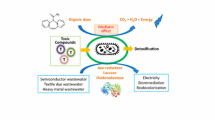
Abstract
Microbial fuel cell (MFC) is a promising technology that utilizes exoelectrogens cultivated in the form of biofilm to generate power from various types of sources supplied. A metal-reducing pathway is utilized by these organisms to transfer electrons obtained from the metabolism of substrate from anaerobic respiration extracellularly. A widely established model organism that is capable of extracellular electron transfer (EET) is Shewanella oneidensis. This review highlights the strategies used in the transformation of S. oneidensis and the recent development of MFC in terms of intervention through genetic modifications. S. oneidensis was genetically engineered for several aims including the study on the underlying mechanisms of EET, and the enhancement of power generation and wastewater treating potential when used in an MFC. Through engineering S. oneidensis, genes responsible for EET are identified and strategies on enhancing the EET efficiency are studied. Overexpressing genes related to EET to enhance biofilm formation, mediator biosynthesis, and respiration appears as one of the common approaches.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Research are being focused on microbial fuel cell (MFC) technology in recent years due to its potential in generating electricity from organic matters such as wastewater, as an alternative strategy for sustainable energy production. An MFC consists of an anode chamber and a cathode chamber, separated by a proton-exchange membrane (PEM). Microbes grow on the electrode surface in the anode chamber which is kept in anaerobic condition and consume substrates such as glucose via oxidation, producing carbon dioxide, protons, and electrons. The electrons are channeled to the cathode chamber, which is usually kept in aerobic condition, via an external circuit, and protons diffuse through the PEM to combine with the final electron acceptor such as molecular oxygen to form water (reduction process). The redox reaction thus generates electricity. MFC uses microorganisms in the form of biofilm that is capable of transferring electrons produced during anaerobic respiration extracellularly to an anode, known as extracellular electron transfer (EET). Due to the ability to transfer electrons, these microorganisms are termed as exoelectrogens.
Exoelectrogens that are widely studied with established genome database are Shewanella spp. and Geobacter spp., more specifically Shewanella oneidensis and Geobacter sulfurreducens [1]. The former is well known for its bidirectional flow of electrons across the cellular membranes [2,3,4,5,6], while the latter exchanges electrons through conductive nanowires to various electron acceptors [7]. Different substrates can be utilized depending on the strain of microorganisms used, where lactate, acetate, and glucose are the typical substrates used by exoelectrogens for anaerobic respiration [1]. Shewanella oneidensis was also found to be a potential exoelectrogen for generating bioelectricity from MFCs by degrading chitin and biomass hydrolysate, thus providing a new opportunity for the food waste and biomass industries [8, 9]. Additionally, more organisms that confer exoelectrogenic capabilities are being identified [10, 11]. Recent studies discovered that different exoelectrogens could be used to liberate electrons from cellulose, which is a complex polymer [12]. This process is achieved through a syntrophic consortium where interspecies communication leads to decomposition of cellulose into volatile acids through metabolic cooperation. Recently, Ueoka et al. [13] demonstrated an electrode plate-culture (EPC) method that allowed selective isolation of exoelectrogens specifically through utilizing an electrode plate covered with medium as a sole electron acceptor. The isolation of exoelectrogens in a complex microbiome was solely based on their capabilities to undergo EET, and not on their specific nutrient requirement commonly practiced at present. This method could thus potentially lead to more different exoelectrogens being isolated and subsequently identified.
Along with the refinement of techniques in molecular biology and the rapid advancement of biotechnology, it is only a matter of time where the metabolic pathways of various organisms are identified and cloned into a strain where it could be ubiquitously utilized as an exoelectrogen for efficient power generation in an MFC. At the current stage of research, the relationship and interactions among various exoelectrogens in a mixed-cultured biofilm are still not well defined. This signifies the importance to look into the metabolic pathway of various exoelectrogens at the DNA level and to also further establish the biological interactions between different species in a biofilm and how each of them plays a role in substrate oxidation. In this review, research studies on S. oneidensis leading to the discovery of the respective gene functions for EET are introduced. Comparison on overexpression of these genes for increased efficiency when applied in MFC is also conducted. Coupled with foreign gene conferring additional utilities such as substrate metabolism and mediator regeneration, these approaches had allowed manipulations of S. oneidensis from the biological aspects. These experimental findings when complemented with electrochemical manipulations will potentially allow the MFC technology to be applicable in an actual treatment plant for sustainable energy production.
Biofilm Formation and Metal-Reducing Pathway of S. oneidensis
Biofilms are referred to as a complex of embedded microorganisms that could be of a pure or mixed culture within an extracellular matrix composed of self-produced extracellular polymeric substances [14, 15]. Electroactive biofilms are imperative for exoelectrogens to efficiently carry out EET for bioelectricity generation. Bioelectricity generation is directly proportional to the biofilm and type of electrode surface where negatively charged and hydrophobic anode surfaces are less favorable for electroactive biofilm formation. An optimum thickness is preferable to achieve substantial current densities as very thick deposition of biofilm was shown to limit electron flow [15, 16]. The electron transfer processes are mediated by the c-type cytochromes (c-Cyts), also known as the redox-active proteins [17, 18]. Microbial proteins (such as type IV pili) and electron shuttles (such as flavins and pyocyanin) also enhance the conductivity of the biofilm, thus increasing electron transfer [15, 19, 20].
Electron transfer of S. oneidensis biofilm is based on both direct electron transfer (DET) and mediated electron transfer (MET). Marsili et al. [21] showed that the removal of electron shuttle flavin resulted in more than 70% loss in electron transfer in the biofilm. Three factors, including electrode material [22, 23], environmental parameters [23, 24], and biofilm thickness [25], affect the characteristics of biofilms formed by S. oneidensis. The genes msh (encodes mannose-sensitive hemagglutinin)/pilT and mxd (encodes a putative carbohydrate containing cell-associated component) are involved in surface attachment and biofilm formation [26, 27]. A cluster of chemotaxis genes allow the strain to sense the electron acceptors. Genes within the clusters which are essential for chemotactic response to electron acceptor during anaerobic respiration are cheA-3 and cheA-1, where CheA-3 histidine protein kinase will be expressed when the electron acceptors are present [28, 29]. The downregulation of motility protein flagellin and proteins associated with oxidative stress, along with the upregulation of quorum sensing proteins, riboflavin synthesizing proteins, heme, and transporter components ABC, causes metabolic shifts in S. oneidensis biofilms [30].
For S. oneidensis, a metal-reducing (mtr) conduit is responsible for the flow of current from the interior of cells to the outer membrane and subsequently to the extracellular anodes, and to minerals such as Fe(III), Mn(III), or Mn(IV) [31] via the outer membrane c-type cytochromes (OMCs) [32]. A six multi-haem c-Cyts consist of (i) CymA (inner membrane tetraheme c-Cyts), (ii) Fcc3, (iii) MtrA (periplasmic decaheme c-Cyts), (iv) MtrC, (v) OmcA and a small tetrahaem cytochrome (STC), and lastly (vi) porin-like outer membrane protein MtrB [33,34,35,36,37]. Electrons from the organic matters and the electron donor, typically lactate in the case of S. oneidensis, go through an intracellular electron transfer pathway, from NADH through the quinol pool in the inner membrane, subsequently through CymA where quinol is oxidized and the electrons are transferred to STC and Fcc3 [38,39,40,41], then to a terminal reductase complex consisting of MtrA, MtrB, and MtrC. While the exact pathway of the electrons being transferred from STC and Fcc3 is unsure, a study involving mutant STC and Fcc3 showed reduced efficiency in Fe(III) oxides or oxyhydroxides-reducing activities, indicating that they may play a role in the transportation of electrons from CymA to MtrA [35, 42]. This complex then transfers electrons from the periplasmic proteins to the surface of bacteria [43,44,45,46], where physical interactions of MtrC and OmcA enable electron transfer directly to Fe(III)-containing minerals (Fig. 1). To transport electrons exogenously to the surface of electrodes, S. oneidensis secretes co-factors to OMC, riboflavins (RF) for OmcA, and flavin mononucleotide (FMN) for MtrC, illustrated by the exhibition of complementary binding sites for these molecules on the cytochromes [17, 18, 48]. These flavins also act as an “electron shuttle” that mediates the transfer of electrons between the cells and the electrode. Recent studies showed that adding these electron shuttles exogenously or through amplifying their expression in S. oneidensis improved power generation [17, 18, 49].
Electrons produced from the oxidation of substrate catalyzed by CymA are transported to the MtrA-MtrB-MtrC complex, then to the surface of the terminal electron acceptor extracellularly, and subsequently for power generation. Reprinted with permission from Min et al. [47]. Copyright (2017) American Chemical Society
Transformation of S. oneidensis
Transformation of S. oneidensis is mainly carried out to study the genes essential for EET by gene repression or overexpression, with the aim of enhancing the overall power generation capabilities and wastewater treating potential of MFC. The important factors in a typical gene transformation experiment on S. oneidensis include promoter, plasmid, and transformation method. In most studies, researchers tend to use an isopropyl β-d-1-thiogalactopyranoside (IPTG)–inducible promoter element, PlacIq-lacIq-Ptac, to induce overexpression of inserted gene in the plasmid vector. Table 1 summarizes the recent approaches in engineering S. oneidensis for improved power generation in an MFC. West et al. [61] demonstrated the utilization of trimethylamine N-oxide (TMAO) as an inducer for EET by replacing the native mtrCAB promoter, which controls the expression of metal-reduction pathway responsible for EET with PtorF. PtorF regulates the expression of operon to encode the TMAO respiratory system in S. oneidensis. This transformation enables the overexpression mtr pathway in S. oneidensis to be induced by a range of TMAO concentrations, albeit with comparatively lower current produced than that of the WT strain.
Cao et al. [51] developed a synthetic plasmid toolkit for gene transformation experiments designed for S. oneidensis. In the study, different elements within the plasmid vector such as promoters, antibiotic resistant cassette, and replication origins which are typically used for a gene transformation procedure in S. oneidensis were evaluated and characterized using green fluorescent protein (GFP) as a reporter. This was further fine-tuned through the investigation by overexpressing the mtrCAB gene cluster using promoters of different strengths. Expression of mtrCAB at a moderate level showed 134% improvement in the maximum power density of an MFC, demonstrating the convenience of the toolkit for fine-tuning of gene expressions.
Similarly, Ng et al. [62] evaluated the performance of various plasmids and promoters through the expression of GFP proteins, and subsequently mtr pathway genes, in recombinant S. oneidensis. They reported that the replication origin repB along with pLacl promoter was able to drive expression of inserted genes with higher enzymatic activity as compared with the controlled strain. Plasmid DNA is delivered into cells most commonly through conjugation from Escherichia coli or electroporation. The latter is more convenient for transfer of both linear and circular DNA. Gene editing tools are recombineering (Lambda-red, Cre/lox) and via clusters of regularly interspaced short palindromic repeats (CRISPR). Lambda-red protein is a protein derived from phage lambda and consists of three parts: Exo (lambda exonuclease), Gam, and Beta. Exo degrades dsDNA from 5′ to 3′ ends, leaving a ssDNA at the recessed region; Gam prevents nucleases produced by E. coli from degrading linear dsDNA; and Beta binds to ssDNA regions produced by Exo and facilitates the recombination process by promoting the incorporation of product to target site (Fig. 2) [63].
a The three components of Lambda-red recombineering system with Exo, Beta, and Gam. Gam prevents digestion of linear DNA by endogenous RecBCD and SbcCD, Exo degrades linear dsDNA from 5′, and Beta promotes annealing of ssDNA created by Exo to a complementary target in the cell. b An overview of using this system to knockout a gene of interest with an ampicillin-resistant cassette
While this recombineering technique is generally employed in programming E. coli, Corts et al. [64] established the optimum conditions for making electrocompetent cells and demonstrated the application of Lambda-red recombineering in S. oneidensis by expressing recombinases under an arabinose inducible promoter pBAD and targeted the integrated lacZ gene from E. coli in the chromosome of S. oneidensis, then subsequently the native rpsL gene. The Cre/lox system is a site-specific recombinase containing the Cre protein originated from P1 bacteriophage which catalyzes the recombination at the loxP sequence [65]. So far, no research has utilized the Cre/lox system for programming S. oneidensis. However, Enyeart et al. [66] established the system of genome editing via targetrons and recombinases (GETR) where mobile group II introns, termed “targetrons,” worked in tandem with Cre/lox recombination in S. oneidensis. Retargetable mobile group II introns have been developed recently as a tool that can be designed to be inserted into a given DNA site at high efficiency. Intron-encoded proteins (IEP) are involved in self-splicing and in the process of “retrohoming” where the intron site reverses splices into DNA specifically [66]. Enyeart et al. [66] designed a targetron containing a loxP site that allowed gene insertion into ribosomal rrs in S. oneidensis. It was reported that the insertion was found in most copies of rrs gene in the genome, showing that the system could be potentially utilized for reprogramming S. oneidensis.
The CRISPR system allows targeted genome editing in a range of different organisms (Fig. 3) [68]. The first usage of the CRISPRi system was reported to repress mtr and biofilm formation genes, speF and uvrY, in order to establish a CRISPRi-sRNA method that allowed transcriptional and translational gene regulation in S. oneidensis [50]. Through CRISPRi, the repression of speF (encodes a putrescine biosynthesis gene that produces putrescene as a strategy employed by prokaryotes for active cell dispersal from biofilms) and uvrY, which is related to synthesis of surface polysaccharide that deters formation of biofilms, enhanced power generation of MFC by 65% when compared with the control strain. Li et al. [69] then further developed a similar system on gene interference based on CRISPR in which several genes within S. oneidensis were repressed and subsequently resulted in enhanced EET. This system assumed that electrons flux would split to diverse terminal electron acceptors during energy metabolism. Thus, not all electron flux would contribute to the extracellular electron acceptor for EET. By targeting gene sequences related to these competitive electron transfer pathways for interference, it allows the electron flux to be redirected to the desired EET pathway which can elevate the efficiency of EET. However, this synthetic S. oneidensis was not tested in MFC for power generation. These various tools in engineering S. oneidensis could be further exploited for more discoveries on EET-related genes, and manipulations could be performed to enhance the effect of the said genes with the objectives of increasing the overall potential of an MFC.
Mechanism of genome editing using the CRISPR/Cas9 system adopted from [67]. This editing system requires a single-guide RNA (sgRNA) that directs the Cas9 endonuclease to the specific site of interest in the genomic DNA, resulting in a double-strand break. With the presence of a donor DNA in trans, a transgenic DNA can be synthesized, whereas with the absence of a donor DNA, double-strand break will be repaired by the host cell, resulting in an insertion or deletion that will disrupt the ORF of a gene
Enhancing MFC Performance Through Engineering Biofilm Formation
One of the strategies in enhancing MFC performance is to promote biofilm formation in exoelectrogens through biotechnology. Bis-(3′-5′)-cyclic dimeric guanosine monophosphate (c-di-GMP) was reported in multiple literatures that they play a central role in bacterial biofilm formation by promoting expression of adhesive matrix components which in turn facilitate formation of bacterial biofilm [70]. Liu et al. [70] demonstrated an enhancement in power generation of MFC by S. oneidensis via the expression of heterologous YdeH, which is a diguanylate cyclase encoded by ydeH in E. coli, to catalyze the biosynthesis of c-di-GMP. It was reported that the YdeH-engineered strain with a constitutive promoter was able to produce maximum power density 2.8 times higher than that of the controlled strain [70].
Lin et al. [52] showed that the expression of porin gene, oprF from Pseudomonas aeruginosa in S. oneidensis, coupled with the addition of recombinant Saccharomyces cerevisiae that produces lactate with ethanol synthesis pathway knocked-out, was able to enhance power generation of MFC by up to 73% compared with the control. These studies showed that biofilm formation affects MFC performance, and heterologous expression of genes related to biofilm formation could be one of the potential strategies in enhancing its performance. One of the major bottlenecks in MFC technology remains to be the long-term robustness and efficiency of the biofilms. Ding et al. [71] demonstrated that the disruption of the putrescine biosynthesis gene, speF, allowed S. oneidensis to form highly cohesive biofilms. Studies on the detachment of S. oneidensis found out that in-frame deletions of the global transcriptional regulators ArcA and CRP resulted in severe impacts on the detachment of the mutants in arcA and crp. This suggested a role for the genes to play in the formation of biofilm [25]. These mutants were not tested on an MFC and could be a potential subject for consideration in future experimental design. Quorum sensing (QS) is a process which is often characterized alongside biofilm formation processes. As of now, QS in S. oneidensis is not fully characterized yet. This could be a topic to be further explored by transcriptomic profiling as it could serve as a potential process to be exploited to enhance biofilm formation.
Enhancing MFC Performance Through Engineering EET Pathway
It was previously described that c-Cyts CymA mediates EET and acts as a redox-active protein. Overexpression of the cymA gene in S. oneidensis MR-1 showed higher maximum power generated in MFC and higher growth rate of the synthetic MR-1 in the anodic chamber compared with wild type MR-1 [53]. The overexpression of cymA facilitates more electrons generated from anaerobic respiration to be transported extracellularly to the electron acceptor. Cyclic adenosine 3′,5′-monophosphate (cAMP) plays a very important role in a multitude of biological processes. Additionally, Cheng et al. [72] also reported that cyclic adenosine 3′,5′-monophosphate receptor protein, cAMP-CRP complex, regulates EET through upregulation of expression levels of c-Cyts and flavin synthetic pathway–related genes. EET performance was enhanced via elevating the cAMP level in synthetic S. oneidensis performed through the expression of an exogenous gene encoding adenylate cyclase from Beggiatoa sp. However, these strains were not tested in an MFC reactor.
The expression of the EET mediators, flavins, is able to increase the performance of MFC. As previously stated, flavins act as an electron shuttle for EET where S. oneidensis secretes electrons as a product of anaerobic respiration. These flavins facilitate the transportation of these electrons to the electrode. Riboflavin secretion by S. oneidensis was proposed to aid in taxis of the organisms towards insoluble electron acceptor through sensing the redox gradient in the environment [73]. Yang et al. [54] showed that flavin synthetic pathway from Bacillus subtilis, ribADEHC, expressed heterologously in S. oneidensis resulted in 13.2 times increase in power generation compared with WT. Min et al. [47] reported an increase in power generation of approximately 1.1 times when using expression of S. oneidensis in which both the flavin synthetic gene cluster, ribD-ribC-ribBA-ribE, and the mtr pathway gene cluster, mtrC-mtrA-mtrB, were co-expressed in S. oneidensis. Detachment of S. oneidensis cells from biofilms was also studied. It was reported that rapid decrease in the concentration of oxygen stimulated cell detachment. In-frame deletions of individual genes arcA, crp, and etrA were reported to allow the transformed S. oneidensis to be defective in the detachment response, where most profound effect was observed in the transformed cell with crp deleted [25].
Coupling both genetic modification with mixed microbial community was shown to further enhance MFC performance through mutualistic relationship. In [55], glycerol was firstly converted to lactate by Klebsiella pneumoniae and subsequently fed to S. oneidensis as the carbon source for power generation. Alcohol dehydrogenase gene adhE in K. pneumoniae was knocked out, and foreign lactate dehydrogenase gene (lhdD) and lactate transporter gene (lldP) gene were introduced. This modified K. pneumoniae was then co-cultured with modified S. oneidensis expressing a foreign gene (ribADEHC) from B. subtilis which encodes a biosynthetic flavin pathway. This combination of organisms when inoculated into MFC resulted in 217% improvement in power density as compared with the wild type consortium [55].
Enhancing MFC Performance Through Engineering Respiratory Mechanisms of S. oneidensis—NAD+
Sources of electrons transported through the mtr pathway are derived from dehydrogenases such as lactate dehydrogenase, or NAD(H/+) dehydrogenases [74]. In the past, the role of NADH in EET and under anoxic conditions was not well characterized. Recent research found out that S. oneidensis utilizes Nuo or Nqr1, which are NADH dehydrogenases, for growth and EET via anaerobic respiration of lactate in high-potential electron acceptor [74, 75].
Li et al. [56] recently demonstrated that modifications in expression of NADH enhanced power generation capabilities of MFC. This is due to the role of NADH in respiration and as an intracellular electron carrier and source for EET in S. oneidensis. NADH is regenerated by essential redox enzymes, such as GapA2, to serve as an electron acceptor in glycolysis. In Barrangou et al. [68], several genes were identified from carbon metabolism pathways and were overexpressed together, resulting in an overall enhancement in power generated from MFC and also increased regeneration of NADH in the mutant S. oneidensis. These genes were namely gapA2 and pflB from glycolysis, and mdh from TCA cycle. pflB encodes enzyme pyruvate–formate lyase which catalyzes the conversion of acetyl-CoA and formate from pyruvate. As formate is a reduced metabolite, it can be utilized as an electron donor to S. oneidensis for power generation purposes. To enhance the release of stored electrons from formate to anode, fdh* from Candida boidinii, which encodes a NAD+-dependent formate dehydrogenase, was also expressed exogenously, to facilitate the conversion of NADH and CO2 from formate. TCA cycle, also known as the Krebs cycle, is essential for ATP production in most organisms. The overexpression of the native gene mdh in S. oneidensis that encodes a NAD+-dependent malate dehydrogenase to convert oxaloacetate from malate showed promising results in enhancing power generating ability of an MFC. Overexpressing each of these genes individually in S. oneidensis was able to increase voltage output significantly when compared with a wild type strain by 26%, 28%, 9%, and 29% for gapA2, pflB, fdh*, and mdh respectively. Li et al. [56] then generated a mutant S. oneidensis with all these genes identified for the performance improvement of MFC. This mutant strain was reported to be able to enhance power generation by three times compared with the controlled WT S. oneidensis.
In a separate study, Li et al. [57] identified genes in the NAD+ biosynthetic pathway and subsequently overexpressed them to increase power generated from MFC through increasing NADH synthesized. Engineered genes were mainly identified from the salvage pathway through assimilating Nm, utilizing nicotinic acid (Na) as a precursor, and the universal biosynthesis pathway synthesizing NAD+. S. oneidensis was unable to utilize Na as a precursor due to the absence of Na transporter and nicotinate phosphoribosyltransferase that converts nicotinic acid mononucleotide (NaMN) from Na. Hence, Na assimilation pathway was reconstructed through ycel from B. subtilis that encodes a bifunctional niaP transporter for Na and Nm. Endogenous Nm metabolic pathway was constructed through gene pncB which encodes the nicotinate phosphoribosyltransferase from Salmonella typhimurium for the conversion of Na to NaMN and was introduced to create a mutant S. oneidensis. It was reported that these mutants harboring both ycel and pncB were able to generate voltage higher than that of the WT strain, showing increased intracellular NADH, NAD+, and NAD(H/+) levels at 78%, 44%, and 54% times, respectively. Li et al. [57] then subsequently overexpressed endogenous gene nadV (which encodes a Nm phosphoribosyltransferase) and pnc (which encodes the Nm mononucleotide deaminase) in the Nm utilization pathway. It was reported that both mutants harboring ycel, and with the genes ycel, nadV, and pncC, showed superior power outputs. For the universal biosynthesis pathway, nadE* and nadD* from E. coli were expressed heterogeneously along with native nadM in S. oneidensis. All genes identified from both pathways were assembled, resulting in mutant S. oneidensis strains that were able to produce maximum power density 4.4 times more as compared with the control strain. Additionally, intracellular NAD(H/+) pool size of the mutant S. oneidensis was increased by 2.1 times when compared with the WT strain [57].
Engineering S. oneidensis for Enhanced Utilities
For the enhancement of wastewater treating capabilities in MFC, several studies were conducted to engineer S. oneidensis, including the incorporation of foreign metabolic pathway to allow the organism to metabolize substrates that they normally do not utilize. An example of such cases would be the incorporation of xylose metabolic pathway into S. oneidensis which allowed xylose to be used as a substrate in an MFC, resulting in an improved power generation up to 2.1 mW/m2 [58]. This demonstrated the potential in reprogramming S. oneidensis with foreign metabolic pathways for utilization of different kinds of substrates commonly found in wastewater.
Microbial communities exhibit mutualistic properties in which the presence of mixed culture enhances the growth of biofilm in MFC [52, 55, 59]. This leads to the studies of interactions between different organisms and subsequently the development of engineering approaches to allow enhanced performance, or utilization of substrate for power generation that would otherwise not be possible. Parallel fermentation was carried out in which the co-cultivation of Streptococcus bovis with S. oneidensis allowed simultaneous fermentation of lactic acid from starch by S. bovis, followed by power generation by S. oneidensis up to 50 mW/m2 [60]. These experimental findings indicate that the combination of different organisms, coupled with genetic modifications in an MFC, can be further exploited to enhance the mutualistic interactions for improved power generation.
Computational Modeling
Computational modeling has been utilized to facilitate understanding of metabolism by S. oneidensis at the system level over the past decade. Pinchuk et al. [76] developed a genome-scale constraint-based model (referred as iSO783) for metabolism in S. oneidensis MR-1. This model investigated MR-1’s capabilities and metabolic behavior in the presence of oxygen and estimated ATP requirements in both growth and non-growth phases. This study integrated both computational and experimental approaches to determine that lactate-grown S. oneidensis aerobically used less efficient enzymes to stimulate electron flux for the generation of proton motive force. The modeling approach demonstrated the predictions of cell behavior in response to genetic or external environmental interventions, and enhanced the current understanding of microbial metabolism by integrating experimental data to refine metabolic pathways reconstructed from a genome annotation [76]. Ong et al. [77] then expanded the model based on updated genome annotation. They developed a core model for Shewanella sp. (referred as Core) using genome annotations of the 21 sequenced strains which represented the conserved metabolic functionalities of all strains of Shewanella. Besides, the study utilized a computational algorithm, Comparison of Networks by Gene Alignment (CONGA), to identify the functional difference in developed metabolic networks in order to reveal unique genetic and metabolic differences among different Shewanella strains. Recently, Gaffney et al. [78] published a thorough review on the exploration of computational approaches on microbial electrochemical cells, allowing a deeper understanding of the complex EET mechanisms.
Conclusions
The advancement in the field of molecular biology has enabled rapid development of strategies in engineering S. oneidensis which is important for the future development of MFC. There are several obstacles that need to be addressed from the biological, electrochemical, and physical aspects of the MFC technology. Coupling modifications on the electrochemical parameters with genetic manipulations and computational modeling can be the potential aspects where future research can venture into. The reviewed improvement technologies would certainly be beneficial to the evaluation studies, development, and integration of microbial fuel cells to the real industries for sustainable energy generation.
References
Santoro, C., Arbizzani, C., Erable, B., & Ieropoulos, I. (2017). Microbial fuel cells: from fundamentals to applications. A review. Journal of Power Sources, 356, 225–244. https://doi.org/10.1016/j.jpowsour.2017.03.109.
Jiao, Y., Qian, F., Li, Y., Wang, G., Saltikov, C. W., & Gralnick, J. A. (2011). Deciphering the electron transport pathway for graphene oxide reduction by Shewanella oneidensis MR-1. Journal of Bacteriology, 193(14), 3662–3665. https://doi.org/10.1128/JB.00201-11.
Schuetz, B., Schicklberger, M., Kuermann, J., Spormann, A. M., & Gescher, J. (2009). Periplasmic electron transfer via the c-type cytochromes MtrA and FccA of Shewanella oneidensis MR-1. Applied and Environmental Microbiology, 75(24), 7789–7796. https://doi.org/10.1128/AEM.01834-09.
Coursolle, D., Baron, B. D., Bond, R. D., & Gralnick, A. J. (2009). The Mtr respiratory pathway is essential for reducing flavins and electrodes in Shewanella oneidensis. Journal of Bacteriology, 192(2), 467–474. https://doi.org/10.1128/JB.00925-09.
Bretschger, O., Obraztsova, A., Sturm, C. A., Chang, I. S., Gorby, Y. A., Reed, S. B., Culley, D. E., Reardon, C. L., Barua, S., Romine, M. F., Zhou, J., Beliaev, A. S., Bouhenni, R., Saffarini, D., Mansfeld, F., Kim, B.-H., Fredrickson, J. K., & Nealson, K. H. (2007). Current production and metal oxide reduction by Shewanella oneidensis MR-1 wild type and mutants. Applied and Environmental Microbiology, 73(21), 7003–7012. https://doi.org/10.1128/AEM.01087-07.
Lies, D. P., Hernandez, M. E., Kappler, A., Mielke, R. E., Gralnick, J. A., & Newman, D. K. (2005). Shewanella oneidensis MR-1 uses overlapping pathways for iron reduction at a distance and by direct contact under conditions relevant for biofilms. Applied and Environmental Microbiology, 71(8), 4414–4426. https://doi.org/10.1128/AEM.71.8.4414-4426.2005.
Reguera, G., McCarthy, K. D., Mehta, T., Nicoll, J. S., Tuominen, M. T., & Lovley, D. R. (2005). Extracellular electron transfer via microbial nanowires. Nature, 435(7045), 1098–1101.
Li, S.-W., Zeng, R. J., & Sheng, G.-P. (2017). An excellent anaerobic respiration mode for chitin degradation by Shewanella oneidensis MR-1 in microbial fuel cells. Biochemical Engineering Journal, 118, 20–24.
Wang, Y.-Z., Shen, Y., Gao, L., Liao, Z.-H., Sun, J.-Z., & Yong, Y.-C. (2017). Improving the extracellular electron transfer of Shewanella oneidensis MR-1 for enhanced bioelectricity production from biomass hydrolysate. RSC Advances, 7(48), 30488–30494.
Sharma, S. C. D., Feng, C., Li, J., Hu, A., Wang, H., Qin, D., & Yu, C.-P. (2016). Electrochemical characterization of a novel exoelectrogenic bacterium strain SCS5, isolated from a mediator-less microbial fuel cell and phylogenetically related to Aeromonas jandaei. Microbes and Environments, 31(3), 213–225. https://doi.org/10.1264/jsme2.ME15185.
Sacco, N. J., Bonetto, M. C., & Cortón, E. (2017). Isolation and characterization of a novel electrogenic bacterium, Dietzia sp. RNV-4. PLoS One, 12(2), e0169955–e0169955. https://doi.org/10.1371/journal.pone.0169955.
Toczyłowska-Mamińska, R., Szymona, K., Król, P., Gliniewicz, K., Pielech-Przybylska, K., Kloch, M., & Logan, B. E. (2018). Evolving microbial communities in cellulose-fed microbial fuel cell. Energies, 11(1), 1–12. https://doi.org/10.3390/en11010124.
Ueoka, N., Kouzuma, A., & Watanabe, K. (2018). Electrode plate-culture methods for colony isolation of exoelectrogens from anode microbiomes. Bioelectrochemistry, 124, 1–6. https://doi.org/10.1016/j.bioelechem.2018.06.008.
Snider, R. M., Strycharz-Glaven, S. M., Tsoi, S. D., Erickson, J. S., & Tender, L. M. (2012). Long-range electron transport in Geobacter sulfurreducens biofilms is redox gradient-driven. Proceedings of the National Academy of Sciences, 109(38), 15467–15472. https://doi.org/10.1073/pnas.1209829109.
Parameswaran, P., Bry, T., Popat, S. C., Lusk, B. G., Rittmann, B. E., & Torres, C. I. (2013). Kinetic, electrochemical, and microscopic characterization of the thermophilic, anode-respiring bacterium Thermincola ferriacetica. Environmental Science & Technology, 47(9), 4934–4940. https://doi.org/10.1021/es400321c.
Wrighton, K. C., Thrash, J. C., Melnyk, R. A., Bigi, J. P., Byrne-Bailey, K. G., Remis, J. P., Schichnes, D., Auer, M., Chang, C. J., & Coates, J. D. (2011). Evidence for direct electron transfer by a Gram-positive bacterium isolated from a microbial fuel cell. Applied and Environmental Microbiology, 77(21), 7633–7639. https://doi.org/10.1128/AEM.05365-11.
Brutinel, E. D., & Gralnick, J. A. (2012). Shuttling happens: soluble flavin mediators of extracellular electron transfer in Shewanella. Applied Microbiology and Biotechnology, 93(1), 41–48. https://doi.org/10.1007/s00253-011-3653-0.
Kotloski, N. J., & Gralnick, J. A. (2013). Flavin electron shuttles dominate extracellular electron transfer by Shewanella oneidensis. mBio, 4. https://doi.org/10.1128/mBio.00553-12.
Malvankar, N. S., & Lovley, D. R. (2012). Microbial nanowires: a new paradigm for biological electron transfer and bioelectronics. ChemSusChem, 5(6), 1039–1046. https://doi.org/10.1002/cssc.201100733.
Orellana, R., Leavitt, J. J., Comolli, L. R., Csencsits, R., Janot, N., Flanagan, K. A., Gray, A. S., Leang, C., Izallalen, M., Mester, T., & Lovley, D. R. (2013). U(VI) reduction by diverse outer surface c-type cytochromes of Geobacter sulfurreducens. Applied and Environmental Microbiology, 79(20), 6369–6374. https://doi.org/10.1128/AEM.02551-13.
Marsili, E., Baron, D. B., Shikhare, I. D., Coursolle, D., Gralnick, J. A., & Bond, D. R. (2008). Shewanella secretes flavins that mediate extracellular electron transfer. Proceedings of the National Academy of Sciences, 105(10), 3968–3973. https://doi.org/10.1073/pnas.0710525105.
Dolch, K., Danzer, J., Kabbeck, T., Bierer, B., Erben, J., Förster, A. H., Maisch, J., Nick, P., Kerzenmacher, S., & Gescher, J. (2014). Characterization of microbial current production as a function of microbe–electrode-interaction. Bioresource Technology, 157, 284–292. https://doi.org/10.1016/j.biortech.2014.01.112.
Kipf, E., Koch, J., Geiger, B., Erben, J., Richter, K., Gescher, J., Zengerle, R., & Kerzenmacher, S. (2013). Systematic screening of carbon-based anode materials for microbial fuel cells with Shewanella oneidensis MR-1. Bioresource Technology, 146, 386–392. https://doi.org/10.1016/j.biortech.2013.07.076.
Rosenbaum, M., Cotta, M. A., & Angenent, L. T. (2010). Aerated Shewanella oneidensis in continuously fed bioelectrochemical systems for power and hydrogen production. Biotechnology and Bioengineering, 105, 880–888. https://doi.org/10.1002/bit.22621.
Thormann, K. M., Saville, R. M., Shukla, S., & Spormann, A. M. (2005). Induction of rapid detachment in Shewanella oneidensis MR-1 biofilms. Journal of Bacteriology, 187(3), 1014–1021. https://doi.org/10.1128/JB.187.3.1014-1021.2005.
Saville, R. M., Dieckmann, N., & Spormann, A. M. (2010). Spatiotemporal activity of the mshA gene system in Shewanella oneidensis MR-1 biofilms. FEMS Microbiology Letters, 308(1), 76–83. https://doi.org/10.1111/j.1574-6968.2010.01995.x.
Thormann, K. M., Saville, R. M., Shukla, S., Pelletier, D. A., & Spormann, A. M. (2004). Initial phases of biofilm formation in Shewanella oneidensis MR-1. Journal of Bacteriology, 186(23), 8096–8104. https://doi.org/10.1128/JB.186.23.8096-8104.2004.
Li, J., Romine, M. F., & Ward, M. J. (2007). Identification and analysis of a highly conserved chemotaxis gene cluster in Shewanella species. FEMS Microbiology Letters, 273(2), 180–186. https://doi.org/10.1111/j.1574-6968.2007.00810.x.
Tai, S.-K., Wu, G., Yuan, S., & Li, K.-C. (2010). Genome-wide expression links the electron transfer pathway of Shewanella oneidensis to chemotaxis. BMC Genomics, 11(1), 319. https://doi.org/10.1186/1471-2164-11-319.
De Vriendt, K., Theunissen, S., Carpentier, W., De Smet, L., Devreese, B., & Van Beeumen, J. (2005). Proteomics of Shewanella oneidensis MR-1 biofilm reveals differentially expressed proteins, including AggA and RibB. Proteomics, 5(5), 1308–1316. https://doi.org/10.1002/pmic.200400989.
Myers, C. R., & Nealson, K. H. (1988). Bacterial manganese reduction and growth with manganese oxide as the sole electron acceptor. Science, 240(4857), 1319–1321. https://doi.org/10.1126/science.240.4857.1319.
Lovley, D. R. (2012). Electromicrobiology. Annual Review of Microbiology, 66(1), 391–409. https://doi.org/10.1146/annurev-micro-092611-150104.
Myers, J. M., & Myers, C. R. (2000). Role of the tetraheme cytochrome CymA in anaerobic electron transport in cells of Shewanella putrefaciens MR-1 with normal levels of menaquinone. Journal of Bacteriology, 182(1), 67–75. https://doi.org/10.1128/JB.182.1.67-75.2000.
Myers, C. R., & Myers, J. M. (2002). MtrB is required for proper incorporation of the cytochromes OmcA and OmcB into the outer membrane of Shewanella putrefaciens MR-1. Applied and Environmental Microbiology, 68(11), 5585–5594. https://doi.org/10.1128/AEM.68.11.5585-5594.2002.
Sturm, G., Richter, K., Doetsch, A., Heide, H., Louro, R. O., & Gescher, J. (2015). A dynamic periplasmic electron transfer network enables respiratory flexibility beyond a thermodynamic regulatory regime. The ISME Journal, 9(8), 1802–1811.
Beliaev, A. S., Saffarini, D. A., McLaughlin, J. L., & Hunnicutt, D. (2001). MtrC, an outer membrane decahaem c cytochrome required for metal reduction in Shewanella putrefaciens MR-1. Molecular Microbiology, 39(3), 722–730. https://doi.org/10.1046/j.1365-2958.2001.02257.x.
Coursolle, D., & Gralnick, J. A. (2010). Modularity of the Mtr respiratory pathway of Shewanella oneidensis strain MR-1. Molecular Microbiology, 77, 995–1008. https://doi.org/10.1111/j.1365-2958.2010.07266.x.
Firer-Sherwood, M. A., Bewley, K. D., Mock, J.-Y., & Elliott, S. J. (2011). Tools for resolving complexity in the electron transfer networks of multiheme cytochromes c. Metallomics, 3(4), 344–348. https://doi.org/10.1039/C0MT00097C.
Marritt, S. J., Lowe, T. G., Bye, J., McMillan, D. G. G., Shi, L., Fredrickson, J., Zachara, J., Richardson, D. J., Cheesman, M. R., Jeuken, L. J. C., & Butt, J. N. (2012). A functional description of CymA, an electron-transfer hub supporting anaerobic respiratory flexibility in Shewanella. Biochemical Journal, 444(3), 465–474. https://doi.org/10.1042/BJ20120197.
McMillan, D. G. G., Marritt, S. J., Butt, J. N., & Jeuken, L. J. C. (2012). Menaquinone-7 is specific cofactor in tetraheme quinol dehydrogenase CymA. Journal of Biological Chemistry, 287(17), 14215–14225. https://doi.org/10.1074/jbc.M112.348813.
McMillan, D. G. G., Marritt, S. J., Firer-Sherwood, M. A., Shi, L., Richardson, D. J., Evans, S. D., Elliott, S. J., Butt, J. N., & Jeuken, L. J. C. (2013). Protein–protein interaction regulates the direction of catalysis and electron transfer in a redox enzyme complex. Journal of the American Chemical Society, 135(28), 10550–10556. https://doi.org/10.1021/ja405072z.
Leys, D., Meyer, T. E., Tsapin, A. S., Nealson, K. H., Cusanovich, M. A., & Van Beeumen, J. J. (2002). Crystal structures at atomic resolution reveal the novel concept of “electron-harvesting” as a role for the small tetraheme cytochrome c. Journal of Biological Chemistry, 277(38), 35703–35711. https://doi.org/10.1074/jbc.M203866200.
Hartshorne, R. S., Reardon, C. L., Ross, D., Nuester, J., Clarke, T. A., Gates, A. J., Mills, P. C., Fredrickson, J. K., Zachara, J. M., Shi, L., Beliaev, A. S., Marshall, M. J., Tien, M., Brantley, S., Butt, J. N., & Richardson, D. J. (2009). Characterization of an electron conduit between bacteria and the extracellular environment. Proceedings of the National Academy of Sciences, 106(52), 22169–22174. https://doi.org/10.1073/pnas.0900086106.
Ross, D. E., Ruebush, S. S., Brantley, S. L., Hartshorne, R. S., Clarke, T. A., Richardson, D. J., & Tien, M. (2007). Characterization of protein-protein interactions involved in iron reduction by Shewanella oneidensis MR-1. Applied and Environmental Microbiology, 73(18), 5797–5808. https://doi.org/10.1128/AEM.00146-07.
White, G. F., Shi, Z., Shi, L., Wang, Z., Dohnalkova, A. C., Marshall, M. J., Fredrickson, J. K., Zachara, J. M., Butt, J. N., Richardson, D. J., & Clarke, T. A. (2013). Rapid electron exchange between surface-exposed bacterial cytochromes and Fe(III) minerals. Proceedings of the National Academy of Sciences, 110(16), 6346–6351. https://doi.org/10.1073/pnas.1220074110.
Richardson, D. J., Butt, J. N., Fredrickson, J. K., Zachara, J. M., Shi, L., Edwards, M. J., White, G., Baiden, N., Gates, A. J., Marritt, S. J., & Clarke, T. A. (2012). The ‘porin–cytochrome’ model for microbe-to-mineral electron transfer. Molecular Microbiology, 85(2), 201–212. https://doi.org/10.1111/j.1365-2958.2012.08088.x.
Min, D., Cheng, L., Zhang, F., Huang, X. N., Li, D. B., Liu, D. F., Lau, T. C., Mu, Y., & Yu, H. Q. (2017). Enhancing extracellular electron transfer of Shewanella oneidensis MR-1 through coupling improved flavin synthesis and metal-reducing conduit for pollutant degradation. Environmental Science and Technology, 51(9), 5082–5089. https://doi.org/10.1021/acs.est.6b04640.
Okamoto, A., Kalathil, S., Deng, X., Hashimoto, K., Nakamura, R., & Nealson, K. H. (2014). Cell-secreted flavins bound to membrane cytochromes dictate electron transfer reactions to surfaces with diverse charge and pH. Scientific Reports, 4, 5628.
Kumar, R., Singh, L., & Zularisam, A. W. (2016). Exoelectrogens: recent advances in molecular drivers involved in extracellular electron transfer and strategies used to improve it for microbial fuel cell applications. Renewable and Sustainable Energy Reviews, 56, 1322–1336. https://doi.org/10.1016/j.rser.2015.12.029.
Cao, Y., Li, X., Li, F., & Song, H. (2017). CRISPRi-sRNA: transcriptional-translational regulation of extracellular electron transfer in Shewanella oneidensis. ACS Synthetic Biology, 6(9), 1679–1690. https://doi.org/10.1021/acssynbio.6b00374.
Cao, Y., Song, M., Li, F., Li, C., Lin, X., & Song, H. (2019). A synthetic plasmid toolkit for Shewanella oneidensis MR-1. Frontiers in Microbiology, 10, 410.
Lin, T., Bai, X., Hu, Y., Li, B., Yuan, Y. J., Song, H., Yang, Y., & Wang, J. (2017). Synthetic Saccharomyces cerevisiae-Shewanella oneidensis consortium enables glucose-fed high-performance microbial fuel cell. AICHE Journal, 63(6), 1830–1838. https://doi.org/10.1002/aic.15611.
Vellingiri, A., Song, Y. E., Munussami, G., Kim, C., Park, C., Jeon, B. H., Lee, S. G., & Kim, J. R. (2019). Overexpression of c-type cytochrome, CymA in Shewanella oneidensis MR-1 for enhanced bioelectricity generation and cell growth in a microbial fuel cell. Journal of Chemical Technology & Biotechnology, 94(7), 2115–2122.
Yang, Y., Ding, Y., Hu, Y., Cao, B., Rice, S. A., Kjelleberg, S., & Song, H. (2015). Enhancing bidirectional electron transfer of Shewanella oneidensis by a synthetic flavin pathway. ACS Synthetic Biology, 4(7), 815–823. https://doi.org/10.1021/sb500331x.
Li, F., Yin, C., Sun, L., Li, Y., Guo, X., & Song, H. (2018). Synthetic Klebsiella pneumoniae-Shewanella oneidensis consortium enables glycerol-fed high-performance microbial fuel cells. Biotechnology Journal, 13(5), 1700491.
Li, F., Li, Y., Sun, L., Chen, X., An, X., Yin, C., Cao, Y., Wu, H., & Song, H. (2018). Modular engineering intracellular NADH regeneration boosts extracellular electron transfer of Shewanella oneidensis MR-1. ACS Synthetic Biology, 7(3), 885–895. https://doi.org/10.1021/acssynbio.7b00390.
Li, F., Li, Y.-X., Cao, Y.-X., Wang, L., Liu, C.-G., Shi, L., & Song, H. (2018). Modular engineering to increase intracellular NAD (H/+) promotes rate of extracellular electron transfer of Shewanella oneidensis. Nature Communications, 9(1), 3637.
Li, F., Li, Y., Sun, L., Li, X., Yin, C., An, X., Chen, X., Tian, Y., & Song, H. (2017). Engineering Shewanella oneidensis enables xylose-fed microbial fuel cell. Biotechnology for Biofuels, 10(1), 1–10. https://doi.org/10.1186/s13068-017-0881-2.
Kim, C., Song, Y. E., Lee, C. R., Jeon, B.-H., & Kim, J. R. (2016). Glycerol-fed microbial fuel cell with a co-culture of Shewanella oneidensis MR-1 and Klebsiella pneumonae J2B. Journal of Industrial Microbiology & Biotechnology, 43(10), 1397–1403.
Uno, M., Phansroy, N., Aso, Y., & Ohara, H. (2017). Starch-fueled microbial fuel cells by two-step and parallel fermentation using Shewanella oneidensis MR-1 and Streptococcus bovis 148. Journal of Bioscience and Bioengineering, 124(2), 189–194.
West, E. A., Jain, A., & Gralnick, J. A. (2017). Engineering a native inducible expression system in Shewanella oneidensis to control extracellular electron transfer. ACS Synthetic Biology, 6(9), 1627–1634. https://doi.org/10.1021/acssynbio.6b00349.
Ng, I.-S., Guo, Y., Zhou, Y., Wu, J.-W., Tan, S.-I., & Yi, Y.-C. (2018). Turn on the Mtr pathway genes under pLacI promoter in Shewanella oneidensis MR-1. Bioresources and Bioprocessing, 5(1), 35. https://doi.org/10.1186/s40643-018-0221-9.
Sawitzke, J. A., Thomason, L. C., Costantino, N., Bubunenko, M., Datta, S., & Court, D. L. (2007). Recombineering: in vivo genetic engineering in E. coli, S. enterica, and beyond. Methods in Enzymology, 421. https://doi.org/10.1016/S0076-6879(06)21015-2.
Corts, A. D., Thomason, L. C., Gill, R. T., & Gralnick, J. A. (2019). A new recombineering system for precise genome-editing in Shewanella oneidensis strain MR-1 using single-stranded oligonucleotides. Scientific Reports, 9(1), 1–10. https://doi.org/10.1038/s41598-018-37025-4.
Abremski, K., & Hoess, R. (1984). Bacteriophage P1 site-specific recombination. Purification and properties of the Cre recombinase protein. The Journal of Biological Chemistry, 259(3), 1509–1514.
Enyeart, P. J., Chirieleison, S. M., Dao, M. N., Perutka, J., Quandt, E. M., Yao, J., Whitt, J. T., Keatinge-Clay, A. T., Lambowitz, A. M., & Ellington, A. D. (2013). Generalized bacterial genome editing using mobile group II introns and Cre-lox. Molecular Systems Biology, 9(1), 1–16. https://doi.org/10.1038/msb.2013.41.
Costa JR, Bejcek BE, McGee JE, Fogel AI, Brimacombe KR, Ketteler R (2004) Genome editing using engineered nucleases and their use in genomic screening. Assay Guidance Manual 1–24.
Barrangou, R., Fremaux, C., Deveau, H., Richards, M., Boyaval, P., Moineau, S., Romero, D. A., & Horvath, P. (2007). CRISPR provides acquired resistance against viruses in prokaryotes. Science (New York, N.Y.), 315, 1709–1712. https://doi.org/10.1126/science.1138140.
Li, J., Tang, Q., Li, Y., Fan, Y.-Y., Li, F.-H., Wu, J.-H., Min, D., Li, W.-W., Lam, P. K., & Yu, H.-Q. (2020). Rediverting electron flux with an engineered CRISPR-ddAsCpf1 system to enhance the pollutant degradation capacity of Shewanella oneidensis. Environmental Science & Technology, 54(6), 3599–3608.
Liu, T., Yu, Y. Y., Deng, X. P., Ng, C. K., Cao, B., Wang, J. Y., Rice, S. A., Kjelleberg, S., & Song, H. (2015). Enhanced Shewanella biofilm promotes bioelectricity generation. Biotechnology and Bioengineering, 112(10), 2051–2059.
Ding, Y., Peng, N., Du, Y., Ji, L., & Cao, B. (2014). Disruption of putrescine biosynthesis in Shewanella oneidensis enhances biofilm cohesiveness and performance in Cr(VI) immobilization. Applied and Environmental Microbiology, 80(4), 1498–1506. https://doi.org/10.1128/AEM.03461-13.
Cheng, Z. H., Xiong, J. R., Min, D., Cheng, L., Liu, D. F., Li, W. W., Jin, F., Yang, M., & Yu, H. Q. (2020). Promoting bidirectional extracellular electron transfer of Shewanella oneidensis MR-1 for hexavalent chromium reduction via elevating intracellular cAMP level. Biotechnology and Bioengineering 117(5), 1294–1303. https://doi.org/10.1002/bit.27305.
Oram, J., & Jeuken, L. J. (2019). Tactic response of Shewanella oneidensis MR-1 toward insoluble electron acceptors. MBio, 10(1), e02490–e02418.
Madsen CS, TerAvest MA (2019) NADH dehydrogenases contribute to extracellular electron transfer by Shewanella oneidensis MR-1 in bioelectrochemical systems. bioRxiv:657668.
Hirose, A., Kasai, T., Aoki, M., Umemura, T., Watanabe, K., & Kouzuma, A. (2018). Electrochemically active bacteria sense electrode potentials for regulating catabolic pathways. Nature Communications, 9(1), 1–10. https://doi.org/10.1038/s41467-018-03416-4.
Pinchuk, G. E., Hill, E. A., Geydebrekht, O. V., De Ingeniis, J., Zhang, X., Osterman, A., Scott, J. H., Reed, S. B., Romine, M. F., & Konopka, A. E. (2010). Constraint-based model of Shewanella oneidensis MR-1 metabolism: a tool for data analysis and hypothesis generation. PLoS Computational Biology, 6(6), e1000822.
Ong, W. K., Vu, T. T., Lovendahl, K. N., Llull, J. M., Serres, M. H., Romine, M. F., & Reed, J. L. (2014). Comparisons of Shewanella strains based on genome annotations, modeling, and experiments. BMC Systems Biology, 8(1), 31.
Gaffney, E., Grattieri, M., Rhodes, Z., & Minteer, S. D. (2020). Review—exploration of computational approaches for understanding microbial electrochemical systems: opportunities and future directions. Journal of the Electrochemical Society, 167(6). https://doi.org/10.1149/1945-7111/ab872e.
Author information
Authors and Affiliations
Contributions
Conceptualization and writing—original draft preparation: Dexter Hoi Long Leung; supervision: Yin Sze Lim; data analysis: Kasimayan Uma; resources: Ja-Hon Lin and Guan-Ting Pan; writing—review and editing: Siewhui Chong; funding acquisition: Thomas Chung-Kuang Yang.
Corresponding authors
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Leung, D.H.L., Lim, Y.S., Uma, K. et al. Engineering S. oneidensis for Performance Improvement of Microbial Fuel Cell—a Mini Review. Appl Biochem Biotechnol 193, 1170–1186 (2021). https://doi.org/10.1007/s12010-020-03469-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-020-03469-6