Abstract
Next-generation massive parallel sequencing and bioinformatic techniques were applied to observe the differences in the transcriptome of Litopenaeus vannamei juveniles that were fed with a control diet (T1) or the same diet plus a live Ulva clathrata supplement (T2). The average size of the libraries was of 271 bp for T1 and 275 bp for T2. A total of 7,706,527 raw reads were obtained with a total of 6,591,856 reads after filtering through Q30, from which 3,855,181 corresponded to T1 and 2,736,675 corresponded to T2. Assembled contigs of each library were annotated using BLASTx from NCBI [obtaining 15,861 (42.77 %) unigenes for T1 and 14,246 (45.06 %) for T2]. In addition, the Kyoto Encyclopedia of Genes and Genomes database (KEGG) as well as the Gene Ontology database (GO) were employed. The differential expression analysis generated a total of 396 transcripts with different expression levels between the two treatments, and these were classified into 4 groups in accordance to their possible function; among others, it identified a percentage of transcripts associated with: immune response (12 %), lipid metabolism (34 %), oxidation-reduction processes (34 %), and stress responses (20 %). The study shows a systematic description of the transcriptome analysis of L. vannamei and provides valuable genetic information required for studying the molecular mechanisms operating when shrimp are fed with macroalgae U. clathrata.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Pacific white shrimp, Litopenaeus vannamei, is a crustacean species cultured all around the world. It has a considerable commercial value, and its production through farming has surpassed fisheries production, reaching great economic and social importance, as it represents a source of incomes and jobs. For this reason, research on new functional ingredients capable of improving the organism’s production has increased; some of these ingredients can play a vital role in the organism’s predisposition to fulfill its dietary needs. Several authors have studied different marine algae species as meals, algal extracts, or live algae biomass as supplements in feeds and feeding regimes for shrimp, finding that these ingredients contain bioactive substances such as sulphated polysaccharides associated with immune system stimulation, antiviral, and antioxidant activities (Chotigeat et al. 2004; Rocha de Souza et al. 2007; Wijesekara et al. 2011; Immanuel et al. 2012), polyphenols that are well-known for their antioxidant activity (Bozin et al. 2008; Matanjun et al. 2008; Kumar et al. 2011; Shanab et al. 2011; Simić and Ranković 2012; Trigui et al. 2013; Heffernan et al. 2015) as well as some other lipiphilic compounds such as chlorophyll, carotenoids, and fatty acids associated with antibacterial and antioxidant activities (Simić and Ranković 2012; Awang et al. 2014).
Ulva clathrata is a macroalga containing vitamins, minerals, carotenoids, polysaccharides, antioxidants and which has a higher protein content compared to other algal species (Cruz-Suárez et al. 2009, 2013). It has been demonstrated that inclusion of U. clathrata in the diet of L. vannamei improves growth, food conversion ratio, and pigmentation and modifies fatty acid and sterol content in muscle (Cruz-Suárez et al. 2009, 2010, 2013; Peña-Rodríguez et al. 2010, 2011); nevertheless, there are no studies regarding the effects of the dietary inclusion of this alga at a molecular level. The development of new molecular biological techniques, such as massive parallel sequencing methods, have allowed mapping and transcriptome quantification, enabling us to improve our comprehension about the complexity of genomes even in eukaryotic organisms where there is incomplete genome sequencing (Ekblom and Galindo 2011; Guo et al. 2013). With regard to insects, there are studies demonstrating the feasibility of this kind of technology. Such studies have had the objective to understand the organism’s response related to nutritional changes and diet quality, in an attempt to improve insect diets (Yocum et al. 2006; Alaux et al. 2011; Zou et al. 2013). In crustaceans, massive parallel sequencing has been employed to identify genes related to immune response, after inoculation with a virus (Li et al. 2013; Zeng et al. 2013; Chen et al. 2013a; Sookruksawong et al. 2013); however, this new tool has not been used in nutritional studies. The present study has as an objective to use next-generation sequencing and bioinformatic techniques in order to obtain information about the L. vannamei transcriptome when these shrimp are fed a live U. clathrata as a dietary supplement, in order to find an explanation at the molecular level of the benefits of eating seaweed, i.e., genes that change their expression level and metabolic pathways in which these genes are involved. This study also seeks to describe possible molecular mechanisms involved in the improvement of nutritional parameters, for enriching cDNA databases and identifying the genes involved in immune defense and lipid metabolism.
Materials and methods
Experimental diets and animals
The experiment was conducted at the Programa Maricultura facilities (FCB, UANL, Monterrey, Mexico) using specific pathogen-free (SPF) shrimp L. vannamei (certificate: DGSA-DSAP-CSAMO-080/2011) obtained from a commercial farm located in Veracruz, Mexico (Langostinos y Camarones de Oriente, S.A. de C.V.). The shrimp were acclimated for 2 weeks prior to the experiment in a recirculating sea water system (35 ± 2 ppt salinity, pH 8.2 ± 0.1, 96.7 ± 1.5 % oxygen) at a temperature of 29 ± 2 °C.
The experiment was conducted using 6 replicated aquaria per treatment. Ten shrimp were allocated to each aquarium (average initial weight 3 ± 0.01 g). The control group (T1) was fed twice a day ad libitum with a control diet (0900 and 1600 h) formulated with 35 % protein and 8.0 % lipids. The experimental group (T2) was fed twice a day (0900 and 1600 h) with the same diet and ad libitum on live U. clathrata. Daily, 8 g of living alga was placed in Maya packaging and placed at the bottom of each aquarium at 0900 h and left for 24 h to allow the animals always to have it at their disposal. The next day (before placing the algae again), the remains of algae were weighed and the amount consumed calculated. Both treatment groups were fed over a 20-day period. Feed residue and feces were siphonated out and moults were removed on a daily basis.
Sample collection
Four shrimp per aquarium were sampled after 20 days of feeding. The shrimp that were selected were apparently healthy and in an inter-moult phase. The moulting stage was determined using the technique of Robertson et al. (1987). Subsequently, the hepatopancreas of each shrimp was extracted and individually placed in an Eppendorf tube containing 1 mL RNAlater (Ambion Life Technologies). Approximately 20 mg of each hepatopancreas was sampled in order to perform total extraction of RNA, and the remaining tissue was stored at −80 °C.
RNA extraction and purification
Total RNA from 40 hepatopancreas (20 from each treatment) were extracted using the PureZOL RNA Isolation Reagent (Bio-Rad), in accordance with the manufacturer’s instructions. Total RNA quality and integrity were assessed using an Experion RNA StdSens Analysis Kit (Bio-Rad), in accordance with the manufacturer’s instructions at Experion Automated Electrophoresis Systems. Eleven samples representing the best quality RNA were selected from each treatment and quantified using the Quant-iT RiboGreen RNA (Invirtogen) assay kit, with the purpose of taking equimolar quantities of each sample and forming a pool for each treatment. The mRNA fraction was purified using RNA Purification Beads in accordance with the TruSeq RNA Sample Preparation v2 Guide (Illumina Part #15026495 Rev. C May 2012) (Fig. 1).
Transcriptome sequencing and bioinformatic analysis
Transcriptome library sequencing was performed using a MiSeq Reagent Kit v2 Illumina next generation sequencer with 300 cycles in accordance with the manufacturer’s instructions. After filtering low-quality reads using default parameters of the Trimmomatic program (Bolger et al. 2014), the remaining high-quality reads were assembled using the default parameters of the Trinity program developed at the Broad Institute and the Hebrew University of Jerusalem (Grabherr et al. 2011). This software is specifically designed for assembling de novo transcriptomes. All transcriptome data were subsequently registered in DRYAD (accession number doi:10.5061/dryad.j970q). The assembled contigs of each treatment were annotated using the non redundant protein (Nr) NCBI database and InterProScan database. Data were further analyzed using the BLAST2GO software suite (http://www.BLAST2go.org) (Blüthgen et al. 2005; Götz et al. 2008) to predict functions of individual ESTs assigning Gene Ontology terms (Ye et al. 2006). Data were also analyzed with the KAAS- KEEG Automatic Annotation Server (Kanehisa and Goto 2000; Moriya et al. 2007; Kanehisa et al. 2008).
Identification of differentially expressed genes
Sequence reads were used to generate a single Trinity hybrid assembly (Grabherr et al. 2013) based on combining all reads across both treatments (T1 and T2). Transcript levels were measured as FPKM (Fragments Per Kilobase of exon model per Million mapped reads) for differential gene expression analyses (http://jura.wi.mit.edu/bio/education/hot_topics/RNAseq/RNA_Seq.pdf); these data were normalized using the TMM method (Robinson and Oshlack 2010). Statistical comparisons between the different transcriptomes were conducted using EdgeR Bioconductor software (Robinson et al. 2010, 2011) in accordance to the Trinity instructions. Transcripts were considered differentially expressed in a given library when the p-value was less than 0.05.
Differential expression validation by quantitative PCR
Quantitative PCR (qPCR), to validate differential expression, used the same total RNA pools of each treatment that were used for MiSeq-Illumina sequencing. First strand cDNA was synthesized from 2 μg of total RNA using a High Capacity cDNA Reverse Transcription (Invitrogen, USA) kit, in accordance with the manufacturer’s protocol. Primers were designed using Primer3Plus software (http://biotools.umassmed.edu/cgi-bin/primer3plus/primer3plus.cgi). Seven candidate transcripts related to immune response and lipid metabolism were selected for validation by qPCR: sam domain and hd domain-containing protein 1 [comp8719_c0_seq1], interleukin enhancer-binding factor 2 homolog [comp8600_c0_seq3], solute carrier family 7 (cationic amino acid transporter) [comp10896_c0_seq3], platelet-activating factor acetylhydrolase [comp10353_c0_seq12], serine palmitoyltransferase 1 [comp10732_c0_seq1], elongation factor-1 alpha [comp7540_c0_seq1], and fatty acid synthase [comp45386_c0_seq1] (Table 4). The shrimp beta-actin gene was used as a constitutively expressed control (Cheng et al. 2005); (Chen et al. 2014). Quantitative PCR and data analysis were performed using a LightCycler 480 System (Roche). The qPCR reaction mixture (10 μL) contained 2× Power SYBR Green PCR Master mix, 0.5 μL of each forward and reverse primer, and 2.5 μL of template cDNA. Amplifications were performed under the following conditions: 1 cycle of 95 °C for 2 min; 35 cycles at 95 °C for 1 min, 55 °C for 1 min, 72 °C for 1 min; and a final extension at 72 °C for 5 min. Receptor mRNA relative expression was measured using the method 2–ΔΔCt (Livak and Schmittgen 2001). The relative expression of the seven target genes in treated (T2 U. clathrata) vs control shrimp (T1) was normalized to an endogenous reference gene (beta-actin) and expressed as the Normalized Target Gene Expression Level = 2–ΔΔCt, where ΔΔCt is defined as the difference between ΔCt of treated shrimp and ΔCt of control shrimp, and the ΔCt is defined as the difference between Ct target gene and Ct of reference gene (Brink et al. 2000). Serial dilutions from one of the samples were made to validate the efficiency of the qPCR assay and were tested with the oligonucleotides. The results were analyzed in a calibration curve considering that the regression value must be close to 1 and the slope close to −3.32 (Wong and Medrano 2005). Each qPCR reaction was carried out in triplicate.
Results and discussion
Sequencing and de novo assembly
As described in the aforementioned methods, libraries from cDNA synthesized from mRNAs pools of each treatment (T1 and T2) were sequenced using an Illumina MiSeq, resulting in 7,706,527 of total raw reads. After evaluating the reads using the Trimmomatric trimming program and removing adaptors, a total of 6,591,856 cleaned reads were obtained. The GC content, ≥Q30, and unknown bases (N) were of 48.00, 78.90, and 0.00 %, respectively (Table 1). In the de novo assembly, 37,078 transcripts were generated for (T1) with a 666-bp average length, and a N50 value of 1007 bp; for (T2), 31,615 transcripts were generated with a 592-bp average length, and a N50 value of 800 bp (Table 2). This last result shows us that our transcriptome database has acceptable coverage and the data greatly enrich existing information about the L. vannamei transcriptome.
Functional annotation
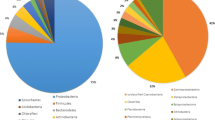
Functional annotation of transcript sequences was performed using BLASTx, along with the non-redundant protein NR sequence database of the NCBI (E-value threshold of 1e-5) (Guo et al. 2013) and analyzed using the Blast2GO program. A total of 15,861 transcripts for T1 (42.77 %) and 14,246 for T2 (45.06 %) were annotated in accordance to their homolog function (Table 3). Results on the distribution showing the best matches for each sequence are given in Fig. 2. Annotated unigenes showed matches with unigenes of Daphnia pulex (12 %), Tribolium castaneum (6 %), Pediculus humanus (5 %), Branchiostoma floridae (3 %), Crassostrea gigas (3 %), Ixodes scapularis (3 %), Nasonia vitripennis (3 %), Capitella teleta (3 %), followed by Litopenaeus vannamei (3 %). Our results agree with those of Guo et al. (2013) and Xiaohan Chen et al. (2013b), indicating that mostly all or the majority of unigenes are similar to those of Daphnia pulex, Tribolium castaneum, and Pediculus humanus; this might be attributed to the close phylogenetic relationship to these species, for which a great deal of genetic information is available over all of the annoted genes presented. On the other hand, the remaining annotated unigenes show certain similarity with other species, a fact which might be attributed to the relatively limited amount of the L. vannamei protein sequences available in the database.
Gene ontology (GO) analysis was performed using the Blast2GO program (Blüthgen et al. 2005; Götz et al. 2008) to determine the functional categories (Ye et al. 2006) represented by the L. vannamei transcriptome. The transcriptome comparison between T1 and T2 showed high similarity between transcript distributions in the three GO categories. We were able to observe that 18,682 (50.40 %) and 15,929 (50.30 %) of transcripts corresponded to biological processes, 14,466 (39.03 %) and 12,335 (39.01 %) to cellular components, and 7980 (21.25 %) and 6804 (21.52 %) to molecular functions classified in 49 subcategories (Fig. 3). With regard to the biological processes category, the two subcategories that were most represented were: organic substance metabolic processes and primary metabolic processes. Concerning the cellular components, the most represented subcategories were extracellular space components and cellular parts. Lastly, for the molecular function category, the most represented subcategories were binding function and catalytic activity. These subcategories were the most represented for both treatments in general, the functional transcript distributions were extremely similar between the two experimental groups of shrimp, and the number of transcripts expressed in each GO category was not significantly affected by U. clathrata consumption.
Subsequently, sequences were annotated using the KAAS-KEEG Automatic Annotation Server (Kanehisa and Goto 2000; Moriya et al. 2007; Kanehisa et al. 2008), with the purpose of determining the described metabolic pathways and homologue functions. Using KAAS-KEEG annotation, we characterized 9649 (26.02 %) and 8655 (23.37 %) transcripts, respectively in each treatment (T1 and T2).
Identification of differently expressed transcripts
A total of 396 differently expressed transcripts were identified between the two treatments (high expression and low expression). For functional annotation, all the differentially expressed transcripts were annotated in the following gene databases: BLASTx, InterPro, GO, and KAAS-KEEG Automatic Annotation Server (Kanehisa and Goto 2000; Moriya et al. 2007; Kanehisa et al. 2008) and using the Blast2GO program (E-value threshold of 1e-5) (Guo et al. 2013). Of this total, 347 (87.4 %) were annotated in BLASTx, 284 (71.7 %) by GO, and 319 (80.5 %) by InterPro. An average 87 % of differentially expressed transcripts were annotated with the different databases; nevertheless, 13 % had low sequence homology with the sequences that were already known through public databases. These data matched with the aforementioned data presented by Zeng et al. (2013) and by Chen et al. (2013b). In the same way, they obtained 80 % similarity with currently reported transcripts. In contrast to these authors who obtained about 1300 differentially expressed transcripts, we observed only 396 differentially expressed transcripts. This may be due to the fact that the effector that was used in our study for inducing differential expression was the diet supplementation with Ulva. It is important to take into account that the animals were always under the same conditions (stress, health, nutritional, etc.); meanwhile, in the aforementioned studies, the effector which produced changes in gene expression was viral infection (WSSV in the article of Chen et al. 2013b and TSV in Zeng et al. 2013), which is obviously a more drastic effector as it causes great differences between healthy and diseased animals.
The relative expressions of seven candidate transcripts involved in immune response and lipid metabolism were determined in hepatopancreas by qPCR. Results showed that all these transcripts were significantly up-regulated in response to the consumption of U. clathrata as a dietary supplement (T2). The relative expression of T2 (Ulva) to T1 (control) was: 1.3-fold change of sam domain and hd domain-containing protein 1, 0.42-fold change of interleukin enhancer-binding factor 2 homolog, 0.60-fold change of solute carrier family 7 (cationic amino acid transporter), 0.47-fold change platelet-activating factor acetylhydrolase, 0.54-fold change of serine palmitoyltransferase 1, 0.49-fold change of elongation factor-1 alpha, and 22.7-fold change of fatty acid synthase. These results also further confirmed the reliability of RNA-seq and the accuracy of the Trinity assembly. It has been demonstrated that massive parallel sequencing technologies are a competent method that can be employed for detecting differentially expressed transcripts (Marioni et al. 2008).
KAAS- KEGG analysis of the differentially expressed transcripts
All differentially expressed transcripts were assigned to reference canonic pathways in the KAAS-KEEG Automatic Annotation Server with the purpose of establishing modified metabolic pathways. The top 15 pathways are presented in Fig. 4. The distribution of high-expression or low-expression transcripts between treatments was different: T1 presented the highest quantity of high-expression transcripts in the “Metabolic pathways,” “Biosynthesis of secondary metabolites,” “Microbial metabolism in diverse environments,” “Carbon metabolism,” “Glycerolipid metabolism,” “Glycerophospholipids metabolism,” “Ether lipid metabolism,” “Amino sugar and nucleotide sugar metabolism,” “Pathways in cancer,” “MicroRNAs in cancer,” “Influenza A,” and “Pancreatic secretion” pathways; meanwhile, in T2, the high-expression transcripts were found in the “Spliceosome,” “RNA transport,” and “Lysosome” pathways; this indicates some changes in the physiological state of shrimp which consumed U. clathrata (Table 4).
Candidate genes involved in the lipid metabolism and the immune response of L. vannamei
Concerning the metabolic pathways associated with immune response, the following pathways were affected: toll-like receptor signaling pathway, apoptosis, MAPK signaling pathway, melanogenesis, drug metabolism - cytochrome P450, natural killer cell-mediated cytotoxicity, and TNF signaling pathways (Table 5) (Cohen 1993; Dobrovolskaia et al. 2003; Andrade et al. 2004; Yang et al. 2008; Barakat et al. 2009). A few studies have reported that dietary supplementation with algae meals or extracts might improve the immune response, resistance, and survival of shrimp challenged with bacteria or viruses due to the presence of certain immune-stimulating compounds such as fucoidan, alginates, laminarins, and carragenans (Selvin et al. 2004; Cheng et al. 2008; Cruz-Suárez et al. 2009). At a molecular level, it has been demonstrated that the addition of sulfated polysaccharides of Ulva (Enteromorpha) prolifera stimulates the production of macrophages and increases cytokine expression (Kim et al. 2011). Other studies have shown that polysaccharides and extracts from marine algae might have a certain influence on the immune response genes (Leiro et al. 2007; Reilly et al. 2008; Yoon et al. 2009; Bahar et al. 2012; Kandasamy et al. 2012).
The following modified metabolic pathways were associated with lipid metabolism: ether lipid metabolism, fat digestion and absorption, glycerophospholipid metabolism, fatty acid degradation, alpha-linolenic acid metabolism, fatty acid biosynthesis, linoleic acid metabolism, biosynthesis of unsaturated fatty acids, or arachidonic acid metabolism (Table 5). These results confirm the effect at a molecular level on the lipid metabolism and agree with previously reported studies (Maeda et al. 2005; Seo et al. 2013). These studies also demonstrated that U. clathrata had significant effects on lipid composition of different fishes and terrestrial animal species (Nakagawa et al. 1987; Nakagawa 1997; Wong et al. 1999; Pengzhan et al. 2003; Ortiz et al. 2006; Carillo et al. 2008; Ergün et al. 2008). In shrimp, U. clathrata promotes the production of certain 18 carbon polyunsaturated fatty acids (PUFAs) (Kumari et al. 2010), as well as favoring an increase in the content of 18:0, 18:3n3, 20:5n3 (Ortiz et al. 2006; Carillo et al. 2008; Peña-Rodríguez et al. 2010). In contrast, it has been demonstrated that fucoidan treatment (polysaccharide extracted from brown seaweed) affects adipogenesis in 3T3-L1 cell lines (Kim et al. 2010) as it suppresses the adipocites differentiation and inhibits the expression of adipogenic transcription factors α (C / EBPAα), γ (PPARγ) and AP2. In the same way, the inclusion of Ulva linza and Lessonia trabeculata in diets for rats significantly reduces the content of intra abdominal fat, cholesterol, and triglyceride serum levels in healthy animals, as well as in animals showing signs of metabolic syndrome, and at the same time, there is a reduction in catalase gene expression in animals with metabolic syndrome (Ramirez-Higuera et al. 2014). In our study, it was possible to demonstrate these modifications (at genomic level) in the lipid metabolism of shrimp fed with Ulva.
Conclusion
In this study, massive parallel sequencing of RNA has been employed with the intention of examining the L. vannamei transcriptome in response to the consumption of U. clathrata as a dietary supplement. This is considered a first work at the molecular level regarding nutrigenomics related to the consumption of marine algae by L. vannamei. Differentially expressed genes were detected between both nutritional conditions, and the transcriptional reactions mainly described the immune response pathway, lipid metabolism, stress and stimulus response, and oxidation–reduction processes. The identified genes and/or the described metabolic pathways suggest that Ulva consumption modified the molecular mechanisms involved in shrimp immune system, as well as in the deposition and movement of different lipid classes in the tissues of shrimp.
References
Alaux C, Dantec C, Parrinello H, Le Conte Y (2011) Nutrigenomics in honey bees : digital gene expression analysis of pollen’ s nutritive effects on healthy and varroa-parasitized bees. BMC Genomics 12:496
Andrade AA, Silva NG, Pereira ACTC et al (2004) The vaccinia virus-stimulated mitogen-activated protein kinase (MAPK) pathway is required for virus multiplication. 446:437–446
Awang AN, Ng JL, Matanjun P, Sulaiman MR, Tan TS, Ooi YBH (2014) Anti-obesity property of the brown seaweed, Sargassum polycystum using an in vivo animal model. J Appl Phycol 26:1043–1048
Bahar B, O’Doherty JV, Hayes M, Sweeney T (2012) Extracts of brown seaweeds can attenuate the bacterial lipopolysaccharide-induced pro-inflammatory response in the porcine colon ex vivo. J Anim Sci 90:46–48
Barakat A, DiLoreto DS, Zhang Y, Smith C, Baier K, Powell WA, Wheeler N, Sederoff R, Carlson JE (2009) Comparison of the transcriptomes of American chestnut (Castanea dentata) and Chinese chestnut (Castanea mollissima) in response to the chestnut blight infection. BMC Plant Biol 9:51
Blüthgen N, Brand K, Cajavec B, Swat M, Herzel H, Beule D (2005) Biological profiling of gene groups utilizing gene ontology. Genome Inform 16:106–115
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 1–7. doi:10.1093/bioinformatics/btu170
Bozin B, Mimica-Dukic N, Samojlik I, Goran A, Igic R (2008) Phenolics as antioxidants in garlic (Allium sativum L., Alliaceae). Food Chem 111:925–929
Brink N, Szamel M, Young AR, Wittern KP, Bergemann J (2000) Comparative quantification of IL-1beta, IL-10, IL-10r, TNFalpha and IL-7 mRNA levels in UV-irradiated human skin in vivo. Inflamm Res 49:290–296
Carillo S, López E, Casas M, Avila E, Castillo RM, Carranco ME, Calvo C, Perez-Gil F (2008) Potential use of seaweeds in the laying hen ration to improve the quality of n-3 fatty acid enriched eggs. J Appl Phycol 20:271–278
Chen X, Lin H-Z, Jiang S-G, Wu KC, Liu YJ, Tian LX, Zhang YQ, Niu J (2013a) Dietary supplementation of honeysuckle improves the growth, survival and immunity of Penaeus monodon. Fish Shellfish Immunol 35:161–169
Chen X, Zeng D, Chen X, Xie D, Zhao Y, Yang C, Li Y, Ma N, Li M, Yang Q, Liao Z, Wang (2013b) Transcriptome analysis of Litopenaeus vannamei in response to white spot syndrome virus infection. PLoS ONE 8:e73218
Chen K, Li E, Gan L, Wang X, Xu C, Lin H, Qin JG, Chen L (2014) Growth and lipid metabolism of the Pacific white shrimp Litopenaeus vannamei at different salinities. J Shellfish Res 33:825–832
Cheng W, Liu C-H, Tsai C-H, Chen J-C (2005) Molecular cloning and characterisation of a pattern recognition molecule, lipopolysaccharide- and β-1,3-glucan binding protein (LGBP) from the white shrimp Litopenaeus vannamei. Fish Shellfish Immunol 18:297–310
Cheng A-C, Chen Y-Y, Chen J-C (2008) Dietary administration of sodium alginate and k-carrageenan enhances the innate immune response of brown-marbled grouper Epinephelus fuscoguttatus and its resistance against Vibrio alginolyticus. Vet Immunol Immunopathol 121:206–215
Chotigeat W, Tongsupa S, Supamataya K, Phongdara A (2004) Effect of fucoidan on disease resistance of black tiger shrimp. Aquaculture 233:23–30
Cohen JJ (1993) Apoptosis: the physiologic pathway of cell death. Hosp Pract (Off Ed) 28:35–43
Cruz-Suárez LE, Tapia-Salazar M, Nieto-López MG, Guajardo-Barbosa C, Ricque-Marie D (2009) Comparison of Ulva clathrata and the kelps Macrocystis pyrifera and Ascophyllum nodosum as ingredients in shrimp feeds. Aquac Nutr 15:421–430
Cruz-Suárez LE, León A, Peña-Rodríguez A, Rodríguez-Peña G, Moll B, Ricque-Marie D (2010) Shrimp/Ulva co-culture: a sustainable alternative to diminish the need for artificial feed and improve shrimp quality. Aquaculture 301:64–68
Cruz-Suárez LE, Nieto-López M, Tapia-Salazar M, Gamboa-Delgado J, Maldonado-Muniz M, Villareal-Cavazos D, Ricque-Marie D (2013) Avances en la Valoración de Macroalgas del Género Ulva como Nutracéutico en Litopenaeus vannamei. In:Cruz-Suárez, L.E., Ricque-Marie, D., Tapia-Salazar, M., Nieto-López, M.G., Villarreal-Cavazos, D. A., Gamboa-Delgado, J., Alvarez-González C (ed) Contribuciones Recientes en Alimentación y Nutrición Acuícola, Universidad Autónoma de Nuevo León, pp 553–597
Dobrovolskaia MA, Medvedev AE, Thomas KE, Cuesta N, Toshchakov V, Ren T, Cody MJ, Michalek SM, Rice NR, Vogel SN (2003) Induction of in vitro reprogramming by Toll-like receptor (TLR)2 and TLR4 agonists in murine macrophages: effects of TLR “homotolerance” versus “heterotolerance” on NF-kB signaling pathway components. J Immunol 170:508–519
Ekblom R, Galindo J (2011) Applications of next generation sequencing in molecular ecology of non-model organisms. Heredity (Edinb) 107:1–15
Ergün S, Soyutürk M, Güroy B, Güroy D, Merrifield D (2008) Influence of Ulva meal on growth, feed utilization, and body composition of juvenile Nile tilapia (Oreochromis niloticus) at two levels of dietary lipid. Aquac Int 17:355–361
Götz S, García-Gómez JM, Terol J, Williams TD, Nagaraj SH, Nueda MJ, Robles M, Talón M, Dopazo J, Conesa A (2008) High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res 36:3420–3435
Grabherr MG, Haas BJ, Yassour M et al (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–652
Grabherr MG, Haas BJ, Yassour M et al (2013) Trinity: reconstructing a full-length transcriptome without a genome from RNA-Seq data. Nat Biotechnol 29:644–652
Guo H, Ye CX, Wang AL, Xian JA, Liao SA, Miao YT, Zhang SP (2013) Trascriptome analysis of the Pacific white shrimp Litopenaeus vannamei exposed to nitrite by RNA-seq. Fish Shellfish Immunol 35:2008–2016
Heffernan N, Smyth TJ, Soler-Villa A, Fitzgerald RJ, Brunton NP (2015) Phenolic content and antioxidant activity of fractions obtained from selected Irish macroalgae species (Laminaria digitata, Fucus serratus, Gracilaria gracilis and Codium fragile). J Appl Phycol 27:519–530
Immanuel G, Sivagnanavelmurugan M, Marudhupandi T, Radhakrishnan S, Palavesam A (2012) The effect of fucoidan from brown seaweed Sargassum wightii on WSSV resistance and immune activity in shrimp Penaeus monodon (Fab). Fish Shellfish Immunol 32:551–564
Kandasamy S, Khan W, Evans F, Critchley AT, Prithiviraj B (2012) Tasco®: a product of Ascophyllum nodosum enhances immune response of Caenorhabditis elegans against Pseudomonas aeruginosa infection. Mar Drugs 10:84–105
Kanehisa M, Goto S (2000) KEGG : Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36:D480–D484
Kim K-J, Lee O-H, Lee B-Y (2010) Fucoidan, a sulfated polysaccharide, inhibits adipogenesis through the mitogen-activated protein kinase pathway in 3T3-L1 preadipocytes. Life Sci 86:791–797
Kim J-K, Cho ML, Karnjanapratum S, Shin IS, You SG (2011) In vitro and in vivo immunomodulatory activity of sulfated polysaccharides from Enteromorpha prolifera. Int J Biol Macromol 49:1051–1058
Kumar M, Kumari P, Trivedi N, Shukla MK, Gupta V, Reddy CRK, Jha B (2011) Minerals, PUFAs and antioxidant properties of some tropical seaweeds from Saurashtra coast of India. J Appl Phycol 23:797–810
Kumari P, Kumar M, Gupta V, Reddy CRK, Jha B (2010) Tropical marine macroalgae as potential sources of nutritionally important PUFAs. Food Chem 120:749–757
Leiro JM, Castro R, Arranz A, Lamas J (2007) Immunomodulating activities of acidic sulphated polysaccharides obtained from the seaweed Ulva rigida C. Agardh. Int Immunopharmacol 7:879–888
Li X, Cui Z, Liu Y, Song C, Shi G (2013) Transcriptome analysis and discovery of genes involved in immune pathways from hepatopancreas of microbial challenged mitten crab Eriocheir sinensis. PLoS ONE 8:e68233
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Maeda H, Hosokawa M, Sashima T, Funayama K, Miyashita K (2005) Fucoxanthin from edible seaweed, Undaria pinnatifida, shows antiobesity effect through UCP1 expression in white adipose tissues. Biochem Biophys Res Commun 332:392–397
Marioni JC, Mason CE, Mane SM, Stephens M, Gilad Y (2008) RNA-seq : an assessment of technical reproducibility and comparison with gene expression arrays. Genome Res 18:1509–1517
Matanjun P, Mohamed S, Mustapha NM, Muhammad K, Ming CH (2008) Antioxidant activities and phenolics content of eight species of seaweeds from north Borneo. J Appl Phycol 20:367–373
Moriya Y, Itoh M, Okuda S, Yoshizawa AC, Kanehisa M (2007) KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res 35:W182–W185
Nakagawa H (1997) Effect of dietary algae on improvement of lipid metabolism in fish. Biomed Pharmacother 51:345–348
Nakagawa H, Kasahara S, Sugiyama T (1987) Effect of Ulva meal supplementation on lipid metabolism of black sea bream, Acanthopagrus schlegeli (Bleeker). Aquaculture 62:109–121
Ortiz J, Romero N, Robert P, Araya J, Lopez-Hernández J, Bozzo C, Navarrete E, Osorio A, Rios A (2006) Dietary fiber, amino acid, fatty acid and tocopherol contents of the edible seaweeds Ulva lactuca and Durvillaea antarctica. Food Chem 99:98–104
Peña-Rodríguez A, León A, Moll B, Tapia-Salazar M, Nieto-Lopez MG, Villareal-cavazos D, Ricque-Marie D, Cruz-Suarez LE (2010) Uso de Ulva clathrata en la nutrición del camarón blanco : revisión. In: Cruz-Suarez, L.E., Ricque-Marie, D., Tapia-Salazar, M., Nieto-López, M.G., Villarreal-Cavazos, D. A., Gamboa-Delgado, J. (eds) Avances en Nutrición Acuícola X - Memorias del X Simposio Internacional de Nutrición Acuícola, 8–10 de Noviembre, San Nicolás de los Garza, N. L., México. Universidad Autónoma de Nuevo León, Monterrey, pp 700–712
Peña-Rodríguez A, Mawhinney TP, Ricque-Marie D, Cruz-Suárez LE (2011) Chemical composition of cultivated seaweed Ulva clathrata (Roth) C. Agardh. Food Chem 129:491–498
Pengzhan Y, Ning L, Xiguang L, Gefei Z, Quanbin Z, Pengcheng L (2003) Antihyperlipidemic effects of different molecular weight sulfated polysaccharides from Ulva pertusa (Chlorophyta). Pharmacol Res 48:543–549
Ramirez-Higuera A, Quevedo-Corona L, Paniagua-Castro N, Chamorro-Ceballos G, Milliar-Garcia A, Jaramillo-Flores ME (2014) Antioxidant enzymes gene expression and antihypertensive effects of seaweeds Ulva linza and Lessonia trabeculata in rats fed a high-fat and high-sucrose diet. J Appl Phycol 26:597–605
Reilly P, O’Doherty JV, Pierce KM, Callan JJ, O’Sullivan JT, Sweeney T (2008) The effects of seaweed extract inclusion on gut morphology, selected intestinal microbiota, nutrient digestibility, volatile fatty acid concentrations and the immune status of the weaned pig. Animal 2:1465–1473
Robertson L, Bray W, Leung-Trujillo J, Lawrence A (1987) Practical molt staging of Penaeus setiferus and Penaeus stylirostris. J World Aquacult Soc 18:180–185
Robinson MD, Oshlack A (2010) A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol 11(3):R25
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140
Robinson M, McCarthy D, Chen Y, Smyth GK (2011) edgeR: differential expression analysis of digital gene expression data user’s guide. Most 23:1–77
Rocha de Souza MC, Marques CT, Guerra Dore CM, Frerreira da Silva FR, Rocha HAO, Leite EL (2007) Antioxidant activities of sulfated polysaccharides from brown and red seaweeds. J Appl Phycol 19:153–160
Selvin J, Huxley AJ, Lipton AP (2004) Immunomodulatory potential of marine secondary metabolites against bacterial diseases of shrimp. Aquaculture 230:241–248
Seo MJ, Choi HS, Lee OH, Lee BY (2013) Grateloupia lanceolata (Okamura) Kawaguchi, the edible red seaweed, inhibits lipid accumulation and reactive oxygen species production during differentiation in 3T3-L1 cells. Phytother Res 27:655–663
Shanab SMM, Shalaby EA, El-Fayoumy EA (2011) Enteromorpha compressa exhibits potent antioxidant activity. J Biomed Biotechnol 2011:726405. doi:10.1155/2011/726405
Simić S, Ranković B (2012) Evaluation of in vitro antioxidant and antimicrobial activities of green microalgae Trentepohlia umbrina. Not Bot Horti Agrobot Cluj-Napoca 40:86–91
Sookruksawong S, Sun F, Liu Z, Tassanakajon A (2013) RNA-Seq analysis reveals genes associated with resistance to Taura syndrome virus (TSV) in the Pacific white shrimp Litopenaeus vannamei. Dev Comp Immunol 41:523–533
Trigui M, Gasmi L, Zouari I, Tounsi S (2013) Seasonal variation in phenolic composition, antibacterial and antioxidant activities of Ulva rigida (Chlorophyta) and assessment of antiacetylcholinesterase potential. J Appl Phycol 25:319–328
Wijesekara I, Pangestuti R, Kim S-K (2011) Biological activities and potential health benefits of sulfated polysaccharides derived from marine algae. Carbohydr Polym 84:14–21
Wong ML, Medrano JF (2005) Real-time PCR for mRNA quantitation. Biotechniques 39:75–85
Wong KH, Sam SW, Cheung PCK, Ang PO (1999) changes in lipid profiles of rats fed with seaweed-based diets. Nutr Res 19:1519–1527
Yang C, Zhang J, Li F, Ma H, Zhang Q, Jose Priya TA, Zhang X, Xiang J (2008) A toll receptor from Chinese shrimp Fenneropenaeus chinensis is responsive to Vibrio anguillarum infection. Fish Shellfish Immunol 24:564–574
Ye J, Fang L, Zheng H, Zhang Y, Chen J, Zhang Z, Wang J, Li S, Li R, Bolund L, Wang J (2006) WEGO: a web tool for plotting GO annotations. Nucleic Acids Res 34:W293–W397
Yocum GD, Coudron TA, Brandt SL (2006) Differential gene expression in Perillus bioculatus nymphs fed a suboptimal artificial diet. J Insect Physiol 52:586–592
Yoon W-J, Ham YM, Kim S-S, Yoo B-S, Moon J-Y, Baik JS, Lee NH, Hyun C-G (2009) Suppression of pro-inflammatory cytokines, iNOS, and COX-2 expression by brown algae Sargassum micracanthum in RAW 264.7 macrophages. EurAsian J Biosci 143:130–143
Zeng D, Chen X, Xie D, Zhao Y, Yang C, Li Y, Ma N, Peng M, Yang Q, Liao Z, Wang H, Chen X (2013) Transcriptome analysis of Pacific white shrimp (Litopenaeus vannamei) hepatopancreas in response to Taura syndrome Virus (TSV) experimental infection. PLoS ONE 8:e57515
Zou D, Coudron TA, Liu C et al (2013) Nutrigenomics in Arma chinensis: transcriptome analysis of Arma chinensis fed on artificial diet and Chinese oak silk moth Antheraea pernyi pupae. PLoS ONE 8:e60881. doi:10.1371/journal.pone.0060881
Acknowledgments
We wish to thank Julian Gamboa for providing very valuable comments on earlier versions of this paper. We are grateful to David Villarreal for helping in the hepatopancreas sampling and moults stage determination. Thanks as well to Manuel Vazques Ruiz for the support in the relative quantification of the expression level of genes.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Elizondo-Reyna, E., Medina-González, R., Nieto-López, M.G. et al. Consumption of Ulva clathrata as a dietary supplement stimulates immune and lipid metabolism genes in Pacific white shrimp Litopenaeus vannamei . J Appl Phycol 28, 3667–3677 (2016). https://doi.org/10.1007/s10811-016-0889-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-016-0889-1