified, and races 0 and 1 have caused the most serious NCLB outbreaks in China (Gao et al. 2011; Liu et al. 1996). Reduced conservation tillage and application of chemical fungicides may diminish incidence of NCLB, but these methods are neither economical nor environmentally friendly for grain production. Thus, the use of resistance QTL or genes to improve the resistance of maize to NCLB is a cost-effective and environmentally friendly approach to reducing yield loss caused by this disease (Rish 2000). Substantial numbers of maize germplasm resources have been evaluated for NCLB resistance (Wang et al. 2012). Thus, the detection and pyramiding of quantitative trait loci (QTLs) for NCLB resistance will greatly improve the efficiency of marker-assisted breeding of maize for this trait (Pratt and Gordon 2005).
Many QTLs for NCLB resistance have been identified across the ten maize chromosomes. Using the F2:3 lines derived from a cross between Mo17 and B52, five QTL were detected for resistance to NCLB that explained 48% of the total phenotypic variation in that population (Freymark et al. 1994; Dingerdissen et al. 1996). QTLs located on chromosome bin 1.06/1.07, 3.07, 4.03, 5.04, and 6.05/6.06 were detected using F3 families derived from a cross between the early maturing European inbred lines D32 and D145 (Welz and Schechert 1999). One major QTL, qNLB8.06 DK888 , had been fine-mapped and delimited to an interval of 0.46 Mb including three candidate genes (Chung et al. 2010). Using a genetic map containing high-density single-nucleotide polymorphisms (SNPs) spaced at an average genetic distance of 0.74 cM, Chen et al. (2016) analyzed QTL for NCLB score and lesion size and found that qNCLB5.04 accounted for 19 and 20% of the phenotypic variation in these traits, respectively. In addition, several NCLB resistance-related loci have been detected in a genome-wide association study (GWAS) with a nested association mapping (NAM) population. An association panel consisting of 999 inbred lines fingerprinted with 56,110 SNP markers was used to find 81 SNPs significantly associated with NCLB resistance (Ding et al. 2015). Poland et al. (2011) evaluated a NAM population composed of 5000 inbred lines for resistance to NCLB and identified 29 QTLs with multiple alleles. Although numerous QTL that control resistance to NCLB have been identified in maize, current understanding of the genetic architecture and the molecular mechanisms of resistance to this disease remains rudimentary.
Ht1 is a single dominant gene that was identified in a cross between inbred lines GE440 and Ladyfinger popcorn (Hooker 1961). The dominant genes Ht1, Ht2, Ht3, Htm1, Htn1, N, and HtP and the recessive genes ht4 and rt have been identified (Bentolila et al. 1991; Van 2001; Simcox et al. 1993; Ogliari et al. 2005, 2007). They are independent of each other and resistant to specific physiological races, respectively. The genes Ht1, Ht2, and Ht3, which are located on chromosomes 2L, 8L, and 7L, respectively, convey resistance by inhibiting the expansion of chlorotic spots and decreasing the production of spores by the pathogen (Hilu and Hooker 1963; Hooker et al. 1963; Hooker and Kim 1973; Hooker 1981). Resistance conferred by HtN (Simcox and Bennetzen 1993) and Htm1 (Welz and Geiger 2000), which are located on chromosome 8L, is characterized by slower onset of epiphytotics after inoculation (Gevers 1975; Raymundo et al. 1981). Ht4 was located on chromosome 1L and showed the point of circular of fading green halo (Carson 1995). The Htn1 gene was cloned and found to code for a plant cell wall-associated receptor-like kinase with a pathogen and elicitor perception roles in the innate immune system of plants (Hurni et al. 2015).
Studies of the genetic architecture of complex traits and variations in maize have generally been performed using linkage analysis. However, most of the genetic maps based on low-throughput molecular markers have been of low density, which limits the efficiency and accuracy of QTL mapping and reduces coverage by genetic markers (Holland 2007). Compared to mapping using SSR markers, genotyping by sequencing (GBS) with next-generation sequencing technology is a powerful tool for SNP discovery and high-density genetic map construction. The development of ultra-high-density genetic maps using large populations of advanced RILs is an efficient way to identify QTL for complex agronomic traits (Zhou et al. 2016; Zhang et al. 2017). Three high-density integrated genetic linkage maps for maize that composite the SNP data from F2, RIL, and US-NAM mapping populations have been developed, which each comprise 6533, 2496, and 5296 markers with average distances between neighboring bins of 0.2, 0.72, and 0.28 cM, respectively (Chen et al. 2014; Li et al. 2015; Wen et al. 2015).
The genetic backgrounds of mapping populations can be complex and vulnerable to environmental effects (Eshed and Zamir 1994; Alonso-Blanco and Koornneef 2000). Secondary mapping populations such as chromosome segment substitution line (CSSL) can reduce such interference from the genetic background and are now being widely used for the mapping and cloning of genes and QTL that control important agronomic traits in maize. The use of CSSLs can improve the accuracy of QTL mapping by isolating a single or several chromosomal fragments from the donor parent in a genetic background identical to the recurrent parent. CSSLs have been useful for detecting QTL with smaller additive effects that had been masked by QTLs with larger additive effects in primary populations (Shim et al. 2010). These QTLs could then be resolved into individual Mendelian factors for further genetic analysis (Wang et al. 2012; Yuan et al. 2012). To date, several genes or QTL have been identified using CSSLs in maize (Zhang et al. 2011a, b; Marathi and Jena 2015). For example, a set of 130 CSSLs were developed by repeated backcrossing and selfing with Nongxi531 as the donor and H21 as the recipient. In total, 11 QTLs for kernel row number (KRN) were detected in three environments using stepwise regression, with between 9.87 and 19.44% of the phenotypic variation in KRN explained by a single QTL (Li et al. 2014).
In the present study, a maize RIL population was evaluated for resistance to NCLB race 0, and QTL mapping was conducted using an ultra-high-density bin map with 4183 bin markers constructed using these RILs. Furthermore, a set of CSSLs was developed for fine mapping and evaluated for resistance to NCLB to determine the potential of novel QTL. These results provide valuable information about maize resistance to NCLB, and markers developed within the qNCLB7.02 region may be useful in breeding for resistance to this disease.
Materials and methods
Plant materials and field design
A RIL population consisting of 314 F11 lines was developed by single-seed descent from a cross between the maize inbred lines Qi319 and Ye478. The RILs and both parents were planted in the field at Xin Zhou (43° 52′ N, 124° 82′ E), Shanxi Province, China, in 2015 and 2016. The resistant line Qi319 was derived from the hybrid 78599. The susceptible line Ye478 was derived from the cross U8112 × 5003 and developed into an elite inbred line that has been used widely in maize breeding programs in China. All of the RILs were arranged in the field in a randomized incomplete block design with two replications. Qi319 and Ye478 were planted in each block as the resistant and susceptible controls, respectively. Lines were grown in single 4-m rows spaced 0.25 m apart with a planting density of 60,000 plants/ha. Standard agricultural management practices were employed throughout each growing season in each location.
Evaluation of northern corn leaf blight resistance
Artificial inoculation was performed using crushed leaf material infected with Setosphaeria turcica as inoculum, belonging to the dominant race 0 through the identification of physiological races, that had been collected during the previous year at the same location. Experimental plots were inoculated at the V10–V12 growth stages by placing 20–30 pathogen-colonized sorghum seeds and crushing the diseased leaves into the leaf whorl of each plant (Carson 1995). The first evaluation of disease symptoms was conducted approximately 4 weeks after inoculation, and the second and subsequent evaluations were conducted at approximately 10-day intervals. Plants were scored on a scale for NCLB severity from 1 to 9 depending on the characteristics of the blight lesions (Supplemental Table S1) (Madden et al. 2007).
Analysis of phenotypic data
All descriptive statistics (e.g., mean, range, skewness, and kurtosis) for the parental lines and RILs across 2 years were calculated using Statistical Analysis System (SAS) software 9.0 (SAS Institute, Inc., Cary, NC Year). The broad-sense heritability (H2) of NCLB resistance across two environments was calculated according to Knapp et al. (1985). Heritability was calculated as H2 = δ2 g / (δ2 g + δ2 ge /e + δ2/er), where δ2 g is the genetic variance, δ2 ge is genotype × environment interaction, δ2 is the error variance, e is the number of environments, and r is the number of replications per environment. The estimates for δ2 g , δ2 ge , and δ2 were obtained using standard analysis of variance (ANOVA) with the general linear model procedure (PROC GLM) in SAS 9.0.
Linkage mapping for resistance to NCLB in RIL population
The RIL population was genotyped using a genotyping-by-sequencing (GBS) approach on an Illumina HiSeq2500 platform. A total of 137,699,000 reads with an average of 357,376 reads per individual RIL were generated, which is equivalent to approximately 0.07-fold coverage of the maize B73 RefGen_V3 genome for each individual RIL. A total of 88,268 SNPs were retained to identify bin markers to construct a high-density genetic map using 4183 bin markers. This map of the RIL population covers all ten maize chromosomes with a total genetic distance of 1545.65 cM and average distance between adjacent markers of 0.37 cM with a physical distance of about 0.51 Mb. The detailed genetic map used in this study was described in a previous study (Zhou et al. 2016).
QTL analysis for resistance to NCLB was conducted using the method of inclusive composite interval mapping (ICIM) in QTL IciMapping software 4.0 (Wang et al. 2014). ICIM applies a two-step strategy to effectively separate cofactor selection from the interval mapping process to more effectively control the background additive and dominance effects and improves mapping of QTL with additive effects compared to the composite interval mapping (CIM) method (Zeng 1994). For each of the datasets (2015, 2016, and BLUP), a significant threshold to affirm a putative QTL was obtained from 1000 permutations at P < 0.05 with a logarithm of odds (LOD) score > 3.5 (Doerge and Churchill 1996).
Validation of the qNCLB7.02, qNCLB8-1, and qNCLB8-2 loci in CSSL populations
Two hundred CSSLs were developed by using a combination of crossing, backcrossing, and molecular marker-assisted selection (MAS) between maize inbred lines Qi319 as the donor parent, and Ye478 as the recurrent parent. SSR detection was performed using the method described by Wang et al. (2009). The substituted segment length was estimated based on graphical genotypes (Young and Tanksley 1989). A chromosome segment that is flanked by two donor type markers (DD) is considered a100% donor type, a chromosome segment that is flanked by two recipient type markers (RR) is considered a 0% donor type and a chromosome segment that is flanked by one donor and one recipient type marker (DR) is considered a 50% donor type. The sum of the DD length and the two DR lengths was considered as the estimated length of a chromosome substituted segment. The length of the donor-substituted segments was calculated based on the marker’s location on the maize SSR linkage map (Zhu et al. 2009).
Each CSSL comprising evenly two to four introgression segments was genotyped based on the physical locations and genotypes of 201 SSRs distributed evenly across the ten chromosomes at an average marker interval of 9.94 Mb. To validate the effects of qNCLB7.02 and qNCLB8-1, 19 CSSLs that covered entire chromosomes 7 and 8 were developed. CSSL types 1 to 19 were developed by five cycles of backcrossing followed by two cycles of selfing (BC 5 F 2 ), using Ye478 as the recurrent parent and Qi319 as the donor of the randomized allele monitored by molecular marker-assisted selection. In 2016, CSSL types 1 to 19, together with the parental lines, were evaluated for NCLB resistance in Xin Zhou, Shanxi Province, China. The design for this field trial was consistent with that for the RIL population above. Mean separations for genotypes and NCLB resistance were performed using Student’s t test.
Results
Evaluation of phenotypic variation
Data for the phenotypic variation in resistance to race 0 of NCLB in these 314 RIL populations and their parental lines in 2015 and 2016 are presented in Table 1 and Supplemental Fig. S1A. Significant differences in resistance to NCLB were detected between Qi319 and Ye478 (P = 0.0016 in 2015 and P = 0.00058 in 2016), with average values of 2.1 and 7.32, respectively (Supplemental Fig. S1B). The average scale of RIL was used to represent the disease scale of its corresponding individual. A continuous distribution from highly resistant to complete susceptible implied that maize resistance to NCLB was quantitatively inherited (Supplemental Fig. S2). ANOVA revealed significant differences (P < 0.05) among genotypes and environments for NCLB in the RILs. The estimated H2 for NCLB resistance in the RIL population was 57.7% (Table 2).
QTL mapping of resistance to NCLB in RIL populations
A total of 11 QTL identified for resistance to NCLB were detected in the RIL mapping population, explaining 3.53–15.29% of the total phenotypic variation across two environments and BLUP (Table 3 and Fig. 1). In 2015, five additive QTLs were detected and the phenotypic variation explained by each QTL ranged from 3.78 to 10.11%. The most significant QTL, qNCLB7.02, was detected on chromosome 7 with a LOD score of 6.77. Among these QTL, the resistance alleles at qNCLB4.04, qNCLB7.02, qNCLB8-1, and qNCLB9-1 were derived from the resistant parent Qi319, while the resistance allele at qNCLB1.01 originated from the susceptible parent Ye478. In 2016, six QTL for resistance to NCLB were detected, including qNCLB5.06, qNCLB6.05, qNCLB7.02, qNCLB8-2, qNCLB9-2, and qNCLB10.04, each of which explained 3.53–15.29% of the total phenotypic variance. In addition, the CIs for these QTL spanned physical distances from 0.3 to 1.4 Mb, with an average of 0.63 Mb, compared to the B73 RefGen_v3 genome. Among these 11 QTL, qNCLB7.02, which explained 10.11 or 15.29% of the phenotypic variation in disease score in two environments and BLUP, was consistently detected and mapped to a position between bin markers Mk3399 and Mk3400, which spans about 1.4 Mb on the B73 RefGen_v3 sequence. The CI for qNCLB8-1 was 97.7–98.5 Mb in 2015 and that for qNCLB8-2 was 80.75–81.05 Mb in 2016, and these QTL explained 8.38 and 10.67% of the phenotypic variation in disease score in the RIL population in 2015 and 2106, respectively.
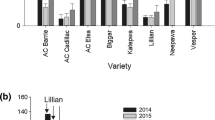
Detection of NCLB resistance QTL in two environments. a Diagram of QTL for NCLB on the whole genome in 314 RILs of Qi319×Ye478. The logarithm of odds profile, the relative position, and relevant markers are displayed using QTL cartographer version 3.5. b The major qNCLB7.02. c The minor qNCLB8-1 and qNCLB8-2. E1: the year of 2015 in Xin Zhou, Shanxi Province. E2: the year of 2016 in Xin Zhou, Shanxi Province
Validation of the qNCLB7.02, qNCLB8-1, and qNCLB8-2 loci in CSSL population
To validate qNCLB7.02, qNCLB8-1, and qNCLB8-2, 19 CSSLs covering chromosomes 7 and 8 were developed from a cross between maize inbred lines Qi319 as the donor parent, and Ye478 as the recurrent parent. Among these CSSLs, two CSSLs, type 6 and type 7, harbored qNCLB7.02 and one CSSL, type 13, harbored the CI for qNCLB8-1 and qNCLB8-2. The lengths of the introgressed fragments in CSSL type 6, type 7, and type 13 were 15, 8, and 77 Mb, respectively. Meanwhile, the background recovery rates in CSSL type 6, type 7, and type 13 were 91.6, 91.4, and 89.3%, respectively (Fig. 2 and Supplemental Table S2).
Combined with field phenotypic characteristics, CSSL type 6, which harbors the qNCLB7.02 introgression by linked marker bnlg1808, was found to be associated with resistance to NCLB with a disease score of 3.5. The mean disease scores for type 6 were lower than that of the susceptible parent Ye478, which has a disease score of 7.5 and differs significantly in resistance to NCLB from the other CSSLs at P = 0.0008 (Fig. 2, Supplemental Table S2 and Fig. S3). The interval sizes of the two QTLs on chromosome 8, qNCLB8-1 and qNCLB8-2, were 0.8 and 0.3 Mb on the B73 RefGen_v3 sequence, respectively, in 2015 and 2016. In the present study, the introgression fragment of CSSL type 13 that harbors qNCLB8-1 and qNCLB8-2 was associated with resistance to NCLB with a disease score of 5 at P = 0.002. Alleles at qNCLB8-1 or qNCLB8-2 might contribute improved disease resistance, so further fine-mapping in subsequent trials will be required (Fig. 2).
However, CSSLs type 6, type 10, and type 11 also harbor overlapping introgression fragments from the donor Qi319 inbred line. These overlapping regions conferred no significant difference in resistance to NCLB when comparing type 10 (P = 0.06) or type 11 (P = 0.053) with the susceptible parent Ye478, which suggests that the overlapping introgression fragments in type 10 and type 11 play no role in resistance to NCLB (Fig. 2 and Supplemental Table S2). Therefore, these results indicate that qNCLB7.02, qNCLB8-1, and qNCLB8-2 could improve resistance to NCLB in these CSSLs, and that CSSL type 6 and type 13 could be used to construct secondary backcrossing populations for fine-mapping and cloning of qNCLB7.02 and qNCLB8-1 or qNCLB8-2.
Discussion
A novel QTL for resistance to NCLB
QTL mapping has been an efficient strategy for the dissection of quantitative traits in maize breeding (Chen et al. 2014). However, the quality of genetic maps significantly affects the accuracy of QTL mapping. With the development of next-generation sequencing technology and high-density markers will greatly facilitate the identification of recombination events and the exact recombination breakpoints, which increases the power to detect QTL (Varshney et al. 2009; Schnable et al. 2009). In the present study, 11 QTL for resistance to NCLB were detected, and qNCLB7.02, which was consistently identified in all test environments and explained 10.11–15.29% of the phenotypic variation in disease score in the RIL population, was mapped to a position between bin markers of Mk3399 and Mk3400, which spans about 1.4 Mb on the B73 RefGen_v3 sequence (from physical position 137.05 to 138.45 Mb). Compared with the locations of QTL for NCLB resistance that have been identified in previous studies (Chen et al. 2016; Simcox and Bennetzen 1993), qNCLB7.02 from line Qi319 is likely to be a new locus for resistance to NCLB. Poland et al. (2011) detected 29 quantitative trait loci for resistance to NCLB in a NAM population. Their full QTL model could explain 77% of the variation in this trait, and subsequent association mapping identified one candidate gene (GRMZM2G043600) located near qNCLB7.02, which was 1.69 Mb downstream of the confidence interval for qNCLB7.02. Also, the confidence interval of qNCLB8-1, qNCLB8-2 does not include Ht2 or Htn1, which implicates qNCLB8-1 or qNCLB8-2 as new minor QTL on chromosome 8 that could improve resistance to NCLB (Zaitlin et al. 1992; Hurni et al. 2015; Chen et al. 2016).
Verification of stable QTL for NCLB resistance in CSSLs after artificial inoculation
Because the genetic backgrounds of mapping populations can obscure the effects of QTL in various environments (Eshed and Zamir 1994), identical genetic backgrounds in mapping population can allow QTL to be resolved into typical Mendelian factors and thereby greatly enhance the efficiency of fine mapping (Alonso-Blanco and Koornneef 2000). The construction of CSSLs from a cross between the inbred lines Qi319 and Ye478 has been a powerful strategy for precise QTL mapping of a genomic region. Our analysis using these CSSLs was convenient and complemented use of the RIL population, which allowed quick confirmation of individual QTL by identifying links between markers and QTL (Wang et al. 2012). The CSSLs have allowed more accurate estimates of genetic effects in a specific background (Yuan et al. 2012). Further, NCLB resistance is a complex, quantitatively inherited trait that is easily influenced by environmental conditions (Parlevliet 2002). To deal with this issue, experiments were performed in regions with severe NCLB outbreaks using artificial inoculation to increase the accuracy of phenotyping. After applying those strategies, 19 CSSLs covering chromosomes 7 and 8 were selected to validate the novel qNCLB7.02 identified for resistance to NCLB. The strategy we used was quite feasible, and will lay the foundation for future fine-mapping and cloning of stable and significant QTL for NCLB resistance.
Future application of NCLB resistance QTL in breeding programs
Many physiological races of S. trucica occur in China. The emergence of a new dominant race can be the prelude to maize varieties that were previously resistant becoming susceptible to NCLB. At present, a total of 12 physiological races have been identified, including races 0, 1, 2, 3, N, 12, 13, 1N, 2N, 123, 12N, and 123N. Among these, races 0, 1, and 0/1 dominate in the corn-growing areas of China (Gao et al. 2011; Zhang et al. 2011a, b; He et al. 2011; Sun et al. 2005). To date, accurate and rapid detection of the pathogen combined with quick deployment of resistant varieties have been the most effective ways to control NCLB. For example, DK888, a hybrid with resistance to multiple diseases, has been used as a donor of resistance alleles when constructing near-isogenic lines (NILs). The resistance of qNLB8.06 DK888 has been characterized by its specificity for race 0 and race 1 (Chung et al. 2010). Mo17 (as the resistant parent) has been used to perform linkage mapping of NCLB resistance after artificial inoculation with race 1 (Zheng et al. 2007). Further studies have shown race 1 to be the dominant race in the northeastern spring maize-growing area in China. Meanwhile, studies to identify and analyze disease resistance in five heterotic groups used in maize production and breeding in China have concluded that PN78599 germplasm from the PB heterotic group is highly resistant to NCLB and that hybrids including this germplasm also generally exhibit high degrees of disease resistance (Tong et al. 2005; Yu et al. 2011). In our study, the resistant parent Qi319 was also derived from the PB heterotic group, which implies that Qi319 should be resistant to race 0 and race 1.
Some genes that confer resistance to NCLB act quantitatively (horizontal resistance) and others act qualitatively (vertical resistance). Qualitative resistance is typically race-specific and controlled by single genes, whereas quantitative resistance is generally race-nonspecific and oligogenically or polygenically controlled (Geiger and Heun 1989; Mackay et al. 2009; Raihan et al. 2016). However, qualitative resistance should not simply be equated to major gene resistance, nor should quantitative resistance simply be equated to minor gene resistance. Because of the influences of the environment, materials, and physiological races of pathogens, qualitative resistance of maize to S. turcica might have a partial effect due to specificity against one or a few races, while quantitative resistance may have a more complete effect against a broader spectrum of races (Welz et al. 2000). For example, Htl, Ht2, and Ht3 confer qualitative resistance, while Htn1 confers quantitative resistance (Bentolila et al. 1991; Van 2001; Simcox et al. 1993; Ogliari et al. 2005; Hurni et al. 2015). Further, maize inbred line Ye8112 showed a high level of resistance to NCLB to race 0 but was susceptible to races 1 and N (Zhang 2013). Thus, stacking resistance alleles at multiple QTL into a single line could and should be achieved using phenotypic or MAS. Unlike morphological traits that can be directly selected by breeders, resistance can only be assessed under pathogen stress conditions, which are quite variable across years and locations and thus make phenotypic selection challenging. MAS can be effective if the genetic architecture controlling resistance to a given disease is clearly understood. A few of the genes or QTL underlying disease resistance in maize have already been applied in breeding resistant lines. For example, a resistance allele of the major head smut resistance QTL qHSR2.09 has been introgressed via MAS into 10 susceptible maize inbred lines. The 10 inbred lines that were converted to resistant phenotypes, and the hybrids derived from them, have all shown substantial improvement in resistance to head smut (Zhao et al. 2012; Zuo et al. 2014; Yang et al. 2017). Considering the genes or QTL underlying high qualitative resistance or quantitative resistance, several resistance genes have been introduced by MAS into multiline varieties that could be used in production and are resistant to multiple physiological races of NCLB. Our results provide important information for further fine mapping and ultimately cloning of genes controlling quantitative disease resistance traits. Future insights into the molecular mechanisms of disease resistance will in turn aid development of functional molecular markers and elite inbred lines, genomic selection in maize breeding populations, and breeding of NCLB-resistant hybrids.
References
Alonso-Blanco C, Koornneef M (2000) Naturally occurring variation in Arabidopsis: an underexploited resource for plant genetics. Trends Plant Sci 5(1):22–29. https://doi.org/10.1016/S1360-1385(99)01510-1
Bentolila S, Guitton C, Bouvet N, Sailland A, Nykaza S, Freyssinet G (1991) Identification of an RFLP marker tightly linked to the Ht1 gene in maize. Theor Appl Genet 82:393–398
Carson ML (1995) A new gene in maize conferring the chlorotic halo reaction to infection by Exserohilum turcicum. Plant Dis 79(7):717–720. https://doi.org/10.1094/PD-79-0717
Chen ZL, Wang BB, Dong XM, Liu H, Ren LH, Chen J, Hauck A, Song W, Lai JS (2014) An ultra-high density bin-map for rapid QTL mapping for tassel and ear architecture in a large F2 maize population. BMC Genomics 15(1):433–443. https://doi.org/10.1186/1471-2164-15-433
Chen GS, Wang XM, Long SS, Jaqueth J, Li BL, Yan JB, Ding JQ (2016) Mapping of QTL conferring resistance to northern corn leaf blight using high-density SNPs in maize. Mol Breeding 36(1):4–13. https://doi.org/10.1007/s11032-015-0421-3
Chung CL, Jamann T, Longfellow J, Nelson R (2010) Characterization and fine-mapping of a resistance locus for northern leaf blight in maize bin 8.06. Theor Appl Genet 121(2):205–227. https://doi.org/10.1007/s00122-010-1303-z
Ding JQ, Ali F, Chen GS, Li HH, Mahu G, Yang N, Narro L, Yan JB (2015) Genome-wide association mapping reveals novel sources of resistance to northern corn leaf blight in maize. BMC Plant Biol 15(1):206–217. https://doi.org/10.1186/s12870-015-0589-z
Dingerdissen AL, Geiger HH, Lee M, Schechert A, Welz HG (1996) Interval mapping of genes for quantitative resistance of maize to Setosphaeria turcica, cause of northern leaf blight in a tropical environment. Mol Breeding 2(2):143–156. https://doi.org/10.1007/BF00441429
Doerge RW, Churchill GA (1996) Permutation tests for multiple loci affecting a quantitative character. Genetics 142:285–294
Eshed Y, Zamir D (1994) A genomic library of Lycopersicon pennellii in L. esculentum: a tool for fine mapping of genes. Euphytica 79(3):175–179. https://doi.org/10.1007/BF00022516
Freymark PJ, Lee M, Martinson CA, Woodman WL (1994) Molecular-marker-facilitated investigation of host-plant response to Exserohilum turcicum in maize (Zea mays L.): components of resistance. Theor Appl Genet 88(3-4):305–313. https://doi.org/10.1007/BF00223637
Gao JX, Shu X, Gao ZG, Zhuang JH, Zhang XF, Zhang S (2011) Identification and dynamic analysis of physiological races of Exserohilum turcicum in Northeastern China in 2009 (in Chinese). J Maize Sci 19(3):138–140
Geiger HH, Heun M (1989) Genetics of quantitative resistance to fungal diseases. Annu Rev Phytopathol 27(1):317–341. https://doi.org/10.1146/annurev.py.27.090189.001533
Gevers HO (1975) A new major gene for resistance to Helminthosporium turcicum leaf blight of maize. Plant Dis Rep 59:296–299
He ZM, Li WC, Wang XM (2011) Identification of Setosphaeria turcica races in Yunnan and Guizhou provinces (in Chinese). Plant Prot 37:112–115
Hilu HM, Hooker AL (1963) Monogenic chlorotic-lesion resistance to Helminthosporium turcicum in corn seedlings. Phytopathology 53:909–914
Holland JB (2007) Genetic architecture of complex traits in plants. Curr Opin Plant Biol 10(2):156–161. https://doi.org/10.1016/j.pbi.2007.01.003
Hooker AL (1961) A new type of resistance in corn Helminthosporium turcicum. Plant Dis Rep 45:780–781
Hooker AL. (1981) Citation classic-reaction of corn seedings with male-sterile cytoplasm to Helminthosporium-turcicum[J]. Current Contents/Agriculture Biology & Environmental Sciences. 52:18–18
Hooker AL, Kim SK (1973) Monogenic and multigenic resistance to Helminthosporium turcicum in corn. Plant Dis Rep 57:586–589
Hooker AL, Johnson PE, Shurtleff MC (1963) Soil fertility and northern cornleaf blight infection. Agron J 55(4):411–417. https://doi.org/10.2134/agronj1963.00021962005500040039x
Hurni S, Scheuermann D, Krattinger SG, Kessel B, Wicker T, Herren G, Fitze MN, Breen J, Presterl T, Ouzunova M, Keller B (2015) The maize disease resistance gene Htn1 against northern corn leaf blight encodes a wall-associated receptor-like kinase. Proc Natl Acad Sci U S A 112:8781–8785
Knapp SJ, Stroup WW, Ross WM (1985) Exact confidence intervals for heritability on a progeny mean basis. Crop Sci 25(1):192–194. https://doi.org/10.2135/cropsci1985.0011183X002500010046x
Li CH, Li YX, Bradbury PJ, Wu X, Shi YS, Song YC (2015) Construction of high-quality recombination maps with low-coverage genomic sequencing for joint linkage analysis in maize. BMC Biol 13(1):78–90. https://doi.org/10.1186/s12915-015-0187-4
Li F, Jia HT, Liu L, Zhang CX, Liu J, Zhang ZX (2014) Quantitative trait loci mapping for kernel row number using chromosome segment substitution lines in maize GENET MOL RES 13 (1):1707–1716
Liu GS, Dong JG, Deng FY, Guo A, Zhang F, Zang MH (1996) Preliminary study of physiologic specialization and new nomenclature for Exserohilum turcicum of corn in China (in Chinese). Acta Phytophysiol Sin 26:305–310
Mackay TF, Stone EA, Ayroles JF (2009) The genetics of quantitative traits: challenges and prospects. Nat Rev Genet 10(8):565–577. https://doi.org/10.1038/nrg2612
Madden LV, Hughes G, van den Bosch F (2007) The study of plant disease epidemics. APS press, St. Paul, MN, p432
Marathi B, Jena KK (2015) Floral traits to enhance outcrossing for higher hybrid seed production in rice: present status and future prospects. Euphytica 201(1):1–14. https://doi.org/10.1007/s10681-014-1251-9
Ogliari JB, Guimaraes MA, Geraldi IO, Camargo LEA (2005) New resistance genes in the Zea mays L.-Exserohilum turcicum pathosystem. Genet Mol Biol 28(3):435–439. https://doi.org/10.1590/S1415-47572005000300017
Ogliari JB, Guirnaraes MA, Aranha Carnargo LE (2007) Chromosomal locations of the maize (Zea mays L.) HtP and rt genes that confer resistance to Exserohilum turcicum. Genet Mol Biol 30(3):630–634. https://doi.org/10.1590/S1415-47572007000400021
Parlevliet JE (2002) Durability of resistance against fungal, bacterial and viral pathogens: present situation. Euphytica 124(2):147–156. https://doi.org/10.1023/A:1015601731446
Perkins JM, Pedersen WL (1987) Disease development and yield losses associated with northern leaf blight on corn. Plant Dis 71(10):940–943. https://doi.org/10.1094/PD-71-0940
Poland JA, Bradbury PJ, Buckler ES, Nelson RJ (2011) Genome-wide nested association mapping of quantitative resistance to northern leaf blight in maize. Proc Natl Acad Sci U S A 108(17):6893–6898. https://doi.org/10.1073/pnas.1010894108
Pratt RC, Gordon SG (2005) Breeding for resistance to maize foliar pathogens. Plant Breed Rev 119–173
Raihan MS, Liu J, Huang J, Guo H, Pan QC, Yan JB (2016) Multi-environment QTL analysis of grain morphology traits and fine mapping of a kernel-width QTL in Zheng58 × SK maize population. Theor Appl Genet 129(8):1465–1477. https://doi.org/10.1007/s00122-016-2717-z
Raymundo AD, Hooker AL, Perkins JM (1981) Effect of gene HtN on development corn leaf blight epidemics. Plant Dis 65(4):327–330. https://doi.org/10.1094/PD-65-327
Rish NJ (2000) Searching for genetic deteminants in the new millennum. Nature 405(6788):847–856. https://doi.org/10.1038/35015718
Schnable PS, Ware D, Fulton RS, Stein JC, Wei F, Pasternak S et al (2009) The B73 maize genome: complexity, diversity, and dynamics. Science 326(5956):1112–1115. https://doi.org/10.1126/science.1178534
Shim RA, Angeles ER, Ashikari M, Takashi T (2010) Development and evaluation of Oryza glaberrima Steud. Chromosome segment substitution lines (CSSLs) in the background of O. sativa L. cv. ‘Koshihikari’. Breed Sci 60(5):613–619. https://doi.org/10.1270/jsbbs.60.613
Simcox KD, Bennetzen JL (1993) The use of molecular markers to study Setosphaeria turcica resistance in maize. Phytopathology 83(12):1326–1330. https://doi.org/10.1094/Phyto-83-1326
Simcox KD, Jeflrey L, Bennetzen (1993) Mapping the HtN resistance gene to the long arm of chromosome 8. Maize Genet Coop Newsl 67:118–119
Sun SQ, Wen LL, Gao DJ (2005) Identification of physiological races and mating type of Exserohilum turcicum (in Chinese). J Maize Sci 13:112–113
Tefferi A, Hulluka M, Welz HG (1996) Assessment of damage and grain yield loss in maize caused by northern leaf blight in western Ethiopia. J Plant Dis Prot 103:353–363
Tong SH, Chen G, Wang XJ, Wang ZY, Chen L, Yue H (2005) Resistance identification and evaluation of maize heterosis groups to the main disease in China. Rain Fed Crops 25:101–103
Van SD (2001) SCAR markers for the Ht1, Ht2, Ht3 and HtN1 resistance genes in maize. Proceedings of the 43rd Annual Maize Genetics Conference, Lake Geneva, WI, 43:134-136
Varshney RK, Nayak SN, May GD, Jackson SA (2009) Next-generation sequencing technologies and their implications for crop genetics and breeding. Trends Biotechnol 27:522–530
Wang BT, Shen CT, Zhang JZ, Li XH, Xi ZY (2009) Study of a micro-PCR reaction system in maize. J Maize Sci 17(4):29–31 (in Chinese, English abstract)
Wang ZQ, Yu CY, Liu X, Liu SJ, Yin CB, Liu LL, Lei JG, Jiang L, Yang C, Chen LM, Zhai HQ, Wan JM (2012) Identification of Indica rice chromosome segments for the improvement of Japonica inbreds and hybrids. Theor Appl Genet 124(7):1351–1364. https://doi.org/10.1007/s00122-012-1792-z
Wang JK, Li HH, ZHang LY Z, and Meng L (2014) Genetic resources program international maize and wheat improvement center (CIMMYT) Apdo Postal 6-641
Welz HG, Geiger HH (2000) Genes for resistance to northern corn leaf blight in diverse maize populations. Plant Breed 119:1–14
Welz HG, Schechert AW (1999) Dynamic gene action at QTL for resistance to Setosphaeria turcica in maize. Theor Appl Genet 98(6-7):1036–1045. https://doi.org/10.1007/s001220051165
Wen W, Li K, Alseek S, Omranian N, Zhao L, Zhou Y, Xiao YJ, Jin M, Yang N, Liu HJ, Florian A, Li WQ, Pan CQ, Yan JB, Fernie AR (2015) Genetic determinants of the network of primary metabolism and their relationships to plant performance in a maize recombinant inbred line population. Plant Cell 27(7):1839–1856. https://doi.org/10.1105/tpc.15.00208
Yang Q, Balint-Kurti P, Xu ML (2017) Quantitative disease resistance: dissection and adoption in maize. Mol Plant 10(3):402–413. https://doi.org/10.1016/j.molp.2017.02.004
Young ND, Tanksley SD (1989) Restriction fragment length polymorphism maps and the concept of graphical genotypes. Theor Appl Genet 77(1):95–101. https://doi.org/10.1007/BF00292322
Yu HY, Fu JF, Zhou RJ, Yan XR, Kang XJ (2011) Monitoring and causal analysis on epidemic of northern corn leaf blight in Liaoning Province (in Chinese). Hubei Agric Sci 0439-8114, 07-1375-02
Yuan L, Ding D, Li WH, Xie HL, Tang JH, Fu ZY (2012) Construction of single segment substitution lines (SSSLs) of the elite inbred lines in maize (in Chinese). J Maize Sci 20:52–55
Zaitlin D, Demars SJ, Gupta M (1992) Linkage of a second gene for NCLB resistance to molecular markers in maize. Maize Genet Coop Newsl 66:69–70
Zeng ZB (1994) Precision mapping of quantitative trait loci. Genetics 136(4):1457–1468
Zhang XL (2013) Study on the resistance of maize to northern corn leaf blight and southern corn rust. Doctorate Chinese Academy of Agricultural Sciences
Zhang H, Zhao Q, Sun ZZ, Zhang CQ, Feng Q, Tang SZ, Liang GH, Gu MH, Han B, Liu QQ (2011a) Development and high-throughput genotyping of substitution lines carrying the chromosome segments of indica 93-11 in the background of japonica Nipponbare. J Genet Genomics 38(12):603–611. https://doi.org/10.1016/j.jgg.2011.11.004
Zhang MH, Xu XD, Liu KJ, Dong HY, Jiang Y, Hu L (2011b) Physiological differentiation and races distribution of Exserohilum turcicum in China (in Chinese). J Maize Sci 19:138–141
Zhang CS, Zhou ZQ, Yong HJ, Zhang XC, Hao ZF, Zhang FJ, Li MS, Zhang DG, Li XH, Wang ZH, Weng JF (2017) Analysis of the genetic architecture of maize ear and grain morphological traits by combined linkage and association mapping. Theor Appl Genet 130(5):1011–1029. https://doi.org/10.1007/s00122-017-2867-7
Zhao XR, Tan GQ, Xing Y, Wei L, Chao Q, Zuo WL, Lubberstedt T, Xu ML (2012) Marker-assisted introgression of qHSR1 to improve maize resistance to head smut. Mol Breeding 30(2):1077–1088. https://doi.org/10.1007/s11032-011-9694-3
Zheng ZP, Liu XH, Huang YB, Li Z, He C, Tan ZB (2007) QTL mapping for resistance to northern corn leaf blight in maize Southwest China Journal of Agricultural Sciences 04:634–637
Zhu WY, Lin J, Yang DW, Zhao L, Zhang YD, Zhu Z, Chrn T, Wang CL (2009) Development of chromosome segment substitution lines derived from backcross between two sequenced rice cultivars, indica recipient93-11 and japonica donor nipponbare. Plant Mol Biol Rep 27(2):126–131. https://doi.org/10.1007/s11105-008-0054-3
Zhou ZQ ,Zhang CS, Zhou Y, Hao ZF, Wang ZH, Zeng X, Di H, Li MS, Zhang DG, Yong HJ, Zhang SH, Weng JF, Li XH (2016) Genetic dissection of maize plant architecture with an ultra-high density bin map based on recombinant inbred lines BMC Genomicsc 17:178
Zuo WL, Chao Q, Zhang N, Ye JR, Tan GQ, Li BL, Xing YX, Zhang BQ, Liu HJ, Fengler KA, Zhao J, Zhao XR, Chen YS, Lai JS, Yan JB, Xu ML (2014) A maize wall-associated kinase confers quantitative resistance to head smut. Nat Genet 47(2):151–157. https://doi.org/10.1038/ng.3170
Acknowledgments
This work was supported by a grant from the National Key Research and Development Program of China (2016YFD0101201) and the Chinese Academy of Agricultural Sciences (CAAS) Innovation Project.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(DOC 4.56 mb).
Rights and permissions
About this article
Cite this article
Wang, J., Xu, Z., Yang, J. et al. qNCLB7.02, a novel QTL for resistance to northern corn leaf blight in maize. Mol Breeding 38, 54 (2018). https://doi.org/10.1007/s11032-017-0770-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-017-0770-1