Abstract
ECF transporters are a family of active transporters for vitamins. They are composed of four subunits: a membrane-embedded substrate-binding subunit (EcfS), a transmembrane coupling subunit (EcfT) and two ATP-binding-cassette ATPases (EcfA and EcfA′). We have investigated the mechanism of the ECF transporter for riboflavin from the pathogen Listeria monocytogenes, LmECF–RibU. Using structural and biochemical approaches, we found that ATP binding to the EcfAA′ ATPases drives a conformational change that dissociates the S subunit from the EcfAA′T ECF module. Upon release from the ECF module, the RibU S subunit then binds the riboflavin transport substrate. We also find that S subunits for distinct substrates compete for the ATP-bound state of the ECF module. Our results explain how ECF transporters capture the transport substrate and reproduce the in vivo observations on S-subunit competition for which the family was named.







Similar content being viewed by others
References
Rodionov, D.A. et al. A novel class of modular transporters for vitamins in prokaryotes. J. Bacteriol. 191, 42–51 (2009).
Henderson, G.B. & Zevely, E.M. Binding and transport of thiamine by Lactobacillus casei. J. Bacteriol. 133, 1190–1196 (1978).
Henderson, G.B., Zevely, E.M. & Huennekens, F.M. Coupling of energy to folate transport in Lactobacillus casei. J. Bacteriol. 139, 552–559 (1979).
Henderson, G.B., Zevely, E.M. & Huennekens, F.M. Mechanism of folate transport in Lactobacillus casei: evidence for a component shared with the thiamine and biotin transport systems. J. Bacteriol. 137, 1308–1314 (1979).
Lewinson, O., Lee, A.T., Locher, K.P. & Rees, D.C. A distinct mechanism for the ABC transporter BtuCD–BtuF revealed by the dynamics of complex formation. Nat. Struct. Mol. Biol. 17, 332–338 (2010).
Hebbeln, P., Rodionov, D.A., Alfandega, A. & Eitinger, T. Biotin uptake in prokaryotes by solute transporters with an optional ATP-binding cassette-containing module. Proc. Natl. Acad. Sci. USA 104, 2909–2914 (2007).
Eitinger, T., Rodionov, D.A., Grote, M. & Schneider, E. Canonical and ECF-type ATP-binding cassette importers in prokaryotes: diversity in modular organization and cellular functions. FEMS Microbiol. Rev. 35, 3–67 (2011).
Erkens, G.B., Majsnerowska, M., Ter Beek, J. & Slotboom, D.J. Energy coupling factor-type ABC transporters for vitamin uptake in prokaryotes. Biochemistry 51, 4390–4396 (2012).
ter Beek, J., Duurkens, R.H., Erkens, G.B. & Slotboom, D.J. Quaternary structure and functional unit of energy coupling factor (ECF)-type transporters. J. Biol. Chem. 286, 5471–5475 (2011).
Xu, K. et al. Crystal structure of a folate energy-coupling factor transporter from Lactobacillus brevis. Nature 497, 268–271 (2013).
Wang, T. et al. Structure of a bacterial energy-coupling factor transporter. Nature 497, 272–276 (2013).
Zhang, M. et al. Structure of a pantothenate transporter and implications for ECF module sharing and energy coupling of group II ECF transporters. Proc. Natl. Acad. Sci. USA 111, 18560–18565 (2014).
Zhang, P. Structure and mechanism of energy-coupling factor transporters. Trends Microbiol. 21, 652–659 (2013).
Slotboom, D.J. Structural and mechanistic insights into prokaryotic energy-coupling factor transporters. Nat. Rev. Microbiol. 12, 79–87 (2014).
Erkens, G.B. et al. The structural basis of modularity in ECF-type ABC transporters. Nat. Struct. Mol. Biol. 18, 755–760 (2011).
Neubauer, O. et al. Two essential arginine residues in the T components of energy-coupling factor transporters. J. Bacteriol. 191, 6482–6488 (2009).
Neubauer, O., Reiffler, C., Behrendt, L. & Eitinger, T. Interactions among the A and T units of an ECF-type biotin transporter analyzed by site-specific crosslinking. PLoS ONE 6, e29087 (2011).
Duurkens, R.H., Tol, M.B., Geertsma, E.R., Permentier, H.P. & Slotboom, D.J. Flavin binding to the high affinity riboflavin transporter RibU. J. Biol. Chem. 282, 10380–10386 (2007).
Zhang, P., Wang, J. & Shi, Y. Structure and mechanism of the S component of a bacterial ECF transporter. Nature 468, 717–720 (2010).
Folta-Stogniew, E. Oligomeric states of proteins determined by size-exclusion chromatography coupled with light scattering, absorbance, and refractive index detectors. Methods Mol. Biol. 328, 97–112 (2006).
Berntsson, R.P. et al. Structural divergence of paralogous S components from ECF-type ABC transporters. Proc. Natl. Acad. Sci. USA 109, 13990–13995 (2012).
Kaback, H.R. et al. Site-directed alkylation and the alternating access model for LacY. Proc. Natl. Acad. Sci. USA 104, 491–494 (2007).
Korkhov, V.M., Mireku, S.A. & Locher, K.P. Structure of AMP-PNP-bound vitamin B12 transporter BtuCD–F. Nature 490, 367–372 (2012).
Majsnerowska, M. et al. Substrate-induced conformational changes in the S-component ThiT from an energy coupling factor transporter. Structure 21, 861–867 (2013).
Karpowich, N.K. & Wang, D.N. Assembly and mechanism of a group II ECF transporter. Proc. Natl. Acad. Sci. USA 110, 2534–2539 (2013).
Hopfner, K.P. et al. Structural biology of Rad50 ATPase: ATP-driven conformational control in DNA double-strand break repair and the ABC-ATPase superfamily. Cell 101, 789–800 (2000).
Smith, P.C. et al. ATP binding to the motor domain from an ABC transporter drives formation of a nucleotide sandwich dimer. Mol. Cell 10, 139–149 (2002).
Chen, J., Lu, G., Lin, J., Davidson, A.L. & Quiocho, F.A. A tweezers-like motion of the ATP-binding cassette dimer in an ABC transport cycle. Mol. Cell 12, 651–661 (2003).
Benabdelhak, H. et al. Positive co-operative activity and dimerization of the isolated ABC ATPase domain of HlyB from Escherichia coli. Biochem. J. 386, 489–495 (2005).
Davidson, A.L. & Sharma, S. Mutation of a single MalK subunit severely impairs maltose transport activity in Escherichia coli. J. Bacteriol. 179, 5458–5464 (1997).
Nikaido, K. & Ames, G.F. One intact ATP-binding subunit is sufficient to support ATP hydrolysis and translocation in an ABC transporter, the histidine permease. J. Biol. Chem. 274, 26727–26735 (1999).
Zaitseva, J., Jenewein, S., Jumpertz, T., Holland, I.B. & Schmitt, L. H662 is the linchpin of ATP hydrolysis in the nucleotide-binding domain of the ABC transporter HlyB. EMBO J. 24, 1901–1910 (2005).
Zaitseva, J. et al. Functional characterization and ATP-induced dimerization of the isolated ABC-domain of the haemolysin B transporter. Biochemistry 44, 9680–9690 (2005).
Chai, C. et al. Structural basis for a homodimeric ATPase subunit of an ECF transporter. Prot. Cell 4, 793–801 (2013).
Kawate, T. & Gouaux, E. Fluorescence-detection size-exclusion chromatography for precrystallization screening of integral membrane proteins. Structure 14, 673–681 (2006).
Erkens, G.B. & Slotboom, D.J. Biochemical characterization of ThiT from Lactococcus lactis: a thiamin transporter with picomolar substrate binding affinity. Biochemistry 49, 3203–3212 (2010).
Eudes, A. et al. Identification of genes encoding the folate- and thiamine-binding membrane proteins in Firmicutes. J. Bacteriol. 190, 7591–7594 (2008).
Oldham, M.L., Khare, D., Quiocho, F.A., Davidson, A.L. & Chen, J. Crystal structure of a catalytic intermediate of the maltose transporter. Nature 450, 515–521 (2007).
Davidson, A.L., Shuman, H.A. & Nikaido, H. Mechanism of maltose transport in Escherichia coli: transmembrane signaling by periplasmic binding proteins. Proc. Natl. Acad. Sci. USA 89, 2360–2364 (1992).
Burgess, C.M. et al. The riboflavin transporter RibU in Lactococcus lactis: molecular characterization of gene expression and the transport mechanism. J. Bacteriol. 188, 2752–2760 (2006).
Ott, E., Stolz, J., Lehmann, M. & Mack, M. The RFN riboswitch of Bacillus subtilis is a target for the antibiotic roseoflavin produced by Streptomyces davawensis. RNA Biol. 6, 276–280 (2009).
Vogl, C. et al. Characterization of riboflavin (vitamin B2) transport proteins from Bacillus subtilis and Corynebacterium glutamicum. J. Bacteriol. 189, 7367–7375 (2007).
Andersson, H. & von Heijne, G. Membrane protein topology: effects of delta mu H+ on the translocation of charged residues explain the 'positive inside' rule. EMBO J. 13, 2267–2272 (1994).
Studier, F.W. Protein production by auto-induction in high density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Geertsma, E.R., Nik Mahmood, N.A., Schuurman-Wolters, G.K. & Poolman, B. Membrane reconstitution of ABC transporters and assays of translocator function. Nat. Protoc. 3, 256–266 (2008).
Kodama, T., Fukui, K. & Kometani, K. The initial phosphate burst in ATP hydrolysis by myosin and subfragment-1 as studied by a modified malachite green method for determination of inorganic phosphate. J. Biochem. 99, 1465–1472 (1986).
Acknowledgements
We are grateful to the staff at beamlines X25 and X29 of the National Synchrotron Light Source in the Brookhaven National Laboratory for assistance in X-ray diffraction experiments. We thank R. Mancusso, D. Sauer and C. Huang for technical assistance and J. Marden for comments on the manuscript. N.K.K. thanks the American Heart Association and the US National Institutes of Health (NIH) (F32-HL091618) for postdoctoral fellowships. This work was also financially supported by NIH grants R01DK099023, R01-DK073973, R01-GM093825 and R01- MH083840 to D.-N.W.
Author information
Authors and Affiliations
Contributions
N.K.K. designed and performed the experiments with J.M.S. and N.C.; N.K.K. and D.-N.W. analyzed the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Characterization of the purified ECF riboflavin transporter from L. monocytogenes, LmECF–RibU.
(a) Preparative size exclusion chromatography (SEC) trace of purified LmECF–RibU and SDS-PAGE of the peak fractions (inset). (b) The molecular mass (Mw) of the protein complex in detergent was determined by analytical SEC and multi-angle laser light scattering (MALLS).
Supplementary Figure 2 TMRM labeling of WT and cysteine mutants of LmRibU.
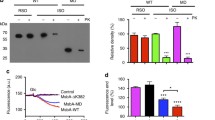
(a) Purified LmRibU variants were incubated with the thiol reactive probe, TMRM, for the indicated time points. After termination of the reaction by DTT, samples were subjected to SDS-PAGE and TMRM labeling was visualized on a fluorescence scanner. (b) Initial rates of TMRM labeling were calculated from the data shown in (a).
Supplementary Figure 3 TMRM labeling of WT and cysteine mutants of LmECF–RibU.
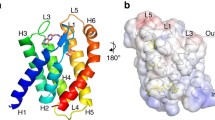
(a) Ribbon diagram of the LmECF–RibU homology model with each subunit colored as indicated and cysteine residues shown as red spheres. The position of the membrane is represented as black lines. (b) Purified LmECF–RibU variants were labeled with the thiol reactive probe, TMRM, for the indicated time points. After SDS-PAGE (top panels), the labeling of the individual subunits was visualized on a fluorescence scanner (bottom panels). (c) Initial rates of TMRM labeling were calculated from these data.
Supplementary Figure 4 AMP-PNP is a low-affinity competitive inhibitor of LmECF–RibU.
(a) ATPase activity of LmECF–RibU in the presence of increasing concentrations of AMPPNP. Analytical SEC traces of LmECF–RibU measured in the presence of the vitamin B2 transport substrate and several concentrations of AMPPNP detected by UV absorbance (b) or riboflavin fluorescence (c).
Supplementary Figure 5 Characterization of the H199′A H204A ATPase-deficient mutant of LmECF–RibU.
(a) Preparative SEC trace of the purified protein and SDS-PAGE of the peak fractions (inset). (b) The molecular mass of the protein complex in detergent was determined by analytical SEC and MALLS.
Supplementary Figure 6 Schematic illustrating the experimental design to test for S-subunit exchange by EcfAA′T and calibration of the S-subunit exchange assay with free RibU*.
(a) First, a mixture of LmECF–RibU and isolated substrate-bound RibU that has been labeled with a fluorophore (S*) was prepared. Next either ADP, ATP, or no nucleotide is added and the samples are incubated for 10 min at 25 °C. Subsequently, the samples are separated by SEC, and binding of S* by EcfAA′T is detected by monitoring the S* fluorescence relative to elution position. SEC traces of several concentrations of RibU* detected by UV absorbance (b) and TMRM fluorescence(c). The integrated peak areas from (c) were used to calibrate the assay, as shown in (d).
Supplementary Figure 7 The TMRM label attached to RibU* is not transferred to LmECF–RibU during the incubation step of the exchange reaction.
LmECF–RibU (1), RibU* (2), or an equimolar mixture of the two in the absence (3) or presence of MgATP (4) were incubated under the same conditions of the exchange reaction. Instead of SEC, these samples were subject to SDS-PAGE (left), and the TMRM labeling was visualized on a fluorescence scanner (right). If the TMRM molecule was transferred from RibU* to LmECF–RibU, we would expect to see a fluorescent band in (4) corresponding to EcfA, which is labeled by TMRM on Cys11.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Table 1 and Supplementary Notes 1–3 (PDF 609 kb)
Supplementary Data Set 1
Complete gels for Figure 5c,f (PDF 2753 kb)
Rights and permissions
About this article
Cite this article
Karpowich, N., Song, J., Cocco, N. et al. ATP binding drives substrate capture in an ECF transporter by a release-and-catch mechanism. Nat Struct Mol Biol 22, 565–571 (2015). https://doi.org/10.1038/nsmb.3040
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3040
- Springer Nature America, Inc.
This article is cited by
-
Structural basis for inhibition of the drug efflux pump NorA from Staphylococcus aureus
Nature Chemical Biology (2022)
-
Membrane mediated toppling mechanism of the folate energy coupling factor transporter
Nature Communications (2020)
-
Cu Transport by the Extended Family of CcoA-like Transporters (CalT) in Proteobacteria
Scientific Reports (2019)
-
A flavin-based extracellular electron transfer mechanism in diverse Gram-positive bacteria
Nature (2018)
-
Structural insight in the toppling mechanism of an energy-coupling factor transporter
Nature Communications (2016)