Abstract
The effect of central metabolic activity of Escherichia coli cells acting as biocatalysts on the performance of microbial fuel cells (MFCs) was studied with glucose used as the energy source. Milliliter-scale two-chambered MFCs were used with 2-hydroxy-1,4-naphthoquinone (HNQ) as an electron mediator. Among the single-gene deletions examined, frdA, pdhR, ldhA, and adhE increased the average power output of the constructed MFC. Next, multiple-gene knockout mutants were constructed using P1 transduction. The Δ5 (ΔfrdAΔpdhRΔldhAΔadhEΔpta) strain showed the highest ave. power output (1.82 mW) and coulombic efficiency (21.3%). Our results show that the combination of multiple-gene knockout in E. coli cells leads to the development of an excellent catalyst for MFCs. Finally, preventing a decrease in the pH of the anodic solution was a key factor for improving the power output of the Δ5 strain, and a maximum ave. power output of 2.21 mW was achieved with 5% NaHCO3 in the buffer. The ave. power density of the constructed MFC was 0.27 mW/cm3, which is comparable to an enzymatic fuel cell of a Milliliter-scale using glucose dehydrogenase.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Clean energy production from renewable substrates is essential for achieving sustainable energy supplies. Microbial fuel cells (MFCs) are devices that convert the chemical energy of biological fuels into electrical energy using microorganisms as biocatalysts [1,2,3]. MFCs have several key advantages over conventional fuel cells. For example, MFCs can be operated under mild conditions and do not require expensive inorganic catalysts or purification of the biocatalysts [4], which allows for large-scale, long-lifetime operation.
Looking back at the long history of MFC research, the idea of using microorganisms to produce electricity was, remarkably, first conceived and reported in 1911, using E. coli cells as the biocatalyst [5]. E. coli shows promise as a biocatalyst for MFCs because this bacterium can utilize a broad spectrum of organic compounds. However, electron mediators are necessary for E. coli cells to be used in MFC systems [6,7,8], because E. coli cells do not possess the extracellular electron transfer (EET) pathways found in electricigenic bacteria such as Shewanella oneidensis or Geobacter sulfurreducens [9, 10]. Due to the lack of EET pathways in E. coli cells, many researchers have attempted to promote the performance of E. coli-based MFCs by endowing E. coli cells with an EET pathway either via gene modification [11,12,13,14,15] or by modifying the electrode or the MFC system [16,17,18] for more efficient electron transfer. However, modifying the central metabolic activity of E. coli on the performance of MFCs have not been well studied.
In the present study, the performance of E. coli-based MFCs using E. coli cells that had multiple central metabolism genes knocked out was examined. In the first step, single-gene knockout mutants were compared in terms of their effect on the output of a constructed MFC system. Next, multiple-gene knockout mutants were constructed by sequential knockout of genes that had increased the MFC power output, and the combination of multiple genes knocked out with highest power output was determined. Furthermore, the effect of 2-hydroxy-1,4-naphthoquinone (HNQ) and sodium hydrogen carbonate concentrations on the output of the Δ5 strain was examined to consider the electron transfer efficiency and pH value at the anode.
Materials and methods
Bacterial strains
The E. coli strains used in this study are listed in Table 1. E. coli K-12 strain BW25113 and its derivatives [19] were obtained from the National BioResource Project (NIG, Mishima, Japan). The BW25113(DE3) strain was constructed in previous work [20]. In our study, multiple-deficiency mutant strains were constructed from the BW25113(DE3) strain by P1 transduction using P1kc phage [20]. Briefly, P1 transduction was performed on a kanamycin (Km)-sensitive recipient strain, with the donor strain harboring a Km cassette between flippase recognition target sites (FRT-Km-FRT cassette), which was inserted at the desired gene on the chromosome. Km resistance was used to select the recipient strain. The Km cassette was eliminated from the recipient strain by FLP/FRT (Flippase/Flippase Recognition Target) recombination. For multiple-deficiency mutant strains, the P1 transduction and elimination of the Km cassette steps were repeated. The deletion was confirmed by PCR using loci-specific primers (Supplementary Table S1).
Construction of MFCs
Milliliter-scale two-chambered MFCs were fabricated according to a method described previously with minor modifications [21]. Briefly, anode and cathode tanks of 8.3 mL capacity were constructed from an acrylic rod of diameter 3.4 cm. Commercial carbon rods with diameters of 0.5 cm were used as cathodes. Carbon fiber bundles (Mitsubishi Rayon Co. Ltd, Japan) were used as high-performance anodes. The diameter of one filament was 7 µm and one bundle consisted of approximately 12,000 filaments. For one anode, 10 bundles approximately 8 cm long were used to maintain contact with E. coli cells in the anode solution. A cation-specific membrane, GORE-SELECT (Japan Gore-Tex Inc. Japan), of thickness 30 µm, was used to separate the two electrode chambers.
Operation of MFCs
The fundamental anode solution (7.5 mL) was composed of 50 g/L glucose, 10 g/L sodium hydrogen carbonate (NaHCO3), 0.2 g/L ammonium sulfate, and 0.5 M phosphate buffer (pH 8.0) in reverse osmosis water. This solution also contained 1 g/L 2-hydroxyl-1,4-naphthoquinone (HNQ) as an electron transfer mediator [21]. The cathodic chamber was filled with 7.5 mL of deionized water containing 158 g/L potassium ferricyanide, K3[Fe(CN)6]. Each E. coli strain was pre-cultured in lysogeny broth (LB) medium at 30 °C. After 24 h, glucose was added to the culture broth at a concentration of 50 g/L and cultured for 1 h as a warm-up culture for the MFC operation.
E. coli cells were harvested by centrifugation (10,000×g, 5 min). The harvested E. coli cells were resuspended using the anode solution to an OD600 = 64 and added to the anodic chamber. The MFC was connected to an ammeter and a voltmeter (Sanwa Electric Instrument, Tokyo, Japan) and operated at 37 °C. Current and voltage were recorded via a computer using PC Kink 7 software (Sanwa Electric Instrument) through a connection with a 100 Ω resistor.
Analyses
Samples of 0.2 mL were withdrawn from the anode solution of a fuel cell for measuring the glucose concentration. After centrifugation (10,000×g, 5 min), the glucose concentration was determined by HPLC (GL Sciences Inc. Tokyo, Japan) using an ULTRON PS-80 N column and ultrapure water as the solvent with a flow rate of 0.6 mL/min at 60 °C. Glucose was detected using a refractive index detector.
The coulombic efficiency was calculated as 100 × Cp/CT [22], where Cp is the total coulombs calculated by integrating the current over time, and CT is the theoretical amount of coulombs available from glucose, which is given by the following equation:
where F is Faraday’s constant (98,485 C per mol of electrons), N is the number of mol of electrons generated per mol of glucose (24 electrons per mol), \(\Delta S\) is the concentration of glucose consumed, v is the volume of liquid, and M is the molecular mass of glucose.
Results and discussion
Performance of the MFC with single-gene knockout mutants
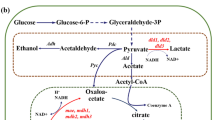
Figure 1 shows an overview of the modification of E. coli central metabolism as a biocatalyst for improving the power output in a constructed MFC system. In total, 13 genes associated with the central metabolism of E. coli cells were selected for single-gene deletion. Eight genes, ldhA, adhE, adhP, pta, poxB, ackA, acs and pflB, are involved in the synthesis or degradation of organic acids generated by branching from glycolysis [23]. Four genes, pdhR, arcA, arcB and rpoS, are regulators to repress the activity of the TCA cycle under microaerobic or anaerobic conditions [24]. The frdA gene encodes the enzyme which catalyzes the reduction of fumarate to succinate in the TCA cycle [25]. We expected that knockouts of these genes would lead to more NADH (i.e., electrons) generation from the TCA cycle.
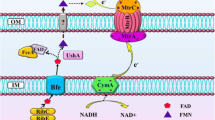
Overview of the modification of E. coli central metabolism as a biocatalyst for improving power output in a constructed MFC system. This overview includes the hypothetical mechanism of electron transfer, in which HNQ obtains electrons from a quinone pool such as ubiquinone in the cell membrane of E. coli cells
Figure 2 shows the time course of power outputs from whole fuel cells composed of various single-gene knockout mutants of E. coli strains. For the wild-type (WT) BW25113 strain, the power started at 1.5 mW and reached a peak value of 2.3 mW after 60 min. This value is comparable to that of a similar MFC system that used yeast cells [21], indicating that E. coli can be used as a biocatalyst in the examined MFC system. After 60 min, the power output decreased in a linear manner to 0.5 mW at the end of the reaction (1080 min). Furthermore, for the single-gene knockout mutant strains, while the peak values and the slopes of decreasing power differed, the time courses showed a similar tendency. Among the mutant strains, the ΔfrdA, ΔpdhR, ΔldhA and ΔadhE strains had higher peak values of power (approximately 2.5 mW) when compared with that of the WT stain, and these values were maintained at a higher level throughout the MFC operation period. As for the genes involved in acetate metabolism (Δpta, ΔackA, ΔpoxB and Δacs), the ΔpoxB strain showed a relatively higher peak value; however, the decrease in power output was much steeper than for other strains, especially during the latter stages of the operating period. In contrast, the Δpta strain showed the smallest peak value of 2.0 mW at 60 min. The deletion of genes related to formate and ethanol production (ΔadhP and ΔpflB) and regulators of the TCA cycle did not improve the power output drastically.
The ave. power outputs of whole fuel cells for 18 h are summarized in Fig. 3. The ave. power of the WT strain was 1.25 mW, whereas four strains (ΔfrdA, ΔpdhR, ΔldhA, and ΔadhE) showed significantly higher ave. power values. The maximum ave. value was 1.49 mW, which was achieved by the ΔadhE strain. The power outputs of the other nine strains (Δpta, ΔackA, ΔpoxB, Δacs, ΔadhP, ΔpflB, ΔarcA, ΔarcB and ΔrpoS) were the same or less than the WT strain. The ave. power output of the ΔrpoS strain was significantly lower than the other strains, at 1.09 mW.
Ave. power output of whole fuel cells for 18 h operation with each knockout mutant strain of E. coli. Data were obtained from more than three independent experiments. Vertical bars indicate standard deviations. Statistically significant differences from the WT strain (p < 0.05) are marked with asterisks
The anodic chamber of the MFC constructed was considered to be under oxygen-limited conditions because there was minimal oxygen supply and the cell concentration was extremely high (OD600 = 64). In E. coli, under oxygen-limited conditions, the frdA gene encodes fumarate reductase, which catalyzes the reduction of fumarate to succinate accompanied by the oxidation of reduced menaquinone [25]. Deletion of the frdA gene has been reported to significantly suppress succinate production and slightly increases lactate production [26, 27]. These results suggest that the accumulation of reduced compounds arising from frdA deletion promoted the performance of MFC. The pyruvate dehydrogenase complex regulator (PdhR) is a transcriptional regulator that negatively controls the formation of the pyruvate dehydrogenase complex (PDHc), NADH dehydrogenase (NDH)-2 and cytochrome bo3 oxidase in E. coli. Under oxygen-limited conditions, it has been reported that pdhR-deficient mutants grow and consume glucose more efficiently than the WT strain, with enhanced respiration because of the increased activities of PDHc and NDH-2 [28]. These reports support our result of improved MFC output following pdhR deletion. The ldhA and adhE genes encode lactate dehydrogenase and alcohol dehydrogenase, respectively [23]. Given that these enzymes consume NADH to produce anaerobic byproducts, it is not unexpected that the deletion of these genes increased the power output of E. coli MFCs (Fig. 1).
Improvement of MFC performance with multiple-knockout mutants
Next, the cumulative effects of multiple-gene deletions in E. coli cells on MFC performance were examined. The BW25113(DE3) strain containing the λDE3 lysogen was used as the parent strain in a series of experiments. First, four genes (frdA, pdhR, ldhA and adhE) were selected for multiple-gene deletions, because deletion of each of these genes increased MFC the power. Although single-gene deletions of pta, poxB and pflB had no great effect on MFC power output, these enzymes are involved in the acetate and formate production pathways and play an important role in central, anaerobic carbon metabolism reactions. These genes were also selected for deletion because the consequences may differ when these genes are cumulatively deleted. The arcA gene was also chosen for deletion because a previous report described improved MFC performance using the ΔarcA strain [6]. In that report, the ΔarcA strain increased the power output because of the activation of key enzymes in the TCA cycle.
Figure 4 shows the time course of MFC power outputs using multiple-knockout mutants. Compared with the WT (BW25113(DE3)) strain, all multiple-knockout mutants showed much higher peak values of power output. While the double-knockout mutant Δ2 (ΔfrdAΔpdhR) showed a rapid decrease in power output after 200 min, the triple-knockout mutant Δ3 (ΔfrdAΔpdhRΔldhA) slightly suppressed the power drop during the latter stages of the operation period. The Δ4 (ΔfrdAΔpdhRΔldhAΔadhE) and Δ5 (ΔfrdAΔpdhRΔldhAΔadhEΔpta) strains maintained relatively high power outputs and their time profiles overlapped. The sixth mutation of each of pflB, poxB, and arcA in the Δ5 strain did not improve the power profile.
Time courses of power outputs of whole fuel cells composed of various multiple-gene knockout mutant strains of E. coli. The definitions of the strains are given in Table 1. The lines show the average value of the power obtained from more than three independent experiments
The ave. power outputs are summarized in Fig. 5. The ave. power of BW25113(DE3) was 1.23 mW, which was almost equal to that of BW25113 (1.25 mW). This result suggests that the insertion of λDE3 lysogen into the genomic DNA of BW25113 did not affect its performance in the MFC. The ave. power of Δ2 was 1.46 mW, which was slightly higher than that of the single-knockout mutants, ΔfrdA (1.41 mW) and ΔpdhR (1.40 mW). The additional deletion of ldhA (Δ3) increased the ave. power to 1.57 mW. A cumulative effect of multiple-gene deletions on power output was observed. Further deletion of the adhE gene (Δ4) significantly promoted the performance of the MFC (1.81 mW) due to the slower decrease in power during the latter stages of operation period. Thus, the deletion of four genes (frdA, pdhR, ldhA, and adhE) resulted in a cumulative effect that increased the MFC power output. In addition, the ave. power of the Δ5 strain was 1.82 mW, and the additional deletion of the pta gene did not increase the MFC performance. Furthermore, a sixth mutation of each of pflB, poxB, and arcA in the Δ5 strain negatively affected the power output. These results suggest that the cumulative effects of multiple-gene deletions on MFC performance became saturated. The coulombic efficiency of these multiple-gene mutants was calculated based on the amount of glucose consumed (Fig. 5). The coulombic efficiencies of the WT, Δ2, and Δ3 strains were 8.1%, 9.5% and 9.1%, respectively, which were not significant. In contrast, the coulombic efficiency of Δ4 was 14%, which was significantly higher than that of Δ3. In addition to the increased ave. power, the repression of glucose consumption in the Δ4 strain is also a key reason for the higher coulombic efficiency (Fig. 5). Furthermore, the coulombic efficiency of the Δ5 strain reached 21.3%, even though the ave. power of this strain was almost equal to that of Δ4. This result is also because of the further decrease in glucose consumption by the Δ5 strain. In terms of fuel economy, Δ5 is a superior strain when compared with that of Δ4, although their power outputs are at the same level. Further multiple-gene deletion did not drastically increase coulombic efficiency. Considering both ave. power and the coulombic efficiency, it was concluded that the Δ5 strain with multiple-gene knockout is the best biocatalyst among the examined strains.
Ave. power output, glucose consumption, and coulombic efficiency of whole fuel cells for 18 h operation with various multiple-gene knockout mutant strains of E. coli. The definitions of the strains are given in Table 1. Data were obtained from more than three independent experiments. Vertical bars indicate standard deviations. Asterisks show statistically significant differences (p < 0.05)
Effect of HNQ concentration on the performance of the Δ5 strain
Although the Δ5 strain was selected as the best multiple-gene knockout E. coli for MFC, repression of glucose consumption still remained as a serious problem. We hypothesized that the reason for the decrease in glucose consumption was that the electron transfer efficiency was not sufficient for E. coli to regenerate the intracellular oxidation state at the anode. To confirm the effect of the mediator concentration on the power and glucose consumption for the Δ5 strain, lower (0.05%) and higher (0.20%) concentrations of HNQ (basal condition 0.10%) were added to the anodic chamber. Figure 6a shows the time course of power outputs using different mediator concentrations. In the presence of 0.05% HNQ, the power was clearly lower than that obtained with 0.10% HNQ. The ave. power with 0.05% HNQ was 1.56 mW (Fig. 6a), which was significantly lower when compared with that of 0.10% HNQ (1.82 mW). These results indicate that 0.05% HNQ is not sufficient to transfer all electrons generated from the Δ5 cells. In contrast, the time courses of power output using 0.10% and 0.20% HNQ almost overlapped and the ave. power did not increase with 0.20% HNQ. Based on these results, we concluded that the basal condition (0.10% HNQ) provided sufficient electron transfer efficiency and further increases in the HNQ concentration did not improve the power output and glucose consumption of Δ5 cells.
Effect of the mediator (HNQ) concentration on the performance of MFCs with the E. coli Δ5 strain. a Power output of MFCs with 0.05%, 0.10% and 0.20% HNQ. b Ave. power outputs of MFCs for 18 h operation with different HNQ concentrations. Data were obtained from more than three independent experiments and vertical bars indicate standard deviations. Asterisks show statistically significant differences (p < 0.05)
Effect of the NaHCO3 concentration on the pH value of the anodic solution and performance of the Δ5 strain
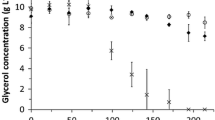
The pH of the anodic solution with the Δ5 strain was found to decrease considerably during the operating period. Although the pH of the anodic solution was initially adjusted with phosphate buffer (pH 8.0) and 1% NaHCO3, the pH value was observed to decrease to 5.1 over the 18 h operation period (Fig. 7a). This result suggests that the considerable pH drop decreased the activity of the Δ5 strain, which caused a decrease in power and glucose consumption. Higher concentrations of NaHCO3 were added to the anode (3–7%) to prevent the pH drop in the anodic solution of Δ5 strain. As shown in Fig. 7a, the pH of the anodic solution over 18 h gradually increased with increasing NaHCO3 concentrations used. The pH with 3% and 5% NaHCO3 was 6.1 and 6.8, respectively, and reached 7.4 with 7% NaHCO3 added, indicating that a drop in pH was prevented with higher NaHCO3 concentrations. Figure 7b shows the time course of MFC power outputs with different NaHCO3 concentrations. Compared with the basal condition (1% NaHCO3), the power with 3% NaHCO3 showed the same level of peak value and the power drop was strongly prevented during the latter stages of the operation period. With 5% NaHCO3, the value was maintained at a higher level throughout the MFC operation period. These results show that preventing the decrease in pH in the anodic solution effectively improved the power output of the Δ5 strain. A further increase of the NaHCO3 concentration to 7% showed a smaller peak value of 2.5 mW at 60 min and the power output was maintained at ~ 2.0 mW throughout the operation period. Therefore, the power output was more constant as the concentration of NaHCO3 increased. The values of the ave. power output, glucose consumption and coulombic efficiency are summarized in Fig. 7c. The ave. power with 3% NaHCO3 was 2.08 mW, which was significantly higher than that with 1% NaHCO3 (1.82 mW). With 5% NaHCO3, the ave. power increased slightly to 2.21 mW and was saturated with 7% NaHCO3 (2.20 mW). Glucose consumption with 3% NaHCO3 was 180 mg, which was two times higher than that with 1% NaHCO3. These results suggested that glucose consumption was strongly promoted by preventing a decrease in pH and this improved the power output of the Δ5 strain. Coulombic efficiency decreased with an increase in NaHCO3 concentration, which was inversely proportional to glucose consumption (Fig. 7c). Thus, controlling the pH was a key factor to improve the power output of the Δ5 strain and a maximum output of 2.21 mW was achieved with 5% NaHCO3.
Effect of NaHCO3 concentration on the pH value of the anodic solution and performance of the E. coli Δ5 strain. a pH value of the anodic solution over 18 h. b Time courses of power outputs of the E. coli Δ5 strain. c Ave. power output, glucose consumption, and coulombic efficiency of whole fuel cells for 18 h operation. Data were obtained from more than three independent experiments and vertical bars indicate standard deviations. Asterisks show statistically significant differences (p < 0.05)
Comparison of the performance of an enzymatic biofuel cell and the MFC prepared in this study
Biofuel cells are categorized based on the type of catalyst used; enzymes or microorganisms. In general, an enzymatic fuel cell (EFC) using glucose oxidase (GOx) or glucose dehydrogenase (GDH) shows a higher power density than a MFC because of the efficient reaction provided by the condensed pure enzymes. In contrast, a MFC can obtain a higher coulombic efficiency because microbial glucose degradation can theoretically obtain 24 electrons, whereas enzymatic glucose degradation is a 2-electron reaction. The MFC system in the present study has a simple anode component using glucose as a single substrate and that is very similar to EFCs. The results of the MFC used in the this study were compared with an enzymatic fuel cell with comparable cell volume (6.5 cm3), operating time (16.6 h) and glucose concentration (1.2 M) [29]. Table 2 shows the comparison of the performance of the EFC and MFC used in this study. The calculated ave. power density of the Δ5 strain with 5% NaHCO3 was 0.27 mW/cm3, which reached approximately 35% that obtained by the in EFC (0.77 mW/cm3) of a previous report [29]. The highest coulombic efficiency was 21.3% obtained from the Δ5 strain with 1% NaHCO3, which is five times or more when compared with that of the EFC (3.7%). Thus, although the power output is lower when compared with EFC, the constructed MFC provides a considerable improvement on previous MFCs and has reached a comparable level to EFCs.
Conclusions
In this study, the effect of central metabolic activity of E. coli cells, using glucose as an energy source on the performance of a MFC was studied. Single-gene deletions of frdA, pdhR, ldhA, and adhE increased the power output of the constructed MFC. Multiple-gene knockout mutants created using P1 transduction showed that the Δ5 (ΔfrdAΔpdhRΔldhAΔadhEΔpta) strain resulted in the highest ave. power (1.82 mW) and coulombic efficiency (21.3%). Finally, controlling the pH value was found to be a key factor in improving the power output of the Δ5 strain and a maximum ave. power of 2.21 mW was achieved with 5% NaHCO3. The ave. power density of the constructed MFC was 0.27 mW/cm3, which is comparable in performance to an enzymatic fuel cell (0.77 mW/cm3).
References
Choudhury P, Prasad Uday US, Bandyopadhyay TK, Ray RN, Bhunia B (2017) Performance improvement of microbial fuel cell (MFC) using suitable electrode and Bioengineered organisms: a review. Bioengineered 8:471–487
Li M, Zhou M, Tian X, Tan C, McDaniel CT, Hassett DJ, Gu T (2018) Microbial fuel cell (MFC) power performance improvement through enhanced microbial electrogenicity. Biotechnol Adv 36:1316–1327
Santoro C, Arbizzani C, Erable B, Ieropoulos I (2017) Microbial fuel cells: From fundamentals to applications A review. J Power Sources 356:225–244
Oliveira VB, Simões M, Melo LF, Pinto AMFR (2013) Overview on the developments of microbial fuel cells. Biochem Eng J 73:53–64
Potter MC (1911) Electrical effects accompanying the decomposition of organic compounds. Proc R Soc Lond B Biol Sci 84:260–276
Liu J, Yong Y-C, Song H, Li CM (2012) Activation enhancement of citric acid cycle to promote bioelectrocatalytic activity of arcA knockout Escherichia coli toward high-performance microbial fuel cell. ACS Catal 2:1749–1752
Ojima Y, Kawata T, Matsuo N, Nishinoue Y, Taya M (2014) Recovery of electric energy from formate by using a recombinant strain of Escherichia coli. Bioprocess Biosyst Eng 37:2005–2008
Wang C-T, Chen W-J, Huang R-Y (2010) Influence of growth curve phase on electricity performance of microbial fuel cell by Escherichia coli. Int J Hydrogen Energ 35:7217–7223
Logan BE, Regan JM (2006) Electricity-producing bacterial communities in microbial fuel cells. Trends Microbiol 14:512–518
Lovley DR (2006) Microbial fuel cells: novel microbial physiologies and engineering approaches. Curr Opin Biotechnol 17:327–332
Feng J, Qian Y, Wang Z, Wang X, Xu S, Chen K, Ouyang P (2018) Enhancing the performance of Escherichia coli-inoculated microbial fuel cells by introduction of the phenazine-1-carboxylic acid pathway. J Biotechnol 275:1–6
Jensen HM, Albers AE, Malley KR, Londer YY, Cohen BE, Helms BA, Weigele P, Groves JT, Ajo-Franklin CM (2010) Engineering of a synthetic electron conduit in living cells. Proc Natl Acad Sci USA 107:19213–19218
Jensen HM, TerAvest MA, Kokish MG, Ajo-Franklin CM (2016) CymA and exogenous flavins improve extracellular electron transfer and couple it to cell growth in Mtr-expressing Escherichia coli. ACS Synth Biol 5:679–688
Sturm-Richter K, Golitsch F, Sturm G, Kipf E, Dittrich A, Beblawy S, Kerzenmacher S, Gescher J (2015) Unbalanced fermentation of glycerol in Escherichia coli via heterologous production of an electron transport chain and electrode interaction in microbial electrochemical cells. Bioresour Technol 186:89–96
Yong YC, Yu YY, Yang Y, Liu J, Wang JY, Song H (2013) Enhancement of extracellular electron transfer and bioelectricity output by synthetic porin. Biotechnol Bioeng 110:408–416
Park IH, Heo YH, Kim P, Nahm KS (2013) Direct electron transfer in E. coli catalyzed MFC with a magnetite/MWCNT modified anode. RSC Adv 3:16665–16671
Singh S, Modi A, Verma N (2016) Enhanced power generation using a novel polymer-coated nanoparticles dispersed-carbon micro-nanofibers-based air-cathode in a membrane-less single chamber microbial fuel cell. Int J Hydrogen Energ 41:1237–1247
Zhou X, Chen X, Li H, Xiong J, Li X, Li W (2016) Surface oxygen-rich titanium as anode for high performance microbial fuel cell. Electrochim Acta 209:582–590
Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, Datsenko KA, Tomita M, Wanner BL, Mori H (2006) Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: The Keio collection. Mol Syst Biol 2(2006):0008
Koma D, Yamanaka H, Moriyoshi K, Ohmoto T, Sakai K (2012) A convenient method for multiple insertions of desired genes into target loci on the Escherichia coli chromosome. Appl Microbiol Biotechnol 93:815–829
Kaneshiro H, Takano K, Takada Y, Wakisaka T, Tachibana T, Azuma M (2014) A milliliter-scale yeast-based fuel cell with high performance. Biochem Eng J 83:90–96
Liu H, Logan BE (2004) Electricity generation using an air-cathode single chamber microbial fuel cell in the presence and absence of a proton exchange membrane. Environ Sci Technol 38:4040–4046
Clark DP (1989) The fermentation pathways of Escherichia coli. FEMS Microbiol Rev 63:223–234
Partridge JD, Sanguinetti G, Dibden DP, Roberts RE, Poole RK, Green J (2007) Transition of Escherichia coli from aerobic to micro-aerobic conditions involves fast and slow reacting regulatory components. J Biol Chem 282:11230–11237
Maklashina E, Berthold DA, Cecchini G (1998) Anaerobic expression of Escherichia coli succinate dehydrogenase: functional replacement of fumarate reductase in the respiratory chain during anaerobic growth. J Bacteriol 180:5989–5996
Mazumdar S, Clomburg JM, Gonzalez R (2010) Escherichia coli strains engineered for homofermentative production of D-lactic acid from glycerol. Appl Environ Microbiol 76:4327–4336
Zhou L, Zuo ZR, Chen XZ, Niu DD, Tian KM, Prior BA, Shen W, Shi GY, Singh S, Wang ZX (2011) Evaluation of genetic manipulation strategies on D-lactate production by Escherichia coli. Curr Microbiol 62:981–989
Maeda S, Shimizu K, Kihira C, Iwabu Y, Kato R, Sugimoto M, Fukiya S, Wada M, Yokota A (2017) Pyruvate dehydrogenase complex regulator (PdhR) gene deletion boosts glucose metabolism in Escherichia coli under oxygen-limited culture conditions. J Biosci Bioeng 123:437–443
Sakai H, Mita H, Sugiyama T, Tokita Y, Shirai O, Kano K (2014) Construction of a multi-stacked sheet-type enzymatic biofuel cell. Electrochemistry 82:156–161
Acknowledgements
This work was supported by a Grant-in-Aid for Scientific Research (C) Grant number 17K06932 from the Japan Society for the Promotion of Science. We thank the Edanz Group (www.edanzediting.com/ac) for editing a draft of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ojima, Y., Kawaguchi, T., Fukui, S. et al. Promoted performance of microbial fuel cells using Escherichia coli cells with multiple-knockout of central metabolism genes. Bioprocess Biosyst Eng 43, 323–332 (2020). https://doi.org/10.1007/s00449-019-02229-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-019-02229-z