Abstract
We recently found that Saccharomyces cerevisiae (strain CCMI 885) secretes antimicrobial peptides (AMPs) derived from the glycolytic enzyme glyceraldehyde 3-phosphate dehydrogenase (GAPDH) that are active against various wine-related yeast and bacteria. Here, we show that several other S. cerevisiae strains also secrete natural biocide fractions during alcoholic fermentation, although at different levels, which correlates with the antagonistic effect exerted against non-Saccharomyces yeasts. We, therefore, term this biocide saccharomycin. The native AMPs were purified by gel-filtration chromatography and its antimicrobial activity was compared to that exhibited by chemically synthesized analogues (AMP1 and AMP2/3). Results show that the antimicrobial activity of the native AMPs is significantly higher than that of the synthetic analogues (AMP1 and AMP2/3), but a conjugated action of the two synthetic peptides is observed. Moreover, while the natural AMPs are active at pH 3.5, the synthetic peptides are not, since they are anionic and cannot dissolve at this acidic pH. These findings suggest that the molecular structure of the native biocide probably involves the formation of aggregates of several peptides that render them soluble under acidic conditions. The death mechanisms induced by the AMPs were also evaluated by means of epifluorescence microscopy-based methods. Sensitive yeast cells treated with the synthetic AMPs show cell membrane disruption, apoptotic molecular markers, and internalization of the AMPs. In conclusion, our work shows that saccharomycin is a natural biocide secreted by S. cerevisiae whose activity depends on the conjugated action of GAPDH-derived peptides. This study also reveals that S. cerevisiae secretes GAPDH-derived peptides as a strategy to combat other microbial species during alcoholic fermentations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The antagonism exerted by Saccharomyces cerevisiae against wine-related yeast and bacteria during alcoholic fermentation has been related to the secretion of antimicrobial peptides (AMPs) (Albergaria et al. 2010; Branco et al. 2014; Kemsawasd et al. 2015). In a recent work, Branco et al. (2014) isolated a peptidic fraction from S. cerevisiae fermentation supernatants (strain CCMI 885) containing AMPs derived from the glycolytic enzyme glyceraldehyde 3-phosphate dehydrogenase (GAPDH). Two main peptides were identified in that bioactive fraction: the AMP2/3 and the AMP1, with the amino acid residues VSWYDNEYGYSTR and ISWYDNEYGYSAR, molecular masses of 1.638 and 1.622 kDa, respectively, and a theoretical isoelectric point (pI) of 4.37 (Branco et al. 2014). These anionic peptides correspond to fragments of the S. cerevisiae GAPDH2/3 (AMP2/3) and GAPDH1 (AMP1) isoenzymes.
AMPs are evolutionarily conserved components of the innate immune system and constitute the first line of antimicrobial defense in organisms across the eukaryotic kingdom (Sang and Blecha 2009; Wong et al. 2007). In the majority of cases, AMPs are cationic in nature and kill microbes by interacting with the anionic components of target cell membranes (Brogden 2005). Nevertheless, several anionic AMPs have also been found in animals and plants and, in recent years, it has become clear that they are also involved in the innate immune response of different organisms (Harris et al. 2009). The minimum inhibitory concentration (MIC) of anionic AMPs is usually weaker (MIC > 600 μM) than that of cationic AMPs (MIC ranging 10–100 μM) (Matsuzaki 2009). But the activity of anionic AMPs can be enhanced by several factors such as by the action of divalent metal cations (Dashper et al. 2005) or by additional peptides, as it was reported for lactococcin G (Nissen-Meyer et al. 1992). Anionic AMPs use a diverse range of antimicrobial mechanisms such as translocation across the membrane and permeabilization of cell membranes via pore formation (Harris et al. 2009).
Apoptosis in yeast was first discovered by Madeo et al. (1997) and was considered to be an unexpected finding since unicellular organisms seem to have no advantage in committing suicide. Nevertheless, apoptosis in yeast is now firmly confirmed, and several intrinsic, as well as exogenous stresses such as H2O2, UV irradiation, and acetic acid, have been described as apoptosis inducers in yeast cells (Madeo et al. 1999; Laun et al. 2001; Ludovico et al. 2001). Moreover, AMPs have been found to induce apoptosis in different sensitive microorganisms (Jin et al. 2010; Reiter et al. 2005). For instance, in S. cerevisiae, the virally encoded killer toxins K1 and K28 induce an apoptotic cell response in sensitive yeast strains (Reiter et al. 2005).
The aim of the present work was to characterize the antimicrobial properties and the death-inducing mechanisms of the GAPDH-derived AMPs, and to evaluate the role they play in the ability of S. cerevisiae strains to combat other microbial species during wine fermentation. With that purpose, several S. cerevisiae strains were screened regarding the levels of the natural biocide secreted during mixed-culture alcoholic fermentations and the antagonistic effect exerted against a sensitive non-Saccharomyces strain. Chemically synthesized analogues of the two main peptides (AMP2/3 and AMP1) that compose the native biocide were used to evaluate its antimicrobial activity and death-inducing mechanisms (e.g., cell membrane disruption, death by apoptosis, and internalization of AMPs).
Materials and methods
Strains and growth conditions
In this work we used the following S. cerevisiae strains: CCMI 885 (Culture Collection of Industrial Microorganisms of ex-INETI, Portugal); ISA 1000 (Culture collection of Instituto Superior de Agronomia, Portugal), ISA 1028, ISA 1029, ISA 1046, ISA 1063, ISA 1200; S101 (Saint Georges S101, Bio Springer, France); and ATCC 6269 (American Type Culture Collection). The non-Saccharomyces yeast strains used were as follows: Dekkera bruxellensis ISA 2211; Hanseniaspora guilliermondii NCYC 2380 (National Collection of Yeast Cultures, Norwich, United Kingdom); Kluyveromyces marxianus PYCC 2671 (Portuguese Yeast Culture Collection, FCT/UNL, Caparica, Portugal); Lachancea thermotolerans PYCC 2908; and Torulaspora delbrueckii PYCC 4478. Yeast strains were maintained on yeast extract peptone dextrose (YEPD)-agar slants (20 g/l of glucose, 20 g/l of peptone, 10 g/l yeast extract, 20 g/l agar) and stored at 4 °C. Inoculums were prepared by transferring biomass from one YEPD-agar slant (pre-grown at 30 °C for 48 h) into in 250-ml flasks with 100 ml of YEPD and incubating flasks at 30 °C and 150 rpm, for 16 h. All media were autoclaved at 120 °C for 20 min.
Mixed-culture alcoholic fermentations
Synthetic grape juice (SGJ) (110 g/l of glucose plus 110 g/l of fructose, pH 3.5, prepared as described in Pérez-Nevado et al. (2006)) fermentations were performed with mixed cultures of H. guilliermondii and each of the following S. cerevisiae strains: CCMI 885, ISA 1000, ISA 1028, ISA 1029, ISA 1046, ISA 1063, ISA 1200, S101, and ATCC 6269. One SGJ-fermentation was performed with H. guilliermondii in single-culture and used as negative control of the antagonism exerted by the S. cerevisiae strains. All fermentations were carried out in 500-ml flasks containing 300 ml of SGJ that were inoculated with 105 cells per milliliter of each yeast and incubated at 25 °C, under gentle agitation (80 rpm). Fermentations were carried out in duplicates and daily samples were taken to determine cell growth, sugars consumption, and ethanol production. Cell growth was assessed by colony forming units (CFU) counts. Briefly, 100 μl of culture sample were spread onto YEPD-agar plates, after appropriate dilution, and incubated at 30 °C in a vertical incubator (Infors, Anjou, Canada) for 2–6 days. In the mixed-culture fermentations, CFU counts of H. guilliermondii were obtained on YEPD-agar plates with 0.001 % of cycloheximide and the CFU counts of S. cerevisiae determined as the difference between the total number of CFU on YEPD-agar plates (both species grow) and the number of CFU on cycloheximide-YEPD-agar plates (only H. guilliermondii grows). Sugars (glucose and fructose) and ethanol concentrations were determined by high-performance liquid chromatography (HPLC) using an HPLC apparatus (Merck Hitachi, Darmstadt, Germany) equipped with a refractive index detector (L-7490, Merck Hitachi, Darmstadt, Germany). Cell-free samples (filtration by 0.45 μm Millipore membranes) were injected into a Sugar-Pak column (Waters Hitachi, Milford, USA) and eluted with a degassed CaEDTA (50 mg/l) aqueous mobile phase at 90 °C and 0.5 ml/min.
Purification of native biocide fractions by gel-filtration chromatography
Cell-free supernatants (7-day-old) from each of the mixed-culture fermentations performed were ultrafiltrated by centrifugal filter units (Vivaspin 15R, Sartorius, Gottingen, Germany) equipped with 10 and 2 kDa cutoff membranes. Peptidic fractions (2–10 kDa) were obtained by first passing the fermentation supernatants through 10 kDa centrifugal filter units and then concentrating (10-fold) those permeates in 2 kDa centrifugal filter units. These peptidic fractions (2–10 kDa) were then fractionated by gel-filtration chromatography using a Superdex-Peptide column (10/300 GL, GE Healthcare, London, UK) coupled to an HPLC system (Merck Hitachi, Darmstadt, Germany) equipped with a UV detector (Merck Hitachi, Darmstadt, Germany). The peptidic supernatant fractions (2–10 kDa) were eluted with 0.1 M ammonium acetate at a flow rate of 0.7 ml/min. The eluate fractions between the retention time 27–29 min were collected and lyophilized.
Spectrum of action and antimicrobial properties of the native biocide
The minimum inhibitory concentration (MIC) and half inhibitory concentration (IC50) of the native biocide (i.e., GAPDH-derived AMPs) were determined against H. guilliermondii, L. thermotolerans, K. marxianus, T. delbrueckii and D. bruxellensis. The gel-filtration lyophilized fraction-II obtained from the S. cerevisiae strain CCMI 885 fermentation supernatant was resuspended in YEPD with 30 g/l of ethanol and pH 3.5. Growth inhibitory assays were performed in 96-well microplates containing 100 μl of YEPD medium, without fraction-II (control) and with fraction-II at final protein concentrations of 125, 250, 500, and 1000 μg/ml. Media were inoculated with 105 cells per milliliter of each of the above-mentioned non-Saccharomyces yeasts, and the microplates incubated in a Thermo-Shaker (Infors HT, Bottmingen, Switzerland) at 30 °C, under strong agitation (700 rpm). Cell growth was followed by optical density measurements (at 590 nm) in a Microplate Reader (Dinex Technologies Inc., Chantilly, USA) and by CFU counts. The MIC was defined as the minimum concentration of biocide that completely inhibited the growth of the sensitive yeast, and the IC50 as the concentration of biocide that induced a growth reduction of 50 % as compared with growth in the respective control assay.
Antimicrobial activity of synthetic peptide analogues (AMP2/3 and AMP1)
Analogues of the AMP2/3 (amino acids residues: VSWYDNEYGYSTR) and AMP1 (amino acids residues: ISWYDNEYGYSAR) were chemically synthetized according to standard procedures and purchased from GenScript Inc. Company (GenScript HK Limited, Hong Kong). The synthetic peptides were obtained in lyophilized form; stock-solutions of each peptide were prepared by dissolving 2 mg of lyophilized powder in 1 ml of deionized water and the pH was adjusted to 8.0 with a sodium hydroxide solution until total solubilization was attained. The antimicrobial activity of the synthetic peptides AMP2/3 and AMP1 was determined against H. guilliermondii in growth inhibitory assays performed as described in the “Spectrum of action and antimicrobial properties of the native biocide” section. Briefly, a 50-μl aliquot of each AMP stock solution was mixed with 50 μl of 2 × YEPD (two-fold concentrated YEPD with 60 g/l ethanol) and the final pH was adjusted to 6.0. The AMPs solutions were used in growth assays at the following concentrations (μg/ml): 125, 250, 500, and 1000. Media were inoculated with 105 cells per milliliter of H. guilliermondii and cultures were incubated in a Thermo-Shaker (Infors HT, Bottmingen, Switzerland) at 30 °C, under strong shaking (700 rpm). The combined action of the two synthetic peptides was also tested against the same yeast strain, using mixtures of the synthetic AMP2/3 + AMP1 at the ratios of 1:1; 2:1; 4:1, and 6:1, to a final concentration of 1000 μg/ml.
Internalization of AMPs fluorescently labeled with fluorescein
Exponentially grown cells of H. guilliermondii and D. bruxellensis were separately incubated in deionized water and in YEPD medium (pH = 6.0) at room temperature (ca. 20–25 °C) with synthetic peptides (AMP2/3 and AMP1) fluorescently labeled with fluorescein (FITC). The AMPs were chemically synthesized and fluorescently labeled with FITC according to standard procedures and purchased from GenScript Inc. Company (GenScript HK Limited, Hong Kong). The AMPs-FITC were added to deionized water and to YEPD medium using a mixture of AMP2/3 + AMP1 in a ratio of (4:1) to a final concentration of 1000 μg/ml. The initial cell density in the assays was 106 cells per milliliter, and four different media were used as follows: deionized water and YEPD, without and with ethanol (30 g/l). Each assay was performed in duplicates. After 1 h of incubation with the AMPs-FITC, cells were harvested by centrifugation (7000×g, for 5 min), stained with 10 μl of propidium iodide (PI) solution (1 mg/ml) and incubated for 30 min in the dark. Finally, cells were visualized in an epifluorescent microscope (Zeiss Axiovert 50, Oberkochen, Germany) equipped with a Zeiss Neofluor ×40 objective (numerical aperture 0.75) and the number of cells emitting green fluorescence (internalization of the AMPs-FITC) and red fluorescence (PI-stained cells) was quantified to determine the percentage of cells that were able to internalize the AMPs-FITC and those with permeabilized membranes (PI-stained cells).
Analyses of apoptotic and necrotic markers
Apoptotic and/or necrotic markers induced by the synthetic AMPs in sensitive yeast cells were assessed in H. guilliermondii incubated (105 cells/ml) for 2 h in YEPD (with 30 g/l of ethanol at pH 6.0) with the synthetic AMPs. The AMPs were added to the YEPD medium using mixtures of the two synthetic peptides (i.e., AMP2/3 + AMP1) in a ratio of 4:1, to final concentration of 100 μg/ml. H. guilliermondii cells incubated in YEPD without the AMPs were used as negative control, and H. guilliermondii cells incubated in YEPD with 5 mM of peroxide of hydrogen (H2O2) were used as positive control. Since cycloheximide inhibits the protein synthesis in yeast, this antibiotic is typically used to validate the apoptosis-inducing ability of a given stress (e.g., H2O2). Thus, to confirm the ability of these AMPs to induce apoptosis, H. guilliermondii was incubated in YEPD with 0.01 % of cycloheximide and 100 μg/ml of the synthetic AMPs. Apoptotic cellular markers (i.e., DNA strand breaks, phosphatidylserine exposure at the surface of the cytoplasmatic membrane, and chromatin condensation) were detected in the AMP-treated cells by the epifluorescent microscopic methods described in the following section.
TUNEL method
DNA strand breaks were confirmed by the incorporation of modified dUTPs at the 3ʹ-OH ends of fragmented DNA using the enzyme terminal deoxynucleotidyl transferase (TdT). Modifications were directly detected by epifluorescent microscopy using a fluorescently modified nucleotide (i.e., fluorescein-dUTP) (Click-iT TUNEL Alexa Fluor imaging Assay, Invitrogen, USA) and the following procedure: firstly, yeast cells were fixed during 1 h with 4 % paraformaldehyde, digested with zymolyase and β-glucuronidase during 1 h and 30 min at 37 °C under agitation (150 rpm) and permeabilized with sodium citrate 0.1 M for 30 min at 70 °C. Then, cells were washed with PBS buffer and incubated with 20 μl of TUNEL reaction mixture (60 U/ml of terminal deoxynucleotidyl transferase, 1 μl of EdUTP nucleotide mixture, and 47 μl of reaction buffer) for 1 h at 37 °C in the dark; finally, cells were washed with PBS and spotted onto a Neubauer chamber to enumerate the cells exhibiting green fluorescence by epifluorescence microscopy (Olympus BX-60 microscope, Tokyo, Japan).
ANNEXIN V/PI staining
The exposure of phosphatidylserine at the surface of the cytoplasmatic membrane in apoptotic cells was detected by applying fluorescein conjugated with Annexin V (Alexa fluor 488, Invitrogen, Paisley, UK) together with PI. Briefly, cells were washed in sorbitol buffer (2 M sorbitol, 0.5 mM MgCl2, 35 mM potassium phosphate, pH 6.8), digested with zymolyase and β-glucuronidase during 1 h and 30 min at 37 °C under agitation (150 rpm) and after this washed with biding buffer/sorbitol (10 mM HEPES/NaOH, pH 7.4, 140 mM NaCl, 2.5 mM CaCl2). Then, 5 μl of Annexin V and 2 μl of PI (0.5 μg/ml) were added and cells were incubated for 20 min at room temperature in the dark. These cells were harvested by centrifugation, resuspended in binding buffer/sorbitol and spotted onto a Neubauer chamber to enumerate cells stained with PI (necrotic cells) and cells emitting green fluorescence at the surface of the cytoplasmatic membrane (apoptotic cells) by epifluorescence microscopy (Olympus BX-60 microscope, Tokyo, Japan).
DAPI staining
Chromatin condensation was accessed by microscopic observations of cells stained with the fluorescent dye 4,6 diamidino-2-phenylindole (DAPI). Briefly, AMP-treated cells were incubated for 20 min with 1 mg/ml of DAPI (Invitrogen, Paisley, UK) in the dark, at room temperature. Cells were harvested by centrifugation and resuspended in PBS and spotted onto a Neubauer chamber to enumerate cells exhibiting blue fluorescence (DAPI-stained cells) by epifluorescence microscopy (Olympus BX-60 microscope, Tokyo, Japan).
Results
Antagonism of S. cerevisiae strains and secretion of native biocide fractions
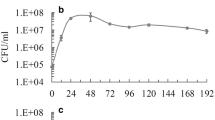
The antagonism exerted by different S. cerevisiae strains against non-Saccharomyces yeast was assessed by performing synthetic grape juice (SGJ) fermentations with mixed-cultures of H. guilliermondii and several S. cerevisiae strains. SGJ-fermentation performed with H. guilliermondii in single culture was used as negative control. Comparing the growth profiles of H. guilliermondii during the mixed-culture fermentations (Fig. 1a–i) with the single-culture fermentation (Fig. 1j), it is clear that all S. cerevisiae strains induced death of H. guilliermondii, although at different rates. While the cell viability of H. guilliermondii was entirely lost within the first 72 h in the mixed-culture fermentations performed with the S. cerevisiae strains CCMI 885, ISA 1028, and ISA 1046, a similar effect occurred only after 96 h with the strains ISA 1000, ISA 1063, and S101 and only after 168 h with the strains ISA 1029, ISA 1200, and ATCC 6269. In all mixed-culture fermentations, the initial sugars (220 g/l of glucose + fructose) were entirely consumed within 3–7 days, whereas in the single-culture fermentation H. guilliermondii consumed only 72 % of the initial sugars, leaving 62 g/l of residual sugars (glucose + fructose) after 7 days and producing 66 g/l of ethanol (Fig. S1 in the Electronic Supplementary Material (ESM)).
Cell density profiles (CFU/ml) of S. cerevisiae (diamonds) and H. guilliermondii (squares) during mixed-culture fermentations performed with S. cerevisiae strains CCMI 885 (a), ISA 1000 (b), ISA 1028 (c), ISA1029 (d), ISA 1046 (e), ISA 1063 (f), ISA 1200 (g), S101 (h), ATCC 6269 (i), and during the single-culture fermentation of H. guilliermondii (j). Values represented are means of triplicate measurements ± SD (error bars) of two independent biological experiments
The peptidic fractions (2–10 kDa) of cell-free supernatants (7-day-old) obtained from every fermentation were fractionated by gel-filtration chromatography to isolate the natural biocide fraction in which the GAPDH-derived AMPs were previously identified by Branco et al. (2014) (i.e., peak-II indicated in Fig. 2). Death rates of H. guilliermondii during the mixed-culture fermentations show a positive correlation with the relative amount of the natural biocide fraction present in each supernatant (Table 1). Indeed, the supernatants from the fermentations where H. guilliermondii died within 72 h (strains CCMI 885, ISA 1028, and ISA 1046) showed the largest peak II areas, while the supernatants from the fermentations where H. guilliermondii took 168 h to die off (strains ISA 1029, ISA 1200. and ATCC 6269) showed the smallest peak-II areas (Table 1). The area of peak-II in these supernatants varied by two-fold, with the supernatants from the S. cerevisiae strains CCMI 885 and ISA 1029 exhibiting the largest area and the smallest area, respectively.
Gel-filtration chromatographic profiles of the peptidic fractions (2–10 kDa) of supernatants obtained from the mixed-culture fermentations performed with H. guilliermondii and different S. cerevisiae strains (CCMI 885, S101, and ISA 1029), and from the single-culture fermentation of H. guilliermondii (Hg). Fractions indicated as peak-II correspond to the bioactive fraction in which the GAPDH-derived AMPs were previously identified by Branco et al. (2014)
Sequence alignments of GAPDH isoenzymes for wine-related yeasts
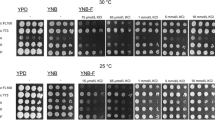
In S. cerevisiae, three related but not identical GAPDH isoenzymes (GAPDH1, GAPDH2, and GAPDH3) are encoded by unlinked genes designated TDH1, TDH2, and TDH3 (McAlister and Holland 1985). The GAPDH-derived peptides (AMP2/3 and AMP1) identified in S. cerevisiae fermentation supernatants match the C-terminal (309–321) sequence of the isoenzymes GAPDH2/3 (amino acids residues: VSWYDNEYGYSTR) and GAPDH1 (amino acids residues: ISWYDNEYGYSAR), respectively. To investigate the species-specificity of the amino acid sequences of these AMPs, we performed the sequence alignment of GAPDH isoenzymes of several non-Saccharomyces yeasts, in the region containing the AMP2/3 and the AMP1 fragments (Fig. S2 in the ESM). Results show a high homology among the GAPDH sequences in the AMP2/3’s region. However, the amino acid sequence of the AMP1 fragment seems to be quite unique, since it does not fully match any of the GAPDH sequences of the non-Saccharomyces yeasts analyzed (at least one amino acid is always different). It should be mentioned that we did not use H. guilliermondii in the sequence alignments because the genome of this species has not been sequenced yet.
Antimicrobial properties of the native and synthetic AMPs
Minimum inhibitory concentrations (MICs) of the native biocide (GAPDH-derived AMPs) against H. guilliermondii, K. marxianus, and L. thermotolerans were 250 μg/ml, while against T. delbrueckii and D. bruxellensis higher values were observed (500 and 1000 μg/ml, respectively) (Table 2). Half inhibitory concentrations (IC50) agree well with MICs for the same non-Saccharomyces yeasts (Table 2). The fungicidal effect of the native AMPs against these non-Saccharomyces yeasts was quantified as the number of LOGs of [CFU/ml] reduction (Table 3). The native AMPs show a strong fungicidal effect against H. guilliermondii, reducing its cell density by 4.2 and 5.2 orders of magnitude at 250 and 500 μg/ml, respectively (Table 3). Against T. delbrueckii and D. bruxellensis, the fungicidal effect of the native AMPs was lower, with the cell density of these yeasts being reduced by 3.6 and 2.9 orders of magnitude, respectively, at 500 and 2000 μg/ml (Table 3).
To further investigate the mode of action of the GAPDH-derived AMPs, synthetic analogues of the two main peptides that were isolated from the native biocide fraction (i.e., AMP1 and AMP2/3) were used to assess their antimicrobial effect against the sensitive yeast H. guilliermondii. However, due to the anionic nature of these synthetic peptides (pI = 4.35) it was not possible to test their inhibitory effect at the same acidic conditions used for the native biocide (i.e., YEPD at pH = 3.5), since they did not dissolve at this acidic pH. In fact, the synthetic AMPs contain a majority of acidic amino acids in their primary structure, which prevents its solubilization at pH = 3.5. Therefore, the antimicrobial activity of the synthetic peptides was assessed in YEPD at pH 6.0, using increasing concentrations (0, 125, 250, 500, and 1000 μg/ml) of AMP2/3 and AMP1, either alone or mixed at different ratios. Results showed (Fig. 3) that both AMPs inhibited the growth of H. guilliermondii, although the AMP1 exhibited a much stronger effect than the AMP2/3 (76 % of inhibition for the AMP1 and only 30 % for AMP2/3, both at 1000 μg/ml). Besides, none of the synthetic AMPs (used either alone or mixed in a 1:1 ratio) was able to kill H. guilliermondii with the same efficiency of the natural AMPs (Table 3). Nevertheless, an increased antimicrobial effect was observed when the two peptides were used together (Fig. 3). Given these observations, we evaluated the effect of mixing the AMP2/3 with the AMP1 at different proportions, namely at the ratios of 2:1, 4:1, and 6:1 (final concentrations of 1000 μg/ml), on the growth inhibition of H. guilliermondii. These ratios were chosen based on the fact that the GAPDH2/3 isoenzymes from which the AMP2/3 derives are produced by S. cerevisiae cells at much higher proportions than the GAPDH1 isoenzyme, from which the AMP1 derives. In fact, McAlister and Holland (1985) found that the contribution of the TDH1, TDH2, and TDH3 gene products to the total GAPDH activity in S. cerevisiae cells is 10–15, 25–30, and 50–60 %, respectively. Results revealed that, under such conditions, the AMPs were able to kill H. guilliermondii (cell viability was reduced by about 1–2 orders of magnitude), with the strongest fungicidal effect being achieved at the ratio of 4:1 (Fig. 4). The conjugated action of the two synthetic peptides can also be confirmed by comparing the inhibitory capacity of the AMP1 used alone or mixed with the AMP2/3 at a ratio of 4:1 (AMP2/3:AMP1), for the same concentration of AMP1 in each situation (Fig. S3). However, regardless of the way the synthetic AMPs were used (alone or mixed at any ratio) its antimicrobial activity was always lower than that of the natural biocide. Indeed, the synthetic AMPs were only able to reduce the cell viability of H. guilliermondii by ca. two orders of magnitude (using mixtures of AMP2/3 + AMP1 in a ratio of 4:1) at 1000 μg/ml (Fig. 4), while the natural AMPs reduced the cell viability of the same yeast strain by ca four orders of magnitude (Table 3) at a much lower concentration (at 250 μg/ml).
Growth inhibition of H. guilliermondii (relative to control) in the assays performed in YEPD (pH = 6.0) with increasing concentrations of the synthetic AMP2/3 alone (triangles), of the synthetic AMP1 alone (circles) and of a mixture of AMP2/3 and AMP1 (squares) at a ratio of 1:1. Values represented are means of triplicate measurements ± SD (error bars) of two independent biological experiments
Cell viability (CFU/ml) of H. guilliermondii after 18 h of incubation in YEPD (pH = 6.0) without AMPs (control) and with mixtures of the synthetic AMP2/3 and AMP1 (final concentrations of 1000 μg/ml) at ratios of 6:1, 4:1, and 2:1. The initial cell density was 105 CFU/ml in all the assays. Values are means of triplicate measurements ± SD (error bars) of two biological independent assays
Internalization of the synthetic AMPs by sensitive yeast cells
To investigate the internalization ability of the AMPs, exponentially grown cells of H. guilliermondii and D. bruxellensis were separately incubated in deionized water and in YEPD (both media with and without ethanol) in the presence of the AMPs fluorescently labeled with FITC (AMPs-FITC). Results showed that the synthetic AMPs were able to enter in cells of both yeasts (Figs. 5 and 6). However, the percentage of cells that internalized the AMPs significantly increased when cells were incubated in YEPD (ca. 25–30 %) instead of water (less than 10 %) (Fig. 6a, b). On the other hand, ethanol had no impact on the ability of the AMPs to penetrate H. guilliermondii cells (Fig. 6a), whereas an increased internalization was observed in D. bruxellensis cells (Fig. 6b). The membrane integrity of H. guilliermondii and D. bruxellensis cells was assessed by staining cells with PI, revealing that all cells that internalized the AMPs (AMPs-FITC) also showed compromised cell membranes (PI-stained) (Fig. 6a, b).
Percentage of H. guilliermondii (a) and D. bruxellensis (b) cells that internalized the synthetic AMPs fluorescently labeled with FITC (AMPs-FITC) and that lost membrane integrity (PI-stained) after incubation in deionized water and in YEPD, without ethanol (AMPs-FITC/PI-stained) and with 30 g/l of ethanol (AMPs-FITC + EthOH/ PI-stained + EthOH). Values correspond to means of triplicate measurements ± SD (error bars) from two biological independent assays
Apoptotic/necrotic molecular markers in AMPs-treated cells
Apoptotic cell death induced by AMPs has been reported by several authors (Jin et al. 2010) and (Reiter et al. 2005). This led us to investigate whether an apoptosis-like process occurs in sensitive yeast cells exposed to the synthetic AMPs. Cells dying by apoptosis display typical molecular markers such as the following: DNA strand breaks, detectable by the TUNEL-assay; chromatin condensation, detectable by DAPI-staining; and exposure of phosphatidylserine at the outer cell membrane, detectable by Annexin V-FITC staining. In the latter assay, apoptotic and necrotic cells can be distinguished by double staining cells with Annexin V (green fluorescence) and PI (red fluorescence), which is a membrane-impermeant fluorescent dye. These cellular markers were assessed by epifluorescent microscopy in H. guilliermondii cells incubated in YEPD without the AMPs (control) and with 100 μg/ml of synthetic AMPs. Cells were also incubated in YEPD with 5 mM of H2O2 (positive control) and in YEPD with 100 μg/ml of synthetic AMPs plus cycloheximide (negative control). Results (Fig. 7) showed that H. guilliermondii cells treated with 100 μg/ml of AMPs exhibited 28 % of cells with DNA strand breaks (TUNEL-positive), 4 % of cells with phosphatidylserine exposure at the membrane surface (Annexin+/PI−), 1 % of necrotic cells (Annexin+/PI+), and no cells with chromatin condensation (DAPI-positive). Prior to these assays, H. guilliermondii cells were incubated in YEPD medium in the presence of 3.0, 5.0, and 180 mM of H2O2 and the above-mentioned apoptotic molecular markers were assessed, following the procedure described by Madeo et al. (1999). Results revealed that H. guilliermondii cells exposed to 5 mM of H2O2 exhibited a higher percentage of apoptotic cells than when exposed to 3 mM of a H2O2, while 180 mM of H2O2 induced necrosis (PI-stained cells) in 95 % of cells (data not shown). Thus, H2O2 at 5.0 mM was used as positive control of death by apoptosis in H. guilliermondii cells (Fig. 7). Conversely, cycloheximide inhibits apoptosis in yeast since it blocks the protein synthesis machinery required to execute the programmed cell death mechanism. By comparing the apoptosis molecular markers exhibited by H. guilliermondii cells treated with the synthetic AMPs in the absence and in the presence of cycloheximide (Fig. 7), our results suggest that the AMPs indeed induce apoptosis in the sensitive yeast H. guilliermondii.
Percentage of H. guilliermondii cells exhibiting apoptotic/necrotic molecular markers after incubation in YEPD medium, for 2 h, at different conditions: without AMPs (control); with 5 mM of H2O2 (positive control); with 100 μg/ml of AMPs plus 0.01 % cycloheximide (negative control); with 100 μg/ml of AMPs. DAPI apoptotic cells with chromatin condensation; Annexin+/PI+ necrotic cells with compromised membranes; TUNEL apoptotic cells with DNA-strand breaks; Annexin+/PI− apoptotic cells with phosphatidylserine exposed at the surface of cytoplasmatic membrane. Values correspond to means of triplicate measurements ± SD (error bars) from two biological independent assays
Discussion
We had previously reported that S. cerevisiae (strain CCMI 885) secretes AMPs derived from the glycolytic enzyme GAPDH (Albergaria et al. 2010; Branco et al. 2014), active against several wine-related non-Saccharomyces yeasts (e.g., D. bruxellensis, K. marxianus, L. thermotolerans, and T. delbrueckii) and bacteria (e.g., Oenococcus oeni). In the present work, we show that several other S. cerevisiae strains also secrete these AMPs during alcoholic fermentation and, therefore, we term the native biocide saccharomycin. In addition, we demonstrate that there is a positive correlation between the death rates of H. guilliermondii during mixed-culture fermentations performed with different S. cerevisiae strains and the levels of saccharomycin excreted to the extracellular medium. Since saccharomycin exhibited a fungicidal effect against several wine-related non-Saccharomyces yeasts, our results strongly suggest that secretion of GAPDH-derived peptides is a defensive strategy used by S. cerevisiae strains to combat other microbial species during wine fermentation.
The involvement of GAPDH in the defense system of S. cerevisiae seems surprising, since this protein is mainly associated with its glycolytic role. However, recent studies have shown that GAPDH also displays several other activities in different subcellular locations (Nakajima et al. 2009; Silva et al. 2011; Sirover 2005, 2011). For example, GAPDH is a cell-wall-associated protein with adhesion properties in bacteria (Izquierdo et al. 2009) and in the yeast Candida albicans where it plays a role in virulence (Gil et al. 1999). GAPDH has also been found on the cell surface of different yeasts such as in K. marxianus, involved in cell flocculation (Fernandes et al. 1992), and in S. cerevisiae, with unknown functions (Delgado et al. 2001, 2003). In mammalian cells, GAPDH is overexpressed in neuronal apoptotic cells and involved in Alzheimer’s disease (Sunaga et al. 1995), while, in parasite, GAPDH is an immuno-suppressor (Sahoo et al 2013). Due to its diverse activities, GAPDH has been called a “moonlighting protein” (Sirover 2011). Besides, two different GAPDH-derived peptides with antifungal activity were recently isolated: one from the human placental tissue (Wagener et al. 2013) and the other from the skin of yellowfin tuna (Seo et al. 2012). On the other hand, GAPDH is a highly conserved protein, which means that its amino acid sequence should not vary significantly among close-related species. Indeed, our sequence alignments of the GAPDH isoenzymes for S. cerevisiae and some wine-related non-Saccharomyces yeasts show huge homology within the region that contains the AMP2/3. Nevertheless, the amino acid sequence of the AMP1 varies for, at least, one amino acid within the GAPDH sequences of those non-Saccharomyces yeasts. Interestingly, our results show that the antimicrobial activity of the AMP1 is much higher than that of the AMP2/3. The AMP1 originates from the GAPDH1 isoenzyme, which is only synthesized when S. cerevisiae cells enter the stationary growth phase (Boucherie 1995). Taken together, these findings could explain why the non-Saccharomyces yeasts invariable die off more intensely after S. cerevisiae attains the stationary growth phase during alcoholic fermentations (Albergaria et al. 2010; Nissen and Arneborg 2003; Pérez-Nevado et al. 2006).
Using synthetic analogues of the main peptides that compose the natural biocide (i.e., AMP2/3 and AMP1) we found that the antimicrobial activity of the native AMPs depends on the conjugated action of these GAPDH-derived peptides. Besides, a maximal antimicrobial effect was found when the AMP2/3 was mixed with the AMP1 at a ratio of 4:1. It is worth noting that, if the naturally secreted GAPDH-derived peptides are able to form aggregates of five molecules, the global molecular weight (MW) of those aggregates would be of about 8.0 kDa (MW of each peptide is ca. 1.6 kDa), which agrees with the apparent MW of the bioactive fractions isolated from the gel-filtration chromatography (data not shown). Moreover, while saccharomycin is active at acidic conditions (YEPD at pH = 3.5), the synthetic peptides are not. These findings prompt us to propose that the natural biocide may adopt a molecular structure involving the formation of aggregates of several peptide molecules (probably, five peptides) which render them soluble and bioactive at acidic conditions. In fact, different studies have shown that the activity of some AMPs depends on the conjugated action of several molecules (Nissen-Meyer et al. 1992; Straus and Hancock 2006). That is the case of lactococcin G, a bacteriocin whose activity depends on the complementary action of two peptides at approximately equal proportions (Nissen-Meyer et al. 1992). Also, daptomycin is an anionic AMP that was first isolated from Streptomyces roseosporus (Debono et al. 1987) and whose activity depends on the formation of aggregates of 14–16 daptomycin molecules that form a micelle-like structure by the action of calcium cations (Ca2+) (Straus and Hancock 2006).
Most AMPs induce death of sensitive cells by interacting with cell membranes and permeabilizing them (Pandey et al. 2011). However, some AMPs have developed unique mechanisms to translocate across membranes and to act on cytoplasmic targets without disrupting cell membranes (Powers and Hancock 2003). Indeed, translocation across membranes by a micellar aggregate mechanism was first proposed for the frog-derived antimicrobial peptide buforin II (Park et al. 2000), which rather than causing large membrane perturbations induces a transient disruption without permanent permeabilization (Powers and Hancock 2003). Once present in the cytoplasm, AMPs are thought to interact with DNA, RNA, and/or cellular proteins and to inhibit synthesis of these compounds (Brown and Hancock 2006). In the present work, we investigated the death mechanisms induced by the S. cerevisiae AMPs, by treating sensitive yeast cells with synthetic AMPs fluorescently labeled with FITC (internalization of peptides) and by staining those cells with PI (loss of cell membrane integrity). Results showed that all sensitive cells that internalized the AMPs, also exhibited cell membrane permeabilization. In a previous work (Branco et al. 2015), we had already found that the natural S. cerevisiae AMPs induce membrane permeabilization of H. guilliermondii cells. Here, we show that the synthetic AMPs are also able to cross the cell membrane and enter in the cytoplasm of sensitive yeast cells (both H. guilliermondii and D. bruxellensis), which means these AMPs are cell-penetrating peptides. Besides, both of these cellular effects were much more pronounced when sensitive cells were treated with the AMPs in YEPD (25–30 % of cells with disrupted membranes and AMPs internalization) than in deionized water (less than 10 % of cells with disrupted membranes and AMPs internalization). These results suggest that some component of the YEPD medium, probably a metal cation (e.g., Fe2+, Mn2+, Mg2+, etc.), may enhance the activity of these AMPs. Although the effect of metal cations on the internalization of AMPs has never been reported, several studies (Dashper et al. 2005, 2007) have shown that the antimicrobial activity of anionic AMPs can be enhanced by the action of metal cations, since they promote the biding with the negatively charged cell wall. That is the case of kappacins, isolated from bovine milk and the first anionic AMPs to be investigated (Malkoski et al. 2001). In fact, Dashper et al. (2005) demonstrated that the antibacterial effect of kappacins is enhanced by the presence of divalent metal cations (both Zn2+ and Ca2+). In addition, those authors found that under acidic conditions the membranolytic ability of kappacins in the presence of Zn2+ was higher, which could promote the influx of hydrogen ions, lower the intracellular pH, and thus increase the antibacterial activity (Dashper et al. 2005, 2007).
AMPs are known to trigger cell death by inducing molecular markers typical of death by apoptosis (Jin et al. 2010; Reiter et al. 2005). Here, we show that sensitive yeast cells treated with sub-lethal concentrations (i.e., 100 μg/ml) of synthetic analogues of the GAPDH-derived AMPs exhibit cellular markers characteristic of death by apoptosis such as DNA fragmentation, a typical late apoptosis phenomenon.
In conclusion, our work shows that saccharomycin is a natural biocide secreted by different S. cerevisiae strains that induces the death of several wine-related non-Saccharomyces yeasts, and whose activity depends on the conjugated action of GAPDH-derived peptides. The death mechanisms induced by these AMPs on sensitive yeasts involve cell membrane permeabilization, internalization of peptides and induction of apoptotic molecular markers.
References
Albergaria H, Francisco D, Gori K, Arneborg N, Gírio F (2010) Saccharomyces cerevisiae CCMI 885 secretes peptides that inhibit the growth of some non-Saccharomyces wine-related strains. Appl Microbiol Biotechnol 86:965–972. doi:10.1007/s00253-009-2409-6
Boucherie H (1995) Differential synthesis of glyceraldehyde-3-phosphate dehydrogenase polypeptides in stressed yeast cells. FEMS Microbiol Lett 125:127–133. doi:10.1016/0378-1097(94)00484-9
Branco P, Francisco D, Chambon C, Hébraud M, Arneborg N, Almeida MG, Caldeira J, Albergaria H (2014) Identification of novel GAPDH-derived antimicrobial peptides secreted by Saccharomyces cerevisiae and involved in wine microbial interactions. Appl Microbiol Biotechnol 98:843–853. doi:10.1007/s00253-013-5411-y
Branco P, Viana T, Albergaria H, Arneborg N (2015) Antimicrobial peptides (AMPs) produced by Saccharomyces cerevisiae induce alterations in the intracellular pH, membrane permeability and culturability of Hanseniaspora guilliermondii cells. Int J Food Microbiol 205:112–118. doi:10.1016/j.ijfoodmicro.2015.04.015
Brogden KA (2005) Antimicrobial peptides: pore formers or metabolic inhibitors in bacteria? Nat Rev Microbiol 3:238–250. doi:10.1038/nrmicro1098
Brown KL, Hancock RE (2006) Cationic host defense (antimicrobial) peptides. Curr Opin Immunol 18:24–30. doi:10.1016/j.coi.2005.11.004
Dashper SG, O’Brien-Simpson NM, Cross KJ, Paolini RA, Hoffmann B, Catmull DV, Malkoski M, Reynolds EC (2005) Divalent metal cations increase the activity of the antimicrobial peptide kappacin divalent metal. Antimicrob Agents Chemother 49:2322–2328. doi:10.1128/AAC.49.6.2322
Dashper SG, Liu SW, Reynolds EC (2007) Antimicrobial peptides and their potential as oral therapeutic agents. Int J Pept Res Ther 13:505–516. doi:10.1007/s10989-007-9094-z
Debono M, Barnhart M, Carrell CB, Hoffmann JA, Occolowitz JL, Abbott BJ, Fukuda DS, Hamill RL, Biemann K, Herlihy WC (1987) A21978C, a complex of new acidic peptide antibiotics: isolation, chemistry, and mass spectral structure elucidation. J Antibiot (Tokyo) 40:761–777. doi:10.7164/antibiotics.40.761
Delgado ML, O’Connor JE, Azorı I, Renau-piqueras J, Gil ML, Gozalbo D (2001) The glyceraldehyde-3-phosphate dehydrogenase polypeptides encoded by the genes are also cell wall proteins. Microbiol 147:411–417. doi:10.1099/00221287-147-2-411
Delgado ML, Gil ML, Gozalbo D (2003) Starvation and temperature upshift cause an increase in the enzymatically active cell wall-associated glyceraldehyde-3-phosphate dehydrogenase protein in yeast. FEMS Yeast Res 4:297–303. doi:10.1016/S1567-1356(03)00159-4
Fernandes PA, Keen JN, Findlay JBC, Moradas-Ferreira P (1992) A protein homologous to glyceraldehyde-3-phosphate dehydrogenase is induced in the cell wall of a flocculent Kluyveromyces marxianus. Biochim Biophys Acta 1159:67–73. doi:10.1016/0167-4838(92)90076-P
Gil ML, Villamón E, Monteagudo C, Gozalbo D, Martínez JP (1999) Clinical strains of Candida albicans express the surface antigen glyceraldehyde 3-phosphate dehydrogenase in vitro and in infected tissues. FEMS Immunol Med Microbiol 23:229–234, doi:http://dx.doi.org/10.1111/j.1574-695X.1999.tb01243.x
Harris F, Dennison SR, Phoenix DA (2009) Anionic antimicrobial peptides from eukaryotic organisms. Curr Protein Pept Sci 10:585–606. doi:10.2174/138920309789630589
Izquierdo E, Horvatovich P, Marchioni E, Aoude-Werner D, Sanz Y, Ennahar S (2009) 2-DE and MS analysis of key proteins in the adhesion of Lactobacillus plantarum, a first step toward early selection of probiotics based on bacterial biomarkers. Electroph 30:949–956. doi:10.1002/elps.200800399
Jin X, Mei H, Li X, Ma Y, Zeng A, Wang Y, Lu X, Chu F (2010) Apoptosis-inducing activity of the antimicrobial peptide cecropin of Musca domestica in human hepatocellular carcinoma cell line BEL-7402 and the possible mechanism. Acta Biochim Biophys Sin 42:259–265. doi:10.1093/abbs/gmq021.Advance
Kemsawasd V, Branco P, Almeida MG, Caldeira J, Albergaria H, Arneborg N (2015) Cell-to-cell contact and antimicrobial peptides play a combined role in the death of Lachanchea thermotolerans during mixed-culture alcoholic fermentation with Saccharomyces cerevisiae. FEMS Microbiol Lett 362:1–8. doi:10.1093/femsle/fnv103
Laun P, Pichova A, Madeo F, Fuchs J, Ellinger A, Kohlwein S, Dawes I, Fröhlich KU, Breitenbach M (2001) Aged mother cells of Saccharomyces cerevisiae show markers of oxidative stress and apoptosis. Mol Microbiol 39:1166–1173. doi:10.1046/j.1365-2958.2001.02317.x
Ludovico P, Sousa MJ, Silva MT, Leão C, Côrte-Real M (2001) Saccharomyces cerevisiae commits to a programmed cell death process in response to acetic acid. Microbiol 147:2409–2415. doi:10.1099/00221287-147-9-2409
Madeo F, Fröhlich E, Fröhlich KU (1997) A yeast mutant showing diagnostic markers of early and late apoptosis. J Cell Biol 139:729–734. doi:10.1083/jcb.139.3.729
Madeo F, Fröhlich E, Ligr M, Grey M, Sigrist SJ, Wolf DH, Fröhlich KU (1999) Oxygen stress : a regulator of apoptosis in yeast. J Cell Biol 145:757–767. doi:10.1083/jcb.145.4.757
Malkoski M, Dashper SG, O’Brien-Simpson NM, Neil M, Talbo GH, Macris M, Cross KJ, Reynolds EC (2001) Kappacin, a novel antibacterial peptide from bovine milk. Antimicrob Agents Chemother 45:2309–2315. doi:10.1128/AAC.45.8.2309
Matsuzaki K (2009) Control of cell selectivity of antimicrobial peptides. Biochim Biophys Acta 1788:1687–1692. doi:10.1016/j.bbamem.2008.09.013
McAlister L, Holland MJ (1985) Differential expression of the three yeast glyceraldehyde-3- phosphate dehydrogenase genes. J Biol Chem 260:15019–15027
Nakajima H, Amano W, Kubo T, Fukuhara A, Ihara H, Azuma YT, Tajima H, Inui T, Sawa A, Takeuchi T (2009) Glyceraldehyde-3-phosphate dehydrogenase aggregate formation participates in oxidative stress-induced cell death. J Biol Chem 284:34331–34341. doi:10.1074/jbc.M109.027698
Nissen P, Arneborg N (2003) Characterization of early deaths of non- Saccharomyces yeasts in mixed cultures with Saccharomyces cerevisiae. Arch Microbiol 180:257–263. doi:10.1007/s00203-003-0585-9
Nissen-Meyer J, Holo H, Havarstein LS, Sletten K, Nes IF (1992) A novel lactococcal bacteriocin whose activity depends on the complementary action of two peptides. J Bacteriol 174:5686–5692
Pandey BK, Srivastava S, Singh M, Ghosh JK (2011) Inducing toxicity by introducing a leucine-zipper-like motif in frog antimicrobial peptide, magainin 2. Biochem J 436:609–620. doi:10.1042/BJ20110056
Park CB, Yi KS, Matsuzaki K, Kim MS, Kim SC (2000) Structure-activity analysis of buforin II, a histone H2A-derived antimicrobial peptide: the proline hinge is responsible for the cell-penetrating ability of buforin II. Proc Natl Acad Sci USA 97:8245–8250. doi:10.1073/pnas.150518097
Pérez-Nevado F, Albergaria H, Hogg T, Girio F (2006) Cellular death of two non-Saccharomyces wine-related yeasts during mixed fermentations with Saccharomyces cerevisiae. Int J Food Microbiol 108:336–345. doi:10.1016/j.ijfoodmicro.2005.12.012
Powers JP, Hancock RE (2003) The relationship between peptide structure and antibacterial activity. Peptides 24:1681–1691. doi:10.1016/j.peptides.2003.08.023
Reiter J, Herker E, Madeo F, Schmitt MJ (2005) Viral killer toxins induce caspase-mediated apoptosis in yeast. J Cell Biol 168:353–358. doi:10.1083/jcb.200408071
Sahoo S, Murgavel S, Devi IK, Vedamurthy GV, Gupta SC, Singh BP, Joshi P (2013) Glyceraldehyde-3-phosphate dehydrogenase of the parasitic nematode Haemonchus contortus binds to complement C3 and inhibits its activity. Parasite Immunol 35:457–467. doi:10.1111/pim.12058
Sang Y, Blecha F (2009) Porcine host defense peptides: expanding repertoire and functions. Dev Comp Immunol 33:334–343. doi:10.1016/j.dci.2008.05.006
Seo JK, Lee MJ, Go HJ, Park TH, Park NG (2012) Purification and characterization of YFGAP, a GAPDH-related novel antimicrobial peptide, from the skin of yellowfin tuna, Thunnus albacares. Fish Shellfish Immunol 33:743–752. doi:10.1016/j.fsi.2012.06.023
Silva A, Almeida B, Sampaio-Marques B, Reis MI, Ohlmeier S, Rodrigues F, Vale AD, Ludovico P (2011) Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) is a specific substrate of yeast metacaspase. Biochim Biophys Acta 1813:2044–2049. doi:10.1016/j.bbamcr.2011.09.010
Sirover MA (2005) New nuclear functions of the glycolytic protein, glyceraldehyde-3-phosphate dehydrogenase, in mammalian cells. J Cell Biochem 95:45–52. doi:10.1002/jcb.20399
Sirover MA (2011) On the functional diversity of glyceraldehyde-3-phosphate dehydrogenase: biochemical mechanisms and regulatory control. Biochim Biophys Acta 1810:741–751. doi:10.1016/j.bbagen.2011.05.010
Straus SK, Hancock RE (2006) Mode of action of the new antibiotic for Gram-positive pathogens daptomycin: comparison with cationic antimicrobial peptides and lipopeptides. Biochim Biophys Acta 1758:1215–1223. doi:10.1016/j.bbamem.2006.02.009
Sunaga K, Takahashi H, Chuang DM, Ishitani R (1995) Glyceraldehyde 3-phosphate dehydrogenase is over-expressed during apoptotic death of neuronal cultures and is recognized by a monoclonal-antibody against amyloid plaques from Alzheimer’s brain. Neurosci Lett 200:133–136. doi:10.1016/0304-3940(95)12098-O
Wagener J, Schneider JJ, Baxmann S, Kalbacher H, Borelli C, Nuding S, Küchler R, Wehkamp J, Kaeser MD, Mailänder-Sanchez D, Braunsdorf C, Hube B, Schild L, Forssmann WG, Korting HC, Liepke C, Schaller M (2013) A Peptide derived from the highly conserved protein GAPDH is involved in tissue protection by different antifungal strategies and epithelial immunomodulation. J Investigat Dermatol 133:144–153. doi:10.1038/jid.2012.254
Wong JH, Xia L, Ng TB (2007) A review of defensins of diverse origins. Curr Protein Pept Sci 8:446–459. doi:10.2174/138920307782411446
Acknowledgments
The present work was financed by FEDER funds through POFC-COMPETE and by national funds through Fundação para a Ciência e a Tecnologia (FCT) in the scope of project FCOMP-01-0124-FEDER-014055. P. Branco is the recipient of a PhD fellowship (SFRH/BD/89673/2012) funded by FCT, Portugal.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
The present work was funded by FCT (project grant number FCOMP-01-0124-FEDER-014055).
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 369 kb)
Rights and permissions
About this article
Cite this article
Branco, P., Francisco, D., Monteiro, M. et al. Antimicrobial properties and death-inducing mechanisms of saccharomycin, a biocide secreted by Saccharomyces cerevisiae . Appl Microbiol Biotechnol 101, 159–171 (2017). https://doi.org/10.1007/s00253-016-7755-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7755-6