Abstract
In strawberry (Fragaria × ananassa Duch.), auxin has been recognized as the main signal molecule coordinating the growth and initiation of ripening of fruits. The molecular mechanism regulating auxin biosynthesis in strawberry remains unknown. This project reports two YUCCA flavin monooxygenase genes FaYUC1-2 isolated from cultivated strawberry. FaYUC1 and FaYUC2 are most homologous to AtYUC6 and AtYUC4, respectively. Significant expression of FaYUC1-2 is found in vegetative meristems and reproductive organs, with overlapping but distinct patterns. During fruit development, both transcripts of FaYUC1 and FaYUC2 in achenes reach a peak around large green fruit (G2) stage, but the sudden rise in FaYUC2 transcript level is much steeper and begins earlier than that in FaYUC1. FaYUC2 is also obviously expressed in the receptacles from green fruits, hinting another auxin source for receptacle development, other than achenes. FaYUC1 over-expression Arabidopsis exhibits typical auxin hyper-accumulation phenotype in many aspects, such as the narrow and downward curled leaves, strong apical dominance, short and hairy root. It is also severely sterile, due to the disruption of floral meristems initiation and floral organs development. Transgenic analysis indicates that strawberry YUC gene may hold conserved role in auxin biosynthesis like their homologs in other plants. Integrated with the spatiotemporal expression features, these results led us to propose that FaYUC1-2 may involve in many developmental processes including flower and fruit development in strawberry.
Key message This paper is the first report of isolation and characterization of strawberry auxin biosynthesis genes. And their conserved functions in auxin biosynthesis were confirmed after ectopic expression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Auxin is involved either directly or indirectly in every aspect of plant development, virtually playing the integrative role in coordinating plant growth and development (Delker et al. 2008; Leyser 2010, 2011). It is required for initiation and polar growth of all organ primordia: for leaf and lateral root during vegetative development, and for floral meristems and floral organs during reproductive development (Davies 2004; Teale et al. 2006). Particularly, auxin plays a key role in fruit initiation, seed development, and fruit ripening (Pandolfini et al. 2007; Sundberg and Østergaard 2009). Auxin exerts its effects both locally and distally. Recent findings show that localized auxin biosynthesis, in particular, plays essential roles in many developmental processes including gamogenesis, embryogenesis, seedling growth, vascular patterning, and flower development (Zhao 2010).
Auxin biosynthesis occurs via several interconnecting pathways. So far, no single complete pathway for de novo auxin biosynthesis in plants has been firmly established. However, recent studies have led to the discoveries of several genes in tryptophan-dependent pathways. Of these, YUCCA (YUC) genes are key auxin biosynthesis genes, which encode flavin monooxygenases (FMO) catalyzing a rate-limiting step in a tryptophan-dependent auxin biosynthesis pathway (Zhao et al. 2001). The YUC genes appear to have a broad existence and be highly conserved throughout the plant kingdom. Functional characterization of some YUC genes in Arabidopsis (Cheng et al. 2006; Kim et al. 2007, 2011; Zhao et al. 2001), petunia (Tobeña-Santamaria et al. 2002), rice (Woo et al. 2007; Yamamoto et al. 2007), maize (Gallavotti et al. 2008), and tomato (Expósito-Rodríguez et al. 2007, 2011) indicates that they play similar roles in auxin biosynthesis during plant development.
The strawberry genus, Fragaria, belongs to the Rosaceae family, an economically important plant family including many fruit tree crops (apple, pear, peach, plum, apricot, cherry), herbaceous fruit plants (strawberry, brambles), and ornamental plants (rose, Photinia serrulata, Chinese flowering crab apple). Cultivated strawberry (Fragaria × ananassa Duch.) is a highly valued fruit crop species. Genomically, F. × ananassa is among the most complex of crop plants, harboring eight sets of chromosomes (2n = 8x = 56) derived from as many as four different diploid ancestors (Folta and Davis 2006). However, genomes of octoploid and diploid Fragaria species share a high level of macrosynteny and colinearity (Rousseau-Gueutin et al. 2008). In addition, both the octoploid and diploid strawberry are amenable to genetic transformation, small in size, with rapid growth and generous seed set (Folta and Davis 2006; Shulaev et al. 2008). Furthermore, the genome of the diploid woodland strawberry is available recently (Shulaev et al. 2011). These properties render strawberry an attractive surrogate for testing gene function for many plants in the Rosaceae family.
Botanically, the strawberry fruit is actually a ‘false fruit’, a modified receptacle with one-seeded fruits (achenes) located on the surface in spirally arranged rows. Nitsch (1950) first demonstrated that achenes control the growth of strawberry fruit. This control is exerted through auxin, and the concentration of auxin in the achenes varies greatly with the stage of development (Nitsch 1950, 1955). It was revealed that the decline in auxin concentration in the achenes as fruits mature modulates the rate of strawberry fruit ripening (Given et al. 1988). Later, auxin has been recognized as the main signal molecule coordinating the growth and initiation of ripening in strawberry fruit (Perkins-Veazie 1995). Fruit ripening processes that appear to be auxin dependent include pathways related to pigmentation, stress/defense, cell wall metabolism, cell structure, fatty acid metabolism, and flavor/aroma synthesis (Aharoni et al. 2002). For example, expression of the flavor compound biosynthesis-related gene FaQR1 correlates inversely with auxin levels (Raab et al. 2006). It has been reported that transgenic modification of the DefH9-iaaM auxin-synthesizing gene greatly improved fruit yield in both wild and cultivated strawberry plants (Mezzetti et al. 2004). However, the knowledge about molecular mechanism of auxin biosynthesis is entirely elusive in strawberry.
To bridge this gap, we had sought to isolate and characterize the genes related to auxin biosynthesis in strawberry long before the genome of woodland strawberry being available. Here, we report that two YUC genes FaYUC1-2 were isolated from cultivated strawberry and characterized in terms of their structural, phylogenic, and transcriptional features. The conserved function of strawberry YUC gene in auxin biosynthesis was further exemplified via ectopic expression of FaYUC1 in Arabidopsis. Our results suggest that FaYUC1-2 may involve in many developmental processes including flower and fruit development in strawberry.
Materials and methods
Plant materials and growth conditions
The elite strawberry cultivar ‘Jiuxiang’ (Fragaria × ananassa Duch. cv. Jiuxiang), used in this study was the crossing progeny of ‘Kunouwase’ and ‘Toyonoka’, bred by the Forest and Fruit Tree Research Institute (FFRI) of Shanghai Academy of Agricultural Sciences (SAAS) (Chinese New Variety Right No. CNA004064E). ‘Jiuxiang’ strawberry was grown in a plastic-mulched cultivation system located at the trial station of SAAS in Zhuanghang Town, Fengxian District, Shanghai. In this system, ‘Jiuxiang’ strawberry produces fruits from first December to late May the next year. For expression analysis in different organs, sampling of young roots (about 1 cm in length, root apical tips from mature fruiting plants), young stolons (about 1 cm in length, the young apical stolon tips), shoot apical meristems (SAM, about 0.5 cm in length), fully expanded functional leaves, full-blown flowers (including sepals), and ripe fruits (completely red ripe fruits, receptacles with achenes) was independently conducted three times in January 2012. In particular, flowers and fruits were selected and dissected at eight consecutive developmental stages: F1, young flowers before pollination (floral buds); F2, mature flowers with partially withered petals (about 4–7 days after pollination); G1, small green fruits (about 1–2 cm in diameter); G2, large green fruits (about 2.5–3.5 cm in diameter); W1, white receptacles with green achenes; W2, white receptacles with some rosy and brown achenes; T, turning fruits being half white and half red; R, completely ripe fruits (Fig. 1). The flowers and fruits of eight stages were concurrently sampled on February 18th, which was independently repeated on April 2, 2011.
The fruit development process of ‘Jiuxiang’ (Fragaria × ananassa Duch. cv. Jiuxiang). This process is divided into eight consecutive stages: F1 floral bud stage, F2 mature flower stage (about 4–7 days after pollination), G1 small green fruit stage, G2 large green fruit stage, W1 white receptacle with green achenes stage, W2 white receptacle with some rosy and brown achenes stage, T turning stage, R ripe stage. Scale bars 1 cm
Cloning of FaYUC1-2 genes
Twelve pairs of degenerate primers (Table 1) composed of three sense primers in FAD-binding site and four antisense primers in NADP binding site of YUCCA FMOs were used to amplify the conserved regions of strawberry YUCCA genes with mixed cDNAs templates from SAMs and green fruits. Based on the partial cDNA clones for strawberry YUC, rapid amplification of cDNA ends (RACE) was performed to isolate the full-length cDNAs. The 3′ cDNA ends were isolated following the CLONTECH SMART™ RACE protocol (Clontech, USA, PT3269-1), and the 5′ cDNA ends were isolated using the Takara 5′-FULL RACE Kit (Takara, Dalian, China, CAT#D315). Accordingly, primersets (Table 1) spanning full open reading frames (ORFs) were designed to directly amplify full-length clones from cDNA and DNA templates, respectively. All amplicons were cloned into the pUCm-T vector (Sangon, Shanghai, China, SK2212) for sequencing.
RNA extraction and reverse-transcription PCR (RT-PCR)
Total RNA was isolated from strawberry tissues with a CTAB extraction buffer following a previously reported method (Chang et al. 1993). The quality and quantity of RNA extracts were evaluated using the Nanodrop ND-1000 Spectrophotometer (Thermo Scientific, Wilmington, USA) and visualized by 1 % agarose gel electrophoresis under denaturing conditions.
Total RNA samples were treated with DNaseI (Fermentas, Burlington, CA, Cat.EN0521) at 37 °C for 30 min to remove genomic DNA contamination, and then were utilized to synthesize the first-strand cDNAs using M-MLV reverse transcriptase (Promega, Madison, USA, Cat. M170A).
For expression analysis of FaYUC genes in different organs and during fruit development, quantitative RT-PCR was conducted in an ABI 7500 Real-Time PCR System (Applied Biosystems, Forster City, USA) using a SYBR Premix Ex Taq™ (Perfect Real Time) Kit (Takara, Dalian, China, DRR041A). Fluorescence data collected during the 72 °C step were analyzed using ABI 7500 analysis software. The amount of cDNA templates in each sample was normalized using the amplification of FaActin. Annealing temperature for all RT-PCR primersets (Table 1) was 58 °C. In each biological test, three repeats were used for expression analysis of certain gene in one sample (PCR template). Two or three biological replicates were performed.
Construction of 35S-FaYUC1 plasmid
A 1,315-bp fragment spanning FaYUC1 ORF was amplified from ‘Jiuxiang’ strawberry cDNAs with the primer pair Fyu1-F (5′-CCAACATGGACTACACAACAATACAAGG-3′, underlined: the start code) and Fyu1-S (5′-CACTATGGCGACGACGATAGAG-3′, underlined: the terminal code) using Ex-Taq DNA polymerase (Takara, Dalian, DRR100C). The product was cloned into the pUCm-T vector and sequenced for confirmation. After the T-vector with proper insertion orientation was selected via the release of a 1,087-bp fragment by EcoRI digestion, FaYUC1 cDNA was transferred from pUCm-T to the pCambia1300-mod vector (Duan et al. 2008) between the KpnI and BamHI sites downstream of the 35S cauliflower mosaic virus promoter. Gene-specific PCR combined with restriction digestion analysis was used to confirm the pCambia1300-mod vector harboring 35S:FaYUC1. Agrobacterium tumefaciens strain EHA105 was used as host for the resultant construct.
Arabidopsis thaliana transformation and phenotypic analysis
The above construct harboring 35S:FaYUC1 was transformed into wild type Col-0 Arabidopsis plants using a floral-dip method (Clough and Bent 1998). The seeds of transgenic plants were selected on solid half MS medium containing 20 mg L−1 hygromycin (Hyg). Plates were sealed with parafilm and placed vertically in a growth chamber (SANYO, Japan, MLR-351H), with a 16-h light/8-h dark cycle, 120 μmol m−2 s−1 photo flux density, at 22 ± 2 °C for 10 days before transplanting to soil. Ectopic expression of strawberry YUCCA gene was confirmed via RT-PCR using gene-specific primers (see Supplementary Fig. S1). Whole plants were photographed with a digital camera (Fujifilm, FinePix S602 Zoom, Japan). Young seedlings of wild type and transgenic plants were viewed with a dissecting microscope (Shanghai Optical Instrument Factory, XTL3400), and image data were collected with a digital camera (Panasonic, DMC-LX3, Japan) under light-field.
Results
Isolation of the full-length cDNA and DNA clones for FaYUC1-2
Two cDNA fragments (527 and 506 bp, respectively) spanning the conserved region of YUCCA-type genes were isolated from the mixed cDNAs from green fruits and SAMs. Based on these partial cDNA clones, RACE experiments were performed and led to the isolation of the corresponding full-length cDNA (1,315 and 1,235 bp) and DNA (2,552 and 2,442 bp) clones. The 1,315 bp cDNA fragment harboring a 1,308 bp ORF encoding a 435-aa peptide was named FaYUC1, while the 1,235 bp cDNA with an 1,224 bp ORF encoding a 407-aa peptide named FaYUC2. These two genes were deposited into GenBank under the accession number JF898837 (FaYUC1) and JF898838 (FaYUC2), respectively.
Structural and homologous characterization of FaYUC1-2
Gene structural analysis shows that both FaYUC1 and FaYUC2 consist of four exons and three introns. The position and size of the four exons are fairly well conserved (Fig. 2a). This pattern of exon–intron organization is quite similar to that in Arabidopsis YUC1, YUC2, YUC4, YUC6, and YUC10. The obvious divergence between the structures of FaYUC1 and FaYUC2 is the relative lengths of their first and third introns. The general structure of FaYUC1 is most similar to that of AtYUC6, with the first intron being the largest. By contrast, the structure of FaYUC2 is most similar to that of AtYUC4, with the third intron being the largest.
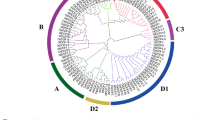
Structural and phylogenic analysis of FaYUC1-2. a Intron (white box) and exon (black box) organization of FaYUC1-2. The numbers indicate the length in base pairs (bp) of the corresponding exons or introns. b Distribution and alignment of various motifs in FaYUC1-2, AtYUC1, AtYUC4, and AtYUC6 proteins. The exact amino acid numbering and the consensus are shown in the parentheses and on the top of the alignment, respectively. Conserved residues identical to the consensus are shaded in black (conserved in all FMOs) or in gray (additional residues conserved among Arabidopsis YUC proteins). c Phylogenic tree of FaYUC1-2 and homologous proteins from other plants. Multiple sequence alignment was first performed using the MUSCLE program at http://www.ebi.ac.uk/Tools/msa/muscle/ (Chenna et al. 2003). Based on the alignment result, phylogenic tree was generated with MEGA ver. 5.02 software (Tamura et al. 2011). In addition to FaYUC1-2, other YUCCA proteins shown are YUC1-11 (AT4G32540, AT4G13260, AT1G04610, AT5G11320, AT5G43890, AT5G25620, AT2G33230, AT4G28720, AT1G04610, AT1G48910 and AT1G21430) from Arabidopsis, OsYUC1-7 (BAD68007, AAU43964, BAD87432, BAD 32703, ABA99096, BAC80117, CAE76085) from rice, and two putative VvYUCs (XP 002281015 and XP 002281597) from grape
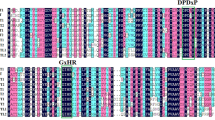
Pairwise comparison of the deduced protein sequences revealed that FaYUC1 and FaYUC2 share 61.1 % identity to each other. In addition, FaYUC1 shares 75.4 and 61.5 % identity to AtYUC6 and AtYUC2, respectively. FaYUC2 shows 74.1 and 68.8 % identity to AtYUC4 and AtYUC1, respectively. Sequence alignment identified the highly conserved motifs of plant YUCCA-type FMOs (Hou et al. 2011; Schlaich 2007) in FaYUC1-2 (Fig. 2b). The FAD-binding motif near the N terminus and the NADPH-binding motif in the middle are completely identical among FaYUC1-2 and their Arabidopsis homologs. The GC motif near the FAD-binding site, which has been suggested to further stabilize FAD binding, is nearly identical in FaYUC1 and AtYUC6. The ATG-containing motif 2 at the C-terminal region, which may link the NADPH site to the active site, is the same in FaYUC2 and AtYUC4. The consensus residues in ATG-containing motif 1 (the putative linker between the FAD site and NADPH site) and in the FMO-identifying sequence motif near the NADPH motif (which is suggested to contribute to NADPH binding) are conserved in all present YUC FMOs (Fig. 2b).
To determine the evolutionary relationships among FaYUC1-2 and their homologs from grape, rice and Arabidopsis, we performed phylogenic analysis using a neighbor-joining method (Fig. 2c). FaYUC1 is revealed to be most homologous to AtYUC6. This protein together with AtYUC6 and AtYUC2 from Arabidopsis, OsYUC2-4 from rice, and two grape putative YUC proteins, constitute one group. By contrast, FaYUC2, most homologous to AtYUC4, is included in another group which contains AtYUC1, AtYUC4, OsYUC1 and OsYUC5 (Fig. 2c). The phylogenic relationships between FaYUC and Arabidopsis YUC proteins are coincident with the conservation in their exon–intron organization.
The spatiotemporal expression patterns of FaYUC1 and FaYUC2 in Fragaria × ananassa Duch. cv. Jiuxiang
The expression features of FaYUC1-2 in different organs were investigated by quantitative RT-PCR (Fig. 3). The results show that FaYUC1 is relatively preferentially expressed in the apical meristems from both roots and shoots (SAM), moderately expressed in young stolon tips and full-blown flowers. Transcripts of FaYUC1 were not detected in mature leaves and ripe fruits. By contrast, FaYUC2 was found to be expressed in all tissues examined. The transcript level of FaYUC2 is moderate in mature leaves and flowers, higher in SAMs, and highest in young stolon tips. This gene is also weakly expressed in root meristem tips and ripe fruits.
Quantitative real-time PCR analysis of FaYUC1 and FaYUC2 transcripts in different tissues from strawberry (Fragaria × ananassa Duch. cv. Jiuxiang). Amplification of FaActin transcripts was used for internal control. Root young roots (about 1 cm apical tip) from mature fruiting plants, leaf fully expanded functional leaves, SAM shoot apical meristems (about 0.5 cm upper part of the primary shoot), stolon young part (about 1 cm apical part) of the stolons, without leaves, flower full-blown flowers (including sepals), fruit completely ripe fruits (receptacles with achenes)
During fruit development (Fig. 1), dynamic expression of FaYUC1-2 in the receptacles and the achenes was investigated via quantitative real-time PCR (Fig. 4). It should be noted that the results are expressed in the transcription levels relative to that in whole floral buds before pollination (F1 stage) for each gene. In achenes, accumulation of two FaYUC genes is dynamically changed and reaches a peak around G2 stage (large green fruits). However, obvious divergences exist in their expression profiles during fruit development. In achenes, the sudden rise in their transcript levels occurs at different stage and is not as steep. The FaYUC1 transcript level is lower in mature flowers (F2 stage) and small green fruits (G1 stage), in comparison to that in floral buds (F1 stage) (Fig. 4a). FaYUC1 transcript level is moderately increased to the peak at G2 stage, approaching 2.3-fold of that at F1 stage. Since the W1 stage, FaYUC1 transcription is significantly reduced, even reaching 10 % at T stage. By contrast, the prominent feature for FaYUC2 transcript accumulation is the very steep increase occurring at the G1 stage (small green fruits) (Fig. 4c). This expression soon reaches a peak at G2 stage, when the achenes of the strawberry fruits contain 110 times more FaYUC2 transcripts than does the floral buds before pollination (F1 stage). Later, accumulation of FaYUC2 transcripts is decreased and fluctuated, but still obviously present in achenes of ripe fruits (R stage).
Quantitative real-time PCR analysis of FaYUC1-2 transcript level in achenes and receptacles during strawberry fruit development. a Dynamic expression of FaYUC1 in achenes relative to that in floral buds. b Dynamic expression of FaYUC1 in receptacles relative to that in floral buds. c Dynamic expression of FaYUC2 in achenes relative to that in floral buds. d Dynamic expression of FaYUC2 in receptacles relative to that in floral buds. The amount of cDNA templates in each sample was normalized by the amplification of FaActin. Y axis in all panels indicates gene expression levels relative to that in floral buds before pollination (F1 stage). X axis in all panels shows developmental stages (as indicated in Fig. 1). The achenes and receptacles of 5–10 fruits at certain developmental stages were dissected for RNA extraction and expression analysis. The experiment was independently conducted twice. Errors bars represent the standard deviations within the repeats. Note that different scales are used for different Y axes
Expression levels of both two FaYUC genes are very low in receptacles (Fig. 4b, d). FaYUC1 was only examined in receptacles from mature flowers (F2 stage), but FaYUC2 was obviously detected in receptacles from fruits at F2, G1 and G2 stages.
FaYUC1 over-expression plants share phenotypes typical of auxin hyper-accumulation
All FaYUC1 over-expression plants display long and narrow rosette leaves with elongated petioles (Fig. 5a). The cauline leaves in most transgenic plants are also long and narrow with downward curled edges (Fig. 5b). Although FaYUC1 over-expression does not uniformly increase plant height, all transgenic plants show strong apical dominance (Fig. 5c). Notably, the strong apical dominance is accompanied by severe disruption of reproductive development in most conditions (Fig. 5d, e). Indeed, total seven lines of independent transgenic plants over-expressing FaYUC1 are obtained, but only one of these plants is partially fertile, while the remaining six lines are completely sterile.
FaYUC1 over-expression plants display severe abnormality in both vegetative and reproductive organs. a Rosette leaves from wild type (Col) and FaYUC1 over-expression (FaYUC1-OX) plants. b Cauline leaves from wild type and FaYUC1-OX plants. c 1.5-month-old wild type and FaYUC1-OX plants (two independent transgenic lines). d The upper parts of main shoots from wild type and FaYUC1-OX plants. e The mature siliques from wild type or similar structures from FaYUC1-OX plants. T1 progeny of 35S:FaYUC1 transgenic plants is shown
The progeny of the unique fertile transgenic plant is further studied along with wild type control. Under normal light conditions, 7-day-old transgenic seedlings display epinastic cotyledons as well as short and hairy roots (Fig. 6a). Short, hookless, etiolated hypocotyls are found in 3-day-old transgenic seedlings under dark conditions (Fig. 6b). The differences between wild type and FaYUC1 over-expression seedlings could be phenocopied by exogenous 2,4-D (a synthetic auxin) treatment on wild type (see Supplementary Fig. S2).
Over-expression of FaYUC1 results in stocky and hairy phenotype in seedlings under both light and dark conditions. a Wild type (Col) and FaYUC1-OX seedlings grown on half MS medium for 7 days under normal light conditions. b Wild type (Col) and FaYUC1-OX seedlings grown on half MS medium for 3 days under dark conditions. T1 progeny of 35S:FaYUC1 transgenic plants is studied
Discussion
A new insight into plant auxin biosynthesis emerged when two dominant Arabidopsis mutants, yucca, were isolated by activation tagging screen, and an auxin biosynthesis pathway was proposed, in which the YUC gene is a rate-limiting step (Zhao et al. 2001). Using a homologous cloning strategy, two YUCCA-like genes FaYUC1-2 were isolated from cultivated strawberry in this study. It was revealed that the strawberry YUC genes are highly conserved. FaYUC1 is most homologous to AtYUC6, while FaYUC2 most homologous to AtYUC4, both in gene organization and protein module. By the way, our isolation of YUC genes from diploid strawberry was facilitated by the release of genomic information in January 2011. It was revealed that the woodland strawberry genome harbors nine loci for YUC genes and eight of them encode functional protein (unpublished work of our lab).
Quantitative RT-PCR analysis revealed the overlapping but distinct expression patterns for FaYUC1-2. FaYUC1 is preferentially expressed in the apical meristems from both roots and shoots, while FaYUC2 is expressed in all the tissues examined, with the highest level in stolon apical meristem tips. However, previous studies showed that plant YUCCA family is expressed in nearly all organs including flowers, leaves and roots in Arabidopsis, but the expression of most members in certain organ is restricted to a small group of discrete cells (Cheng et al. 2006). Similarly, OsYUCCA1 transcription is also found to be limited to small groups of discrete cells, including the tips of leaves and roots, and parenchyma cells surrounding large vascular stem tissues (Yamamoto et al. 2007). Promoter-driven GUS analysis and in situ hybridization study may provide an integrated understanding for FaYUC1-2 expression sites in the future.
Recently, it was found that the expression levels of early auxin-responsive genes, FaIAA1 and FaIAA2, are very high at early stage during fruit development, and decreased sharply at ripening stage in strawberry (Liu et al. 2011). Expression of these auxin-responsive genes may be the indicators of auxin contents. More than half a century ago, it was reported that the free auxin content in the achenes of the ‘Marshall’ strawberry fruits reached a steep peak 12 days after pollination (Nitsch 1950, 1955). Later, the dissection analysis showed that peaks of free auxin both in the receptacles and in the achenes appeared just before the white stage (Archbold and Dennis 1984). During ‘Jiuxiang’ strawberry fruit development, both FaYUC1-2 were abundantly expressed in achenes at G2 stage, although the sudden rise in the transcript level of FaYUC2 occurs earlier and steeper than that of FaYUC1. Dynamic expression of FaYUC1-2 in achenes is somewhat coincident with previous studies on auxin contents in strawberry fruit, suggesting that these two genes may play some roles in auxin biosynthesis for fruit development. As strawberry ripens, transcript level of FaYUC1-2 both declined gradually. This observation reinforced the previous hypothesis that the decline in the content of auxin in achenes as the fruits mature may modulate the rate of fruit ripening (Dreher and Poovaiah 1982; Given et al. 1988; Perkins-Veazie 1995). It is unknown why FaYUC2 is still significantly expressed in achenes from the ripe fruits when the growth and development has already been accomplished. Can it be related with stress responses for maintaining post-harvest quality? Besides, it is intriguing to find the expression of FaYUC1-2 in the strawberry receptacles at earlier development stages (Fig. 4b, d). In particular, FaYUC2 transcripts were detected in receptacles from mature flower to large green fruit stage. Thus, auxin synthesized through the YUC pathway in strawberry receptacles at earlier stages could be the second IAA source supporting fruit development. The previous opinion that the receptacles did not yield any free auxin (Nitsch 1950), henceforth should be reevaluated carefully. It may be metabolized, or the auxin content may be too low to be detected.
FaYUC1 over-expression in Arabidopsis results in severe defects in many aspects. Obviously, both the proper level and the proper distribution of auxin are crucial for plant development. For example, seedlings are unable to establish an apical hook, when the normal auxin balance in cells is disrupted (Lehman et al. 1996). Floral organ boundaries have been suggested to require low auxin levels, allowing for organ boundary gene activity and growth repression (Heisler et al. 2005). The abnormal phenotypes in FaYUC1 over-expression plants could also result from regulation of undesirable auxin through crosstalk with other hormones. Redundant auxin arrests bud outgrowth and enhances apical dominance by modifying local levels of cytokinin (Swarup et al. 2002). Narrow and curled leaves and strong apical dominance are well-known indicators of auxin hyper-accumulation (Kim et al. 2011). It is therefore reasonable that the phenotype of FaYUC1 over-expression plants can be attributed wholly or partly to altered growth and development due to their higher auxin contents. To conclude, the diverse high-auxin phenotypes in leaves, flowers, seedlings, and in plant stature indicate that strawberry YUC gene holds conserved function in auxin biosynthesis as its homologs do in Arabidopsis.
Several lines of evidences in this study suggest that auxin biosynthesized by YUC FMOs actively roles in the development of strawberry flowers and fruits. Is the YUCCA-involved auxin biosynthesis pathway crucial for strawberry fruit development? This notion, however, remains to be explored via reverse genetics study in strawberry. Such work is on the way in our lab. The accumulation of knowledge about auxin biosynthesis in strawberry will allow us to develop powerful tools for controlling auxin to improve certain agronomic traits (e.g. fruit yield and homogeneity).
Abbreviations
- FMO:
-
Flavin monooxygenase
- ORF:
-
Open reading frame
- RACE:
-
Rapid amplification of cDNA ends
- RT-PCR:
-
Reverse transcription-PCR
- SAM:
-
Shoot apical meristem
References
Aharoni A, Keizer LC, Van Den Broeck HC, Blanco-Portales R, Muñoz-Blanco J, Bois G, Smit P, De Vos RC, O’Connell AP (2002) Novel insight into vascular, stress, and auxin-dependent and -independent gene expression programs in strawberry, a non-climacteric fruit. Plant Physiol 129:1019–1031
Archbold DD, Dennis FG (1984) Quantification of free ABA and free and conjugated IAA in strawberry achenes and receptacles issue during fruit development. J Am Soc Hort Sci 109:30–35
Chang S, Puryear J, Cairney J (1993) A simple and efficient method for isolating RNA from pine trees. Plant Mol Biol Rep 11:113–116
Cheng Y, Dai X, Zhao Y (2006) Auxin biosynthesis by the YUCCA flavin monooxygenases controls the formation of floral organs and vascular tissues in Arabidopsis. Genes Dev 20:1790–1799
Chenna R, Sugawara H, Koike T, Lopez R, Gibson TJ, Higgins DG, Thompson JD (2003) Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res 31:3497–3500
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Davies PJ (2004) Plant hormones: biosynthesis, signal transduction, action, 3rd edn. Kluwer, Dordrecht, pp 4–6
Delker C, Raschke A, Quint M (2008) Auxin dynamics: the dazzling complexity of a small molecule’s message. Planta 227:929–941
Dreher TW, Poovaiah BW (1982) Changes in auxin content during development in strawberry fruits. J Plant Growth Regul 1:267–276
Duan K, Yi K, Dang L, Huang H, Wu W, Wu P (2008) Characterization of a sub-family of Arabidopsis genes with the SPX domain reveals their diverse functions in plant tolerance to phosphorus starvation. Plant J 54:965–975
Expósito-Rodríguez M, Borges AA, Borges-Pérez A, Hernández M, Pérez JA et al (2007) Cloning and biochemical characterization of ToFZY, a tomato gene encoding a flavin monooxygenase involved in a tryptophan-dependent auxin biosynthesis pathway. J Plant Growth Regul 26:329–340
Expósito-Rodríguez M, Borges AA, Borges-Pérez A, Pérez JA (2011) Gene structure and spatiotemporal expression profile of tomato genes encoding YUCCA-like flavin monooxygenases: the ToFZY gene family. Plant Physiol Biochem 49:782–791
Folta KM, Davis TM (2006) Strawberry genes and genomics. Crit Rev Plant Sci 25(5):399–415
Gallavotti A, Barazesh S, Malcomber S, Hall D, Jackson D, Schmidt RJ, McSteen P (2008) sparse inflorescence1 encodes a monocot-specific YUCCA-like gene required for vegetative and reproductive development in maize. Proc Natl Acad Sci 105:15196–15201
Given NK, Veins MA, Grierson D (1988) Hormonal regulation of ripening in the strawberry, a non-climacteric fruit. Planta 174:402–406
Heisler M, Ohno C, Das P, Sieber P, Reddy GV, Long JA, Meyerowitz EM (2005) Patterns of auxin transport and gene expression during primordium development revealed by live imaging of the Arabidopsis inflorescence meristem. Curr Biol 15:1899–1911
Hou X, Liu S, Pierri F, Dai X, Qu LJ, Zhao Y (2011) Allelic analyses of the Arabidopsis YUC1 locus reveal residues and domains essential for the functions of YUC family of flavin monooxygenases. J Integr Plant Biol 53:54–62
Kim JI, Sharkhuu A, Jin JB, Li P, Jeong JC, Baek D, Lee SY, Blakeslee JJ, Murphy AS, Bohnert HJ, Hasegawa PM, Yun DJ, Bressan RA (2007) yucca6, a dominant mutation in Arabidopsis, affects auxin accumulation and auxin-related phenotypes. Plant Physiol 145:722–735
Kim JI, Murphy AS, Baek D, Lee SW, Yun DJ, Bressan RA, Narasimhan ML (2011) YUCCA6 over-expression demonstrates auxin function in delaying leaf senescence in Arabidopsis thaliana. J Exp Bot 62:3981–3992
Lehman A, Black R, Ecker JR (1996) HOOKLESS1, an ethylene response gene, is required for differential cell elongation in the Arabidopsis hypocotyl. Cell 85:183–194
Leyser O (2010) The power of auxin in plants. Plant Physiol 154:501–505
Leyser O (2011) Auxin, self-organization, and the colonial nature of plants. Curr Biol 21:R331–R337
Liu DJ, Chen JY, Lu WJ (2011) Expression and regulation of the early auxin-responsive Aux/IAA genes during strawberry fruit development. Mol Biol Rep 38:1187–1193
Mezzetti B, Landi L, Pandolfini T, Spena A (2004) The defH9-iaaM auxin synthesizing gene increases plant fecundity and fruit production in strawberry and raspberry. BMC Biotechnol 4:4
Nitsch JP (1950) Growth and morphogenesis of the strawberry as related to auxin. Am J Bot 37:211–215
Nitsch JP (1955) Free auxins and free tryptophane in the strawberry. Plant Physiol 30:33–39
Pandolfini T, Molesini B, Spena A (2007) Molecular dissection of the role of auxin in fruit initiation. Trends Plant Sci 12:327–329
Perkins-Veazie P (1995) Growth and ripening of strawberry fruit. Hortic Rev 17:267–297
Raab T, López-Ráez JA, Klein D, Caballero JL, Moyano E, Schwab W, Muñoz-Blanco J (2006) FaQR, required for the biosynthesis of the strawberry flavor compound 4-hydroxy-2,5-dimethyl-3(2H)-furanone, encodes an enone oxidoreductase. Plant Cell 18:1023–1037
Rousseau-Gueutin M, Lerceteau-Köhler E, Barrot L, Sargent DJ, Monfort A, Simpson D, Arús P, Guérin G, Denoyes-Rothan B (2008) Comparative genetic mapping between octoploid and diploid Fragaria species reveals a high level of colinearity between their genomes and the essentially disomic behavior of the cultivated octoploid strawberry. Genetics 179:2045–2060
Schlaich NL (2007) Flavin-containing monooxygenases in plants: looking beyond detox. Trends Plant Sci 12:412–418
Shulaev V, Korban SS, Sosinski B, Abbott AG, Aldwinckle HS, Folta KM, Iezzoni A, Main D, Arús P, Dandekar AM, Lewers K, Brown SK, Davis TM, Gardiner SE, Potter D, Veilleux RE (2008) Multiple models for Rosaceae genomics. Plant Physiol 147:985–1003
Shulaev V, Sargent DJ, Crowhurst RN et al (2011) The genome of woodland strawberry (Fragaria vesca). Nat Genet 43:109–116
Sundberg E, Østergaard L (2009) Distinct and dynamic auxin activities during reproductive development. Cold Spring Harb Perspect Biol 1:a001628
Swarup R, Parry G, Graham N, Allen T, Bennett M (2002) Auxin cross-talk: integration of signalling pathways to control plant development. Plant Mol Biol 49:411–426
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol Biol Evol. doi:10.1093/molbev/msr121
Teale WD, Paponov IA, Palme K (2006) Auxin in action: signaling, transport and the control of plant growth and development. Nat Rev Mol Cell Biol 7:847–859
Tobeña-Santamaria R, Bliek M, Ljung K, Sandberg G, Mol JN, Souer E, Koes R (2002) FLOOZY of petunia is a flavin mono-oxygenase-like protein required for the specification of leaf and flower architecture. Genes Dev 16:753–763
Woo YM, Park HJ, Su’udi M, Yang JI, Park JJ, Back K, Park YM, An G (2007) Constitutively wilted 1, a member of the rice YUCCA gene family, is required for maintaining water homeostasis and an appropriate root to shoot ratio. Plant Mol Biol 65:125–136
Yamamoto Y, Kamiya N, Morinaka Y, Matsuoka M, Sazuka T (2007) Auxin biosynthesis by the YUCCA genes in rice. Plant Physiol 143:1362–1371
Zhao Y (2010) Auxin biosynthesis and its role in plant development. Annu Rev Plant Biol 61:49–64
Zhao Y, Christensen SK, Fankhauser C, Cashman JR, Cohen JD, Weigel D, Chory J (2001) A role for flavin monooxygenase-like enzymes in auxin biosynthesis. Science 291:306–309
Acknowledgments
The authors would like to thank anonymous referees for comments on the manuscript. This work was supported by funds from the Science and Technology Commission of Shanghai Municipality (Shanghai Rising-Star Program, 09QA1405300) and the Agricultural Commission of Shanghai Municipality (Key Program, 2008-No.1-4).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by H. Judelson.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, H., Ying, YY., Zhang, L. et al. Isolation and characterization of two YUCCA flavin monooxygenase genes from cultivated strawberry (Fragaria × ananassa Duch.). Plant Cell Rep 31, 1425–1435 (2012). https://doi.org/10.1007/s00299-012-1258-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-012-1258-4