Abstract
Genomic analyses have become an important tool to identify new avenues for therapy. This is especially true for cancer types with extremely poor outcomes, since our lack of effective therapies offers no tangible clinical starting point to build upon. The highly malignant brain tumor glioblastoma (GBM) exemplifies such a refractory cancer, with only 15 month average patient survival. Analyses of several hundred GBM samples compiled by the TCGA (The Cancer Genome Atlas) have produced an extensive transcriptomic map, identified prevalent chromosomal alterations, and defined important driver mutations. Unfortunately, clinical trials based on these results have not yet delivered an improvement on outcome. It is, therefore, necessary to characterize other regulatory routes known for playing a role in tumor relapse and response to treatment. Alternative splicing affects more than 90% of the human coding genes and it is an important source for transcript variation and gene regulation. Mutations and alterations in splicing factors are highly prevalent in multiple cancers, demonstrating the potential for splicing to act as a tumor driver. As a result, numerous genes are expressed as cancer-specific splicing isoforms that are functionally distinct from the canonical isoforms found in normal tissue. These include genes that regulate cancer-critical pathways such as apoptosis, DNA repair, cell proliferation, and migration. Splicing defects can even induce genomic instability, a common characteristic of cancer, and a driver of tumor evolution. Importantly, components of the splicing machinery are targetable; multiple drugs can inhibit splicing factors or promote changes in splicing which could be exploited to begin improving clinical outcomes. Here, we review the current literature and present a case for exploring RNA processing as therapeutic route for the treatment of GBM.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Glioblastoma genomics
Glioblastoma (GBM) is the most common and most lethal cerebral malignancy. According to Word Health Organization (WHO 2016), a GBM patients’ life expectancy is about 15 month after diagnosis, and less than 5%, of patients survive more than 5 years (WHO). Currently, GBM is diagnosed through advanced imaging techniques, including computing tomography (CT) and magnetic resonance imaging (MRI). The primary treatment regimen is surgical removal of the tumor, followed by radiotherapy and/or chemotherapy (WHO, ABTA 2016). Unfortunately, many other factors such as high vascularity, aggressive proliferation and invasion, and the notorious heterogeneity that characterize GBM contribute to therapeutic resistance (Sottoriva et al. 2012; Xie and Mittal 2014; Motaln et al. 2015). This inefficacy of current therapies emphasizes urgent need for new discoveries leading to more efficient routes of treatment.
The Cancer Genome Atlas (TCGA) is a massive genomic and transcriptomic project started in 2005 by the National Cancer Institute (NCI) and the National Human Genome Research Institute (NHGRI) with the purpose of assembling genomic information about multiple tumor types, including GBM. The TCGA classifies GBM into four subgroups: classical, mesenchymal, neural, and pro-neural. Tumors in the classical subgroup present the most common genomic abnormalities. They are characterized by high expression of the epidermal growth factor receptor EGFR, but do not contain TP53 mutations, commonly observed in other GBM subgroups. Mesenchymal tumors tend to be more aggressive and often display mutations in NFI, PTEN, and TP53 and display high expression of CHI3L1 and MET. The neural subgroup typically expresses several genes commonly observed in normal cerebral tissue, non-tumor nerve cells, or neurons, whereas the pro-neural subtype is associated with younger patient and is characterized by frequent mutations in IDH1, PDGFRA, and TP53 (Verhaak et al. 2010). Other studies have categorized GBM mutations based on survival time (Kim et al. 2013, Bao et al. 2013). In long survivor patients, the amplification of CDK4 and of EGFR, deletion of CDKN2A, increased expression of DNM1 and mitogen-activated protein kinase 1 (MAPK1), and decreased expression of HSPA9, PSMD3, and CANX have been observed (Brennan et al. 2013; Patel et al. 2013). Overexpression of PI3K/AKT pathway-associated genes, including NEK9 and PI3KCB, of ribosomal proteins, including RPS11 and RPS20, and of some survival kinase genes are genomic characteristics of treatment-resistant patients. Mutations in the telomerase reverse transcriptase gene, TERT, were also observed in resistant tumors (Yong et al. 2015; Simon et al. 2015; Varghese et al. 2016). Interestingly, these mutations can occur in the non-protein-coding space; in fact, TERT promoter mutations may be the most common mutations in GBM.
This set of studies and findings is building the profile of differentially expressed and mutated genes in GBM. In addition to these items, scientists are now focusing their efforts upon comprehension of epigenetic and RNA processing changes, including DNA methylation, aberrant miRNA expression, and more recently alterations in splicing pattern and polyadenylation [poly(A)] site usage.
RNA-processing regulators in glioblastoma
Changes in the expression levels and activity of RNA binding proteins can trigger multiple alterations in RNA processing, thus contributing to the acquisition of cancer-relevant phenotypes. Expression analysis of more than 300 RBPs in normal vs. tumor tissues showed that 36 of them are upregulated in brain tumors (Galante et al. 2009). More recently, using TCGA data, Kechavarzi and Janga (2014) have studied approximately 850 RBPs in 16 different tissues from the Human BodyMap 2.0 Project. First, they showed that RBPs are differentially expressed at significantly higher levels compared to other classes of genes, including regulators, such as transcription factors, suggesting a key role in controlling gene expression. Next, the authors demonstrated that a set of 30 RBPs is strongly upregulated across at least two-thirds of the nine cancers profiled. Highly upregulated RBPs were related to gene expression, transcriptional deregulation and transport of biomolecules, cellular regulation, and proliferation (Kechavarzi and Janga 2014). Cheung and collaborators (2008) compared ten GBM and ten non-tumor tissues via microarray analysis and identified three splicing regulators associated with GBM-specific splicing events. In a study focusing exclusively on glioblastoma, we studied over 1500 RBPs and their expression profile in tumor versus normal brain samples (Correa et al. 2016). We identified 223 upregulated and 135 downregulated RBPs in tumors compared to normal brain, as well as 275 upregulated and 85 downregulated RBPs found in glioma stem cells (GSCs) compared to normal neural progenitor cells. We determined 58 RBPs to be upregulated in tumor cells in both analyses, 21 of which were associated with poor prognosis. Their impact on cell viability, proliferation, and apoptosis revealed the small nuclear ribonucleoprotein-associated proteins B (SNRPB), a core component of the spliceosome, as the main effectors. Knockdown of SNRPB had a strong impact on alternative splicing events, preferentially affecting genes involved in RNA processing, DNA repair, and chromatin modulation. The exon junction complex proteins MAGOH and MAGOHB and the splicing component SNRPG were also implicated as possible oncogenic factors (Correa et al. 2016). A similar effect has been observed in lung cancer, where high expression of SNRPB along with other RNA-processing regulators was significantly associated with reduced overall survival (Valles et al. 2012).
A major player in RNA processing and cancer development is the Polypyrimidine-tract-binding protein, PTB (Kafasla et al. 2012) which functions at the intersection between neurogenesis and brain tumor development. PTB is poorly expressed in cerebral tissues, but highly expressed in low-grade astrocytoma, anaplastic astrocytoma, glioblastoma multiforme, medulloblastoma, paraganglioma, and the glial population of both ganglioglioma and dysplastic gangliocytoma (McCutcheon et al. 2004; Izaguirre et al. 2012; Fontana et al. 2015). High expression of PTB is also observed in tumors outside the nervous system. In ovarian and breast cancers, for instance, it is overexpressed and its knockdown led to a decrease in growth, colony formation, and invasiveness (He et al. 2007, 2014). PTB binds to polypyrimidine tracts located mainly in introns and can influence exon inclusion (Kafasla et al. 2012). PTB overexpression in glioma tissues and GSCs alters the regulation of microtubule dynamics through MARK4 by causing an increase in the production of the MARK4L isoform, thus enhancing cell proliferation (Fontana et al. 2015). Neoplastic transformation of glial cells affects the inclusion of the alpha exon in human FGFR-1 mRNA (Jin et al. 1999). This alpha exon is included in FGFR-1 mRNA transcripts in normal cells, but not in GBM cells thanks to the action of PTB. The new isoform generates a high-affinity receptor that confers a cell growth advantage (Jin et al. 2000).
A member of the large family of heterogeneous nuclear ribonucleoproteins, hnRNPH, is upregulated in gliomas (Lefave et al. 2011; Golan-Gerstl et al. 2011). It contributes to invasion and survival of tumor cells and impacts tumor growth. In GBM, the death-domain adaptor protein insuloma–glucagonoma protein 20 (IG20) is consistently aberrantly spliced and generates an antagonist, anti-apoptotic isoform (MAP-kinase activating death-domain protein, MADD), redirecting TNF-α/TRAIL-induced death signaling to promote survival and proliferation. This switch in splicing is regulated by hnRNPH. Similarly, hnRNPH regulates the splicing of RON tyrosine kinase receptor, generating a variant that promotes migration and invasion (Lefave et al. 2011; Golan-Gerstl et al. 2011). hnRNPH participation in tumorigenesis is likely very complex, since RNA-seq, CLIP, and proteomic analyses revealed an elaborate network containing a large number of hnRNPH targets regulated at multiple levels. Although splicing regulation is the main mechanism by which hnRNPH regulates gene expression, poly(A) site selection, and translation are also employed (Uren et al. 2016).
HuR (ELAVL1) is probably the most well-characterized RNA binding protein and is the only member of the ELAV family that is ubiquitously expressed (Grammatikakis et al. 2017). HuR is a polyvalent RNA binding protein, acting at the level of RNA stability, translation, and RNA processing and multiple binding sites for HuR have been identified in intronic regions. Several research groups performed genomic analyses of HuR (RNA-seq and CLIP), establishing a large list of targets that corroborate its involvement in multiple biological processes, including apoptosis, proliferation, RNA processing and metabolism, cell cycle, angiogenesis, and inflammation (Lebedeva et al. 2011; Mukherjee et al. 2011; Uren et al. 2011). RNA-seq analysis of HuR knockdown cells indicating changes in splicing, supporting then an additional role for HuR as an RNA-processing regulator (Srikantan and Gorospe 2011). HuR is overexpressed in highly malignant tumors, including GBM, and is related to poor survival (Abdelmohsen and Gorospe 2010; Vo et al. 2012; Grammatikakis et al. 2017). HuR knockdown in GBM cells decreased anchorage-independent growth and cell proliferation, induced apoptosis, and reduced tumor volume in a xenograft assay. Conversely, overexpression of HuR induced chemo-resistance to standard glioma therapies (Filippova et al. 2011).
RBM14 is an RNA binding protein that interacts with the transcriptional co-regulator TRBP and regulates transcription and splicing in a promoter-preferential manner, affecting the expression of steroid hormones (Auboeuf et al. 2002). It is homologous to the oncoproteins EWS and TLS. The RBM14 gene is amplified at the chromosome 11q13 locus in a subset of primary human cancers, including non-small cell lung carcinoma, squamous cell skin carcinoma, and lymphoma, and RBM14 has been shown to have transforming activities in soft agar assays (Su et al. 2007). On the other hand, RBM14 is significantly decreased in human renal cell carcinoma when compared with normal kidney. In this context, it seems to function as tumor suppressor. RBM14 inhibits G(1)-S transition in human kidney cells and suppresses anchorage-independent growth and xenograft tumor formation, in part via inhibition of MYC and its downstream effectors CCND1 and SKP2 (Kang et al. 2008). RBM14 is highly expressed in embryonic tissues and stem cells and is necessary to maintain the stem-like state of GBM spheres. RBM14 knockdown affects GBM sphere size, reduces tumorigenesis, and increases the sensibility of GBM stem-like cells in vivo (Yuan et al. 2014). RBM14 was identified in a screen as an inhibitor of neurite outgrowth (Simpson et al. 2015). This multi-functional protein has also been recently connected to genomic instability, DNA repair, and radio-resistance in the context of GBM (Kai 2016). RBM14 was also identified as a novel suppressor of assembly of centriolar protein complexes, where its depletion induces ectopic formation of centriolar protein complexes via the STIL/CPAP complex. A reduction in RBM14 levels makes GBM cells more sensitive to radiation, since it stimulates DNA repair by controlling the DNA-PK-dependent non-homologous end-joining (NHEJ) pathway (Yuan et al. 2014; Shiratsuchi et al. 2015).
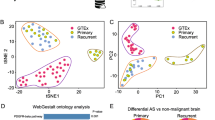
Figure 1 summarizes the regulatory roles of the RBPs here described. In Fig. 2, we show their expression profile in gliomas and survival plots. High expression of hnRNPH1, PTBP1, SNRPB, and ELAVL1 is linked to poor survival. As expected, the expression of these RBPs is higher in GBM in respect to gliomas grades II and III, while there is decreased expression of NUDT in higher grades.
RNA-processing regulators and relevance to GBM development. Expression levels of RBM14, hnRNPH1, PTB, SNRPB, HuR (ELAVL1), and NUDT21 (CFIm25) were assessed in grade II (red boxplot), III (green boxplot), and IV (blue boxplot) gliomas, as well as patient survival time according to gene expression level. Boxplots and survival plots were created using GlioVis portal (Bowman et al. 2017). The data set used in those analyses was from adult human data from TCGA_GBMLGG multiple data sets combined. On the right: survival curve analysis was based on Kaplan–Meier estimation and was considered significant if long-rank and Wilcoxon p values were less than or equal to 0.05. We considered all histology groups (oligodendroglioma, oligoastrocytoma, astrocytoma, and GBM) of all subtypes (mesenchymal, classical, and pro-neural) and based the chosen cutoff on the median value. The survival curve represents the number of survivors (Y axis) along the time (X axis). On the left: the box plots were based on mRNA expression data from RNA-seq platform. We used a predefined plot based on histology and compared glioma grades II, III, and IV
A recent article from Bi and collaborators (2017) has reinforced the importance of RNA-processing factors in glioblastoma development. Using label-free quantitative proteomics, they have detected 136 differentially expressed proteins between GBM and low-grade gliomas. Among them are several proteins implicated in RNA processing (SNRPA1, SNRPB2, SF3A1, hnRNPL, EFTUD2, SF3B3, PRPF8, and SF3B2). Through network analysis, the authors have linked RNA-processing regulators to EGFR, STAT1, and MAPK1 and signal transduction pathways implicated in GBM development. It is important to emphasize that these splicing factors were found to be in the central position of the network connecting RNA processing and neuronal structure and function.
All examples listed above refer to RNA-processing factors implicated in oncogenic activities. Unfortunately, regulators acting as tumor suppressors in the context of GBM have not received much attention. Analysis of TCGA data indicates that several RBPs implicated in RNA processing are strongly downregulated in GBM (Correa et al. 2016), and in many cases, their low levels of expression are associated with poor prognosis. Table 1 shows 63 of those RBPs downregulated in GBM compared to normal tissue. These are labeled according to function and we indicate the relation of each to alternative splicing, RNA processing, and binding to poly(A) tail. This list is likely ripe for functional screening to identify new players in gliomagenesis.
Alternative splicing events in glioblastoma development
RNA-processing regulation occurs in a specific manner by a complex RNA–protein network according to cellular and tissue context. Changes in this delicate balance can lead to disease states and cancer (Yeo et al. 2016). In fact, alternative and aberrant splicing can affect several important players in tumor initiation and growth (Yu et al. 2007; Danan-Gotthold et al. 2015; Tsai et al. 2015; Yeo et al. 2016). For example, changes in expression or function of RNA-processing factors as well as mutations in splice sites, regulatory elements, and RNA binding protein sites are responsible for most of the alterations observed in cancer cells (Brooks et al. 2014; Kechavarzi and Janga 2014; Weinhold et al. 2014; Darman et al. 2015).
Large-scale expression analyses in multiple tumor types have produced a catalogue of splicing isoforms related to malignant transformation (Dorman et al. 2014; Brooks et al. 2014, Tsai et al. 2015; DiFeo et al. 2009, Dargahi et al. 2014). Similarly, splicing regulators displaying mutations or differential expression in cancer cells have been identified. The most common examples include SF3B1 SRSF1, RBM4, RBM5/6/10, U2AF1, and the splicing kinases clk/STY and CLK2 (Tsai et al. 2015; Brooks et al. 2014; Garcia-Sacristan et al. 2005; Yoshida et al. 2015). Interestingly, an examination of TCGA RNA-seq data from eight distinct cancer types showed several shared splicing alterations produced by the RNA binding proteins (RBPs) RBFOX2, QKI, MBNL1/2, PTBP1, and CELF2 (Danan-Gotthold et al. 2015), suggesting common malignant routes via splicing regulation.
Changes in splicing mechanisms have also been widely associated with GBM malignancy. Genome-wide analysis from exon expression array defined a set of 14 genes with splicing alterations prevalent in GBM samples (Cheung et al. 2008). In another large-scale study, 117 genes have shown to display both splicing and expression alterations in both GBM and oligodendroma. Many of these genes belong to categories regulating neuron differentiation, exocytosis, and regulation of neurotransmitter secretion. Several of the most significantly upregulated and downregulated genes were suggested by the authors as possible GBM biomarkers (Yu and Fu 2015). In another study, expression analysis of 250 GBM patients identified 2477 genes with alternative exon usage in tumors. Overall, the genes displaying alternative exon usage were related to multiple pathways involved in cancer establishment, including cell adhesion, regulation of Ras and Rho signaling, cytoskeleton organization, chromatin modification, and oxidative phosphorylation (Sadeque et al. 2012).
Another series of studies focused on splice alterations of specific GBM relevant genes, particularly those of the epidermal growth factor (EGF) signaling pathway. Several genes associated with EGF are regulated by alternative splicing and play relevant functions in GBM development. EGF itself undergoes alternative exon usage in GBM resulting in variants that increase GBM survival or overall increased EGF expression that increases GBM survival and resistance (Sadeque et al. 2012). In a related example, splicing of a brain-enriched cassette exon in the membrane-binding tumor suppressor annexin A7 (ANXA7) decreases endosomal targeting of EGFR, enhancing EGFR signaling during glioblastoma development (Ferrarese et al. 2014). The EGFRvIII mutant isoform is highly prevalent in GBM (Padfield et al. 2015) and its expression results in genome-wide alterations in alternative splicing patterns. One such EGFRvIII-induced change upregulates the heterogeneous nuclear ribonucleoprotein (hnRNP) A1, which in turn promotes the splicing of the transcript encoding the MYC-interacting partner Max, thus generating Delta Max. Delta Max promotes glioma cell proliferation and GBM growth by enhancing tumorigenic functions of MYC (Babic et al. 2013). In addition, CD97, a member of the epidermal growth factor seven-span transmembrane (EGF-TM7) family, is implicated in cell adhesion and migration, and is overexpressed in classical and mesenchymal GBM subtypes. Five extracellular EGF-like domains can be alternatively spliced to generate different isoforms. Two of CD97 isoforms, known as EGF (1,2,5) and EGF (1,2,3,5), are found in GBM and participate in growth, migration, and metastasis of cancer cells, as well as binding integrins which enable GBM invasiveness and angiogenesis. The proportion of these isoforms present in GBM is striking enough to put one of them, EGF (1,2,5), on the list of GBM prognostic candidates (Safaee et al. 2015). Table 2 summarizes prominent alternatively spliced genes in GBM, including their respective GO terms related to their molecular functions.
Alternative splicing isoforms can have very distinct biological functions. A switch in splicing isoform balance or generation of a new isoform can contribute to cancer phenotypes. For instance, the C-CBL gene encodes an E3 ubiquitin-protein ligase involved in cell signaling and protein ubiquitination. Although this gene normally functions as a tumor suppressor, a C-CBL splicing variant generated by exon skipping in glioma cells contributes to malignant behavior (Seong et al. 2015). Similarly, the expression of RSU1, a gene that normally inhibits oncogenic Ras signaling in GBM cells, can undergo exon skipping resulting in a loss of tumor suppressor function and subsequent tumor growth (Chunduru et al. 2002). Another example is KAP, a cyclin-dependent kinase-associated phosphatase that dephosphorylates CDK2, inhibiting cell cycle progression. In astrocytomas, KAP aberrant splicing leads to the production of a dominant negative variant that decreased KAP protein, promotes cell cycle progression, and increases cell migration (Yu et al. 2007). The transcription activator Krüppel-like factor 6 (KLF6) gene encodes multiple protein isoforms derived from alternative mRNA splicing. As many as 16 alternatively spliced variants with divergent or even opposing functions can be produced. The full-length KLF6 (KLF6-FL) is a tumor suppressor gene, while the KLF6 splice variant 1 (KLF6-SV1) is an oncogenic isoform prevalent in GBM cells (Tchirkov et al. 2010). KLF6-SV1 reduction decreased cell proliferation by about 50% (Camacho-Vanegas et al. 2007). GLI1 encodes a transcription factor that promotes stem cell proliferation, but its activity is inhibited by TP53. GLI1’s truncated isoform, TGLI1, has gain-of-function in relation to the parental form, is highly expressed in GBM, promotes tumor cell migration, invasion, and malignancy, and fuels tumor growth by increasing angiogenesis (Lo et al. 2009). Carbonic anhydrase XII (CA-XII) is a transmembrane enzyme that is associated with hypoxic tumor growth by creating an acidic environment preferred by some tumor cells. The CA-XII isoform present in astrocytomas is primarily a shorter mRNA variant, where the absence of 11 amino acids may cause changes in structure and protein function augmenting tumor cell growth (Haapasalo et al. 2008). The growth hormone-releasing hormone (GHRH) is a member of the glucagon family of proteins that is produced in the hypothalamus and later cleaved to generate somatoliberin, which stimulates growth hormone release from the pituitary gland. GHRH receptor has two splice isoforms, the functional SV1 and the non-functional SV2. GBM patients, whose tumors lack GHRH expression, present poor prognosis, while GHRH positive and SV1 negative patients showed a better prognosis (Mezey et al. 2014).
In addition to oncogenic isoforms, alternative splicing can create tumor suppressor isoforms as well. USP5 maintains chromatin structure and degradation of abnormal proteins, whereas USP5 isoform 1 inhibits cellular growth and migration (Izaguirre et al. 2012). The tumor suppressor INK4b is a cyclin-dependent kinase inhibitor which forms a complex with CDK4 or CDK6 to prevent the activation of the CDK kinases by cyclin D. Although INK4b is frequently deleted in GBM (Kim and Sharpless 2006; Solomon et al. 2008), Simon and collaborators (2001) found wild-type INK4b in 34% of GBM cell lines analyzed. The INK4b gene displays two splice variants, p15 and p10; however, only the full product (p15) displays tumor suppression properties (Simon et al. 2001). RECK is a cysteine-rich, extracellular protein with protease inhibitor-like domains that was defined as tumor suppressor and metastases inhibitor in many contexts. RECK suppresses tumor invasion by negatively regulating members of the matrix metalloproteinase family: MMP-9, MMP-2, and MT1-MMP. Higher canonical RECK expression in combination with higher canonical to alternative transcript expression ratio positively correlates with higher overall survival rate after chemotherapeutic treatment of GBM patients. Moreover, glioblastoma cells transfected with RECK-B alternative splice variant showed higher anchorage-independent clonal growth (Trombetta-Lima et al. 2015). Some splicing variants of CYP27B1, a gene that participates on D3 vitamin metabolism, retain intron 1 leading to transcripts with premature stop codon. Those splicing variants result in truncated proteins without enzymatic activity. These variants contribute to decreased CYP27B1 activity, thereby decreasing GBM tumorigenicity (Diesel et al. 2005).
Similar to alternative splicing, alternative poly(A) (APA) contributes to transcript variation by generating mRNAs with shorter or longer 3′ untranslated regions (UTRs). These APA variations can have dramatic impact on gene expression levels, by creating or eliminating target sites for miRNAs and RNA binding proteins, which affect mRNAs translational efficiency and stability. RNA-seq analyses have established that APA is more prevalent than anticipated. Moreover, comparisons between normal and tumor tissue revealed great differences in respect to poly(A) site usage (Mayr and Bartel 2009). A large-scale study identified 4530 APA isoforms for 2733 genes in GBM, of which 182 APA isoforms from 148 genes were differentially expressed between GBM and normal brain tissue (Shao et al. 2013). A bona fide example of how APA can influence tumor progression is the gene O6-methylguanine-DNA methyltransferase (MGMT) which encodes a protein that can reverse the damage of alkylating agents, including the GBM standard-of-care therapeutic temozolomide. Elongation of the 3′ UTR of MGMT mRNA leads to miRNA-mediated silencing, impacting therapy outcome (Kreth et al. 2013). Factors implicated in poly(A) are proposed to act as suppressor agents. This is the case of CFIm25 (NUDT21) gene; its depletion in U251 GBM cells produced 3′ UTR shortening, activation of oncogenic pathways, and alterations in cell proliferation and growth. In HeLa cells, CFIm25 knockdown generated 1450 transcripts with shorter 3′ UTR (Masamha et al. 2014). We observed that the expression of CFIm25 is lower in GBM in comparison with LGG. Moreover, brain tumor patients displaying low expression of CFIm25 have worse prognosis.
Cancer therapy via splicing modulation
Since defects in mRNA splicing can lead to the development of several diseases, including cancer, modulation of splicing has the potential to be a promising therapeutic route. A critical consideration for any potential therapy is the therapeutic window—the difference between the dose that kills the targeted cancer cells and the dose that harms the normal tissues of the body. One of the most exciting outcomes of the cancer-specific biological functions described above is that cancer may be uniquely susceptible to splicing modulation therapy, where normal body tissues may not be. The first study to demonstrate this transformation-specific dependence upon RNA splicing machinery focused on the SF3b-complex protein PHF5A. Genome-wide functional screens using patient-derived models demonstrated that PHF5A is required for GBM cell survival but not for proliferating normal neural stem cells (NSCs). PHF5A loss or drug inhibition of its complex caused massive cell death only in the cancer cells (Hubert et al. 2013). RNA sequencing confirmed genome-wide splicing alterations and aberrant splicing events such as intron inclusion after PHF5A loss only in the cancer cells, not in normal NSCs. Since NSCs share many features with their transformed GBM counterparts, the authors iteratively transformed the resistant NSCs to become tumorigenic and pinpointed MYC overexpression as the trigger for splicing inhibitor sensitivity (Hubert et al. 2013). Studies in other cancer types have since confirmed that the spliceosome is a therapeutic vulnerability in MYC-driven cancers (Adler et al. 2014; Hsu et al. 2015). MYC’s ability to function as a global amplifier of transcription (Lin et al. 2012) could confer an overall increase in cellular RNA flux due to MYC activation specifically in cancer cells. Such increased demand might explain how MYC-driven cancer cells can acquire a unique sensitivity to perturbations of an otherwise universal cellular process. These studies have underscored the need for therapeutically tractable inhibitors of the RNA splicing machinery.
One of the first splicing modulators studied in cancer therapy was spliceostatin A, a methyl ketal derivative of FR901464, a potent anti-tumor compound isolated from a culture broth of Pseudomonas sp. No. 2663. Spliceostatin A functions as an inhibitor of splicing component SF3B1, which is involved in 3′ splice site recognition (Darman et al. 2015; Alsafadi et al. 2016). In leukemia, SF3B1 mutations were found in advanced stage of the disease and were related to poor prognosis (Cazzola et al. 2013). Mutations in SF3B1 gene were also identified in breast cancer and linked to ER-positive diseases, AKT1 mutations, and copy number variations. According to the same study, SF3B1 mutant cell lines were sensitive to spliceostatin A and the treatment has altered splicing signature (Maguire et al. 2015).
In the particular case of GBM, metformin has been shown to act synergistically with temozolomide (TMZ) to inhibit the proliferation of glioblastoma cells. Metformin downregulated SOX2 expression in TMZ-resistant glioma cells, reduced neurosphere formation capacity of glioblastoma cells, and inhibited GBM xenograft growth. Gene expression profiling data revealed that metformin’s impact on GBM cells mainly involves RNA binding and splicing pathways (Yang 2016); however, this may be an indirect effect through diverse cellular machinery. Another relevant compound is the anti-hypertensive agent amiloride, which can lead to apoptosis and radio-sensitivity through the modulation of APAF1 splicing (Tang et al. 2013). Among the analogous of FR901464, Pladienolide B reduced tumor size in glioblastoma U251 xenografts (Mizui et al. 2004). Importantly, Pladienolide B synthetic analog E7107 is in phase 1 clinical trial and has been shown good results by controlling tumor growth in several types of cancer (Eskens et al. 2013; Hong et al. 2014; Fan et al. 2011). Another interesting example is AR-A 014418, a selective GSK-3 inhibitor. Treatment of U373 and U87 glioblastoma lines with AR-A 014418 activated the apoptotic signaling pathway and reduced cell viability accompanied by downregulation of SRSF1, SRSF5, PTPB1, and hnRNPs (Yadav et al. 2014). Another compound which results in altered RNA splicing is hydrogen peroxide (H2O2). Treatment of BE2 neuroblastoma and MDA-MD-468 adenocarcinoma cells with H2O2 decreased expression of important splicing regulators such as PTBP1 and hnRNP A2/B1 and induced expression of the alternative spliced isoform of sGC, an anti-oxidant subunit activated in response to oxidative stress (Cote et al. 2012). As discussed above, PTBP1 and several hnRNPs are overexpressed in tumors, including glioblastoma, and the downregulation of these factors can be an important target to new anti-tumor therapies development. Finally, cross-talk between miRNAs and the RBPs from splicing machinery is an under-explored level of regulation with many interesting possibilities. Teplyuk and collaborators (2016) showed that miR-10b is a promising candidate for the development of new therapies; modulation of miR-10b function affected cell cycle and splicing regulation in glioma stem cells (GSC) and attenuated the growth of intracranial GBM xenografts.
Attempts to use highly targeted therapies, including RTK inhibitors, have thus far produced disappointing results in GBM. This could be due to a number of factors such as cellular heterogeneity, including controversial stem-like cell populations, barriers to drug delivery, including the blood–brain-barrier, cellular plasticity, and challenges of modeling GBM through traditional culture models. However, targeting components of oncogenic splicing machinery causes widespread alteration of thousands of transcripts and hundreds of proteins simultaneously, reducing the chance for therapeutic escape due to heterogeneity or adaptation. This approach has also been effective using both traditional cell lines as well as primary patient-derived GBM models in vitro and in vivo (Hubert et al. 2013). These findings suggest that the modulation of RNA splicing in refractory diseases such as GBM may hold therapeutic promise, where narrower targeting of single, easily replaceable proteins has been less effective.
Conclusions
Twenty years of genomic research have established RNA processing as a main contributor of transcript variation and gene expression regulation as well as its weight as a player in numerous diseases and cancer. In the last 5 years, cancer researchers increased their interest in RNA processing. Numerous mutations and aberrant expression of splicing regulators and RBPs have been observed in a variety of tumor types. These can function as tumor drivers and as major contributors of cancer-relevant phenotypes. Moreover, the growing list of inhibitors and molecules capable of interfering with splicing decisions has made RNA processing a target in the map of cancer therapy. Nowhere is this needed more than in the case of GBM. GBM therapies based on targeting a single gene or pathway have not shown dramatic improvement in clinical outcome despite specific genomic mutation data. Less targeted strategies using inhibitors of major regulatory hubs such as RBPs and splicing factors could have more potent effects and may be harder for tumor cells to evade.
Although the many examples of RNA-processing events and regulators implicated in GBM compiled here touch a variety of biological processes, it is clear that these many studies have just scratched the surface of this field. Genomic analyses of RNA processing in GBM need to be expanded. The TCGA and similar databases are rich depositories that can be explored more extensively. Each new study reports new splicing mRNA isoforms and poly(A) site alterations in GBM, but we have no mechanism for incorporating new findings into a unified database such as the TCGA, so we are very far from having a comprehensive catalogue of RNA isoform profiles. Similarly, the global impact of most RNA-processing regulators implicated in GBM is not known, since RNA-seq and CLIP analyses were not performed in most studies so far. Fortunately, in recent years, we have seen an expansion and improvement of both genomic methods and bioinformatics tools to analyze splicing and poly(A) site events, as well as an increase in RNA-seq read length and a decreasing of sequencing price, which allow better mapping of splicing and poly(A) site selection. Furthermore, the improvement of GBM models such as xenografts, organoids, and direct patient samples is opening new territory for exploration by allowing RNA processing to be studied amidst greater cellular diversity. All of these advancements combined and the development of new drugs are creating an unprecedented opportunity to illuminate the roles of RNA-processing regulation in GBM (and cancer) and to convert this knowledge into new therapeutic advancements, increasing the rate of patient survival.
References
Abdelmohsen K, Gorospe M (2010) Posttranscriptional regulation of cancer traits by HuR. Wiley Interdiscip Rev RNA 1(2):214–229. doi:10.1002/wrna.4
ABTA (2016) American Brain Tumor Association. www.abta.org. Accessed 15 Nov 2016
Adler AS, McCleland ML, Yee S, Yaylaoglu M, Hussain S, Cosino E, Quinones G, Modrusan Z, Seshagiri S, Torres E, Chopra VS, Haley B, Zhang Z, Blackwood EM, Singh M, Junttila M, Stephan JP, Liu J, Pau G, Fearon ER, Jiang Z, Firestein R (2014) An integrative analysis of colon cancer identifies an essential function for PRPF6 in tumor growth. Genes Dev 28(10):1068–1084. doi:10.1101/gad.237206.113
Alsafadi S, Houy A, Battistella A, Popova T, Wassef M, Henry E, Tirode F, Constantinou A, Piperno-Neumann S, Roman-Roman S, Dutertre M, Stern MH (2016) Cancer-associated SF3B1 mutations affect alternative splicing by promoting alternative branchpoint usage. Nat Commun. doi:10.1038/ncomms10615 (Article Number 10615)
Auboeuf D, Hönig A, Berget SM, O’Malley BW (2002) Coordinate regulation of transcription and splicing by steroid receptor coregulators. Science 298(5592):416–419
Babic I, Anderson ES, Tanaka K, Guo D, Masui K, Li B, Zhu S, Gu Y, Villa GR, Akhavan D, Nathanson D, Gini B, Mareninov S, Li R, Camacho CE, Kurdistani SK, Eskin A, Nelson SF, Yong WH, Cavenee WK, Cloughesy TF, Christofk HR, Black DL, Mischel PS (2013) EGFR mutation-induced alternative splicing of Max contributes to growth of glycolytic tumors in brain cancer. Cell Metab 17(6):1000–1008. doi:10.1016/j.cmet.2013.04.013
Bao ZS, Zhang CB, Wang HJ, Yan W, Liu YW, Li MY, Zhang W (2013) Whole-genome mRNA expression profiling identifies functional and prognostic signatures in patients with mesenchymal glioblastoma multiforme. CNS Neurosci Ther 19(9):714–720. doi:10.1111/cns.12118
Bi B, Li F, Guo J, Li C, Jing R, Lv X, Chen X, Wang F, Azadzoi KM, Wang L, Liu Y, Yang J (2017) Label -free quantitative proteomics unravels the importance of RNA processing in glioma malignancy. Neuroscience 351:84–95
Bowman RL, Wang Q, Carro A, Verhaak RG, Squatrito M (2017) GlioVis data portal for visualization and analysis of brain tumor expression datasets. Neuro Oncol 19(1):139–141. doi:10.1093/neuonc/now247 (Epub 9 Nov 2016)
Brennan CW, Verhaak RG, McKenna A, Campos B, Noushmehr H, Salama SR, Zheng S, Chakravarty D, Sanborn JZ, Berman SH, Beroukhim R, Bernard B, Wu CJ, Genovese G, Shmulevich I, Barnholtz-Sloan J, Zou L, Vegesna R, Shukla SA, Ciriello G, Yung WK, Zhang W, Sougnez C, Mikkelsen T, Aldape K, Bigner DD, Van Meir EG, Prados M, Sloan A, Black KL, Eschbacher J, Finocchiaro G, Friedman W, Andrews DW, Guha A, Iacocca M, O’Neill BP, Foltz G, Myers J, Weisenberger DJ, Penny R, Kucherlapati R, Perou CM, Hayes DN, Gibbs R, Marra M, Mills GB, Lander E, Spellman P, Wilson R, Sander C, Weinstein J, Meyerson M, Gabriel S, Laird PW, Haussler D, Getz G, Chin L, TCGA Research Network (2013) The somatic genomic landscape of glioblastoma. Cell 155(2):462–477. doi:10.1016/j.cell.2013.09.034
Brooks AN, Choi PS, de Waal L, Sharifnia T, Imielinski M, Saksena G, Pedamallu CS, Sivachenko A, Rosenberg M, Chmielecki J, Lawrence MS, DeLuca DS, Getz G, Meyerson M (2014) A pan-cancer analysis of transcriptome changes associated with somatic mutations in U2AF1 reveals commonly altered splicing events. PLoS One 9(1):e87361. doi:10.1371/journal.pone.0087361
Camacho-Vanegas O, Narla G, Teixeira MS, DiFeo A, Misra A, Singh G, Chan AM, Friedman SL, Feuerstein BG, Martignetti JA (2007) Functional inactivation of the KLF6 tumor suppressor gene by loss of heterozygosity and increased alternative splicing in glioblastoma. Int J Cancer 121(6):1390–1395
Cazzola M, Rossi M, Malcovati L (2013) Biologic and clinical significance of somatic mutations of SF3B1 in myeloid and lymphoid neoplasms. Blood 121(2):260–269. doi:10.1182/blood-2012-09-399725
Cheung HC, Baggerly KA, Tsavachidis S, Bachinski LL, Neubauer VL, Nixon TJ, Aldape KD, Cote GJ, Krahe R (2008) Global analysis of aberrant pre-mRNA splicing in glioblastoma using exon expression arrays. BMC Genom 9:216. doi:10.1186/1471-2164-9-216
Chunduru S, Kawami H, Gullick R, Monacci WJ, Dougherty G, Cutler ML (2002) Identification of an alternatively spliced RNA for the Ras suppressor RSU-1 in human gliomas. J Neurooncol 60(3):201–211
Correa BR, de Araujo PR, Qiao M, Burns SC, Chen C, Schlegel R, Agarwal S, Galante PA, Penalva LO (2016) Functional genomics analyses of RNA-binding proteins reveal the splicing regulator SNRPB as an oncogenic candidate in glioblastoma. Genome Biol 17(1):125. doi:10.1186/s13059-016-0990-4
Cote GJ, Zhu W, Thomas A, Martin E, Murad F, Sharina IG (2012) Hydrogen peroxide alters splicing of soluble guanylyl cyclase and selectively modulates expression of splicing regulators in human cancer cells. Plos One 7(7):1–9
Danan-Gotthold M, Golan-Gerstl R, Eisenberg E, Meir K, Karni R, Levanon EY (2015) Identification of recurrent regulated alternative splicing events across human solid tumors. Nucleic Acids Res 43(10):5130–5144. doi:10.1093/nar/gkv210
Dargahi D, Swayze RD, Yee L, Bergqvist PJ, Hedberg BJ, Heravi-Moussavi A, Dullaghan EM, Dercho R, An J, Babcook JS, Jones SJ (2014) A pan-cancer analysis of alternative splicing events reveals novel tumor-associated splice variants of matriptase. Cancer Inform 13:167–177. doi:10.4137/CIN.S19435
Darman RC, Seiler M, Agrawal AA, Lim KH, Peng S, Aird D, Bailey SL, Bhavsar EB, Chan B, Colla S, Corson L, Feala J, Fekkes P, Ichikawa K, Keaney GF, Lee L, Kumar P, Kunii K, MacKenzie C, Matijevic M, Mizui Y, Myint K, Park ES, Puyang X, Selvaraj A, Thomas MP, Tsai J, Wang JY, Warmuth M, Yang H, Zhu P (2015) Cancer-associated SF3B1 hotspot mutations induce cryptic 3′ splice site selection through use of a different branch point. Cell Reports 13(5):1033–1045
Diesel B, Radermacher J, Bureik M, Bernhardt R, Seifert M, Reichrath J, Fischer U, Meese E (2005) Vitamin D(3) metabolism in human glioblastoma multiforme: functionality of CYP27B1 splice variants, metabolism of calcidiol, and effect of calcitriol. Clin Cancer Res 11(15):5370–5380
DiFeo A, Martignetti JA, Narla G (2009) The role of KLF6 and its splice variants in cancer therapy. Drug Resist Updat 12(1–2):1–7. doi:10.1016/j.drup.2008.11.001
Dorman SN, Viner C, Rogan PK (2014) Splicing mutation analysis reveals previously unrecognized pathways in lymph node-invasive breast cancer. Sci Rep 4:7063. doi:10.1038/srep07063
Eskens FALM, Ramos FJ, Burger H, O’Brien JP, Piera A, Jonge MJA, Mizui Y, Wiemer EAC, Carreras MJ, Baselga J, Tabernero J (2013) Phase I pharmacokinetic and pharmacodynamic study of the first-in-class spliceosome inhibitor E7107 in patients with advanced solid tumors. Clin Cancer Res 19(22):6296–6304. doi:10.1158/1078-0432.CCR-13-0485
Fan L, Lagisetti C, Edwards CC, Webb TR, Potter PM (2011) Sudemycins, novel small molecule analogues of FR901464, induce alternative gene splicing. ACS Chem Biol 6(6):582–589. doi:10.1021/cb100356k
Ferrarese R, Harsh GR 4th, Yadav AK, Bug E, Maticzka D, Reichardt W, Dombrowski SM, Miller TE, Masilamani AP, Dai F, Kim H, Hadler M, Scholtens DM, Yu IL, Beck J, Srinivasasainagendra V, Costa F, Baxan N, Pfeifer D, von Elverfeldt D, Backofen R, Weyerbrock A, Duarte CW, He X, Prinz M, Chandler JP, Vogel H, Chakravarti A, Rich JN, Carro MS, Bredel M (2014) Lineage-specific splicing of a brain-enriched alternative exon promotes glioblastoma progression. J Clin Invest 124(7):2861–2876. doi:10.1172/JCI68836
Filippova N, Yang X, Wang Y, Gillespie GY, Langford C, King PH, Wheeler C, Nabors LB (2011) The RNA-binding protein HuR promotes glioma growth and treatment resistance. Mol Cancer Res 9(5):648–659. doi:10.1158/1541-7786.MCR-10-0325
Fontana L, Rovina D, Novielli C, Maffioli E, Tedeschi G, Magnani I, Larizza L (2015) Suggestive evidence on the involvement of polypyrimidine-tract binding protein in regulating alternative splicing of MAP/microtubule affinity-regulating kinase 4 in glioma. Cancer Lett 359(1):87–96. doi:10.1016/j.canlet.2014.12.049
Galante PAF, Sandhu D, Abreu RS, Gradassi M, Slager N, Vogel C, Souza SJ, Penalva LOF (2009) A comprehensive in silico expression analysis of RNA binding proteins in normal and tumor tissue. RNA Biol 6(4):426–433
García-Sacristán A, Fernández-Nestosa MJ, Hernández P, Schvartzman JB, Krimer DB (2005) Protein kinase clk/STY is differentially regulated during erythroleukemia cell differentiation: a bias toward the skipped splice variant characterizes postcommitment stages. Cell Res 15(7):495–503
Golan-Gerstl R, Cohen M, Shilo A, Suh SS, Bakàcs A, Coppola L, Karni R (2011) Splicing factor hnRNP A2/B1 regulates tumor suppressor gene splicing and is an oncogenic driver in glioblastoma. Cancer Res 71(13):4464–4472. doi:10.1158/0008-5472.CAN-10-4410
Grammatikakis I, Abdelmohsen K, Gorospe M (2017) Posttranslational control of HuR function. Wiley Interdiscip Rev RNA. doi:10.1002/wrna.1372
Haapasalo J, Hilvo M, Nordfors K, Haapasalo H, Parkkila S, Hyrskyluoto A, Rantala I, Waheed A, Sly WS, Pastorekova S, Pastorek J, Parkkila AK (2008) Neuro Oncol 10(2):131–138. doi:10.1215/15228517-2007-065
He X, Pool M, Darcy KM, Lim SB, Auersperg N, Coon JS et al (2007) Knockdown of polypyrimidine tract-binding protein suppresses ovarian tumor cell growth and invasiveness in vitro. Oncogene 26:4961–4968
He X, Arslan AD, Ho T-T, Yuan C, Stampfer MR, Beck WT (2014) Involvement of polypyrimidine tract-binding protein (PTBP1) in maintaining breast cancer cell growth and malignant properties. Oncogenesis 3(1):e84. doi:10.1038/oncsis.2013.47
Hishiki T, Kawamoto S, Morishita S, Okubo K (2000) BodyMap: a human and mouse gene expression database. Nucleic Acids Res 28(1):136–138
Hong DS, Kurzrock R, Naing A, Wheler JJ, Falchook GS, Schiffman JS, Faulkner N, Pilat MJ, O’Brien J, LoRusso P (2014) A phase I, open-label, single-arm, dose-escalation study of E7107, a precursor messenger ribonucleic acid (pre-mRNA) splicesome inhibitor administered intravenously on days 1 and 8 every 21 days to patients with solid tumors. Investig New Drug 32(3):436–444
Hsu TY, Simon LM, Neill NJ, Marcotte R, Sayad A, Bland CS, Echeverria GV, Sun T, Kurley SJ, Tyagi S, Karlin KL, Dominguez-Vidaña R, Hartman JD, Renwick A, Scorsone K, Bernardi RJ, Skinner SO, Jain A, Orellana M, Lagisetti C, Golding I, Jung SY, Neilson JR, Zhang XH, Cooper TA, Webb TR, Neel BG, Shaw CA, Westbrook TF (2015) The spliceosome is a therapeutic vulnerability in MYC-driven cancer. Nature 525(7569):384–388. doi:10.1038/nature14985
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4(1):44–57
Hubert CG, Bradley RK, Ding Y, Toledo CM, Herman J, Skutt-Kakaria K, Girard EJ, Davison J, Berndt J, Corrin P, Hardcastle J, Basom R, Delrow JJ, Webb T, Pollard SM, Lee J, Olson JM, Paddison PJ (2013) Genome-wide RNAi screens in human brain tumor isolates reveal a novel viability requirement for PHF5A. Genes Dev 27(9):1032–1045. doi:10.1101/gad.212548.112
Izaguirre DI, Zhu W, Hai T, Cheung HC, Krahe R, Cote GJ (2012) PTBP1-dependent regulation of USP5 alternative RNA splicing plays a role in glioblastoma tumorigenesis. Mol Carcinog 51(11):895–906. doi:10.1002/mc.20859
Jin W, Bi W, Huang ES, Cote GJ (1999) Glioblastoma cell-specific expression of fibroblast growth factor receptor-1beta requires an intronic repressor of RNA splicing. Cancer Res 59(2):316–319
Jin W, McCutcheon IE, Fuller GN, Huang ES, Cote GJ (2000) Fibroblast growth factor receptor-1 alpha-exon exclusion and polypyrimidine tract-binding protein in glioblastoma multiforme tumors. Cancer Res 60(5):1221–1224
Kafasla P, Mickleburgh I, Llorian M, Coelho M, Gooding C, Cherny D, Joshi A, Kotik-Kogan O, Curry S, Eperon IC, Jackson RJ, Smith CW (2012) Defining the roles and interactions of PTB. Biochem Soc Trans 40(4):815–820. doi:10.1042/BST20120044
Kai M (2016). Roles of RNA-binding proteins in DNA damage response. Int J Mol Sci 17(3):310. doi:10.3390/ijms17030310 (Review). Erratum in: Int J Mol Sci. doi:10.3390/ijms17040604
Kang YK, Schiff R, Ko L, Wang T, Tsai SY, Tsai MJ, O’Malley BW (2008) Dual roles for coactivator activator and its counterbalancing isoform coactivator modulator in human kidney cell tumorigenesis. Cancer Res 68(19):7887–7896. doi:10.1158/0008-5472.CAN-08-1734
Kechavarzi B, Janga SC (2014) Dissecting the expression landscape of RNA-binding proteins in human cancers. Genome Biol 15(1):R14. doi:10.1186/gb-2014-15-1-r14
Kim WY, Sharpless NE (2006) The regulation of INK4/ARF in cancer and aging. Cell 127(2):265–275
Kim YW, Koul D, Kim SH, Lucio-Eterovic AK, Freire PR, Yao J, Wang J, Almeida JS, Aldape K, Yung WK (2013) Identification of prognostic gene signatures of glioblastoma: a study based on TCGA data analysis. Neuro Oncol 15(7):829–839. doi:10.1093/neuonc/not024
Kreth S, Limbeck E, Hinske LC, Schütz SV, Thon N, Hoefig K, Egensperger R, Kreth FW (2013) In human glioblastomas transcript elongation by alternative polyadenylation and miRNA targeting is a potent mechanism of MGMT silencing. Acta Neuropathol 125(5):671–681. doi:10.1007/s00401-013-1081-1
Lebedeva S, Jens M, Theil K, Schwanhäusser B, Selbach M, Landthaler M, Rajewsky N (2011) Transcriptome-wide analysis of regulatory interactions of the RNA-binding protein HuR. Mol Cell 43(3):340–352. doi:10.1016/j.molcel.2011.06.008
Lefave CV, Squatrito M, Vorlova S, Rocco GL, Brennan CW, Holland EC, Pan YX, Cartegni L (2011) Splicing factor hnRNPH drives an oncogenic splicing switch in gliomas. EMBO J 30(19):4084–4097. doi:10.1038/emboj.2011.259
Lin CY, Lovén J, Rahl PB, Paranal RM, Burge CB, Bradner JE, Lee TI, Young RA (2012) Transcriptional amplification in tumor cells with elevated c-Myc. Cell 151(1):56–67. doi:10.1016/j.cell.2012.08.026
Lo HW, Zhu H, Cao X, Aldrich A, Ali-Osman F (2009) A novel splice variant of GLI1 that promotes glioblastoma cell migration and invasion. Cancer Res 69(17):6790–6798. doi:10.1158/0008-5472.CAN-09-0886
Maguire SL, Leonidou A, Wai P, Marchiò C, Ng CKY, Sapino A, Salomon AV, Reis-Filho JS, Weigelt B, Natrajan RC (2015) SF3B1 mutations constitute a novel therapeutic target in breast cancer. J Pathol 235(4):571–580. doi:10.1002/path.4483
Masamha CP, Xia Z, Yang J, Albrecht TR, Li M, Shyu A, Li W, Wagner EJ (2014) CFIm25 links alternative polyadenylation to glioblastoma tumor suppression. Nature 510(7505):412–416. doi:10.1038/nature13261
Mayr C, Bartel DP (2009) Widespread shortening of 3′ UTRs by alternative cleavage and polyadenylation activates oncogenes in cancer cells. Cell 138(4):673. doi:10.1016/j.cell.2009.06.016
Mccutcheon IE, Hentschel SJ, Fuller GN, Jin W, Cote GJ (2004) Expression of the splicing regulator polypyrimidine tract-binding protein in normal and neoplastic brain. Neuro Oncol 6(1):9–14
Mezey G, Treszl A, Schally AV, Block NL, Vízkeleti L, Juhász A, Klekner A, Nagy J, Balázs M, Halmos G, Bognár L (2014) Prognosis in human glioblastoma based on expression of ligand growth hormone-releasing hormone, pituitary-type growth hormone-releasing hormone receptor, its splicing variant receptors, EGF receptor and PTEN genes. J Cancer Res Clin Oncol 140(10):1641–1649. doi:10.1007/s00432-014-1716-1
Mizui Y, Sakai T, Iwata M, Uenaka T, Okamoto K, Shimizu H, Yamori T, Yoshimatsu K, Asada M (2004) Pladienolides, new substances from culture of Streptomyces platensis Mer-11107III. In vitro and in vivo antitumor activities. J Antibiot 57(3):188–196
Motaln H, Koren A, Gruden K, Ramšak Z, Schichor C, Lah T (2015) Heterogeneous glioblastoma cell cross-talk promotes phenotype alterations and enhanced drug resistance. Oncotarget 6(38):40998–41017. doi:10.18632/oncotarget.5701
Mukherjee N, Corcoran DL, Nusbaum JD, Reid DW, Georgiev S, Hafner M, Ascano M Jr, Tuschl T, Ohler U, Keene JD (2011) Integrative regulatory mapping indicates that the RNA-binding protein HuR couples pre-mRNA processing and mRNA stability. Mol Cell 43(3):327–339. doi:10.1016/j.molcel.2011.06.007
Padfield E, Ellis HP, Kurian KM (2015) Current therapeutic advances targeting EGFR and EGFRvIII in glioblastoma. Front Oncol. doi:10.3389/fonc.2015.00005
Patel VN, Gokulrangan G, Chowdhury SA, Chen Y, Sloan AE, Koyutürk M, Barnholtz-Sloan J, Chance MR (2013) Network signatures of survival in glioblastoma multiforme. PLoS Comput Biol 9(9):e1003237. doi:10.1371/journal.pcbi.1003237
Sadeque A, Serão NV, Southey BR, Delfino KR, Rodriguez-Zas SL (2012) Identification and characterization of alternative exon usage linked glioblastoma multiforme survival. BMC Med Genom 5:59. doi:10.1186/1755-8794-5-59
Safaee M, Fakurnejad S, Bloch O, Clark AJ, Ivan ME, Sun MZ, Oh T, Phillips JJ, Parsa AT (2015) Proportional upregulation of CD97 isoforms in glioblastoma and glioblastoma-derived brain tumor initiating cells. PLoS One 10(2):e0111532. doi:10.1371/journal.pone.0111532
Seong MW, Ka SH, Park JH, Park JH, Yoo HM, Yang SW, Park JM, Park D, Lee ST, Seol JH, Chung CH (2015) Deleterious c-Cbl exon skipping contributes to human glioma. Neoplasia 17(6):518–524. doi:10.1016/j.neo.2015.06.003
Shao J, Zhang J, Zhang Z, Jiang H, Lou X, Huang B, Foltz G, Lan Q, Huang Q, Lin B (2013) Alternative polyadenylation in glioblastoma multiforme and changes in predicted RNA binding protein profiles. OMICS 17(3):136–149. doi:10.1089/omi.2012.0098
Shiratsuchi G, Takaoka K, Ashikawa T, Hamada H, Kitagawa D (2015) RBM14 prevents assembly of centriolar protein complexes and maintains mitotic spindle integrity. EMBO J 34(1):97–114. doi:10.15252/embj.201488979
Simon M, Köster G, Ludwig M, Mahlberg R, Rho S, Watzka M, Schramm J (2001) Alternative splicing of the p15 cdk inhibitor in glioblastoma multiforme. Acta Neuropathol 102(2):167–174
Simon M, Hosen I, Gousias K, Rachakonda S, Heidenreich B, Gessi M, Schramm J, Hemminki K, Waha A, Kumar R (2015) TERT promoter mutations: a novel independent prognostic factor in primary glioblastomas. Neuro Oncol 17(1):45–52. doi:10.1093/neuonc/nou158
Simpson MT, Venkatesh I, Callif BL, Thiel LK, Coley DM, Winsor KN, Wang Z, Kramer AA, Lerch JK, Blackmore MG (2015) The tumor suppressor HHEX inhibits axon growth when prematurely expressed in developing central nervous system neurons. Mol Cell Neurosci 68:272–283. doi:10.1016/j.mcn.2015.08.008
Solomon DA, Kim JS, Jean W, Waldman T (2008) Conspirators in a capital crime: co-deletion of p18INK4c and p16INK4a/p14ARF/p15INK4b in glioblastoma multiforme. Cancer Res 68(21):8657–8660. doi:10.1158/0008-5472.CAN-08-2084
Sottoriva A, Spiteri I, Piccirillo SGM, Touloumis A, Collins P, Marioni JC, Curtisc C, Watts C, Tavaré S (2012) Intratumor heterogeneity in human glioblastoma reflects cancer evolutionary dynamics. PNAS 110(10):4009–4014. doi:10.1073/pnas.1219747110
Srikantan S, Gorospe M (2011) UneCLIPsing HuR nuclear function. Mol Cell 43(3):319–321. doi:10.1016/j.molcel.2011.07.016
Su Y, Yang Z, Xiong S, Zhang L, Blanchard KL, Peiper SC, Dynan WS, Tuan D, Ko L (2007) Gene amplification and associated loss of 5′ regulatory sequences of CoAA in human cancers. Oncogene 26(6):822–835
Tang JY, Chang HW, Chang JG (2013) Modulating roles of amiloride in irradiation-induced antiproliferative effects in glioblastoma multiforme cells involving Akt phosphorylation and the alternative splicing of apoptotic genes. DNA Cell Biol 32(9):504–510. doi:10.1089/dna.2013.1998
Tchirkov A, Sapin V, Marceau G, Chautard E, Narla G, Veronese L, Friedman S, Khalil T, Vago P, Kemeny JL, Verrelle P (2010) Increased expression of the oncogenic KLF6-SV1 transcript in human glioblastoma. Clin Chem Lab Med 48(8):1167–1170. doi:10.1515/CCLM.2010.219
Teplyuk NM, Uhlmann EJ, Gabriely G, Volfovsky N, Wang Y, Teng J, Karmali P, Marcusson E, Peter M, Mohan A, Kraytsberg Y, Cialic R, Chiocca EA, Godlewski J, Tannous B, Krichevsky AM (2016) Therapeutic potential of targeting microRNA-10b in established intracranial glioblastoma: first steps toward the clinic. EMBO Mol Med 8(3):268–287. doi:10.15252/emmm.201505495
Trombetta-Lima M, Winnischofer SM, Demasi MA, Astorino Filho R, Carreira AC, Wei B, de Assis-Ribas T, Konig MS, Bowman-Colin C, Oba-Shinjo SM, Marie SK, Stetler-Stevenson W, Sogayar MC (2015) Isolation and characterization of novel RECK tumor suppressor gene splice variants. Oncotarget 6(32):33120–33133. doi:10.18632/oncotarget.5305
Tsai YS, Dominguez D, Gomez SM, Wang Z (2015) Transcriptome-wide identification and study of cancer-specific splicing events across multiple tumors. Oncotarget 6(9):6825–6839
Uren PJ, Burns SC, Ruan J, Singh KK, Smith AD, Penalva LO (2011) Genomic analyses of the RNA-binding protein Hu antigen R (HuR) identify a complex network of target genes and novel characteristics of its binding sites. J Biol Chem 286(43):37063–37066. doi:10.1074/jbc.C111.266882
Uren PJ, Bahrami-Samani E, de Araujo PR, Vogel C, Qiao M, Burns SC, Smith AD, Penalva LO (2016) High-throughput analyses of hnRNP H1 dissects its multi-functional aspect. RNA Biol 13(4):400–411. doi:10.1080/15476286.2015.1138030
Valles I, Pajares MJ, Segura V, Guruceaga E, Gomez-Roman J, Blanco D, Tamura A, Montuenga LM, Pio R (2012) Identification of novel deregulated RNA metabolism-related genes in non-small cell lung cancer. PLoS One 7(8):e42086. doi:10.1371/journal.pone.0042086
Varghese RT, Liang Y, Guan T, Franck CT, Kelly DF, Sheng Z (2016) Survival kinase genes present prognostic significance in glioblastoma. Oncotarget 7(15):20140–20151. doi:10.18632/oncotarget.7917
Verhaak RG, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, Miller CR, Ding L, Golub T, Mesirov JP, Alexe G et al (2010) Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17(1):98–110
Vo DT, Abdelmohsen K, Martindale JL, Qiao M, Tominaga K, Burton TL, Gelfond JA, Brenner AJ, Patel V, Trageser D, Scheffler B, Gorospe M, Penalva LO (2012) The oncogenic RNA-binding protein Musashi1 is regulated by HuR via mRNA translation and stability in glioblastoma cells. Mol Cancer Res 10(1):143–155. doi:10.1158/1541-7786.MCR-11-0208
Weinhold N, Jacobsen A, Schultz N, Sander C, Lee W (2014) Genome-wide analysis of noncoding regulatory mutations in cancer. Nat Genet 46:1160–1165. doi:10.1038/ng.3101
World Health Organization (WHO) (2016) www.who.int. Accessed 15 Nov 2016
Xie Q, Mittal Berens ME (2014) Targeting adaptive glioblastoma: an overview of proliferation and invasion. Neuro Oncol 16(12):1575–1584. doi:10.1093/neuonc/nou147
Yadav AK, Vashishta V, Joshi N, Taneja P (2014) AR-A 014418 used against GSK3beta downregulates expression of hnRNPA1 and SF2/ASF splicing factors. J Oncol 2014:695325. doi:10.1155/2014/695325
Yang SH, Li S, Lu G, Xue H, Kim DH, Zhu JJ, Liu Y (2016) Metformin treatment reduces temozolomide resistance of glioblastoma cells. Oncotarget. doi:10.18632/oncotarget.12859
Yeo GW, Bjar R, Benegiamo G, Aigner S, Bengtson MH, Berkowitz ND, Bos TJ, Brown SA, Buac K, Calarco JA, Fan AC, Gosai SJ, Gracida X, Gregory BD, Hattori A, Huelga SC, Hundley HA, Ito T, Leung AKL, Licatalosi DD, Lovci MT, Massirer KB, Moore MJ, Norris AD, Nostrand ELV, Nussbacher JK, Panda S, Serebrov V, Silverman IM, Washburn MC (2016) RNA processing—disease and genome-wide probing. Springer, San Diego
Yong WH, Shabihkhani M, Telesca D, Yang S, Tso JL, Menjivar JC, Wei B, Lucey GM, Mareninov S, Chen Z, Liau LM, Lai A, Nelson SF, Cloughesy TF, Tso CL (2015) Ribosomal proteins RPS11 and RPS20, two stress-response markers of glioblastoma stem cells, are novel predictors of poor prognosis in glioblastoma patients. PLoS One 10(10):e0141334. doi:10.1371/journal.pone.0141334
Yoshida T, Kim JH, Carver K, Su Y, Weremowicz S, Mulvey L, Yamamoto S, Brennan C, Mei S, Long H, Yao J, Polyak K (2015) CLK2 is an oncogenic kinase and splicing regulator in breast cancer. Cancer Res 75(7):1516–1526. doi:10.1158/0008-5472.CAN-14-2443
Yu F, Fu WM (2015) Identification of differential splicing genes in gliomas using exon expression profiling. Mol Med Rep 11(2):843–850. doi:10.3892/mmr.2014.2775
Yu Y, Jiang X, Schoch BS, Carroll RS, Black PM, Johnson MD (2007) Aberrant splicing of cyclin-dependent kinase-associated protein phosphatase KAP increases proliferation and migration in glioblastoma. Cancer Res 67(1):130–138
Yuan M, Eberhart CG, Kai M (2014) RNA binding protein RBM14 promotes radio-resistance in glioblastoma by regulating DNA repair and cell differentiation. Oncotarget 5(9):2820–2826
Acknowledgements
Work on glioblastoma in LOP and PAFG labs is supported by the Conselho Nacional de desenvolvimento científico e tecnológico (CNPq—Process Number 400262/2014-2, Brazil) and NIH (5R21CA205475 and 1R21CA175875). CGH is supported by an NIH F32 (CA189647) fellowship. FMM is sponsored by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), Brazil.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Rights and permissions
About this article
Cite this article
Marcelino Meliso, F., Hubert, C.G., Favoretto Galante, P.A. et al. RNA processing as an alternative route to attack glioblastoma. Hum Genet 136, 1129–1141 (2017). https://doi.org/10.1007/s00439-017-1819-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-017-1819-2