Abstract
The structure and function of a cadaverine–lysine antiporter CadB and a putrescine–ornithine antiporter PotE in Escherichia coli were evaluated using model structures based on the crystal structure of AdiC, an agmatine–arginine antiporter, and the activities of various CadB and PotE mutants. The central cavity of CadB, containing the substrate binding site, was wider than that of PotE, mirroring the different sizes of cadaverine and putrescine. The size of the central cavity of CadB and PotE was dependent on the angle of transmembrane helix 6 (TM6) against the periplasm. Tyr73, Tyr89, Tyr90, Glu204, Tyr235, Asp303, and Tyr423 of CadB, and Cys62, Trp201, Glu207, Trp292, and Tyr425 of PotE were strongly involved in the antiport activities. In addition, Trp43, Tyr57, Tyr107, Tyr366, and Tyr368 of CadB were involved preferentially in cadaverine uptake at neutral pH, while only Tyr90 of PotE was involved preferentially in putrescine uptake. The results indicate that the central cavity of CadB consists of TMs 2, 3, 6, 7, 8, and 10, and that of PotE consists of TMs 2, 3, 6, and 8. These results also suggest that several amino acid residues are necessary for recognition of cadaverine in the periplasm because the level of cadaverine is much lower than that of putrescine in the periplasm at neutral pH. All the amino acid residues identified as being strongly involved in both the antiport and uptake activities were located on the surface of the transport path consisting of the central cavity and TM12.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Polyamines are necessary for normal cell growth of bacteria at both neutral and acidic pH, and polyamine levels are elaborately regulated by biosynthesis, degradation, and transport (Igarashi and Kashiwagi 2010b). As for polyamine transport systems in Escherichia coli, there are two polyamine uptake systems, PotABCD, a spermidine-preferential uptake system, and PotFGHI, a putrescine-specific uptake system, which belong to ATP-binding cassette transporters (Igarashi and Kashiwagi 1996, 1999, 2010a). It is also shown that spermidine excretion protein complex, MdtJI, and another putrescine uptake protein PuuP during the utilization of putrescine as energy source exist in E. coli (Higashi et al. 2008; Kurihara et al. 2009).
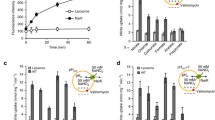
Furthermore, unique transporters such as CadB, a cadaverine–lysine antiporter (Soksawatmaekhin et al. 2004, 2006), PotE, a putrescine–ornithine antiporter (Kashiwagi et al. 1992, 1997, 2000), and AdiC, an agmatine–arginine antiporter (Gong et al. 2003; Iyer et al. 2003), which belong to the amino acid/polyamine/organocation (APC) superfamily of transporters (Jack et al. 2000), are necessary for cell growth of Escherichia coli at acidic pH (Fig. 1). These transporters function as electrogenic antiporters, and increase the pH in medium and nucleotide biosynthesis by producing CO2, together with the respective amino acid decarboxylases (Takayama et al. 1994; Soksawatmaekhin et al. 2004). So, cadB is transcribed together with cadA encoding the inducible lysine decarboxylase (Meng and Bennett 1992; Watson et al. 1992), and potE is transcribed together with speF encoding the inducible ornithine decarboxylase (Kashiwagi et al. 1991). In the case of adi genes, adiC and adiA encoding inducible arginine decarboxylase are coordinately regulated but independently transcribed (Gong et al. 2003). CadB and PotE also catalyze the proton motive force-dependent uptake of cadaverine and putrescine at neutral pH, as indicated by the fact that the potE gene in a medium-copy number vector was initially identified as a gene for putrescine transporter at neutral pH (Kashiwagi et al. 1990; Soksawatmaekhin et al. 2004).
Physiological functions of AdiC, CadB and PotE in E. coli. In acidic conditions, the three proteins function as electrogenic diamine-amino acid antiporters, and pH in the medium is increased by excretion of the diamines (Soksawatmaekhin et al. 2004). Amino acid decarboxylases generates a pH gradient by consuming a cytoplasmic proton. This process causes the increase in the level of ATP in cells (Soksawatmaekhin et al. 2004)
Recently, the crystal structures of AdiC were reported at 3.6 Å resolution (Gao et al. 2009) and 3.2 Å resolution (Fang et al. 2009). Furthermore, the crystal structure of the AdiC-arginine complex was reported at 3.0 Å resolution (Gao et al. 2010). Thus, the structural models of CadB and PotE were constructed with SWISS-MODEL (Arnold et al. 2006; Bordoli et al. 2009) using the ternary structure of AdiC as a template. Correlations and comparisons of structure and function for CadB and PotE were made based on the model and on the activities of various mutants of CadB and PotE.
Comparison of structures of CadB and PotE
Based on the structure of AdiC, models of the ternary structure of CadB and PotE were constructed (Fig. 2). The overall sizes (vertical size × horizontal size vs membrane) of CadB and PotE were 60 × 40 and 56 × 40 Å, respectively. The most significant structural difference among AdiC, CadB, and PotE was observed in transmembrane helix 6a (TM6a)—specifically, angle of TM6a against the central cavity constituting the substrate binding site was different in three transporters. The difference in the angle of TM6a between CadB and PotE was small, but there was a marked difference in the position of TM6a relative to the other TM domain in CadB and PotE (Fig. 2). This was due to a structural difference at the region of variable loop between TM5 and TM6 and the length of TM6a and TM6b (Fig. 3). In addition, the relative position of TM2 in AdiC was different from the position of TM2 in CadB and PotE (Fig. 2). This was due to the difference in the size and position of the variable loop between TM1 and TM2 (Fig. 3). The difference of the relative position of TM2 in CadB and PotE was also small, but it was caused by difference in the size of TM1a and TM1b in CadB and PotE (Fig. 3). Another difference between AdiC and CadB, and PotE was observed in the variable loop between TM7 and TM8 (Figs. 2, 3). Thus, the relative position of TM7 and TM8 in AdiC was different from that in CadB and PotE. Since the variable loop between TM8 and TM9 is different in CadB and PotE, the relative position of TM8 and TM9 in CadB and PotE may be different. Such a difference of TM8 and TM9 in CadB and PotE may also influence the structure of the adjacent TMs, i.e., TM7 and TM10. Taken together, the structural difference between CadB and PotE was mainly due to the difference of TM6, the loop structure between TM5 and TM6, and TM1 (Figs. 2, 3).
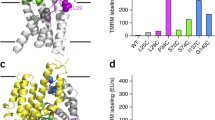
Structural model of AdiC, CadB, and PotE. Model structure of CadB and PotE was constructed using AdiC as a template (Gao et al. 2009) using SWISS-MODEL Workspace, a web-based integrated service dedicated to protein structure homology modeling (http://swissmodel.expasy.org/) (Arnold et al. 2006; Bordoli et al. 2009). The structures were visualized using PyMOL viewer v 0.99 (http://www.pymol.org). TMs shown with a black number in a white square constitute the central cavity
Amino acid alignment of AdiC, CadB, and PotE. Sequence alignment was performed using CLUSTAL-W version 1.83 (http://clustalw.ddbj.nig.ac.jp/top-j.html). TM is shown by a black box with white lettering, and α-helix on a loop by a grey box with white lettering
Although the overall ternary structures of CadB and PotE are similar, there is a clear difference in the central cavity, where substrates are recognized. The size of the central cavity was bigger in CadB than in PotE, which reflects the size of the substrate, cadaverine (7.4 Å) and putrescine (6.2 Å) (Fig. 4). This was largely due to the differences in the size and the angle of TM6a. The largest width of the central cavity of CadB was 27 Å, while that of PotE was 23 Å. Thus it is expected that the substrates (cadaverine and putrescine) could easily access the central cavity.
Conformational difference of the central cavity between CadB and PotE. The structure of the central cavity of CadB and PotE from the extracellular side and its oblique view was constructed as described in the legend of Fig. 2 and shown together with the size of cadaverine and putrescine. TMs shown with a black number in a white square constitute the central cavity. Differences of angle between TM10 and TM6a of CadB and PotE were emphasized in the right figures of the central cavities. Surface structure of CadB and PotE was visualized using PyMOL viewer v. 0.99 (http://www.pymol.org)
Alignment of functional amino acid residues at the central cavity of CadB and PotE
We have previously identified functional amino acids which are involved in the activities of CadB and PotE (Kashiwagi et al. 1997, 2000; Soksawatmaekhin et al. 2006). These amino acid residues are aligned on the newly constructed secondary structure of CadB and PotE based on the model ternary structure (Fig. 5). Since Tyr73, Tyr89, Tyr90, Glu204, Tyr235, Asp303, and Tyr423 of CadB, and Cys62, Trp201, Glu207, Trp292, and Tyr425 of PotE were strongly, and Tyr55, Glu76, Tyr246, Tyr310, Cys370, and Glu377 of CadB, and Glu77, Tyr92, Cys210, Cys285, Cys286, and Glu433 of PotE were moderately involved in the antiport activities together with uptake activities, it is thought that the transport path of CadB consists of TMs 2, 3, 6, 7, 8, 10, and 12, and that of PotE consists of TMs 2, 3, 6, 8, and 12 (Figs. 5, 6). The results confirm that the central cavity is bigger in CadB than in PotE. Since the functional amino acid residues were not found on TM1 of CadB and PotE, the structural difference of TM1 in CadB and PotE is thought to influence the relative position of TM2 and TM3 in CadB and PotE. It is also thought that TM12 of CadB and PotE constitutes the entrance of the transport path of cadaverine and putrescine from the cytoplasm.
Amino acid residues involved in the activities of CadB and PotE. A model of the secondary structure of proteins was constructed according to the ternary structure model shown in Fig. 2. Putative transmembrane segments are shown in large boxes. Amino acid residues involved in both uptake and antiport activities, diamine uptake activity, and antiport activity only are classified by symbols shown in the figure. The large and small circles indicate strong and moderate involvement in the activities, respectively. Mutated amino acids with no effect are shown in small grey boxes
Amino acid residues involved in transport activity located on the surface of transport path of CadB and PotE. Amino acids strongly involved in both uptake and antiport activities, and uptake activity only are shown together with amino acids moderately involved in antiport activity only (see Fig. 5). The central cavity was modeled based on the results shown in Figs. 4 and 5. TM12 is involved in the recognition of cadaverine and putrescine
Furthermore, Trp43, Tyr57, Tyr107, Tyr366 and Tyr368 of CadB, and only Tyr90 of PotE were strongly involved in cadaverine and putrescine uptake at neutral pH, and Trp41, Tyr174, Glu185 and Glu408 of CadB, and Tyr78 and Trp422 of PotE were moderately involved in the uptake at the neutral pH without affecting the antiport activities (Fig. 5). These results suggest that certain residues of amino acids are necessary for recognition of cadaverine in the periplasm because the level of cadaverine is much lower than that of putrescine in the periplasm at neutral pH. In addition, Arg299 of CadB and Lys301 and Tyr308 of PotE were involved in cadaverine–lysine and putrescine–ornithine antiport activities without affecting uptake activities. It may be that a basic amino acid is necessary for recognition of the COOH group of lysine and ornithine. These amino acids are also located in the central cavity, i.e., both Arg299 of CadB and Lys301 of PotE in TM8 (Fig. 6). Although two tyrosine residues in TM10 of CadB were strongly involved in the uptake activities, no functional amino acid was found in TM10 of PotE. Such a functional difference may be caused by the difference of the variable loop between TM8 and TM9 in CadB and PotE, as mentioned in the previous section.
The transport path consisting of the central cavity and TM12 was modeled according to the ternary structure of CadB and PotE (Fig. 6). Functional amino acid residues were located on the surface of transport path, and the number of functional amino acid residues correlated well with the size of the central cavity.
Working hypothesis for the function of CadB and PotE
It has been shown that the expression of cadB mRNA is greatly enhanced at acidic pH by lysine and moderately by ornithine, and that of potE mRNA is enhanced greatly by ornithine and moderately by lysine (Soksawatmaekhin et al. 2004). It was also shown that PotE catalyzes putrescine–lysine exchange (Kashiwagi et al. 1992). It is expected that lysine is much more abundant than ornithine in the external environment of bacteria, judging from the amino acid composition in bacteria and mammals (Herbert et al. 1966; Gitlitz et al. 1974; Kashiwagi and Igarashi 1988). Cadaverine content of E. coli at neutral pH is <2% of putrescine (Igarashi et al. 1986), and it takes time to synthesize cadaverine with inducible lysine decarboxylase at acidic pH.
A model for agmatine–arginine exchange has been proposed (Gao et al. 2010). According to the model, it is thought that CadB and PotE also exist as an outward-open form. Since putrescine is abundant inside cells due to a combination of unbound putrescine (Miyamoto et al. 1993) and putrescine released from RNA at acidic pH, probably PotE functions first. Either ornithine or lysine binds to PotE, and the structure of PotE changes to an inward-open form. Then, putrescine exchanges with ornithine or lysine due to the higher affinity of putrescine than ornithine or lysine for PotE (Kashiwagi et al. 1992). As a result, ornithine or lysine is moved into the cytoplasm, and putrescine is moved into the periplasm through interaction with aromatic amino acids, Trp201 and Trp292 on TM6a and TM8. After this, cadaverine is accumulated through inducible lysine decarboxylase. Subsequently, when cadaverine levels rise due to the activity of inducible lysine decarboxylase, the cadaverine–lysine antiporter of CadB is initiated. CadB functions similarly to PotE. First lysine binds to the central cavity, and it is exchanged with cadaverine. Then, bound cadaverine moves into the periplasm through interaction with aromatic amino acids Tyr89 and Tyr90 on TM3 and Tyr235 on TM7. The Km value of putrescine for antiport activity of PotE was 73 μM (Kashiwagi et al. 1992), and that of cadaverine for antiport activity of CadB was 303 μM (Soksawatmaekhin et al. 2004). In this way, both CadB and PotE function as electrogenic cadaverine–lysine antiporter and putrescine–ornithine or lysine antiporter, leading to an increase in pH in medium, and stimulation of cell growth as reported previously (Takayama et al. 1994; Soksawatmaekhin et al. 2004) (Fig. 1). When the cad operon was inactivated, expression of potE mRNA was greatly enhanced (Soksawatmaekhin et al. 2004), confirming that CadB and PotE function together. Neutralization of the external environment is an important function of free polyamines, although polyamines usually function through their interaction with nucleic acids, especially with RNA (Igarashi and Kashiwagi 2010b).
Concluding remarks and future perspectives
Based on the crystal structure of AdiC, model structures of CadB and PotE were constructed. Although the model structures of CadB and PotE were similar, the width of the central cavity was different, reflecting differences in the size of the substrate. Accordingly, TMs constituting the central cavity were different in CadB and PotE. The cavity of CadB consisted mainly of TMs 2, 3, 6, 7, 8, and 10, and that of PotE consisted of TMs 2, 3, 6, and 8 (see Fig. 6). As for the working hypothesis for the molecular mechanism of antiport activity, further study is necessary to obtain clear evidence for that. These two transporters contribute to cell growth by creating membrane potential and increasing pH in the external medium (see Fig. 1). If free diamines (cadaverine and putrescine) have another unique function outside the cells in addition to elevating the pH, the importance of these transporters increase greatly. Thus, it may be interesting to look for another function of cadaverine and putrescine in the external medium.
Abbreviations
- AdiC:
-
Agmatine–arginine antiporter
- CadB:
-
Cadaverine–lysine antiporter
- PotE:
-
Putrescine–ornithine antiporter
- APC superfamily:
-
Amino acid/polyamine/organocation superfamily
- TM:
-
Transmembrane helix
References
Arnold K, Bordoli L, Kopp J, Schwede T (2006) The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22:195–201
Bordoli L, Kiefer F, Arnold K, Benkert P, Battey J, Schwede T (2009) Protein structure homology modeling using SWISS-MODEL workspace. Nat Protoc 4:1–13
Fang Y, Jayaram H, Shane T, Kolmakova-Partensky L, Wu F, Williams C, Xiong Y, Miller C (2009) Structure of a prokaryotic virtual proton pump at 3.2 Å resolution. Nature 460:1040–1043
Gao X, Lu F, Zhou L, Dang S, Sun L, Li X, Wang J, Shi Y (2009) Structure and mechanism of an amino acid antiporter. Science 324:1565–1568
Gao X, Zhou L, Jiao X, Lu F, Yan C, Zeng X, Wang J, Shi Y (2010) Mechanism of substrate recognition and transport by an amino acid antiporter. Nature 463:828–832
Gitlitz PH, Sunderman FW Jr, Hohnadel DC (1974) Ion-exchange chromatography of amino acids in sweat collected from healthy subjects during sauna bathing. Clin Chem 20:1305–1312
Gong S, Richard H, Foster JW (2003) YjdE (AdiC) is the arginine: agmatine antiporter essential for arginine-dependent acid resistance in Escherichia coli. J Bacteriol 185:4402–4409
Herbert JD, Coulson RA, Hernandez T (1966) Free amino acids in the caiman and rat. Comp Biochem Physiol 17:583–598
Higashi K, Ishigure H, Demizu R, Uemura T, Nishino K, Yamaguchi A, Kashiwagi K, Igarashi K (2008) Identification of a spermidine excretion protein complex (MdtJI) in Escherichia coli. J Bacteriol 190:872–878
Igarashi K, Kashiwagi K (1996) Polyamine transport in Escherichia coli. Amino Acids 10:83–97
Igarashi K, Kashiwagi K (1999) Polyamine transport in bacteria and yeast. Biochem J 344:633–642
Igarashi K, Kashiwagi K (2010a) Characteristics of cellular polyamine transport in prokaryotes and eukaryotes. Plant Physiol Biochem 48:506–512
Igarashi K, Kashiwagi K (2010b) Modulation of cellular function by polyamines. Int J Biochem Cell Biol 42:39–51
Igarashi K, Kashiwagi K, Hamasaki H, Miura A, Kakegawa T, Hirose S, Matsuzaki S (1986) Formation of a compensatory polyamine by Escherichia coli polyamine-requiring mutants during growth in the absence of polyamines. J Bacteriol 166:128–134
Iyer R, Williams C, Miller C (2003) Arginine-agmatine antiporter in extreme acid resistance in Escherichia coli. J Bacteriol 185:6556–6561
Jack DL, Paulsen IT, Saier MH Jr (2000) The amino acid/polyamine/organocation (APC) superfamily of transporters specific for amino acids, polyamines and organocations. Microbiology 146:1797–1814
Kashiwagi K, Igarashi K (1988) Adjustment of polyamine contents in Escherichia coli. J Bacteriol 170:3131–3135
Kashiwagi K, Hosokawa N, Furuchi T, Kobayashi H, Sasakawa C, Yoshikawa M, Igarashi K (1990) Isolation of polyamine transport-deficient mutants of Escherichia coli and cloning of the genes for polyamine transport proteins. J Biol Chem 265:20893–20897
Kashiwagi K, Suzuki T, Suzuki F, Furuchi T, Kobayashi H, Igarashi K (1991) Coexistence of the genes for putrescine transport protein and ornithine decarboxylase at 16 min on Escherichia coli chromosome. J Biol Chem 266:20922–20927
Kashiwagi K, Miyamoto S, Suzuki F, Kobayashi H, Igarashi K (1992) Excretion of putrescine by the putrescine–ornithine antiporter encoded by the potE gene of Escherichia coli. Proc Natl Acad Sci U S A 89:4529–4533
Kashiwagi K, Shibuya S, Tomitori H, Kuraishi A, Igarashi K (1997) Excretion and uptake of putrescine by the PotE protein in Escherichia coli. J Biol Chem 272:6318–6323
Kashiwagi K, Kuraishi A, Tomitori H, Igarashi A, Nishimura K, Shirahata A, Igarashi K (2000) Identification of the putrescine recognition site on polyamine transport protein PotE. J Biol Chem 275:36007–36012
Kurihara S, Tsuboi Y, Oda S, Kim HG, Kumagai H, Suzuki H (2009) The putrescine importer PuuP of Escherichia coli K-12. J Bacteriol 191:2776–2782
Meng SY, Bennett GN (1992) Nucleotide sequence of the Escherichia coli cad operon: a system for neutralization of low extracellular pH. J Bacteriol 174:2659–2669
Miyamoto S, Kashiwagi K, Ito K, Watanabe S, Igarashi K (1993) Estimation of polyamine distribution and polyamine stimulation of protein synthesis in Escherichia coli. Arch Biochem Biophys 300:63–68
Soksawatmaekhin W, Kuraishi A, Sakata K, Kashiwagi K, Igarashi K (2004) Excretion and uptake of cadaverine by CadB and its physiological functions in Escherichia coli. Mol Microbiol 51:1401–1412
Soksawatmaekhin W, Uemura T, Fukiwake N, Kashiwagi K, Igarashi K (2006) Identification of the cadaverine recognition site on the cadaverine–lysine antiporter CadB. J Biol Chem 281:29213–29220
Takayama M, Ohyama T, Igarashi K, Kobayashi H (1994) Escherichia coli cad operon functions as a supplier of carbon dioxide. Mol Microbiol 11:913–918
Watson N, Dunyak DS, Rosey EL, Slonczewski JL, Olson ER (1992) Identification of elements involved in transcriptional regulation of the Escherichia coli cad operon by external pH. J Bacteriol 174:530–540
Acknowledgments
We thank Drs. A. J. Michael and K. Williams for their help in preparing this manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tomitori, H., Kashiwagi, K. & Igarashi, K. Structure and function of polyamine-amino acid antiporters CadB and PotE in Escherichia coli . Amino Acids 42, 733–740 (2012). https://doi.org/10.1007/s00726-011-0989-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-011-0989-9