Abstract
Among various types of skin diseases, skin cancer is the deadliest form of the disease. This paper classifies seven types of skin diseases: Actinic keratosis and intraepithelial carcinoma, Basal cell carcinoma, Benign keratosis, Dermatofibroma, Melanoma, Melanocytic type, and Vascular lesions. The primary objective of this paper is to evaluate the performance of these deep learning networks on skin lesion images. The lesion classification is implemented through transfer learning on fourteen deep learning networks: AlexNet, GoogleNet, ResNet50, VGG16, VGG19, ResNet101, InceptionV3, InceptionResNetV2, SqueezeNet, DenseNet201, ResNet18, MobileNetV2, ShuffleNet and NasNetMobile. The dataset used for these experiments are from ISIC 2018 of about 10,154 images. The results show that DenseNet201 performs best with 0.825 accuracy and improves skin lesion classification under multiple diseases. The proposed work shows the various parameters, including the accuracy of all fourteen deep learning networks, which helped build an efficient automated classification model for multiple skin lesions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Human skin is the biggest organ in the body and is loaded with various factors, including sun rays (Ultraviolet), sunburn, lifestyle, smoking, liquor utilization, physical action, viruses, and the workplace. These factors compromise its trustworthiness and have real and dangerous impacts on human health. Diseases that influence the skin directly are the common reasons for every human disease, influencing almost 33% of the total population of around 1.9 billion at once, leading to research in this discipline. Skin diseases added to roughly 1.79% of the global problem of diseases, estimated by balanced life years. In Great Britain, 60% of the population experiences skin diseases for the duration of their lives. Skin issues can be harmful, incendiary, or infectious and influence individuals, all things considered, particularly older people and young children. There are various consequences of skin issues, for example, death (on account of melanoma), loss of connections, effect on day-to-day activity and damage to internal organs. Besides, they likewise represent a danger to humans, affecting them mentally, prompting loneliness, misery, and even rashness. Dermal diseases should be treated at the beginning to decrease related outcomes, costs, mortality, and morbidity rates. As indicated by Dr Macrene Alexiades-Armenakas, cancer and eczema are among the five most basic skin syndromes [1]. In this manner, our fundamental emphasis is on building a system that can automatically diagnose for analyses and grouping these types of diseases.
The visual assessment with the naked eye during a skin disease health check-up makes it hard to contrast skin injuries and typical tissue, producing a misdiagnosis. Indeed, dermoscopy is a mainly solid conceptual method for skin injuries. The input thought of dermoscopy is to obtain an enlarged high–resolution image while removing reflections from the skin's surface. The utilization of dermoscopy imaging tools improves the general perception of skin lesions, sensitivity (correct finding of disease) and specificity (correct finding of doubtful disease) contrasted with visual assessment; further, it needs improvement to analyze skin disease. However, since physical examination from dermoscopy images are frequently time-consuming, error-prone, complex, and personal (i.e., might create different diagnostic outcome). Hence, an automated and consistent computer-aided diagnostic (CAD) framework to recognize skin cancer growth has developed a significant evaluation device that offers dermatologists a subsequent input to help and aid their decisions [2].
There are numerous investigations on diagnosing skin disease utilizing deep learning and machine learning strategies in the literature. In recent times, complex medical problems are handled through deep learning convolutional neural networks (CNN) [3,4,5,6], particularly dermoscopic image analysis [7,8,9,10,11] to melanoma recognition. Overall, there are existing and tested convolutional neural network classifiers based on deep learning, for example, AlexNet [12], VGGNet [13], GoogLeNet [14], ResNet [15], and DenseNet [16]. These CNNs go further to expand for more accurately identifying the five challenging tasks [17, 18]. The research goes to a two-course model that utilizes a deep residual network to segment and classify skin lesions [11]. The method of moving a portioned mask of skin lesions, alongside descriptions of clinical rules, such as, shading, texture, and morphological qualities, to the CNN through U-Net [19] has been used. Later, it introduced a two-advance model for the classification of skin lesions [20]. Shading, texture, and shape qualities were removed from images through U-Net and sent to a Support Vector Machine (SVM) classifier to analyze favourable or dangerous dermoscopy pictures. The hybrid system for classifying the skin lesion is introduced, a blend of CNN and SVM with sparse coding(SC) [21]. In recent years, researchers have also built a deep learning network based on a hybrid approach through encoding local descriptors [11]. The various features, including deep and statistical from ResNet, utilized the SVM classifier and the chi-square kernel to recognize the distinctive skin lesions. The usage of both handcrafted through traditional approach and learned features using deep learning with ResNet-50 for improvement in classification was proposed. It consolidates high-quality conventional capacities through picture handling and ResNet-50 profound, getting the hang of learning capacities [8]. In the latest, the authors detect seven classes of cancer using a support vector machine with an artificial bee colony method. RNA sequencing is also utilized [9].
Apart from the literature, we have classified the seven different skin diseases, including Actinic keratoses and intraepithelial carcinoma, Basal cell carcinoma, Benign keratosis, Dermatofibroma, Melanoma, Melanocytic type, and Vascular lesions, using fourteen types of deep learning networks. The objective of this paper is to evaluate the performance of these networks based on performance measures. These networks are trained, validated, and tested on ISIC 2018 dataset using Matlab.
In the following, we list contributions of this study:
-
1.
We used transfer learning by adding one fully connected layer, softmax layer and cross-entropy layer as pre-trained networks. These three layers in total can classify skin lesions into seven classes.
-
2.
To improve the classification performance, we added pre-processing steps as data normalization, resizing and augmentation.
-
3.
The comparison of fourteen types of deep learning networks is analysed to build an automated classification model.
The remainder of this paper is outlined as follows. Section 2 presents the methodology with description of dataset and detail of fourteen deep learning models. Section 3 shows the experiments and results with implementation. Section 4 deliberates the performance of deep learning networks for classification of multiple skin lesion. Finally, the paper ends with a conclusion in Sect. 5.
2 Methodology
This section briefly explains the dataset description, its pre-processing and different deep convolutional neural networks. Then, all these networks are compared using the same number of epochs, learning rate and batch size.
2.1 Dataset
This study used the ISIC 2018 dataset for dermoscopy images [22, 23]. It contains pigmented lesions in different populations. This data set consists of 10,015 images in total. The image sizes are scaled to 224 × 224 × 3 or 227 × 227 × 3 or as per the network requirement. The diagnostic classes in this dataset are given as follows [22, 23].
-
(1)
Actinic keratoses and intraepithelial carcinoma abbreviated as ‘akiec’.
-
(2)
Basal cell carcinoma abbreviated as ‘bcc’.
-
(3)
Benign keratosis abbreviated as ‘bkl’.
-
(4)
Dermatofibroma abbreviated as ‘df’.
-
(5)
Melanoma abbreviated as ‘mel’.
-
(6)
Melanocytic type abbreviated as ‘nv’.
-
(7)
Vascular lesions abbreviated as ‘vasc’.
Figure 1 illustrate the examples for akiec. Figure 2 shows the examples for bcc. Figure 3 shows the examples for bkl. Figure 4 shows the examples for df. Figure 5 shows the examples for mel. Figure 6 shows the examples for nv. Figure 7 shows the examples for vasc. All examples are taken from [22, 23].
The number of data contained in image set for each class is defined in Table 1.
As shown in Table 1, most of the data in the dataset belong to the sixth grade, and there is no equal distribution between classes. Therefore, the training phase will lead to more learning of the 6th class and the network to learn this information.
2.2 Data pre-processing
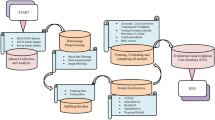
In our approach, the pre-processing steps are reduced to guarantee better generalizability when tried on the dermoscopic skin lesion dataset. In this way, just the three standard pre-processing steps are applied, generally utilized for transfer learning. Initially, the images are normalized by taking away the typical RGB values from the dataset. After that, the images are resized utilizing bicubic interpolation for the networks input (227 × 227 and 224 × 224). At last, the training set is augmented by arbitrarily flipping the training images along the vertical axis and changing it up to 30 pixels horizontally and vertically. Data augmentation keeps the network away from overfitting and memorizing the exact details of the training images. The block diagram of the classification model is shown in Fig. 8.
2.3 AlexNet
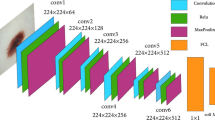
This network contains 8 learned layers (5 convolutional and 3 fully connected layers) [24, 25]. The convolution filters and an activation non-linear function (ReLU) is applied after every layer. Due to fully connected layers, the input size is fixed to RGB images of 224 × 224 × 3, i.e., 150,528 values. In the FCN and cross-entropy layer, there are 4096 neurons; each neuron is a different feature of the image. Dropouts are utilized at regular intervals to avoid overfitting issues.
2.4 GoogleNet
The GoogleNet architecture comprises 22 convolutional layers containing 9 Inception modules [26]. The Inception module has three kernel varieties with various sizes: 5 × 5, 3 × 3 and 1 × 1 for convolution and 3 × 3 filters for pooling. These small convolutions help reduce the depth of the feature map, and many inception modules also helps reduce the number of computations. The receptive field size for this network is 224 × 224 × 3 utilizing the RGB colors gap with the predefined parameter. Like different CNNs, this network also learn convolutional filters contributions through stochastic gradient descent (SGD) algorithms throughout the training stage to retrieve compelling features as the image crosses the hierarchical structure of the network.
2.5 ResNet50
This network consists of a stem used for input, followed by four stages and an output layer [27]. It consists of 50 layers and an input image size fixed to 224 × 224. The stem is input through a 7 × 7 convolution by a return of 64 channels besides a pitch of 2, trailed through a 3 × 3 max pooling layer besides a pitch of 2. The width and height of the input decrease by stem through a factor of 4, and rises a size of channel to 64. The individual stage starts with a downsampling block, further trailed by various residual blocks at the starting of stage2. This block consists of path A and path B. Path A consists of three convolutions with a kernel size of 1 × 1, 3 × 3, and 1 × 1. Path B utilizes a pitch of 2 with a 1 × 1 convolution to convert the input into the output of path A. Finally, the sum of these two paths is used to get the output of the downsampling block.
2.6 VGG16/VGG19
The VGG-16 architecture is reasonable for GPUs with local memory [28]. The architecture consists of 16 layers, among which there are 13 convolutional layers, 5 Max Pooling layers and 3 fully connected layers, which sums up to 21 layers but only 16 weight layers. The information layer is thought to be a 224 × 224 × 3 pixel RGB picture. All convolutional layers are followed by the Relu function with a size of 3 × 3. The VGG19 network [29] contains 19 trained layers combining convolutional and FC layers, max-pooling, and dropout layers.
2.7 ResNet101
The ResNet101 network structure consists of residual connections and generally used for classification purposes. The gradients flow throughout the layers directly and prevent gradient as zero after the chain rule applications [30]. A total of 104 convolutional layers are present in the network. Alongside, there are 33 blocks of layers, and the output of the previous block is used among these 29 blocks directly as residual connections above. To receive the input of the other blocks, these residual connections are being utilized at the termination of every block using the initial value of the sigma operator.
2.8 InceptionV3
This network is based on inception modules and consists of 48 layers deep [31]. These inception blocks contain convolutions corresponding with diverse kernel sizes to abstract features and finally summative the results. Initially, the input image is passed through convolution, batch normalization, Relu and this sequence are followed by pooling and various inception layers for feature extraction. Finally, in the classification part, the dropout layer is applied to reduce overfitting, and softmax with cross-entropy as output layers.
2.9 InceptionResNetV2
This network [32] consists of 164 layers in deep and inception blocks with residual connections. The input size of an image acceptable to the network is of size 299 × 299. The network consists of the main module as stem, followed by Inception resnet-A, Reduction-A, Inception resnet-B module, reduction B and inception resnet-C modules. The stem module consists of series of 3 × 3 convolution, 3 × 3 maxpool, and filter concat to give input to inception resnet-A block. The Relu activation function is applied in each inception block and the reduction block contains 1 × 1, 3 × 3 convolution with a pooling layer. Inception-ResNet used batch-normalization only on top of the traditional layers but not on top of the summations.
2.9.1 SqueezeNet
This network starts with [33] a separate convolution layer (conv1), trailed by 8 Fire modules (fire2-9), and termination with a final convolution layer (conv10). The filters numbers get increased as per fire module from starting till the end of the network. The fire module is composed with a squeezed convolution layer of 1 × 1 filters only, serving into an enlarge layer with a combination of 1 × 1 and 3 × 3 size convolution filters. The maxpooling operation is applied after convolution1, fire4, fire8, and convolution10 layers.
2.9.2 DenseNet201
The connections are directly between the convolutional layers in this network (DenseNet) [34, 35]. In every network layer, the feature map of the continuing layer acts as input, whereas feature maps that are generated are used by way of inputs to the next layer. This network consists of four blocks with layers in the same amount. These blocks have feature maps with various sizes of 56-by-56, 28-by-28, 14-by-14, and 7-by-7. The number of input feature maps are reduced to advance the computational efficiency by using 1 × 1 convolution before each 3 × 3 convolution, known as the bottleneck layer.
2.9.3 ResNet18
This network starts with 7 × 7 convolution, 3 × 3 max pooling, followed by 4 convolutional layers in each module [36]. There are total 18 layers which include the first convolution layer and the last FC layer. This network has an image input size of 224-by-224. The initial 2 layers of this network are similar to the GoogLeNet: the 64 output channels with stride 2 based 7 × 7 convolutional layer trailed by 3 × 3 max pooling layer, stride 2 and Relu function. The change is the batch wise normalization layer additional next to every convolutional layer in the network. It consists of residual blocks with 3 × 3 convolutional layers having a similar amount of output channels.
2.9.4 Mobile NetV2
The MobileNetV2 model [37] is based on the concept of separable convolutions, which can be depth wise (dw) or pointwise (pw) convolution. In the case of the former, a filter is applied to every channel used for input. Then to combine an output of former 1 × 1 convolution is applied by later convolution. A one-step process is required to get the original block of outputs by filtering and pooling together by convolution. The process is separated in two layers, filtering and combining. This factorization results into significantly reducing computation and size.
2.9.5 ShuffleNet
The ShuffleNet [38] utilizes pointwise group convolution and channel shuffle to reduce computation cost while maintaining accuracy. By shuffling the channels, ShuffleNet outperforms MobileNetV1. The network contains a heap of network units assembled in three stages. It has two units as bottleneck unit through depthwise convolution and shufflenet unit through pointwise group convolution.
2.9.6 NasNetMobile
Nasnet [39] is an architecture constitute of basic building blocks i.e. cells optimized through reinforcement learning. The network has an image input size of 224-by-224. A cell composed of convolution and pooling operations, depends upon the strength of the network to repeat these operations at n number of times. This network contains 12 cells and millions of multiply-accumulates (MACs). There are two cells: Normal cell, which returns a feature map of the same dimension and Reduction cell, which returns a feature map where the feature map height and width are reduced by a factor of two. The various combinations of operations or layers applied to the network in the form of identity, convolution (1 × 7, 7 × 1), average pooling (3 × 3), max pooling (5 × 5), depth wise convolution (3 × 3, 7 × 7, 5 × 5) and dilated convolution (3 × 3).
3 Experiments and results
The experiments are performed with Dell compatible computer equipped with a Core i7 processor. The ISIC 2018 dataset defined in RGB space is used. The datasets are divided into seven classes, Actinic keratoses and intraepithelial carcinoma, Basal cell carcinoma, Benign keratosis, Dermatofibroma, Melanoma, Melanocytic type, and Vascular lesions. The fourteen types of convolution deep learning networks are used; in which the last fully connected layer is replaced with FC7, softmax and cross-entropy for classification. The experiments are performed with all types of networks using 10,015 dermoscopic images.
Each experiment consisted of the number of runs where the batch size, epochs and the initial learning rate were 64, 10 and 0.0001, respectively, and these values are fixed for all experiments. The images are randomly divided: 70% of the images are used for training and 30% are used for validation and testing. All the images are pre-processed using re-sizing, normalization, and augmentation. The networks are evaluated using the following metric described in Eqs. 1, 2, 3 and 4:
where, tp, tn, fp and fn refer to true positive, true negative, false positive and false negative, respectively.
The alexnet training Progress with accuracy and error loss over 10 Epochs is shown in Fig. 9. The performance of alexnet in which class wise recall, precision, f1 score and overall accuracy based on confusion matrix are shown in Fig. 10 and Table 2, the best result is highlighted in bold. The Predictions on Random samples made by Transfer Learning on AlexNet is shown in Fig. 11.
The GoogleNet training Progress with accuracy and error loss over 10 Epochs is shown in Fig. 12. The performance of GoogleNet in which class wise recall, precision, f1 score and overall accuracy based on confusion matrix are shown in Fig. 13 and Table 3, the best result is highlighted in bold. The Predictions on Random samples made by Transfer Learning on GoogleNet is shown in Fig. 14.
The ResNet50 training Progress with accuracy and error loss over 10 Epochs is shown in Fig. 15. The performance of ResNet50 in which class wise recall, precision, f1 score and overall accuracy based on confusion matrix are shown in Fig. 16 and Table 4, the best result is highlighted in bold. The Predictions on Random samples made by Transfer Learning on ResNet50 is shown in Fig. 17.
The VGG16 training Progress with accuracy and error loss over 10 Epochs is shown in Fig. 18. The performance of VGG16 in which class wise recall, precision, f1 score and overall accuracy based on confusion matrix are shown in Fig. 19 and Table 5, the best result is highlighted in bold. The Predictions on Random samples made by Transfer Learning on VGG16 is shown in Fig. 20.
The VGG19 training Progress with accuracy and error loss over 10 Epochs is shown in Fig. 21. The performance of VGG19 in which class wise recall, precision, f1 score and overall accuracy based on confusion matrix are shown in Fig. 22 and Table 6, the best result is highlighted in bold. The Predictions on Random samples made by Transfer Learning on VGG19 is shown in Fig. 23.
The ResNet101 training Progress with accuracy and error loss over 10 Epochs is shown in Fig. 24. The performance of ResNet101 in which class wise recall, precision, f1 score and overall accuracy based on confusion matrix are shown in Fig. 25 and Table 7, the best result is highlighted in bold. The Predictions on Random samples made by Transfer Learning on ResNet101 is shown in Fig. 26.
The InceptionV3 training Progress with accuracy and error loss over 10 Epochs is shown in Fig. 27. The performance of InceptionV3 in which class wise recall, precision, f1 score and overall accuracy based on confusion matrix are shown in Fig. 28 and Table 8, the best result is highlighted in bold. The Predictions on Random samples made by Transfer Learning on InceptionV3 is shown in Fig. 29.
The InceptionResNetV2 training Progress with accuracy and error loss over 10 Epochs is shown in Fig. 30. The performance of InceptionResNetV2 in which class wise recall, precision, f1 score and overall accuracy based on confusion matrix are shown in Fig. 31 and Table 9, the best result is highlighted in bold. The Predictions on Random samples made by Transfer Learning on InceptionResNetV2 is shown in Fig. 32.
The SqueezeNet training Progress with accuracy and error loss over 10 Epochs is shown in Fig. 33. The performance of SqueezeNet in which class wise recall, precision, f1 score and overall accuracy based on confusion matrix are shown in Fig. 34 and Table 10, the best result is highlighted in bold. The Predictions on Random samples made by Transfer Learning on SqueezeNet is shown in Fig. 35.
The DenseNet201 training Progress with accuracy and error loss over 10 Epochs is shown in Fig. 36. The performance of DenseNet201 in which class wise recall, precision, f1 score and overall accuracy based on confusion matrix are shown in Fig. 37 and Table 11, the best result is highlighted in bold. The Predictions on Random samples made by Transfer Learning on DenseNet201 is shown in Fig. 38.
The ResNet18 training Progress with accuracy and error loss over 10 Epochs is shown in Fig. 39. The performance of ResNet18 in which class wise recall, precision, f1 score and overall accuracy based on confusion matrix are shown in Fig. 40 and Table 12, the best result is highlighted in bold. The Predictions on Random samples made by Transfer Learning on ResNet18 is shown in Fig. 41.
The MobileNetV2 training Progress with accuracy and error loss over 10 Epochs is shown in Fig. 42. The performance of MobileNetV2 in which class wise recall, precision, f1 score and overall accuracy based on confusion matrix are shown in Fig. 43 and Table 13, the best result is highlighted in bold. The Predictions on Random samples made by Transfer Learning on MobileNetV2 is shown in Fig. 44.
The ShuffleNet training Progress with accuracy and error loss over 10 Epochs is shown in Fig. 45. The performance of ShuffleNet in which class wise recall, precision, f1 score and overall accuracy based on confusion matrix are shown in Fig. 46 and Table 14, the best result is highlighted in bold. The Predictions on Random samples made by Transfer Learning on ShuffleNet is shown in Fig. 47.
The NasNetMobile training Progress with accuracy and error loss over 10 Epochs is shown in Fig. 48. The performance of NasNetMobile in which class wise recall, precision, f1 score and overall accuracy based on confusion matrix are shown in Fig. 49 and Table 15, the best result is highlighted in bold. The Predictions on Random samples made by Transfer Learning on NasNetMobile is shown in Fig. 50.
The fourteen deep convolutional networks with number of layers, size, parameters, input size, time, accuracy, recall, precision and f1 score, the average out for a single matrix, are presented in Table 16. DenseNet201 performs better in performance measures and highlighted bold in the Table 16.
4 Discussion
It is noted that the results obtained from the fourteen deep convolutional networks are comparable in terms of accuracy, recall, precision and f1 score performance measures. By comparing the performance measures, DenseNet201 network can perform best with 0.825 accuracy. Furthermore, this network gives good efficiency by eliminating the vanishing gradient problem, supporting feature propagation, encouraging reuse of features, and reducing the number of parameters. Besides improved efficiency, another best point is that the information flow and gradients movement is smooth in this network, which helps them in training and reduces the overfitting problem.
The performance of the networks is evaluated on the ISIC 2018 dataset consists of 10,015 dermoscopic images. The training progress, confusion matrix for each type of class and testing images results show the best performance of DenseNet201. The ResNet50 model takes the second position, which is taking less time and good accuracy. Similarly, the other model's performance is measurable with time parameters complexity.
5 Conclusion
This study compared the performance of fourteen deep convolutional neural networks for classifying seven types of diseases. This model's performance is measured using ISIC 2018 dataset. The pre-trained networks are modified by replacing their last fully connected layer with our fully connected-7, softmax and cross-entropy layer for classification into seven classes. The results are summarized in Table 16. The results show the best performance of DenseNet201 for classification. In the upcoming years, computer-aided diagnostic systems will be more for classifying a skin lesion by looking at the inspiring results of our study and the former studies using deep learning techniques. The models designed with this baseline are expected to give high accuracy with minimum computation time. In future, our goal is to achieve more effective results by increasing the number of epochs and skin lesions categories.
Data Availability
ISIC database.
References
Hameed N, Shabut AM, Ghosh MK, Hossain MA (2020) Multi-class multi-level classification algorithm for skin lesions classification using machine learning techniques. Expert Syst with Appl 141:112961. https://doi.org/10.1016/j.eswa.2019.112961
Al-Masni MA, Kim DH, Kim TS (2020) Multiple skin lesions diagnostics via integrated deep convolutional networks for segmentation and classification. Comput Methods Programs Biomed 190:105351. https://doi.org/10.1016/j.cmpb.2020.105351
Nasiri S, Helsper J, Jung M, Fathi M (2020) DePicT Melanoma Deep-CLASS: a deep convolutional neural networks approach to classify skin lesion images. BMC bioinform 21:1–3. https://doi.org/10.1186/s12859-020-3351-y
Mukherjee S, Adhikari A, Roy M (2019) Malignant melanoma classification using cross-platform dataset with deep learning CNN architecture. In: Bhattacharyya S, Pal SK, Pan I, Das A (eds) Recent trends in signal and image processing 2019. Springer, Singapore, pp 31–41
Seeja RD, Suresh A (2019) Deep learning based skin lesion segmentation and classification of melanoma using support vector machine (SVM). Asian Pac J of Cancer Prev APJCP 20(5):1555
Chung YM, Hu CS, Lawson A, Smyth C (2019) Toporesnet: A hybrid deep learning architecture and its application to skin lesion classification. arXiv preprint, arXiv:1905.08607
Al-Antari MA, Al-Masni MA, Kim TS (2020) Deep learning computer-aided diagnosis for breast lesion in digital mammogram. In: Lee G, Fujita H (eds) Deep Learning in Medical Image Analysis. Springer, Cham, pp 59–72
Ali R, Hardie RC, De Silva MS, Kebede TM (2019) Skin lesion segmentation and classification for ISIC 2018 by combining deep CNN and handcrafted features. arXiv preprint, arXiv:1908.05730
Al-Obeidat F, Rocha Á, Akram M et al (2021) (CDRGI)-Cancer detection through relevant genes identification. Neural Comput Applic. https://doi.org/10.1007/s00521-021-05739-8
Yilmaz E, Trocan M (2020) Benign and malignant skin lesion classification comparison for three deep-learning architectures. In: Thanh Nguyen N, Jearanaitanakij K, Selamat A, Trawiński B, Chittayasothorn S (eds) Asian Conf on Intell Inf and Database Syst. Springer, Cham, pp 514–524
Saba T, Khan MA, Rehman A, Marie-Sainte SL (2019) Region extraction and classification of skin cancer: a heterogeneous framework of deep CNN features fusion and reduction. J Med Syst 43:289. https://doi.org/10.1007/s10916-019-1413-3
Amin J, Sharif A, Gul N, Anjum MA, Nisar MW, Azam F, Bukhari SA (2020) Integrated design of deep features fusion for localization and classification of skin cancer. Pattern Recognit Lett 131:63–70. https://doi.org/10.1016/j.patrec.2019.11.042
Delibasis K, Georgakopoulos SV, Tasoulis SK, Maglogiannis I, Plagianakos VP (2020) On image prefiltering for skin lesion characterization utilizing deep transfer learning. In: Iliadis L, Parvanov Angelov P, Jayne C, Pimenidis E (eds) International conf on engineering appl of neural networks. Springer, Cham, pp 377–388
Ballester P, Araujo R (2016) On the performance of GoogLeNet and AlexNet applied to sketches. In :Proceedings of the AAAI Conference on Artificial Intelligence: 30(1)
Chen M, Chen W, Chen W, Cai L, Chai G (2020) Skin cancer classification with deep convolutional neural networks. J of Med Imaging and Health Info 10(7):1707–1713. https://doi.org/10.1166/jmihi.2020.3078
Gessert N, Nielsen M, Shaikh M, Werner R, Schlaefer A (2020) Skin lesion classification using ensembles of multi-resolution EfficientNets with meta data. MethodsX 7:100864. https://doi.org/10.1016/j.mex.2020.100864
Carcagnì P, Leo M, Cuna A, Mazzeo PL, Spagnolo P, Celeste G, Distante C (2019) Classification of skin lesions by combining multilevel learnings in a DenseNet architecture. In: Ricci E, Bulò SR, Snoek C, Lanz O, Messelodi S, Sebe N (eds) International Conf on Image Analysis and Processing. Springer, Cham, pp 335–344. https://doi.org/10.1007/978-3-030-30642-7_30
Chen EZ, Dong X, Li X, Jiang H, Rong R, Wu J (2019) Lesion attributes segmentation for melanoma detection with multi-task u-net. In: Proceedings of the 2019 IEEE 16th international symp on biomedical imaging (ISBI 2019), pp 485–488, IEEE. https://doi.org/10.1109/ISBI.2019.8759483
Yang J, Sun X, Liang J, Rosin PL (2018) Clinical skin lesion diagnosis using representations inspired by dermatologist criteria. In: Proceedings of the IEEE conf on computer vision and pattern recognition, pp 1258–1266, IEEE.
Nazi ZA, Abir TA (2020) Automatic skin lesion segmentation and melanoma detection: Transfer learning approach with u-net and dcnn-svm. In: Uddin MS, Bansal JC (eds) Proc of international joint conf on computational intelligence. Springer, Singapore, pp 371–381. https://doi.org/10.1007/978-981-13-7564-4_32
Rastgoo M, Lemaître G, Morel O, Massich J, Garcia R, Meriaudeau F, Marzani F, Sidibé D (2016) Classification of melanoma lesions using sparse coded features and random forests. Med Imaging 2016: Comput-Aided Diagn 9785:97850
Codella N, Rotemberg V, Tschandl P, Celebi ME, Dusza S, Gutman D, Helba B, Kalloo A, Liopyris K, Marchetti M, Kittler H (2019) Skin lesion analysis toward melanoma detection 2018: A challenge hosted by the international skin imaging collaboration (isic). arXiv preprint arXiv:1902.03368
Tschandl P, Rosendahl C, Kittler H (2018) The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Sci Data 5(1):1–9. https://doi.org/10.1038/sdata.2018.161
Krizhevsky A, Sutskever I, Hinton GE (2012) Imagenet classification with deep convolutional neural networks. Adv Neural Inf Process Syst 25:1097–1105
Negrete PDM, Iano Y, Monteiro ACB, França RP, Gomes G, de Oliveira D, Pajuelo (2021) Classification of dermoscopy skin images with the application of deep learning techniques. In: Iano Y, Arthur R, Saotome O, Kemper G, Monteiro ACB (eds) Proc of the 5th Brazilian technology symp. Springer, Cham, pp 73–81. https://doi.org/10.1007/978-3-030-57566-3_7
Balazs H (2018) Skin lesion classification with ensembles of deep convolutional neural networks. J of Biomed Inf 86:25–32. https://doi.org/10.1016/j.jbi.2018.08.006
Szegedy C, Liu W, Jia Y, Sermanet P, Reed S, Anguelov D, Erhan D, Vanhoucke V, Rabinovich A (2015) Going deeper with convolutions. In: Proceedings of the IEEE conf on computer vision and pattern recognition, pp 1–9
Quang NH (2017) Automatic skin lesion analysis towards melanoma detection. In: proceedings of the 2017 21st Asia Pacific symposium on intelligent and evolutionary systems (IES), pp 106–111. https://doi.org/10.1109/iesys.2017.8233570
Kwasigroch A, Mikołajczyk A, Grochowski M (2017) Deep neural networks approach to skin lesions classification—A comparative analysis. In: proceedings of the 2017 22nd int conf on methods and models in automation and robotics (MMAR), pp 1069–1074. https://doi.org/10.1109/mmar.2017.8046978
Demir A, Yilmaz F, Kose O (2019) Early detection of skin cancer using deep learning architectures: Resnet-101 and inception-v3. In 2019 Medical Technologies Congress (TIPTEKNO), pp 1–4, IEEE. https://doi.org/10.1109/tiptekno47231.2019.8972045
Shahin AH, Kamal A, Elattar MA (2018) Deep ensemble learning for skin lesion classification from dermoscopic images. In: proceedings of the 2018 9th cairo international biomedical engineering conference (CIBEC), pp. 150–153. https://doi.org/10.1109/cibec.2018.8641815
Szegedy C, Ioffe S, Vanhoucke V, Alemi A (2017) Inception-v4, inception-resnet and the impact of residual connections on learning. In: Proceedings of the AAAI Conf on Artificial Intelligence:31(1)
Iandola FN, Han S, Moskewicz MW, Ashraf K, Dally WJ, Keutzer K (2016) SqueezeNet: AlexNet-level accuracy with 50x fewer parameters and< 0.5 MB model size. arXiv preprint arXiv:1602.07360
Al-Masni MA, Kim DH, Kim TS (2020) Multiple skin lesions diagnostics via integrated deep convolutional networks for segmentation and classification. Comput Methods Programs Biomed 190:105351. https://doi.org/10.1016/j.cmpb.2020.105351
Huang G, Liu Z, Van Der Maaten L, Weinberger KQ (2017). Densely connected convolutional networks. In: Proceedings of the IEEE conf on computer vision and pattern recognition, pp 4700–4708
Czum JM (2020) Dive into deep learning. J of the American College of Radiology 17(5):637–638. https://doi.org/10.1016/j.jacr.2020.02.005
Howard AG, Zhu M, Chen B, Kalenichenko D, Wang W, Weyand T, Andreetto M, Adam H (2017) Mobilenets: Efficient convolutional neural networks for mobile vision applications. arXiv preprint arXiv:1704.04861
Zhang X, Zhou X, Lin M, Sun J (2018) Shufflenet: An extremely efficient convolutional neural network for mobile devices. In: Proceedings of the IEEE conf on computer vision and pattern recognition, pp 6848–6856
Zoph B, Vasudevan V, Shlens J, Le QV (2018) Learning transferable architectures for scalable image recognition. In: Proceedings of the IEEE conf on computer vision and pattern recognition, pp 8697–8710
Funding
None.
Author information
Authors and Affiliations
Contributions
Ginni Arora: Conceptualization, Methodology, Software, Data curation, Validation, Writing- Original draft preparation. Ashwani Kumar Dubey: Conceptualization, Methodology, Supervision, Reviewing and Editing. Zainul Abdin Jaffery: Conceptualization, Supervision, Reviewing and Editing. Alvaro Rocha: Reviewing and Editing.
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict of interest.
Consent to participate
I consent to participate.
Consent for publication
I consent for publication.
Code availability
Custom code.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Arora, G., Dubey, A.K., Jaffery, Z.A. et al. A comparative study of fourteen deep learning networks for multi skin lesion classification (MSLC) on unbalanced data. Neural Comput & Applic 35, 7989–8015 (2023). https://doi.org/10.1007/s00521-022-06922-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00521-022-06922-1