Abstract
Many plant species excrete organic acids into the rhizosphere in response to aluminum stress to protect sensitive cells from aluminum rhizotoxicity. When the roots of Eucalyptus camaldulensis, a major source of pulp production, were incubated in aluminum-toxic medium, citrate released into the solution increased as a function of time. Citrate excretion was inducible by aluminum, but not by copper or sodium chloride stresses. This indicated that citrate is the major responsive organic acid released from the roots of this plant species to protect the root tips from aluminum damage. Four genes highly homologs to known citrate-transporting multidrugs and toxic compounds exclusion proteins, named EcMATE1–4, were isolated using polymerase chain reaction-based cloning techniques. Their predicted proteins included 12 membrane spanning domains, a common structural feature of citrate-transporting MATE proteins, and consisted of 502–579 amino acids with >60 % homology to orthologous genes in other plant species. One of the homologs, designated EcMATE1, was expressed in the roots more abundantly than in the shoots and in response to both Al and low pH stresses. Ectopic expression of EcMATE1 and 3 in tobacco hairy roots enhanced Al-responsive citrate excretion. Pharmacological characterization indicated that Al-responsive citrate excretion involved a protein phosphorylation/dephosphorylation process. These results indicate that citrate excretion through citrate-transporting multidrugs and toxic compounds exclusion proteins is one of the important aluminum-tolerance mechanisms in Eucalyptus camaldulensis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Eucalyptus sp., a typical fast growing tree species, is a major resource for pulp production (e.g., Greaves et al. 1997; Campinhos 1999). Over 187 million ha of plantations are currently operated throughout the world (FAO 2001). This species naturally adapts to various environmental conditions, but the established clones (i.e., elite clones) for plantation carrying suitable properties for pulp production such as long cellulose fiber length and high productivity of wood tips sometimes show poor growth at some sites. Molecular breeding such as marker-assisted selection and transgenic breeding would be one reasonable approach to confer tolerance to environment stresses to clones for plantation (Raymond and Apiolaza 2004). In fact, a genome sequence for establishing molecular markers is being developed by an international consortium (DOE Joint Genome Institute and the Eucalyptus Genome Network, EUCAGEN) and procedures for gene transformation have been established in some species (Ito et al. 1996). Identification of critical genes that regulate particular traits is important and remaining issues for realizing efficient breeding strategies.
Soil stress, such as salinity or acidity, is one of the major causes of poor growth in Eucalyptus plantations (White et al. 2009; Feikema and Baker 2011). Since acid soils are widely distributed in subtropical regions, identification of genes critical for resistance to acid soil stresses, such as aluminum (Al) stress, is important for developing molecular markers for breeding. Various resistance mechanisms, such as UDP-glucose-mediated resistance (Fukuda et al. 2007) and a lower negative charge of the plasma membrane surface (Wagatsuma et al. 1995), have been identified in various plant species, but organic acid (OA) excretion is the most common Al-resistance mechanism in many plant species (Ma et al. 2001; Kochian et al. 2004). OA excretion from the roots protects the sensitive cells in the root apex from aluminum (Al) rhizotoxicity (Delhaize et al. 1993). Inactivation of Al rhizotoxicity by phosphorus was observed in buckwheat (Zheng et al. 2005), but organic acids namely malate, citrate and oxalate excretion from the roots may be the most important mechanism of Al tolerance of this species (e.g. Ma et al. 1998; Zheng et al. 1998). Several studies have indicated that alterations in OA metabolism play a role in enhanced OA excretion in Al tolerance (e.g., canola, Anoop et al. 2003; alfalfa, Tesfaya et al. 2001), while transport capacity through the plasma membrane is also a regulating factor of this trait (Furukawa et al. 2007; Magalhaes et al. 2007). Similar mechanisms were identified in OA excretion enhanced by P-deprivation such as white lupin (Neumann et al. 1999; Kihara et al. 2003) and a mutant carrot cells (Takita et al. 1999; Ohno et al. 2003). Critical genes involved in these processes would be candidate molecular makers for marker-assisted selection to improve acid soil resistance.
An Al-responsive malate transporter, named Aluminum-activated malate transporter 1 (ALMT1), was identified in wheat (Sasaki et al. 2004) and other plant species such as Arabidopsis thaliana (Hoekenga et al. 2006). Citrate transporters, which are distinct from ALMT1 and belong to the multidrug and toxic compound extrusion (MATE) transporter family, have been identified in various plant species (Rogers and Guerinot 2002. Some citrate-transporting MATEs are Al responsive and determine the Al tolerance of crops such as sorghum (Magalhaes et al. 2007; Maron et al. 2010) and barley (HvAACT1: Hordeum vulgare aluminum-activated citrate transporter 1, Furukawa et al. 2007). Genetic analysis has revealed that the differences in Al tolerance of some varieties by OA excretion could be explained by differences in transcript levels (Sasaki et al. 2006; Furukawa et al. 2007) and protein transport capacity (Raman et al. 2008). This suggests that genes encoding these transporters would be reasonable targets for marker-assisted selection to improve acid soil resistance.
In this study, we characterized OA release from Eucalyptus roots and identified genes encoding putative citrate-transporting MATE transporters by molecular cloning. Because citrate is the major OA released from Eucalyptus roots, genes encoding putative citrate-transporting MATEs were isolated. One of four isolates was inducible by Al treatment and had high homology to previously isolated genes for citrate-transporting MATEs in other plant species. Further, characterization confirmed that the gene encodes an Al-responsive citrate-transporting MATE in Eucalyptus.
Materials and methods
Plant materials and bacterial strains
Eucalyptus camaldulensis var. obtuse (Location: EMU CREEK PETFORD) obtained from the Australian Tree Seed Centre was used throughout the experiments. Tobacco (Nicotiana tabacum, BY-2 (Bright-Yellow 2)) and the hyper-virulent Agrobacterium rhizogenes strain ATCC15834 (American type culture collection, VA, USA) carrying a native Ri plasmid were used for hairy root development.
Rhizotoxic treatments
Eucalyptus seeds were surface sterilized with sodium hypochlorite (1 % available chlorine; 20 min at 10 °C) and then pre-cultured under aseptic conditions using the method that was originally developed for Arabidopsis in vitro culture by Kobayashi et al. (2007), with minor modifications. Four nylon mesh sheets (1.3 cm square of 50 mesh per inch; 20 seedlings per sheet) were placed on a floating plastic mesh (5 cm square) in a plastic pot containing 150 ml of pre-culture medium. The pre-culture medium contained modified Hoagland–Arnon nutrients [0.4 mM Ca(NO3)2, 0.1 mM NH4H2PO4, 0.2 mM MgSO4, 40 μM KCl, 5.4 μM EDTA-Fe, 1 μM MnCl2, 4.6 μM H3BO3, 0.076 μM ZnSO4, 0.032 μM CuSO4, 0.001 μM (NH4)6Mo7O24] with 1 % sucrose at an initial pH of 5.6. The pots were kept at 23 °C with a 16-h photoperiod (20 μmol E m−2 s−1).
After 6 days of pre-culture, the floating plastic meshes supporting seedlings were transferred to a new plastic pot containing control solution (with P i removed from the pre-culture medium and the pH adjusted to 4.8) and pre-incubated for 1 h. For isolating RNA after rhizotoxic treatments, the pre-incubated seedlings were again transferred to new pots containing control, Cu-toxic (control solution + 1 μM CuSO4, pH 4.8), NaCl-toxic (control solution + 30 mM NaCl, pH 4.8), Al-toxic (control solution + 50 μM AlCl3, pH 4.6) or low pH (control solution at pH 4.0) solution. For collecting excreted organic acids from the roots, each nylon mesh growing pre-incubated seedlings on the floating plastic was transferred to a 6-well plate containing 3 ml of rhizotoxic and control solutions. To test the effect of inhibitors of protein phosphorylation and di-phosphorylation, calyculin, K-252a, staurosporin or cyclosporin were added to give a final concentration of 5 μM. All treatments were carried out under continuous light (20 μmol E m−2 s−1) at 25 °C.
RNA isolation, reverse transcription, DNA sequencing and sequence analysis
Total RNA was isolated using the method developed by Suzuki et al. (2004). The total RNA was then reverse transcribed using the Transcriptor High Fidelity cDNA synthesis kit (Roche Applied Science, Tokyo, Japan). The ABI PRISM3130xl DNA sequencer and ABI BigDye terminator system (ver3.1) were used for DNA sequencing analysis according to the manufacturer’s recommended protocols. Sequence analyses, amino acid alignment and phylogenetic tree analysis were carried out in CLUSTALW (http://www.ebi.ac.uk/Tools/msa/clustalw2/), and prediction of membrane spanning domains was carried out using HMMTOP (http://www.enzim.hu/hmmtop/).
Isolation of Eucalyptus homologues of citrate-transporting MATE family genes
Partial cDNA fragments of putative citrate-transporting MATEs were obtained by nested PCR from the cDNA of Al-treated roots using degenerate primers. The degenerate primers were designed from the conserved domains of citrate-transporting MATEs from Arabidopsis thaliana (At3g08040), Oryza sativa (Os03g11734.1) and Lupinus albus (Q3T7F5) as follows; 1st PCR: forward, 5′-GCIGCIGAYCCIYTIGCI, reverse, 5′-RCARAAIGTIACIGCIACIAC, 2nd PCR: forward, 5′-GAYACIGCITTYATHGGI reverse, 5′-RTCYTTRAAICCICKRAA. The resulting amplicons were sub-cloned and sequenced. After the sequencing analysis, the 3′ and 5′ ends of each gene were obtained according to the manufacturer’s recommended protocols of the SMART RACE cDNA Amplification Kit (Takara-bio, Ohtsu, Japan) and then sequenced. The sequences of the isolated clones were deposited in the Genbank database (Accession numbers: EcMATE1, AB725912; EcMATE2, AB725913; EcMATE3, AB725914; EcMATE4, AB725915).
Transcript analysis of the EcMATE gene family
The expression levels of the EcMATE genes were analyzed by real-time PCR using the LightCycler 480 SYBR Green I Master kit (Roche) and a Light Cycler 480 (Roche) according to manufacturer’s protocols with specific primer pairs; EcMATE1: forward 5′-AGTCTCCCTTATCAGCATTGCTTCA, reverse 5′-TAAACGTTGTGGAAGAAGTCCTTCTCTAAT, EcMATE2: forward 5′-ATGCCAGAGGACAGTGTTCAGCATCT, reverse 5′-TGCAGTGTCAATTAGGGAAGCAACAGGATC, EcMATE3: forward 5′-GCGTTGAATCTTTCTTGATTTTG, reverse 5′-CAGTCTCCCCACTTCAAGAATTA, EcMATE4: forward: 5′-CACAGGCGGCTTTGCTGCAA, reverse 5′-AGGCTGATAACTATTGGCGCTG. EcActin (forward 5′-GTTGCACCTCCTGAGAGAAAGT, reverse 5′-TAGCTCACCAACAAAGACCTTGC) was used as an endogenous control.
Vector construction and Agrobacterium rhizogenes-mediated transformation
Mini-Ti plasmid vectors carrying EcMATE1–4 were constructed using one of the special Gateway® vectors, pGWA2, for plant transformation (Nakagawa et al. 2009). Briefly, pGWA2 has a pBI121 backbone with a Gateway cloning site in which a gene can be inserted between the cauliflower mosaic virus 35s promoter (CaMV35s) and Agrobacterium nopaline synthase terminator (NOS-T). This cassette was in the T-DNA region and was flanked by gene cassettes for hygromycin resistance and kanamycin resistance. The genes for EcMATE1–4 were PCR amplified with primer pairs attaching the consensus clonase sequences and then inserted into the Gateway cloning site of the T-DNA region, resulting in pGWA2-EcMATE1–4 constructs for CaMV35s-driven overexpression of each gene in transformed hairy roots. All cloning procedures followed the standard protocols of the Gateway system (Invitrogen). Primer pairs used for directional TOPO cloning were as follows; EcMATE1: forward 5′-CACCATGGCCGAGGACTCTGATGTTCGTG, reverse 5′-TCATAAACGTTGTGGAAGAAGTCCTTCTCTAAT, EcMATE2: forward 5′-CACCATGCCAGAGGACAGTGTTCAGCATCT, reverse 5′-TTATGAAGCCTGTGGTGTACGTTGACCC, EcMATE3: forward 5′-CACCATGCCTCTGTCTATGTTCTTCAAGGA, reverse 5′-TTAGTTATTTAGAAATCCCCAAGGTCCCATGCC, EcMATE4: forward: 5′-CACCATGGAGCCTCTGGAGGGTTCG, reverse 5′-TCAGCCCCAGAGAAAATTCCAAGGACCC.
pGWA2-EcMATE1 was introduced into Agrobacterium rhizogenes ATCC15834 by electroporation, and the strain was infected to aseptically grown tobacco leaves (BY-2) to produce transgenic hairy roots. Infected tobacco leaves (1 cm square) were kept on 1/4 strength B5 agar (1 % w/v) medium containing galactose (1 % w/v) and acetosyringone (20 μg/ml) for 2 days, and then washed in 6.25 mg/l of meropenem (Sumitomo pharmaceutical Co., Japan, Tokyo) solution. The washed leaves were further incubated on 1/4 strength B5 agar medium (1 % w/v agar) containing sucrose (1 % w/v), meropenem (6.25 mg/l), and 1 ppm of naphthylacetic acid (NAA). The medium was renewed every 2 weeks until hairy root formation. Transgenic hairy roots were then selected by further culture on the same agar containing 50 μg of hygromycin instead of meropenem. All cultures were kept at 22 °C with a 16-h photoperiod (20 μmol quanta m−2 s−1).
Intracellular localization of EcMATE1 by Agrobacterium-mediated transient assay
Intracellular localization of EcMATE1 was analyzed by Agrobacterium-mediated transient assay in tobacco roots based on the method developed by Sparkes et al. (2006) that was used for transient assay in tobacco leaves using Agrobacterium tumefaciens. A cDNA encoding synthetic green fluorescent protein (sGFP; see Sawaki et al. 2009) was connected to the C-terminus of EcMATE1 by PCR reactions. Each gene was first amplified separately using primer pairs for EcMATE1 (forward 5′-TTAACCCGGGATGGCCGAGGACTCTGATGT-3′, reverse 5′-TCGCCCTTGCTCACCATTAAACGTTGTGGAAGAAGT-3′) and sGFP (forward 5′-ACTTCTTCCACAACGTTTAATGGTGAGCAAGGGCGA-3′ and reverse 5′-CGAAGAGCTCTTACTTGTACAGCTCGTCCA-3′), then jointed EcMATE1-sGFP by PCR, using the EcMATE1 forward and sGFP reverse primers. The joined DNA was replaced β-glucuronidase of pBI121 then transformed to a hyper-virulence Agrobacterium tumefaciens EHA101. Root of tobacco, grown on a agar medium [1/2 strength MS medium (Murashige and Skoog 1962) containing 1 % (w/v) sucrose] for 4 weeks, were removed and rinsed in distilled water. Then immersed to the Agrobacterium buffer (pH 5.6 of 10 mM MES/KOH, 10 mM of MgCl2, OD660 = 0.2) containing 150 μM of acetosyringone. The seedlings were kept 2 days on moistened filter paper, then the root cells were observed GFP by a fluorescent microscope (Axio, Zeiss-Japan, Tokyo).
Quantification of malate and citrate in root exudates
The citrate and malate in the solutions were measured enzymatically by the NAD+/NADH cycling method (Hampp et al. 1984) as described previously (Kihara et al. 2003). Briefly, malate and citrate were stoichiometrically converted to NADH by malate dehydrogenase or citrate lyase and then the NADH was quantified by the cycling method.
Organic acid excretion from transgenic tobacco hairy roots
Transgenic tobacco hairy roots were pre-grown for 1 week in 25 % strength MS solution (pH 5.6) with 2 % sucrose. The excised root tips (3 cm; 100 root tips) were pre-treated for 2 h in 30 ml of control treatment solution in 100 ml flasks containing 1/20 strength MS solution with 1 % (w/v) sucrose at pH 4.8 but lacking phosphate. After pre-treatment, the roots were each transferred to 30 ml of control or Al-toxic solution (control-treatment solution + 25 μM AlCl3). The medium was renewed at 24 h then incubation continued until 48 h. All flasks were kept on a rotary shaker (50 rpm) in dark room at 25 °C. The medium after 24-h treatment was used for organic acids analysis. Growth of the transgenic hairy roots was examined by root elongation for 3 days in control and Al-toxic solutions used for organic excretion assay.
Statistical analysis and reproducibility
All experiments were repeated at least twice, and confirmed reproducibility. Statistical analysis such as student t test was performed using a software Microsoft excel.
Results
Characteristics of organic acid excretion from Eucalyptus
Various plants excrete organic acids from the roots in response to Al, but there are differences between species in the organic acids excreted and the patterns of excretion in terms of induction period. In preliminary experiments, we detected both malate and citrate excretion, but not oxalate from the roots of E. camaldulensis among the major organic acids that detoxify Al toxicity. It was thus both malate and citrate in the culture solutions were quantified in various conditions.
To determine whether malate or citrate was the major organic acid in the root exudates, the roots of 1-week-old E. camaldulensis seedlings (root length 5–10 mm) were exposed to Al solutions. Citrate excretion was increased when the Al concentration was greater than 25 μM, while malate excretion showed no response to external Al (Fig. 1a). The citrate excretion in Al solution was time-dependent (Fig. 1b). The excretion was determined every 3 h and increased for at least the first 12 h. These results suggested that E. camaldulensis possesses Al-inducible citrate excretion capacity.
Citrate and malate release profile in Eucalyptus camaldulensis. a Response to Al concentration. b Time-course response to Al (50 μM) stress. c Growth of roots with various stressors (50 μM AlCl3, low pH (pH 4.0), 1.0 μM CuSO4 and 30 mM NaCl) for 3 days. d Citrate release with various stressors (identical to conditions used for c). Each solution was adjusted to pH 4.6 except the low pH treatment. Each experiment was replicated three times and mean ± SD are shown
To further compare citrate and malate excretion from the roots, both organic acids were quantified after incubation in other rhizotoxic treatments, including 50 μM Al, low pH (pH 4.0), 1.0 μM Cu or 30 mM NaCl. Each stress treatment caused 50 % inhibition of root elongation compared with controls (Fig. 1c). Similar levels of malate excretion were detected in all treatments, but citrate excretion was only detected in the Al treatment, in which it was greater than malate excretion (Fig. 1d). These results indicated that citrate excretion is specific to Al stress.
Isolation of putative EcMATE genes by PCR-based cloning
Using degenerate primers designed from the conserved amino acid sequence shared by the reported citrate-transporting MATE genes, about 40 partial cDNA fragments encoding EcMATE family genes were obtained. From subsequent 3′ and 5′ RACE using specific primers designed from the partial fragments, four different genes encoding putative EcMATE proteins were isolated. The cDNA clones showed conserved amino acid sequences at the positions of the degenerate primers, and encoded proteins of 502 (EcMATE3) to 579 (EcMATE1) amino acids (Table 1). These genes contained both 5′ and 3′ UTR regions.
The amino acid sequences of the EcMATEs shared typical features of citrate-transporting MATE proteins in other plant species (Fig. 2a). All of the EcMATE proteins had 12 predicted trans-membrane domains, which was identical to other citrate-transporting MATE proteins. Phylogenetic tree analysis grouped the EcMATE1 and two proteins in the same group, while EcMATE 3 and 4 fell into another group (Fig. 2b). All of these proteins were closer to the dicotyledonous plants (Arabidopsis and white lupin) than to homologs in monocotyledonous plants. AtFRD3 (Arabidopsis thaliana ferric redictase defective3) a citrate transporter involved in iron-acquisition, was closest to EcMATE1 and 2, while AtMATE, an Al-responsive citrate-transporting Arabidopsis MATE, was closest to EcMATE4.
Amino acid alignment (a) and phylogenetic tree (b) of citrate-transporting MATE proteins from Eucalyptus and other plant species. EcMATE1–4, Eucalyptus camaldulensis; AtMATE (At1g51340) and AtFRD3 (At3g08040), Arabidopsis thaliana; OsFRDL1 (Os03g0216700), Oryza sativa; HvAACT1 (BAF75822), Hordeum vulgare; SbMATE (ABS89149), Sorghum bicolor; LaMATE (AAW30733), Lupinus albus; ZmMATE1 (FJ015156.1), Zea maize. TM1–12 in a indicates transmembrane regions predicted by HMMTOP, and the box indicates the highly conserved region. The scale bar in b indicates amino acid substitutions per site. The actual value depends on the branch lengths in the tree
Expression profile of the EcMATE genes in response to rhizotoxic treatments
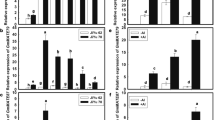
To characterize EcMATE gene expression, transcripts levels of the genes were analyzed in both shoots and roots under various rhizotoxic treatments. The EcMATE4 transcript was expressed in both shoots and roots, while the transcript levels of EcMATE1, 2 and 3 were greater in the roots under various treatments (Suppl Fig. S1). Among the root-abundant EcMATE homologs, EcMATE1 showed Al- and low pH-responsive expression, when expression levels were compared among stress treatments causing 50 % growth inhibition (Fig. 1c). EcMATE1 expression was induced by Al and low pH, while both NaCl (30 mM) and Cu (1.0 μM) did not induce (Fig. 3a). The greatest fold change (i.e., Al/control) of EcMATE1 was 6.4 after 4-h Al treatment, when the roots began citrate excretion (Figs. 1b, 3b). EcMATE1 expression was induced by Al and low pH stress in a dose-dependent manner (Fig. 3c, d). In addition, EcMATE1 expression was greater in the root tip than the mature part (Fig. 3e), which was the site of OA excretion (Suppl Fig. S2). These results suggested that EcMATE1 plays an important role in Al-inducible citrate excretion in E. camaldulensis.
MATE gene expression analysis of E. camaldulensis roots treated with ion stress. a Expression of MATE1–4 to various stressors for 24 h (see conditions in Fig. 1c), b time-course analysis of EcMATE1 expression treated with 50 μM Al by real-time PCR. Dose response analysis of Al (c) and low pH (d). e Relative expression of EcMATE1 in root tip (0–5 mm) and mature part of roots (>5 mm) with or without 50 μM Al for 24 h. Expression level was quantified by real-time PCR. Means of three replicates and the error bar ± SD are indicated
Subcellular localization of EcMATE1
To visualize subcellular localization of EcMATE1, we analyzed localization of EcMATE1:sGFP by Agrobacterium-mediated transient expression assay. When cytosolic sGFP was transformed to the tobacco roots, all part of cells fluoresced brightly (Fig. 4a, c). On the other hand, plant expressing EcMATE1:sGFP fusion protein carried many cells that showed green fluoresce at the edge of the cell (Fig. 4b, d). It suggested that EcMATE1 protein localized at the plasma membrane.
Subcellular localization of EcMATE1:: sGFP fusion protein in tobacco root cells. Genes for fusion protein of EcMATE1:: sGFP (a, b) and cytosol-localizing sGFP (c, d) were introduced to tobacco by Agrobacterium transformation then visualized by a fluorescent microscope at 3 days after infection (a, c). Bright field images (b, d) are also shown. Bar indicates 50 μm
Ectopic expression of EcMATE1 in tobacco hairy roots
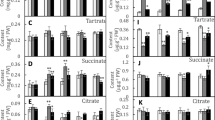
To characterize the function of EcMATE1 in Al-responsive citrate efflux, transgenic hairy roots of tobacco were obtained using a hyper-virulent A. rhizogenes strain carrying native Ri plasmids and Ti plasmids containing EcMATE and KmR genes in the T-DNA region. Transgenic hairy roots carrying CaMV35s:: EcMATE1, which was selected as KmR, excreted citrate into the Al medium, at levels greater than the GUS (β-glucuronidase)-transformed control. The EcMATE1 transgenic did not show enhanced citrate excretion in the absence of Al (Fig. 5a). In addition, the transgenic hairy roots carrying EcMATE1 grew better than the control transgenic carrying GUS (Fig. 5b). Among the other homologs, EcMATE3 showed Al-responsive citrate excretion (Suppl Fig. S3), while it was not inducible by Al (Fig. 1c). These results indicated that both EcMATE1 encodes Al-responsive citrate-transporting MATE in E. camaldurensis.
Citrate release and growth of transgenic tobacco hairy roots carrying EcMATE1. a Citrate release at 24 h in Al (25 μM at pH 4.8) and control (pH 4.8) media. b Relative root elongation (25 μM Al at pH 4.8–0 Al control at pH 4.8). GUS transgenic lines were used as a control experiment. Mean ± SD are shown (n = 3). Asterisks indicate significant difference compared with each control (GUS) line by student t test (P < 0.05)
Response of EcMATE1 expression to protein kinase and phosphatase inhibitors
To determine whether the citrate release in Eucalyptus is regulated by a protein phosphorylation/dephosphorylation process, Al-responsive citrate excretion was characterized in the presence of various protein kinase and phosphatase inhibitors (Fig. 6). Al-responsive citrate excretion was affected by both protein kinase (K-252a and staurosporine) and phosphatase inhibitors (calyculin A and cyclosporin A). Cyclosporin A inhibited excretion by 40 %, while the others inhibited excretion by over 70 % (Fig. 6a). These effective inhibitors, K-252A, staurosporine and calyculin, repressed the expression of EcMATE1 (Fig. 6b), while a constitutively expressing EcMATE2 maintained expression (Suppl Fig. S4). These results suggested that Al-responsive citrate excretion through EcMATE1 involves a protein phosphorylation/dephosphorylation process for inducing citrate excretion.
Effect of protein kinase and phosphatase inhibitors on citrate release and EcMATE1 expression. a Citrate release to Al solution (50 μM, pH 4.6) from E. camaldulensis roots in the presence of protein kinase inhibitors (K-252a and staurosporin A) and protein phosphatase inhibitors (calyculin A and cyclosporin A) or the absence of protein phosphorylation/dephosphorylation inhibitors. b Transcripts of EcMATE1 in roots treated with Al solution (50 μM, pH 4.6) alone or supplemented with calyculin A or K-252a. Citrate released into the medium and the transcript levels of EcMATE1 were analyzed after 24-h treatment. Each experiment was replicated three times and mean ± SD are shown. Calyculin A and K-252a greatly repressed Al-responsive citrate release from the roots
Discussion
Many plant species release OA from the roots to protect Al-sensitive cells in the apex (Ma et al. 2001; Kochian et al. 2004). This strategy is common in herbaceous plants in both monocot and dicots. In this study, we determined that Eucalyptus releases citric acids in response to Al (Fig. 1a). This indicates that OA excretion from the roots also plays a role in the Al tolerance of some woody plant species. Eucalyptus excreted malate and citrate into the rhizosphere under control conditions (Al stress free conditions), but Al toxicity only enhanced the citrate excretion (Fig. 1d). This can be explained by the Al-inducible expression of EcMATE transporter(s) closely homologous to known plant citrate-transporting MATE proteins.
It has been reported that MATEs have 12 transmembrane domains and one conserved region linked to citrate-transporting capacity (Yang et al. 2011). Four MATE homologs were isolated from E. camaldulensis. All genes shared the consensus features of citrate-transporting MATEs, namely the 12 transmembrane domains and the conserved region among the known citrate-transporting MATEs (Fig. 2). Among the homologs, EcMATE1 was uniquely up-regulated during Al treatment and nearly specific to Al and low pH (Fig. 3), which is a typical expression pattern of Al-responsive OA transporters. For example, AtALMT1 regulating Al-responsive malate excretion in Arabidopsis thaliana is inducible by Al in an ion-specific manner (Kobayashi et al. 2007). Although the contribution to Al tolerance was less than AtALMT1, a citrate-transporting MATE (AtMATE) showed a similar Al-induced pattern in Arabidopsis (Liu et al. 2009).
Ectopic expression of EcMATE1 in tobacco by Agrobacterium rhizogenesis-mediated transformation enhanced Al-responsive citrate excretion in the hairy roots (Fig. 3). This indicated that the citrate-transporting EcMATE1 could function in tobacco cells. This was similar to results where transgenic Arabidopsis carrying SbMATE displayed Al tolerance by enhanced citrate excretion (Magalhaes et al. 2007). The citrate transport functionality of these ectopically expressed MATE proteins could be explained by the activation of the protein by Al. Oocyte studies have indicated that MATE proteins could be activated by direct interaction with Al (e.g., Magalhaes et al. 2007). The involvement of protein phosphorylation in this Al-activating process was suggested by the Al-responsive malate excretion of an ALMT1-type malate transporter in Arabidopsis (Kobayashi et al. 2007). Similar protein phosphorylation might have activated EcMATE1 in the Eucalyptus. In fact, inhibitors of protein phosphorylation/di-phosphorylation affected Al-responsive citrate excretion.
Organic acid excretion plays a role in various plant stress tolerances, including Al tolerance, efficient P i (Neumann et al. 1999) and Fe acquisition (Miethke and Marahiel 2007), and rhizo-bacterium-mediated pathogen resistance (e.g., Rudrappa et al. 2008). Molecular studies of OA excretion have revealed that transporters and their regulation are critical factors for determining organic acid species and stress responses. Recently, promoter evolution was proposed as the mechanism by which barley has obtained Al-responsive HvAACT1-mediated citrate excretion. Promoter analysis of HvAACT1 indicated that evolutionary changes in the promoter of a citrate-transporting MATE, possibly for Fe-acquisition, resulted in altered expression patterns of the gene to protect the sensitive root tip from Al toxicity (Fujii et al. 2012). EcMATE1 expression was slightly greater in the root tip than mature root tissue (Fig. 3e) and would localize plasma membrane (Fig. 4) similar to HvAACT1. Although genetic association of EcMATE1 and Al tolerance has not been clarified yet, further analysis of EcMATE1 would be interesting to understand the evolution of Al-responsive citrate excretion in woody plant species.
The direction of further studies will be towards enhanced Al tolerance of Eucalyptus. Overexpression of Al-tolerance genes such as EcMATE1 would be one approach. In this case, the combination of EcMATE1 overexpression and the alteration of OA metabolism (e.g., over-expression of citrate synthase; in canola by Anoop et al. 2003, in Nicotiana benthamiana by Deng et al. 2009) might be useful to efficiently enhance Al tolerance via citrate excretion. Recently, key transcription factors for Al tolerance were isolated in Arabidopsis (STOP1: sensitive to proton rhizotoxicity1, Iuchi et al. 2007) and rice (ART1: Al-resistance transcription factor 1, Yamaji et al. 2009). These proteins have high homology to Cys2–His2 zinc finger proteins and regulate multiple genes controlling Al tolerance. Although regulation of EcMATE1 by a STOP1 homolog has not been investigated in Eucalyptus, co-regulation of multiple Al-tolerance genes, including AtMATE (Liu et al. 2009), was associated with Arabidopsis STOP1 (Sawaki et al. 2009) and rice ART1 (Yamaji et al. 2009). Further analysis to identify STOP1-homolog regulatory genes would be an interesting approach for identifying co-regulated Al-tolerance genes in Eucalyptus. Such genes could also be utilized for developing molecular markers in Eucalyptus for efficient molecular breeding of Al tolerance.
Abbreviations
- Al:
-
Aluminum
- ALMT1:
-
Aluminum-activated malate transporter 1
- ART1:
-
Al-resistance transcription factor 1
- AtFRD3:
-
Arabidopsis thaliana ferric redictase defective3
- HvAACT1:
-
Hordeum vulgare aluminum-activated citrate transporter 1
- GUS:
-
β-Glucuronidase
- MATE:
-
Multidrugs and toxic compounds exclusion
- MS:
-
Murashige and Skoog
- OAs:
-
Organic acids
- STOP1:
-
Sensitive to proton rhizotoxicity1
References
Anoop VM, Basu U, McCammon MT, McAlister-Henn L, Taylor GJ (2003) Modulation of citrate metabolism alters aluminum tolerance in yeast and transgenic canola overexpressing a mitochondrial citrate synthase. Plant Physiol 132:2205–2217
Campinhos E (1999) Sustainable plantations of high-yield Eucalyptus trees for production of fiber: the Aracruz case. New Forest 17:129–143
Delhaize E, Ryan PR, Randall PJ (1993) Aluminum tolerance in wheat (Triticum asestivum L.) II. Aluminum-stimulated excretion of malic acid from root apices. Plant Physiol 103:695–702
Deng W, Luo K, Li Z, Yang Y, Hu N, Wu Y (2009) Overexpression of Citrus junos mitochondrial citrate synthase gene in Nicotiana benthamiana confers aluminum tolerance. Planta 230:355–365
FAO (2001) Global data on forest plantations resources in Forest genetic resources, FAO, Rome, http://www.fao.org/docrep/004/Y2316E/y2316e0b.htm#bm11
Feikema P, Baker T (2011) Effect of soil salinity on growth of irrigated plantation Eucalyptus in south-eastern Australia. Agric Water Manage 98(7):1180–1188
Fujii M, Yokosho K, Yamaji N, Saisho D, Yamane M, Takahashi H, Sato K, Nakazono M, Ma JF (2012) Acquisition of aluminium tolerance by modification of a single gene in barley. Nat Commun 3:713
Fukuda T, Saito A, Wasaki J, Shinano T, Osaki M (2007) Metabolic alterations proposed by proteome in rice roots grown under low P and high Al concentration under low pH. Plant Sci 172:1157–1165
Furukawa J, Yamaji N, Wang H, Mitani N, Murata Y, Sato K, Katsuhara M, Takeda K, Ma JF (2007) An aluminum-activated citrate transporter in barley. Plant Cell Physiol 48:1081–1091
Greaves BL, Borralho NMG, Raymond CA (1997) Breeding objective for plantation eucalypts grown for production of kraft pulp. For Sci 43:465–472
Hampp R, Goller M, Füllgraf H (1984) Determination of compartmented metabolite pools by a combination of rapid fractionation of oat mesophyll protoplasts and enzymic cycling. Plant Physiol 75:1017–1021
Hoekenga OA, Maron LG, Pineros MA, Cancado GM, Shaff J, Kobayashi Y, Ryan PR, Dong B, Delhaize E, Sasaki T, Matsumoto H, Yamamoto Y, Koyama H, Kochian LV (2006) AtALMT1, which encodes a malate transporter, is identified as one of several genes critical for aluminum tolerance in Arabidopsis. Proc Natl Acad Sci USA 103:9738–9743
Ito K, Doi K, Tatemichi Y, Shibata M (1996) Plant regeneration of eucalypts from rotating nodule cultures. Plant Cell Rep 16:42–45
Iuchi S, Koyama H, Iuchi A, Kobayashi Y, Kitabayashi S, Kobayashi Y, Ikka T, Hirayama T, Shinozaki K, Kobayashi M (2007) Zinc finger protein STOP1 is critical for proton tolerance in Arabidopsis and coregulates a key gene in aluminum tolerance. Proc Natl Acad Sci USA 104:9900
Kihara T, Wada T, Suzuki Y, Hara T, Koyama H (2003) Alteration of citrate metabolism in cluster roots of white lupin. Plant Cell Physiol 44:901–908
Kobayashi Y, Hoekenga OA, Itoh H, Nakashima M, Saito S, Shaff JE, Maron LG, Pineros MA, Kochian LV, Koyama H (2007) Characterization of AtALMT1 expression in aluminum-inducible malate release and its role for rhizotoxic stress tolerance in Arabidopsis. Plant Physiol 145:843–852
Kochian LV, Hoekenga OA, Pineros MA (2004) How do crop plants tolerate acid soils? Mechanisms of aluminum tolerance and phosphorous efficiency. Annu Rev Plant Biol 55:459–493
Liu J, Magalhaes JV, Shaff J, Kochian LV (2009) Aluminum-activated citrate and malate transporters from the MATE and ALMT families function independently to confer Arabidopsis aluminum tolerance. Plant J 57:389–399
Ma JF, Hiradate S, Matsumoto H (1998) High aluminum resistance in buckwheat. Plant Physiol 117:753–759
Ma JF, Ryan PR, Delhaize E (2001) Aluminium tolerance in plants and the complexing role of organic acids. Trends Plant Sci 6:273–278
Magalhaes JV, Liu J, Guimaraes CT, Lana UG, Alves VM, Wang YH, Schaffert RE, Hoekenga OA, Pineros MA, Shaff JE, Klein PE, Carneiro NP, Coelho CM, Trick HN, Kochian LV (2007) A gene in the multidrug and toxic compound extrusion (MATE) family confers aluminum tolerance in sorghum. Nat Genet 39:1156–1161
Maron LG, Piñeros MA, Guimarães CT, Magalhaes JV, Pleiman JK, Mao C, Shaff J, Belicuas SNJ, Kochian LV (2010) Two functionally distinct members of the MATE (multi-drug and toxic compound extrusion) family of transporters potentially underlie two major aluminum tolerance QTLs in maize. Plant J 61:728–740
Miethke M, Marahiel MA (2007) Siderophore-based iron acquisition and pathogen control. Microbiol Mol Biol Rev 71:413–451
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497
Nakagawa T, Ishiguro S, Kimura T (2009) Gateway vectors for plant transformation. Plant biotechnol 26:275–284
Neumann G, Massonneau A, Martinoia E, Römheld V (1999) Physiological adaptations to phosphorus deficiency during proteoid root development in white lupin. Planta 208:373–382
Ohno T, Koyama H, Hara T (2003) Characterization of citrate transport through the plasma membrane in a carrot mutant cell line with enhanced citrate excretion. Plant Cell Physiol 44:156–162
Raman H, Ryan PR, Raman R, Stodart BJ, Zhang K, Martin P, Wood R, Sasaki T, Yamamoto Y, Mackay M (2008) Analysis of TaALMT1 traces the transmission of aluminum resistance in cultivated common wheat (Triticum aestivum L.). Theor Appl Genet 116:343–354
Raymond CA, Apiolaza LA (2004) Incorporating wood quality and deployment traits in Eucalyptus globulus and Eucalyptus nitens. In: Plantation forest biotechnology for the 21st century. Forest Research New Zealand, Rotorua, pp 87–99
Rogers EE, Guerinot ML (2002) FRD3, a member of the multidrug and toxin efflux family, controls iron deficiency responses in Arabidopsis. Plant Cell 14:1787–1799
Rudrappa T, Czymmek KJ, Pare PW, Bais HP (2008) Root-secreted malic acid recruits beneficial soil bacteria. Plant Physiol 148:1547–1556
Sasaki T, Yamamoto Y, Ezaki B, Katsuhara M, Ahn SJ, Ryan PR, Delhaize E, Matsumoto H (2004) A wheat gene encoding an aluminum-activated malate transporter. Plant J 37:645–653
Sasaki T, Ryan PR, Delhaize E, Hebb DM, Ogihara Y, Kawaura K, Noda K, Kojima T, Toyoda A, Matsumoto H (2006) Sequence upstream of the wheat (Triticum aestivum L.) ALMT1 gene and its relationship to aluminum resistance. Plant Cell Physiol 47:1343–1354
Sawaki Y, Iuchi S, Kobayashi Y, Kobayashi Y, Ikka T, Sakurai N, Fujita M, Shinozaki K, Shibata D, Kobayashi M, Koyama H (2009) STOP1 regulates multiple genes that protect Arabidopsis from proton and aluminum toxicities. Plant Physiol 150:281–294
Sparkes IA, Runions J, Kearns A, Hawes C (2006) Rapid, transient expression of fluorescent fusion proteins in tobacco plants and generation of stably transformed plants. Nat Protoc 1:2019–2025
Suzuki Y, Kawazu T, Koyama H (2004) RNA isolation from siliques, dry seeds, and other tissues of Arabidopsis thaliana. BioTechniques 37:542–544
Takita E, Koyama H, Hara T (1999) Organic acid metabolism in aluminum-phosphate utilizing cells of carrot (Daucus carota L.). Plant Cell Physiol 40:489–495
Tesfaya M, Temple SJ, Allan DL, Vance CP, Samac DA (2001) Overexpression of malate dehydrogenase in transgenic alfalfa enhances organic acid synthesis and confers tolerance to aluminum. Plant Physiol 127:1836–1844
Wagatsuma T, Jujo K, Ishikawa S, Nakashima T (1995) Aluminum-tolerant protoplasts from roots can be collected with positively charged silica microbeads: a method based on differences in surface negativity. Plant Cell Physiol 36:1493–1502
White DA, Crombie DS, Kinal J, Battaglia M, McGrath JF, Mendham DS, Walker SN (2009) Managing productivity and drought risk in Eucalyptus globulus plantations in south-western Australia. For Ecol Manage 259:33–44
Yamaji N, Huang CF, Nagao S, Yano M, Sato Y, Nagamura Y, Ma JF (2009) A zinc finger transcription factor ART1 regulates multiple genes implicated in aluminum tolerance in rice. Plant Cell Online 21:3339–3349
Yang XY, Yang JL, Zhou Y, Pineros MA, Kochian LV, Li GX, Zheng SJ (2011) A de novo synthesis citrate transporter, Vigna umbellata multidrug and toxic compound extrusion, implicates in Al-activated citrate efflux in rice bean (Vigna umbellata) root apex. Plant, Cell Environ 34:2138–2148
Zheng SJ, Ma JF, Matsumoto H (1998) High aluminum resistance in buckwheat. I. Al-induced specific secretion of oxalic acid from root apex. Plant Physiol 117:745–751
Zheng SJ, Yang JL, He YF, Yu XH, Zhang L, You JF, Shen RF, Matsumoto H (2005) Immobilization of aluminum with phosphorus in roots is associated with high aluminum resistance in buckwheat. Plant Physiol 138:297–303
Acknowledgments
We thank Dr Tsuyoshi Nakagawa (Shimane University) for providing the gateway vector, pGWB2. This work was supported by the METI (Ministry of Economy, Trade and Industry) Grant Program, Japan.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sawaki, Y., Kihara-Doi, T., Kobayashi, Y. et al. Characterization of Al-responsive citrate excretion and citrate-transporting MATEs in Eucalyptus camaldulensis . Planta 237, 979–989 (2013). https://doi.org/10.1007/s00425-012-1810-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-012-1810-z