Abstract
Allele diversities of four markers specific to intron three, exon four and promoter regions of the aluminum (Al) resistance gene of wheat (Triticum aestivum L.) TaALMT1 were compared in 179 common wheat cultivars used in international wheat breeding programs. In wheat cultivars released during the last 93 years, six different promoter types were identified on the basis of allele size. A previous study showed that Al resistance was not associated with a particular coding allele for TaALMT1 but was correlated with blocks of repeated sequence upstream of the coding sequence. We verified the linkage between these promoter alleles and Al resistance in three doubled haploid and one intercross populations segregating for Al resistance. Molecular and pedigree analysis suggest that Al resistance in modern wheat germplasm is derived from several independent sources. Analysis of a population of 278 landraces and subspecies of wheat showed that most of the promoter alleles associated with Al resistance pre-existed in Europe, the Middle East and Asia prior to dispersal of cultivated germplasm around the world. Furthermore, several new promoter alleles were identified among the landraces surveyed. The TaALMT1 promoter alleles found within the spelt wheats were consistent with the hypothesis that these spelts arose on several independent occasions from hybridisations between non-free-threshing tetraploid wheats and Al-resistant hexaploid bread wheats. The strong correlation between Al resistance and Al-stimulated malate efflux from the root apices of 49 diverse wheat genotypes examined was consistent with the previous finding that Al resistance in wheat is conditioned primarily by malate efflux. These results demonstrate that the markers based on intron, exon and promoter regions of TaALMT1 can trace the inheritance of the Al resistance locus within wheat pedigrees and track Al resistance in breeding programmes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Although a significant variation in Al resistance occurs in wheat, it is generally considered to be one of the more Al-sensitive members of the Poaceae (Garvin and Carver 2003). This variation has been used to improve yields on acid soils and Al resistance present in many modern cultivars originates from Central and South America where the strongly acidic soils imposed a severe selection pressure. The genes for Al resistance were transmitted through a small number of genotypes for subsequent cultivar development (De Sousa 1998). For instance, Brazilian wheat cultivars such as Frontana, Carazinho, BH1146, Maringa and Cotipora have been used as donor sources of Al resistance genes to breed wheat varieties adapted to acidic soils worldwide (Hettel 1989; http://genbank.vurv.cz/wheat/pedigree/).
In many wheat populations, genetic analysis indicates that Al resistance is controlled by a major gene (Kerridge and Kronstad 1968; Delhaize et al. 1993a; Somers and Gustafson 1995; Somers et al. 1996; Basu et al. 1997; Raman et al. 2005). A major locus conditioning Al resistance in wheat has been mapped in many segregating populations to the long arm of chromosome 4D (4DL) (Luo and Dvorák 1996; Riede and Anderson 1996; Rodriguez-Milla and Gustafson 2001; Raman et al. 2005). Multigenic inheritance for Al resistance in wheat has also been reported. For instance, Berzonsky (1992) concluded that the Al resistance of Atlas 66 was encoded by more than two genes and that the genes were not all located on the D-genome chromosomes. Ma et al. (2005) identified a single major QTL for Al resistance in Atlas 66 located on 4DL and more recently Zhou et al. (2007) reported on the presence of an additional minor QTL for Al resistance in Atlas 66 located on chromosome 3BL.
The major mechanism for Al resistance in wheat is now well understood. Compared to Al-sensitive wheat, Al resistant lines release greater amounts of malate from their root apices (Delhaize et al. 1993b; Ryan et al. 1995b; Tang et al. 2002; Raman et al. 2005) which chelate and detoxify the harmful Al cations. This is further supported by studies showing that Al ions activate anion currents at the root apices of Al-resistant seedlings (Ryan et al. 1997) that are due to malate efflux (Zhang et al, 2001). Furthermore, a study demonstrated that Al-resistance and the malate efflux phenotype in wheat cosegregate and map to the same position on chromosome 4DL (Raman et al. 2005). Recently, TaALMT1 (originally named ALMT1) was cloned (Sasaki et al. 2004) and mapped to the same location on chromosome 4DL (Ma et al. 2005; Raman et al. 2005). TaALMT1 encodes for a membrane-localised protein (Yamaguchi et al. 2005) which confers an Al-activated malate efflux and greater Al resistance to transgenic tobacco cells and intact barley plants (Sasaki et al. 2004; Delhaize et al. 2004). Furthermore the level of TaALMT1 expression is correlated with Al resistance in a range of wheat genotypes. These results provide a strong evidence that the TaALMT1 is the major Al resistance gene of wheat.
Analysis of the exon sequences of TaALMT1 has, to date, only identified two alleles neither of which is diagnostic of Al resistance (Sasaki et al. 2004; Raman et al. 2005). In contrast, intron three and the promoter region show considerable allelic variability. The variation in intron three is due to a simple sequence region (SSR) with variable copy numbers and the presence of indels (Raman et al. 2005, 2006). Variation in the promoter region consists of single nucleotide polymorphisms (SNPs) as well as single or multiple tandem repeats of sequence 31–803 bp long (Sasaki et al. 2006). Sasaki et al. (2006) suggested that the variations in the promoter sequence are responsible, at least in part, for the different levels of TaALMT1 expression in a range of wheat cultivars of non-Japanese origin.
The availability of markers for exon four (Sasaki et al. 2004), intron three (Raman et al. 2006) and the promoter region (Sasaki et al. 2006) of TaALMT1, has now made it possible to study the genetic relationships of the TaALMT1 gene in a range of wheat genotypes. We have employed these markers to characterize wheat germplasm currently used in breeding programs internationally and in a selection of landraces that included subspecies of common wheat (T. aestivum L.). Our objectives were to (1) determine the allele diversity of the TaALMT1 gene among a large number of cultivated wheat varieties and in a selection of landraces, (2) determine whether certain promoter types co-segregate with Al resistance, (3) investigate the transmittance of TaALMT1 alleles in some wheat varieties released over the last 93 years and (4) assess the relative merits of the various markers for their application in marker-assisted selection (MAS).
Materials and methods
Plant materials
Seed of 179 common wheat cultivars of known pedigree and 278 landraces derived from Europe, the Middle East and Asia including 28 accessions of the T. aestivum subspecies (also referred to as major morphological groups: spelta, macha, vavilovii, compactum and sphaerococcum) were provided by the Australian Winter Cereal Collection in Tamworth, Australia and the wheat breeding node of Enterprise Grains Australia at Wagga Wagga. Pedigree information and year of release of the cultivars were obtained from Whiting (2001) or from the website (http://genbank.vurv.cz/wheat/pedigree/) and are presented in Supplementary Table 1.
Three doubled haploid (DH) and one intercross mapping populations segregating for Al resistance were used to investigate linkage relationships between the Al resistance phenotype and the TaALMT1 promoter marker. These populations were derived from Diamondbird (Al-resistant)/Janz (Al-sensitive), Spica (Al-sensitive)/Maringa (Al-resistant) and Currawong (Al-resistant)/CD87 (Al-sensitive) (Raman et al. 2005). F3 families derived from an ET8 (resistant)/ES8 (sensitive) that had previously been scored for Al resistance (Sasaki et al. 2004) were also used to investigate the linkage between Al resistance and alleles in the promoter region of TaALMT1.
Evaluation of aluminum resistance
The germplasm accessions including wheat cultivars and mapping populations were grown in a nutrient solution following the methods described by Raman et al. (2002). Seeds were surface-sterilised with a 1.0% NaOCl solution for 10 min, submerged in an aerated solution of Vitavax (0.005 g/L), and incubated at 20°C for approximately 16 h in darkness. Eight germinated seeds from each cultivar or line of a mapping population showing similar growth were laid on a plastic mesh placed in contact with aerated nutrient solution containing (μM): Ca 1,000; Mg 400; K 1,000; NO3 3,400; NH4 600; PO4 100; SO4 401.1; Cl 78; Na 40.2; Fe 20; B 23; Mn 9; Zn 0.8; Cu 0.30; and Mo 0.1. Seedlings were exposed to a photoperiod of 16/8 h (day/night) and incubated at 20°C for a further 48 h. Seedlings were then transferred to aerated nutrient solution containing 80 μM Al for 24 h, under the same growth conditions. The pH of all solutions was maintained at 4.2 ± 0.1 throughout the experiment. Seedlings were removed from the nutrient + Al solution and the roots were washed in trays containing distilled water for 10 min. Roots were stained with hematoxylin solution containing 0.2% (w/v) hematoxylin and 0.02% (w/v) KIO3 for 15 min (Polle et al. 1978). Four wheat cultivars with known Al resistance response: Carazinho and Dollarbird (Al-resistant), and Rosella and Banks (Al-sensitive) were used in all experiments as controls. The experiment was performed twice.
Measurement of malate efflux
Malate efflux from excised root apices was measured as described previously (Ryan et al. 1995b). Briefly, surface-sterilised seed were grown for 4 days over aerated 0.2 mM CaCl2 (pH 4.2). Twenty root apices (3–4 mm) per cultivar were excised and placed in 5 mL sterilised glass vials containing 1 mL CaCl2 solution (0.2 mM, pH 4.5), sealed with Parafilm™ and placed on a shaker (∼60 rpm) for 40–60 min. The vials were removed from the shaker, the tissue washed twice with 1 mL of the same CaCl2 solution, then 1 mL of treatment solution (0.1 mM AlCl3, 0.2 mM CaCl2; pH 4.5) was added to each vial and returned to the shaker for 60 min. The solution was collected and malate quantified using an enzyme assay (Delhaize et al. 1993b). Briefly, 0.75 mL of glycine buffer (0.5 M glycine, 0.4 M hydrazine hydrate; pH 9.0) was mixed with 30 μL of NAD+ solution (30 mg/mL) and 0.68 mL of sample. The reaction mixture was pre-incubated for 30 min to stabilise chemical reactions and then 5 μL of L-malate dehydrogenase (5 mg/mL, Boehringer–Mannheim) was added to oxidise malate to oxaloacetate. The appearance of NADH was measured using the increase in absorbance at 340 nm and the final change was proportional to the initial malate in the sample. To determine the relationship between Al resistance and level of malate efflux in diverse wheat cultivars, relative root length was used to evaluate Al resistance which was calculated as the mean root length in nutrient solution containing 80 μM Al divided by the mean root length in nutrient solution without Al. The lengths of the longest roots of at least eight seedlings per treatment were averaged.
DNA extraction
After seedlings had been evaluated for Al resistance by hematoxylin staining, leaf samples were collected in 96-well formatted 1.2 mL tubes. The samples were frozen in liquid nitrogen and ground to a fine powder using a mixer mill (Retsch GmbH & Co, Germany) and DNA was extracted using the DNeasy 96 kit (Qiagen, Hilden, Germany). The DNA was quantified using a biophotometer (Eppendorf AG, Hamburg, Germany) and 100 ng of it was used as template for PCR amplifications.
PCR amplification for evaluation of allelic diversity
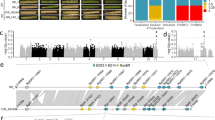
Primer pairs that defined markers targeting the promoter (Sasaki et al. 2006), coding (cleaved amplified polymorphic sequence—CAPS; Sasaki et al. 2004) and repetitive-indel regions of TaALMT1 (Raman et al. 2006; Fig. 1) were used to detect polymorphisms. Primers for the markers were synthesised from published sequences (Genbank accessions DQ072260, DQ072261 and AB243162). Sequences of the primers were: (1) long promoter fragment (LPF) marker 5′-CCTGGTTTTCTTGATGGGGGCACA-3′ (forward) and 5′-TGCCCACCATCTCGCCGTCGCTCTCTCT (reverse), (2) short promoter fragment (SPF) marker 5′-GCTCCTACCACTATGGTTGCG-3′ (forward) and 5′-CCAGGCCGACTTTGAGCGAG-3′ (reverse), (3) CAPS marker 5′-GGAATGGAATTCAACTGCTTTGGCG-3′ (forward) and 5′-TCCTCAGTGGCCTTCGAATTAAGG-3′ (reverse), (4) simple sequence repeats (SSR) marker (TaALMT1-SSR3a pair described by Raman et al. 2006) 5′-CTCGTGACAAAAGCCACTCA-3′ (forward) and 5′-GACGCAATCAAGGGGAATAA-3′ (reverse) and (5) indel marker (TaALMT1-SSR3b pair described by Raman et al. 2006) 5′-ATGCCATTTCTTCTGTACTGACA-3′ (forward) and 5′-AAAGAGTCCTCAGTGGCCTTCGAA-3′ (reverse). Amplifications were undertaken in 12.5 μL containing 1× reaction buffer, 0.3 μM of each primer, 0.5 units of Hot Start Taq™ DNA polymerase (Qiagen, Hilden, Germany) and 100 ng of gDNA in 96-well plates (Edwards Instruments, Australia). PCR cycling profiles for the TaALMT1 amplifications were as described elsewhere (Sasaki et al. 2004; Raman et al. 2006; Sasaki et al. 2006). The PCR amplifications were carried-out in GeneAmp® PCR System 2700 (Applied Biosystem, USA). The amplified fragments were resolved on 1% agarose gel containing ethidium bromide and run at 100 V for 1 and 1.5 h in 1× TAE buffer. Analysis of CAPS (Sasaki et al. 2004) and repetitive indel (Raman et al. 2006) markers was performed as described previously. Allele scoring and estimation of expected heterozygosity (polymorphism information content: PIC value) was calculated as described previously (Raman et al. 2003). Products from PCR that used LPF and SPF primer pairs in separate reactions (Fig. 2) identified a range of promoter alleles classified in to Types I–VI according to the size of products (Sasaki et al. 2006). Sequencing further identified Type I′ promoters which are variants of Type I with several SNPs (Sasaki et al. 2006) and are named here as Type I′a (Genbank accession: AB243169) and Type I′b (Genbank accession: AB243168).
Structure of TaALMT1 showing the approximate regions amplified by the various primers used as markers (thin black lines). Grey block arrow promoter region upstream of the first exon, white block arrows exons and black blocks introns. The hatched region within the promoter denotes a region where repeated blocks of sequence associated with Al resistance most commonly occur (Sasaki et al. 2006). The CAPS marker identifies one SNP in exon four while the SSR and indel markers both identify polymorphisms in the third intron. The thin line labelled SPF (short promoter fragment) and LPF (long promoter fragment) denote regions amplified by the promoter markers to identify the presence of repeated blocks of sequence. The vertical arrows indicate the locations where the polymorphisms occur for the SSR, indel and CAPS markers. The LPF primers identify most promoter types including Type III of Chinese Spring that encompasses a larger repeat than the hatched region. The SPF primers allow the Type I and II promoters, that only differ by 31 bp, to be distinguished from one another. The presence of various repeats determines the fragment sizes when the promoter marker primers are used to amplify genomic DNA as shown in Fig. 2
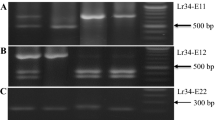
PCR fragments amplified from a range of wheat cultivars demonstrate the allelic variation in the promoter region of TaALMT1. The LPF primer pair A can detect all the promoter types except Types I and II which are distinguished with the SPF primer pair B. Genomic DNA was amplified with primers that span a region where repeated blocks of sequence occur (Fig. 1). For A the genotypes, promoter types and sizes of the amplified products are as follows: M markers (shown as kB); lane 1 ES8 (Type I; 1,190 bp ); lane 2 Currawong (Type IV; 1,470 bp); lane 3 Maringa (Type VI; 1,600 bp); lane 4 ET8 (Type V; 1,750 bp) and lane 5 Chinese Spring (Type III; 1,993 bp). For B the genotypes, promoter types and sizes of the amplified products are as follows: M markers (0.1 kb ladder); lane 1 ES8 (Type I, 612 bp) and lane 2 Trintecinco (Type II, 643 bp)
Sequencing of PCR products
Genomic DNA was isolated as described above from leaf tissue of a single seedling previously screened for Al resistance. PCR amplification was performed in a 50 μL reaction that contained 100 ng of genomic DNA template and appropriate primers, as described earlier. The PCR amplification was carried out for 15 min at 95°C (1 cycle), followed by 40 cycles of 45 s at 94°C, 45 s at 65°C and 1 min at 72°C, with a final 10 min incubation at 72°C. Amplification products were separated onto 1% TAE buffered agarose gels. The desired fragments were cut into slices and purified using a QIA gel extraction kit (Qiagen). Sequence analysis was performed by the dye termination method (Big dye, version1.1) using ABI prism 3,100 sequencer at AGRF Brisbane, Australia using 6 pmole of forward and reverse primers. Both the DNA strands of PCR products were sequenced directly from amplification products twice for quality control.
Linkage analysis
Segregation data obtained from the promoter marker were integrated with SSR markers that showed linkage with Al resistance in a previous study (Raman et al. 2005). Marker data were scored as “A” for maternal allele, “B” for paternal allele and “H” for heterozygous genotypes as appropriate. χ2 tests were undertaken to test the goodness of fit of marker and Al resistance segregation data to Mendelian ratios.
Results
Allele diversity at the TaALMT1 locus within cultivated wheat cultivars
Genomic DNA from 179 wheat cultivars (including Chinese Spring: a landrace) with known pedigrees derived from international breeding programs (Supplementary Table 1) was assessed for allelic diversity at the four TaALMT1 regions defined by separate markers (Fig. 1). Table 1 shows that within this wheat germplasm the number of alleles for the various markers ranged from two for the CAPS and indel markers to eight for the SSR marker. The PIC values of the LPF/SPF, SSR, CAPS and indel markers were 0.44, 0.52, 0.37 and 0.37, respectively and were correlated with the number of alleles.
The CAPS marker distinguished the two alleles (TaALMT1-1 and TaALMT1-2) found in the coding region but, as reported previously, neither of these was found to be consistently associated with Al resistance (Table 1; Sasaki et al. 2004; Raman et al. 2005). The indel marker identified positive and null alleles for the presence and absence of a 14 bp insertion in intron three and once again there was no relationship between this polymorphism and Al resistance. All the cultivars carrying the TaALMT1-1 allele, as identified by the CAPS marker, possessed the insertion and cultivars with the TaALMT1-2 allele did not, which is consistent with the close proximity of these markers (Fig. 1). The primers for the SSR marker amplified a fragment by PCR that ranged from 225 to 241 bp and identified seven alleles (Table 1). The promoter markers (LPF/SPF) identified six alleles (Table 1; Fig 2) as previously described by Sasaki et al. (2006). These alleles were generated by the presence of repeated blocks of sequence (see Fig. 2 in Sasaki et al. 2006). The Type II allele in the promoter region possesses a 31 bp duplication that can not be distinguished from the Type I allele with the LPF primers. Therefore, all cultivars scored as Type I with the LPF primers were subsequently re-analysed with the SPF primers which allowed these two promoter alleles to be distinguished from one another (Fig. 2b).
The relationships between alleles identified with each marker and Al resistance were established by hematoxylin staining. No single marker, including the repetitive indels and promoter markers, was able to identify all Al-resistant cultivars. Of the four markers tested, the promoter marker most closely predicted Al resistance. Of the 108 Al-resistant cultivars examined 100 (92.6%) possessed promoter Type V (Table 1; Supplementary Table 3) and 107, including those with the Type V promoter, had promoter types that contained one or more repeated blocks of sequence (Supplementary Table 2) consistent with a previous report that assessed a smaller group of cultivars (Sasaki et al. 2006). The single Al-resistant cultivar that lacked any repeats was Seneca (Type I′b) while Trintecinco (Type II; Table 1), also Al-resistant, possessed a relatively small repeat of only 31 bp. Although most cultivars with the 235 bp SSR marker were Al-resistant, many other Al-resistant cultivars possessed a 225 bp SSR fragment which was most common among Al-sensitive cultivars (Table 1).
Segregation of the TaALMT1 promoter alleles with Al resistance
Sasaki et al. (2006) previously showed a general correlation between promoter type and Al resistance within non-Japanese cultivars. Here we sought to establish whether the different promoter types could be used to track Al resistance in four wheat populations segregating for Al resistance. Two DH populations (Spica/Maringa and Currawong/CD87) were of particular interest because a previous study that scored the lines for Al resistance showed that the parental lines possessed the same TaALMT1 coding allele which prevented them from being differentiated with the CAPS marker (Raman et al. 2005, 2006). We found that the promoter alleles of the Al-resistant parents co-segregated with Al resistance within three doubled haploid mapping populations (Table 2) and an intercross population generated from ET8 (promoter allele Type V) and ES8 (promoter allele Type I). The F2:3 families of the ET8/ES8 population segregated in a 1:2:1 ratio for Type V homozygotes (16 families): Type V/Type I heterozygotes (36 families): Type I homozygotes (15 families) for a total of 67 families analysed and a χ2 (1:2:1) of 0.42 (P = 0.8–0.9). In all instances the Al resistance phenotypes of the F2:3 families determined previously (Sasaki et al. 2004), co-segregated with the Type V promoter allele.
Transmission and distribution of Al resistance
To trace the transmission of Al resistance, a selection of landraces was also analysed (Supplementary Table 2) which included samples of the major morphological groups/subspecies within T. aestivum. From this analysis, 22 haplotypes within cultivated genotypes and landraces were identified from the combinations of various alleles as identified by the four markers (Table 1). Some of the haplotypes may have arisen as a result of recombinations within the TaALMT1 locus. For example, the simplest explanation for the derivation of haplotypes 5 and 17 is that they resulted from a recombination event between the promoter and SSR regions of haplotypes 1 and 18 (or vice versa). Other haplotypes differed only in the length of the SSR marker (for example haplotypes 1–3) but were otherwise identical.
Haplotype 18 possessed the Type V promoter and was the most common of Al-resistant cultivars within the cultivated genotypes. This haplotype was present in the Brazilian cultivars Fronteira and Frontana that are thought to be an important source of Al resistance worldwide. However, the substantial differences in haplotype structure present in other Al-resistant cultivars suggest that Al resistance arose on more than one occasion. For instance, haplotypes 8, 10, 11, 13, 19 and 20 were also associated with Al-resistant cultivars although they were infrequent with each being represented by three or fewer cultivars in this study.
All promoter types associated with Al resistance in the cultivated wheat genotypes were also found within the landraces (Table 1; Fig. 3). Furthermore, sequence analysis identified three promoter types within the landraces not previously described. These promoter types were related to those already identified with one being a variant of Type I with a 2 bp deletion (Type I′c: Genbank accession EF446133) another being a variant of Type II with SNPs (Type II′: Genbank accession EF446134) and a third being a variant of Type VI in that it possessed two repeats instead of three (Type VII: Genbank accession EF446135). Haplotype 1, which was the most common haplotype in the landraces, was mostly associated with Al-sensitive accessions. However, a group of accessions with haplotype 1, predominately those originating from Nepal, were Al-resistant (Table 1; Fig. 3). A number of the haplotypes were common to both the landraces and cultivated genotypes (bold numbers on Table 1).
Geographical locations of TaALMT1 promoter alleles associated with Al resistance found in landraces of hexaploid wheat. Total number of accessions and number of Al-resistant accessions identified are given (e.g. India: 33, 4 represents 33 landrace accessions screened of which 4 were Al-resistant). Intermediate Al-resistant accessions (highlighted with asterisks in Supplementary Table 2) in addition to those reported by Stodart et al. (2007) are included as Al-resistant. Promoter types for the Al-resistant landraces are indicated in Roman numerals
Relationship of malate efflux to Al resistance
To determine whether the principal mechanism of Al resistance in the wheat cultivars examined here relied on malate efflux from root apices, a subpopulation of cultivars representing most of the different haplotypes was evaluated for Al resistance and malate efflux. Aluminum-resistant cultivars tended to release more malate than Al-sensitive cultivars and a positive correlation was observed between malate efflux and Al resistance among the 49 cultivars examined (r = 0.81; Fig. 4). This relationship held true for the Al-resistant cultivars Seneca and Trintecinco, which possessed promoter types (I and II) that are associated with low malate efflux in some cultivars (Sasaki et al. 2006).
Correlation between malate efflux from root apices and Al resistance determined by relative root growth in wheat cultivars. Relative root growth (RRG) of diverse wheat accessions representing a range of haplotypes at the TaALMT1 locus was determined for seedlings exposed to 80 μM AlCl3. The identity and malate efflux of individual cultivars is shown in Supplementary Fig. 1
Discussion
Allele diversity
Sasaki et al. (2006) previously reported the existence of six promoter types in the upstream region of the TaALMT1 gene based on fragment size and SNPs in PCR products. The alleles differ from one another in number and arrangement of tandem sequence repeats which are thought to influence the level of TaALMT1 expression and hence Al resistance. The origin of these tandem repeats is unclear but may have arisen by inadvertent replication of genomic DNA by the “rolling circle” machinery of viruses and transposons as suggested for the mlo locus in barley (Piffanelli et al. 2004). The relationship between Type V and Type VI promoters both of which possess three different but related repeats can be explained if the original replications that gave rise to these repeats occurred in opposite directions (Delhaize et al. 2007). Promoters that possess three tandem repeats but are otherwise identical to those with two tandem repeats could have arisen by unequal cross-over events during recombination. Here, we have identified these same promoter types and confirmed their identities by sequencing. We have extended the work of Sasaki et al. (2006) by using four different markers to assess the allelic variation of TaALMT1 in diverse hexaploid germplasm and identified three new promoter variants within landraces. Of the germplasm analysed here, all those possessing repeated blocks of sequence in their promoters were Al-resistant although a number of landraces lacking these repeats were also Al-resistant. The SSR marker also showed high allelic diversity both within cultivated germplasm and landraces, and identified eight alleles including all six described by Raman et al. (2006).
Transmittance of Al resistance
Most Al-resistant cultivars in Table 1 are grouped as haplotype 18. This group included Fronteira, the acid-soil resistant cultivar released in Brazil in 1932 (Supplementary Table 1; De Sousa 1998). Frontiera is the Al-resistant parent and Mentana the Al-sensitive parent of Frontana, which in turn, is the progenitor of many other Al-resistant cultivars developed at the Centro Internacional de Mejoramiento de Maíz y Trigo (CIMMYT). Hettel (1989) described that 30% of the wheat germplasm in CIMMYT includes Brazilian germplasm selected primarily for their Al resistance and disease resistance. The close interaction of breeding programs around the world with CIMMYT is likely to have resulted in the extensive use of Frontana and its derivatives, as a source of Al resistance. The predominance of this haplotype in Al-resistant cultivars around the world is consistent with the presence of Frontana and Fronteira in their pedigrees (Supplementary Table 1). One of the potential problems with germplasm evaluation is that it relies on correct labelling, documentation and maintenance of seed stocks. The finding that genotypes of the cultivars Gutha, Frontana, and Oasis existed as Al-resistant and Al-sensitive accessions (Supplementary Table 1) can also be explained by the expected genetic heterogeneity when a cultivar is developed. Varieties are typically derived from a single plant selection between F4 and F6, and at these generations an individual plant will be heterozygous at a small number of loci. This will subsequently result in a cultivar that consists of mixed but related genotypes. The Al-resistant accessions of these cultivars possessed haplotype 18, whereas the Al-sensitive accessions possessed haplotype 1 indicating mixed populations for the original sources of these accessions with the same cultivar name. In addition, some of the landrace accessions were also found to be segregating or mixed as would be expected for heterogenous populations (Supplementary Table 2).
Although haplotype 18 occurred in a large number of Al-resistant cultivars, it is also apparent that Al resistance in cultivated wheats based on TaALMT1 may have at least four other origins. By considering the different sequence patterns upstream of TaALMT1 (Sasaki et al. 2006), we conclude that the Al-resistant cultivars with promoter Types I′b (Seneca), II (Trintecinco), III (Chinese Spring), and VI (Maringa) are likely to have different origins from genotypes with promoter Type V (Sasaki et al. 2006). By contrast promoter Type IV is likely derived from Type V by unequal cross-over events as discussed above. The haplotype structures of these Al-resistant cultivars also differ from one another. Chinese Spring and Maringa have a 225 bp SSR marker and the null allele for the indel marker, whereas Seneca and Trintecinco have a 241 or 239 bp SSR marker and the positive allele for the indel marker (Table 1). Maringa has a Brazilian lineage but Chinese Spring does not. The unexpected finding is that Maringa (Type VI) and Toropi (Type VI) have Frontana (Type V) in their pedigrees even though they do not have a Type V promoter. This could be accounted for by the multiple sources of Al resistance used in early crosses in Brazilian wheat breeding. De Sousa (1998) explains that the Al-resistant genotypes Polyssú and Alfred Chaves 6–21 were likely to have both contributed to the development of cultivar Frontiera and subsequently Frontana. Furthermore, both Maringa and Toropi included other Al-resistant genotypes in their pedigrees (Petiblanco 8 and Ponta Grossa) and it is possible that one or other of these genotypes contributed the alternate Type VI promoter allele. Trintecinco was developed from different Al-resistant sources (Alfredo Chaves 3–21/Afredo Chaves 4–21) and this may explain it to be having the Type II promoter allele associated with its Al resistance. It could have been a matter of chance then that the Type V promoter finally dominated in the Al-resistant cultivars derived from Brazilian germplasm since all three promoter types are associated with similar levels of Al resistance. Alternatively, early breeders may have inadvertently favoured one allele over another if they were selecting for some other desirable phenotype linked to one of these alleles.
It is now clear that most of the promoter alleles associated with Al resistance pre-existed in landraces prior to the development of elite wheat varieties. For instance, the promoter alleles associated with Al resistance in Brazilian cultivars already existed within European landraces (Table 1; Fig. 3) and it is plausible that they were carried by Italian and Portuguese immigrants who settled in Brazil. The subsequent strong selection pressure for Al resistance on Brazilian soils ensured that these alleles predominated in subsequent cultivar development. The presence of Al resistance in non-Brazilian cultivars such as Bencubbin (Australia); Chinese Spring (China); Seneca (USA but derived from a Mediterranean variety) and landraces from Bulgaria, Croatia, India, Italy, Nepal, Spain, Tunisia and Turkey (Stodart et al. 2007) also support the notion that resistance had arisen prior to the severe selection pressure applied in South America. Although the Type I promoter was predominately associated with Al-sensitive genotypes, a subpopulation of Al-resistant landraces throughout these regions also possessed the Type I promoter. Preliminary analysis of these Al-resistant genotypes indicates that the resistance mechanism relies on Al-activated efflux of malate (data not shown) as found for the cultivar Seneca which also possesses the Type I promoter (Supplementary Fig. 1). The association of Type I promoters with Al resistance may result if elements that enhance TaALMT1 expression in these genotypes occur further upstream to the region analysed or if unlinked loci encode trans-acting factors that interact with the TaALMT1 promoter to increase expression level. In addition, although Al resistance conditioned by TaALMT1 appears to be the predominant mechanism in cultivated hexaploid wheat, it is also conceivable that different genes control Al resistance within some landraces with the Type I promoters.
The presence of several promoter types within the spelts (non-free-threshing hexaploids) is also of interest. Molecular evidence indicates that the spelts are derived from hybridisation events between non-free-threshing tetraploid wheat (2N = AABB) and free-threshing hexaploid wheat (Blatter et al. 2004). Furthermore, analysis of the A and B genomes of spelt indicates that this type of hybridization has occurred on several occasions (Blatter et al. 2004). Since TaALMT1 is located on the D-genome, the presence of several different promoter types (I, IV, and VI) in the spelts with identical sequence to those found in cultivated common wheats is consistent with the idea that the spelts arose from several independent hybridizations. Two of these hybridizations involved promoter alleles associated with Al resistance which in the landraces are either absent (promoter Type VI) or relatively infrequent (promoter Type IV; Table 1; Supplementary Table 2). This suggests that the hybridization events that gave rise to the Al-resistant spelts originated on acid soils where the Al-resistant landraces would have predominated.
Utility of markers for breeding
We have used PCR-based markers that target different regions of the TaALMT1 gene. The CAPS and indel markers have limited utility in only being able to identify two alleles neither of which is diagnostic of Al resistance, and in the case of the CAPS marker, requires an additional enzyme digestion step. Raman et al. (2006) discussed the advantages of different marker types in MAS for Al resistance and identified PCR-based markers such as those targeting SSRs within the TaALMT1 gene as being suitable for high throughput screening and multiplexing. Previously described SSR markers are not based on TaALMT1 and map at least 2 cm apart from the Al resistance locus (Rodriguez-Milla and Gustafson 2001; Ma et al. 2005; Raman et al. 2005). The repetitive indel SSR marker (Raman et al. 2006) is based on the TaALMT1 gene but is prone to “stuttering” and small variations (1–4 bp) prevalent in mono- and di-nucleotide repeats in the intron three SSR region can be difficult to resolve where the differences between the parental genotypes are small (Raman et al. 2006). In many cases the SSR fragments from intron three of TaALMT1 differ by 10 bp or more (Table 1) thus avoiding this problem. Despite the advantages of the TaALMT1-based SSR marker it cannot be used in instances where the alleles do not differ sufficiently. For instance, the SSR marker was not polymorphic between the parental cultivars of the DH lines derived from Spica/Maringa and Currawong/CD87, whereas both of these populations could be scored with the promoter marker.
The marker based on the promoter region is not a “universal” marker in the sense that it is able to identify all Al-resistant types. This is illustrated by those Al-resistant cultivars with Type I and II promoter alleles mostly associated with Al sensitivity. Furthermore, Sasaki et al. (2006) identified Al-sensitive Japanese cultivars with promoter types usually associated with resistance (for example, Kitakami Komugi and Kitakei 1354). Nevertheless, the PCR marker based on the promoter region is codominant, highly polymorphic, amenable for multiplexing and targets the region of TaALMT1 that is thought to control the level of Al resistance in many genotypes.
References
Basu U, McDonald-Stephens JL, Archambault DJ, Good AG, Briggs KG, Taing-Aung, Taylor GJ (1997) Genetic and physiological analysis of doubled-haploid, aluminium-resistant lines of wheat provide evidence for the involvement of a 23 kD, root exudate polypeptide in mediating resistance. Plant Soil 196:283–288
Berzonsky WA (1992) The genomic inheritance of aluminium tolerance in “Atlas 66” wheat. Genome 35:689–693
Blatter RHE, Jacomet S, Schlumbaum A (2004) About the origin of European spelt (Triticum spelta L.): allelic differentiation of the HMW Glutenin B1–1 and A1–2 subunit genes. Theor Appl Genet 108:360–367
De Sousa CNA (1998) Classification of Brazilian wheat cultivars for aluminium toxicity in acid soils. Plant Breed 117:217–221
Delhaize E, Craig S, Beaton CB, Bennet RJ, Jagadish VC, Randall PJ (1993a) Aluminum tolerance in wheat (Triticum aestivum L.) I. Uptake and distribution of aluminum in root apices. Plant Physiol 103:685–693
Delhaize E, Ryan PR, Randall PJ (1993b) Aluminum tolerance in wheat (Triticum aestivum L.) II. Aluminum stimulated excretion of malic acid from root apices. Plant Physiol 103:695–702
Delhaize E, Ryan PR, Hebb DM, Yamamoto Y, Sasaki T, Matsumoto H (2004) Engineering high level aluminum tolerance in barley with the ALMT1 gene. Proc Natl Acad Sci USA 101:15249–15254
Delhaize E, Gruber BD, Ryan PR (2007) The roles of organic anion permeases in aluminium resistance and mineral nutrition. FEBS Lett 581:2255–2262
Garvin DF, Carver BF (2003) Role of the genotype in tolerance to acidity and aluminium toxicity. In: Rengel Z (ed) Handbook of soil acidity. Dekker, New York, pp 387–406
Hettel GP (1989) Wheat production advances in South America’s Colossus: the gains from 20 years of Brazilian/CIMMYT collaboration. CIMMYT, Mexico
Kerridge PC, Kronstad WE (1968) Evidence of genetic resistance to aluminum toxicity in wheat (Triticum aestivum Vill., Host). Agron J 60:710–711
Luo M-C, Dvorák J (1996) Molecular mapping of an aluminum tolerance locus on chromosome 4D of Chinese Spring wheat. Euphytica 91:31–35
Ma H-X, Bai G-H, Carver BF, Zhou L-L (2005) Molecular mapping of a quantitative trait locus for aluminum tolerance in wheat cultivar Atlas 66. Theor Appl Genet 112:51–57
Piffanelli P, Ramsay L, Waugh R, Benabdelmouna A, D’Hont A, Hollricher K, JØrgensen JH, Schulze-Lefert P, Panstruga R (2004) A barley cultivation-associated polymorphism conveys resistance to powdery mildew. Nature 430:887–891
Polle E, Konzak CF, Kittrick JA (1978) Visual detection of aluminum tolerance levels in wheat by hematoxylin staining of seedling roots. Crop Sci 18:823–827
Raman H, Moroni JS, Sato K, Read BJ, BJ Scott (2002) Identification of AFLP and microsatellite markers tightly linked with an aluminium tolerance gene in barley. Theor Appl Genet 105:458–464
Raman H, Karakousis A, Moroni JS, Raman R, Read B, Garvin DF, Kochian LV, Sorrells ME (2003) Development and allele diversity of microsatellite markers linked to the aluminium tolerance gene Alp in barley. Aust J Agri Res 54:1315–1321
Raman H, Zhang K, Cakir M, Appels R, Garvin DF, Maron LG, Kochian LV, Moroni JS, Raman R, Imtiaz M, Drake-Brockman F, Waters I, Martin P, Sasaki T, Yamamoto Y, Matsumoto H, Hebb DM, Delhaize E, Ryan PR (2005) Molecular mapping and characterization of ALMT1, the aluminium-tolerance gene of bread wheat (Triticum aestivum L.). Genome 48:781–791
Raman H, Raman R, Wood R, Martin P (2006) Repetitive indel markers within the ALMT1 gene controlling aluminium tolerance in wheat (Triticum aestivum L). Mol Breed 18:171–183
Riede CR, Anderson JA (1996) Linkage of RFLP markers to an aluminum tolerance gene in wheat. Crop Sci 36:905–909
Rodriguez Milla MA, Gustafson JP (2001) Genetic and physical characterization of chromosome 4DL in wheat. Genome 44:883–892
Ryan PR, Delhaize E, Randall PJ (1995a) Malate efflux from root apices and tolerance to aluminium are highly correlated in wheat. Aust J Plant Physiol 22:531–536
Ryan PR, Delhaize E, Randall PJ (1995b) Characterisation of Al-stimulated efflux of malate from the apices of Al-tolerant wheat roots. Planta 196:103–111
Ryan PR, Skerrett M, Findlay G, Delhaize E, Tyerman SD (1997) Aluminium activates an anion channel in the apical cells of wheat roots. Proc Natl Acad Sci USA 94:6547–6552
Sasaki T, Yamamoto Y, Ezaki B, Katsuhara M, Ahn SJ, Ryan PR, Delhaize E, Matsumoto H (2004) A wheat gene encoding an aluminum-activated malate transporter. Plant J 37:645–653
Sasaki T, Ryan PR, Delhaize E, Hebb DM, Ogihara Y, Noda K, Matsumoto H, Yamamoto Y (2006) Analysis of the sequence upstream of the wheat (Triticum aestivum L.) ALMT1 gene and its relationship to aluminium tolerance. Plant Cell Physiol 47:1343–1354
Somers DJ, Gustafson JP (1995) The expression of aluminum stress induced polypeptides in a population segregating for aluminum tolerance in wheat (Triticum aestivum L.). Genome 38:1213–1220
Somers DJ, Briggs KG, Gustafson JP (1996) Aluminum stress and protein synthesis in near isogenic lines of Triticum aestivum differing in aluminum tolerance. Physiol Plant 97:694–700
Stodart BJ, Raman H, Coombes N, Mackay M (2007) Evaluating landraces of bread wheat for tolerance to aluminium under low pH. Genet Resour Crop Evol 54:759–766
Tang Y, Garvin DF, Kochian LV, Sorrells ME, Carver BF (2002) Physiological genetics of aluminum tolerance in the wheat cultivar Atlas 66. Crop Sci 42:1541–1546
Whiting D (2001) In: Wheat varieties in Australia 1968–2001, pp 130, compiled by Rural Solutions SA, 35 Frances Tce, Kadina SA 5554
Yamaguchi M, Sasaki T, Sivaguru M, Yamamoto Y, Osawa H, Ahn SJ, Matsumoto H (2005) Evidence for the plasma membrane localization of Al-activated malate transporter (ALMT1). Plant Cell Physiol 46:812–816
Zhang W-H, Ryan PR, Tyerman SD (2001) Malate-permeable channels and cation channels activated by aluminum in the apical cells of wheat roots. Plant Physiol 125:1459–1472
Zhou L-L, Bai G-H, Ma H-X, Carver BF (2007) Quantitative trait loci for aluminum resistance in wheat. Mol Breed 19:153–161
Acknowledgments
This work was supported by the NSW Agricultural Genomics Centre of BioFirst Initiative of NSW Government of Australia and CSIRO Plant Industry. We thank Dr JS Moroni, Ms Donna Seberry, and Mrs Fiona Wray for their help in Al resistance screening. The Spica/Maringa mapping population was provided by Dr Daryl Mares from University of Adelaide, Australia.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Bohn
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Raman, H., Ryan, P.R., Raman, R. et al. Analysis of TaALMT1 traces the transmission of aluminum resistance in cultivated common wheat (Triticum aestivum L.). Theor Appl Genet 116, 343–354 (2008). https://doi.org/10.1007/s00122-007-0672-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-007-0672-4