Abstract
Structural genes of phospholipid biosynthesis in the yeast S. cerevisiae are activated by the heterodimeric transcription factor Ino2 + Ino4, binding to ICRE (inositol/choline-responsive element) promoter motifs. In the presence of phospholipid precursors inositol and choline, Ino2-dependent activation is inhibited by the Opi1 repressor which interacts with Ino2. In this work, we systematically investigated the importance of regulatory mechanisms possibly affecting ICRE-dependent gene expression. Autoregulatory expression of INO2, INO4 and OPI1 was abolished by promoter exchange experiments, showing that autoregulation of regulators contributes to the degree of differential gene expression but is not responsible for it. Using GFP fusion proteins, Ino2 and Ino4 were found to localize to the nucleus under conditions of repression and derepression. Interestingly, nuclear localization of Ino2 required a functional INO4 gene. Targeting of a lexA–Ino2 fusion to a heterologous promoter containing lexA operator motifs revealed a constitutive gene activation which was not influenced by phospholipid precursors. We could show that Ino2-dependent activation of a lexA–Ino4 fusion is affected by inositol and choline. Since gene activation required interaction of Ino2 and Ino4 mediated by their helix–loop–helix domains, formation/dissociation of the heterodimer must be considered as an important step of target gene regulation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A question of central importance for studies of eukaryotic gene expression is to understand how an extra- or intracellular signal is ultimately executed to increase or diminish transcriptional initiation; in other words: How are the regulators regulated? (Falvey and Schibler 1991). The differential expression of yeast structural genes involved in phospholipid biosynthesis by the availability of metabolic precursor molecules is an attractive model system for investigating the transduction of metabolic signals (reviewed by Chen et al. 2007). Genes such as INO1 (biosynthesis of inositol; Lopes and Henry 1991), CHO1 (choline; Bailis et al. 1992), FAS1 and FAS2 (fatty acids; Schüller et al. 1992a) are coordinately activated by ICRE promoter sequences (inositol/choline-responsive element = UASINO; consensus sequence: WYTTCAYRTG; a comprehensive compilation of motifs is given in Hoppen et al. 2005). ICRE motifs are bound by a heterodimer of positive regulators Ino2 and Ino4 (Schüller et al. 1992b; Ambroziak and Henry 1994; Schwank et al. 1995), both containing a basic helix–loop–helix (bHLH) structural motif (Hoshizaki et al. 1990; Nikoloff et al. 1992) which are necessary and sufficient for dimer formation and specific interaction with the ICRE (Dietz et al. 2003). Recent studies have shown that expression of several genes probably unrelated to phospholipid metabolism is also affected by Ino2 and Ino4 (Hoppen et al. 2005; Jesch et al. 2005; Chen and Lopes 2007). Importantly, Ino2 (but not Ino4) contains two separate transcriptional activation domains TAD1 and TAD2 (Schwank et al. 1995) as well as a repressor interaction domain (RID) which is required for functional deactivation of Ino2 by the repressor Opi1 (Wagner et al. 2001; Heyken et al. 2005). Opi1 is necessary for repression of ICRE-dependent transcription when inositol and choline (IC) are present in excess (Greenberg et al. 1982; Wagner et al. 1999). Under these conditions, Opi1 is localized in the nucleus (Loewen et al. 2003) and prevents Ino2 from activation of target genes by recruitment of the pleiotropic corepressor Sin3 (Wagner et al. 2001). Consequently, Ino2 variants defective for interaction with Opi1 mimic the opi1 mutant phenotype and allow constitutive transcription of ICRE-dependent genes (Heyken et al. 2005). Similarly, depletion of IC (derepressing conditions) causes accumulation of phosphatidic acid (PA) which binds to Opi1, leading to its retention at the endoplasmic reticulum or nuclear membrane (Loewen et al. 2003, 2004). Although Opi1 contains a basic leucine zipper motif (Hoshizaki et al. 1990), no evidence for DNA-binding by Opi1 has been obtained yet. In the absence of Opi1, ICRE-bound Ino2 can contact coactivator complexes (Snf1 with its histone kinase function, Lo et al. 2005; SAGA, TFIIB, Dietz et al. 2003) and activates transcription of target genes.
Although previous work has shown that the balance of Opi1 and Ino2 is critical for activation and repression (Hosaka et al. 1994; Schwank et al. 1997; Wagner et al. 1999), the mechanism of Ino2 regulation remained unknown. Eukaryotic activators may be controlled by several single or combined mechanisms: (1) biosynthetic control of the activator itself (e. g. Gal4, Gcn4; Griggs and Johnston 1991; Hinnebusch 1984); (2) control of intracellular localization (e. g. Pho4, Swi5; O’Neill et al. 1996; Moll et al. 1991); (3) regulation of DNA binding of the activator (e. g. Ace1, MyoD; Fürst and Hamer 1989; Benezra et al. 1990); (4) regulation of transcriptional activation (e. g. Gal4, Cat8; Ma and Ptashne 1987; Rahner et al. 1999). We thus wished to uncouple various mechanisms affecting transcriptional regulators Ino2, Ino4 and Opi1 and to separately evaluate their contribution to differential expression of ICRE-containing structural genes. It turned out that autoregulation of INO2, INO4 and OPI1 (mediated by ICRE upstream motifs; Schüller et al. 1992b; Ashburner and Lopes 1995; Wagner et al. 1999) contributes to the regulation of target genes but is not essential. We show that dimerization of Ino2 and Ino4 is regulated by IC, requiring a functional RID (but not an intact DNA-binding domain) within Ino2. We conclude that the integrity of the Ino2–Ino4 heterodimer must be considered as an important trigger of IC-mediated transcriptional regulation.
Materials and methods
Yeast strains, media and growth conditions
All strains of S. cerevisiae used in this work (compiled in Table 1) are isogenic to strain JS91.15–23 (Schwank et al. 1995). Variants of INO2, INO4 and OPI1 were introduced into strains JKY5 and JKY53 by integrative transformation. Synthetic complete (SC) media used for selective growth of transformants and conditions of gene regulation by IC have been described (Schüller et al. 1992a).
Plasmid constructions
Plasmids constructed and used for this study are listed in Table 2. Promoter variants of INO2, INO4 and OPI1 were inserted into integrative plasmids pRS403 (HIS3; Sikorski and Hieter 1989), YIplac204 (TRP1; Gietz and Sugino 1988) and YIplac128 (LEU2), respectively. MET25 promoter fusions were obtained by insertion of reading frame cassettes amplified by PCR into p413-MET25 (Mumberg et al. 1994) and subsequent transfer of the expression cassette into integrative vectors. To modify promoter sequences (CANNTG core sequences within ICRE motifs upstream of INO2, INO4 and OPI1 were converted into unrelated cleavage sites of restriction enzymes) or specific residues in the coding regions (ino2 E245A, ino4 E54A and ino4 L94A), the QuikChange site-directed mutagenesis kit from Strategene was used. The INO2 L118A mutation has been described previously (Heyken et al. 2005).
The artificial (lexAOp)8-GAL1 upstream region containing eight copies of the lexA operator was amplified from strain L40 (Hollenberg et al. 1995) and fused with lacZ of plasmid YEp356R (Myers et al. 1986). Reporter plasmid pJK119 was obtained by transfer of the (lexAOp)8–GAL1–lacZ fusion to integrative vector YIp352 (Hill et al. 1986). To obtain fusions of the DNA-binding domain of lexA repressor (residues 1–87) with INO2 and INO4, plasmid pSH2-1 (Hanes and Brent 1989) was used as a template for PCR amplifications. Similarly, GFPUV cassettes fused with regulatory genes were amplified from plasmid pBAD–GFPUV (Clontech). Epitope-tagged regulators were constructed by insertion of reading frame cassettes for OPI1, INO2 and INO4 into HA fusion plasmid p426-MET25HA (Mumberg et al. 1994) to give pCW116 (MET25–HA3–OPI1), pMD2 (MET25–HA3–INO2) and pJK51 (MET25–HA3–lexA DBD–INO4), respectively.
Fluorescence microscopy of GFP fusions
Fluorescence microscopy followed the procedure described by Taheri et al. (2000). Yeast strains containing integrated GFPUV fusions (Crameri et al. 1996) with INO2, INO4 or OPI1 were grown to log phase in selective medium (SCD-His) with 1 mM methionine under conditions of IC repression or derepression. Cells from 1 ml of the cultures were concentrated by centrifugation and immediately viewed on a Zeiss Axiovert microscope by either differential interference contrast microscopy (DIC, Nomarski optics) or fluorescence microscopy using a GFP filter set (AHF Analysentechnik AG; Tübingen, Germany). Images were taken using a Xillix Microimager digital camera and the Improvision Openlab software (Improvision; Coventry, UK).
Miscellaneous procedures
Transformation of S. cerevisiae strains, immunoblot analysis, PCR amplification and β-galactosidase assays were performed as previously described (Schwank et al. 1995; Wagner et al. 2001). Enzyme assays were repeated three times, using five independent transformants each. Standard deviations observed with integrated reporter genes ICRE–CYC1–lacZ and (lexAOp)8-GAL1–lacZ were 20% of the mean value or below.
Results
Influence of autoregulation of INO2, INO4 and OPI1 on expression of target genes
Regulatory genes INO2, INO4 and OPI1 each contain a single ICRE motif in their respective upstream regions and therefore control their own expression (Schüller et al. 1992b; Ashburner and Lopes 1995; Schwank et al. 1997; Wagner et al. 1999). Thus, the ratio of positive factors Ino2 + Ino4 and repressor Opi1 may vary under repressing and derepressing conditions, respectively. However, it remained unclear whether autoregulation of regulatory genes subsequently affects the expression pattern of ICRE-dependent target genes. We therefore wished to systematically compare the regulation of an ICRE-activated reporter gene in the presence of INO2, INO4 and OPI1 controlled by upstream regions devoid of their natural ICREs.
For each regulatory gene, three integrating plasmids were constructed: (1) a plasmid containing the entire wild-type control region (autoregulated); (2) a plasmid of identical length but mutated at the ICRE core element upstream of the respective gene (autoregulation eliminated); (3) a plasmid containing the coding region downstream of the heterologous MET25 promoter (cf. Fig. 1a). All regulatory genes controlled by promoter variants were confirmed as functional (not shown). We also constructed the triple mutant JKY5 (Δopi1 Δino2 Δino4, integrated ICRE–CYC1–lacZ reporter gene) and re-introduced regulatory genes OPI1, INO2 and INO4 driven by different control regions.
Variation of INO2, INO4 and OPI1 promoters and influence on the regulation of an ICRE-dependent reporter gene. a Comparison of promoter variants upstream of regulatory genes INO2, INO4 and OPI1. b INO2, INO4 and OPI1 promoter variants on plasmids containing selection markers HIS3, TRP1 and LEU2 were integrated at the respective loci of strain JKY5 (contains chromosomal null mutations Δino2::kanMX Δino4::kanMX Δopi1::kanMX and an integrated ICRE–CYC1–lacZ reporter gene). Transformants were cultivated under selective conditions in the presence of 1 mM methionine. Specific β-galactosidase activities are given in nmoles oNPG hydrolyzed per min per mg of protein (U/mg). R conditions of inositol/choline repression (200 μM inositol + 2 mM choline); D conditions of inositol/choline derepression (5 μM inositol + 5 μM choline); D/R ratio of enzyme activities under derepressing and repressing conditions, respectively; SD standard deviation of the mean value
The resulting strains were cultivated under conditions of IC repression and derepression, respectively, and assayed for expression of the ICRE-dependent reporter gene. As shown in Fig. 1b, loss of autoregulation partially weakened regulation by IC, especially when INO2 variants with a mutated ICRE were used. Nevertheless, even when autoregulation was completely eliminated (ΔICRE triple mutant), target genes were clearly regulated by IC (derepression factor D/R = 6.1 compared with D/R = 14.0 when wild-type promoters controlled INO2, INO4 and OPI1). We conclude that autoregulation of regulators affects ICRE-dependent gene expression as a mechanism of signal amplification but is not responsible for triggering IC repression. To rule out other contributions of the natural control regions, we also investigated ICRE-mediated gene regulation with INO2, INO4 and OPI1 each driven by the MET25 promotor. This promoter is of moderate strength and can be further weakened by addition of methionine (Mumberg et al. 1994), allowing an expression level which is typical of regulatory genes. As is also apparant from Fig. 1b, expression and regulation of the ICRE-dependent reporter gene is almost identical when regulatory genes are activated by the MET25 promoter or by ICRE-mutated natural promoters (D/R = 6.5 and 6.1).
Influence of phospholipid precursors on intracellular localization of Ino2, Ino4 and Opi1
Several transcriptional regulators are controlled at the level of nuclear import/export, thereby allowing or preventing access to their DNA target sites. For Opi1, anchoring at the endoplasmic reticulum in the absence of IC has been previously shown (Loewen et al. 2003). For a systematic comparison of intracellular localization of Ino2, Ino4 and Opi1 under conditions of IC repression and derepression, we constructed bifunctional GFP fusion genes driven by the MET25 promoter which were subsequently integrated into the genome of the respective null mutants. To obtain fluorescent signals with weakly expressed fusion genes, the GFPUV variant of the green fluorescent protein was used (Crameri et al. 1996). The resulting strains MDY50 (GFP-INO2), MDY49 (INO4-GFP) and MDY54 (OPI1-GFP) were able to grow in the absence of inositol (not shown) and showed regulated expression of an ICRE-dependent reporter gene with a weakened MET25 promoter (cf. Fig. 2a). Thus, fluorescence of GFP should reliably correspond with the localization of the respective regulator under both conditions of inositol supply.
a GFP fusion proteins with full-length Ino2, Ino4 and Opi1, respectively, allow regulated expression of an ICRE-dependent reporter gene. GFP fusion genes on integrating HIS3 plasmids (pMD122: INO4-GFPUV, pMD155: GFPUV-INO2, pMD160: OPI1-GFPUV) were expressed under control of the MET25 promoter. Plasmids were transformed into derivatives of strain SIRP3 (integrated ICRE–CYC1–lacZ reporter gene), deleted for the chromosomal copy of the regulatory gene fused to GFP. To avoid overproduction of regulatory proteins, transformants were grown in the presence of 1 mM methionine under repressing (200 μM inositol + 2 mM choline) and derepressing conditions (5 μM inositol + 5 μM choline), respectively. Specific β-galactosidase activities are given in nmoles oNPG hydrolyzed per min per mg of protein (U/mg). M25Pr, MET25 promoter; SD standard deviation of the mean value. b Subcellular localization of regulatory proteins Ino2, Ino4 and Opi1 under conditions of inositol/choline repression and derepression. Representative cells of the strains indicated expressing fusion proteins GFP-Ino2, Ino4-GFP or Opi1-GFP were viewed by differential interference contrast microscopy (DIC, Nomarski optics) or by fluorescence microscopy (GFP)
As shown in Fig. 2b, nuclear fluorescence could be observed with the GFP–Ino2 fusion protein under repressing and derepressing conditions (strain MDY50). Interestingly, a uniform fluorescence signal in the cytoplasm was observed in the absence of INO4 (MDY53). Similar to Ino2, Ino4-GFP constitutively localized to the nucleus (MDY49), even in the absence of INO2 (not shown). These results indicate that Ino4 is required for nuclear import of Ino2 while, in contrast, Ino2 does not affect intracellular localization of Ino4. Opi1-GFP fluorescence is clearly concentrated in the nucleus under repressing conditions while the signal is much more distributed under derepressing conditions. This agrees with previous results, arguing for a retention of the Opi1 repressor at the endoplasmic reticulum or the nuclear membrane when maximal ICRE-dependent gene expression is required (Loewen et al. 2003).
Influence of phospholipid precursors on transcriptional activation by Ino2
Even when activator proteins constitutively localize to the nucleus, binding to their DNA target sites or transcriptional activation may be regulated. We thus fused INO2 to the DNA-binding domain of bacterial lexA repressor and assayed Ino2-dependent gene activation of a lexAOp-containing reporter gene. These tests were performed with three strains (wild-type, Δino4, Δopi1), each cultivated under repressing and derepressing conditions, respectively. Nuclear import of the resulting lexADBD–Ino2 fusion protein in the absence of Ino4 should be guaranteed by an intrinsic nuclear localization sequence within lexA (Rhee et al. 2000).
With constitutive DNA-binding mediated by lexA, Ino2-dependent activation was not influenced by IC supply (cf. Fig. 3). Although TAD2 of Ino2 partially overlaps with the Opi1 repressor interaction domain (RID; Heyken et al. 2005), no increase of transcriptional activation was observed with an isogenic opi1 null mutant. Similarly, absence of Ino4 did not alter Ino2-dependent activation. We conclude that activation mediated by Ino2 occurs constitutively, at least in the context of the full-length protein as analyzed here.
Influence of phospholipid precursors on transcriptional activation. Isogenic strains containing an integrated lexAop–GAL1–lacZ reporter gene were transformed with plasmid pJS519, containing a lexA DBD–INO2 fusion gene (INO2 full length). Specific β-galactosidase activities are given in nmoles oNPG hydrolyzed per min per mg of protein (U/mg). R conditions of inositol/choline repression (200 μM inositol + 2 mM choline); D conditions of inositol/choline derepression (5 μM inositol + 5 μM choline); D/R ratio of enzyme activities under derepressing and repressing conditions, respectively. bHLH basic helix–loop–helix; DBD DNA-binding domain; SD standard deviation of the mean value; TAD transcriptional activation domain
Influence of phospholipid precursors on dimerization of Ino2 and Ino4
For basic helix–loop–helix proteins, dimerization is a prerequisite for DNA binding (Littlewood and Evan 1998). Since Ino2 must be considered as a constitutive activator, we wished to investigate whether dimerization of Ino2 and Ino4 is affected by the availability of phospholipid precursors. We used a simplified two-hybrid system with a lexAOp-dependent reporter gene and a lexA–Ino4 fusion protein. Transcriptional activation of the reporter gene is entirely mediated by full-length Ino2 which is recruited to promoter-bound lexA-Ino4 via its HLH domain.
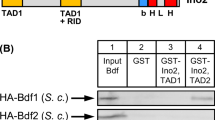
To rule out a biosynthetic variation of regulators under our experimental conditions, MET25–HA fusions of INO2, lexA–INO4 and OPI1 were transformed into the corresponding single mutants. After cultivation of transformants under repressing and derepressing conditions, protein extracts were analyzed by immunoblot analysis. As is apparent from Fig. 4, biosynthesis of regulatory proteins Ino2, lexA–Ino4 and Opi1 was indeed unaffected by the variation of phospholipid precursors.
Constitutive biosynthesis of Ino2, lexA–Ino4 and Opi1 regulators expressed under MET25 promoter control. Plasmids pMD2 (MET25–HA3–INO2), pJK51 (MET25–HA3–lexA DBD–INO4) and pCW116 (MET25–HA3–OPI1) were transformed into mutants SS92.3-1 (Δino2), JS91.15-9 (Δino4) and MBY1 (Δopi1), respectively. Yeast transformants were grown in selective media under conditions of inositol/choline repression (R 200 μM inositol + 2 mM choline) and derepression (D 5 μM inositol + 5 μM choline), respectively. Fifty micrograms of total yeast protein was separated in each lane and immunoblot analyses were performed using anti-HA monoclonal antibody. Two antigenic signals are usually obtained for Ino2 (Ambroziak and Henry 1994)
For a direct comparison of fusion constructs, we used the authentic ICRE-dependent reporter gene (requiring DNA-binding by Ino2 and Ino4) as well as the heterologous lexAOp-dependent reporter gene (requiring formation of the Ino2/Ino4 heterodimer). To avoid deregulation as a result of unphysiological levels of Ino2 and Opi1 (Schwank et al. 1997; Wagner et al. 1999), all regulatory genes were controlled by the MET25 promoter, weakened by supplementation of media with 1 mM methionine.
As expected, activation of either reporter gene failed in the absence of INO2 or INO4 while regulated activation was found with authentic INO2, INO4 and OPI1. Although concentration of these regulators was no longer influenced by phospholipid precursors, the ICRE-dependent reporter gene was still regulated 6,5-fold (Fig. 5a). An almost identical regulatory pattern was observed with a lexA–Ino4 fusion (6,1-fold derepression). Importantly, a considerable degree of derepression was also found with the lexAOp-driven reporter gene (4,0-fold increase; Fig. 5b), indicating that regulation could be transferred to a heterologous cis-acting element. Next, we replaced functional genes INO2 and INO4 by variants with a detrimental mutation in the basic region of the DNA-binding bHLH-domain (mutant alleles ino2E245A and ino4E54A; R. Rohde et al. unpublished). Indeed, activation of the ICRE-dependent reporter gene was completely abolished (Fig. 5a) while the lexAOp-dependent reporter gene showed an even stronger expression compared to wild-type INO regulators. This increase can be explained by the failure of mutant Ino2(E245A) and lexA-Ino4(E54A) to bind to chromosomal ICRE target sites, allowing a higher degree of occupation at the lexAOp. In contrast, wild-type lexA–Ino4 is a bi-functional protein which can bind to both ICRE (together with Ino2) and lexAOp. Importantly, activation with b-domain mutants of Ino2 and Ino4 was still fully regulated, indicating that DNA-binding of Ino2/Ino4 is not required for the IC-response. As a control, we also used mutant allele ino4L94A, encoding an Ino4 variant which is no longer able to interact with Ino2 (unpublished). Indeed, regulated activation was abolished with both reporter genes. Finally, we investigated the influence of a repression-resistant Ino2 variant which is no longer bound by the Opi1 repressor due to a replacement at a critical position of the RID (INO2 mutant allele L118A; Heyken et al. 2005). In the presence of this Ino2 variant, repression of either reporter gene by phospholipid precursors was no longer detectable, mimicing the effect of an opi1 null mutation. While permanent recruitment of lexA–Ino2 to a lexAOp-dependent promoter lead to constitutive gene activation (one-hybrid situation, Fig. 3, see above), regulated activation was observed with promoter-bound lexA–Ino4 and its dimerization partner Ino2 (simplified two-hybrid situation, Fig. 5b). From these results we conclude that dimerization of Ino2 and Ino4 is regulated by phospholipid precursors via Opi1-interaction with Ino2. Moreover, regulated dimerization is not influenced by the ability of Ino2 and Ino4 to bind DNA.
Regulation of ICRE- and lexAOp-dependent reporter genes by phospholipid precursors. Regulatory genes INO2, INO4 and OPI1 were controlled by the MET25 promoter. Variants of MET25-INO2, MET25-(lexA DBD)-INO4 and MET25-OPI1 were integrated at loci his3, trp1 and leu2, respectively. Transformants were uniformly cultivated under selective conditions in the presence of 1 mM methionine to achieve an adequate level of regulators. Interactions among promoter elements and regulatory proteins are depicted for each type of experiment. a Use of an integrated ICRE–CYC1–lacZ reporter gene (strain JKY5). b Use of an integrated lexAop–GAL1–lacZ reporter gene (strain JKY53). Specific β-galactosidase activities are given in nmoles oNPG hydrolyzed per min per mg of protein (U/mg). R conditions of inositol/choline repression (200 μM inositol + 2 mM choline); D conditions of inositol/choline derepression (5 μM inositol + 5 μM choline); D/R ratio of enzyme activities under derepressing and repressing conditions, respectively. bHLH basic helix–loop–helix; DBD DNA-binding domain; SD standard deviation of the mean value; TAD transcriptional activation domain
Discussion
The analysis of signalling systems of transcriptional control in eukaryotes has shown that the observed regulatory phenotype often results from several partial mechanisms which may act additively, cooperatively and/or redundantly (e. g. glucose repression and galactose induction of GAL structural genes; Johnston et al. 1994; Bhat and Murthy 2001). In this work, we wished to systematically analyse whether biosynthetic variation, intracellular localization, transcriptional activation and DNA binding of regulators Ino2, Ino4 and Opi1 contribute to IC-dependent expression of target genes.
We and others have previously shown that INO2, INO4 and OPI1 genes each contain a functional ICRE upstream sequence, arguing for autoregulation of regulators (Schüller et al. 1992b; Ashburner and Lopes 1995; Wagner et al. 1999). It should be mentioned that the ICRE variants mediating IC-regulated expresson of INO2 (AATTCACATGT), INO4 (TATTCACATGT) and OPI1 (TCTTCATATGC) deviate from the optimal UAS sequence (WYTTCACATGS; Schüller et al. 1995). Nevertheless, Ino2 + Ino4 could efficiently bind to these motifs in vitro (Schwank et al. 1997; Wagner et al. 1999). It should be mentioned that autoregulation of INO4 has been questioned by others, suggesting a posttranscriptional mechanism of IC control of INO4 expression, instead (Robinson and Lopes 2000). However, using the MET25 promoter, we found constant levels of HA-tagged Ino4 (not shown) and lexA-Ino4 (this work) under repressing and derepressing conditions. We conclude that IC-mediated regulation of INO4 expression (about threefold) is a result of its specific upstream region and the ICRE therein. In this work, we replaced regulatory genes of phospholipid biosynthesis by variants mutated at their ICRE promoter motifs. The resulting variants were able to functionally complement their respective null mutations and led to regulation of an ICRE-dependent reporter gene by IC which was weakened but still effective (14-fold derepression vs. sixfold derepression observed in the presence of three promoter mutants). Loss of INO2 autoregulation (8–10-fold with its wild-type control region) showed the most significant contribution. Almost identical results were obtained with regulatory genes driven by methionine-repressed MET25 fusions. We conclude that target genes are differentially expressed upon variation of IC supply even with Ino2, Ino4 and Opi1 regulators at a constitutive steady-state level.
The functional compartmentation of a eukaryotic cell provides a regulatory means of transcriptional control so that many activators and repressors show signal-dependent regulation by nuclear import/export. A related mechanism has been shown for Opi1 which interacts with the Scs2 protein of the endoplasmic reticulum (ER) and/or nuclear membrane in the presence of phosphatidic acid (PA) via its FFAT structural motif (two phenylalanine residues in an acidic tract; Loewen et al. 2003, 2004). Our results with a bifunctional OPI1–GFP fusion are in agreement with these findings. While the signal observed with repressed cells concentrated in the nucleus, fluorescence under derepressing conditions was no longer exclusively nuclear. A cluster of basic amino acids (residues 109–138) extending the leucine zipper domain of Opi1 may be responsible for PA binding as well as for nuclear import. While Opi1 shows signal-dependent shuttling between ER and nucleus, Ino4-GFP was localized in the nucleus constitutively. Ino4 contains an extremely basic region (residues 29–40) which may function as a nuclear localization sequence (NLS). In contrast, the GFP–Ino2 fusion protein was nuclear only when INO4 was functional. A basic cluster of amino acids as a conventional NLS is absent from the Ino2 sequence. We thus assume that Ino4 heterodimerizes with Ino2 in the cytoplasm and may subsequently function as a mediator of nuclear import for the resulting complex. In conclusion, our data argue for a nuclear translocation of Ino2 which depends on Ino4 but is not affected by phospholipid precursors.
We have previously shown that the Opi1 repressor interaction domain within Ino2 partially overlaps transcriptional activation domain TAD2 (Heyken et al. 2005). This appears reminiscent of Gal80 binding to the Gal4 TAD prior to galactose induction (Ma and Ptashne 1987) and Dig1 inhibiting activation by Ste12 in the absence of a pheromone signal (Olson et al. 2000). However, transcriptional activation by a lexA–Ino2 fusion was not significantly influenced by IC availability or by deletion of OPI1 and INO4, respectively (cf. Fig. 3). Nevertheless, it should be mentioned that Gardenour et al. (2004) isolated INO2 missense mutants mapping to TAD1 (D20H, F21L) which led to partially increased target gene activation (threefold) under repressing conditions.
We finally investigated whether interaction between Ino2 and Ino4 is influenced by IC regulation. To do this, we fused Ino4 to the DNA-binding domain of bacterial lexA repressor and used a lexAOp-dependent reporter gene to monitor gene expression. Since Ino4 lacks a transcriptional activation domain by itself, activation of the reporter gene should occur only if Ino2 forms a heterodimer with the promoter-recruited lexA–Ino4. No biosynthetic variation of Ino2, lexA–Ino4 and Opi1 under repressing and derepressing conditions was found with our experimental set-up (MET25-driven regulatory genes). In a similar one-hybrid experiment, transcriptional activation by lexA–Ino2 turned out as constitutive (see above). Our experimental approach further requires that DNA-binding of lexA fusion proteins is not affected by IC regulation. Although it is known that fusion domains may affect DNA-binding by lexA (Golemis and Brent 1992), previous results from our group showed that DNA-binding at least by lexA–Ino4 was not regulated by IC (Schwank et al. 1995).
Importantly, lexA–Ino4 + Ino2 led to a fourfold regulation of the lexAOp-dependent reporter gene. With a reporter gene containing the ICRE as the authentic binding site of Ino2 + Ino4 and the same set of regulators, a sixfold derepression was observed (cf. Fig. 5a, b). Gene regulation by IC of the lexAOp-containing promoter was completely maintained with missense mutants of Ino2 and Ino4, each defective for DNA-binding due to alteration of an essential glutamate residue of both basic regions (E245 and E54). Thus, differential activation of the heterologous reporter gene does not require binding of Ino2 + Ino4 to the homologous cis-element, arguing for their regulated dimerization as a critical means of target gene control. Expectedly, disruption of the Ino2–Ino4 interaction by mutation of an essential residue within helix II (L94) completely abolished activation by either promoter motif. In contrast, deletion of OPI1 or mutation of L118 necessary for Opi1 binding to the Ino2 RID similarly caused constitutive gene expression with both reporter genes.
Since dimerization is a prerequisite for DNA-binding by bHLH proteins, occupancy of the ICRE would be also affected by regulated formation of the Ino2 + Ino4 heterodimer in a native promoter situation. As a conclusion from our findings we hypothesize that interaction of Opi1 with Ino2 RID may trigger a conformational change within Ino2 which interferes with Ino4 binding, finally leading to less occupation of genomic ICRE target sites and gene repression. In a previous study, Brickner and Walter (2004) investigated the influence of the unfolded protein response (UPR) on INO1 transcription. Using chromatin immunoprecipitation, these authors describe constitutive binding of Ino2 + Ino4 to the INO1 promoter. Instead, intranuclear relocation of the INO1 locus from the nucleoplasm to the nuclear periphery is postulated as a critical step of gene derepression. However, to our surprise, we find that the original data of Brickner and Walter clearly show a twofold increase of Ino2 binding to the INO1 promoter under derepressing conditions which is in partial agreement with our results. It should be also emphasized that both hypotheses assuming regulated heterodimerization of Ino2 and Ino4 (this work) and subsequent alteration of nuclear localization of INO1 (Brickner and Walter 2004) are compatible with each other. A model which summarizes regulatory transitions based on findings from this and previous work is shown in Fig. 6. In this model, we distinguish between an early phase of repression in which a chain of interactions (ICRE–Ino2–Opi1–Sin3–HDACs) leads to a specific local alteration of chromatin structure and a late phase (steady state conditions) with Opi1 weakening formation of the Ino2–Ino4 heterodimer.
Hypothesis on regulatory transitions upon establishment of IC repression. a Under conditions of IC limitation, accumulation of phosphatidic acid (PA) leads to retention of Opi1 at the ER and/or nuclear membrane via Scs2 (Loewen et al. 2003, 2004), allowing Ino2 + Ino4 to fully activate ICRE-driven target genes. b Opi1 is released from its membrane anchoring upon increase of IC concentration and subsequent consumption of PA. Interaction with the RID of Ino2 specifically targets Opi1 to ICRE-containing genes (Wagner et al. 2001; Heyken et al. 2005). Since Opi1 simultaneously contacts the pleiotropic corepressor Sin3, local modification of chromatin via histone deacetylases (HDACs) initiates the repressed state (Kadosh and Struhl 1997). c Opi1 also weakens interaction of Ino2 and Ino4 (this work), leading to release of the heterodimer from its ICRE target sites in late repression and under steady state conditions. Autoregulation of regulators as a putative means of signal amplification is not considered here. DBD DNA-binding domain; NLS nuclear localization sequence; RID repressor interaction domain; TAD transcriptional activation domain
References
Ambroziak J, Henry SA (1994) INO2 and INO4 gene products, positive regulators of phospholipid biosynthesis in Saccharomyces cerevisiae, form a complex that binds to the INO1 promoter. J Biol Chem 269:15344–15349
Ashburner BP, Lopes JM (1995) Autoregulated expression of the yeast INO2 and INO4 helix–loop–helix activator genes effects cooperative regulation on their target genes. Mol Cell Biol 15:1709–1715
Bailis AM, Lopes JM, Kohlwein SD, Henry SA (1992) Cis and trans regulatory elements required for regulation of the CHO1 gene of Saccharomyces cerevisiae. Nucleic Acids Res 20:1411–1418
Benezra R, Davis RL, Lockshon D, Turner DL, Weintraub H (1990) The protein Id: a negative regulator of helix–loop–helix DNA binding proteins. Cell 61:49–59
Bhat PJ, Murthy TV (2001) Transcriptional control of the GAL/MEL regulon of yeast Saccharomyces cerevisiae: mechanism of galactose-mediated signal transduction. Mol Microbiol 40:1059–1066
Brickner JH, Walter P (2004) Gene recruitment of the activated INO1 locus to the nuclear membrane. PLoS Biol 2:e342
Chen M, Lopes JM (2007) Multiple basic helix–loop–helix proteins regulate expression of the ENO1 gene of Saccharomyces cerevisiae. Eukaryot Cell 6:786–796
Chen M, Hancock LC, Lopes JM (2007) Transcriptional regulation of yeast phospholipid biosynthetic genes. Biochim Biophys Acta 1771:310–321
Crameri A, Whitehorn EA, Tate E, Stemmer WP (1996) Improved green fluorescent protein by molecular evolution using DNA shuffling. Nat Biotechnol 14:315–319
Dietz M, Heyken WT, Hoppen J, Geburtig S, Schüller HJ (2003) TFIIB and subunits of the SAGA complex are involved in transcriptional activation of phospholipid biosynthetic genes by the regulatory protein Ino2 in the yeast Saccharomyces cerevisiae. Mol Microbiol 48:1119–1130
Falvey E, Schibler U (1991) How are the regulators regulated? FASEB J 5:309–314
Fürst P, Hamer D (1989) Cooperative activation of a eukaryotic transcription factor: interaction between Cu(I) and yeast ACE1 protein. Proc Natl Acad Sci USA 86:5267–5271
Gardenour KR, Levy J, Lopes JM (2004) Identification of novel dominant INO2 c mutants with an Opi− phenotype. Mol Microbiol 52:1271–1280
Gietz RD, Sugino A (1988) New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene 74:527–534
Golemis EA, Brent R (1992) Fused protein domains inhibit DNA binding by LexA. Mol Cell Biol 12:3006–3014
Greenberg ML, Goldwasser P, Henry SA (1982) Characterization of a yeast regulatory mutant constitutive for synthesis of inositol-1-phosphate synthase. Mol Gen Genet 186:157–163
Griggs DW, Johnston M (1991) Regulated expression of the GAL4 activator gene in yeast provides a sensitive genetic switch for glucose repression. Proc Natl Acad Sci USA 88:8597–8601
Hanes SD, Brent R (1989) DNA specificity of the bicoid activator protein is determined by homeodomain recognition helix residue 9. Cell 57:1275–1283
Heyken WT, Repenning A, Kumme J, Schüller HJ (2005) Constitutive expression of yeast phospholipid biosynthetic genes by variants of Ino2 activator defective for interaction with Opi1 repressor. Mol Microbiol 56:696–707
Hill JE, Myers AM, Koerner TJ, Tzagoloff A (1986) Yeast/E. coli shuttle vectors with multiple unique restriction sites. Yeast 2:163–167
Hinnebusch AG (1984) Evidence for translational regulation of the activator of general amino acid control in yeast. Proc Natl Acad Sci USA 81:6442–6446
Hollenberg SM, Sternglanz R, Cheng PF, Weintraub H (1995) Identification of a new family of tissue-specific basic helix–loop–helix proteins with a two-hybrid system. Mol Cell Biol 15:3813–3822
Hoppen J, Repenning A, Albrecht A, Geburtig S, Schüller HJ (2005) Comparative analysis of promoter regions containing binding sites of the heterodimeric transcription factor Ino2/Ino4 involved in yeast phospholipid biosynthesis. Yeast 22:601–613
Hosaka K, Nikawa JI, Kosaki T, Yamashita S (1994) Cloning and characterization of the SCS1 gene required for the expression of genes in yeast phospholipid synthesis. J Biochem 115:131–136
Hoshizaki DK, Hill JE, Henry SA (1990) The Saccharomyces cerevisiae INO4 gene encodes a small, highly basic protein required for derepression of phospholipid biosynthesis enzymes. J Biol Chem 265:4736–4745
Jesch SA, Zhao X, Wells MT, Henry SA (2005) Genome-wide analysis reveals inositol, not choline, as the major effector of Ino2p–Ino4p and unfolded protein response target gene expression in yeast. J Biol Chem 280:9106–9118
Johnston M, Flick JS, Pexton T (1994) Multiple mechanisms provide rapid and stringent glucose repression of GAL gene expression in Saccharomyces cerevisiae. Mol Cell Biol 14:3834–3841
Kadosh D, Struhl K (1997) Repression by Ume6 involves recruitment of a complex containing Sin3 corepressor and Rpd3 histone deacetylase to target promoters. Cell 89:365–371
Littlewood TD, Evan GI (1998) Helix–loop–helix transcription factors, 3rd edn. Oxford University Press, Oxford
Lo WS, Gamache ER, Henry KW, Yang D, Pillus L, Berger SL (2005) Histone H3 phosphorylation can promote TBP recruitment through distinct promoter-specific mechanisms. EMBO J 24:997–1008
Loewen CJ, Roy A, Levine TP (2003) A conserved ER targeting motif in three families of lipid binding proteins and in Opi1p binds VAP. EMBO J 22:2025–2035
Loewen CJ, Gaspar ML, Jesch SA, Delon C, Ktistakis NT, Henry SA, Levine TP (2004) Phospholipid metabolism regulated by a transcription factor sensing phosphatidic acid. Science 304:1644–1647
Lopes JM, Henry SA (1991) Interaction of trans and cis regulatory elements in the INO1 promoter of Saccharomyces cerevisiae. Nucleic Acids Res 19:3987–3994
Ma J, Ptashne M (1987) The carboxy-terminal 30 amino acids of GAL4 are recognized by GAL80. Cell 50:137–142
Moll T, Tebb G, Surana U, Robitsch H, Nasmyth K (1991) The role of phosphorylation and the CDC28 protein kinase in cell cycle-regulated nuclear import of the S. cerevisiae transcription factor SWI5. Cell 66:743–758
Mumberg D, Müller R, Funk M (1994) Regulatable promoters of Saccharomyces cerevisiae: comparison of transcriptional activity and their use for heterologous expression. Nucleic Acids Res 22:5767–5768
Myers AM, Tzagoloff A, Kinney DM, Lusty CJ (1986) Yeast shuttle and integrative vectors with multiple cloning sites suitable for construction of lacZ fusions. Gene 45:299–310
Nikoloff DM, McGraw P, Henry SA (1992) The INO2 gene of Saccharomyces cerevisiae encodes a helix–loop–helix protein that is required for activation of phospholipid synthesis. Nucleic Acids Res 20:3253
O’Neill EM, Kaffman A, Jolly ER, O’Shea EK (1996) Regulation of PHO4 nuclear localization by the PHO80–PHO85 cyclin-CDK complex. Science 271:209–212
Olson KA, Nelson C, Tai G, Hung W, Yong C, Astell C, Sadowski I (2000) Two regulators of Ste12p inhibit pheromone-responsive transcription by separate mechanisms. Mol Cell Biol 20:4199–4209
Rahner A, Hiesinger M, Schüller HJ (1999) Deregulation of gluconeogenic structural genes by variants of the transcriptional activator Cat8p of the yeast Saccharomyces cerevisiae. Mol Microbiol 34:146–156
Rhee Y, Gurel F, Gafni Y, Dingwall C, Citovsky V (2000) A genetic system for detection of protein nuclear import and export. Nat Biotechnol 18:433–437
Robinson KA, Lopes JM (2000) The promoter of the yeast INO4 regulatory gene: a model of the simplest yeast promoter. J Bacteriol 182:2746–2752
Schüller HJ, Hahn A, Tröster F, Schütz A, Schweizer E (1992a) Coordinate genetic control of yeast fatty acid synthetase genes FAS1 and FAS2 by an upstream activation site common to genes involved in the membrane lipid biosysnthesis. EMBO J 11:107–114
Schüller HJ, Schorr R, Hoffmann B, Schweizer E (1992b) Regulatory gene INO4 of yeast phospholipid biosynthesis is positively autoregulated and functions as a trans-activator of fatty acid synthase genes FAS1 and FAS2 from Saccharomyces cerevisiae. Nucleic Acids Res 20:5955–5961
Schüller HJ, Richter K, Hoffmann B, Ebbert R, Schweizer E (1995) DNA binding site of the yeast heteromeric Ino2p/Ino4p basic helix–loop–helix transcription factor: structural requirements as defined by saturation mutagenesis. FEBS Lett 370:149–152
Schwank S, Ebbert R, Rautenstrauss K, Schweizer E, Schüller HJ (1995) Yeast transcriptional activator INO2 interacts as an Ino2p/Ino4p basic helix–loop–helix heteromeric complex with the inositol/choline-responsive element necessary for expression of phospholipid biosynthetic genes in Saccharomyces cerevisiae. Nucleic Acids Res 23:230–237
Schwank S, Hoffmann B, Schüller HJ (1997) Influence of gene dosage and autoregulation of the regulatory genes INO2 and INO4 on inositol/choline-repressible gene transcription in the yeast Saccharomyces cerevisiae. Curr Genet 31:462–468
Sikorski RS, Hieter P (1989) A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122:19–27
Taheri N, Köhler T, Braus GH, Mösch HU (2000) Asymmetrically localized Bud8p and Bud9p proteins control yeast cell polarity and development. EMBO J 19:6686–6696
Wagner C, Blank M, Strohmann B, Schüller HJ (1999) Overproduction of the Opi1 repressor inhibits transcriptional activation of structural genes required for phospholipid biosynthesis in the yeast Saccharomyces cerevisiae. Yeast 15:843–854
Wagner C, Dietz M, Wittmann J, Albrecht A, Schüller HJ (2001) The negative regulator Opi1 of phospholipid biosynthesis in yeast contacts the pleiotropic repressor Sin3 and the transcriptional activator Ino2. Mol Microbiol 41:155–166
Acknowledgments
This work has been supported by the Deutsche Forschungsgemeinschaft (DFG). We thank Prof. H. -U. Mösch, Prof. G. Braus and Dr. N. Taheri for several suggestions and invaluable support with fluorescence microscopy.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by G. Braus.
Rights and permissions
About this article
Cite this article
Kumme, J., Dietz, M., Wagner, C. et al. Dimerization of yeast transcription factors Ino2 and Ino4 is regulated by precursors of phospholipid biosynthesis mediated by Opi1 repressor. Curr Genet 54, 35–45 (2008). https://doi.org/10.1007/s00294-008-0197-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-008-0197-7