Abstract
In eukaryotes, the precise transcriptional and post-transcriptional regulations of gene expression are crucial for the developmental processes. More than 100 types of post-transcriptional RNA modifications have been identified in eukaryotes. The deposition of N6-methyladenosine (m6A) into mRNA is among the most common post-transcriptional RNA modifications known in eukaryotes. It has been reported that m6A RNA modification can regulate gene expression. The role of yeast m6A methyltransferase (Ime4) in meiosis and sporulation in diploid cells is very well proven, but its physiological role in haploid cells has remained unknown until recently. Previously, we have shown that Ime4 epitranscriptionally regulates triacylglycerol (TAG) metabolism and vacuolar morphology in haploid cells. Mitochondrial dysfunction leads to TAG accumulation as lipid droplets (LDs) in the cells; besides, LDs are physically connected to the mitochondria. As of now there are no reports on the role of Ime4 in mitochondrial biology. Here we report the important role played by Ime4 in the mitochondrial morphology and functions in Saccharomyces cerevisiae. The confocal microscopic analysis showed that IME4 gene deletion causes mitochondrial fragmentation; besides, the ime4Δ cells showed a significant decrease in cytochrome c oxidase and citrate synthase activities compared to the wild-type cells. IME4 gene deletion causes mitochondrial dysfunction, and it will be interesting to find out the target genes of Ime4 related to the mitochondrial biology. The determination of the role of Ime4 and its targets in mitochondrial biology could probably help in formulating potential cures for the mitochondria-linked rare genetic disorders.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In eukaryotes, the precise and controlled regulation of gene expression is crucial for the developmental processes. The post-transcriptional regulation that acts at the translational level is complementary to transcriptional regulation. The RNA-binding proteins play a key role in post-transcriptional regulation by controlling the mRNA levels (Jin and Neiman 2016; Berchowitz et al. 2013). Not only specific regulatory mechanisms but also cellular growth govern gene expression because both synthesis and degradation simultaneously determine mRNA levels (Chavez et al. 2016). The cellular growth depends on different metabolites. In eukaryotes, specific metabolites play key regulatory roles and thus directly connect metabolic status and cellular functions (Saint-Marc et al. 2015). S-Adenosylmethionine (SAM), an important metabolite, serves as the sole methyl donor for the methylation of histones, nucleic acids, and phospholipids (Ding et al. 2015). In addition, SAM is physiologically linked with lipid accumulation. In the previous study, we have shown that yeast m6A methyltransferases Ime4 epitranscriptionally regulates TAG metabolism and vacuolar morphology in haploid cells (Yadav and Rajasekharan 2017). Recently, studies have shown that mitochondrial dysfunction causes TAG accumulation and formation of LDs in the cells (Singh et al. 2016; Lee et al. 2013). The LDs play an important role in lipid metabolism and energy homeostasis through their interaction with mitochondria (Pu et al. 2011). The most obvious question that arises from these findings is, does TAG metabolism and mitochondria are physiologically linked via methylation? Therefore, in the present study, we studied the role of m6A methyltransferases Ime4 in the mitochondrial functions. Our data showed that IME4 gene deletion produces mitochondrial dysfunction. It will be interesting to find out the target genes of Ime4 which are directly involved in the mitochondrial biogenesis, morphology, and functions.
The m6A methyltransferases and their physiological significance
In eukaryotes, post-synthetic modifications of proteins, DNA, and RNA are common features. Unlike protein and DNA modifications, RNA modifications are not well studied (Schwartz et al. 2013). There are more than 100 types of post-synthetic RNA modifications known, yet our knowledge about their function and physiological significance is limited (Blanco and Frye 2014). The recent development of transcriptome-wide methods to identify RNA modifications such as 5-methylcytidine (m5C) and N6-methyladenosine (m6A) has created a new research field, the ‘epitranscriptome’. It became evident that these post-transcriptional RNA modifications regulate different fundamental cellular processes (Blanco and Frye 2014). The deposition of m6A into mRNA is among the most common post-transcriptional RNA modifications known in eukaryotes (Schwartz et al. 2013; Yue et al. 2015). N6-Adenosyl methyltransferases that introduce a tightly controlled deposition m6A into mRNA are found in almost all kingdoms of eukaryotic life (Yue et al. 2015; Dominissini et al. 2012). Like DNA methylation, it has been reported that m6A RNA modification can regulate gene expression (Fu et al. 2014; Zheng et al. 2013). Role of m6A RNA modification has also been suggested in other key functions such as RNA splicing (Jia et al. 2011), mRNA degradation (Harigaya et al. 2006), RNA stability (Zhang et al. 2010; Brennan and Steitz 2001), translational control (Tuck et al. 1999), meiosis (Schwartz et al. 2013), and cellular differentiation (Zhong et al. 2008).
The m6A methyltransferase in Saccharomyces cerevisiae
Since the m6A methyltransferases are fundamentally conserved throughout eukaryotes, S. cerevisiae is used as a model system to understand the biological relevance of m6A methylation. In S. cerevisiae, Ime4 is a counterpart of mammalian m6A methyltransferase (METTL3). The IME4 gene locus is transcribed into IME4 (sense RNA) and RME2 (the antisense RNA) transcripts depending on the cell type. The role of m6A methyltransferase (Ime4) in meiosis and sporulation in diploid cells is very well studied (Agarwala et al. 2012; Clancy et al. 2002), but its physiological role in haploid cells has remained unknown until recently. In the previous study (Yadav and Rajasekharan 2017), we have shown that Ime4 epitranscriptionally regulates triacylglycerol (TAG) metabolism and vacuolar morphology in haploid cells, independently of the MIS complex (RNA methylation complex consists of the Ime4, Mum2, and Slz1 proteins). The RNA modification-mediated regulation of genes is termed as the epitranscriptional regulation.
A few studies have demonstrated that yeast Ime4 has stationary phase-related functions when glucose is exhausted (Yadav and Rajasekharan 2017; Hongay et al. 2006). In yeast, TAG synthesis is a characteristic feature of the stationary growth phase (Horvath et al. 2011). Recently, studies have shown that defects in mitochondria cause TAG accumulation and formation of lipid droplets (LDs) in the cells (Singh et al. 2016; Lee et al. 2013). LDs are dynamic organelles and consist of a core of TAG and steryl esters surrounded by a phospholipids monolayer that has peripheral and embedded proteins (Farese and Walther 2009). LDs play an important role in the storage and mobilization of TAG. Several studies have shown that LDs are physically connected to the endoplasmic reticulum, mitochondria, and peroxisomes (Martin and Parton 2006; Goodman 2008; Murphy et al. 2009). The LDs play an important role in lipid metabolism and energy homeostasis through their physical connection with mitochondria (Pu et al. 2011). However, the physiological function of these contacts remains poorly understood. Studies of the interaction of LDs with mitochondria will provide new insights into the lipid metabolism and energy homeostasis as well as the pathophysiology of metabolic disorders. LDs have been seen attached to the mitochondria in adipocytes, hepatocytes, and skeletal muscle cells (Novikoff et al. 1980; Kalashnikova and Fadeeva 2006; Shaw et al. 2008). Recently, a study proposed that diacylglycerol (DAG), an intermediate of TAG metabolism, promotes mitochondrial fission (Frohman 2015). As it seems that TAG is metabolically linked with mitochondria and a recent study identified a physiological link between vacuole and mitochondria in yeast (Hughes and Gottschling 2012), we hypothesized a possible role of Ime4 in the mitochondrial functions.
The yeast m6A methyltransferase and mitochondrial functions
As glucose is being used up, the diauxic shift occurs, involving an increase in the expression of nuclear genes responsible for mitochondrial biogenesis (Ocampo et al. 2012). In yeast, fully developed and enlarged tubular mitochondria are characteristic features of stationary phase when glucose is exhausted (Aung-Htut et al. 2013). Therefore, in the present study, to understand the role of Ime4 in mitochondrial functions, the yeast cells were harvested from the stationary phase.
The yeast strains used in this work were procured from Euroscarf. The mitochondrial targeting pVT100U-mtGFP plasmid is the same as reported previously (Westermann and Neupert 2000). The growth and culture conditions were the same as reported previously (Yadav and Rajasekharan 2016). Briefly, a single colony of the yeast strains was precultured in 5 ml of YPD (yeast extract peptone dextrose) liquid medium (1% yeast extract, 2% peptone, 2% glucose, pH 6.5) in a 50-ml culture tube at 30 °C and was allowed to grow with constant shaking overnight. An equal quantity of yeast cells was taken from the preculture and subcultured in the required volume of synthetic minimal medium (SM, yeast nitrogen base: 6.7 g; amino acids drop-out mixture: 1.92 g; uracil: 76 mg/l; and 2% glucose; pH 6.5) at 30 °C. The yeast transformants harboring the pVT100U-mtGFP plasmids were precultured and subcultured in the SM − U + 2% glucose medium at 30 °C. The materials used in media preparation were purchased from HiMedia, Sigma-Aldrich, and Difco. The yeast transformation kit was obtained from Clontech.
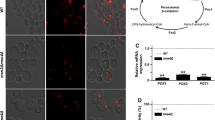
The stationary-phase cells grown in SM were harvested to assess the mitochondrial morphology and functions. The cells from different genetic backgrounds overexpressing the pVT100U-mtGFP plasmids were collected, washed, and stained with the mitochondrial dye MitoTracker Orange CMTMRos (Life Technologies). Dye at the concentration of 100 nM in 10 mM HEPES buffer containing 5% glucose (pH 7.4) was used for the staining purpose. The images were captured with a Zeiss LSM 700 confocal laser scanning microscope. The microscopic imaging showed that IME4 gene deletion causes mitochondrial fragmentation (Fig. 1a). To assess the effect of IME4 gene deletion on mitochondrial functions, cytochrome c oxidase and citrate synthase activities were also measured. The kits for Yeast mitochondria isolation (MITOISO3), cytochrome c oxidase activity (CYTOCOX1), and citrate synthase activity (CS0720) were purchased from Sigma-Aldrich. The cytochrome c oxidase and citrate synthase activities were measured according to the manufacturer’s protocol. The ime4Δ strain showed a significant decrease in cytochrome c oxidase and citrate synthase activities compared to the wild-type cells (Fig. 1b, c).
Effect of IME4 gene deletion on mitochondrial morphology and functions. a IME4 gene deletion and mitochondrial morphology. Stationary-phase yeast cells expressing mitochondrial targeting GFP (mtGFP) protein were stained with the mitochondrial dye MitoTracker Orange CMTMRos and images were taken by a confocal microscope. The optimum brightness and contrast were adjusted in the images as per requirement. Merged, superimposed panel of fluorescence and DIC (differential interference contrast); bar, 5 μm; mtGFP, pVT100U-mtGFP. b IME4 gene deletion and cytochrome c oxidase activity. c IME4 gene deletion and citrate synthase activity. In both the cases, the cells were collected from the stationary-phase and activities were measured, and the obtained ime4Δ strain values were represented in comparison with the obtained wild-type values which were set to 100%. The values are presented as the mean ± SEM (n = 3). Significance was determined at **p < 0.01
Discussion and future perspectives
In the present study, we report that IME4 gene deletion causes mitochondrial dysfunction. Identification and determination of the target genes of Ime4 that are directly involved in the mitochondrial biogenesis, morphology, and functions will pave the way to understand the role of m6A methylation in mitochondrial biology. The mitochondria are vital organelles of the living cell. Essential metabolic reactions and the regulation of some signaling cascades take place in the mitochondria (Dimmer and Scorrano 2006). Defects in the mitochondrial morphology and functions and lipid metabolism have also been associated with the neurodegenerative disorders, such as Alzheimer’s disease, and Huntington’s disease (Lin and Beal 2006). The hereditary spastic paraplegia (HSP, also known as Strumpell-Lorrain disease), a heterogeneous group of genetic neurodegenerative disorders, has also been associated with the mitochondrial dysfunction. In humans, many mitochondrial-linked rare genetic diseases are associated with mutations in poorly characterized genes. Determination and characterization of these genes and their regulation are crucial for understanding and formulating cures for the rare genetic diseases. The determination of the role of Ime4 and its targets in mitochondrial morphology and functions probably could help in the development of therapeutic strategies focused on the m6A mRNA methylation.
References
Agarwala SD, Blitzblau HG, Hochwagen A, Fink GR (2012) RNA methylation by the MIS complex regulates a cell fate decision in yeast. PLoS Genet 8:e1002732. doi:10.1371/journal.pgen.1002732
Aung-Htut MT, Lam YT, Lim YL, Rinnerthaler M, Gelling CL, Yang H, Breitenbach M, Dawes IW (2013) Maintenance of mitochondrial morphology by autophagy and its role in high glucose effects on chronological lifespan of Saccharomyces cerevisiae. Oxid Med Cell Longev 2013:636287. doi:10.1155/2013/636287
Berchowitz LE, Gajadhar AS, van Werven FJ, De Rosa AA, Samoylova ML, Brar GA, Xu Y, Xiao C, Futcher B, Weissman JS, White FM, Amon A (2013) A developmentally regulated translational control pathway establishes the meiotic chromosome segregation pattern. Genes Dev 27:2147–2163. doi:10.1101/gad.224253.113
Blanco S, Frye M (2014) Role of RNA methyltransferases in tissue renewal and pathology. Curr Opin Cell Biol 31:1–7. doi:10.1016/j.ceb.2014.06.006
Brennan CM, Steitz JA (2001) HuR and mRNA stability. Cell Mol Life Sci 58:266–277. doi:10.1007/pl00000854
Chavez S, Garcia-Martinez J, Delgado-Ramos L, Perez-Ortin JE (2016) The importance of controlling mRNA turnover during cell proliferation. Curr Genet 62:701–710. doi:10.1007/s00294-016-0594-2
Clancy MJ, Shambaugh ME, Timpte CS, Bokar JA (2002) Induction of sporulation in Saccharomyces cerevisiae leads to the formation of N6-methyladenosine in mRNA: a potential mechanism for the activity of the IME4 gene. Nucleic Acids Res 30:4509–4518
Dimmer KS, Scorrano L (2006) (De)constructing mitochondria: what for? Physiology (Bethesda Md) 21:233–241. doi:10.1152/physiol.00010.2006
Ding W, Smulan LJ, Hou NS, Taubert S, Watts JL, Walker AK (2015) s-Adenosylmethionine levels govern innate immunity through distinct methylation-dependent pathways. Cell Metab 22:633–645. doi:10.1016/j.cmet.2015.07.013
Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, Cesarkas K, Jacob-Hirsch J, Amariglio N, Kupiec M, Sorek R, Rechavi G (2012) Topology of the human and mouse m6A RNA methylomes revealed by m6A-sEq. Nature 485:201–206. doi:10.1038/nature11112
Farese RV Jr, Walther TC (2009) Lipid droplets finally get a little R-E-S-P-E-C-T. Cell 139:855–860. doi:10.1016/j.cell.2009.11.005
Frohman MA (2015) Role of mitochondrial lipids in guiding fission and fusion. J Mol Med (Berl) 93:263–269. doi:10.1007/s00109-014-1237-z
Fu Y, Dominissini D, Rechavi G, He C (2014) Gene expression regulation mediated through reversible m(6)A RNA methylation. Nat Rev Genet 15:293–306. doi:10.1038/nrg3724
Goodman JM (2008) The gregarious lipid droplet. J Biol Chem 283:28005–28009. doi:10.1074/jbc.R800042200
Harigaya Y, Tanaka H, Yamanaka S, Tanaka K, Watanabe Y, Tsutsumi C, Chikashige Y, Hiraoka Y, Yamashita A, Yamamoto M (2006) Selective elimination of messenger RNA prevents an incidence of untimely meiosis. Nature 442:45–50. doi:10.1038/nature04881
Hongay CF, Grisafi PL, Galitski T, Fink GR (2006) Antisense transcription controls cell fate in Saccharomyces cerevisiae. Cell 127:735–745. doi:10.1016/j.cell.2006.09.038
Horvath SE, Wagner A, Steyrer E, Daum G (2011) Metabolic link between phosphatidylethanolamine and triacylglycerol metabolism in the yeast Saccharomyces cerevisiae. Biochim Biophys Acta 1811:1030–1037. doi:10.1016/j.bbalip.2011.08.007
Hughes AL, Gottschling DE (2012) An early age increase in vacuolar pH limits mitochondrial function and lifespan in yeast. Nature 492:261–265. doi:10.1038/nature11654
Jia G, Fu Y, Zhao X, Dai Q, Zheng G, Yang Y, Yi C, Lindahl T, Pan T, Yang YG, He C (2011) N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol 7:885–887. doi:10.1038/nchembio.687
Jin L, Neiman AM (2016) Post-transcriptional regulation in budding yeast meiosis. Curr Genet 62:313–315. doi:10.1007/s00294-015-0546-2
Kalashnikova MM, Fadeeva EO (2006) Ultrastructural study of liver cells from rooks living in ecologically unfavorable areas. Izv Akad Nauk Ser Biol. doi:10.1134/S106235900
Lee SJ, Zhang J, Choi AM, Kim HP (2013) Mitochondrial dysfunction induces formation of lipid droplets as a generalized response to stress. Oxid Med Cell Longev 2013:327167. doi:10.1155/2013/327167
Lin MT, Beal MF (2006) Mitochondrial dysfunction and oxidative stress in neurodegenerative diseases. Nature 443:787–795. doi:10.1038/nature05292
Martin S, Parton RG (2006) Lipid droplets: a unified view of a dynamic organelle. Nat Rev Mol Cell Biol 7:373–378. doi:10.1038/nrm1912
Murphy S, Martin S, Parton RG (2009) Lipid droplet-organelle interactions; sharing the fats. Biochim Biophys Acta 1791:441–447. doi:10.1016/j.bbalip.2008.07.004
Novikoff AB, Novikoff PM, Rosen OM, Rubin CS (1980) Organelle relationships in cultured 3T3-L1 preadipocytes. J Cell Biol 87:180–196
Ocampo A, Liu J, Schroeder EA, Shadel GS, Barrientos A (2012) Mitochondrial respiratory thresholds regulate yeast chronological lifespan and its extension by caloric restriction. Cell Metab 16:55–67. doi:10.1016/j.cmet.2012.05.013
Pu J, Ha CW, Zhang S, Jung JP, Huh WK, Liu P (2011) Interactomic study on interaction between lipid droplets and mitochondria. Protein cell 2:487–496. doi:10.1007/s13238-011-1061-y
Saint-Marc C, Hurlimann HC, Daignan-Fornier B, Pinson B (2015) Serine hydroxymethyltransferase: a key player connecting purine, folate and methionine metabolism in Saccharomyces cerevisiae. Curr Genet 61:633–640. doi:10.1007/s00294-015-0489-7
Schwartz S, Agarwala SD, Mumbach MR, Jovanovic M, Mertins P, Shishkin A, Tabach Y, Mikkelsen TS, Satija R, Ruvkun G, Carr SA, Lander ES, Fink GR, Regev A (2013) High-resolution mapping reveals a conserved, widespread, dynamic mRNA methylation program in yeast meiosis. Cell 155:1409–1421. doi:10.1016/j.cell.2013.10.047
Shaw CS, Jones DA, Wagenmakers AJ (2008) Network distribution of mitochondria and lipid droplets in human muscle fibres. Histochem Cell Biol 129:65–72. doi:10.1007/s00418-007-0349-8
Singh N, Yadav KK, Rajasekharan R (2016) ZAP1-mediated modulation of triacylglycerol levels in yeast by transcriptional control of mitochondrial fatty acid biosynthesis. Mol Microbiol 100:55–75. doi:10.1111/mmi.13298
Tuck MT, Wiehl PE, Pan T (1999) Inhibition of 6-methyladenine formation decreases the translation efficiency of dihydrofolate reductase transcripts. Int J Biochem Cell Biol 31:837–851. doi:10.1016/S1357-2725(99)00041-2
Westermann B, Neupert W (2000) Mitochondria-targeted green fluorescent proteins: convenient tools for the study of organelle biogenesis in Saccharomyces cerevisiae. Yeast 16:1421–1427. doi:10.1002/1097-0061(200011)16:15<1421::aid-yea624>3.0.co;2-u
Yadav PK, Rajasekharan R (2016) Misregulation of a DDHD domain-containing lipase causes mitochondrial dysfunction in yeast. J Biol Chem 291:18562–18581. doi:10.1074/jbc.M116.733378
Yadav PK, Rajasekharan R (2017) The m6A methyltransferase Ime4 epitranscriptionally regulates triacylglycerol metabolism and vacuolar morphology in haploid yeast cells. J Biol Chem 292:13727–13744. doi:10.1074/jbc.M117.783761
Yue Y, Liu J, He C (2015) RNA N6-methyladenosine methylation in post-transcriptional gene expression regulation. Genes Dev 29:1343–1355. doi:10.1101/gad.262766.115
Zhang Z, Theler D, Kaminska KH, Hiller M, de la Grange P, Pudimat R, Rafalska I, Heinrich B, Bujnicki JM, Allain FH, Stamm S (2010) The YTH domain is a novel RNA binding domain. J Biol Chem 285:14701–14710. doi:10.1074/jbc.M110.104711
Zheng G, Dahl JA, Niu Y, Fu Y, Klungland A, Yang YG, He C (2013) Sprouts of RNA epigenetics: the discovery of mammalian RNA demethylases. RNA Biol 10:915–918. doi:10.4161/rna.24711
Zhong S, Li H, Bodi Z, Button J, Vespa L, Herzog M, Fray RG (2008) MTA is an Arabidopsis messenger RNA adenosine methylase and interacts with a homolog of a sex-specific splicing factor. Plant Cell 20:1278–1288. doi:10.1105/tpc.108.058883
Acknowledgements
This work was supported by the Council of Scientific and Industrial Research (CSIR), New Delhi, under the 12th five-year plan project LIPIC. P.K.Y. was supported by the student fellowship from CSIR, New Delhi. The corresponding author is a recipient of the JC Bose National Fellowship.
Author information
Authors and Affiliations
Contributions
RR conceived and initiated the project. RR and PKY designed the experiments. PKY executed the experiments and analyzed the data. PKY and RR discussed the data and wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Yadav, P.K., Rajasekharan, R. The m6A methyltransferase Ime4 and mitochondrial functions in yeast. Curr Genet 64, 353–357 (2018). https://doi.org/10.1007/s00294-017-0758-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-017-0758-8