Abstract
Single-nucleotide polymorphisms (SNPs) are the most prevalent type of variation in genomes that are increasingly being used as molecular markers in diversity analyses, mapping and cloning of genes, and germplasm characterization. However, only a few studies reported large-scale SNP discovery in Aegilops tauschii, restricting their potential use as markers for the low-polymorphic D genome. Here, we report 68,592 SNPs found on the gene-related sequences of the 5D chromosome of Ae. tauschii genotype MvGB589 using genomic and transcriptomic sequences from seven Ae. tauschii accessions, including AL8/78, the only genotype for which a draft genome sequence is available at present. We also suggest a workflow to compare SNP positions in homologous regions on the 5D chromosome of Triticum aestivum, bread wheat, to mark single nucleotide variations between these closely related species. Overall, the identified SNPs define a density of 4.49 SNPs per kilobyte, among the highest reported for the genic regions of Ae. tauschii so far. To our knowledge, this study also presents the first chromosome-specific SNP catalog in Ae. tauschii that should facilitate the association of these SNPs with morphological traits on chromosome 5D to be ultimately targeted for wheat improvement.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wild goatgrass, Aegilops tauschii, is the D-genome progenitor of bread wheat, Triticum aestivum. The hexaploid bread wheat (2n = 6× = 42; AABBDD) originated around 8000 years ago through a spontaneous hybridization between the tetraploid emmer wheat, Triticum turgidum (2n = 4× = 28; AABB), and the diploid Ae. tauschii (2n = 2× = 14, DD), and thereafter assumed most of the global wheat production (Jia et al. 2013). Cultivated wheat species cover the arable lands more than any other food crops; yet, in terms of production and yield, wheat ranks the fourth after sugarcane, maize, and rice (http://faostat.fao.org/). Estimations on the growing world population and the climate change patterns may require up to 70% increase in wheat production by 2050 (Mayer et al. 2014). To meet this challenge, wild germplasms, including relatives and progenitors, will be key to explore favorable alleles and small RNAs such as microRNAs for wheat improvement (Nevo and Chen 2010; Lucas et al. 2011a, b; Kuzuoglu-Ozturk et al. 2012; Budak et al. 2013a; Budak et al. 2013b; Mochida and Shinozaki 2013; Akpinar et al. 2015a, b, c, d).
Molecular breeding allows the development and release of improved varieties much faster than classical breeding strategies. In particular, molecular markers have been pivotal in the screening of genetic resources for favorable alleles and in characterization of germplasms and introgressed lines (Budak et al. 2004; Budak et al. 2005; Castillo et al. 2010; Mizuno et al. 2010; Paux et al. 2012; Lucas et al. 2012; Budak et al. 2015; Winfield et al. 2015). While initially sequence-independent markers, such as restriction fragment length polymorphisms (RFLPs) and random amplified polymorphic DNA (RAPD), aided in genetic mapping and genetic diversity studies, molecular breeding efforts have mostly benefited from sequence-based markers which enable high-throughput, rapid, and cost-effective characterization of large numbers of samples (Lucas et al. 2012; Paux et al. 2012). Of these sequence-based markers, single-nucleotide polymorphisms (SNPs) have become the marker of choice as they overcome shortages of expressed sequence tags (ESTs) and simple sequence repeats (SSRs) (Berkman et al. 2012; Paux et al. 2012; Thomson 2014; Akpinar et al. 2015c). SNPs are the most widespread markers as they involve changes in the nucleotide level, which are transitions (C/T or G/A), transversions (C/G, C/A, or T/A, T/G), or small insertions/deletions (indels), and they can be linked to other marker types, such as EST-SNPs (Paux et al. 2012). Although homoeologous SNPs found in polyploid genomes complicate the identification of varietal SNPs, flexibility, speed, and cost-efficiency of SNP-based markers and platforms have greatly increased the popularity of SNPs in genetic and genomic studies. SNP-based marker systems have the potential to accelerate breeding programs not only by enabling the construction of highly saturated genetic maps but also by facilitating fine mapping of regions of interest and effective utilization of a germplasm by developing genome specific assays (Mammadov et al. 2012).
Ae. tauschii is a promising resource for wheat breeding due to its direct ancestry to bread wheat and genetically diverse populations (Dvorák et al. 1998; Wang et al. 2013). This is particularly important for the D genome traits, as bread wheat D genomes exhibit low level of polymorphism (Paux et al. 2012). Accordingly, this resource has begun to be realized particularly since the release of its whole genome physical map and draft genome sequence (Luo et al. 2013; Jia et al. 2013). SNPs have been utilized to map a powdery mildew resistance gene (Wang et al. 2015), to assess genetic diversity (Akhunov et al. 2010), to evaluate genetic and physical maps (You et al. 2010; Kumar et al. 2015), and to identify associations with morphological traits (Liu et al. 2015a; Liu et al. 2015b). Additionally, a few studies reported genome-wide large-scale SNP identification in Ae. tauschii (You et al. 2011; Iehisa et al. 2012; Wang et al. 2013; Winfield et al. 2015).
Here, we report the chromosome-specific discovery of SNPs in Ae. tauschii 5D chromosome using next-generation genomic and transcriptomic sequences of a total of seven Ae. tauschii accessions. Wheat 5D chromosome is known to carry alleles for important agronomic traits, such as dough strength (HMW-GS, Blechl and Anderson 1996), grain texture (Ha locus, Chantret et al. 2005), grain hardness (Pina, Pinb loci, Morris 2002), free-threshing (WAP2, Ning et al. 2009), flowering (Vrn loci, Quarrie et al. 2005; Yoshida et al. 2010; Zhang et al. 2012; Kippes et al. 2015), disease resistance (Lr1, Cloutier et al. 2007), abiotic stress tolerance (Galaeva et al. 2013), and putative microRNA target genes (Kurtoglu et al. 2013). Here, we also investigated single nucleotide variations (SNVs) between Ae. tauschii and T. aestivum 5D chromosomes by comparing SNVs found on T. aestivum 5D chromosome by mapping Ae. tauschii AL8/78 for which the draft genome sequence is available (Jia et al. 2013) and SNVs found on Ae. tauschii 5D chromosome by mapping T. aestivum D-genome reads published by the International Wheat Genome Sequencing Consortium (Mayer et al. 2014). SNPs and SNVs reported in this study provide an extensive collection on the D genome which should contribute to genetic and genomic efforts such as small RNAs (Budak and Kantar 2015).
Materials and Methods
Sequencing of Flow-Sorted Chromosomes
Flow sorting and sequencing of isolated chromosomes 5DS and 5DL from T. aestivum, 5B from Triticum dicoccoides, and 5D from Ae. tauschii were described previously (Akpinar et al. 2015a, b; Lucas et al. 2014). Briefly, suspensions of intact mitotic chromosomes were isolated from T. aestivum L. var. Chinese Spring using a double ditelosomic line (2n = 40 + 2t5DS + 2t5DL) and from Ae. tauschii genotype MvGB589, and the 5D chromosomes were selected by flow sorting, evaluated for purity, and amplified for direct sequencing, as described previously in detail (Lucas et al. 2014; Akpinar et al. 2015b; Akpinar and Budak 2016).
Shotgun libraries were prepared from 0.5 μg of amplified DNA from each flow-sorted chromosome/chromosome arm, quantified, and sequenced using GS FLX Titanium kits according to the manufacturer’s protocols (all Roche Life Sciences) as previously described (Akpinar et al. 2015a; Lucas et al. 2014). Sequence reads are submitted to the EBI European Nucleotide Archive under the primary accession numbers: ERP002330 (T. aestivum 5D) and PRJEB5993 (Ae. tauschii 5D).
Additional Datasets and Software
Whole genome shotgun sequence reads from Ae. tauschii accession AL8/78 (Jia et al. 2013) were downloaded from the NCBI Sequence Read Archive (BioProject ID: PRJNA182898; run accession numbers: SRR124169-SRR124252). Illumina sequence reads produced from flow-sorted chromosome arms of T. aestivum L. var. Chinese Spring (Mayer et al. 2014) were downloaded from the EBI European Nucleotide Archive in .fastq format (run accession numbers: ERR277143–ERR277149).
RNA-sequencing reads from six Ae. tauschii accessions, IG47182 (DRR001933), PI476874 (DRR001934), 2220007 (SRR947843-SRR947850), AS2404 (SRR1222442-SRR1222447), TQ20 (SRR043333-SRR043335), and D2220009 (SRR949823), were retrieved from the NCBI Sequence Read Archive. Illumina and 454 reads were checked for read quality and composition using FastQC (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and PRINSEQ (http://prinseq.sourceforge.net/), respectively. Reads containing tag, adapter, or primer sequences were trimmed. Illumina reads of 49–51-nucleotide length containing >10% Ns and 454 reads containing >1% Ns were discarded to avoid ambiguity.
Illumina reads were aligned to reference sequences using Burrows-Wheeler Alignment (bwa) software (Li and Durbin 2009). Identification of sequence variations and initial filtering of potential SNPs were carried out using SAMtools (Li et al. 2009). Routine comparisons of small numbers of sequences were carried out using standalone BLAST+ applications, v.2.2.27 (Camacho et al. 2009). Sequence annotations were performed using Blast2GO software with an initial blast step run locally against all known Viridiplantae proteins (Conesa and Götz 2008).
Defining 5D Chromosome Gene-Related Reference Sequences
In order to identify SNPs present on the 5D chromosomes, we used a strategy adapted from that described by You et al. (2011) for calling SNPs using NGS data in the absence of a finished quality genome sequence. First of all, the low-coverage 454 reads from each chromosome (arm) were filtered against mitochondrial/chloroplast DNA contamination and rRNA sequences and masked against known repetitive elements. The masked sequences were then assembled using gsAssembler (Roche 454 Life Sciences), and contigs with higher than expected depth (defined as 3× the average sequencing coverage of the chromosome) were excluded as potential unknown repeat elements. From the remaining low-copy-number (LCN) contigs and singletons, those associated with coding sequences were identified using sequence similarity searches against annotated protein sequences of Brachypodium distachyon, Sorghum bicolor, and Oryza sativa and high-confidence predicted protein sequences of Hordeum vulgare. These procedures are described in detail in our previous papers (Akpinar et al. 2015a,b; Lucas et al. 2014). The gene-associated, LCN contigs and singletons were combined into a single fasta file for each chromosome/chromosome arm and used as references for identifying sequence variations.
SNP Calling and Filtering
Read alignment, mapping, and variant calling steps were carried out on the high-performance computing cluster. For SNP identification from Ae. tauschii AL8/78, run files containing paired-end reads with nominal insert sizes ranging from 167 to 510 nt giving a total average genome coverage of >50× were aligned to the references using the bwa align and sampe modules (reads from libraries with smaller insert sizes were preferred, as both halves of each pair were more likely to map to the same 454 singleton or contig). For SNP identification from T. aestivum var. Chinese Spring, paired-end reads from the flow-sorted chromosomes of the D genome were used for a comparable data volume (Mayer et al. 2014). The resulting alignment files in Sequence Alignment/Map (.sam) format were converted to sorted binary (.bam) files using samtools view and sort. After sorting, the .bam files generated from all runs from the same biological sample were combined, using “samtools mpileup”; the command “bcftools view—vcg” was then used to extract all sequence variations in the variant call format (.vcf). Filtering criteria were then applied to eliminate apparent variations resulting from alignment errors as follows: sites with a read depth of <10 or >3 times the average depth of coverage of the chromosome, or a mapping quality score of <30, and sites within 3 nt of an alignment gap were eliminated using vcfutils.pl VarFilter. Finally, in cases where two sequence variations were detected separated by 3 nt or fewer, both were removed as probable products of low-quality sequence data. After these criteria had been applied, the remaining variations were accepted as putative SNPs or SNVs. SNPs/SNVs identified from different genotypes were compared and collated using in-house Perl scripts that are available upon request. The summarized workflow is given in Fig. 1.
Results and Discussion
Identification of SNPs in Multiple Ae. tauschii Genotypes
Prior to the identification of SNPs on genic sequences from Ae. tauschii chromosome 5D, our SNP calling and filtering workflow was tested using two independently generated sequence datasets in the Ae. tauschii AL8/78 genotype. The validity of the conditions for initial SNP filtering was confirmed by mapping the Ae. tauschii AL8/78 whole genome shotgun reads from Jia et al. (2013), against preliminary sequence scaffolds generated independently from pooled BAC clones of the same accession by Dvorak et al. (http://aegilops.wheat.ucdavis.edu/ATGSP/index.php). Any variants detected between these two datasets should be the result of errors in sequencing or alignment, as both derive from the same genetic material. Accordingly, while initial alignment and variant calling gave a putative 274,286 SNVs and indels, 274,163 of these (99.96%) were eliminated by filtering for read depth and mapping quality as described above. Elimination of all but 123 out of a total 274,286 variations indicated that filtering criteria employed in our workflow can differentiate most sequencing errors from genuine SNPs. This gives confidence that the filtering criteria used are stringent enough to remove the vast majority of erroneous SNP calls.
The filtered SNP calls generated for each genotype under test were compared and collated using in-house Perl scripts, with the workflow summarized in Fig. 1. Briefly, .vcf files containing SNP calls for different genotypes aligned to the Ae. tauschii 5D gene-related reference sequences from accession MvGB589 were combined, sorted, and assigned unique SNP IDs based on their position in the reference sequences. As the most comprehensive dataset available, the genomic DNA reads from Ae. tauschii AL8/78 were used to filter the resulting summary table. Only positions in the reference to which AL8/78 reads aligned at a depth of 10 or more were retained, and the genotype of AL8/78 at every position called as a SNP in one or more of the other accessions but not AL8/78 was re-checked. In most cases, AL8/78 matched the reference sequence, but in some positions, AL8/78 also had indications of sequence variation but had not been called as an SNP owing to high sequence depth or low mapping quality. These instances were accepted as genuine SNPs based on the strength of the evidence from other accessions.
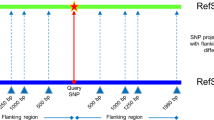
Furthermore, each Ae. tauschii SNP was compared with its corresponding position in T. aestivum var. Chinese Spring. Firstly, gene-related sequences from Chinese Spring 5D chromosome (Lucas et al. 2014) were used to construct a BLAST database. Then, a segment of sequence centered on each SNP was excised from the Ae. tauschii MvGB589 reference dataset and searched for the best match in the Chinese Spring 5D sequences using blastn, requiring a minimum of 90% sequence identity over at least 60 nt. The optimum length of the SNP-centered sequence segment to search for was determined empirically by carrying out searches with segment lengths of 60, 80, 100, and 120 nt and comparing the number of matches to these segments at different percentage sequence identities. Most matches were the same in all of these searches; however, the total number of matches increased slightly with segment length. It was observed that up to 100 nt, the increase was primarily at high identity (>98%) whereas at 120 nt, most of the increase was at 90–96% identity, suggesting that less reliable alignments were beginning to be included (Fig. 2). Therefore, 100 nt was selected as the optimum segment length. For the matches obtained to these segments, the equivalent base in Chinese Spring 5D for each Ae. tauschii SNP position was added to the summary table of all SNPs (Supplementary Table S1).
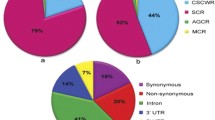
In total, compared to gene-related sequences from chromosome 5D of Ae. tauschii, 68,592 SNPs were identified in one or more accessions, corresponding to an average density of 4.49 SNPs per kilobyte in these sequences. As expected, more SNPs were identified in genomic shotgun than RNA-Seq datasets, with 31,250 identified in AL8/78 and a total of 25,543 in chromosome arms 5DS and 5DL of T. aestivum var. Chinese Spring (Table 1). A small number of SNP positions (550) were called in both 5DS and 5DL; these could be explained by sequence reads from genes with highly similar paralogs on both chromosome arms matching each other. Although majority of the gene-related sequences carrying those SNPs lacked known orthologs in Viridiplantae, others that do exhibit orthologous relationships to known plant proteins suggested involvement in core functions and/or affiliation to large gene families or repeat elements. For instance, seven such contigs and singletons were associated with serine-threonine kinases and phosphatases as core components of cellular signal transduction. Interestingly, 16 more contigs and singletons revealed significant similarity to the far-red-impaired response (FAR) family of transcription factors (TFs) and mutator proteins that are related to the Mutator-like element (MULE) superfamily of transposases (Lin et al. 2007). Two contigs had significant similarity to the SKP1-Cullin-F-box (SCF) E3 ubiquitin ligase complex, while two more sequences were also related to components of the protein turnover machinery (pumilio and ubiquitin carrier E2). Both SKP1-like and pumilio-like genes have multiple copies in plant genomes (Tam et al. 2010; Hong et al. 2013). These observations support the view that SNP positions commonly identified both in 5DS and 5DL may indeed correspond to genes with paralogous coding regions on both chromosome arms or genes associated with repetitive elements. Majority of the SNP-containing Aegilops 5D contigs and singletons had only a single SNP (Fig. 3a). While almost 68% of the contigs, which are much longer than singleton reads, contained one or more SNPs, only 32% of the singletons carried at least one SNP. Overall, Aegilops 5D contigs also had higher numbers of SNPs per contig, compared to singletons (Fig. 3b).
Transcriptome-Guided SNP Identification
Gene-associated sequences from Ae. tauschii (MvGB589) 5D chromosome were aligned with next-generation RNA-Seq reads from six Ae. tauschii genotypes (Table 2). Cumulatively, 17,553 SNPs were identified between the MvGB589 genotype and the six accessions, corresponding to an average density of 1.15 SNPs per kilobyte (corresponding to 1 SNP every ∼870 bp), close to the SNP density (1 SNP every ∼876 bp) in genic sequences between Ae. tauschii AL8/78 and AS75 genotypes (You et al. 2011). However, higher SNP densities, as high as 1 in every 202 bp, were reported for the genic regions on the wheat D genome (Akhunov et al. 2010). This figure is more similar to the overall SNP density identified from all genotypes (including T. aestivum Chinese Spring), which is close to 1 SNP per 223 bp, suggesting that RNA-Seq reads may not be sufficient to discover all SNPs linked to gene-associated regions on the genome. Since the gene-associated sequences used in the identification of SNPs were of genomic source, these sequences may include intergenic regions, such as introns and flanking regulatory elements, which may not be fully covered by RNA-Seq reads, resulting in the underestimation of the overall SNPs in these sequences.
The number of identified SNPs in individual accessions was largely dependent on the size of the transcriptomic data available, ranging from 33 (accession TQ20) to 11,585 (accession D2220009). With larger datasets, however, the number of SNPs appeared to level off around 12,000 which may indicate the maximum number of SNPs that can be identified by transcriptomic sequences of a single accession (Fig. 4). For the accession D2220009, for which size of the RNA-Seq data was the largest, an SNP density of 1 in every 1318 bp was observed on gene-related 5D sequences, which is considerably lower than that of the estimated SNP density for any random Ae. tauschii haplotypes (corresponding to 1 SNP in every 409 bp; Dvorak et al. 2011). This may suggest a closer evolutionary relationship between D2220009 and MvGB589, assuming similar evolutionary dynamics for the remaining chromosomes. Interestingly, ∼3-Gb RNA-Seq reads of AS2404 genotype covered almost 80% of all gene-associated 5D sequences, similar to much larger datasets of 2220007 (12.4 Gb) and D2220009 (31.8 Gb) accessions, although far fewer SNPs could be identified in this genotype (2486 vs 9752 and 11,585, respectively). This may also suggest a closer relationship between MvGB589 and AS2404, compared to 2220007 and D2220009 accessions. Additionally, accessions IG47182 and PI476874 had comparable transcriptomic sequences mapping against the gene-related 5D sequences. While IG47182 revealed 203 SNPs in 71 gene-related sequences from the 5D chromosome, PI476874 had 228 SNPs in 84 gene-related 5D sequences. Of these, 31 SNPs were common to IG47182 and PI476874, with 29 having the same alternative bases in these accessions.
Among the SNPs from all samples, the ratio of transitions (Ti; C<>T or A<>G) to transversions (Tv; Purine <> Pyrimidine) was 1.66 ± 0.25 (excluding the smallest dataset TQ20, 1.75 ± 0.12). A bias towards transitions is commonly observed in SNP analysis due to the high rate at which methylated cytosine residues are deaminated to uracil in biological conditions (Shen et al. 1994). This bias increases confidence that the variants detected are real SNPs; conversely, random sequencing errors would be expected to have a Ti/Tv ratio of 0.5, as for any base substitution, there are two possible transversions and only one transition. Curiously, the Ti/Tv ratio of genic SNPs between PI476874 and MvGB589 5D chromosome was considerably higher (1.92) than all others, including that of the highly similar dataset of IG47182 (Table 2). A previous study of SNPs in the group of seven chromosomes of T. aestivum (Lorenc et al. 2012) observed a Ti/Tv ratio of 2.12. This higher value could be explained by the bread wheat genome being more highly methylated than Ae. tauschii, owing to its hexaploidy and higher repeat content.
Polymorphic SNPs in Ae. tauschii Accessions
In total, 17,553 SNPs were identified by mapping over 40-Gb transcriptomic sequences from all six Ae. tauschii accessions on gene-related 5D sequences of genotype MvGB589. When SNPs identified from individual samples were combined, however, the total number of SNPs was 26,575, likely indicating that almost one third of the SNPs identified by the combined transcriptome sequences lacked sufficient depth in individual samples. This may be due to the limited data size of certain accessions and low representations of transcripts that are not highly expressed in the tissues or under the conditions that the samples are from.
A total of 5732 SNPs were covered by two or more genotypes (Fig. 5). Of these, 4179 (72%) have the same alternative base in all genotypes covering the SNP position, differing from MvGB589. Similarly, of the 965 SNPs covered by three or more genotypes, 776 had the same alternative base in all genotypes covering the SNP position, whereas among the remaining, some genotypes had the same reference base. The SNPs found between the gene-related sequences of the 5D chromosome of MvGB589 and transcriptome sequences of six Ae. tauschii accessions were exclusively biallelic, assuming only two out of the four (A, G, T, C) possibilities.
References
Akhunov ED, Akhunova AR, Anderson OD et al (2010) Nucleotide diversity maps reveal variation in diversity among wheat genomes and chromosomes. BMC Genomics 11:702. doi:10.1186/1471-2164-11-702
Akpinar BA, Budak H (2016) Dissecting miRNAs in wheat D genome progenitor, Aegilops tauschii. Front Plant Sci 7:1–17. doi:10.3389/fpls.2016.00606
Akpinar BA, Yuce M, Lucas SJ et al (2015a) Molecular organization and comparative analysis of chromosome 5B of the wild wheat ancestor Triticum dicoccoides. Scientific Reports 5:Article number: 10763. doi:10.1038/srep10763
Akpinar BA, Lucas SJ, Vr J et al (2015b) Sequencing chromosome 5D of Aegilops tauschii and comparison with its allopolyploid descendant bread wheat ( Triticum aestivum ). Plant Biotechnol J 13:740–752. doi:10.1111/pbi.12302
Akpinar BA, Magni F, Yuce M et al (2015c) The physical map of wheat chromosome 5DS revealed gene duplications and small rearrangements. BMC Genomics 16:453. doi:10.1186/s12864-015-1641-y
Akpinar BA, Kantar M, Budak H (2015d) Root precursors of microRNAs in wild emmer and modern wheats show major differences in response to drought stress. Funct Integr Genomics 15:587. doi:10.1007/s10142-015-0453-0
Berkman PJ, Lai K, Lorenc MT, Edwards D (2012) Next-generation sequencing applications for wheat crop improvement. Am J Bot 99:365–371. doi:10.3732/ajb.1100309
Blechl AE, Anderson OD (1996) Expression of a novel high-molecular-weight glutenin subunit gene in transgenic wheat. Nat Biotechnol 14:875–879. doi:10.1038/nbt0796-875
Budak H, Kantar M (2015) Harnessing NGS and big data optimally: comparison of miRNA prediction from assembled versus non-assembled sequencing data—the case of the grass Aegilops tauschii complex genome. OMICS: A J Integr Biol 19(7):407–415. doi:10.1089/omi.2015.0038
Budak H, Baenziger PS, Beecher et al (2004) The effect of introgressions of wheat D-genome chromosomes of into ‘Presto’ tricale. Euphytica 137:261–270. doi:10.1023/B:EUPH.0000041590.55511.d0
Budak H, Shearman RC, Gulsen O et al (2005) Understanding ploidy complex and geographic origin of the Buchloe dactyloides genome using cytoplasmic and nuclear marker systems. Theor Appl Genet 111:1545–1552. doi:10.1007/s00122-005-0083-3
Budak H, Kantar M, Kurtoglu KY (2013a) Drought tolerance in modern and wild wheat. ScientificWorldJournal 2013:548246. doi:10.1155/2013/548246
Budak H, Akpinar BA, Unver T, Turktas M (2013b) Proteome changes in wild and modern wheat leaves upon drought stress by two-dimensional electrophoresis and nanoLC-ESI–MS/MS. Plant Mol Biol 83(1–2):89–103. doi:10.1007/s11103-013-0024-5
Budak H, Hussain B, Khan Z et al (2015) From genetics to functional genomics: improvement in drought signaling and tolerance in wheat. Front Plant Sci 6:1–13. doi:10.3389/fpls.2015.01012
Camacho C, Coulouris G, Avagyan V et al (2009) BLAST+: architecture and applications. BMC Bioinforma 10:421. doi:10.1186/1471-2105-10-421
Castillo A, Dorado G, Feuillet C, Sourdille P, Hernandez P (2010) Genetic structure and ecogeographical adaptation in wild barley (Hordeum chilense Roemer et Schultes) as revealed by microsatellite markers. BMC Plant Biol 10:266. doi:10.1186/1471-2229-10-266
Chantret N, Salse J, Sabot F et al (2005) Molecular basis of evolutionary events that shaped the hardness locus in diploid and polyploid wheat species (Triticum and Aegilops). Plant Cell 17:1033–1045. doi:10.1105/tpc.104.029181
Cloutier S, McCallum BD, Loutre C et al (2007) Leaf rust resistance gene Lr1, isolated from bread wheat (Triticum aestivum L.) is a member of the large psr567 gene family. Plant Mol Biol 65:93–106. doi:10.1007/s11103-007-9201-8
Conesa A, Götz S (2008) Blast2GO: a comprehensive suite for functional analysis in plant genomics. Int J Plant Genomics 2008:619832. doi:10.1155/2008/619832
Dvorák J, Luo MC, Yang ZL (1998) Restriction fragment length polymorphism and divergence in the genomic regions of high and low recombination in self-fertilizing and cross-fertilizing aegilops species. Genetics 148:423–434
Dvorak J, Luo MC, Akhunov ED (2011) N.I. Vavilov’s theory of centres of diversity in the light of current understanding of wheat diversity, domestication and evolution
Galaeva MV, Fayt VI, Chebotar SV et al (2013) Association of microsatellite loci alleles of the group-5 of chromosomes and the frost resistance of winter wheat. Cytol Genet 47:261–267. doi:10.3103/S0095452713050046
Hong MJ, Kim DY, Seo YW (2013) SKP1-like-related genes interact with various F-box proteins and may form SCF complexes with Cullin–F-box proteins in wheat. Mol Biol Rep 40:969–981. doi:10.1007/s11033-012-2139-1
Iehisa JCM, Shimizu A, Sato K et al (2012) Discovery of high-confidence single nucleotide polymorphisms from large-scale de novo analysis of leaf transcripts of Aegilops tauschii, a wild wheat progenitor. DNA Res 19:487–497. doi:10.1093/dnares/dss028
Jia J, Zhao S, Kong X et al (2013) Aegilops tauschii draft genome sequence reveals a gene repertoire for wheat adaptation. Nature 496:91–95. doi:10.1038/nature12028
Kippes N, Debernardi JM, Vasquez-Gross HA et al (2015) Identification of the VERNALIZATION 4 gene reveals the origin of spring growth habit in ancient wheats from South Asia. Proc Natl Acad Sci U S A 112:E5401–E5410. doi:10.1073/pnas.1514883112
Kumar A, Seetan R, Mergoum M et al (2015) Radiation hybrid maps of the D-genome of Aegilops tauschii and their application in sequence assembly of large and complex plant genomes. BMC Genomics 16:800. doi:10.1186/s12864-015-2030-2
Kurtoglu KY, Kantar M, Lucas SJ et al (2013) Unique and conserved microRNAs in wheat chromosome 5D revealed by next-generation sequencing. PLoS One 8(7):e69801. doi:10.1371/journal.pone.0069801
Kuzuoglu-Ozturk D, Yalcinkaya O, Akpinar BA et al (2012) Autophagy-related gene, TdAtg8, in wild emmer wheat plays a role in drought and osmotic stress response. Planta 236:1081–1092. doi:10.1007/s00425-012-1657-3
Leach LJ, Belfield EJ, Jiang C, Brown C, Mithani A, Harberd NP (2014) Patterns of homoeologous gene expression shown by RNA sequencing in hexaploid bread wheat. BMC Genomics 15:276. doi:10.1186/1471-2164-15-276
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760. doi:10.1093/bioinformatics/btp324
Li H, Handsaker B, Wysoker A et al (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079. doi:10.1093/bioinformatics/btp352
Li A, Liu D, Wu J et al (2014) mRNA and small RNA transcriptomes reveal insights into dynamic homoeolog regulation of allopolyploid heterosis in nascent hexaploid wheat. Plant Cell 26(5):1878–1900. doi:10.1105/tpc.114.124388
Lin R, Ding L, Casola C, et al. (2007) Transposase-derived transcription factors regulate light signaling in Arabidopsis
Liu Y, Wang L, Deng M et al (2015a) Genome-wide association study of phosphorus-deficiency-tolerance traits in Aegilops tauschii. Theor Appl Genet 128:2203–2212. doi:10.1007/s00122-015-2578-x
Liu Y, Wang L, Mao S et al (2015b) Genome-wide association study of 29 morphological traits in Aegilops tauschii. Sci Rep 5:15562. doi:10.1038/srep15562
Lorenc MT, Hayashi S, Stiller J et al (2012) Discovery of single nucleotide polymorphisms in complex genomes using SGSautoSNP. Biology (Basel) 1:370–382. doi:10.3390/biology1020370
Lucas SJ, Durmaz E, Akpinar BA et al (2011a) The drought response displayed by a DRE binding protein from Triticum dicocoides. Plant Physiol Bioechem 49:346–352. doi:10.1016/j.plaphy.2011.01.016
Lucas SJ, Dogan E, Budak H (2011b) TMPIT1 from wild emmer wheat: first characterisation of a stress-inducible integral membrane protein. Gene 483:22–28. doi:10.1016/j.gene.2011.05.003
Lucas SJ, Simkova H, Safar J et al (2012) Functional features of a single chromosome arm in wheat (1AL) determined from its structure. Funct Integ Genomics 12:173. doi:10.1007/s10142-011-0250-3
Lucas SJ, Akpınar BA, Simková H et al (2014) Next-generation sequencing of flow-sorted wheat chromosome 5D reveals lineage-specific translocations and widespread gene duplications. BMC Genomics 15:1–18. doi:10.1186/1471-2164-15-1080
Luo M, Gu YQ, You FM, et al. (2013) A 4-gigabase physical map unlocks the structure and evolution of the complex genome of Aegilops tauschii, the wheat D-genome progenitor. PNAS. doi:10.1073/pnas.1219082110
Mammadov J, Aggarwal R, Buyyarapu R et al (2012) SNP markers and their impact on plant breeding. Int J Plant Genomics 2012:728398. doi:10.1155/2012/728398
Mayer KFX, Rogers J, Dolezel J et al (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345(6194):1251788–1251788. doi:10.1126/science.1251788
Mizuno N, Yamasaki M, Matsuoka Y et al (2010) Population structure of wild wheat D-genome progenitor Aegilops tauschii Coss.: implications for intraspecific lineage diversification and evolution of common wheat. Mol Ecol 19:999–1013. doi:10.1111/j.1365-294X.2010.04537.x
Mochida K, Shinozaki K (2013) Unlocking Triticeae genomics to sustainably feed the future. Plant Cell Physiol 54:1931–1950. doi:10.1093/pcp/pct163
Morris CF (2002) Puroindolines: the molecular genetic basis of wheat grain hardness. Plant Mol Biol 48:633–647
Nevo E, Chen G (2010) Drought and salt tolerances in wild relatives for wheat and barley improvement. Plant Cell Environ 33:670–685. doi:10.1111/j.1365-3040.2009.02107.x
Ning S-Z, Chen Q-J, Yuan Z-W et al (2009) Characterization of WAP2 gene in Aegilops tauschii and comparison with homoeologous loci in wheat. J Syst Evol 47:543–551. doi:10.1111/j.1759-6831.2009.00048.x
Paux E, Sourdille P, Mackay I, Feuillet C (2012) Sequence-based marker development in wheat: advances and applications to breeding. Biotechnol Adv 30:1071–1088. doi:10.1016/j.biotechadv.2011.09.015
Quarrie SA, Steed A, Calestani C et al (2005) A high-density genetic map of hexaploid wheat (Triticum aestivum L.) from the cross Chinese Spring x SQ1 and its use to compare QTLs for grain yield across a range of environments. Theor Appl Genet 110:865–880. doi:10.1007/s00122-004-1902-7
Shen JC, Rideout WM, Jones PA (1994) The rate of hydrolytic deamination of 5-methylcytosine in double-stranded DNA. Nucleic Acids Res 22:972–976
Tam PP, Barrette-Ng IH, Simon DM et al (2010) The Puf family of RNA-binding proteins in plants: phylogeny, structural modeling, activity and subcellular localization. BMC Plant Biol 10:44. doi:10.1186/1471-2229-10-44
Thomson MJ (2014) High-throughput SNP genotyping to accelerate crop improvement. Plant Breed Biotechnol 2:195–212. doi:10.9787/PBB.2014.2.3.195
Wang J, Luo MC, Chen Z et al (2013) Aegilops tauschii single nucleotide polymorphisms shed light on the origins of wheat D-genome genetic diversity and pinpoint the geographic origin of hexaploid wheat. New Phytol 198:925–937. doi:10.1111/nph.12164
Wang Z, Li H, Zhang D et al (2015) Genetic and physical mapping of powdery mildew resistance gene MlHLT in Chinese wheat landrace Hulutou. Theor Appl Genet 128:365–373. doi:10.1007/s00122-014-2436-2
Winfield MO, Allen AM, Burridge AJ et al (2015) High-density SNP genotyping array for hexaploid wheat and its secondary and tertiary gene pool. Plant Biotechnol J n/a-n/a. doi:10.1111/pbi.12485
Yoshida T, Nishida H, Zhu J et al (2010) Vrn-D4 is a vernalization gene located on the centromeric region of chromosome 5D in hexaploid wheat. Theor Appl Genet 120:543–552. doi:10.1007/s00122-009-1174-3
You FM, Luo M-C, Xu K et al (2010) A new implementation of high-throughput five-dimensional clone pooling strategy for BAC library screening. BMC Genomics 11:692. doi:10.1186/1471-2164-11-692
You FM, Huo N, Deal KR et al (2011) Annotation-based genome-wide SNP discovery in the large and complex Aegilops tauschii genome using next-generation sequencing without a reference genome sequence. BMC Genomics 12:59. doi:10.1186/1471-2164-12-59
Zhang J, Wang Y, Wu S et al (2012) A single nucleotide polymorphism at the Vrn-D1 promoter region in common wheat is associated with vernalization response. Theor Appl Genet 125:1697–1704. doi:10.1007/s00122-012-1946-z
Acknowledgements
We thank Dr. Istvan Molnar (Agricultural Institute, Centre for Agricultural Research, Martonvasar, Hungary) for providing Ae. tauschii accession MvGB589.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work funded by Montana Plant Science Endowment, Montana State University and Sabanci University.
Electronic supplementary material
Supplementary Table S1
(XLSX 14593 kb)
Rights and permissions
About this article
Cite this article
Akpinar, B.A., Lucas, S. & Budak, H. A large-scale chromosome-specific SNP discovery guideline. Funct Integr Genomics 17, 97–105 (2017). https://doi.org/10.1007/s10142-016-0536-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-016-0536-6