Abstract
Natural variation in wheat requirement of long exposures to cold temperatures to accelerate flowering (vernalization) is mainly controlled by the Vrn-1, Vrn-2, Vrn-3, and Vrn-4 loci. The first three loci have been well characterized, but limited information is available for Vrn-4. So far, natural variation for Vrn-4 has been detected only in the D genome (Vrn-D4), and genetic stocks for this gene are available in Triple Dirk (TDF, hereafter). We detected heterogeneity in the Vrn-1 alleles present in different TDF stocks, which may explain inconsistencies among previous studies. A correct TDF seed stock from Japan carrying recessive vrn-A1, vrn-B1, and vrn-D1 alleles was crossed with three different winter cultivars to generate F2 mapping populations. Most of the variation in flowering time in these three populations was controlled by a single locus, Vrn-D4, which was mapped within a 1.8 cM interval flanked by markers Xcfd78 and Xbarc205 in the centromeric region of chromosome 5D. A factorial ANOVA for heading time using Vrn-D4 alleles and vernalization as factors showed a significant interaction (P < 0.0001), which confirmed that the Vrn-D4 effect on flowering time is modulated by vernalization. Comparison of the different Triple Dirk stocks revealed that Vrn-B1, Vrn-D1, and Vrn-D4 all have a small residual response to vernalization, but Vrn-D4 differs from the other two in its response to short vernalization periods. The precise mapping and characterization of Vrn-D4 presented here represent a first step toward the positional cloning of this gene.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Flowering at an optimal time is very important for plant reproductive success. To achieve this, plants monitor seasonal changes using environmental cues, such as differences in day length (photoperiod) and the exposure to low temperatures for extended periods of time (vernalization). These seasonal cues are integrated with additional information from the environment (e.g., water or nutrient stresses, limited root space, etc.) and from internal cues (e.g., age of the plant) to determine the initiation of the reproductive phase. The regulation of this transition is particularly critical for annual plants, such as the temperate cereals, since the transition to the reproductive phase is intimately associated with senescence and plant death.
The requirement for vernalization is particularly important for winter cereals to avoid cold injury of the sensitive floral organs during the winter. In wheat, vernalization requirement is controlled by four major genes designated Vrn-1, Vrn-2, Vrn-3, and Vrn-4 (reviewed in Distelfeld et al. 2009a; Flood and Halloran 1986; Trevaskis et al. 2007; Worland et al. 1987). The first three genes have been identified using map-based cloning approaches and validated using mutants and transgenic plants (Yan et al. 2003, 2004b, 2006).
The Vrn-1 gene encodes a MADS-box transcription factor closely related to the Arabidopsis AP1/FRUITFULL family (Yan et al. 2003), which is essential for the transition from the vegetative to reproductive stage in wheat (Shitsukawa et al. 2007). Natural insertions or deletions (indels) in regulatory regions of the three homoeologous genes found in hexaploid wheat (Vrn-A1, Vrn-B1, and Vrn-D1) are associated with dominant alleles for spring growth habit (Fu et al. 2005; Yan et al. 2004a). During vernalization, these regulatory regions show changes in histone methylation and acetylation associated with the transition between repressed and active chromatin states (Oliver et al. 2009). Different combinations of Vrn-A1, Vrn-B1, and Vrn-D1 dominant alleles are the most common sources of spring growth habit among landraces and commercial cultivars of polyploid wheat around the world (Fu et al. 2005; Iqbal et al. 2007; Iwaki et al. 2000, 2001; Stelmakh 1987b; Yan et al. 2004a; Zhang et al. 2008).
The Vrn-2 locus includes two linked and related proteins designated ZCCT1 and ZCCT2, characterized by the presence of a putative zinc finger and a CCT domain (Yan et al. 2004b). Deletions and mutations involving both ZCCT1 and ZCCT2 genes are frequent in diploid wheat and barley and are associated with recessive alleles for spring growth habit (Dubcovsky et al. 2005; Hemming et al. 2009; Yan et al. 2004a). Among the tetraploid wheat species, the Vrn-B2 gene is generally functional whereas the Vrn-A2 gene is not (Distelfeld et al. 2009b). Since Vrn-2 is the only locus with a dominant winter growth habit, at least one functional copy of Vrn-2 combined with homozygous recessive alleles at all three Vrn-1 loci is required to confer winter growth habit in hexaploid wheat.
The Vrn-B3 locus (formerly known as Vrn-5 or Vrn-B4; McIntosh et al. 2003) is homologous to the Arabidopsis FT gene (Yan et al. 2006). This dominant allele, found in the variety Hope, is associated with the insertion of a transposable element in the Vrn-B3 promoter. Natural variation at the Vrn-A3 and Vrn-D3 loci has also been described in hexaploid wheat (Bonnin et al. 2008). Vrn-3 promotes the transcription of Vrn-1 and accelerates flowering (Li and Dubcovsky 2008; Yan et al. 2006). In several species, it has been shown that FT can travel from the leaves to the shoot apex through the phloem (Corbesier et al. 2007; Lin et al. 2007; Tamaki et al. 2007). In wheat, the VRN3 protein interacts with FDL2, which binds to the Vrn-1 promoter (Li and Dubcovsky 2008).
Current models of flowering regulation in the temperate cereals suggest that, before vernalization, Vrn-3 is repressed by Vrn-2 (Hemming et al. 2008; Yan et al. 2006). Long exposures to cold temperature result in the up-regulation of Vrn-1 and the down-regulation of Vrn-2 in the leaves. The release from the Vrn-2 repression results in higher transcript levels of Vrn-3 and the promotion of Vrn-1 above the threshold levels required for flower induction (Distelfeld et al. 2009a; Trevaskis et al. 2007).
In contrast to the previous three vernalization genes, little is known about Vrn-4. The allele for early flowering was originally identified in the Australian cultivar Gabo (Knott 1959; Pugsley 1972), and was backcrossed into Triple Dirk to develop an isogenic line designated TDF (Pugsley 1972). This locus was assigned to chromosome 5D by monosomic analysis (Kato et al. 1993) and is currently designated as Vrn-D4 (formerly known as Vrn4 or Vrn-D5; McIntosh et al. 2003). This locus was later mapped closely linked to SSR marker Xgdm3 on the centromeric region of chromosome 5D (Kato et al. 2003). Natural variation for flowering time at the centromeric region of homoeologous group 5 chromosomes has been found, so far, only in the D genome. While some studies have questioned the existence of Vrn-D4 (Maystrenko 1980; Stelmakh 1987b) or its chromosome location (Goncharov 2003), abundant evidence is presented here supporting its 5D chromosome location.
Using genetic analyses, Iwaki et al. (2000, 2001) found the Vrn-D4 allele for spring growth habit in many spring wheat landraces from different parts of the world (55 out of 272), with a higher frequency in India and neighboring regions. Therefore, the Vrn-D4 locus appears to be an important contributor to variation in flowering time in the hexaploid wheat germplasm and the identification of the gene responsible for these differences may have practical applications in breeding. In addition, the identification of Vrn-4 is important to advance our understanding of the vernalization pathway in the temperate cereals, which appear to have evolved independently of the vernalization pathway in the dicot species (Yan et al. 2004b). The mapping results from this study represent an initial step toward the identification of this gene.
Materials and methods
Plant materials
Two different stocks of the near isogenic line Triple Dirk F (TDF) were used in this study (Table 1). The first one was obtained from Dr. T. Gotoh and was maintained at Okayama University, Japan (TDF-J, hereafter), and the second one was obtained from K. Campbell at Washington State University, USA (TDF-US, hereafter). TDF-J is the same line used by Kato et al. (2003) for the preliminary map of Vrn-D4. The Vrn-1 alleles present in each stock were determined using available molecular markers (Fu et al. 2005; Yan et al. 2004a).
Three populations were developed for the mapping of Vrn-4. The initial mapping populations included 144 F2 plants from the cross between TDF-J and Akakawaaka, a Japanese winter cultivar (Table 1). The limited level of polymorphism observed between the parental lines of this cross prompted the development of two additional populations. The second population included 258 F2 plants from the cross between the Japanese winter cultivar Hayakomugi (Table 1) and TDF-J. The third population (159 F2 plants) was developed from the cross between TDF-J and a substitution line of chromosome 5D from synthetic wheat 5402 in Chinese Spring, henceforth CS(5D5402) (Table 1). Synthetic RL5402 was generated by Dr. E. R. Kerber (Canada Agriculture Research Station, Winnipeg, Manitoba, Canada) from the cross between Tetra Cantach and Ae. tauschii (Kerber 1964). The CS(5D5402) line was developed by Dr. Jan Dvorak (University of California, Davis, USA), who kindly provided us the seeds. Synthetic 5402 was selected among the nine different synthetic lines characterized in the NSF-Wheat-SNP project (http://wheat.pw.usda.gov/SNP/new/index.shtml) because of its high level of polymorphisms with non-synthetic wheats.
The first two populations were analyzed in Japan, and flowering time was determined as the number of days from sowing to flag leaf unfolding. The third population and the interaction studies were performed in the US and flowering time was determined as number of days from sowing to heading. Progeny tests were conducted using F3 seeds to validate the genotyping of F2 plants with critical recombination events flanking the Vrn-D4 locus or with intermediate flowering times in the F2 generation.
Nulli-tetrasomic lines for chromosome 5D, ditelosomic line Dt5DL, and deletion lines for chromosome 5D with break point 5DS2, 5DS5, 5DS1, 5DL1, 5DL9, and 5DL5 were used to determine the arm location and physical position of the markers in the chromosome (Endo and Gill 1996; Linkiewicz et al. 2004; Sears and Steintz-Sears 1978). The TDF-J stock was compared with other Triple Dirk spring near isogenic lines (NILs) carrying the Vrn-A1 (TDD), Vrn-B1 (TDB), Vrn-D1 (TDE) and the winter NIL with recessive alleles for all the previous genes (TDC) (Table 1).
To study the interaction between Vrn-D4 alleles and vernalization, two F2 plants homozygous for the Vrn-D4 allele (TDF) and two homozygous for the vrn-D4 allele (Hayakomugi) were selected from the TDF × Hayakomugi segregating population. Ten F3 seeds from each plant were sown in individual pots (20 plants for each allele, total 40 plants). Half of the plants for each allele were vernalized for 6 weeks at 4°C and the other half were kept in a greenhouse at 20–25°C under the same photoperiod (16 h light). Heading times were recorded at the time of spike emergence.
Growth conditions
The F2 population from a cross between TDF-J and Akakawaaka was grown at constant temperature 20°C (non-vernalizing condition) and continuous light (24 h) in a growth chamber (LH-350SP, Nippon Medical & Chemical Instruments Co. Ltd., Japan). Light source was fluorescent lamps and photon flux density was ca. 160 μmol/m2/s. Planting density was one plant per 2.8 × 4.3 cm2 in a plastic tray (48 × 33 × 7 cm) filled with the 1:1 mixture of soil and bark compost.
The F2 population from a cross between TDF-J and Hayakomugi and their progeny F3 lines were grown in the same growth chamber using the same conditions as above except for the adjustment of the photoperiod to 16 h of light and 8 h of dark (long day), and planting density 2.8 × 5.9 cm2.
The F2 population from a cross between TDF-J and CS(5D5402) was grown in the greenhouse where air temperature was kept over 20–25°C (non-vernalizing condition) and photoperiod was 16 h. Light source in the day was natural daylight and at night incandescent lamps were used as supplementary light to extend photoperiod. Individual seeds were sown in soil-filled half-gallon pots.
To compare the vernalization response of TDF relative to other Triple Dirk NILs (Table 1), seeds were soaked in water at 4°C for 24 h and subsequently kept at 20°C for 24 h for germination. Six germinated seeds were planted for each of the eight treatments, which varied from 0 to 35 days at 2°C (5-day intervals, long days). After the vernalization treatments, plastic trays were transferred to the growth chambers under the same conditions as described above for the TDF-J × Akakawaaka mapping population until flag leaf unfolding. Plants for the non-vernalization control (0 days) were transferred to the growth chamber immediately after germination. Days from sowing to flag leaf unfolding were calculated as described before (Kato and Yamagata 1988). This method corrects for the slower growth at lower temperatures, so flowering time becomes approximately constant among fully vernalized plants irrespective of the duration of vernalization treatment.
Molecular markers and data analyses
Genomic DNA was extracted from young leaves of individual plants using the CTAB method (Murray and Thompson 1980). The Vrn-1 genotype of different TDF stocks was determined by PCR using primers described before (Fu et al. 2005; Yan et al. 2004a). Marker XBG313707 was developed from EST BG313707. D genome-specific primers BG313707_cpF1 (5′-GCTTCCAGACATCGGTCATT-3′) and BG313707D_R1 (5′-CACCACCAGTAACCCAGCC-3′) were used to sequence the critical recombinant lines and map a single nucleotide polymorphism (SNP).
Seven microsatellite markers, Xcfd81, Xcfd78, Xcfd67, Xgdm68, Xbarc205, Xwmc318, and Xgdm3, were used for genetic mapping (http://wheat.pw.usda.gov). PCR amplifications were performed in a 10 μl volume containing 1 μl of PCR buffer (10 mM Tris–HCl, pH 8.3, 50 mM KCl), 1.5 mM of MgCl2, 0.25 units of Taq polymerase (Sigma, USA), 0.2 mM of dNTP, 0.5 μM of primer, and 50 ng of template DNA.
PCR products for the Vrn-1 alleles were separated in 1.2% agarose gels, and those from the SSR markers were separated in 6–18% polyacrylamide gels. PCR products were visualized with ethidium bromide. PCR conditions for the different microsatellite markers included a 95°C denaturing step for 3 min, followed by 35 cycles of 95°C for 30 s, 58–60°C annealing (depending on microsatellite marker) for 30 s, and 72°C for 1 min, and a final extension step at 72°C for 10 min. Annealing temperatures for the different markers were as follows: 57°C for Xcfd78, Xcfd81, and Xgdm68; 60°C for Xcfd67, Xwmc318, XBG313707, and Xgdm3; 65°C for Xbarc205. Genetic maps were constructed using MAPMAKER/EXP3.0 (Lander et al. 1987).
Flowering data from the experiment to determine the interaction between vernalization and Vrn-D4 alleles were analyzed using a 2 × 2 factorial ANOVA. A logarithmic transformation was used to improve the adjustment of the data to the ANOVA assumptions. Statistical analyses were performed using SAS version 9.1 (SAS Institute Inc. 2006).
Real-time quantitative PCR (Q-PCR)
Total RNA was extracted using TRIZOL (Invitrogen, Carlsbad, CA, USA) and first-strand cDNA was synthesized using the SuperScript™ First-Strand Synthesis System (Invitrogen, Carlsbad, CA, USA). Q-PCR was performed on an ABI PRISM 7000 SDS (Applied Biosystems, Foster City, CA, USA) using SYBR® GREEN. PCR setup and reaction conditions were as reported before (Fu et al. 2007). The 2−ΔΔCT method (Livak and Schmittgen 2001) was used to normalize and calibrate transcript values relative to the wheat translation elongation factor 1 alpha-subunit (TEF1, primers 5′-GCCCTCCTTGCTTTCACTCT-3′ and 5′-AACGCGCCTTTGAGTACTTG-3′, 99% efficiency). The quantitative RT-PCR SYBR® GREEN systems for Vrn-1 (Yan et al. 2003), Vrn-2 (Distelfeld et al. 2009b), and Vrn-3 (Yan et al. 2006) genes have been published before.
Results
Differences between TDF stocks
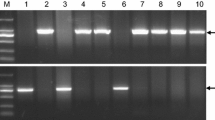
Molecular markers for the Vrn-A1 and Vrn-B1 loci (Fu et al. 2005; Yan et al. 2004a) were used to confirm previous genetic studies suggesting that the original cultivar Triple Dirk has dominant Vrn-A1 and Vrn-B1 alleles (Stelmakh 1987b). The same markers demonstrated that the TDF-J stock carries the expected recessive vrn-A1 and vrn-B1 alleles for winter growth habit (Fig. 1a, b), but also showed that the TDF-US stock carries the dominant Vrn-A1a allele (140-bp insertion in the promoter region) and the dominant Vrn-B1 allele (deletion in intron 1) (Fig. 1a, b). In addition, the TDF-US showed the same alleles for microsatellite markers Xgwm190, Xcfd81, and Xbarc205 in the Vrn-D4 region as TDC (same as original Triple Dirk cultivar), which were different in the TDF-J stock (Fig. 1c).
Heterogeneity of TDF stocks. a PCR analysis of the Vrn-A1 promoter. A 140 bp insertion is present in the Vrn-A1 allele in TDF-US, and absent in the vrn-A1 allele in TDF-J. b PCR analysis of Vrn-B1 first intron. The first lane in the gel shows DNA from TDF-J amplified with primers F and R4 (Fu et al. 2005) that detect the absence of the first intron deletion (vrn-B1 allele), and the second lane in the gel shows DNA from TDF-US amplified with the primers F and R3 (Fu et al. 2005) that detect the presence of the first intron deletion (Vrn-B1 allele). c The TDF stocks have different haplotypes for markers in the Vrn-D4 region. Three SSR markers Xgwm190, Xcfd81, and Xbarc205 showed polymorphisms between TDF-J and TDF-US and no polymorphism between TDF-US and TDC
The absence of Vrn-D4 in the TDF-US stock was confirmed in a population of 118 F2 plants from the cross between TDF-US × CS(5D5402). Segregation for flowering time was associated with the Vrn-A1 and Vrn-B1 regions but no differences in flowering time were associated with marker Xcfd67 from the Vrn-D4 region (data not shown).
Taken together, the previous results suggest that the TDF-US stock is not the original TDF stock described by Pugsley (1972) and is, more likely, a contamination with the original Triple Dirk stock. Therefore, the TDF-US stock was discarded and all further analyses were performed using the TDF-J stock.
Effect of the duration of the vernalization treatment on flowering time in different Triple Dirk NILs
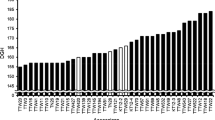
The comparison of the TDF-J with the Triple Dirk NILs for the Vrn-1 dominant alleles revealed differences in the residual effect of vernalization on these alleles for spring growth habit. In the absence of vernalization, the dominant Vrn-A1 allele (TDD) conferred the earliest flowering time and Vrn-D4 was intermediate between Vrn-B1 (TDB) and Vrn-D1 (TDE). A factorial ANOVA including NILs and vernalization treatments as factors showed significant differences among lines (P < 0.0001) and among vernalization treatments (P < 0.0001). The presence of a significant interaction between NILs and vernalization treatments (P < 0.0001) indicated that the different NILs respond in different ways to vernalization treatments of different durations. Pair-wise comparisons among the four isogenic stocks using the Tukey test revealed significant differences for all comparisons (P < 0.0001). Highly significant differences among NILs (P < 0.0001) were also detected in the eight separate ANOVAs for each of the vernalization treatments.
TDD (Vrn-A1) showed no acceleration of flowering time for any of the vernalization treatments and was the earliest to flower for the 0 and 5 days vernalization treatments (Fig. 2). The TDB (Vrn-B1), TDE (Vrn-D1), and TDF-J (Vrn-D4) stocks showed a small residual response to vernalization that was satisfied after 25 days of vernalization (Fig. 2). The difference for TDF-J (Vrn-D4) between the non-vernalized (0 days) and the average of the three saturating vernalization treatments (25, 30, and 35 days) was 1 day, but the difference was significant (P = 0.009). The acceleration of flowering in TDB (Vrn-B1) and TDE (Vrn-D1) was continuous from 5 to 25 vernalization days, but in TDF-J (Vrn-D4) no acceleration of flowering time was observed for the shorter vernalization treatments (5 and 10 days, Fig. 2). Although the profiles for TDF-J (Vrn-D4) and TDD (Vrn-A1) were similar for the 0, 5, 10, and 15 days, Vrn-D4 was approximately 3 days later than Vrn-A1 for each of these treatments. These results suggest that the response of Vrn-D4 to vernalization might be different from the one observed for the dominant Vrn-1 alleles.
Vrn-D4 mapping
The 144 F2 plants from the cross TDF-J × Akakawaaka segregated into 111 spring-type plants and 33 winter-type plants (Fig. 3a), which fits a 3:1 ratio for a single dominant gene segregation (χ2 = 0.33, P = 0.56). In this population, molecular marker Xcfd67 was found to cosegregate with the differences in flowering time (Fig. 3a). However, the low level of polymorphisms found between TDF-J and Akakawaaka precluded the development of a genetic map using this population.
Frequency distribution of the days from sowing to flag leaf unfolding (a and b) or ear emergence (c) in the F2 populations derived from the crosses between TDF-J and a Akakawaaka, b Hayakomugi, and c CS(5D5402). Plants were grown under a non-vernalizing conditions (20°C) and long day photoperiod (a 24 h; b, c 16 h light). Plants were classified by their Xcfd67 genotype (Xcfd67 is linked to Vrn-D4) as follows: gray rectangles correspond to plants homozygous for TDF-J allele, striped rectangles to heterozygous plants, and white rectangles to plants homozygous for the Xcfd67 allele from the other parent
A screen of additional Japanese winter cultivars showed that Hayakomugi was more polymorphic with TDF-J than Akakawaaka. Six microsatellite markers and an EST-derived marker were polymorphic in this TDF-J × Hayakomugi population. The frequency distribution of flowering times in this population was bimodal, but with a small overlap between Xcfd67 classes (Fig. 3b). F3 seeds from the F2 plants with flowering times within the overlapping region as well as from some F2 plants with critical recombination events in the Vrn-D4 region were selected to perform progeny tests and provide a more accurate estimate of the original F2 plants phenotype. All the plants with recombination events between flanking markers Xcfd78 and Xbarc205 showed clear flowering phenotypes (either in the F2 or in the F3 progeny tests), which facilitated a precise mapping of the Vrn-D4 gene within this interval completely linked to Xcfd67. Using these additional data, the plants from this population were classified into 186 spring-type plants and 72 winter-type plants (Fig. 3b). This segregation fits a 3:1 ratio for a single dominant gene segregation (χ2 = 1.16, P = 0.28).
The seven polymorphic markers were confirmed to be from chromosome 5D using the nulli-tetrasomic line missing that chromosome, and were assigned to different chromosome bins as described in Fig. 4a, b. Since Vrn-D4 was completely linked with long arm marker Xcfd67 and short arm marker XBG313707, it was not possible to establish its chromosome arm location. The three linked markers were mapped within a 1.8 cM region flanked by Xcfd78 in the short arm and Xbarc205 in the long arm (Fig. 4c).
Physical and genetic mapping of Vrn-D4. a Example of physical mapping of microsatellite markers Xcfd78 and Xcfd67 using cytogenetic stocks N5D (nulli-tetrasomic line missing chromosome 5D), Dt5DL (ditelosomic line missing the 5DS arm), and 5DS2 to 5DL5 (deletion lines for the short and long arm). b Assignment of markers to chromosome bins. The numbers within each bin indicate the colinear rice chromosome. c Genetic maps of Vrn-D4 relative to molecular markers in the populations from the crosses TDF-J × Hayakomugi and TDF-J × CS(5D5402)
In TDF-J × CS(5D5402) population, there was a clear association between the marker classes and flowering time, with a small number of ambiguous plants (Fig. 3c). However, since all the plants with recombination events between Xcfd81 and Xbarc143 showed unambiguous flowering phenotypes in the F2 or F3 progeny tests, it was possible to map the Vrn-D4 completely linked to markers Xcfd67, XBG313707, Xgdm68, Xbarc205, and Xgdm3 (Fig. 4c). If the genotype of the few plants with intermediate flowering times (and no recombination between flanking markers) is inferred based on the genotype of the Vrn-D4 flanking markers, the 159 F2 plants from the cross TDF-J × CS(5D5402) can be classified into 124 spring-type plants and 35 winter-type plants, which fits a 3:1 segregation ratio for a single dominant gene segregation (χ2 = 0.76, P = 0.38).
All markers that were polymorphic in the TDF-J × Hayakomugi population were also polymorphic in this population and were mapped. In addition, microsatellite marker Xgdm68 not mapped on the previous population was added to this map. The TDF-J × CS(5D5402) population showed lower levels of recombination than the TDF-J × Hayakomugi population, which was reflected in smaller genetic distances (52% reduction) and lower resolution of the markers in the centromeric region. In this population, Vrn-D4 was mapped completely linked to five molecular markers flanked by Xcfd78 in the short arm and Xbarc143 in the long arm (Fig. 4c).
Interaction between Vrn-D4 alleles and vernalization
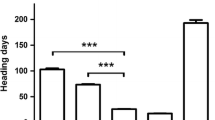
A separate experiment using selected F3 plants from the TDF-J × Hayakomugi population demonstrated significant interactions for flowering time between the Vrn-D4 alleles and the presence or absence of vernalization treatment (2 × 2 factorial ANOVA, P < 0.0001). Significantly larger differences in heading time between Vrn-D4 alleles were detected among unvernalized plants (35 days) than among vernalized plants (10 days). These data confirmed that vernalization modulates the effect of the Vrn-D4 alleles on flowering time.
To see how other vernalization genes were affected by the Vrn-D4 alleles, transcript levels of Vrn-1, Vrn-2, and Vrn-3 were compared between TDF-J (Vrn-D4 allele for spring growth habit) and CS(5D5402) (vrn-D4 allele for winter growth habit) using quantitative RT-PCR. Plants from the two lines were sown at the same time in a greenhouse at non-vernalizing temperatures (20–25°C) under long day conditions (samples were taken at noon). At the time of leaf sample collection for RNA extraction, TDF-J plants were heading and CS(5D5402) plants were still at the vegetative stage. At this stage, TDF-J leaves showed higher transcript levels of the flowering promoting genes Vrn-1 (>4,000-fold increase, P = 0.0002) and Vrn-3 (>60,000-fold increase, P = 0.006) and reduced levels of the flowering repressor Vrn-2 (>80-fold reduction, P = 0.012) than CS(5D5402) (Table 2).
Discussion
Heterogeneity in TDF stocks
Pugsley (1972) identified the spring growth habit gene Vrn-D4 in the cultivar Gabo and showed that it was not allelic to any of the Vrn-1 homoeologs. Gabo was a major cultivar in Australia from the late 1940s to the late 1960s (O’Brien et al. 2001). Gabo’s pedigree includes the Indian cultivar Muzaffar Nagar. Early-maturing forms were introduced from India to avoid rust and drought in Australian breeding programs (Lupton 1987). Since the allelic frequency of Vrn-D4 is relatively high in India compared with other regions (Iwaki et al. 2000, 2001), it was assumed that Muzaffar Nagar might have been Gabo’s donor of Vrn-D4. This hypothesis still needs experimental confirmation.
The Vrn-D4 allele for early flowering from Gabo was transferred by Pugsley (1972) to Triple Dirk C by backcrossing. The resulting line with the dominant Vrn-D4 allele and recessive alleles at all the other vernalization genes was designated TDF. However, several studies have questioned the existence of Vrn-D4 or its chromosome location. Maystrenko (1980) suggested that Gabo has both Vrn-B1 and Vrn-D4 but erroneously assigned them to chromosomes 2B and 5B, respectively. Stelmakh (1987b) initially suggested that TDF and Gabo have both Vrn-A1 and Vrn-B1 but not Vrn-D4. This allelic combination is the same we found in the TDF-US stock and suggests the possibility that Stelmakh used a similar incorrect TDF stock. In his paper, Stelmakh mentioned that the seeds of TDF and Gabo he used were directly provided by Pugsley in 1981 and 1974, respectively. Later, Stelmakh (1998) conducted additional genetic analysis using populations from the cross between a TDF stock from Japan and Vrn-1 tester lines and concluded that TDF-J has Vrn-D4, but that the TDF selection Y used in his 1987 paper had the vrn-D4 allele for winter growth habit.
Gotoh (1979) conducted genetic analyses using a TDF stock provided to him by Pugsley before 1976 and confirmed the existence of Vrn-D4 as a different gene, not allelic to any of the Vrn-1 homoeologs. Kato et al. (2003) confirmed that Vrn-D4 was linked to molecular marker Xgdm3 in the centromeric region of chromosome 5D, and more than 50 cM proximal from the location of the Vrn-D1 locus in the middle of the long arm (Kato et al. 2003). Goncharov (2003) used the same TDF-J and confirmed the existence of Vrn-D4 in TDF and Gabo, although he failed to detect the 5D chromosome location, possibly because of a problem in his monosomic tester line Bersée mono 5D.
In summary, there seems to be some heterogeneity among different TDF stocks, which might be caused by contamination of the TDF seeds by the original Triple Dirk variety. The incorrect TDF stocks can now be readily identified using available molecular markers for Vrn-A1 (Yan et al. 2004a) and Vrn-B1 (Fu et al. 2005).
Vrn-D4 mapping
In this study, the Vrn-D4 gene was mapped in the centromeric region of chromosome 5D, which was consistent with preliminary mapping data generated by Kato et al. (2003). The collinear region in rice chromosome 12 includes several flowering QTLs (Mei et al. 2003; Nagata et al. 2002; Septiningsih et al. 2003; Uga et al. 2007). However, it is currently not possible to determine whether Vrn-D4 corresponds to any of these rice QTL, because the arm location of Vrn-D4 in wheat is not yet known, and therefore, the colinear region in rice chromosome 12 is too large. We are currently expanding the mapping population to generate additional recombination events to delimit better the chromosome location of Vrn-D4 in wheat and its collinear region in rice. Additional sequenced-based markers (such as BG313707) will also be necessary to establish a better correspondence between the two regions.
In the TDF-J × CS(5D5402) population, genetic distances were 2.5-fold smaller than in the TDF-J × Hayakomugi population (Fig. 4). This might be attributed to the high level of polymorphisms detected between chromosomes 5D from CS(5D5402) and from hexaploid wheat. These results are in agreement with previous studies that showed a lower chiasma formation at metaphase I between homologous chromosomes from divergent varieties compared with identical chromosomes from the same variety (Dvorak and McGuire 1981). Particularly relevant to this study is the significant decrease in chromosome pairing detected between chromosome 5D from Chinese Spring and chromosome 5D from Ae. tauschii in a Chinese Spring genetic background relative to the pairing of identical 5D chromosomes (Dvorak 1988).
In summary, a combination of multiple mapping populations, one maximizing recombination and the other one maximizing polymorphisms, seems to be the best strategy to accelerate the development of a high density map of the Vrn-D4 gene. The TDF-J × CS(5D5402) population can be used first to select the closest markers to Vrn-D4, and then, the efforts to find polymorphisms in the TDF-J × Hayakomugi population can be focused in a reduced number of selected markers.
Effect of Vrn-D4 on vernalization response
This study has confirmed the existence of a single locus for early flowering in all three crossing populations between TDF-J and winter lines, and demonstrated that the effect of this gene on flowering time is modulated by vernalization requirement. The significant interaction detected between Vrn-D4 alleles and vernalization is a hallmark of genes that are part of the vernalization pathway. The higher transcript levels of Vrn-1 and Vrn-3 and lower transcript level of Vrn-2 in TDF-J (Vrn-D4 allele) relative to CS(5D5402) (vrn-D4 allele) planted at the same time suggest that Vrn-D4 acts upstream (or is part of) the feedback regulatory loop formed by Vrn-1, Vrn-2, and Vrn-3 (Distelfeld et al. 2009a).
The comparison of the vernalization response of the different Triple Dirk NILs showed that the Vrn-D4 allele for spring growth habit has a residual vernalization response, a phenomenon also observed for the Vrn-B1 and Vrn-D1 alleles, both here and in previous studies (Berry et al. 1980; Pugsley 1972). However, the responses of these two last genes differed slightly from the one observed for Vrn-D4, particularly for plants exposed to short vernalization periods (5–10 days). Flowering in plants carrying the Vrn-B1 and Vrn-D1 alleles was accelerated by 5–10 days exposures to cold temperatures, but no acceleration was detected for Vrn-D4 for similar treatments. These differences may reflect separate roles of these genes in the vernalization pathway, but a final answer to this question will require the cloning of the Vrn-D4 gene.
Spring growth habit gene Vrn-D4 for wheat improvement
Vrn-D4 has not been extensively used in spring wheat breeding programs in North America, Europe, and East Asia including Japan (Goncharov 1998; Gotoh 1979). In Europe and North America, Vrn-A1 and Vrn-B1 are predominant, while in Asia, especially in Japan, Vrn-D1 is frequently found (Goncharov 1998; Gotoh 1979; Stelmakh 1987a). The Vrn-D1 allele is frequent in fall-planted spring wheats, whereas the stronger Vrn-A1 allele is present in high frequency among spring-planted spring varieties (Fu et al. 2005; Iqbal et al. 2007; Iwaki et al. 2000, 2001; Zhang et al. 2008).
Seki et al. (2007) analyzed the effects of Vrn genes on the timing of transition to adult phase using Abukumawase NILs and found that in fully vernalized plants grown in the field the NILs with the Vrn-D4 and Vrn-A1 alleles were earlier than those with the Vrn-B1 and Vrn-D1 alleles. In addition, the results presented here suggest that the Vrn-D4 gene differs from Vrn-B1 and Vrn-D1 in its response to short cold intervals (Fig. 2). A strong Vrn-D4 allele has been reported in the Italian cultivar Mara (Worland et al. 1987), which suggests that there might be multiple alleles of Vrn-D4 with different effects on flowering time. In summary, these results indicate that the Vrn-D4 gene might be useful for fine tuning heading time and vernalization requirement in hexaploid wheat.
References
Berry GJ, Salisbury PA, Halloran GM (1980) Expression of vernalization genes in near-isogenic wheat lines—duration of vernalization period. Ann Bot 46:235–241
Bonnin I, Rousset M, Madur D, Sourdille P, Dupuits C, Brunel D, Goldringer I (2008) FT genome A and D polymorphisms are associated with the variation of earliness components in hexaploid wheat. Theor Appl Genet 116:383–394
Corbesier L, Vincent C, Jang SH, Fornara F, Fan QZ, Searle I, Giakountis A, Farrona S, Gissot L, Turnbull C, Coupland G (2007) FT protein movement contributes to long-distance signaling in floral induction of Arabidopsis. Science 316:1030–1033
Distelfeld A, Li C, Dubcovsky J (2009a) Regulation of flowering in temperate cereals. Curr Opin Plant Biol 12:178–184
Distelfeld A, Tranquilli G, Li C, Yan L, Dubcovsky J (2009b) Genetic and molecular characterization of the VRN2 loci in tetraploid wheat. Plant Physiol 149:245–257
Dubcovsky J, Chen C, Yan L (2005) Molecular characterization of the allelic variation at the VRN-H2 vernalization locus in barley. Mol Breed 15:395–407
Dvorak J (1988) Cytogenetical and molecular inferences about the evolution of wheat. In: Miller TE, Koebner RMD (eds) Proceedings of 7th international wheat genetics symposium, Cambridge, pp 187–192
Dvorak J, McGuire PE (1981) Nonstructural chromosome differentiation among wheat cultivars with special reference to differentiation of chromosomes in related species. Genetics 97:391–414
Endo TR, Gill BS (1996) The deletion stocks of common wheat. J Hered 87:295–307
Flood RG, Halloran GM (1986) Genetics and physiology of vernalization response in wheat. Adv Agron 39:87–125
Fu D, Szűcs P, Yan L, Helguera M, Skinner J, Hayes P, Dubcovsky J (2005) Large deletions in the first intron of the VRN-1 vernalization gene are associated with spring growth habit in barley and polyploid wheat. Mol Genet Genomics 273:54–65
Fu D, Dunbar M, Dubcovsky J (2007) Wheat VIN3-like PHD finger genes are up-regulated by vernalization. Mol Genet Genomics 277:301–313
Goncharov NP (1998) Genetic resources of wheat related species: the Vrn genes controlling growth habit (spring vs. winter). Euphytica 100:371–376
Goncharov NP (2003) Genetics of growth habit (spring vs. winter) in common wheat: confirmation of the existence of dominant gene Vrn4. Theor Appl Genet 107:768–772
Gotoh T (1979) Genetic studies on growth habit of some important spring wheat cultivars in Japan, with special reference to the identification of the spring genes involved. Japan J Breed 29:133–145
Hemming MN, Peacock WJ, Dennis ES, Trevaskis B (2008) Low-temperature and daylength cues are integrated to regulate FLOWERING LOCUS T in barley. Plant Physiol 147:355–366
Hemming MN, Fieg S, Peacock WJ, Dennis ES, Trevaskis B (2009) Regions associated with repression of the barley (Hordeum vulgare) VERNALIZATION1 gene are not required for cold induction. Mol Genet Genomics 282:107–117
Iqbal M, Navabi A, Yang RC, Salmon DF, Spaner D (2007) Molecular characterization of vernalization response genes in Canadian spring wheat. Genome 50:511–516
Iwaki K, Nakagawa K, Kuno H, Kato K (2000) Ecogeographical differentiation in East Asian wheat, revealed from the geographical variation of growth habit and Vrn genotype. Euphytica 111:137–143
Iwaki K, Haruna S, Niwa T, Kato K (2001) Adaptation and ecological differentiation in wheat with special reference to geographical variation of growth habit and Vrn genotype. Pl Breed 120:107–114
Kato K, Yamagata H (1988) Method for evaluation of chilling requirement and narrow-sense earliness of wheat cultivars. Japan J Breed 38:172–186
Kato K, Nakagawa K, Kuno H (1993) Chromosomal location of the vernalization response, Vrn2 and Vrn4, in common wheat, Triticum aestivum L. Wheat Inform Serv 76:53
Kato K, Yamashita M, Ishimoto K, Yoshino H, Fujita M (2003) Genetic analysis of two genes for vernalization response, the former Vrn2 and Vrn4, using PCR based molecular markers. In: Pogna NE, Romano N, Pogna EA, Galterio G (eds) Proceeding of 10th international wheat genetics symposium. Inst Sperimentale per la Cerealcolture, Paestum, Italy, pp 971–973
Kerber ER (1964) Wheat—reconstitution of tetraploid component (AABB) of hexaploids. Science 143:253–255
Knott DR (1959) The inheritance of rust resistance. IV. Monosomic analysis of rust resistance and some other characters in six varieties of wheat including Gabo and Kenya Farmer. Can J Plant Sci 39:215–228
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Li C, Dubcovsky J (2008) Wheat FT protein regulates VRN1 transcription through interactions with FDL2. Plant J 55:543–554
Lin MK, Belanger H, Lee YJ, Varkonyi-Gasic E, Taoka KI, Miura E, Xoconostle-Cazares B, Gendler K, Jorgensene RA, Phinney B, Lough TJ, Lucas WJ (2007) FLOWERING LOCUS T protein may act as the long-distance florigenic signal in the cucurbits. Plant Cell 19:1488–1506
Linkiewicz AM, Qi LL, Gill BS, Ratnasiri A, Echalier B, Chao S, Lazo GR, Hummel DD, Anderson OD, Akhunov ED, Dvorak J, Pathan MS, Nguyen HT, Peng JH, Lapitan NLV, Miftahudin, Gustafson JP, La Rota CM, Sorrells ME, Hossain KG, Kalavacharla V, Kianian SF, Sandhu D, Bondareva SN, Gill KS, Conley EJ, Anderson JA, Fenton RD, Close TJ, McGuire PE, Qualset CO, Dubcovsky J (2004) A 2500-locus bin map of wheat homoeologous group 5 provides insights on gene distribution and colinearity with rice. Genetics 168:665–676
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Lupton FGH (1987) Wheat breeding in Australia. In: Lupton FGH (ed) Wheat breeding. Chapman and Hall, London, pp 63–64
Maystrenko OI (1980) Cytogenetic study of the growth habit and ear emergence time in wheat (Triticum aestivum L.). In: Balyae DK (ed) Proceedings of 14th international congress of genetics, MIR Publishers, Moscow, pp 267–282
McIntosh RA, Yamazaki Y, Devos KM, Dubcovsky J, Rogers WJ, Appels R (2003) Catalogue of Gene Symbols for Wheat. In: Pogna NE, Romano M, Pogna E, Galterio G (eds) Proceedings of 10th international wheat genetics symposium. Inst Sperimentale per la Cerealicoltura, Rome, Paestum, Italy. http://wheat.pw.usda.gov/ggpages/wgc/2000upd.html
Mei HW, Luo LJ, Ying CS, Wang YP, Yu XQ, Guo LB, Paterson AH, Li ZK (2003) Gene actions of QTLs affecting several agronomic traits resolved in a recombinant inbred rice population and two testcross populations. Theor Appl Genet 107:89–101
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucl Acids Res 8:4321–4325
Nagata K, Shimizu H, Terao T (2002) Quantitative trait loci for non-structural carbohydrate accumulation in leaf sheaths and culms of rice (Oryza sativa L.) and their effects on grain filling. Breed Sci 52:275–283
O’Brien L, Morell M, Wrigley C, Appels R (2001) Genetic pool of Australian wheats. In: Bonjean AP, Angus WJ (eds) The world wheat book. Lavoisier, Paris, pp 611–648
Oliver SN, Finnegan EJ, Dennis ES, Peacock WJ, Trevaskis B (2009) Vernalization-induced flowering in cereals is associated with changes in histone methylation at the VERNALIZATION1 gene. Proc Natl Acad Sci USA 106:8386–8391
Pugsley AT (1972) Additional genes inhibiting winter habit in wheat. Euphytica 21:547–552
SAS Institute Inc. (2006) SAS user’s guide, version 9.1. SAS Institute, Inc., Cary
Sears EM, Steintz-Sears LM (1978) The telocentric chromosomes of common wheat. In: Ramanujam S (ed) Proceedings of 5th international wheat genetics symposium, New Delhi, India, pp 389–407
Seki MO, Matsunaka S, Hatta H, Fujita K, Hatano M, Kiribuchi Otobe T, Kawada C, Kato N (2007) Growth and yield of near-isogenic wheat (Triticum aestivum) lines carrying different vernalization response genes. Breed Res 9:125–133
Septiningsih EM, Prasetiyono J, Lubis E, Tai TH, Tjubaryat T, Moeljopawiro S, McCouch SR (2003) Identification of quantitative trait loci for yield and yield components in an advanced backcross population derived from the Oryza sativa variety IR64 and the wild relative O rufipogon. Theor Appl Genet 107:1419–1432
Shitsukawa N, Ikari C, Shimada S, Kitagawa S, Sakamoto K, Saito H, Ryuto H, Fukunishi N, Abe T, Takumi S, Nasuda S, Murai K (2007) The einkorn wheat (Triticum monococcum) mutant, maintained vegetative phase, is caused by a deletion in the VRN1 gene. Genes Genet Syst 82:167–170
Stelmakh AF (1987a) Genetic effects of the Vrn 1–3 loci and specific action of the dominant Vrn-D1 allele in common bread wheat. Physiol Genet 21:278–286
Stelmakh AF (1987b) Growth habit in common wheat (Triticum aestivum L EM. Thell. Euphytica 36:513–519
Stelmakh AF (1998) Genetic systems regulating flowering response in wheat. Euphytica 100:359–369
Tamaki S, Matsuo S, Wong HL, Yokoi S, Shimamoto K (2007) Hd3a protein is a mobile flowering signal in rice. Science 316:1033–1036
Trevaskis B, Hemming MN, Dennis ES, Peacock WJ (2007) The molecular basis of vernalization-induced flowering in cereals. Trends Plant Sci 12:352–357
Uga Y, Nonoue Y, Liang ZW, Lin HX, Yamamoto S, Yamanouchi U, Yano M (2007) Accumulation of additive effects generates a strong photoperiod sensitivity in the extremely late-heading rice cultivar ‘Nona Bokra’. Theor Appl Genet 114:1457–1466
Worland AJ, Gale MD, Law CN (1987) Wheat genetics. In: Lupton FGH (ed) Wheat breeding. Chapman and Hall, London, pp 129–171
Yan L, Loukoianov A, Tranquilli G, Helguera M, Fahima T, Dubcovsky J (2003) Positional cloning of wheat vernalization gene VRN1. Proc Natl Acad Sci USA 100:6263–6268
Yan L, Helguera M, Kato K, Fukuyama S, Sherman J, Dubcovsky J (2004a) Allelic variation at the VRN-1 promoter region in polyploid wheat. Theor Appl Genet 109:1677–1686
Yan L, Loukoianov A, Blechl A, Tranquilli G, Ramakrishna W, SanMiguel P, Bennetzen JL, Echenique V, Dubcovsky J (2004b) The wheat VRN2 gene is a flowering repressor down-regulated by vernalization. Science 303:1640–1644
Yan L, Fu D, Li C, Blechl A, Tranquilli G, Bonafede M, Sanchez A, Valarik M, Dubcovsky J (2006) The wheat and barley vernalization gene VRN3 is an orthologue of FT. Proc Natl Acad Sci USA 103:19581–19586
Zhang XK, Xia XC, Xiao YG, Dubcovsky J, He ZH (2008) Allelic variation at the vernalization genes Vrn-A1, Vrn-B1, Vrn-D1 and Vrn-B3 in Chinese common wheat cultivars and their association with growth habit. Crop Sci 48:458–470
Acknowledgments
This work was supported by the United States Department of Agriculture Cooperative State Research, Education, and Extension Services National Research Initiative competitive grants 2007-35301-17737 and 2007-35301-18188 and Grant-in-Aids for Research Programs of Wheat and Barley Production from the Ministry of Agriculture, Forestry and Fisheries of Japan and for Young Scientists (B) (20780002) from Japan Society for the Promotion of Science (JSPS).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P. Langridge.
T. Yoshida and H. Nishida contributed equally to the work.
Rights and permissions
About this article
Cite this article
Yoshida, T., Nishida, H., Zhu, J. et al. Vrn-D4 is a vernalization gene located on the centromeric region of chromosome 5D in hexaploid wheat. Theor Appl Genet 120, 543–552 (2010). https://doi.org/10.1007/s00122-009-1174-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-009-1174-3