Abstract
Yeast genes of phospholipid biosynthesis are negatively regulated by repressor protein Opi1 when precursor molecules inositol and choline (IC) are available. Opi1-triggered gene repression is mediated by recruitment of the Sin3 corepressor complex. In this study, we systematically investigated the regulatory contribution of subunits of Sin3 complexes and identified Pho23 as important for IC-dependent gene repression. Two non-overlapping regions within Pho23 mediate its direct interaction with Sin3. Previous work has shown that Sin3 recruits the histone deacetylase (HDAC) Rpd3 to execute gene repression. While deletion of SIN3 strongly alleviates gene repression by IC, an rpd3 null mutant shows almost normal regulation. We thus hypothesized that various HDACs may contribute to Sin3-mediated repression of IC-regulated genes. Indeed, a triple mutant lacking HDACs, Rpd3, Hda1 and Hos1, could phenocopy a sin3 single mutant. We show that these proteins are able to contact Sin3 in vitro and in vivo and mapped three distinct HDAC interaction domains, designated HID1, HID2 and HID3. HID3, which is identical to the previously described structural motif PAH4 (paired amphipathic helix), can bind all HDACs tested. Chromatin immunoprecipitation studies finally confirmed that Hda1 and Hos1 are recruited to promoters of phospholipid biosynthetic genes INO1 and CHO2.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Transcriptional corepressor complexes in eukaryotes are of central importance for negative regulation of numerous structural genes by creating a local structure of chromatin inhibitory to gene expression. In the yeast Saccharomyces cerevisiae, two pleiotropic corepressor complexes, Sin3 and Ssn6/Tup1, have been described (reviewed by Grzenda et al. 2009; Silverstein and Ekwall 2005; Malavé and Dent 2006), both of which execute gene repression by recruiting histone deacetylases (Kadosh and Struhl 1997; Watson et al. 2000; Davie et al. 2003). Since neither of these complexes displays DNA binding, promoter targeting is achieved through interactions with sequence-specific DNA-binding proteins.
We have previously shown that the specific repressor Opi1 of yeast phospholipid biosynthesis functions by recruiting Sin3 and Ssn6 (Wagner et al. 2001; Jäschke et al. 2011). Structural genes of phospholipid biosynthesis such as INO1, CHO1 and CHO2 are repressed when precursor molecules inositol and choline (IC) are available in excess. In contrast, limiting concentrations of IC allow activation of these genes which is triggered by the heterodimeric transcription factor Ino2/Ino4 (Schwank et al. 1995; reviewed by Chen et al. 2007). Both Ino2 and Ino4 are members of the basic helix-loop-helix (bHLH) family of DNA-binding proteins and specifically interact with the UAS element ICRE (inositol/choline-responsive element = UASINO; Schüller et al. 1992; Ambroziak and Henry 1994; Hoppen et al. 2005) found upstream of phospholipid biosynthetic genes. While Ino4 is responsible for nuclear import of the heterodimer and its DNA binding (Kumme et al. 2008), Ino2 mediates transcriptional activation (Schwank et al. 1995; Dietz et al. 2003) and regulatory input via its Opi1 interaction domain (Heyken et al. 2005). Thus, Opi1 as the specific repressor of ICRE-dependent genes is indirectly recruited to target promoters by activator Ino2 (Wagner et al. 2001).
Genetic analysis initially has shown that negative regulation of phospholipid biosynthetic genes by IC requires the pleiotropic repressor Sin3, which can bind to the N-terminus of Opi1 (containing OSID: Opi1-Sin3 interaction domain; Slekar and Henry 1995; Wagner et al. 2001). Sin3 contains four paired amphipathic helix motifs (PAH1–PAH4; Wang et al. 1990) which via protein–protein interactions support its function as a scaffold for various unrelated regulatory networks. Interestingly, OSID is responsible for interaction with PAH1 of Sin3 and for contacting tetratricopeptide repeat motifs (TPR) of corepressor Ssn6 (Jäschke et al. 2011). Being recruited to defined control regions via interaction with specific regulators, Sin3 executes repression by associated histone deacetylases (HDACs) such as Rpd3 in yeast (Kadosh and Struhl 1997) and HDAC1 and HDAC2 in mammalian systems (Laherty et al. 1997). Methods of proteomic analysis led to the identification of two multiprotein complexes (designated Sin3/Rpd3L and Sin3/Rpd3S; Loewith et al. 2001; Carrozza et al. 2005b), varying with respect to auxiliary proteins. While Rpd3L preferentially localizes to promoter regions (Rundlett et al. 1998), Rpd3S is associated with transcribed regions (Joshi and Struhl 2005), possibly preventing internal initiations. Importantly, the genome of S. cerevisiae encodes five Zn2+-containing HDACs which deacetylate histones by hydrolysis (Rpd3, Hda1, Hos1, Hos2 and Hos3; Rundlett et al. 1996), while Sir2 is a NAD-dependent enzyme. Based on sequence similarities, HDACs have been classified into four groups (class I, Rpd3, Hos1 and Hos2 in S. cerevisiae; class II, Hda1 and Hos3; class III, Sir2 and related sirtuins; class IV, no member in yeasts; reviewed by Hildmann et al. 2007; Yang and Seto 2008).
Previous work focusing on the negative regulatory element URS1 and its specific binding factor Ume6 proposed the simple regulatory hierarchy URS1-Ume6-Sin3-Rpd3 (Kadosh and Struhl 1997). However, a more complex situation may be effective with other genes repressed by Sin3. Our previous analysis of ICRE-dependent gene regulation showed that repression by IC is severely alleviated in a sin3 null mutant, while loss of Rpd3 causes almost no change (Wagner et al. 2001). We thus hypothesized that Sin3 should be able to recruit not only Rpd3, but also additional HDACs which together may be required to transform chromatin into a repressed state. In this study, we indeed could show that loss of HDACs Rpd3, Hda1 and Hos1 phenocopies deletion of SIN3. This genetic result is further confirmed by demonstration of physical interaction of Sin3 with Rpd3, Hda1 and Hos1 in vitro and in vivo. Finally, we identify two additional HDAC interaction domains within Sin3.
Materials and methods
Yeast strains, media and growth conditions
All strains of the yeast S. cerevisiae used in this study are isogenic to the regulatory wild-type SIRP3 (Schwank et al. 1995), containing a chromosomally integrated ICRE-CYC1-lacZ reporter gene. Strains differ with respect to the status of subunits of Sin3 complexes and HDACs. Strains SIRP3.Δopi1, SIRP3.Δsin3, SIRP3.Δume6 and SIRP3.Δrpd3 have been described (Wagner et al. 2001). Null mutations were introduced into SIRP3 by transformation with deletion mutant alleles described below. Transformants obtained were verified for the presence of the desired mutant allele and the absence of the wild-type allele. Yeast extracts used for protein–protein interaction assays were prepared from strain C13-ABY.S86, lacking four vacuolar proteinases (pra1 prb1 prc1 cps1; De Antoni and Gallwitz 2000). A compilation of strains and genotypes is available as supporting online Table 1.
Synthetic complete (SC) media used for selective growth of transformants and conditions of IC repression or derepression have been described (Schüller et al. 1992). Although doubling times of mutants lacking multiple HDACs were severely delayed with respect to the wild type (275 min for the rpd3 hda1 hos1 triple mutant compared with 100 min for the wild type), strains were uniformly harvested at a cell density of 2 × 107 cells/ml.
Plasmid constructions
The following plasmids were constructed by established procedures to disrupt genes encoding subunits of Sin3 complexes and HDACs: pRH2 (∆sap30::HIS3), pRH4 (∆sds3::HIS3), pRH6 (∆pho23::kanMX), pSW2 (∆dep1::LEU2), pSW21 (∆ume1::LEU2), pTN2 (∆rxt2::LEU2), pTN4 (∆eaf3::LEU2), pYJ11 (Δhos1::TRP1), pYJ16 (Δhos3::LEU2), pYJ18 (Δhda1::HIS3), pYJ20 (Δhos2::hph) and pSS20 (Δino2::LEU2). To construct these plasmids, flanking sequences upstream and downstream of the respective coding regions were amplified by PCR and inserted on both sides of the selection marker, allowing total deletion of reading frames.
To perform interaction assays, Escherichia coli expression plasmids (derived from pGEX-2TK; GE Healthcare) encoding various glutathione S-transferase (GST) fusions were constructed. Length variants of coding regions of PHO23, SDS3, SAP30, RPD3, HDA1 and HOS1 were amplified by PCR and fused behind GST. Similarly, HA-tagged length variants of Sin3 representing PAH and HID domains were expressed in yeast using plasmid p426-MET25HA (Mumberg et al. 1994). For bacterial expression of selected Sin3 variants, plasmid pASK-IBA5 (tetR-regulated; IBA, Göttingen, Germany) was used. Plasmid constructs encoding minimal length variants used for interaction studies were verified by DNA sequencing to confirm authenticity of gene fragments obtained by PCR. Plasmid names and fused sequences are mentioned in the legends of figures.
In vitro interaction assays (GST pull-down)
GST- and HA-tagged proteins used for interaction assays by affinity chromatography were synthesized by E. coli strain BL21 (Stratagene/Agilent). The tac promoter controlling GST fusion genes was induced with 1 mM IPTG. Similarly, tetR-dependent gene expression was activated by 0.2 mg/l anhydrotetracycline. Derepression of MET25-dependent gene fusions was achieved by cultivating yeast transformants in the absence of methionine.
GST fusion proteins synthesized in E. coli were released by sonication, immobilized on glutathione (GSH) Sepharose and subsequently incubated with yeast or bacterial total protein extracts containing HA fusions. To avoid unspecific interactions, protein extracts were pre-cleared by treatment with GSH Sepharose beads prior to incubation with GST fusions. Details on washing steps at intermediary stringency have been described (Wagner et al. 2001). After release of GST fusions with free GSH (10 mM), eluates were separated by SDS/PAGE and proteins transferred to a filter. Following incubation with anti-HA-peroxidase conjugate, HA fusion proteins were detected with POD chemiluminescent substrate (antibody conjugate and substrate from Roche Biochemicals).
Two-hybrid assays
To perform two-hybrid assays, strain PJ69-4A was used (MATa trp1 leu2 ura3 his3Δ gal4Δ gal80Δ GAL2-ADE2 LYS2::GAL1-HIS3 met2::GAL7-lacZ; James et al. 1996). DNA fragments encoding interaction domains of Sin3, Hda1, Hos1 and Rpd3 were inserted into plasmids pGBD-C1 (2 μm GAL4 DBD TRP1) and pGAD-C1 (2 μm GAL4 TAD LEU2), respectively. Double-transformed strains containing both types of fusion plasmids were first selected on medium lacking leucine and tryptophan (-L-T) and subsequently transferred to medium devoid of adenine (-L-T-A).
Chromatin immunoprecipitation
Essentially, chromatin immunoprecipitation (ChIP) analysis followed the procedure described by Cobb and van Attikum (2010). Chromosomal loci HDA1 and HOS1 were modified such that they expressed His-tagged Hda1 and Hos1 without alteration of gene copy number or control region. Tagging was performed by transformation of strain C13-ABY.S86 with a gene-specific modification fragment and selection for resistance against geneticin. The modification fragment was amplified by PCR, using gene-specific primers and plasmid pU6H3HA as a template (contains a His6-HA3-kanMX cassette; De Antoni and Gallwitz 2000). The resulting strains FKY40 (HDA1-His6-HA3) and FKY42 (HOS1-His6-HA3) and their isogenic ino2 derivatives FKY44 and FKY46 grew until mid-log phase and were treated with formaldehyde for 15 min. The cross-linking reaction was subsequently quenched for 5 min by addition of glycine to a final concentration of 125 mM. After lysis, cells were sonicated five times for 30 s to shear chromatin, using a Bandelin Sonoplus UW 70 microtip (35 % power). After sonication, lysates were centrifuged for 10 min at 16,000×g to remove insoluble material and incubated for at least 4 h with His-Tag Dynabeads® (Invitrogen/Dynal®). After elution of affinity-purified proteins and bound DNA with a buffer containing 300 mM imidazole, cross-linking was reversed by heating to 65 °C overnight. DNA was recovered and analyzed by PCR (27 amplification cycles), using specific primers against INO1 (−315/+5) and CHO2 (−360/−40) promoters or ACT1 gene (+841/+1165) as a control.
Miscellaneous procedures
Transformation of S. cerevisiae strains, PCR amplification and β-galactosidase assays have been described (Schwank et al. 1995; Wagner et al. 2001). Oligonucleotides used in this study are presented in supporting online Table 2.
Results
Importance of Sin3 binding proteins and multiple HDACs for ICRE-dependent gene expression
We have previously shown that repression of ICRE-dependent genes requires recruitment of pleiotropic corepressor complexes Sin3 and Ssn6/Tup1 by the pathway-specific repressor Opi1 (Wagner et al. 2001; Jäschke et al. 2011). We thus asked whether additional subunits of Sin3 complexes, Rpd3L and Rpd3S (Carrozza et al. 2005a, b), also contribute to repression of ICRE-dependent genes. The possible importance of subunits Ash1, Raf60, Dot6 and Tod6 (Colina and Young 2005; Shevchenko et al. 2008) also identified in the Rpd3L complex was not characterized in this study. For this investigation, we constructed a set of isogenic single mutants, each containing a synthetic minimal promoter, solely driven by ICRE activating motifs (integrated ICRE-CYC1-lacZ reporter gene; Schwank et al. 1995). To avoid confusion among two regulatory pathways comprising Sin3, we did not use an INO1-lacZ reporter gene, which in addition to ICRE motifs is also affected by the URS1-Ume6-Sin3 pathway (Lopes et al. 1993; Kadosh and Struhl 1997). As a means of deregulation in mutants, we used the ratio of β-galactosidase activities assayed in cells grown under derepressing and repressing conditions, respectively (D/R). As is apparent from data shown in Table 1, subunits of Sin3 complexes unequally affect ICRE-dependent gene expression. In agreement with previous results (Wagner et al. 2001), Sin3 is clearly of outstanding importance (D/R = 2.0), but loss of subunits Pho23 (4.4) and Sap30 (6.1) also weakens the degree of regulation and increases gene expression under repressing conditions at least by 1.5-fold.
It has been suggested that Sin3 executes its regulatory influence by recruiting the HDAC Rpd3 (Kadosh and Struhl 1997). In contrast to what was shown for a sin3 mutant, regulated expression of the reporter gene was not significantly altered in an rpd3 null mutant (Wagner et al. 2001; Table 2). Apparently, Opi1-dependent repression of Ino2-activated ICRE motifs is still effective even in the absence of Rpd3. We thus reasoned that a functional redundancy among yeast HDACs, Rpd3, Hda1, Hos1, Hos2 and Hos3, may exist (Rundlett et al. 1996; no consideration of Sir2 and related NAD-dependent HDACs). Consequently, we constructed deletion cassettes for HDAC genes to obtain all viable combinations of mutations rpd3, hda1, hos1, hos2 and hos3. However, we failed to obtain the following combinations of mutant alleles: triple mutant rpd3 hda1 hos2; quadruple mutants rpd3 hda1 hos1 hos2, rpd3 hda1 hos2 hos3 and rpd3 hos1 hos2 hos3; pentuple mutant rpd3 hda1 hos1 hos2 hos3.
In contrast to HDAC single mutants which showed normal repression, gene expression of the rpd3 hda1 double mutant under repressing conditions was increased (D/R = 3,4; cf. Table 2). Importantly, the triple mutant rpd3 hda1 hos1 (1.9) copied the regulatory phenotype of a sin3 null mutant (2.0), indicating that the corresponding HDACs may be recruited by Sin3 to trigger repression of ICRE-dependent genes.
Two non-overlapping sequences of Pho23 directly interact with Sin3
Pho23 is required for negative regulation of ICRE-dependent gene expression (Table 1). Interestingly, the C-terminal region of Pho23 (amino acids 276-330) comprising the zinc finger-related C4-HC3 plant homeodomain (PHD) shows strong similarity to the mammalian family of ING tumor suppressor proteins (inhibitor of growth) involved in control of cellular proliferation. We thus wished to investigate whether Pho23 binds to Sin3 and to establish a physical map of interacting domains in both proteins. A GST-Pho23 fusion protein (full-length) immobilized at glutathione (GSH) Sepharose was incubated with HA-tagged Sin3, synthesized either in E. coli or in S. cerevisiae. As shown in Fig. 1, Sin3 from either source could bind to Pho23, arguing for a direct interaction of both proteins. For a more precise mapping, GST fusions with Pho23 length variants were incubated with Sin3. Interestingly, two non-overlapping domains of Pho23 outside of ING similarity could bind to Sin3, each independent of the other. These domains were designated PSID1 and PSID2 (Pho23-Sin3 interaction domain; amino acids 80-151 and 208-250, respectively).
Mapping of Pho23 domains responsible for interaction with Sin3. Length variants of Pho23 were fused with GST, immobilized on GSH Sepharose and incubated with full-length HA3-Sin3 in total protein extracts, synthesized by S. cerevisiae (Sc, strain C13-ABY.S86, plasmid pCW117) or E. coli (Ec, strain BL21, plasmid pSW11). GST-Pho23 fusions are encoded by plasmids pSW5 (aa 1-330), pSW23 (aa 1-100), pSW24 (aa 1-151), pSW25 (aa 1-200), pSW26 (aa 251-330), pSW27 (aa 208-330), pSW28 (aa 154-330), pSW32 (aa 208-250), pSW33 (aa 154-200), pSW35 (aa 154-250) and pSW40 (aa 80-151). Input controls are shown at the bottom of the figure (20 % of protein used for the interaction assay). PS1, PS2 (=PSID1, 2): Pho23-Sin3 interaction domains 1 and 2; PHD plant homeodomain
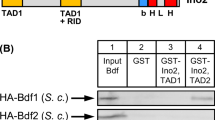
Vice versa, we also mapped Sin3 domains required for Pho23 interaction. GST fusions of Pho23 comprising either PSID1 or PSID2 were incubated with HA-tagged length variants of Sin3. Sin3 fragments used for the interaction assay represent individual structural and functional domains described previously (PAH1-PAH4, HID). As shown in Fig. 2, both PSID sequences could bind aa 801-1100 of Sin3 but not aa 601-950, indicating aa 951-1100 as the Sin3 core domain responsible for Pho23 recruitment. Thus, neither PAH domain is required for binding of Pho23, but instead sequences following the HDAC interaction domain are involved.
Mapping of Sin3 domains interacting with PSID1 and PSID2 of Pho23. GST-Pho23 fusion plasmids pSW25 and pSW32 were used to synthesize residues 1-200 (comprising PSID1) and 208-250 (comprising PSID2). The following expression plasmids representing individual PAH domains were used for synthesis of HA-tagged Sin3 length variants in S. cerevisiae: pCW117 (aa 1-1536), pCW83 (aa 1-300), pYJ91 (aa 301-600), pYJ90 (aa 601-950), pYJ89 (aa 801-1100) and pMP20 (aa 1101-1536). For input controls (shown in the right panel of the figure), 20 % of protein used for the interaction assay was analyzed. P1, P2, P3 and P4 (=PAH1-4): paired amphipathic helices 1–4
Although deregulation of ICRE-dependent genes was less evident for sap30 and sds3 mutants than with a pho23 mutant, we also investigated whether the corresponding proteins could directly interact with Sin3. Similar to what was found for Pho23, GST fusions of full-length Sds3 and Sap30 were able to bind HA-tagged Sin3 from yeast and bacterial protein extracts, arguing for direct interaction (Fig. 3a, b). The C-terminus of Sds3 (aa 201-327) interacts with the same internal domain of Sin3 which is also bound by Pho23 (aa 801-1100). The importance of the C-terminus of Sds3 for binding Sin3 agrees with the existence of a shortened protein in the yeast Saccharomyces kluyveri (173 amino acids; Cliften et al. 2003) lacking aa 1-144 of Sds3 from S. cerevisiae.
Direct interaction of Sds3 (a) and Sap30 (b) with Sin3. Fusion proteins GST-Sds3 (aa 1-327, encoded by pSW4), GST-Sds3 (aa 1-200, pMG4), GST-Sds3 (aa 201-327, pMG6) and GST-Sap30 (aa 1-201, pSW3), respectively, were immobilized on GSH Sepharose and incubated with protein extracts containing epitope-tagged length variants of Sin3. Protein extracts were prepared from transformants of S. cerevisiae (Sc, plasmids pCW117 or pYJ89, 801-1100 of Sin3) or E. coli (Ec, pSW11). Sin3 input controls are shown in Figs. 1 and 2
HDAC Hda1 binds to Sin3 via new interaction domains
As shown above, the absence of Rpd3, Hda1 and Hos1 imitates the regulatory defect observed in a strain lacking Sin3. Thus, in addition to Rpd3, HDACs Hda1 and Hos1 may also directly contact the Sin3 corepressor to induce a more compact chromatin. Class II HDAC Hda1 contains a long C-terminus of unknown function. To investigate a possible interaction of Hda1 with Sin3, we used GST-Hda1 fusions comprising amino acids 1-353 and 354-706, respectively. Indeed, the N-terminus of Hda1 with its deacetylase domain could bind HA-Sin3 full-length protein (cf. Fig. 4a). In subsequent studies, we therefore used the GST-Hda1 (1-353) fusion plasmid to map the Sin3 domain required for interaction. To allow a high resolution of mapping, HA-tagged length variants of Sin3 described above together with smaller fragments each synthesized in yeast were used for the interaction assay. As is shown in Fig. 4a, two non-overlapping Sin3 fragments representing amino acids 301-600 and 1101-1536 (comprising PAH2 and PAH4, respectively) were able to bind to Hda1. Importantly, these fragments do not contain the previously characterized HDAC interaction domain (HID, aa 729-1048). By using N- and C-terminal truncations of the fragment 301-600, we were able to show that it was not PAH2 (406-472), but instead a region in its C-terminus (aa 473-600) that was sufficient to bind Hda1. The same result was obtained with the Sin3 fragment comprising aa 1101-1210. We thus conclude that PAH4 (core sequence: aa 1131-1208) can also bind to Hda1. Obviously, Sin3 contains at least three HID regions which we now designate HID1 (former HID), HID2 and HID3 (cf. Fig. 4).
Mapping of Sin3 domains interacting with histone deacetylases Hda1, Hos1 and Rpd3. a GST-Hda1 fusion protein (residues 1-353 of Hda1, plasmid pMP1) was immobilized on GSH Sepharose and incubated with yeast protein extracts containing epitope-tagged length variants of Sin3. The following plasmids were used for synthesis of Sin3 length variants: pCW117 (aa 1-1536), pCW83 (aa 1-300), pYJ91 (aa 301-600), pYJ90 (aa 601-950), pYJ89 (aa 801-1100), pMP20 (aa 1101-1536), pYJ105 (aa 301-472), pMG7 (aa 473-600), pMG8 (aa 473-575), pMP22 (aa 1101-1300), pMG13 (aa 1101-1210) and pMG15 (aa 1140-1300). b Incubation of immobilized GST-Hos1 full-length fusion protein (470 amino acids, plasmid pYJ26) with yeast protein extracts containing epitope-tagged length variants of Sin3. c Incubation of immobilized GST-Rpd3 fusion protein (residues 141-300 of Rpd3, plasmid pMG17) with yeast protein extracts containing epitope-tagged length variants of Sin3. H1, H2, H3, histone deacetylase interaction domains HID1-3; P1, P2, P3 and P4: paired amphipathic helices PAH1-4
We next wanted to map the region within Hda1 which is required for interaction with Sin3. Length variants of Hda1 were fused with GST and used for interaction assays with Sin3 sequences representing HID2 and HID3 (aa 301-600 and 1101-1300, respectively). Hda1 fragments aa 201-300 and aa 251-353 could interact with HID2 and HID3 of Sin3 (Fig. 5a), indicating that the region of aa 251-300 may function as the core binding domain. Since identical results were obtained with proteins entirely produced in E. coli, we concluded that aa 251-300 of Hda1 mediated its interaction with HID2 and HID3 of Sin3.
Mapping of HDAC domains interacting with Sin3 domains HID1, HID2 and HID3. a Length variants of Hda1 were fused with GST, immobilized on GSH Sepharose and incubated with protein extracts from yeast or E. coli. The following GST expression plasmids were used: pMP1 (aa 1-353), pMP2 (aa 354-706), pMP12 (aa 201-353), pMP14 (aa 201-300) and pMP15 (aa 251-353). Epitope-tagged Sin3 domains representing HID2 and HID3 were expressed in yeast (Sc; plasmids pYJ91, aa 301-600 and pMP22, aa 1101-1300, respectively) or in E. coli (Ec; pMG22 and pMG23, respectively). b Length variants of Hos1 fused with GST were incubated with protein extracts from yeast or E. coli. The following GST expression plasmids were used: pYJ26 (aa 1-470), pMP9 (aa 1-350), pMP4 (aa 236-470) and pYJ85 (aa 236-400). c GST-Rpd3 fusion representing residues 141-300 of Rpd3 (plasmid pMG17) was incubated with protein extracts from yeast or E. coli. The epitope-tagged Sin3 domain representing HID1 was expressed in yeast (Sc; plasmid pYJ92, aa 801-950) or in E. coli (Ec; pMG26). Yeast input controls are shown in Fig. 4. DAC deacetylase core domain, n. t. not tested
HDAC Hos1 also binds to HID2 and HID3 within Sin3
In further studies, we investigated whether class I HDAC Hos1 could also bind to Sin3. Immobilization of GST-Hos1 (full-length) on GSH Sepharose indeed allowed retention of full-length Sin3. Subsequently, HA-tagged length variants of Sin3 described above were also used to map minimal interaction domains. As is shown in Fig. 4b, minimal fragments of Sin3, comprising HID2 and HID3 (=PAH4), were also able to bind to Hos1, while HID1 again failed to interact with Hos1. Thus, an identical interaction pattern was observed for Hda1 and Hos1. Similar to what was found for Hda1, the enzymatic deacetylase core domain of Hos1 (amino acids 236-400) is sufficient for this interaction (Fig. 5b). Again, binding of the Hos1 core region also occurs with HID2 and HID3 of Sin3 synthesized in E. coli, arguing for a direct interaction.
HDACs, Rpd3, Hda1 and Hos1, all bind to HID3/PAH4 of Sin3
Initially, HID(1) was identified as a conserved region of mSin3A (amino acids 524-851, similar to aa 729-1048 of yeast Sin3), which turned out as sufficient to bind the Rpd3-related mammalian HDAC2 (Laherty et al. 1997). We thus wished to test whether Rpd3 can also interact with a second domain of Sin3 and to perform a more precise mapping of the yeast HID1. Assuming that a conserved segment within HDACs may be responsible for Sin3 binding, we constructed a GST-Rpd3 fusion containing amino acids 141-300. Indeed, this fusion protein could interact with two Sin3 fragments covering HID1 (aa 601-950 and aa 801-1100; Fig. 4c). Using proteins entirely produced in E. coli, we could finally confirm that amino acids 801-950 indeed defined the functional core of HID1, which was directly bound by Rpd3 without yeast-specific auxiliary factors (Fig. 5c). In addition, GST-Rpd3 was also able to bind Sin3 fragments containing HID3/PAH4, synthesized either in yeast or in E. coli. Consequently, HID3/PAH4 functions as an interaction domain for at least three HDACs from yeast.
In vivo interaction of Sin3 with various HDACs
In addition to in vitro interaction assays, we also performed two-hybrid analyses to verify binding of Sin3 to HDACs, Hda1, Hos1 and Rpd3. Length variants of Sin3 comprising HID2 (aa 301-600) and HID1, HID2 and HID3 (aa 301-1536), respectively, were fused with the DNA-binding domain (DBD) of Gal4. Core deacetylase domains of Hda1, Hos1 and Rpd3 which have been shown to bind Sin3 in vitro were fused with Gal4 transcriptional activation domain (TAD). Sin3-HDAC interactions in vivo should reconstitute a functional Gal4 activator being able to induce the GAL2-ADE2 reporter gene of the recipient strain, thereby allowing growth of transformants in the absence of adenine. As is shown in Fig. 6, both DBD fusions of Sin3 in combination with empty TAD vector were unable to mediate growth on a medium lacking adenine. In contrast, co-transformation of DBD-Sin3 (aa 301-1536) with TAD fused to HDACs, Hda1, Hos1 or Rpd3, restored growth on adenine-free medium. We conclude that Sin3 comprising HID1-3 is able to interact with these HDACs in vivo. On the other hand, a minimal Sin3 (aa 301-600) with HID2 but devoid of HID1 and HID3 could interact with Hda1 and Hos1, but not with Rpd3.
Interaction of Sin3 with various HDACs shown by two-hybrid assays. The Gal4 DNA-binding domain (DBD) was fused with Sin3 fragments comprising HDAC interaction domains to give plasmids pMG42 (aa 301-600) and pJW31 (aa 301-1536). Correspondingly, Gal4 transcriptional activation domain (TAD) was fused with HDAC fragments to give pMG41 (Hda1, aa 201-353), pMG40 (Hos1, aa 236-400) and pMG39 (Rpd3, aa 141-300). Plasmids encoding fusions DBD-Opi1 and TAD-Sin3 (pJW8 and pJW1, respectively; Wagner et al. 2001) were used as a positive control. As a negative control, empty TAD vector pGAD-C1 was used. DBD and TAD pairs of fusion plasmids (selection markers: TRP1 and LEU2, respectively) were co-transformed into strain PJ69-4A, containing a GAL2-ADE2 fusion which allows growth in the absence of adenine when a functional Gal4 activator is reconstituted. Selection plates (-L -T and -L -T -A; absence of leucine, tryptophan and adenine) were incubated for 48 h
To finally investigate whether Hda1 and Hos1 are indeed recruited to ICRE-containing promoters regulated by Ino2 and Opi1, we performed ChIP analyses. We thus constructed strains containing variants of HDA1 and HOS1 which encode epitope-tagged proteins (His6). To analyze protein occupation of ICRE-containing promoters, we selected INO1 (biosynthesis of inositol; three ICRE motifs) and CHO2 (biosynthesis of choline, three ICRE motifs). The ACT1 gene encoding actin served as a negative control. As shown by pull-down experiments and two-hybrid studies, binding of HDACs follows a defined order of interaction events (ICRE-Ino2-Opi1-Sin3-HDACs; Wagner et al. 2001; this work). We thus also constructed ino2 deletion strains which should no longer allow recruitment of HDACs to ICRE-containing promoters. As shown in Fig. 7, Hda1 and Hos1 indeed occupy ICRE-dependent promoters INO1 and CHO2 under conditions of gene repression by inositol and choline (lines 1 and 3; INO2 intact), while binding to ACT1 is substantially less effective. The figure also shows that binding of HDACs, Hda1 and Hos1, requires a functional INO2 gene (lines 2 and 4, ino2 deletion mutants). These findings agree with results derived from in vitro interaction studies and provide in vivo evidence that ICRE-dependent promoters are affected by more than a single histone deacetylase.
In vivo binding of Hda1 and Hos1 to ICRE-containing promoters INO1 and CHO2 shown by chromatin immunoprecipitation (ChIP). INO2 wild-type strains FKY40 and FKY42 contain HDA1 and HOS1 variants with C-terminal His6 and HA3 epitopes at their natural chromosomal position. FKY44 and FKY46 were constructed by subsequent introduction of an ino2 deletion allele. Strains were grown under conditions of inositol/choline repression. Details of the procedure are given in “Materials and methods”. After reversal of cross-linking, DNA samples were analyzed by PCR using gene-specific primers which allow amplification of ~320 bp fragments. IN input control (analysis of total lysate samples), IP immunoprecipitate (analysis of samples enriched for His-tagged Hda1 and Hos1, respectively)
Discussion
The importance of Sin3 as a pleiotropic corepressor for several regulatory systems of gene expression in eukaryotes is well known (Grzenda et al. 2009). The existence of structural motifs suitable for protein–protein interactions suggested that Sin3 functioned as a recruiting platform for effector enzymes (e.g., HDAC Rpd3), allowing their access to specific promoter regions. In this study, we have investigated the influence of subunits of the Sin3 complex for regulation of phospholipid biosynthetic genes. Characterization of various combinations of HDAC mutations led us to conclude that Rpd3 was not the sole enzyme responsible for gene repression mediated by Sin3. This hypothesis is further confirmed by our demonstration of physical interaction of Sin3 with HDACs, Rpd3, Hda1 and Hos1. The existence of three non-overlapping HDAC interaction domains (HID1, HID2 and HID3) indicates that a greater combinatorial variety of Sin3-containing complexes than previously assumed is able to contribute to target gene repression.
The Sin3/Rpd3L complex comprises at least 12 subunits which may stabilize interactions of Sin3 with specific repressors and thus unequally influences different regulatory pathways. Surprisingly, phenotypes of the corresponding null mutants differ with respect to the regulatory system, leading to enhanced repression of yeast silent mating type loci and at telomeric positions, but increased initiation at the PHO5 promoter and URS1-containing promoters (Vannier et al. 1996; Zhang et al. 1998; Sun and Hampsey 1999; Loewith et al. 2001). To investigate the importance of 13 subunits identified in complexes Rpd3L and Rpd3S for ICRE-dependent gene expression, we used single null mutants for analyzing the regulation of a synthetic minimal promoter which did not contain a URS1 motif such as the widely used INO1 control region. Besides sin3, deregulation was most apparent with a pho23 mutant, which was initially isolated because of increased expression of the PHO5-encoded acid phosphatase (Lau et al. 1998). Pho23 is especially interesting due to the similarity of its C-terminal PHD finger with mammalian tumor suppressors of the ING family. This structural motif strongly binds to trimethylated lysine of histone H3 (H3K4me3) and mediates recruitment of Sin3/Rpd3L to the repressed PHO5 promoter (Shi et al. 2006; Wang et al. 2011). Our results show that Pho23 contains two non-overlapping domains which both can directly bind to a domain of Sin3 close to HID1. In contrast to Rpd3, Pho23 interacts with aa 801-1100 but not with aa 601-950, suggesting that aa 951-1100 are at least necessary for binding of Pho23. Similarly, Sds3 also binds to aa 801-1100 of Sin3 (summarized in Fig. 8a). We could also demonstrate that Sds3 and Sap30 directly interact with Sin3. For the mammalian ING1b protein, association with Sin3 through direct interaction with Sap30 has been shown (Kuzmichev et al. 2002).
a Summary of mapped interaction domains within Sin3. Data were those presented in this work or taken from Wagner et al. (2001) (for Opi1), Washburn and Esposito (2001) (for Ume6) and Xie et al. (2011) (for Sap30). HCR highly conserved region of unknown function, HID histone deacetylase interaction domains; P1, P2, P3 and P4 (=PAH1-4): paired amphipathic helices 1–4. b Hydrophobic-amphipathic pattern in an alignment of sequences from Sin3 HID1 of yeast and H. sapiens. Hydrophobic residues at heptad positions 7/1 and 4/5 are shown in bold. c Hydrophobic-amphipathic pattern in an alignment of sequences from HDACs, Rpd3, Hda1 and Hos1, shown to bind Sin3 HID motifs. #, hydrophobic amino acids
Importantly, comparison of ICRE-dependent gene regulation in sin3 and rpd3 null mutants revealed a striking difference, indicating that Rpd3 could not be responsible for the entire repressing influence of the Sin3 complex. Inspection of the data published by Kadosh and Struhl (1997) on URS1-Ume6 dependent repression showed that similarly loss of gene repression in a sin3 mutant was clearly stronger than in an rpd3 mutant. Both findings can be explained by assuming a separate repressing activity within the Sin3 complex which is not mediated by histone deacetylation or, alternatively, triggered by recruitment of HDAC isoenzymes. Our first evidence supporting the latter view resulted from construction of multiple HDAC null mutants and subsequent analysis of ICRE-dependent gene expression. Although we failed to obtain certain combinations of mutant alleles (indicating that total loss of class I and class II HDACs leads to synthetic lethality), a triple mutant rpd3 hda1 hos1 showed deregulation similar to that observed with a sin3 single mutant.
Our studies with the ICRE-dependent reporter gene revealed that repression and derepression were affected in the sin3 strain and in several mutants lacking multiple HDACs (cf. Table 2). These data could mean that activators as well as repressors may utilize Sin3, as it has been shown for the Ssn6/Tup1 corepressor which also acts as a positive cofactor of Gcn4 activator (Kim et al. 2005). Similarly, a positive role of Sin3-Rpd3 for activation of osmosensitive genes such as HSP12 has been shown (de Nadal et al. 2004). Alternatively, pleiotropic deregulation in a sin3 strain affecting carbon and nitrogen metabolism may cause a general shortage of important metabolites, indirectly leading to less efficient derepression.
Similar regulatory phenotypes of mutants sin3 and rpd3 hda1 hos1 do not necessarily mean that the corresponding HDACs are indeed recruited by Sin3. We have previously shown that repression by Opi1 is also mediated to a certain degree by the Ssn6/Tup1 corepressor (Jäschke et al. 2011), which can recruit HDACs, Rpd3, Hda1, Hos1 and Hos2 (Watson et al. 2000; Wu et al. 2001b; Davie et al. 2003). Thus, glucose repression of the invertase gene SUC2 was abolished in an rpd3 hos1 hos2 triple mutant. These results clearly show that corepressor function may be indeed mediated by more than a single HDAC, supporting the idea of some redundancy among them. Consequently, we performed interaction studies to decide whether Sin3 is also able to recruit additional HDACs. According to the results of our mutational analysis, we concentrated on HDACs, Hda1 and Hos1. Indeed, GST fusions of both enzymes were able to interact with full-length Sin3. In contrast, we could not reproducibly show interaction of Hos2 and Hos3 with Sin3. These in vitro studies were confirmed in vivo by two-hybrid experiments, using fusion plasmids which encode domains of Sin3 and HDACs, Hda1, Hos1 and Rpd3. Using ChIP analysis, we also demonstrate recruitment of Hda1 and Hos1 to promoter regions of INO1 and CHO2. We selected these genes because they show strong regulation by inositol/choline which is mediated by three ICRE upstream motifs (Hoppen et al. 2005; Kodaki et al. 1991). Promoter recruitment of HDACs required a functional Ino2 activator which directly binds ICRE motifs and establishes the core for subsequent interactions with Opi1, Sin3 and finally Hda1/Hos1.
Detailed mapping experiments revealed that Sin3 not only contains the single HDAC interaction domain initially mapped in mammalian proteins (HID, aa 801-950 for the yeast protein), but also two additional segments which we designate HID2 (aa 473-600) and HID3 (aa 1101-1210; cf. Fig. 8a). A hydrophobic-amphipathic sequence pattern which is completely conserved between yeast and human Sin3 proteins could be identified within the renamed HID1 (Fig. 8b). HID2 immediately follows the PAH2 domain and may be specific for yeast Sin3 proteins. This conclusion can be derived from the finding that mSin3 proteins lack sequence motifs convincingly similar to HID2. In contrast, HID3 which is identical to PAH4 is a conserved structural feature of Sin3 proteins. The structural pattern of PAH4 deviates from PAH1, PAH2 and PAH3 and its function remained unclear. To our knowledge, the work reported here is the first demonstration of a physical function for PAH4 within yeast Sin3. For mammalian mSin3A, interaction of PAH4 with TPR motifs of an O-glycosyltransferase (OGT) specific for N-acetylglucosamine (NAG) has been demonstrated (Yang et al. 2002), indicating that NAG transfer on transcription factors may inhibit their activity. In conclusion, the existence of at least three HDAC interaction domains indicates that a single Sin3 protein could be able to simultaneously bind Rpd3, Hda1 and Hos1. Alternatively, Sin3 may only interact with a single HDAC isoenzyme, leading to the coexistence of individual HDAC/Sin3 complexes. This latter view would explain why previous purification procedures using tagged Rpd3 failed to detect Hda1 and Hos1 within the complex (Carrozza et al. 2005b). It should be mentioned that Hda1 also exists in a distinct complex together with non-catalytic subunits Hda2 and Hda3 (Wu et al. 2001a).
Interaction experiments with HDAC length variants showed that Sin3 binding maps within their catalytic deacetylase domains. However, catalytic and binding functions are not identical. Two histidine residues (H247, H248 for Hda1) which are essential for binding the substrates water and acetyllysine via hydrogen bonds lie outside the core interaction domain of Hda1 (aa 251-300). We also searched for hydrophobic-amphipathic sequence motifs within HDACs which possibly could bind Sin3. As shown in Fig. 8c, hydrophobic residues appear at positions 7 or 1 and 4 or 5 of heptad repeats placed above sequence segments of Rpd3, Hda1 and Hos1. By using bacterial protein extracts, we could show that Rpd3, Hda1 and Hos1 are able to directly interact with Sin3 domains. This does not rule out that auxiliary proteins of the Sin3/Rpd3L complex such as Pho23, Sds3 and Sap30 increase Sin3-HDAC interaction in yeast. The interactions shown for yeast Sin3 in this work and previous publications are summarized in Fig. 8a.
References
Ambroziak J, Henry SA (1994) INO2 and INO4 gene products, positive regulators of phospholipid biosynthesis in Saccharomyces cerevisiae, form a complex that binds to the INO1 promoter. J Biol Chem 269:15344–15349
Carrozza MJ, Florens L, Swanson SK, Shia WJ, Anderson S, Yates J, Washburn MP, Workman JL (2005a) Stable incorporation of sequence specific repressors Ash1 and Ume6 into the Rpd3L complex. Biochim Biophys Acta 1731:77–87
Carrozza MJ, Li B, Florens L, Suganuma T, Swanson SK, Lee KK, Shia WJ, Anderson S, Yates J, Washburn MP, Workman JL (2005b) Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell 123:581–592
Chen M, Hancock LC, Lopes JM (2007) Transcriptional regulation of yeast phospholipid biosynthetic genes. Biochim Biophys Acta 1771:310–321
Cliften P, Sudarsanam P, Desikan A, Fulton L, Fulton B, Majors J, Waterston R, Cohen BA, Johnston M (2003) Finding functional features in Saccharomyces genomes by phylogenetic footprinting. Science 301:71–76
Cobb J, van Attikum H (2010) Mapping genomic targets of DNA helicases by chromatin immunoprecipitation in Saccharomyces cerevisiae. Methods Mol Biol 587:113–126
Colina AR, Young D (2005) Raf60, a novel component of the Rpd3 histone deacetylase complex required for Rpd3 activity in Saccharomyces cerevisiae. J Biol Chem 280:42552–42556
Davie JK, Edmondson DG, Coco CB, Dent SY (2003) Tup1-Ssn6 interacts with multiple class I histone deacetylases in vivo. J Biol Chem 278:50158–50162
De Antoni A, Gallwitz D (2000) A novel multi-purpose cassette for repeated integrative epitope tagging of genes in Saccharomyces cerevisiae. Gene 246:179–185
De Nadal E, Zapater M, Alepuz PM, Sumoy L, Mas G, Posas F (2004) The MAPK Hog1 recruits Rpd3 histone deacetylase to activate osmoresponsive genes. Nature 427:370–374
Dietz M, Heyken WT, Hoppen J, Geburtig S, Schüller HJ (2003) TFIIB and subunits of the SAGA complex are involved in transcriptional activation of phospholipid biosynthetic genes by the regulatory protein Ino2 in the yeast Saccharomyces cerevisiae. Mol Microbiol 48:1119–1130
Grzenda A, Lomberk G, Zhang JS, Urrutia R (2009) Sin3: master scaffold and transcriptional corepressor. Biochim Biophys Acta 1789:443–450
Heyken WT, Repenning A, Kumme J, Schüller HJ (2005) Constitutive expression of yeast phospholipid biosynthetic genes by variants of Ino2 activator defective for interaction with Opi1 repressor. Mol Microbiol 56:696–707
Hildmann C, Riester D, Schwienhorst A (2007) Histone deacetylases-an important class of cellular regulators with a variety of functions. Appl Microbiol Biotechnol 75:487–497
Hoppen J, Repenning A, Albrecht A, Geburtig S, Schüller HJ (2005) Comparative analysis of promoter regions containing binding sites of the heterodimeric transcription factor Ino2/Ino4 involved in yeast phospholipid biosynthesis. Yeast 22:601–613
James P, Halladay J, Craig EA (1996) Genomic libraries and a host strain designed for highly efficient two-hybrid selection in yeast. Genetics 144:1425–1436
Jäschke Y, Schwarz J, Clausnitzer D, Müller C, Schüller HJ (2011) Pleiotropic corepressors Sin3 and Ssn6 interact with repressor Opi1 and negatively regulate transcription of genes required for phospholipid biosynthesis in the yeast Saccharomyces cerevisiae. Mol Genet Genomics 285:91–100
Joshi AA, Struhl K (2005) Eaf3 chromodomain interaction with methylated H3-K36 links histone deacetylation to Pol II elongation. Mol Cell 20:971–978
Kadosh D, Struhl K (1997) Repression by Ume6 involves recruitment of a complex containing Sin3 corepressor and Rpd3 histone deacetylase to target promoters. Cell 89:365–371
Kim SJ, Swanson MJ, Qiu H, Govind CK, Hinnebusch AG (2005) Activator Gcn4p and Cyc8p/Tup1p are interdependent for promoter occupancy at ARG1 in vivo. Mol Cell Biol 25:11171–11183
Kodaki T, Hosaka K, Nikawa J, Yamashita S (1991) Identification of the upstream activation sequences responsible for the expression and regulation of the PEM1 and PEM2 genes encoding the enzymes of the phosphatidylethanolamine methylation pathway in Saccharomyces cerevisiae. J Biochem 109:276–287
Kumme J, Dietz M, Wagner C, Schüller HJ (2008) Dimerization of yeast transcription factors Ino2 and Ino4 is regulated by precursors of phospholipid biosynthesis mediated by Opi1 repressor. Curr Genet 54:35–45
Kuzmichev A, Zhang Y, Erdjument-Bromage H, Tempst P, Reinberg D (2002) Role of the Sin3-histone deacetylase complex in growth regulation by the candidate tumor suppressor p33(ING1). Mol Cell Biol 22:835–848
Laherty CD, Yang WM, Sun JM, Davie JR, Seto E, Eisenman RN (1997) Histone deacetylases associated with the mSin3 corepressor mediate mad transcriptional repression. Cell 89:349–356
Lau WTW, Schneider KR, O’Shea EK (1998) A genetic study of signaling processes for repression of PHO5 transcription in Saccharomyces cerevisiae. Genetics 150:1349–1359
Loewith R, Smith JS, Meijer M, Williams TJ, Bachman N, Boeke JD, Young D (2001) Pho23 is associated with the Rpd3 histone deacetylase and is required for its normal function in regulation of gene expression and silencing in Saccharomyces cerevisiae. J Biol Chem 276:24068–24074
Lopes JM, Schulze KL, Yates JW, Hirsch JP, Henry SA (1993) The INO1 promoter of Saccharomyces cerevisiae includes an upstream repressor sequence (URS1) common to a diverse set of yeast genes. J Bacteriol 175:4235–4238
Malavé TM, Dent SY (2006) Transcriptional repression by Tup1-Ssn6. Biochem Cell Biol 84:437–443
Mumberg D, Müller R, Funk M (1994) Regulatable promoters of Saccharomyces cerevisiae: comparison of transcriptional activity and their use for heterologous expression. Nucl Acids Res 22:5767–5768
Rundlett SE, Carmen AA, Kobayashi R, Bavykin S, Turner BM, Grunstein M (1996) HDA1 and RPD3 are members of distinct yeast histone deacetylase complexes that regulate silencing and transcription. Proc Natl Acad Sci USA 93:14503–14508
Rundlett SE, Carmen AA, Suka N, Turner BM, Grunstein M (1998) Transcriptional repression by UME6 involves deacetylation of lysine 5 of histone H4 by RPD3. Nature 392:831–835
Schüller HJ, Hahn A, Tröster F, Schütz A, Schweizer E (1992) Coordinate genetic control of yeast fatty acid synthase genes FAS1 and FAS2 by an upstream activation site common to genes involved in membrane lipid biosynthesis. EMBO J 11:107–114
Schwank S, Ebbert R, Rautenstrauss K, Schweizer E, Schüller HJ (1995) Yeast transcriptional activator INO2 interacts as an Ino2p/Ino4p basic helix-loop-helix heteromeric complex with the inositol/choline-responsive element necessary for expression of phospholipid biosynthetic genes in Saccharomyces cerevisiae. Nucl Acids Res 23:230–237
Shevchenko A, Roguev A, Schaft D, Buchanan L, Habermann B, Sakalar C, Thomas H, Krogan NJ, Shevchenko A, Stewart AF (2008) Chromatin central: towards the comparative proteome by accurate mapping of the yeast proteomic environment. Genome Biol 9:R167
Shi X, Hong T, Walter KL, Ewalt M, Michishita E, Hung T, Carney D, Peña P, Lan F, Kaadige MR, Lacoste N, Cayrou C, Davrazou F, Saha A, Cairns BR, Ayer DE, Kutateladze TG, Shi Y, Côté J, Chua KF, Gozani O (2006) ING2 PHD domain links histone H3 lysine 4 methylation to active gene repression. Nature 442:96–99
Silverstein RA, Ekwall K (2005) Sin3: a flexible regulator of global gene expression and genome stability. Curr Genet 47:1–17
Slekar KH, Henry SA (1995) SIN3 works through two different promoter elements to regulate INO1 gene expression in yeast. Nucl Acids Res 23:1964–1969
Sun ZW, Hampsey M (1999) A general requirement for the Sin3-Rpd3 histone deacetylase complex in regulating silencing in Saccharomyces cerevisiae. Genetics 152:921–932
Vannier D, Balderes D, Shore D (1996) Evidence that the transcriptional regulators SIN3 and RPD3, and a novel gene (SDS3) with similar functions, are involved in transcriptional silencing in S. cerevisiae. Genetics 144:1343–1353
Wagner C, Dietz M, Wittmann J, Albrecht A, Schüller HJ (2001) The negative regulator Opi1 of phospholipid biosynthesis in yeast contacts the pleiotropic repressor Sin3 and the transcriptional activator Ino2. Mol Microbiol 41:155–166
Wang H, Clark I, Nicholson PR, Herskowitz I, Stillman DJ (1990) The Saccharomyces cerevisiae SIN3 gene, a negative regulator of HO, contains four paired amphipathic helix motifs. Mol Cell Biol 10:5927–5936
Wang SS, Zhou BO, Zhou JQ (2011) Histone H3 lysine 4 hypermethylation prevents aberrant nucleosome remodeling at the PHO5 promoter. Mol Cell Biol 31:3171–3181
Washburn BK, Esposito RE (2001) Identification of the Sin3-binding site in Ume6 defines a two-step process for conversion of Ume6 from a transcriptional repressor to an activator in yeast. Mol Cell Biol 21:2057–2069
Watson AD, Edmondson DG, Bone JR, Mukai Y, Yu Y, Du W, Stillman DJ, Roth SY (2000) Ssn6-Tup1 interacts with class I histone deacetylases required for repression. Genes Dev 14:2737–2744
Wu J, Carmen AA, Kobayashi R, Suka N, Grunstein M (2001a) HDA2 and HDA3 are related proteins that interact with and are essential for the activity of the yeast histone deacetylase HDA1. Proc Natl Acad Sci USA 98:4391–4396
Wu J, Suka N, Carlson M, Grunstein M (2001b) TUP1 utilizes histone H3/H2B specific HDA1 deacetylase to repress gene activity in yeast. Mol Cell 7:117–126
Xie T, He Y, Korkeamaki H, Zhang Y, Imhoff R, Lohi O, Radhakrishnan I (2011) Structure of the 30-kDa Sin3-associated protein (SAP30) in complex with the mammalian Sin3A corepressor and Its role in nucleic acid binding. J Biol Chem 286:27814–27824
Yang XJ, Seto E (2008) The Rpd3/Hda1 family of lysine deacetylases: from bacteria and yeast to mice and men. Nat Rev Mol Cell Biol 9:206–218
Yang X, Zhang F, Kudlow JE (2002) Recruitment of O-GlcNAc transferase to promoters by corepressor mSin3A: coupling protein O-GlcNAcylation to transcriptional repression. Cell 110:69–80
Zhang Y, Sun ZW, Iratni R, Erdjument-Bromage H, Tempst P, Hampsey M, Reinberg D (1998) SAP30, a novel protein conserved between human and yeast, is a component of a histone deacetylase complex. Mol Cell 1:1021–1031
Acknowledgments
This work has been supported by the Deutsche Forschungsgemeinschaft (DFG). We thank Marius Wanjek for valuable support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Ronne.
M. Grigat and Y. Jäschke contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Grigat, M., Jäschke, Y., Kliewe, F. et al. Multiple histone deacetylases are recruited by corepressor Sin3 and contribute to gene repression mediated by Opi1 regulator of phospholipid biosynthesis in the yeast Saccharomyces cerevisiae . Mol Genet Genomics 287, 461–472 (2012). https://doi.org/10.1007/s00438-012-0692-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-012-0692-x