Abstract
Repressor protein Opi1 is required to negatively regulate yeast structural genes of phospholipid biosynthesis in the presence of precursor molecules inositol and choline (IC). Opi1 interacts with the paired amphipathic helix 1 (PAH1) of pleiotropic corepressor Sin3, leading to recruitment of histone deacetylases (HDACs). Mutational analysis of the Opi1–Sin3 interaction domain (OSID) revealed that hydrophobic OSID residues L56, V59 and V67 of Opi1 are indispensable for gene repression. Our results also suggested that repression is not executed entirely via Sin3. Indeed, we could show that OSID contacts a second pleiotropic corepressor, Ssn6 (=Cyc8), which together with Tup1 is also able to recruit HDACs. Interestingly, mutations sin3 and ssn6 turned out as synthetically lethal. Our analysis further revealed that OSID not only binds to PAH1 but also interacts with tetratricopeptide repeats (TPR) of Ssn6. This interaction could no longer be observed with Opi1 OSID variants. To trigger gene repression, Opi1 must also interact with activator Ino2, using its activator interaction domain (AID). AID contains a hydrophobic structural motif reminiscent of a leucine zipper. Our mutational analysis of selected positions indeed confirmed that residues L333, L340, V343, V350, L354 and V361 are necessary for repression of Opi1 target genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Transcriptional repression of regulated structural genes in eukaryotes often depends on pleiotropic corepressor complexes which do not bind directly to cis-acting upstream elements but instead are recruited by specific DNA-binding proteins to target promoters. A well-known corepressor conserved from yeast to mammalian systems is Sin3 (reviewed by Grzenda et al. 2009), initially described as a negative regulator of HO expression in S. cerevisiae (Sternberg et al. 1987; Wang et al. 1990) but also required for repression of glucoamylase genes (Yoshimoto et al. 1992), meiotic genes (Strich et al. 1989), HMR/HML loci (Vannier et al. 1996) and phospholipid biosynthetic genes (Hudak et al. 1994). In addition to Sin3, eukaryotic corepressor complexes Ssn6-Tup1 (not found in higher eukaryotes; Malavé and Dent 2006), N-CoR/SMRT (specific for ligand-free nuclear receptors; Jones and Shi 2003), NuRD (in mammalian systems; McDonel et al. 2009), Groucho (in Drosophila and higher metazoa; Gasperowicz and Otto 2005) and Topless (in plants; Liu and Karmarkar 2008) have been identified for which recruitment of various histone deacetylases was demonstrated.
We and others have previously shown that Sin3 is important for repression of genes involved in phospholipid biosynthesis (Slekar and Henry 1995; Wagner et al. 2001). When phospholipid precursors inositol and choline (IC) are limiting (derepressing conditions), a heterodimer of transcription factors Ino2 and Ino4 binds to the regulatory motif ICRE (inositol/choline-responsive element = UASINO; Schüller et al. 1992; Ambroziak and Henry 1994; Schwank et al. 1995) upstream of phospholipid biosynthetic genes such as INO1, CHO1, FAS1 and FAS2 and triggers transcriptional activation (reviewed by Chen et al. 2007). Under these conditions, cellular concentration of phosphatidic acid (PA) as the central metabolite of phospholipid biosynthesis is high, leading to retention of the pathway-specific transcriptional repressor Opi1 at the endoplasmic reticulum (Loewen et al. 2004). In contrast, at a high concentration of IC (repressing conditions), level of PA drops, leading to release of Opi1 repressor which subsequently enters the nucleus and counteracts gene activation mediated by Ino2. Repression is mediated by direct contact of Opi1 domain AID (activator interaction domain) and Ino2 domain RID (repressor interaction domain; Heyken et al. 2005). Via a distinct domain (OSID, Opi1–Sin3 interaction domain; Wagner et al. 2001), Opi1 also binds to corepressor Sin3, allowing recruitment of histone deacetylase Rpd3 (Kadosh and Struhl 1997) which subsequently may modify local chromatin and prevent access of the transcriptional machinery.
Sin3 works as a platform for multiple protein–protein interactions and contains four paired amphipathic helix motifs (PAH1-4) which are not functionally redundant but instead bind to individual partner proteins (Wang et al. 1990; Wang and Stillman 1993). Opi1 specifically contacts PAH1 of Sin3 (Wagner et al. 2001) while PAH2 is the target of repressor/activator Ume6 (Washburn and Esposito 2001), being involved in general repression via URS1 motifs or activation of early meiotic genes (Steber and Esposito 1995). Sin3 is part of at least two protein complexes, designated Rpd3L and Rpd3S, varying with respect of auxiliary proteins (Carrozza et al. 2005b). Specific repressors of other regulatory pathways such as Mig1 (glucose repression), α2 (repression of a-specific genes in α mating type) and Rox1 (repression of hypoxic genes in the presence of oxygen) utilize corepressor complex Ssn6 (=Cyc8) + Tup1 (=Flk1) to execute gene repression (Keleher et al. 1992; reviewed by Malavé and Dent 2006). Ssn6 contains 10 TPR motifs (tetratricopeptide repeat, comprising 34 amino acids; Blatch and Lässle 1999) at its N-terminus which are indispensable for repressor function and mediate protein–protein interactions. A similar function has been shown for 7 WD-40/β-transducin repeats (Smith et al. 1999) present at the C-terminus of Tup1. Importantly, Ssn6 and Tup1 recruit histone deacetylases Rpd3, Hos1 and Hos2 (Davie et al. 2003) while Tup1 also contacts histones H3 and H4 (Edmondson et al. 1996). Thus, a common mechanism of gene repression mediated by Sin3 and Ssn6/Tup1 corepressor complexes is apparent.
Previously, we have shown that expression of ICRE-dependent genes in an opi1Δ mutant is completely insensitive to high concentrations of IC (Wagner et al. 2001). In a sin3Δ mutant, regulation by IC was clearly alleviated but not entirely abolished. We thus hypothesized that a second mechanism of gene repression acting in parallel to Sin3 may be effective for ICRE-dependent genes. In this work, we show that this is indeed the case, with Ssn6/Tup1 also mediating Opi1-triggered repression. A single domain within Opi1 is able to contact both Sin3 and Ssn6. Interestingly, at least one of both corepressor complexes must be functional to guarantee cellular viability.
Materials and methods
Yeast and E. coli strains, media and growth conditions
Strains of the yeast Saccharomyces cerevisiae used in this work are listed in Table 1. Strain PJ69-4A (James et al. 1996) was used for two-hybrid assays. To test for functional complementation by variants of OPI1, opi1 deletion strain CWY1 (derived from wild-type SIRP3; Schwank et al. 1995) was used. Integrative transformation of CWY1 with expression plasmids containing OPI1 variants (see below) led to strains YDC1-YDC12. Strains with single mutations sin3Δ::kanMX and ssn6Δ::LEU2 were obtained by introduction of the respective null mutant alleles (Wagner et al. 2001) into regulatory wild-type strains isogenic to SIRP3. Yeast extracts used for protein–protein interaction assays were prepared from strain C13-ABY.S86, lacking four vacuolar proteinases.
Synthetic complete (SCD) media used for selective growth of transformants and conditions of IC repression or derepression have been described (Schüller et al. 1992). To select against URA3, synthetic complete medium supplemented with 1 mg/ml 5-fluoroorotic acid (5-FOA) was used.
Plasmid constructions
The coding region of OPI1 was mutagenized at selected positions (see below) and gene variants were fused with the natural OPI1 control region (−437/−1). Expression constructs were subsequently inserted as EcoRI/HindIII fragments into integrating plasmid YIplac128 (Gietz and Sugino 1988) to give pDC18–24, pDC40–43, pCM3–4 and pCM17–21 (LEU2 OPI1 var). Deletion variant opi1(Δ2-107) was newly constructed by PCR amplification, fused with its natural promoter and similarly inserted into YIplac128. Plasmids obtained were linearized at a single ClaI site and targeted to the leu2 locus of recipient strain CWY1.
To perform two-hybrid assays, residues 1–106 of Opi1 variants were fused with the DNA-binding domain of Gal4. Thus, specific PCR primers were used to amplify the corresponding region of OPI1, introducing restriction sites EcoRI and BamHI. After activation of these sites, fragments were inserted into plasmid pGBD-C1 (James et al. 1996) to give various Gal4DBD–Opi1(1–106) fusions (pDC29–35, pDC44–47). Plasmid pJW1 [Gal4AD-Sin3(1–300)] has been previously described (Wagner et al. 2001).
To test for synthetic lethality of mutations sin3 and ssn6, single copy plasmids pKH20 (ARS CEN URA3 MET25-SIN3; contains the 4.4 kb SIN3 gene as a BamHI/XhoI fragment) and pYJ37 (ARS CEN URA3 MET25-SSN6; contains SSN6 as a 2.9 kb BamHI/EcoRI fragment) were constructed by insertion of PCR-amplified fragments into expression plasmid p416-MET25 (Mumberg et al. 1994).
E. coli expression plasmids pCW86 [GST–Opi1(1–106)] and pCW136 (tetp/o HA3-OPI1) have been described (Wagner et al. 2001). Bacterial expression of SSN6 length variants representing various TPR motifs was obtained by PCR amplification and subsequent insertion of BamHI/EcoRI fragments into plasmid pGEX-2TK (GE Healthcare) to give GST–Ssn6var plasmids (pJuS5, 11, 13, 14, 15, 16 and 20, pYJ95, 99 and 107).
For yeast expression of Ssn6 length variants, plasmid p426-MET25HA (Mumberg et al. 1994) was used. DNA representing residues 1–398 (comprising TPR motifs) and 399–966 was amplified by PCR (1.2 kb BamHI/EcoRI fragment and 1.7 kb BamHI/EcoRI fragment, respectively) and inserted into the vector cleaved correspondingly to give plasmids pYJ65 [MET25-HA3-SSN6(1–398)] and pYJ64 [MET25-HA3-SSN6(399–966)]. The same vector was used for epitope-tagging of Opi1 missense variants L56A, V59A and V67A. The coding region of OPI1 was cut with BamHI/HindIII from pDC18 (V59A), pDC20 (L56A) and pDC21 (V67A) and inserted into p426-MET25HA to give pJuS7, pYJ108 and pYJ109, respectively. Expression plasmids pCM35-pCM40 containing epitope-tagged Opi1 variants mutated within the Ino2 interaction domain (L333A, L340A, V343A, V350A, L354A and V361A) were constructed correspondingly.
Site-directed mutagenesis
To alter selected residues in the coding region of OPI1, the QuikChange site-directed mutagenesis kit of Stratagene was used. To obtain mutations within OSID and AID, plasmid pWTH99 containing the OPI1 coding region was used (Heyken et al. 2005). To replace selected residues against alanine, we used pairs of mutagenic primers introducing a GCG codon instead of the natural codon, flanked by 15–19 nucleotides on both the sides. DNA sequencing was used to confirm the presence of the desired mutant alleles of opi1 (L53A, L56A, D57A, R58A, V59A, I63A, V67A, F70A, Y71A, L81A, L88A, L333A, L340A, L343A, M347A, V350A, L354A and V361A) and the absence of any other change in the plasmids obtained (pDC11–17, pDC36–39, pCM1–2, pCM12–16).
In vitro-interaction assays (GST pull-down)
To perform interaction assays by affinity chromatography, heterologous expression of GST- and HA-tagged proteins was performed in E. coli strain BL21 (Stratagene/Agilent). Gene expression in transformants containing GST fusions regulated by the tac promoter was induced with 1 mM IPTG. To produce HA-Opi1 in E. coli, tet promoter-dependent expression of pCW136 was activated by 0.2 mg/l anhydrotetracycline. For maximal derepression of MET25-dependent HA fusion gene variants, yeast transformants were cultivated in a selective medium in the absence of methionine.
Following bacterial biosynthesis, GST fusion proteins containing length variants of either Opi1 or Ssn6 were immobilized on glutathione (GSH) Sepharose (GE Healthcare). The procedure of affinity chromatography and subsequent washing steps at intermediary stringency has been described (Wagner et al. 2001). GST fusions and bound HA-tagged proteins were finally eluted with free GSH (10 mM) and subsequently separated by SDS/PAGE. Following transfer to filters, HA fusion proteins were incubated with anti-HA-peroxidase conjugate and detected using POD chemiluminescent substrate (both supplied by Roche Biochemicals).
Miscellaneous procedures
Transformation of S. cerevisiae strains, immunoblot analysis, PCR amplification and β-galactosidase assays was performed as previously described (Schwank et al. 1995; Wagner et al. 2001).
Results
Mutational analysis of Opi1 domain interacting with Sin3
We have previously shown that the N-terminus of Opi1 (residues 1–106) is able to interact with PAH1 of Sin3 in vivo and in vitro (Wagner et al. 2001). Within this domain (designated OSID), an amphipathic pattern of hydrophobic amino acids could be identified (residues 53–88). To investigate the possible importance of these residues for regulated expression of Opi1 target genes, we performed a site-directed mutagenesis at the selected positions, leading to replacement of hydrophobic amino acids to alanine (L53, L56, V59, I63, V67, F70, L81 and L88). As a control, the importance of two charged side chains (D57 and R58) and a hydrophilic amino acid (Y71) was also investigated.
To assay for interaction of Opi1 wild-type and missense variants with Sin3 in vivo, we fused Opi1 residues 1–106 with the DNA-binding domain of Gal4. The resulting plasmids were co-transformed into strain PJ69-4A with a plasmid encoding Sin3 residues 1–300 (comprising PAH1) fused behind Gal4 activation domain. In the case of Opi1-Sin3 interaction, the resulting hybrid constructs should be able to activate the Gal4-dependent reporter gene GAL7-lacZ of strain PJ69-4A. As is shown in Table 2, Opi1 variants L56A, V59A and V67A were defective for wild-type activation of the reporter gene.
We next investigated the consequences of defective interaction of Opi1 and Sin3 for repression of an Opi1-dependent reporter gene. Full-length OPI1 wild-type and missense mutants were transferred into an integrating yeast vector (YIplac128). The resulting plasmids (LEU2 OPI1 variant) were then transformed into strain CWY1 (leu2 opi1Δ::HIS3), containing an integrated ICRE-CYC1-lacZ reporter gene. In agreement with results of the two-hydrid analysis, Opi1 variants L56A, V59A and V67A were unable to restore IC-dependent repression of the reporter gene (Table 3). Opi1 variants which were functional for interaction with Sin3 could also complement the opi1 null mutation of recipient strain CWY1.
In a previous work, we characterized Opi1 length variants for functional complementation of an opi1 null mutation (Wagner et al. 1999) and described that the opi1(Δ2-107) allele lacking OSID could partially restore gene repression. Since this result contradicts the findings with Opi1 missense variants reported above, we re-constructed the deletion allele and repeated the assay for reporter gene expression with transformants obtained. As is also apparent from Table 3, our assays show that loss of repressor function was observed following integration of the new opi1(Δ2-107) allele. The reason for the discrepancy of previous data and results reported in this work remains unclear. Nevertheless, our new findings show that Opi1 lacking residues 2–107 do no longer support repression. This result agrees with the phenotypes of OSID missense mutants.
Synthetic lethality of sin3 and ssn6 null mutants
The phenotype of missense mutants opi1L56A, opi1V59A and opi1V67A mapping to OSID shows a complete loss of function for the respective Opi1 variants. It appears reasonable to assume that a sin3 null mutant should phenocopy this defect. However, this assumption is not entirely supported by the previous finding of severely reduced but, nevertheless, residual gene regulation observed with a sin3 null mutant (Wagner et al. 2001). We thus hypothesized that Opi1 OSID may recruit not only Sin3 but a second mediator of repression as well. Possible candidates are Ssn6 and Tup1 which together affect a large number of target genes repressed by various stimuli (Malavé and Dent 2006). Indeed, previous testing of ICRE-dependent gene regulation in ssn6 and tup1 null mutants also showed an increased gene expression under repressing conditions (Wagner et al. 2001). To further confirm the importance of both corepressor complexes Sin3 and Ssn6/Tup1 for repression of ICRE-dependent genes, we wished to construct a double mutant sin3 ssn6. However, we were unable to obtain this double mutant by introduction of a ssn6Δ::LEU2 disruption cassette into a sin3 deletion strain. Similarly, no meiotic products of the desired genotype were obtained after crossing single mutants sin3 and ssn6 and subsequent sporulation. Thus, mutations sin3 and ssn6 may be synthetically lethal. To confirm this, we constructed plasmids containing markers ARS CEN URA3 SIN3 (pKH20) and ARS CEN URA3 SSN6 (pYJ37) which were subsequently transformed into strain JS05.4 (ura3/ura3 sin3/SIN3 ssn6/SSN6). Following sporulation, we selected for haploid progeny with sin3 ssn6 double mutations, carrying plasmid copies of either SIN3 or SSN6. Similarly, ssn6 and sin3 single mutants were transformed with SSN6 or SIN3 plasmids. While transformants ssn6 + SSN6 and sin3 + SIN3 were still viable after selecting against URA3 plasmids using 5-fluoroorotic acid (5-FOA), no growth was observed after plasmid curing of the double mutant (Fig. 1). Thus, at least one corepressor complex Sin3 or Ssn6 + Tup1 must be functional to support viability of the yeast cell. We conclude that mutations sin3 and ssn6 are indeed synthetically lethal. The same result was obtained for mutations sin3 and tup1 (not shown).
Synthetic lethality of mutations sin3 and ssn6. Serial dilutions of regulatory wild-type strain JS91.15-23 and strains with single or double mutations ssn6 and sin3 were transferred to YEPD, SCD-Ura and SCD + 5-FOA media. The following transformants were analyzed: JS91.15-23 + p416-MET25 (wild-type + empty vector), JS05.5-2B + pYJ37 (ssn6Δ + SSN6 plasmid), JS05.1-3 + pKH20 (sin3Δ + SIN3 plasmid), JS05.4-3C + pYJ37 (ssn6Δ sin3Δ + SSN6 plasmid) and JS05.7-4A + pKH20 (ssn6Δ sin3Δ + SIN3 plasmid). Relevant genotypes and times of incubation (d, days) are indicated
OSID interacts with TPR motifs of Ssn6
To confirm our assumption of Ssn6 being important for Opi1-mediated gene repression, we assayed for in vitro-interaction of GST–Opi1 (1–106) with HA-tagged Ssn6. It has been previously shown that TPR motifs of Ssn6 are responsible for interaction with various repressor proteins (Tzamarias and Struhl 1995; Smith et al. 1995). We thus separately expressed epitope-tagged Ssn6 residues 1–398 (comprising its TPR motifs) and 399–966 in yeast. Although both fragments of Ssn6 could be stably detected in protein extracts (Fig. 2a, lanes 1, 2), only the TPR-containing fragment was able to interact with GST–Opi1 (Fig. 2a, lane 4). No evidence for direct binding of HA-tagged Tup1 to GST–Opi1 could be obtained (not shown).
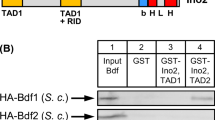
In vitro interaction of Opi1 and Ssn6. a Input control and assay for Opi1-binding of Ssn6 fragments 1–398 and 399–966 (plasmids pYJ65 and pYJ64). GST–Opi1(1–106) was synthesized in E. coli (BL21 transformed with plasmid pCW86), immobilized on GSH Sepharose and incubated with yeast total protein extract. b Length variants of Ssn6 comprising various combinations of TPR motifs were fused with GST and incubated with bacterially synthesized HA-Opi1 (plasmid pCW136). c Comparative analysis of interaction of GST–Ssn6(79–175) comprising TPR motifs 2–3 (pYJ95) with HA-tagged Opi1 wild-type and missense variants (full-length) expressed in S. cerevisiae (plasmids pCW116, pYJ108, pJuS7 and pYJ109). Extracts containing 75 μg of total protein were analyzed for the input control. To achieve comparable amounts of HA-Opi1 variants for the interaction assay, total protein was adjusted accordingly
We next asked whether a combination of specific TPR motifs or a minimal number of them is required for Opi1 OSID to bind. For reasons of protein stability, Ssn6 TPR length variants were fused with GST and subsequently incubated with HA-Opi1. As is shown in Fig. 2b, Ssn6 fragments containing non-overlapping TPR motifs were able to bind Opi1 with comparable efficiency. Similarly, the number of motifs contained in the fusion constructs did not influence binding efficiency. Finally, we arbitrarily selected individual TPR motifs 1, 5 and 8 for fusion with GST and were able to show that each motif was sufficient for interaction with Opi1.
As shown above, Opi1 variants L56A, V59A and V67A led to IC-insensitive expression of the ICRE-dependent reporter gene. We thus compared binding of Opi1 wild-type and missense variants to TPR motifs 2–3 of Ssn6. As is shown in Fig. 2c, Opi1 full-length mutant proteins could be expressed in S. cerevisiae but were unable to interact with bacterially synthesized GST–Ssn6(TPR2–3), explaining the loss of repression observed with Opi1 mutants L56A, V59A and V67A.
Mutational analysis of Opi1–Ino2 interaction
We have previously shown that carboxy-terminal residues of Opi1, designated AID (amino acids 321–380), are necessary and sufficient for binding to Ino2 (Heyken et al. 2005). A closer inspection of AID revealed the existence of a periodic pattern of hydrophobic amino acids, reminiscent of a leucine zipper (L-N6-L-N6-M-N6-L-N6-V). It should be emphasized that this motif is distinct from the genuine leucine zipper located at the core of Opi1 (residues 139–160). Similar to our analysis of OSID, we replaced all hydrophobic residues within AID compatible with an amphipathic α-helix (L333, L340, V343, M347, V350, L354 and V361) by alanine and integrated the OPI1 variants obtained into the chromosomal DNA of an opi1 null mutant. Stability of HA-tagged variants was confirmed by immunoblot analysis (cf. Fig. 3). As is apparent from Table 4, all Opi1 variants investigated were unable to trigger repression of the reporter gene in the presence of IC (exception: missense mutation M347A allowed a substantial repression but did not completely restore wild-type regulation). These findings support the hypothesis of a leucine-zipper-related structural motif as the critical core of Opi1 repressor required for deactivating Ino2.
Control of stable expression of Opi1 variants containing missense mutations within the Ino2 activator interaction domain (AID). Plasmids pCW116 (wild-type) and pCM35–pCM40 were transformed into strain C13-ABY.S86. For Western blot analysis with anti-HA antibodies, extracts containing 75 μg of total protein were used
Discussion
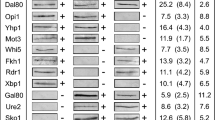
Previous analyses have shown that pathway-specific repressor proteins require interaction with corepressors to finally fulfil their function. In yeast, corepressors Sin3 and Ssn6/Tup1 can be recruited to pleiotropically downregulate a large number of unrelated structural genes (Green and Johnson 2004; Grzenda et al. 2009; Malavé and Dent 2006). Genetic evidence for a role in repression and demonstration of physical interaction with Sin3 has been shown for repressors Ume6 (Kadosh and Struhl 1997), Ash1 (Carrozza et al. 2005a), Whi5, Stb1 (Takahata et al. 2009), Fkh1, Fkh2 (Ho et al. 2002) while Ssn6/Tup1 directly mediate repression by α2, Crt1, Mig1, Rox1 (Malavé and Dent 2006), Nrg1 (Berkey et al. 2004), Sfl1 (Conlan and Tzamarias 2001), Sko1 (Proft and Struhl 2002), Rgt1 (Polish et al. 2005), Hmo1 (Krogan et al. 2006) and Cup9 (Xia et al. 2008). Both corepressors may also interact with activators Pho4, Hac1 (recruiting Sin3; Graumann et al. 2004; Schröder et al. 2004), Hap1 and Aft1 (recruiting Ssn6/Tup1; Zhang and Guarente 1994; Fragiadakis et al. 2004). Consequently, a positive influence on transcription for yeast corepressor complexes Sin3 and Ssn6/Tup1 has been described as well (Vidal et al. 1991; Proft and Struhl 2002).
In this work, we have shown that a single domain of yeast repressor Opi1 (OSID) is able to interact with both corepressor complexes Sin3 and Ssn6/Tup1. Mutational analysis of OSID led to the identification of Opi1 variants L56A, V59A and V67A which could be stably expressed in S. cerevisiae but were defective for interaction with Sin3 and entirely insensitive to gene repression by IC. In contrast, a sin3 deletion mutant was strongly, but not completely deregulated (Wagner et al. 2001). We thus concluded that OSID may be involved in a second pathway of repression acting in parallel to Sin3. Indeed, we could show that OSID not only binds to PAH1 of Sin3 but also interacts with TPR motifs of corepressor Ssn6. No evidence for interaction of Opi1 with Tup1 could be obtained. The functional importance of Opi1–Ssn6 interaction was further supported by our observation that Opi1 variants L56A, V59A and V67A were defective for in vitro interaction with Ssn6. This finding explains why target gene repression in the presence of these variants was completely abolished. Interaction of two corepressor complexes with a single specific repressor may be the exception but not a general rule, at least in yeast. Besides Opi1, recruitment of Sin3 and Ssn6/Tup1 has been described only for Cti6 involved in regulation of metal acquisition by yeast (Papamichos-Chronakis et al. 2002; Puig et al. 2004). While Ssn6 binds to the C-terminus of Cti6, no mapping of its Sin3 binding site has been performed. Thus, it is unclear whether a single domain of Cti6 mediates interaction with both corepressors.
To further confirm the contribution of both corepressors, we wished to assay ICRE-dependent gene expression in a sin3 ssn6 double mutant. However, we were unsuccessful to construct such a mutant and were able to show synthetic lethality of both mutations. The same is true for sin3 and tup1 mutations. These results are not entirely surprising, considering the substantially reduced growth rate of each single mutant. Our finding is in agreement with synthetic lethality of ssn6 and rpd3 described in a study on genetic interactions among factors of histone modification (Lin et al. 2008).
Mapping of the Opi1 binding domain within Ssn6 using GST pull down assays clearly allowed us to demonstrate OSID-TPR interaction which agrees with results previously obtained for repressors such as α2 and Mig1. However, two-hybrid fusion constructs containing Gal4DBD–Opi11–106 and Gal4TAD–Ssn6TPR1–10 could not activate a GAL7-lacZ reporter gene (data not shown). Presumably, the presence of highly potent TPR repression domains which are autonomously able to recruit histone deacetylases (Watson et al. 2000) counteracts gene activation as the working principle of the two-hybrid approach. Mapping studies in vitro using combinations of several TPR motifs and even single TPR motifs revealed a surprising degree of functional redundancy among them. Although a strong conservation at the sequence level is apparently absent, single TPR motifs 1, 5 and 8 chosen arbitrarily could interact with OSID of Opi1. These findings agree with the results previously described for interaction of α2 repressor and TPR6 of Ssn6 (Smith and Johnson 2000), based on identical methods. On the other hand, a combination of defined TPR motifs required for repression of specific genes has been also described (Tzamarias and Struhl 1995). For α2-Ssn6 interaction, the tripeptide motif SRI reminiscent of the peroxisomal targeting sequence PTS1 has been described as important (Smith and Johnson 2000). OSID does not contain a PTS1 consensus motif but the sequence SNV (residues 65–67) is present. Although V67 is indispensable for Opi1–Ssn6 interaction and target gene repression, it is questionable whether SNV may be considered as PTS1-like because asparagine is biochemically too distant from genuine basic amino acids.
Binding of OSID to PAH and TPR motifs suggest that both interaction modules should be able to contact related targets. Solution structures have been determined for PAH1 and PAH2 of mammalian Sin3, both of which generate an amphipathic four-helix-bundle with an up-and-down topology (Brubaker et al. 2000; also designated wedged helical bundle, Spronk et al. 2000). PAH motifs form a hydrophobic pocket which provides a suitable binding surface for a helical partner protein. According to their structural analysis and modelling of NRSF-mSin3B interaction, Sahu et al. (2008) proposed a consensus sequence for binding to PAH1 (ΦXΦΦSXΦS; Φ, bulky hydrophobic residue; S, residue with short side-chain; X, any residue except proline). Within OSID of Opi1, two sequence motifs correspond to this consensus (residues 53–60: LNILDRVS and residues 68–75: VTFYDEIN). Interestingly, the mutant phenotype found for Opi1 variants L56A and V59A nicely fits to this proposition (V67 maps outside the consensus). Mammalian NRSF (neural-restrictive silencer factor) contains identical residues at corresponding positions which are involved in hydrophobic interactions with α-helical segments of PAH1 (L49 and V52 of NRSF with F93 and F58 of mSin3B, respectively). The second motif within OSID may be less important since variants F70A and Y71A showed unimpaired interaction with Sin3.
The 34 amino acids of a TPR motif form a pair of antiparallel α-helices with limited sequence conservation (reviewed by D’Andrea and Regan 2003). Using structural data from TPR motifs of protein phosphatase 5, Gounalaki et al. (2000) presented a model of Ssn6-Tup1 interaction, emphasizing hydrophobic interactions. We thus conclude that interaction modules PAH and TPR both employing helical segments with hydrophobic-apolar surfaces should exhibit sufficient structural flexibility to allow binding of the same partner domain (OSID of Opi1; summarized in Fig. 4).
Summary of regulatory interactions effective under conditions of inositol/choline repression. AID activator interaction domain; bZIP basic leucine zipper; DBD DNA-binding domain; FFAT phenylalanines in acidic tract; HDAC histone deacetylase; HID HDAC interaction domain; ICRE inositol/choline-responsive element; OSID Sin3/Ssn6-Opi1 interaction domain; P1–P4 paired amphipathic helices; RID repressor interaction domain; TAD transcriptional activation domain; TPR tetratricopeptide repeat; WD40 tryptophan/aspartate-containing repeat. Protein–protein interactions are depicted by arrows
We finally performed a mutational analysis of the Opi1 AID, required to silence Ino2. We have previously characterized mutant alleles altered at the position of charged amino acids (double mutants K337 E338 and R372 E373; Heyken et al. 2005) which are strongly conserved among Opi1 proteins from various yeasts. Although these Opi1 variants showed 3–4-fold increase of ICRE-dependent gene expression under repressing conditions, a significant degree of regulation remained. In this work, we describe the regulatory phenotype of variants altered at the position of a sequence motif with similarity to a leucine zipper (residues 333–361: L-N6-L-N2-V-N3-M-N2-V-N3-L-N6-V). Among seven mutant proteins each containing a single alanine mutation at the positions indicated six turned out as completely deregulated. This result is in agreement with the phenotype of an opi1 allele encoding a V343Q mutation (rum1-25; Kaadige and Lopes 2006), isolated by a strategy selecting for loss of Opi1 function. We thus conclude that the hydrophobic-amphipathic motif is indeed the functional core of Opi1 AID which binds to the repressor interaction domain (RID) of Ino2, counteracting target gene activation. Interactions among regulators specific for phospholipid biosynthesis and pleiotropic factors as characterized in this and previous work are summarized in Fig. 4.
References
Ambroziak J, Henry SA (1994) INO2 and INO4 gene products, positive regulators of phospholipid biosynthesis in Saccharomyces cerevisiae, form a complex that binds to the INO1 promoter. J Biol Chem 269:15344–15349
Berkey CD, Vyas VK, Carlson M (2004) Nrg1 and Nrg2 transcriptional repressors are differently regulated in response to carbon source. Eukaryot Cell 3:311–317
Blatch GL, Lässle M (1999) The tetratricopeptide repeat: a structural motif mediating protein–protein interactions. Bioessays 21:932–939
Brubaker K, Cowley SM, Huang K, Loo L, Yochum GS, Ayer DE, Eisenman RN, Radhakrishnan I (2000) Solution structure of the interacting domains of the Mad-Sin3 complex: implications for recruitment of a chromatin-modifying complex. Cell 103:655–665
Carrozza MJ, Florens L, Swanson SK, Shia WJ, Anderson S, Yates J, Washburn MP, Workman JL (2005a) Stable incorporation of sequence specific repressors Ash1 and Ume6 into the Rpd3L complex. Biochim Biophys Acta 1731:77–87
Carrozza MJ, Li B, Florens L, Suganuma T, Swanson SK, Lee KK, Shia WJ, Anderson S, Yates J, Washburn MP, Workman JL (2005b) Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell 123:581–592
Chen M, Hancock LC, Lopes JM (2007) Transcriptional regulation of yeast phospholipid biosynthetic genes. Biochim Biophys Acta 1771:310–321
Conlan RS, Tzamarias D (2001) Sfl1 functions via the co-repressor Ssn6-Tup1 and the cAMP-dependent protein kinase Tpk2. J Mol Biol 309:1007–1015
D’Andrea LD, Regan L (2003) TPR proteins: the versatile helix. Trends Biochem Sci 28:655–662
Davie JK, Edmondson DG, Coco CB, Dent SY (2003) Tup1-Ssn6 interacts with multiple class I histone deacetylases in vivo. J Biol Chem 278:50158–50162
Edmondson DG, Smith MM, Roth SY (1996) Repression domain of the yeast global repressor Tup1 interacts directly with histones H3 and H4. Genes Dev 10:1247–1259
Fragiadakis GS, Tzamarias D, Alexandraki D (2004) Nhp6 facilitates Aft1 binding and Ssn6 recruitment, both essential for FRE2 transcriptional activation. EMBO J 23:333–342
Gasperowicz M, Otto F (2005) Mammalian Groucho homologs: redundancy or specificity? J Cell Biochem 95:670–687
Gietz RD, Sugino A (1988) New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene 74:527–534
Gounalaki N, Tzamarias D, Vlassi M (2000) Identification of residues in the TPR domain of Ssn6 responsible for interaction with the Tup1 protein. FEBS Lett 473:37–41
Graumann J, Dunipace LA, Seol JH, McDonald WH, Yates JR 3rd, Wold BJ, Deshaies RJ (2004) Applicability of tandem affinity purification MudPIT to pathway proteomics in yeast. Mol Cell Proteomics 3:226–237
Green SR, Johnson AD (2004) Promoter-dependent roles for the Srb10 cyclin dependent kinase and the Hda1 deacetylase in Tup1-mediated repression in Saccharomyces cerevisiae. Mol Biol Cell 15:4191–4202
Grzenda A, Lomberk G, Zhang JS, Urrutia R (2009) Sin3: master scaffold and transcriptional corepressor. Biochim Biophys Acta 1789:443–450
Heyken WT, Repenning A, Kumme J, Schüller HJ (2005) Constitutive expression of yeast phospholipid biosynthetic genes by variants of Ino2 activator defective for interaction with Opi1 repressor. Mol Microbiol 56:696–707
Ho Y, Gruhler A, Heilbut A, Bader GD, Moore L, Adams SL, Millar A, Taylor P, Bennett K, Boutilier K, Yang L, Wolting C, Donaldson I, Schandorff S, Shewnarane J, Vo M, Taggart J, Goudreault M, Muskat B, Alfarano C, Dewar D, Lin Z, Michalickova K, Willems AR, Sassi H, Nielsen PA, Rasmussen KJ, Andersen JR, Johansen LE, Hansen LH, Jespersen H, Podtelejnikov A, Nielsen E, Crawford J, Poulsen V, Sorensen BD, Matthiesen J, Hendrickson RC, Gleeson F, Pawson T, Moran MF, Durocher D, Mann M, Hogue CW, Figeys D, Tyers M (2002) Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature 415:180–183
Hudak KA, Lopes JM, Henry SA (1994) A pleiotropic phospholipid biosynthetic regulatory mutation in Saccharomyces cerevisiae is allelic to sin3 (sdi1, ume4, rpd1). Genetics 136:475–483
James P, Halladay J, Craig EA (1996) Genomic libraries and a host strain designed for highly efficient two-hybrid selection in yeast. Genetics 144:1425–1436
Jones PL, Shi YB (2003) N-CoR-HDAC corepressor complexes: roles in transcriptional regulation by nuclear hormone receptors. Curr Top Microbiol Immunol 274:237–268
Kaadige MR, Lopes JM (2006) Analysis of Opi1p repressor mutants. Curr Genet 49:30–38
Kadosh D, Struhl K (1997) Repression by Ume6 involves recruitment of a complex containing Sin3 corepressor and Rpd3 histone deacetylase to target promoters. Cell 89:365–371
Keleher CA, Redd MJ, Schultz J, Carlson M, Johnson AD (1992) Ssn6-Tup1 is a general repressor of transcription in yeast. Cell 68:709–719
Krogan NJ, Cagney G, Yu H, Zhong G, Guo X, Ignatchenko A, Li J, Pu S, Datta N, Tikuisis AP, Punna T, Peregrin-Alvarez JM, Shales M, Zhang X, Davey M, Robinson MD, Paccanaro A, Bray JE, Sheung A, Beattie B, Richards DP, Canadien V, Lalev A, Mena F, Wong P, Starostine A, Canete MM, Vlasblom J, Wu S, Orsi C, Collins SR, Chandran S, Haw R, Rilstone JJ, Gandi K, Thompson NJ, Musso G, St Onge P, Ghanny S, Lam MH, Butland G, Altaf-Ul AM, Kanaya S, Shilatifard A, O’Shea E, Weissman JS, Ingles CJ, Hughes TR, Parkinson J, Gerstein M, Wodak SJ, Emili A, Greenblatt JF (2006) Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 440:637–643
Lin YY, Qi Y, Lu JY, Pan X, Yuan DS, Zhao Y, Bader JS, Boeke JD (2008) A comprehensive synthetic genetic interaction network governing yeast histone acetylation and deacetylation. Genes Dev 22:2062–2074
Liu Z, Karmarkar V (2008) Groucho/Tup1 family co-repressors in plant development. Trends Plant Sci 13:137–144
Loewen CJ, Gaspar ML, Jesch SA, Delon C, Ktistakis NT, Henry SA, Levine TP (2004) Phospholipid metabolism regulated by a transcription factor sensing phosphatidic acid. Science 304:1644–1647
Malavé TM, Dent SY (2006) Transcriptional repression by Tup1-Ssn6. Biochem Cell Biol 84:437–443
McDonel P, Costello I, Hendrich B (2009) Keeping things quiet: roles of NuRD and Sin3 co-repressor complexes during mammalian development. Int J Biochem Cell Biol 41:108–116
Mumberg D, Müller R, Funk M (1994) Regulatable promoters of Saccharomyces cerevisiae: comparison of transcriptional activity and their use for heterologous expression. Nucleic Acids Res 22:5767–5768
Papamichos-Chronakis M, Petrakis T, Ktistaki E, Topalidou I, Tzamarias D (2002) Cti6, a PHD domain protein, bridges the Cyc8-Tup1 corepressor and the SAGA coactivator to overcome repression at GAL1. Mol Cell 9:1297–1305
Polish JA, Kim JH, Johnston M (2005) How the Rgt1 transcription factor of Saccharomyces cerevisiae is regulated by glucose. Genetics 169:583–594
Proft M, Struhl K (2002) Hog1 kinase converts the Sko1-Cyc8-Tup1 repressor complex into an activator that recruits SAGA and SWI/SNF in response to osmotic stress. Mol Cell 9:1307–1317
Puig S, Lau M, Thiele DJ (2004) Cti6 is an Rpd3-Sin3 histone deacetylase associated protein required for growth under iron-limiting conditions in Saccharomyces cerevisiae. J Biol Chem 279:30298–30306
Sahu SC, Swanson KA, Kang RS, Huang K, Brubaker K, Ratcliff K, Radhakrishnan I (2008) Conserved themes in target recognition by the PAH1 and PAH2 domains of the Sin3 transcriptional corepressor. J Mol Biol 375:1444–1456
Schröder M, Clark R, Liu CY, Kaufman RJ (2004) The unfolded protein response represses differentiation through the RPD3-SIN3 histone deacetylase. EMBO J 23:2281–2292
Schüller HJ, Hahn A, Tröster F, Schütz A, Schweizer E (1992) Coordinate genetic control of yeast fatty acid synthase genes FAS1 and FAS2 by an upstream activation site common to genes involved in membrane lipid biosynthesis. EMBO J 11:107–114
Schwank S, Ebbert R, Rautenstrauss K, Schweizer E, Schüller HJ (1995) Yeast transcriptional activator INO2 interacts as an Ino2p/Ino4p basic helix-loop-helix heteromeric complex with the inositol/choline-responsive element necessary for expression of phospholipid biosynthetic genes in Saccharomyces cerevisiae. Nucleic Acids Res 23:230–237
Slekar KH, Henry SA (1995) SIN3 works through two different promoter elements to regulate INO1 gene expression in yeast. Nucleic Acids Res 23:1964–1969
Smith RL, Johnson AD (2000) A sequence resembling a peroxisomal targeting sequence directs the interaction between the tetratricopeptide repeats of Ssn6 and the homeodomain of α2. Proc Natl Acad Sci USA 97:3901–3906
Smith RL, Redd MJ, Johnson AD (1995) The tetratricopeptide repeats of Ssn6 interact with the homeo domain of α2. Genes Dev 9:2903–2910
Smith TF, Gaitatzes C, Saxena K, Neer EJ (1999) The WD repeat: a common architecture for diverse functions. Trends Biochem Sci 24:181–185
Spronk CA, Tessari M, Kaan AM, Jansen JF, Vermeulen M, Stunnenberg HG, Vuister GW (2000) The Mad1-Sin3B interaction involves a novel helical fold. Nat Struct Biol 7:1100–1104
Steber CM, Esposito RE (1995) UME6 is a central component of a developmental regulatory switch controlling meiosis-specific gene expression. Proc Natl Acad Sci USA 92:12490–12494
Sternberg PW, Stern MJ, Clark I, Herskowitz I (1987) Activation of the yeast HO gene by release from multiple negative controls. Cell 48:567–577
Strich R, Slater MR, Esposito RE (1989) Identification of negative regulatory genes that govern the expression of early meiotic genes in yeast. Proc Natl Acad Sci USA 86:10018–10022
Takahata S, Yu Y, Stillman DJ (2009) The E2F functional analogue SBF recruits the Rpd3(L) HDAC, via Whi5 and Stb1, and the FACT chromatin reorganizer, to yeast G1 cyclin promoters. EMBO J 28:3378–3389
Tzamarias D, Struhl K (1995) Distinct TPR motifs of Cyc8 are involved in recruiting the Cyc8-Tup1 corepressor complex to differentially regulated promoters. Genes Dev 9:821–831
Vannier D, Balderes D, Shore D (1996) Evidence that the transcriptional regulators SIN3 and RPD3, and a novel gene (SDS3) with similar functions, are involved in transcriptional silencing in S. cerevisiae. Genetics 144:1343–1353
Vidal M, Strich R, Esposito RE, Gaber RF (1991) RPD1 (SIN3/UME4) is required for maximal activation and repression of diverse yeast genes. Mol Cell Biol 11:6306–6316
Wagner C, Blank M, Strohmann B, Schüller HJ (1999) Overproduction of the Opi1 repressor inhibits transcriptional activation of structural genes required for phospholipid biosynthesis in the yeast Saccharomyces cerevisiae. Yeast 15:843–854
Wagner C, Dietz M, Wittmann J, Albrecht A, Schüller HJ (2001) The negative regulator Opi1 of phospholipid biosynthesis in yeast contacts the pleiotropic repressor Sin3 and the transcriptional activator Ino2. Mol Microbiol 41:155–166
Wang H, Stillman DJ (1993) Transcriptional repression in Saccharomyces cerevisiae by a SIN3-LexA fusion protein. Mol Cell Biol 13:1805–1814
Wang H, Clark I, Nicholson PR, Herskowitz I, Stillman DJ (1990) The Saccharomyces cerevisiae SIN3 gene, a negative regulator of HO, contains four paired amphipathic helix motifs. Mol Cell Biol 10:5927–5936
Washburn BK, Esposito RE (2001) Identification of the Sin3-binding site in Ume6 defines a two-step process for conversion of Ume6 from a transcriptional repressor to an activator in yeast. Mol Cell Biol 21:2057–2069
Watson AD, Edmondson DG, Bone JR, Mukai Y, Yu Y, Du W, Stillman DJ, Roth SY (2000) Ssn6-Tup1 interacts with class I histone deacetylases required for repression. Genes Dev 14:2737–2744
Xia Z, Turner GC, Hwang CS, Byrd C, Varshavsky A (2008) Amino acids induce peptide uptake via accelerated degradation of CUP9, the transcriptional repressor of the PTR2 peptide transporter. J Biol Chem 283:28958–28968
Yoshimoto H, Ohmae M, Yamashita I (1992) The Saccharomyces cerevisiae GAM2/SIN3 protein plays a role in both activation and repression of transcription. Mol Gen Genet 233:327–330
Zhang L, Guarente L (1994) Evidence that TUP1/SSN6 has a positive effect on the activity of the yeast activator HAP1. Genetics 136:813–817
Acknowledgments
This work has been supported by the Deutsche Forschungsgemeinschaft (DFG). We thank Felix Kliewe, Gudrun Ebel and Karola Hahn for valuable support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Collart.
Rights and permissions
About this article
Cite this article
Jäschke, Y., Schwarz, J., Clausnitzer, D. et al. Pleiotropic corepressors Sin3 and Ssn6 interact with repressor Opi1 and negatively regulate transcription of genes required for phospholipid biosynthesis in the yeast Saccharomyces cerevisiae . Mol Genet Genomics 285, 91–100 (2011). https://doi.org/10.1007/s00438-010-0589-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-010-0589-5