Abstract
In this review the authors present an overview of different molecular modeling campaigns dealing with the study and characterization of cyclodextrins (CDs) inclusion complexes with applicability to diverse biomedical and technological domains. The aim of this review is to present in a concise manner the new tendencies towards CDs molecular modeling studies in the context of a scientific computing era characterized by detailed and exhaustively validated molecular modeling protocols combined with and enormous and continuously growing computing power. Therefore, the present review covers research efforts reported in the last 5 years, including the simulation of native and modified CDs in a new and more detailed manner than what was possible in the past as well as their inclusion complexes with bioactive molecules studied by detailed protocols and exhaustive free-energy of binding calculations. Also, particular emphasis is devoted to the molecular modeling simulation of CDs included as part of drug delivery matrixes and intelligent nanodevices such as CD-based molecular motors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
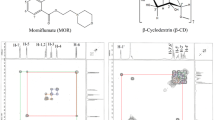
Cyclodextrins (CDs, Fig. 1) are a family of macromolecules constituted of α-1,4-linked D-glucopyranose subunits, which generate a significant interest in the biosciences due to their particularly useful properties [1,2,3,4]. Among the different fields of scientific interest, special focus has been placed on the field of biopharmaceutical applications due to their ability to modulate several properties affecting the performance and therapeutic profiles of drugs such as their solubility, stability, release, bioavailability and toxicity, among others. This behavior is mostly related to the capability of CDs to form inclusion complexes [5, 6], which are three-dimensional arrangements in which a guest molecule interacts through non-covalent forces with the host CD. In most cases, this interaction is a consequence of the inclusion of the guest molecule in the internal cavity of the CD, which can be considered hydrophobic if compared to the bulk of the solvent, and thus able of including whole apolar molecules and/or functional groups. Also, the internal cavity of the CD is able to accommodate buried solvent (water) molecules, meaning that during the inclusion process, the displacement of these solvent molecules is required. Thus, the structural and energetic features driving the inclusion of guest molecules into CDs is a complex phenomenon involving a dynamic network of intermolecular interactions and conformational and solvation/desolvation events.
Considering the above mentioned aspects, there is a great interest in understanding and rationalizing at a molecular level the physicochemical and structural properties of CDs inclusion complexes. In line with this, several experimental techniques such as crystallography, microcalorimetry, phase solubility analysis and nuclear magnetic resonance (NMR) [5, 6], among others, are widely applied as part of the preparation and characterization of these kinds of systems. However, the mentioned experimental techniques are somehow limited to provide detailed atomic resolution of the corresponding complexes as well as to clearly elucidate the key features related to the conformational properties of these systems. Also, performing experimental assays is particularly costly and tedious, and whenever the screening of a large number of compounds is envisioned, it becomes impractical or not feasible.
In this context, a wide range of molecular modeling methods able to characterize at a molecular level the behavior of molecules have been available for a relatively long time [7, 8]. These techniques constitute a set of computational tools and algorithms that are currently used throughout the biosciences community to shade light on the phenomena related to intermolecular interactions, and their conformational and energetic dependence. In particular, molecular modeling methods have found a wide utility for the study and characterization of CDs complexes, with some very good reviews been reported periodically [9,10,11,12]. It is noteworthy that scientific computing has evolved enormously in the last few years and will continue to grow exponentially in the years to come [13, 14]. In this context, tremendous advances are being made towards the development of improved calculation algorithms, refined force-fields, and new implementations of both classical and quantum ways of modeling small molecules and large biomolecules [15, 16]. Also, an enormous advance in computing power and high-performance computing infrastructure available for scientific computing purposes has been seen over the last few years. Additionally, the consolidation of molecular modeling packages and codes ported for execution on graphic processor units (GPU) [17] has opened up new horizons for the large-scale simulation of chemical systems in general and for CDs systems in particular.
The above mentioned scenario is evidenced by the numerous reports published recently on highly elaborated and large-scale molecular modeling simulation studies involving CDs systems. In light of this, and considering the authors´ experience regarding molecular modeling studies applied to CDs inclusion complexes, we present and discuss the state of the art techniques currently used in this scientific field, covering the most important reports presented in the last 5 years. We would like to highlight that our intention is not to exhaustively cover all the reports including some kind of molecular modeling study on CDs systems, since the number of reports in the last years is enormous. Rather than that, the driving force for literature selection was aimed towards presenting selected reports with particular emphasis in the new possibilities provided by modern advances in computational infrastructure and molecular modeling protocols.
The sections below have been organized considering the increasing complexity of the simulated system and the complexity of the technical approaches employed. We start our revision by exploring molecular modeling studies of pure CDs systems and modified CDs macromolecules. Afterwards, we revise molecular modeling tools aimed at the characterization of inclusion complexes, both binary and ternary. We end this revision with comments on complex supramolecular systems that include CDs as key constituents such as nanowires, rotaxanes and molecular engines. We hope the present review can serve as a useful material and as an introduction for scientists with no previous experience in molecular modeling studies regarding CDs as well as for experienced computational chemists who wish to start molecular modeling campaigns on systems involving CDs.
Molecular modeling of native CDs
The guest recognition properties of CDs constitute an important feature of this macromolecule that is frequently subjected to in silico screening and prediction by means of computational modeling methods. It has been widely reported that the selective inclusion of CDs is mostly driven by intermolecular forces that act to achieve and stabilize the resulting inclusion complex. In general, there is agreement on the fact that hydrophobic interactions play an important role in driving the complex formation and stabilization. Thus, the size, shape, and conformational degrees of freedom of the CD cavity are essential features to obtain stable and selective complexation. The most studied natural CDs are α, β and γCD, which consist of six, seven, and eight d-glucopyranose residues. These natural CDs exhibit differences in their hydrophobic cavity sizes, which in turn confers differential binding capacity.
The differential affinity of α, β, and γCD has been studied in detail by Ren et al. [18]. using molecular modeling techniques in light of the possibility of using CDs in soil remediation applications. In this study, not only was the differential capacity of natural CDs to sequester environmental pollutants (i.e. DDT) explored by applying high affinity and selective inclusion in the hydrophobic cavity, but also a detailed understanding of the inclusion mode was possible. As part of the soil remediation strategy, DDT included in the CD cavity was much more soluble in aqueous environments, and thus more bioaccesible for uptake by plant and microorganism able to elicit biodegradation. However, the biodegradable “soft point” within the DDT molecule was required to be exposed for biotransformation to occur, and thus knowing the specific mode of inclusion is critical for the intended use. These studies involved carefully designed simulation protocols, including extensive MD simulations to sample CDs conformational space combined with adaptive biasing force algorithmic approaches and steered molecular dynamics to determine the corresponding potential of mean force (PMF) profiles by describing the guest molecule binding. These theoretical studies suggested that all CDs were able to bind DDT and capture it within their hydrophobic cavity; however, only αCD was capable of further promoting its biodegradability due to the partial inclusion of DDT in the CD cavity. Consequently, an increasing role in the enhancement of solubility and biodegradation of DDT was proposed by these authors as a positive aspect of αCD compared to other natural CDs. The hypothesis was consistent with aqueous phase solubility and biodegradation experimental measurements, supporting the potential of the molecular modeling procedures applied to screen these kinds of properties.
In a homologous study, Cerón-Carrasco [19] explored the potential use of α, β and γCD as enhancers of the activity of the antioxidants carnosol and carnosic acid by formation of inclusion complexes. Combined docking and QM calculations, including the hybrid ONIOM approach, were used to accurately assess these features. These authors concluded that γCD was able to efficiently bind the antioxidant agent, while αCD was discarded in other research efforts. This study also demonstrated that molecular docking was a reliable tool to filter CD molecules based on their cavity volume. To date, a wide range of molecular docking tools are available, [20] many of which include user-friendly graphic user interfaces to set up and run docking experiments, making this tool easily accessible to researchers not specialized in molecular modeling algorithms.
As mentioned before, the execution of computing intensive studies dealing with the conformational sampling of CDs is nowadays possible. In this respect, Suárez et al. [21] performed a 5 ms molecular dynamics study to explore the conformational space accessible to native αCD, βCD and γCD. These authors described specific structural and dynamic features for each CD, including different numbers of water molecules bound to the hydrophobic cavity of the macromolecule. This structural observation is quite relevant to the design of inclusion complexes since the energetic cost associated with the displacement of water molecules buried in the CD cavity is related to the guest molecule affinity. In order to be able to perform these extended MD simulations, the authors employed GPU units, in this particular case through the use of the PMEMD. CUDA code, which is part of the AMBER suite of programs [22, 23].
In addition to natural CDs, large-ring cyclodextrins have attracted the attention in recent years as tools to prepare inclusion complexes with large guest molecules [24,25,26]. Ivanov et al. [27] applied a carefully designed MD protocol to study the conformation of CD38, a cyclodextrin formed by 38 sugar subunits and a principal component analysis to identify stable conformers along the 0.5 µs MD simulation trajectory. Overall deformations in the macroring were observed, with helix-turn shapes being formed at different regions of the macroring. These kinds of CDs present a considerable challenge for molecular modeling methods due to the large number of degrees of conformational freedom. To assess this issue, Khuntawee et al. [28] applied replica exchange molecular dynamics simulations to the conformational study of CD10, finding agreements between calculation and available crystal structures regarding the distortion of two CD helices.
Molecular modeling of modified cyclodextrins
The use of CDs as nanodevices with technological applicability requires not only a fine tuning of their guest inclusion properties, but also the compliance with other physicochemical properties that confer utility on them. Native CDs are often not well-suited to comply with these requirements, and thus their chemical modification is required, which in turn encompasses a very active and attractive area of research. Nowadays, the field of carbohydrate chemistry is rapidly growing, and is now possible to fine tune by means of synthetic method most natural (and simple) carbohydrate to more complex ones. As such, it is possible to monofunctionalize, perfunctionalize, perform simple or multiple capping, just to cite some procedures, on most CDs. The discussion on these procedures lies out of the scope of this review, and interested readers may refer to Chap. 9: “Advances in Cyclodextrin Chemistry”, bt Guiey and Sollogoub, in Modern Synthetic Methods in Carbohydrate Chemistry [29, 30]. By taking advantage of the reactivity of the hydroxyl moieties present in CDs, the chemical space available for chemical modification is enormous. Thus, the possibility of modeling the structural and physicochemical properties of modified CDs arises as a very powerful tool for the design of carefully tuned CDs structures. At this point, it needs to be addressed that from a synthetic point of view, the obtention of monofunctionalized CDs is quite challenging due to synthetic and/or purification limitations, and as such, the modeling of randomly substituted CDs should be carefully addressed before initiation the modeling campaign in an attempt to further explain experimental findings. Also, in silico organic chemistry is a rapidly growing area within molecular modeling methods [31] and is useful for the virtual exploration of the chemical space of a molecule. The discussion of these methods is out of the scope of this review, but we would just like to point out that nowadays it is technically feasible to generate and evaluate in silico a wide range of modified CDs by enumeration of chemical reactions using specific computer languages.
As an example of these kinds of studies, we can mention the research carried out by Shityakov et al. [32], who were able to model a system comprised of a sulfobutyl ether-and hydroxypropyl-β-cyclodextrin and their corresponding inclusion complexes with the general anesthetic propofol. In particular, the inclusion complex formed with sulfobutyl ether- β-cyclodextrin allowed an enhancement not only of the stability of propofol in solution, but also of the blood–brain barrier permeability compared to the non-modified CD. An agreement was found between the properties calculated from the simulated system and those from experimental measurements, demonstrating an adequate parametrization of the chemically modified CD. In this case, the well validated GLYCAM_06j-1 force field specifically developed for carbohydrates was used [33] and combined with the GAFF force field to assess the modified CD parametrization [34]. The modification of CDs with sulfobutyl ether moieties has also been modeled for other therapeutic compounds, finding also agreement between experimental and theoretical methods [35,36,37,38]. In a similar research, Altarsha et al. [39] simulated peracetylated βCD in supercritical carbon dioxide by combining parametrization procedures based on the GLYCAM force field and RESP charge calculation approaches [40, 41]. Again, the calculated structural properties were in agreement with experimental observations, evidencing that a self-closure of the molecular cavity was produced as a consequence of the chemical modifications, and thus elicited significant changes in the CD inclusion properties.
Also, the modification of natural CDs by conjugation with therapeutic compound can be modeled. Kordopati et al. [42] simulated a βCD conjugated with a luteinizing hormone-releasing hormone (LHRH) analogue in an attempt to enhance the biopharmaceutical limitations associated with the therapeutic use of peptides. In this work, explicit solvent molecular dynamics simulations demonstrated that aromatic segments of the therapeutic peptide were included in the interior core of the CD, which in turn might favor its bioavailability and/or biopharmaceutical properties. In this case, the structural behavior of the chemical spacer included in the conjugate was identified as a key feature driving the inclusion of the guest molecule, further supporting the potential of predictive molecular modeling simulations to perform virtual screening campaigns on these kinds of systems.
Structural characterization of CDs complexes
The structural and energetic behavior associated with the binding of guest molecules to CDs has been studied very actively by means of molecular modeling methods, including molecular docking, molecular dynamics, and free energy of binding analyses [43,44,45,46,47,48,49,50,51,52]. In particular, the possibility to calculate free energies of binding based on extended molecular configurations, constitutes fascinating and rapidly evolving area with utmost importance in the field of CDs systems characterization. In this way, the most elaborated modeling workflows frequently includes a final stage of free energy of binding analysis to address consistency with experimental affinity calculations. Unfortunately, it is impossible to address the details of each report presented in this review, but we would like to highlight the importance of this scientific field and refer the reader to recent reviews [53, 54]. The considerable increase in computing power observed in recent years has led to the possibility of applying very detailed and advanced protocols for the simulation of CDs:guest molecules inclusion complexes. Thus, a detailed characterization of the structure and energetic properties of inclusion complexes is now relatively easily accessible by using standard computing infrastructure. Nowadays, the possibility to execute explicit solvent simulation for inclusion complexes, allows the study of the critical role of water molecules in the complexation process. As such, solvent (water) molecules may be part of higher order (i.e. three or four-component) systems. Also, this increase in computing power has made possible the development of high quality simulation protocols able to model not only the interaction of the included molecule, but also the energetic landscape associated with the inclusion process that occurs when the guest molecule diffuses from the core of the solution into the interior of the CD cavity. Among these protocols, carefully designed umbrella sampling [55] schemes have been reported to characterize the inclusion of a wide variety of ligands. Therefore, molecular modeling methods stands as consolidated techniques to complement experimental techniques aimed at the structural characterization of CDs binary complexes and the dynamic features of their formation. In a further perspective, molecular modeling methods open up the possibility of screening large library of compounds as guest molecules, among other new possibilities.
In light of the commented aspects, the structural modeling of CDs host–guest binary systems has been and continues to be actively studied, and nowadays standard protocols and simulation conditions aimed at large scale prediction of inclusion complexes are available [56,57,58,59,60]. In the following subsection, a revision of several works dealing with modern aspects of the molecular modeling simulation of binary and ternary inclusion complexes will be presented.
Structural characterization of CDs binary complexes
As presented in the section above, modern simulation techniques dealing with the modeling of binary systems by means of carefully designed umbrella sampling protocols are able to provide detailed information about the mode of inclusion of guest compounds in CDs. Also, more advanced research efforts include detailed quantum calculations to assess the binding of the guest molecule. Sancho et al. [61], for example, studied the inclusion of chalcone and a chalcone derivative in βCD by determining the corresponding free energy profile obtained through the combination of umbrella sampling protocols and hybrid QM/MM methods. These protocols allowed elucidating the region of the molecule that is first included in the cavity of βCD, with these calculations being in agreement with experimental measurements. To date, the use of quantum calculations to assess the stability of binary complexes has been frequently reported [62,63,64]. In addition, combined methods such as computational calorimetry are possible by using different semi-empirical and quantum calculations based on MD trajectories [53, 65]. Taking a similar approach, Cao et al. [66] determined the PMF to model the encapsulation of fullerene (C60) in the hydrophobic cavity of γCD, describing the critical role of the van der Waals dispersion forces in the encapsulation process. As can be seen in Fig. 2, an interesting feature of CDs is their ability to form inclusion complexes with molecules with guest molecules with larger size than the buried area of the macromolecule cavity.
Finding the best CD system for a specific guest molecule is also feasible by applying molecular modeling methods. In this respect, Toth et al. [67] screened a set of 20 different CDs including natural and modified CDs to find the best host molecule that could act as a chiral selector for ofloxacin at different pH environments. As a result, they concluded that negatively charged CDs were the best host molecules among the whole set of the studied macromolecules. In this regard, molecular modeling techniques have been widely used to assess the chiral selection potential of CDs, which is of great pharmaceutical and industrial relevance [68,69,70,71,72,73,74,75,76]. Suliman et al. [77], for example, studied the inclusion mode of the drug baclofen in α and βCD by using docking and molecular dynamics simulations combined with quantum calculations. They observed a higher chiral discrimination potential of βCD compared to that of αCD, feature that is mostly derived from the different intermolecular hydrogen bonding networks that may be established with a particular guest molecule.
An important aspect that needs to be assessed when preparing CDs inclusion complexes is the corresponding stoichiometry. Although this experimental determination is very tedious, molecular modeling techniques are capable of assessing the number of guest molecules that may be included in the CD hydrophobic cavity [78, 79]. In this respect, Terekhova et al. [79] studied several binding modes and stoichiometries for the binding of different aromatic carboxylic acids to hydroxypropyl-γ-CD. A systematic screening of the ligand positioned in the CD cavity was performed considering different binding modes and calculating the associated energy of complexation. In most cases, a 1:2 stoichiometry was found, which was in accordance with experimental determinations.
Furthermore, a frequent requirement in the pharmaceutical field is the use of cosolvents as part of the designed formulations. It is well known that the structure of inclusion complexes with CDs may be considerably modified by the presence of cosolvent molecules and as such, it is relevant to anticipate its effect on the inclusion behavior. In this regard, Zhang et al. [80] exhaustively modeled the effect of commonly used alcohol cosolvents on the inclusion of two isoflavonoids (puerarin and daidzin), describing an intrinsic structural rigidization elicited by the cosolvent, which in turn diminished the affinity of the guest molecule. The same authors also modeled the effect of other cosolvents on the structural and inclusion properties of CDs [81]. In this way, the possibility of modeling in detail different explicit solvent systems as part of molecular modeling protocols arises as a powerful contribution to the in silico techniques.
To date, most of the studies dealing with the modeling of guest molecules:CDs binary systems have been oriented towards the optimization of the formulation properties. However, the possibility of modifying the pharmacodynamic interaction of drugs and biomacromolecules by encapsulation with CDs has been recently proposed [82,83,84,85,86,87]. In this respect, Sameena et al. [83] determined the binding of raloxifene to DNA and bovine serum albumin in the presence and absence of βCD. Using molecular docking studies, the authors described a clear competition between the CD inclusion and the binding to biomacromolecules.
Characterization of ternary inclusion complexes
Extensive reports have described the possibility of further enhancing the solubilization and complexation elicited by CD through the addition of auxiliary agents able to efficiently interact with binary complexes and form ternary aggregates. To date, a wide range of chemicals have been reported as potential third compounds; among them, the most extensively studied are aminoacids, polymers and hydroxyl agents [88]. Taking into account that each binary complex will be enhanced by specific third agents, molecular modeling methods arise as a very powerful screening technique to select the most adequate adjuvant, and thus making it possible to perform large-scale experimental screenings. Again, the increase in computing power observed in recent years also allows the execution of massive screening campaigns with this specific purpose.
As an example of the use of aminoacids as auxiliary agents, we can mention the study conducted by Sherje et al. [89], who performed molecular docking and molecular dynamics simulations to assess the effect of different auxiliary agents on the binding of Etodolac to HP-β-CD. From the screened compounds (which included aminoacids and polymers), l-arginine turned out to be the most effective compound, which was in line with the latest experimental determinations. From the modeling, it was observed that l-arginine increased the affinity of the included compound by establishing additional hydrogen bonds between the host and guest molecules. These observations are consistent with reports by Sapte et al. [90], who also modeled the effect of l-arginine on the binding of cefuroxime axetil to βCD. The effect of l-arginine in particular and aminoacids in general has been systematically described as a complexation adjuvant in several reports including molecular modeling techniques [91,92,93].
By using molecular docking, molecular dynamics, and free energy of binding analyses, Barbosa et al. [94] demonstrated that the addition of triethanolamine (TEA) to metotrexate:βCD complexes further enhanced the guest molecule solubility, which derived from a significantly increased electrostatic interaction component between the guest molecule and the CD. In this case, TEA was found to establish stable interactions in the region corresponding to the wide rim of βCD. As in the previously described reports, these theoretical observations were consistent with experimental studies.
Study of pH dependent inclusion
A particular case of the design and modeling of modified CDs is the preparation of macromolecules containing pH responsive moieties. These kinds of systems are being actively studied in order to prepare inclusion complexes able to act as pH-selective drug release devices. In this context, molecular modeling techniques are very powerful tools not only to model the thermodynamics properties driving the inclusion phenomena, but also to predict and model the pH dependence behavior of the host molecule, including the structural events on the CDs conformation that may trigger a drug-releasing event. Bearing in mind the different pH environments in living systems, applications in different fields of the biosciences are possible. An actively studied application is the formation of inclusion complexes with anticancer drugs [95,96,97,98,99,100], since it has been demonstrated that the pH in the environment of tumoral cells is lower than that of normal cells [101].
In this regard, Swiech et al. [102] modeled a pH sensitive system containing a lipoic acid derivative of βCD including the anticancer drug doxorubicin as guest molecule. The CD was chemically modified on the narrow rim using a triazole scaffold as linker moiety, which was self-included in the CD hydrophobic cavity upon a downshift of the pH environment. Molecular modeling studies involved standard molecular dynamics simulations using the YASARA software and the AMBER03 force-field. At pH 7.4 the inclusion of DOX was predicted, while at pH 5.5 the formation of a self-inclusion complex in which the lipoic acid was buried in the CD hydrophobic cavity was observed, thus eliciting the release of the included drug. It is noteworthy that these findings were consistent with experimental evidences showing a higher cellular toxicity of the DOX-loaded βCD complex towards immortal (HeLa) cell lines with respect to normal cells. In a previous report [103], these authors also performed molecular modeling studies based on a similar type of chemical modifications, but employing aromatic groups as self-inclusion scaffolds. A similar type of behavior regarding the release of the included drug was observed. These researchers also demonstrated that by mean of molecular dynamics simulations a 2–3 order of magnitude increase was obtained by carefully controlling the flexibility of the linker compared to the native CD [100]. This last work included the simulation of explicit solvent environments (DMSO, water, and DMSO:water mixtures), evidencing differential binding modes depending on the solvent composition.
Additionally, the possibility of obtaining CDs self-assemblies triggered by the presence of guest molecules can be studied in detail by means of molecular modeling methods. For example, De Sousa et al. [104] used molecular dynamics techniques to verify the formation of a nanostructure formed between ampicillin (AMP) and βCD in a 4:4 stoichiometry. This involved the inclusion of two AMP molecules in the βCD combined with the interaction of two additional molecules at the outer surface of the host macromolecule.
Clearly, molecular modeling techniques able to efficiently model the changes in ionization states and its corresponding structural effect on modified CDs constitute a very promising tool for the screening of specifically modified CDs. These modeling strategies can also benefit significantly from modern constant pH molecular dynamics simulation protocols [105,106,107].
Studies on systems containing multiple CD units
In recent years, much interest has been shown in supramolecular systems in which multiple CDs units forms part of the supramolecular system as covalent or non-covalent moieties, making it possible to obtain sophisticated materials able to comply with specific functions [108].
In the simplest example of the simulation of systems containing multiple CD units, Zhang et al. [109] applied molecular modeling techniques to model the dimerization behavior of βCD in different solvents. The authors used molecular dynamics simulations combined with umbrella sampling protocols to obtain the PMF for the association of two CD units. After modeling the dimerization process in explicit solvent environments, the author concluded that the solvents with the lowest hydrogen bond donor and acceptor properties (such as chloroform) strongly favored the dimerization event. Also, the effect of host molecules on the dimer formation was explored, finding that specific ligands may strengthen the stability of the dimer. This work also demonstrated the importance of explicitly modeling the solvent environment when dealing with CDs simulations. In this respect, the same research group [109, 110], studied in detail the necessity of explicitly including the solvent environment to adequately model CDs inclusion phenomena. In another report, Zhang et al. [111] reported the modeling of three inclusion complexes involving isoflavones and βCD dimers, applying molecular dynamics and PMF analyses. The systematic exploration of head to head (HH), head to tail (HT) and tail to tail (TT) oriented dimers combined with detailed thermodynamic analyses made it possible to shade light on the cooperativity of guest molecules regarding the stabilization of the dimeric structure. It is worth mentioning that the association of CD-complexes constitutes a feasible event once the inclusion complex is formed. The possibility of predicting this event is highly relevant, since some physical properties of the system (such as opalescence) may arise as a consequence of associations. Also, the drug/CDs aggregated may result in effective solubility enhancers themselves, which may in turn raise the solubility of drugs through non inclusion effects.
In addition, Staelens et al. [112] exhaustively studied the behavior of a system containing multiple units of β and γCD as nanotubes by means of molecular dynamics simulations, finding different orientations for both CDs. Also, the inclusion behavior of selected ligands towards these tubes was reported. The possibility of modeling the association of multiple CDs units is also feasible. Raffaini and Ganazzoli [113] reported a remarkable molecular dynamics study to describe the self-aggregation behavior of βCD monomers conjugated with porphyrin monomers. Liu et al. [114] also studied the self-inclusion of a modified αCD using molecular modeling methods by applying microsecond MD simulations and free-energy calculations. Notably, by determining the free-energy barrier for two self-inclusion pathways, the corresponding mechanism was elucidated. These kinds of models are of great utility for the design of new rotaxanes based on this kind of behavior.
In line with the use of CDs dimers as encapsulating agents, Wallace et al. [115] modeled γCD dimers covalently linked by succinamide and urea moieties. The binding of curcumin to these dimers was explored by MD simulations, with the free energies of binding being calculated by thermodynamic integration. These authors found agreement between the calculated and experimental results.
One particular device derived from systems containing multiple CD units is the nanosponges [116]. These systems are being actively explored in light of potential applications such as drug delivery devices and environmental detoxification of contaminants, among others. These nanosystems are prepared by reacting CDs with cross-linking agents. The behavior of the resulting nanosponges as delivery devices is highly dependent on the nature of this cross-linking agent, and thus this property can be studied using molecular modeling methods. In these findings, Raffaini et al. [117] applied molecular dynamics simulations, both in vacuo and in an explicit water model, to study nanosponges containing different ratios of βCD and two cross-linking agents (pyromellitic dianhydride and diphenyl carbonate). The resulting models allowed a characterization of the hydration properties of the nanosponge and the identification of water molecule distributions within the material and its surface.
Also, the effect of different physicochemical properties on CD mulimerization may be studied at atomic detail by applying molecular modeling methods. In this respect, Mixcoha et al. [118] reported extended molecular dynamics simulations in which the aggregation of α and βCDs were studied as a function of temperature in the bulk of the solvent. The higher tendency of βCD to form aggregates compared to αCD was modeled and described at an atomistic level. The obtained conclusions were in agreement with experimental observations.
Finally, systems containing multiple CD units may also be useful as chemical reaction platforms and thus merit simulation efforts. For instance, Takayanagi et al. [119] studied by means of molecular modeling methods the utility of CDs as catalysts of polymerization reactions. They observed that the reaction proceeded more efficiently by the cooperation of multiple βCDs molecules.
Modeling CDs in complex systems
It has been remarked many times throughout this review that the enormous increase in computing power observed over the last few years has strengthened in an unprecedented way the possibilities of modeling not only CDs in solution, but also complex systems containing CDs. In this section, some remarkable research efforts on these kinds of simulations are included.
One widely studied property of CDs in the biopharmaceutical field is its possibility of enhancing the bioavailability of drugs, feature that has been reported because it is related to the interaction of CDs with biological membranes [120, 121]. In line with the modeling of these properties, Khuntawee et al. [122] performed a detailed all-atom MD simulation to describe the interaction of βCD with a lipid bilayer system. The whole system was simulated on the microsecond scale, and from the resulting MD trajectories, several features were studied, such as the permeation and insertion of βCD into the lipid bilayer, the energetic of the association between βCD and the membrane, and the conformational changes produced in the βCD truncated cone shape as a consequence of the insertion into the lipid. It was observed that βCD was able to spontaneously migrate towards the lipid surface, but it was not further inserted into the membrane bilayer core region. Also, a significant change in the βCD conformation was observed as a consequence of the interaction with the bilayer, suggesting that it was closely related to the drug-releasing properties of βCD upon interaction with the biological membrane. This report clearly demonstrated how in silico models are of great value to the design of mucoadhesive drug delivery CDs devices.
Another field in which CDs are being actively modeled as part of complex macromolecular systems is the development of molecular motors [123]. Molecular motors are nanoscale devices which transform the free energy of chemical reactions into mechanical work. These devices are being studied for multiple applications, involving molecular recognition, nanosensors, and transducers. Rotaxanes, including CDs, are under active research in this respect [124]. In line with these strategies, Singharoy et al. [125] reported a simulation scheme developed by combining several molecular modeling approaches in order to capture the structural and energetic behavior of CD based on molecular motors. Similarly, Liu et al. [126] modeled by means of MD simulations and free-energy calculations the compression and decompression strokes of a molecular engine prepared using modified βCDs. The obtainment of the corresponding PMF and its subsequent analyses helped to understand, at an atomic level, the forces involved in these kinds of molecular engines and also allowed the design of new ones based on specifically tuned CDs that may exhibit higher efficiencies. The application of modern high throughput molecular modeling techniques to the study and design of molecular motors constitutes an incipient and rapidly growing scientific field [127, 128].
Another field in which complex systems containing multiple CD units are being designed and exhaustively characterized by applying molecular modeling methods is the field of molecular wires. These devices are conductive polymers obtained through the formation of inclusion complexes in a polymer chain [129]. CDs are being actively studied as components for molecular wires, with molecular modeling techniques being of significant interest since the prediction of the CDs orientation and dynamic behavior is crucial for the electronic and conductive properties of the molecular wire. In this respect, Tallury et al. [130] used implicit solvent molecular dynamics simulations to study the orientation of βCD included in a polyaniline polymer, determining that the head-to-tail (HT) orientation of the included CDs tended to be more attractive than other types of orientations, which in turn affected the mobility and motion confinement of the CD along the wire.
One final remark referred to the simulation of complex structures involving CDs is oriented towards the stabilization of biologically active peptides by multiple capping with CDs. In this regard, Muhammad et al. [131] studied the interaction between human insulin and multiple units of βCD by means of molecular docking and molecular dynamics simulations. They described the formation of an 1:3 insulin-βCD complex, which in turn suggested the possibility of preparing insulin formulations with enhanced biopharmaceutical properties, mostly its degradability and self-aggregation behavior [132]. These scientific possibilities are being actively studied nowadays by using atomistic molecular modeling methods [133,134,135,136].
From the discussed research efforts, it can be seen that plenty of molecular modeling tools, including software, forcefields and workflow designs are available. The main details are summarized in Table 1. As can be seen, the selection of the method, calculation protocol and simulation workflow is highly dependent on the type of system that needs to be studied and the kind of hypothesis under study. Clearly, the utilization of theoretical modeling tools are highly complementary with experimental techniques towards the possibility of describing CDs systems at an atomistic level.
Conclusions
The potential of CD as nanodevices able to form inclusion complexes has been studied for a long time by experimental and theoretical approaches. However, in recent years, both computational methods and computing power have evolved enormously. In particular, performing computations on GPU cards has further increased the technical possibility of managing larger systems and achieving longer simulation times. Consequently, as presented in this review, CDs are now being exhaustively modeled as part of sophisticated materials with the aim of exhibiting specifically designed properties. Also, elaborate simulation protocols and state of the art forcefields have proven to be able to adequately parameterize CDs as part of simple and complex systems. In overall, molecular modeling techniques are now able exhaustively describe (and predict) not only the inclusion mode of guest molecules to CDs, but also its conformational evolution under diverse physicochemical environments, such as ionic strength, solvent polarities, membrane interfaces, pH, to mention some examples. The elucidation at an atomistic (and quantitative) detail of these properties forecasts a new era in the research involving CDs as pharmaceutical and industrial scaffolds.
This new horizon in the field of in silico studies will definitely continue to expand in the near future, and will have a great impact on the capability to obtain new materials with applicability in diverse fields, including drug delivery, nanosensors, and transducers, among others. A remarkable example of the research possibilities that molecular modeling methods offer for the simulation of CDs materials is molecular motors. Surely, in the years to come, this area of frontier research, which was awarded the Nobel Prize in Chemistry in 2016, will be a matter of amazing discoveries. In summary, it is clear that CDs have now gained significant attention as nanodevices with technological application in this research era driven by the evolving power of computational chemistry and the state of the art molecular modeling protocols in particular.
References
Crini, G.: Review: a history of cyclodextrins. Chem. Rev. 114(21), 10940–10975 (2014)
Sharma, N., Baldi, A.: Exploring versatile applications of cyclodextrins: an overview. Drug Deliv. 23(3), 729–747 (2016)
Iacovino, R., V Caso, J., Di Donato, C., Malgieri, G., Palmieri, M., Russo, L., Isernia, C.: Cyclodextrins as complexing agents: preparation and applications. Curr. Org. Chem. 21(2), 162–176 (2017)
Adeoye, O., Cabral-Marques, H.: Cyclodextrin nanosystems in oral drug delivery: a mini review. Int. J. Pharmaceut. 531(2), 521–531 (2017)
Barbour, L.: Experimental and computational methods in supramolecular chemistry. In: Gokel, G. W., Barbour, L. (eds.) Comprehensive Supramolecular Chemistry II, vol. 2. Elsevier, United Kingdom (2017)
Atwood, J.L., Lehn, J.M.: Comprehensive Supramolecular Chemistry: Cyclodextrins. Pergamon, Oxford (1996)
Jensen, J.H.: Molecular Modeling Basics. CRC Press, Boca Raton (2010)
Schlick, T.: Molecular Modeling and Simulation: An Interdisciplinary Guide. Springer, New York (2013)
Castro, E., Barbiric, D.: Molecular modeling and cyclodextrins: a relationship strengthened by complexes. Curr. Org. Chem. 10(7), 715–729 (2006)
Davies, J.: Spectroscopic and computational studies of supramolecular systems, vol. 4. Springer, London (2013)
Lipkowitz, K.B.: Applications of computational chemistry to the study of cyclodextrins. Chem. Rev. 98(5), 1829–1873 (1998)
Zhao, Q., Zhang, W., Wang, R., Wang, Y., Ouyang, D.: Research advances in molecular modeling in cyclodextrins. Curr. Pharm. Design. 23(3), 522–531 (2017)
Snir, M.: The future of supercomputing. In: Proceedings of the 2028th ACM International Conference on Supercomputing 14, pp. 261–262, ACM
Xie, X., Fang, X., Hu, S., Wu, D.: Evolution of supercomputers. Front. Comput. Sci. China 4(4), 428–436 (2010)
Maximova, T., Moffatt, R., Ma, B., Nussinov, R., Shehu, A.: Principles and overview of sampling methods for modeling macromolecular structure and dynamics. PLoS Comput. Biol. 12(4), e1004619 (2016)
Lameira, J., Kupchencko, I., Warshel, A.: Enhancing paradynamics for QM/MM sampling of enzymatic reactions. J. Phys. Chem. B 120(9), 2155–2164 (2016)
Cai, Y., See, S.: GPU Computing and Applications. Springer, Singapore (2015)
Ren, B., Zhang, M., Gao, H., Zheng, J., Jia, L.: Atomic elucidation of the cyclodextrin effects on DDT solubility and biodegradation. Phys. Chem. Chem. Phys. 18(26), 17380–17388 (2016)
Cerón-Carrasco, J.P., den-Haan, H., Peña-García, J., Contreras-García, J., Pérez-Sánchez, H.: Exploiting the cyclodextrins ability for antioxidants encapsulation: a computational approach to carnosol and carnosic acid embedding. Comput. Theor. Chem. 1077, 65–73 (2016)
Pagadala, N.S., Syed, K., Tuszynski, J.: Software for molecular docking: a review. Biophys. Rev. 9(2), 91–102 (2017)
Suárez, D., Díaz, N.: Conformational and entropy analyses of extended molecular dynamics simulations of α-, β-and γ-cyclodextrins and of the β-cyclodextrin/nabumetone complex. Phys. Chem. Chem. Phys. 19(2), 1431–1440 (2017)
Salomon-Ferrer, R., Götz, A.W., Poole, D., Le Grand, S., Walker, R.C.: Routine microsecond molecular dynamics simulations with AMBER on GPUs. 2. Explicit solvent particle mesh ewald. J. Chem. Theory Comput. 9(9), 3878–3888 (2013)
Salomon-Ferrer, R., Case, D.A., Walker, R.C.: An overview of the Amber biomolecular simulation package. Wiley Interdiscip. Rev. 3(2), 198–210 (2013)
Ivanov, P.: Further studies on the conformations of large-ring cyclodextrins. Bulg. Chem. Commun. 46(A), 238–245 (2014)
Assaf, K.I., Gabel, D., Zimmermann, W., Nau, W.M.: High-affinity host-guest chemistry of large-ring cyclodextrins. Org. Biomol. Chem. 14(32), 7702–7706 (2016)
Ellouze, F., Ben Amar, N., Deratani, A.: Large ring cyclodextrins: synthesis, purification and applications. C. R. Chim. 14(10), 967–971 (2011)
Ivanov, P., Atanassov, E., Jaime, C.: Computational study on the conformations of CD38 and inclusion complexes of some lower-size large-ring cyclodextrins. J. Mol. Struct. 1056, 238–245 (2014)
Khuntawee, W., Rungrotmongkol, T., Wolschann, P., Pongsawasdi, P., Kungwan, N., Okumura, H., Hannongbua, S.: Conformation study of ε-cyclodextrin: Replica exchange molecular dynamics simulations. Carbohyd. Polym. 141, 99–105 (2016)
Crich, D.: Modern Synthetic Methods in Carbohydrate Chemistry: From Monosaccharides to Complex Glycoconjugates. Wiley, Weinheim (2013)
Alvarez-Dorta, D., León, E.I., Kennedy, A.R., Martín, A., Pérez-Martín, I., Suárez, E.: Easy access to modified cyclodextrins by an intramolecular radical approach. Angew. Chem. Int. Ed. 54(12), 3674–3678 (2015)
Cheng, G.-J., Zhang, X., Chung, L.W., Xu, L., Wu, Y.-D.: Computational organic chemistry: bridging theory and experiment in establishing the mechanisms of chemical reactions. J. Am. Chem. Soc. 137(5), 1706–1725 (2015)
Shityakov, S., Salmas, R.E., Durdagi, S., Salvador, E., Pápai, K., Yáñez-Gascón, M.J., Pérez-Sánchez, H., Puskás, I., Roewer, N., Förster, C.: Characterization, in vivo evaluation, and molecular modeling of different propofol–cyclodextrin complexes to assess their drug delivery potential at the blood–brain barrier level. J. Chem. Inf. Model. 56(10), 1914–1922 (2016)
Kirschner, K.N., Yongye, A.B., Tschampel, S.M., González-Outeiriño, J., Daniels, C.R., Foley, B.L., Woods, R.J.: GLYCAM06: a generalizable biomolecular force field. Carbohydr. J. Comput. Chem. 29(4), 622–655 (2008)
Wang, J., Wolf, R.M., Caldwell, J.W., Kollman, P.A., Case, D.A.: Development and testing of a general amber force field. J. Comput. Chem. 25(9), 1157–1174 (2004)
Devasari, N., Dora, C.P., Singh, C., Paidi, S.R., Kumar, V., Sobhia, M.E., Suresh, S.: Inclusion complex of erlotinib with sulfobutyl ether-β-cyclodextrin: preparation, characterization, in silico, in vitro and in vivo evaluation. Carbohyd. Polym. 134, 547–556 (2015)
Shityakov, S., Puskás, I., Pápai, K., Salvador, E., Roewer, N., Förster, C., Broscheit, J.-A.: Sevoflurane-sulfobutylether-β-cyclodextrin complex: preparation, characterization, cellular toxicity, molecular modeling and blood-brain barrier transport studies. Molecules 20(6), 10264–10279 (2015)
Kulkarni, A.D., Belgamwar, V.S.: Inclusion complex of chrysin with sulfobutyl ether-β-cyclodextrin (Captisol®): Preparation, characterization, molecular modelling and in vitro anticancer activity. J. Mol. Struct. 1128, 563–571 (2017)
Yildiz, Z.I., Celebioglu, A., Uyar, T.: Polymer-free electrospun nanofibers from sulfobutyl ether 7-beta-cyclodextrin (SBE 7-β-CD) inclusion complex with sulfisoxazole: fast-dissolving and enhanced water-solubility of sulfisoxazole. Int. J. Pharmaceut. 531(2), 550–558 (2017)
Altarsha, M., Ingrosso, F., Ruiz-López, M.F.: Cavity closure dynamics of peracetylated β-cyclodextrins in supercritical carbon dioxide. J. Phys. Chem. B 116(13), 3982–3990 (2012)
Bayly, C.I., Cieplak, P., Cornell, W.D., Kollman, P.A.: A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges: the RESP model. J. Phys. Chem. 97(40), 10269–10280 (1993)
Cornell, W.D., Cieplak, P., Bayly, C.I., Gould, I.R., Merz, K.M. Jr., Ferguson, D.M., Spellmeyer, D.C., Fox, T., Caldwell, J.W., Kollman, P.A.: A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J. Am. Chem. Soc. 117(19), 5179–5197 (1995)
Kordopati, G.G., Tselios, T.V., Kellici, T., Merzel, F., Mavromoustakos, T., Grdadolnik, S.G., Tsivgoulis, G.M.: A novel synthetic luteinizing hormone-releasing hormone (LHRH) analogue coupled with modified β-cyclodextrin: insight into its intramolecular interactions. Biochim. Biophys. Acta 1850(1), 159–168 (2015)
Wang, R., Zhou, H., Siu, S.W., Gan, Y., Wang, Y., Ouyang, D.: Comparison of three molecular simulation approaches for cyclodextrin-ibuprofen complexation. J. Nanomater. 16(1), 267 (2015)
Sheng Cai, W., Wang, T., Zhe Liu, Y., Liu, P., Chipot, C., Guang Shao, X.: Free energy calculations for cyclodextrin inclusion complexes. Curr. Org. Chem. 15(6), 839–847 (2011)
Sangpheak, W., Khuntawee, W., Wolschann, P., Pongsawasdi, P., Rungrotmongkol, T.: Enhanced stability of a naringenin/2, 6-dimethyl β-cyclodextrin inclusion complex: molecular dynamics and free energy calculations based on MM-and QM-PBSA/GBSA. J. Mol. Graph. Model. 50, 10–15 (2014)
Rutenberg, R., Leitus, G., Fallik, E., Poverenov, E.: Discovery of a non classic host guest complexation mode in a β-cyclodextrin/propionic acid model. Chem. Commun. 52(12), 2565–2568 (2016)
Rahim, M., Madi, F., Nouar, L., Haiahem, S., Fateh, D., Khatmi, D.: β-Cyclodextrin Interaction with Edaravone: Molecular Modeling Study. Proceedings of MEST 2012: Electronic Structure Methods with Applications to Experimental Chemistry, vol. 68, pp. 269–278 (2014)
Onnainty, R.e., Schenfeld, E.M., Quevedo, M.A., Fernández, M.A., Longhi, M.R., Granero, G.E.: Characterization of the hydrochlorothiazide: β-cyclodextrin inclusion complex. Experimental and theoretical methods. J. Phys. Chem. B 117(1), 206–217 (2012)
Oda, M., Kuroda, M.: Molecular dynamics simulations of inclusion complexation of glycyrrhizic acid and cyclodextrins (1: 1) in water. J. Incl. Phenom. Macrocycl. Chem. 85(3–4), 271–279 (2016)
Nociari, M.M., Lehmann, G.L., Bay, A.E.P., Radu, R.A., Jiang, Z., Goicochea, S., Schreiner, R., Warren, J.D., Shan, J., de Beaumais, S.A.: Beta cyclodextrins bind, stabilize, and remove lipofuscin bisretinoids from retinal pigment epithelium. Proc. Natl. Acad. Sci. USA 111(14), E1402–E1408 (2014)
Kogawa, A.C., Zoppi, A., Quevedo, M.A., Raquel, M.: Complexation between darunavir ethanolate and β-cyclodextrin experimental and theoretical studies. World J. Pharm. Pharmaceut. Sci. 298–309 (2014)
Al-Rawashdeh, N.A.F., Al-Sadeh, K.S., Al-Bitar, M.B.: Inclusion complexes of sunscreen agents with β-cyclodextrin: spectroscopic and molecular modeling studies. J. Spectrosc. 1(1), 1–11 (2013)
Mobley, D.L., Gilson, M.K.: Predicting binding free energies: frontiers and benchmarks. Ann. Rev. Biophys. 46, 531–558 (2017)
Abel, R., Wang, L., Mobley, D.L., Friesner, R.A.: A critical review of validation, blind testing, and real-world use of alchemical protein-ligand binding free energy calculations. Curr. Top. Med. Chem. 17(23), 2577–2585 (2017)
Torrie, G.M., Valleau, J.P.: Nonphysical sampling distributions in Monte Carlo free-energy estimation: umbrella sampling. J. Comput. Phys. 23(2), 187–199 (1977)
Wickstrom, L., He, P., Gallicchio, E., Levy, R.M.: Large scale affinity calculations of cyclodextrin host–guest complexes: understanding the role of reorganization in the molecular recognition process. J. Chem. Theory Comput. 9(7), 3136–3150 (2013)
Zhang, H., Yin, C., Yan, H., van der Spoel, D.: Evaluation of generalized born models for large scale affinity prediction of cyclodextrin host–guest complexes. J. Chem. Inform. Model. 56(10), 2080–2092 (2016)
Veselinović, A.M., Veselinović, J.B., Toropov, A.A., Toropova, A.P., Nikolić, G.M.: In silico prediction of the β-cyclodextrin complexation based on Monte Carlo method. Int. J. Pharmaceut. 495(1), 404–409 (2015)
Tan, Z., Xia, J., Zhang, B.W., Levy, R.M.: Locally weighted histogram analysis and stochastic solution for large-scale multi-state free energy estimation. J. Chem. Phys. 144(3), 034107 (2016)
Sugita, M., Hirata, F.: Predicting the binding free energy of the inclusion process of 2-hydroxypropyl-β-cyclodextrin and small molecules by means of the MM/3D-RISM method. J. Phys. Condens. Mater. 28(38), 384002 (2016)
Sancho, M.I., Andujar, S., Porasso, R.D., Enriz, R.D.: Theoretical and experimental study of inclusion complexes of β-cyclodextrins with chalcone and 2′,4′-dihydroxychalcone. J. Phys. Chem. B 120(12), 3000–3011 (2016)
Sahra, K., Dinar, K., Seridi, A., Kadri, M.: Investigation on the inclusion of diclofenac with β-cyclodextrin: a molecular modeling approach. Struct. Chem. 26(1), 61–69 (2015)
Angelova, S., Nikolova, V., Molla, N., Dudev, T.: Factors Governing the host–guest interactions between IIA/IIB group metal cations and α-cyclodextrin: a DFT/CDM study. Inorg. Chem. 56(4), 1981–1987 (2017)
Ateba, B.A., Lissouck, D., Azébazé, A., Ebelle, C.T., Nassi, A., Ngameni, E., Duportail, G., Mbazé, L., Kenfack, C.A.: Characterization of Mammea A/AA in solution and in interaction with β-cyclodextrin: UV–visible spectroscopy, cyclic voltammetry and DFT-TDDFT/MD study. J. Mol. Liq. 213, 294–303 (2016)
Henriksen, N.M., Fenley, A.T., Gilson, M.K.: Computational calorimetry: high-precision calculation of host–guest binding thermodynamics. J. Chem. Theory Comput. 11(9), 4377–4394 (2015)
Cao, R., Wu, S.: In silico properties characterization of water-soluble γ-cyclodextrin bi-capped C 60 complex: free energy and geometrical insights for stability and solubility. Carbohyd. Polym. 124, 188–195 (2015)
Tóth, G., Mohácsi, R., Rácz, Á., Rusu, A., Horváth, P., Szente, L., Béni, S., Noszál, B.: Equilibrium and structural characterization of ofloxacin–cyclodextrin complexation. J. Incl. Phenom. Macrocycl. Chem. 77(1–4), 291–300 (2013)
Shi, M., Zhang, C., Xie, Y., Xu, D.: Stereoselective inclusion mechanism of ketoprofen into β-cyclodextrin: insights from molecular dynamics simulations and free energy calculations. Theor. Chem. Acc. 133(10), 1556 (2014)
Melani, F., Pasquini, B., Caprini, C., Gotti, R., Orlandini, S., Furlanetto, S.: Combination of capillary electrophoresis, molecular modeling and NMR to study the enantioselective complexation of sulpiride with double cyclodextrin systems. J. Pharmaceut. Biomed. 114, 265–271 (2015)
Li, L., Li, X., Luo, Q., You, T.: A comprehensive study of the enantioseparation of chiral drugs by cyclodextrin using capillary electrophoresis combined with theoretical approaches. Talanta 142, 28–34 (2015)
Ghatee, M.H., Sedghamiz, T.: Chiral recognition of propranolol enantiomers by β-cyclodextrin: quantum chemical calculation and molecular dynamics simulation studies. Chem. Phys. 445, 5–13 (2014)
Amharar, Y., Grandeury, A., Sanselme, M., Petit, S., Coquerel, G.r.: A hybrid mechanism in chiral discrimination induced by crystallization of supramolecular compounds. J. Phys. Chem. B 116(20), 6027–6040 (2012)
Alvira, E.: Molecular dynamics study of the influence of solvents on the chiral discrimination of alanine enantiomers by β-cyclodextrin. Tetrahedron Asymmetr. 24(19), 1198–1206 (2013)
Alvira, E.: Influence of molecular stereochemistry on the continuum model for van der waals interaction between β-cyclodextrin and linear molecules. Curr. Phys. Chem. 3(3), 357–365 (2013)
Alvira, E.: Theoretical study of the separation of valine enantiomers by β-cyclodextrin with different solvents: a molecular mechanics and dynamics simulation. Tetrahedron Asymmetr. 26(15), 853–860 (2015)
Abou-Zeid, L.A., Hefnawy, M.: Molecular modeling study of the chiral recognition of celiprolol enantiomers by a β-cyclodextrin. Pharmaceut. Chem. J. 2(3), 16–23 (2015)
Suliman, F.O., Elbashir, A.A.: Enantiodifferentiation of chiral baclofen by β-cyclodextrin using capillary electrophoresis: a molecular modeling approach. J. Mol. Struct. 1019, 43–49 (2012)
Periasamy, R., Kothainayaki, S., Sivakumar, K.: Encapsulation of dicinnamalacetone in β-cyclodextrin: A physicochemical evaluation and molecular modeling approach on 1: 2 inclusion complex. J. Macromol. Sci. A 53(9), 546–556 (2016)
Terekhova, I., Kumeev, R., Alper, G., Chakraborty, S., Pérez-Sánchez, H., Núñez-Delicado, E.: Molecular recognition of aromatic carboxylic acids by hydroxypropyl-γ-cyclodextrin: experimental and theoretical evidence. RSC Adv. 6(55), 49567–49577 (2016)
Zhang, H., Ge, C., van der Spoel, D., Feng, W., Tan, T.: Insight into the structural deformations of beta-cyclodextrin caused by alcohol cosolvents and guest molecules. J. Phys. Chem. B 116(12), 3880–3889 (2012)
Zhang, H., Feng, W., Li, C., Lv, Y., Tan, T.: A model for the shuttle motions of puerarin and daidzin inside the cavity of β-cyclodextrin in aqueous acetic acid: insights from molecular dynamics simulations. J. Mol. Model. 18(1), 221–227 (2012)
Chandrasekaran, S., Sudha, N., Premnath, D., Enoch, I.V.: Binding of a chromen-4-one Schiff’s base with bovine serum albumin: capping with β-cyclodextrin influences the binding. J. Biomol. Struct. Dyn. 33(9), 1945–1956 (2015)
Sameena, Y., Sudha, N., Chandrasekaran, S., Enoch, I.V.: The role of encapsulation by β-cyclodextrin in the interaction of raloxifene with macromolecular targets: a study by spectroscopy and molecular modeling. J. Biol. Phys. 40(4), 347–367 (2014)
Yan, J., Wu, D., Ma, X., Wang, L., Xu, K., Li, H.: Spectral and molecular modeling studies on the influence of β-cyclodextrin and its derivatives on aripiprazole-human serum albumin binding. Carbohyd. Polym. 131, 65–74 (2015)
Tang, P., Tang, B., Wang, Q., Xu, K., Xiong, X., Li, H.: Effect of hydroxypropyl-β-cyclodextrin on the bounding of salazosulfapyridine to human serum albumin. Int. J. Biol. Macromol. 92, 105–115 (2016)
Natesan, S., Sowrirajan, C., Dhanaraj, P., Enoch, I.V.: Capping of silybin with β-cyclodextrin influences its binding with bovine serum albumin: a study by fluorescence spectroscopy and molecular modeling. Bull. Korean Chem. Soc. 35(7), 2114–2122 (2014)
Mansouri, M., Pirouzi, M., Saberi, M.R., Ghaderabad, M., Chamani, J.: Investigation on the interaction between cyclophosphamide and lysozyme in the presence of three different kind of cyclodextrins: determination of the binding mechanism by spectroscopic and molecular modeling techniques. Molecules 18(1), 789–813 (2013)
Figueiras, A., Sarraguça, J.M., Pais, A.A., Carvalho, R.A., Veiga, J.F.: The role of l-arginine in inclusion complexes of omeprazole with cyclodextrins. AAPS PharmSci Tech. 11(1), 233–240 (2010)
Sherje, A.P., Kulkarni, V., Murahari, M., Nayak, U.Y., Bhat, P., Suvarna, V., Dravyakar, B.: Inclusion complexation of etodolac with hydroxypropyl-beta-cyclodextrin and auxiliary agents: Formulation characterization and molecular modeling studies. Mol. Pharmaceut. 14(4), 1231–1242 (2017)
Sapte, S., Pore, Y.: Inclusion complexes of cefuroxime axetil with β-cyclodextrin: physicochemical characterization, molecular modeling and effect of l-arginine on complexation. J. Pharm. Anal. 6(5), 300–306 (2016)
Suvarna, V., Kajwe, A., Murahari, M., Pujar, G.V., Inturi, B.K., Sherje, A.P.: Inclusion complexes of nateglinide with HP–β–CD and l-arginine for solubility and dissolution enhancement: preparation, characterization, and molecular docking study. J. Pharm. Innov. 12(2), 168–181 (2017)
Méndez, S.G., Otero Espinar, F.J., Alvarez, A.L., Longhi, M.R., Quevedo, M.A., Zoppi, A.: Ternary complexation of benzoic acid with β-cyclodextrin and aminoacids. Experimental and theoretical studies. J. Incl. Phenom. Macrocycl. Chem. 85(1–2), 33–48 (2016). https://doi.org/10.1007/s10847-016-0603-6
Jadhav, P., Petkar, B., Pore, Y., Kulkarni, A., Burade, K.: Physicochemical and molecular modeling studies of cefixime–l-arginine–cyclodextrin ternary inclusion compounds. Carbohyd. Polym. 98(2), 1317–1325 (2013)
Barbosa, J.A.A., Zoppi, A., Quevedo, M.A., de Melo, P.N., de Medeiros, A.S.A., Streck, L., de Oliveira, A.R., Fernandes-Pedrosa, M.F., Longhi, M.R., da Silva-Júnior, A.A.: Triethanolamine stabilization of methotrexate-β-cyclodextrin interactions in ternary complexes. Int. J. Mol. Sci. 15(9), 17077–17099 (2014)
Li, L., Zhao, M., Li, W., Wang, Y., Zhang, Z., An, R., Peng, S.: Self-complexation and complexation-controlled target cancer therapy. MedChemComm 3(9), 1059–1061 (2012)
He, H., Chen, S., Zhou, J., Dou, Y., Song, L., Che, L., Zhou, X., Chen, X., Jia, Y., Zhang, J.: Cyclodextrin-derived pH-responsive nanoparticles for delivery of paclitaxel. Biomaterials 34(21), 5344–5358 (2013)
Shi, Q., Zhang, L., Liu, M., Zhang, X., Zhang, X., Xu, X., Chen, S., Li, X., Zhang, J.: Reversion of multidrug resistance by a pH-responsive cyclodextrin-derived nanomedicine in drug resistant cancer cells. Biomaterials 67, 169–182 (2015)
Xiong, Q., Zhang, M., Zhang, Z., Shen, W., Liu, L., Zhang, Q.: Anti-tumor drug delivery system based on cyclodextrin-containing pH-responsive star polymer: in vitro and in vivo evaluation. Int. J. Pharmaceut. 474(1), 232–240 (2014)
Dan, Z., Cao, H., He, X., Zeng, L., Zou, L., Shen, Q., Zhang, Z.: Biological stimuli-responsive cyclodextrin-based host–guest nanosystems for cancer therapy. Int. J. Pharmaceut. 483(1), 63–68 (2015)
Swiech, O., Mieczkowska, A., Chmurski, K., Bilewicz, R.: Intermolecular interactions between doxorubicin and β-cyclodextrin 4-methoxyphenol conjugates. J. Phys. Chem. B 116(6), 1765–1771 (2012)
Hrubý, M., Koňák, Č., Ulbrich, K.: Polymeric micellar pH-sensitive drug delivery system for doxorubicin. J. Control Release 103(1), 137–148 (2005)
Swiech, O., Majdecki, M., Debinski, A., Krzak, A., Stępkowski, T.M., Wójciuk, G., Kruszewski, M., Bilewicz, R.: Competition between self-inclusion and drug binding explains the pH dependence of the cyclodextrin drug carrier–molecular modelling and electrochemistry studies. Nanoscale 8(37), 16733–16742 (2016)
Swiech, O., Dutkiewicz, P., Wójciuk, K., Chmurski, K., Kruszewski, M., Bilewicz, R.: Cyclodextrin derivatives conjugated with aromatic moieties as pH-responsive drug carriers for anthracycline. J. Phys. Chem. B 117(43), 13444–13450 (2013)
De Sousa, F.B., Lima, A.C., Denadai, Â.M., Anconi, C.P., De Almeida, W.B., Novato, W.T., Dos Santos, H.F., Drum, C.L., Langer, R., Sinisterra, R.D.: Superstructure based on β-CD self-assembly induced by a small guest molecule. Phys. Chem. Chem. Phys. 14(6), 1934–1944 (2012)
Goh, G.B., Hulbert, B.S., Zhou, H., Brooks, C.L.: Constant pH molecular dynamics of proteins in explicit solvent with proton tautomerism. Proteins: structure, function, and bioinformatics 82(7), 1319–1331 (2014)
Swails, J.M., York, D.M., Roitberg, A.E.: Constant pH replica exchange molecular dynamics in explicit solvent using discrete protonation states: implementation, testing, and validation. J. Chem. Theory Comput. 10(3), 1341–1352 (2014)
Arthur, E.J., Brooks, C.L.: Efficient implementation of constant pH molecular dynamics on modern graphics processors. J. Comput. Chem. 37(24), 2171–2180 (2016)
Harada, A., Takashima, Y., Yamaguchi, H.: Cyclodextrin-based supramolecular polymers. Chem. Soc. Rev. 38(4), 875–882 (2009)
Zhang, H., Tan, T., Feng, W., Van Der Spoel, D.: Molecular recognition in different environments: β-cyclodextrin dimer formation in organic solvents. J. Phys. Chem. B 116(42), 12684–12693 (2012)
Zhang, H., Tan, T., Hetényi, C., Van Der Spoel, D.: Quantification of solvent contribution to the stability of noncovalent complexes. J. Chem. Theory Comput. 9(10), 4542–4551 (2013)
Zhang, H., Tan, T., Hetényi, C., Lv, Y., Van Der Spoel, D.: Cooperative binding of cyclodextrin dimers to isoflavone analogues elucidated by free energy calculations. J. Phys. Chem. C 118(13), 7163–7173 (2014)
Staelens, N., Leherte, L., Vercauteren, D.P.: Formation and structural, energetic and dynamic properties of cyclodextrin host tubes and included guest molecules. Supramol. Chem. 27(1–2), 90–109 (2015)
Raffaini, G., Ganazzoli, F.: A molecular modeling study of complex formation and self-aggregation behavior of a porphyrin-β-cyclodextrin conjugate. J. Incl. Phenom. Macrocycl Chem. 76(1–2), 213–221 (2013). https://doi.org/10.1007/s10847-012-0193-x
Liu, Y., Chipot, C., Shao, X., Cai, W.: Threading or tumbling? Insight into the self-inclusion mechanism of an altro-α-cyclodextrin derivative. J. Phys. Chem. C 118(33), 19380–19386 (2014)
Wallace, S.J., Kee, T.W., Huang, D.M.: Molecular basis of binding and stability of curcumin in diamide-linked γ-cyclodextrin dimers. J. Phys. Chem. B 117(41), 12375–12382 (2013)
Trotta, F., Zanetti, M., Cavalli, R.: Cyclodextrin-based nanosponges as drug carriers. Beilstein J. Org. Chem. 8(1), 2091–2099 (2012)
Raffaini, G., Ganazzoli, F., Mele, A., Castiglione, F.: A molecular dynamics study of cyclodextrin nanosponge models. J. Incl. Phenom. Macrocycl. Chem. 75(3–4), 263–268 (2013)
Mixcoha, E., Campos-Terán, J., Piñeiro, A.n.: Surface adsorption and bulk aggregation of cyclodextrins by computational molecular dynamics simulations as a function of temperature: α-CD vs β-CD. J. Phys. Chem. B 118(25), 6999–7011 (2014)
Takayanagi, M., Ito, S., Matsumoto, K., Nagaoka, M.: Formation of reactant complex structure for initiation reaction of lactone ring-opening polymerization by cooperation of multiple cyclodextrin. J. Phys. Chem. B 120(29), 7174–7181 (2016)
Loftsson, T., Brewster, M.E.: Pharmaceutical applications of cyclodextrins: effects on drug permeation through biological membranes. J. Pharm. Pharmacol 63(9), 1119–1135 (2011)
Loftsson, T.: Drug permeation through biomembranes: cyclodextrins and the unstirred water layer. Int. J. Pharmaceut. Sci. 67(5), 363–370 (2012)
Khuntawee, W., Wolschann, P., Rungrotmongkol, T., Wong-Ekkabut, J., Hannongbua, S.: Molecular dynamics simulations of the interaction of beta cyclodextrin with a lipid bilayer. J. Chem. Inf. Model. 55(9), 1894–1902 (2015)
Hashidzume, A., Yamaguchi, H., Harada, A.: Cyclodextrin-based molecular machines. In: Credi, A., Silvi, S., Venturi, M. (eds.) Molecular Machines and Motors, pp. 71–110. Springer, Dordrecht (2014)
Wenz, G., Han, B.-H., Müller, A.: Cyclodextrin rotaxanes and polyrotaxanes. Chem. Rev. 106(3), 782–817 (2006)
Singharoy, A., Chipot, C.: Methodology for the simulation of molecular motors at different scales. J. Phys. Chem. B 121(15), 3502–3514 (2016)
Liu, P., Chipot, C., Cai, W., Shao, X.: Unveiling the underlying mechanism for compression and decompression strokes of a molecular engine. J. Phys. Chem. C 118(23), 12562–12567 (2014)
Bruns, C.J., Stoddart, J.F.: The Nature of the Mechanical Bond: From Molecules to Machines. Wiley, Hoboken (2016)
Liu, P., Chipot, C., Shao, X., Cai, W.: Solvent-controlled shuttling in a molecular switch. J. Phys. Chem. C 116(7), 4471–4476 (2012)
Low, P.J., Marqués-González, S.: Molecular wires: an overview of the building blocks of molecular electronics. In: Kiguchi, M. (ed.) Single-Molecule Electronics, pp. 87–116. Springer, Dordrecht (2016)
Tallury, S.S., Smyth, M.B., Cakmak, E., Pasquinelli, M.A.: Molecular dynamics simulations of interactions between polyanilines in their inclusion complexes with β-cyclodextrins. J. Phys. Chem. B 116(7), 2023–2030 (2012)
Muhammad, E.F., Adnan, R., Latif, M.A.M., Rahman, M.B.A.: Theoretical investigation on insulin dimer-β-cyclodextrin interactions using docking and molecular dynamics simulation. J. Incl. Phenom. Macrocycl. Chem. 84(1–2), 1–10 (2016)
Krauland, A.H., Alonso, M.J.: Chitosan/cyclodextrin nanoparticles as macromolecular drug delivery system. Int. J. Pharmaceut. 340(1–2), 134–142 (2007)
Berhanu, W.M., Masunov, A.E.: Controlling the aggregation and rate of release in order to improve insulin formulation: Molecular dynamics study of full-length insulin amyloid oligomer models. J. Mol. Model. 18(3), 1129–1142 (2012)
Muhammad, E.F.: Docking And Molecular Dynamics Simulation Studies Of Insulin-Β-Cyclodextrin Interactions. Universiti Sains Malaysia, Gelugor (2016)
Panchal, A.: Insulin drug delivery systems: a review. Int. J. Res. Pharmaceut. Sci. 2(4), 484–492 (2016)
Gonzalez-Gaitano, G., Ramon Isasi, J., Velaz, I., Zornoza, A.: Drug carrier systems based on cyclodextrin supramolecular assemblies and polymers: present and perspectives. Curr. Pharm. Design 23(3), 411–432 (2017)
Acknowledgements
The authors gratefully acknowledge financial support from the Secretaria de Ciencia y Técnica of the Universidad Nacional de Córdoba (SECYT-UNC), the Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET), and the Agencia Nacional de Promoción Científica y Técnica (ANPCyT), Argentina.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Quevedo, M.A., Zoppi, A. Current trends in molecular modeling methods applied to the study of cyclodextrin complexes. J Incl Phenom Macrocycl Chem 90, 1–14 (2018). https://doi.org/10.1007/s10847-017-0763-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10847-017-0763-z