Abstract
Mine water is an example of an extreme environment that contains a large number of diverse and specific bacteria. It is imperative to gain an understanding of these bacterial communities in order to develop effective strategies for the bioremediation of polluted aquatic systems. In this study, the high-throughput sequencing approach was used to characterize the bacterial communities in two different mine waters of South Africa: vanadium and gold mine water. Over 2629 operational taxonomic units (OTUs) were recovered from 15,802 reads of the 16S ribosomal RNA (rRNA) gene. They represented 8 phyla, 43 orders, 84 families and 105 genera. Proteobacteria and unclassified bacterial sequences were the most dominant. Apart from these, Firmicutes, Bacteroidetes, Actinobacteria, Candidate phylum OD1, Cyanobacteria, Verrucomicrobia and Deinococcus-Thermus were the recovered phyla, although their relative abundance differed between both the mine-water samples. Yet, diversity indices suggested that the bacterial communities inhabiting the vanadium mine water were more diverse than those in gold mine water. Interestingly, substantial percentages of the reads from either sample (58 % in vanadium and 17 % in gold mine water) could not be assigned to any phylum and remained unclassified, suggesting hitherto unidentified populations, and vast untapped microbial diversity. Overall, the results of this study exhibited bacterial community structures with high diversity in mine water, which can be explored further for their role in bioremediation and environmental management.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The tiny microbes that dominate the planet are ubiquitously distributed all over the biosphere, even tolerating a wide range of physicochemical stresses in extreme habitats. Their abundance and widespread presence in almost all ecosystems make them the most successful organisms on the planet. It is well known that microbes are able to control and sustain all possible life forms on earth by essentially maintaining all the biogeochemical cycles. The potential of microbes to command over metabolic and catabolic processes as well as the ability to adapt to changing environmental conditions or survive in hostile environments are universally accepted (Kyrpides et al. 2014). Studies have shown that mining industries are among the major sources of pollution as they produce large amounts of contaminants, which not only disperse into the surrounding environment through aerial deposition or water erosion but also cause pollution of the areas away from their original site (Maynaud et al. 2013).
Mining activities have played a major role in both economic development and environmental pollution in South Africa (Adler et al. 2007). Despite regulations related to mine-water management in South Africa, the continuing discharge of untreated mine water with varying levels of hazardousness causes major environmental problems (Oelofse 2009). Contamination of South African water sources with heavy metals is a widespread problem, especially in the vicinity of mines. Heavy metal contamination has serious consequences mainly for human and animal health. Heavy metals are dangerous as they tend to bioaccumulate in the tissues of receiving organisms through food chain, resulting in toxicity at higher concentration. South Africa is a semi-arid country, and the management of its precious water resources is essential to counteract pollution caused by mining industries. It is important to note that these unique environments harbour metabolically active microorganisms that are well adapted to the multiple environmental stresses they encounter on a daily basis. Analysis of microbial diversity in the mine waters may allow the quantification of the possible effects of heavy metals on different taxonomic groups and potential effects on the biological properties of the receiving ecosystem. It is now generally accepted that more than 95 % of microbes living in the environment are yet to be discovered and explored in physiological terms (Hennecke et al. 2013). Mine water is an example of an extreme environment that contains a large number of very diverse and specific bacteria (Kamika and Momba 2013). Complex interactions between diverse bacterial populations are known to exist; diverse bacteria coexist to cope with the extreme conditions prevailing in the mine waters. Characterizing bacterial communities in the mine waters containing heavy metals would provide valuable data for assessing the various metal-tolerant microbial communities existing in these extreme environments.
The standard microbiological culturing approaches are effective techniques for assessing the microbial diversity, but the majority of the bacteria cannot be cultured in the laboratory (Hugenholtz et al. 1998; Whitman et al. 1998). The accurate assessment of the microbial communities in any environmental medium could be achieved using molecular biological techniques, and findings significantly promote our understanding of the microbial communities (Liu et al. 1997; Muyzer et al. 1993; Quince et al. 2009). 16S ribosomal RNA (rRNA) gene is considered as molecular signature universally present among all bacteria and is being widely used in next-generation sequencing approach to decipher the bacterial diversity of several habitats, e.g. bacteria associated with Lucilia spp. (Singh et al. 2014), tobacco waste extract (Liu et al. 2014a, b14), styrene-degrading biofilters (Portune et al. 2014), activated sludge (Zhao et al. 2014), high-temperature biogas reactors (Röske et al. 2014), phyllosphere of rice (Ren et al. 2014), river receiving effluents from swine farm and farmhouse restaurant (Lu and Lu 2014), endophytes of sugar beet (Shi et al. 2014), petroleum reservoir (Wang et al. 2014), intestinal environment of rats (An et al. 2014), mine tailings (Liu et al. 2014a, b), heavy metals polluted soil (Gołębiewski et al. 2014), soil horizons and drain sediments of coastal acid sulfate landscape (Stroud et al. 2014). The first 16S rRNA gene-based microbial analysis of mine-water systems was done in the mid-1990s (Goebel and Stackebrandt 1994). Subsequently, some of the mine-water systems from different geographical locations were assayed using molecular techniques, e.g. Iron Mountain in California, USA (Bond et al. 2000; Druschel et al. 2004) and the Rio Tinto basin in south-western Spain (Gonzalez-Toril et al. 2003; Garcia-Moyano et al. 2007). These studies, however, were based on a single mining environment, and only limited sequencing depths were achieved using a standard clone library approach. The high-throughput sequencing can be easily implemented at low cost by using the next-generation sequencing platforms (Ye and Zhang 2013). Recently, the bacterial communities of mine waters have been assessed using the high-throughput approach (Kamika and Momba 2013; Kuang et al. 2013). The Illumina MiSeq personal sequencer is one of the popular high-throughput sequencing systems which can generate up to 4.6 GB of data per run. This approach has been applied to decipher the microbial community in distinct samples, such as biofilm in urban drinking water distribution system (Wu et al. 2014), upflow anaerobic sludge blanket reactor (Zhang et al. 2014), polychlorinated biphenyl contaminated environments (Su et al. 2014), river water (Staley et al. 2013), clinical specimen (Reuter et al. 2013; Loman et al. 2013), beer and wine (Bokulich and Mills 2013), the gut microbiome (Markle et al. 2013), ancient DNA (Gansauge and Meyer 2013), soil, host-associated environments (Caporaso et al. 2012) and the human microbiome (Kuczynski et al. 2012). One of the exciting possibilities provided by this technology is the accurate estimation of the diversity and structure of microbial communities, which may lead to a better understanding of their ecology in the mine water.
Despite an increase in mining-related studies over the past decades, our current knowledge regarding the prevalence and complexity of bacterial communities is still incomplete, partly due to the lack of reproducible screening methods. The present study compares two geographically isolated mine waters using high-throughput sequencing of the 16S rRNA gene using a 500-bp amplicon generated from the universal bacterial primers 27 F and 518R (Lane 1991). Assessment of the microbial communities in mine waters will not only provide information about their ecophysiology and the ecophysiological adaptations required to cope with extreme environments; knowledge of these complex communities will also assist in developing and providing sustainable and reliable remediation solutions (Mohapatra et al. 2011). The objectives of this research were to classify the bacteria present in the mine waters from South Africa and to compare the bacterial communities in the samples across geographic locations. To date, there has been no report of using high-throughput/next-generation sequencing to compare the complexity of the bacterial communities present in the different mine waters of South Africa. To our knowledge, this study describes the first attempt to explore and compare the bacterial diversity and community structure in selected South African vanadium and gold mine waters of South Africa using next next-generation sequencing and to provide useful information on the bacteria inhabiting the mine-water ecosystems. It was hypothesized that bacterial populations would be distinct based on geographic location and physicochemical parameters.
Materials and methods
Chemicals sourced
Bacterial DNA isolation kits were supplied by Epigenetics, USA, while the DreamTaq™ Green PCR Master Mix for PCR and DNA markers were purchased from Fermentas (Thermo Fisher Scientific, Pittsburgh PA, USA). TAE buffer and molecular grade agarose were sourced from Sigma-Aldrich (Steinheim, Germany).
Mine-water sample collection
Mine-water samples from vanadium and gold mines were collected in 1-L sterile plastic sampling bottles in triplicate from the evaporation dam of Vanchem Vanadium Products Ltd., Ferrobank, Mpumalanga and Mogale Gold Mine, Randfontein, Gauteng, South Africa, respectively. No specific permit was needed for the collection of the wastewater samples in the described sample area, and this study did not involve endangered or protected species. However, the responsible officers, namely the Process Development Manager at the chemical plant of Vanchem Vanadium Products Ltd. and the Water Plant Foreman at Mogale Gold Mine, were informed to assist the research team with the collection of mine-water samples. Samples were kept in a cooler box (4 °C) while being transported to the laboratory for further analysis.
Physicochemical analysis
For physicochemical analyses, samples collected were thoroughly mixed prior to filtration and No. 1 filter papers (Whatman) were used. The profile of the filtered samples was determined in terms of the chemical oxygen demand (COD), dissolved oxygen (DO), pH, electrical conductivity (EC) and heavy metals. The COD concentration was measured using closed reflux methods as described in Standard Methods (APHA 2001), while the pH and the DO were analyzed using a pH probe (Model: PHC101, HACH) and DO probe (Model: LDO, HACH), respectively. Heavy metals were determined using the inductively couple plasma optical emission spectrometer (ICP-OES) (Spectro Arcos, Kleve Germany). The limits of detection (LOD) varied between 10 and 60 mg/L depending on the elements.
Metagenomic DNA extraction
For each replicate, a 500-mL sample was filtered using filter paper with a pore size of 0.22 μm (Whatman filter No. 1) and the residues on the filter papers containing the microbes were washed with 1X PBS. The metagenomic DNA was extracted independently for all the samples with the ZR Fungal/Bacterial DNA MiniPrep™ Kit (Zymo Research Corporation), as per manufacturer’s instruction. Finally, the DNA was eluted in 50 μL of MilliQ water. The integration of the metagenomic DNA was assessed on the 0.8 % (w/v) agarose gel, and the DNA was quantified using a NanoDrop spectrophotometer (Nanodrop 2000, Thermo Scientific, Japan).
PCR amplification and sequencing
The PCR reaction was performed in triplicate on all the extracted DNA samples using the universal primer 27F and 518R. These primers have been reported to amplify approximately 500 bp of the 16S rRNA gene sequence targeting variable regions V1 and V3 of the 16S rRNA gene (Lane 1991). Each PCR reaction contained 25 μL of 2X DreamTaq™ Green Master Mix (DNA polymerase, dNTPs and 4 mM MgCl2), 22 μL of nuclease-free water, 1 μL of forward primer (0.2 μM) and 1 μL of reverse primer (0.2 μM), template DNA 50–100 ng up to the final reaction volume of 50 μL. The following cycling parameters were used: initial denaturation step at 94 °C for 5 min, followed by 30 cycles of 94 °C for 1 min, 55 °C for 30 s and 72 °C for 1 min 30 s, with a final 10-min extension at 72 °C, followed by final incubation at 4 °C. The PCR products were loaded onto 1.5 % (w/v) agarose gel and then visualized under a UV transilluminator (InGenius Bio Imaging System, Syngene, Cambridge, UK). The PCR amplicons were pooled together for the respective samples and purified using the QIAquick PCR purification kit (Qiagen, Germany). The DNA concentrations were quantified by using a NanoDrop spectrophotometer. The purified amplicons were pooled at equal concentrations for both different samples. The pooled samples were sequenced at Inqaba Biotechnology Industries, South Africa on the Illumina MiSeq platform.
Sequence analysis
The raw sequences were processed in MG-RAST pipeline (http://metagenomics.anl.gov/) for quality control (QC) assessment which removes artificial replicate sequences produced by sequencing artefacts and low quality sequences (Meyer et al. 2008). Although 16S rRNA-specific primers were used, a few non-ribosomal sequences were obtained. All QC passed sequences were pre-screened using qiime-uclust for at least 70 % identity to ribosomal sequences from the following RNA databases—Greengenes, LSU, SSU and RDP—and the non-ribosomal sequences were removed from the each data set. The ribosomal sequences were screened for chimeric sequences using UCHIME (Edgar et al. 2011). All the chimeric sequences were removed from further analysis. The entire non-chimeric rRNA sequences were analyzed using Ribosomal Database Project (RDP) Classifier tool of RDP pyrosequencing pipeline (Wang et al. 2007) with a confidence threshold of 80 %. These sequences were uploaded on the RDP Align tool and aligned in the same pipeline, and the cluster files were generated for each sample with the RDP Complete Linkage Clustering tool. These cluster files were then used as input to generate rarefaction curves by using the RDP Rarefaction tool. Genetic distance was calculated, and sequences were clustered into operational taxonomic units (OTUs) using 0.03 and 0.20 dissimilarity cut-off levels. The diversity index Shannon and richness estimator Chao1 were calculated for each sample at both distances. The relative abundance (%) of individual taxa within each community was calculated by comparing the number of sequences assigned to a specific taxon against the number of total sequences obtained for that sample. Data was made publicly available at the NCBI Sequence Read Archive under the accession number SRR1584394.
Results
Physicochemical profile of mine waters
The gold mine waters were more acidic and saline than the vanadium mine waters, but the DO and COD values were found to be comparatively lower for gold mine-water samples (Table 1). The following heavy metals were detected in the respective gold mine-water samples: V, Ni, Cd, As, Cr, Hg and Co. However, in vanadium mine-water samples, Cr and Hg could not be detected among these heavy metals.
Sequencing
The DNA preparations were of high quality, as their A260/280 ratios were approximately 1.8 for all samples, indicating high purity, and they performed well as PCR templates. In total, 15,802 reads were obtained (Table 2); out of which, 14,557 (92.13 %) met the quality criteria. These sequences were searched for identity with rRNA sequences, and 14,492 sequences showed ribosomal origin. The 2618 chimeric sequences were removed, and 11,874 (75.14 % of the total number) high-quality non-chimeric rRNA reads were included in the downstream analysis. The mean read length ranged between 290 and 268 bp for vanadium and gold mine-water samples, respectively. All the filtered reads were successfully classified as belonging to the domain bacteria.
Community species richness and diversity indices
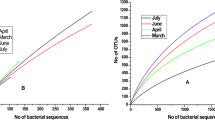
The communities were moderately sampled at the genetic distance level of 0.03, indicating that many more reads would be required to capture all the diversity, but at the distance of 0.2, it can be observed that major phyla were recovered (Fig. 1). More OTUs were observed for the metagenome of vanadium mine-water samples than in the gold mine-water samples (Table 3). Both the observed and estimated richness (Chao1) were higher for the vanadium mine-water samples. The parametric diversity index Shannon also indicated higher bacterial diversity in vanadium mine-water samples.
Bacterial community structure
In total, 8 bacterial phyla were observed apart from unclassified bacterial sequences. Proteobacteria, Firmicutes, Bacteroidetes, Actinobacteria, Candidate phyla OD1 and Cyanobacterial sequences were observed in both types of mine-water samples. Representatives of Verrucomicrobia were only found in vanadium water while Deinococcus-Thermus only in gold mine water (Fig. 2). Proteobacteria was the most abundant phylum in gold mine water (75.59 %), whereas the majority of sequences from vanadium mine-water samples could not be classified even up to the phylum level and unclassified bacterial sequences dominated the vanadium mine-water sample (58.37 %) (Fig. 2, Table S1). Notable differences in the proportions of these phyla (i.e. composition) were observed between both mine-water samples assayed (Fig. 2). For instance, in gold mine water, the unclassified bacteria were the second most dominant category (17.64 %) followed by Actinobacteria (3.37 %), Bacteroidetes (1.84 %), Firmicutes (1.43 %), OD1 (0.04 %) and Deinococcus-Thermus (0.02 %). However, unclassified bacteria were found to be dominant in vanadium mine water, while the Proteobacteria (23.28 %) were the second most dominant phyla followed by Bacteroidetes (10.38 %), OD1 (5.58 %), Cyanobacteria (1.84 %), Firmicutes (0.15 %) and Verrucomicrobia (0.01 %). The most abundant class was Alphaproteobacteria accounting for 35.51 % in gold mine water and 11.75 % in vanadium mine water. Gammaproteobacteria and Betaproteobacteria were found in both types of mine-water samples, whereas Deltaproteobacteria was found exclusively in gold mine water. Classification of the reads into lower taxonomic levels revealed extremely diverse bacterial communities in both types of mine-water samples, with up to 20 orders, 39 families and 48 genera detected in the vanadium mine-water samples, and 23 orders, 45 families and 57 genera detected in the gold mine-water samples (Table S1).
Taxonomic distribution of different bacterial phylogenetic groups in vanadium and gold mine water. Analysis of 16S rRNA gene sequences was done in comparison with the RDP II database. The percentages of the phylogenetically classified sequences are plotted on the y-axis. VMW vanadium mine water, GMW gold mine water
The rank abundance curve represented taxonomic richness and evenness showing the abundance of previously unclassified sequences (Fig. 3, Table S2). In the vanadium mine water, the unclassified members of bacteria were most abundant followed by unclassified Flavobacteriaceae, genus OD1_genera_incertae_sedis, Marinobacter sp. and Thalassospira sp. and these groups accounted for 77.71 % of total sequences. The five most abundant categories in gold mine waters were Brevundimonas sp., Pseudomonas sp., unclassified bacteria, unclassified Proteobacteria and unclassified Gammaproteobacteria, which covered 79.48 % of total sequences. In total, 82 % of sequences in vanadium mine water could not be classified up to genus level. However, in case of gold mine water, 42 % of sequences could not be assigned to any genus (Table S2). The most dominant families among classified sequences in vanadium mine water were Alteromonadaceae (Gammaproteobacteria) and Rhodospirillaceae (Alphaproteobacteria) whereas Caulobacteraceae (Alphaproteobacteria) and Pseudomonadaceae (Gammaproteobacteria) were most abundant in gold mine-water samples. The analysis based on the occurrence and relative abundance of different families suggested a strong dissimilarity among the bacterial communities in both types of mine-water samples (Fig. 3, Table S2). It was observed that many different bacterial communities resided in both mine waters with least community overlap between these two samples. Only 25 sequences were found to be common to both mine waters. These common sequences belonged to the categories unclassified bacteria (11), Pseudomonas sp. (4), Brevundimonas sp. (3), unclassified Gammaproteobacteria (2), OD1_genera_incertae_sedis (2), unclassified Flavobacteriaceae (1), unclassified Rhodobacteraceae (1) and unclassified Proteobacteria (1).
Discussion
In this study, the bacterial community structure and species richness in the metagenome of two mine waters located in the Gauteng and Mpumalanga Provinces of the Republic of South Africa were analyzed. The physicochemical analysis revealed that both types of mine-water samples were more acidic than the permissible minimum limit of pH 5.5 set by the South African National Water Act, No. 36 of 1998. The electrical conductivities of the mine waters were found to be higher than the maximum permissible limit of 250 mS/cm for effluent discharged into the receiving water bodies. If this water is discharged directly into water bodies, it might pose a major risk factor to existing community structure and diversity of microbial populations (Kuang et al. 2013). It has been reported that the low pH in mine waters increases the solubility of many metals, and coupled with low availability of organic carbon, it presents extremely challenging and hostile environmental growth conditions (Dopson et al. 2003; Slonczewski et al. 2009). The heavy metals such as lead, cadmium, mercury, silver and chromium have been reported to be toxic even at low concentrations and have no known beneficial effects on bacterial cells (Nies 2004). Microbial community composition is highly correlated to physicochemical parameters such as pH and metal ion concentrations, as described by previous investigators (Johnson and Hallberg 2003; Imarla et al. 2006).
The rarefaction analysis suggests that microbial diversity could be even higher than observed and similar results were reported in other extreme ecosystems, e.g. hydrothermal chimneys (Brazelton et al. 2010), acidic hot spring (Bohorquez et al. 2012) and acid mine drainage (Kuang et al. 2013). The diversity indices (Table 3) indicated higher bacterial diversity in vanadium mine-water samples, and this could be due to the fact that vanadium mine waters were less acidic and saline than the gold mine waters.
Microbial survival in mine waters depends on intrinsic physiological and genetic adaptation. Various types of resistance mechanisms are used by microbes in order to cope with heavy metals (Nies 2004). The present study is the first report to present a comprehensive comparative sequence analysis of the bacterial diversity from different mine waters of South Africa by high-throughput sequencing method. The member of the phylum Proteobacteria are frequently observed in mines as the gold mine water is dominated by Proteobacteria, and in vanadium mine water, they are the second most dominant after unclassified bacterial sequences. The abundance of Proteobacteria was also found in a Chinese copper mine (He et al. 2007) as well as in deep mines in South Africa (Raji et al. 2008). The dominance of Proteobacteria was also reported in a previous study on the microbial diversity of the effluents of the same vanadium mines (Kamika and Momba 2013) as well as on acid mine drainage sites across Southeast China (Kuang et al. 2013). Nevertheless, in this study, we found that unclassified bacterial sequences were more abundant than the Proteobacteria in this vanadium mine water which supports the finding that there is a high abundance of unculturable microbes in any environment (Aslam et al. 2010). Compared with highly diverse Proteobacteria communities, the other communities in both the mine-water samples were less complex and only Bacteroidetes accounted for more than 10 % of the reads in vanadium mine water (Table S1). It has been stated that most of the bacteria inhabiting the metal-contaminated water systems are able to develop various types of resistance mechanisms to adapt to and resist metal toxicity; as a result, microbial diversity remains high (Gough and Stahl 2011; Berg et al. 2012). In the vanadium mine water, the occurrence of Marinobacter sp. supported this fact as the Marinobacter sp. isolated from vanadium mine effluents showed high tolerance to vanadium ions (Kamika and Momba 2013). It should be mentioned that the use of culture-independent methodologies to assess the microbial diversity of any gold mine water in South Africa has never been reported previously.
The difference in microbial communities between these mine waters (Fig. 3, Table S2) can probably be attributed to the effects of the particular physicochemical parameters of the mine waters. These data are indicative of substantial heterogeneity in microbial community composition and provide valuable information about the impacts of various stresses on the biological integrity of ecosystems. The mine waters harbour indigenous microorganisms that are physiologically adapted to tolerate the prevailing extreme conditions and are able to maintain their viability.
The present study reports the comparative analysis of geographically isolated mine water microbes using a high-throughput sequencing approach to elucidate the diversity and community structure of the bacteria that are physiologically adapted to tolerate high concentrations of heavy metals. In summary, mine-water samples were found to harbour unique bacterial communities that are well adapted to this extreme environment. This study provides an inventory of the metal-tolerant bacteria which will be helpful in the determination of culturing conditions if the intention is to isolate and identify them by culture techniques in laboratory settings to determine specific metabolic capabilities. Future investigations related to the functional analysis of the microbial community in mine waters based on metatranscriptomics will likely reveal a more pronounced understanding of their ecophysiological role.
References
Adler RA, Claassen M, Godfrey L, Turton AR (2007) Water, mining, and waste: an historical and economic perspective on conflict management in South Africa. Econ Peace Secur J 2:33–44
An C, Kuda T, Yazaki T, Takahashi H, Kimura B (2014) Caecal environment of rats fed far East Asian-modelled diets. Appl Microbiol Biotechnol. doi:10.1007/s00253-014-5535-5538
APHA (2001) Standard methods for the examination of water and wastewater, 20th edn. American Public Health Association (APHA), Washington DC
Aslam Z, Yasir M, Khaliq A, Matsui K, Chung YR (2010) Too much bacteria still unculturable. Crop Environ 1:59–60
Berg J, Brandt KK, Al-Soud WA, Holm PE, Hansen LH, Sørensen SJ, Nybroe O (2012) Long-term Cu exposure selects for Cu-tolerant bacterial communities with altered composition, but unaltered richness. Appl Environ Microbiol 78:7438–7446
Bohorquez LC, Delgado-Serrano L, López G, Osorio-Forero C, Klepac-Ceraj V, Kolter R, Junca H, Baena S, Zambrano MM (2012) In-depth characterization via complementing culture-independent approaches of the microbial community in an acidic hot spring of the Colombian Andes. Microb Ecol 63:103–115
Bokulich NA, Mills DA (2013) Improved selection of internal transcribed spacer-specific primers enables quantitative, ultra-high-throughput profiling of fungal communities. Appl Environ Microbiol 79:2519–2526
Bond PL, Smriga SP, Banfield JF (2000) Phylogeny of microorganisms populating a thick, subaerial, predominantly lithotrophic biofilm at an extreme acid mine drainage site. Appl Environ Microbiol 66:3842–3849
Brazelton WJ, Ludwig KA, Sogin ML, Andreishcheva EN, Kelley DS, Shen C-C, Edwards RL, Baross JA (2010) Archaea and bacteria with surprising microdiversity show shifts in dominance over 1,000-year time scales in hydrothermal chimneys. Proc Natl Acad Sci U S A 107:1612–1617
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N, Owens SM, Betley J, Fraser L, Bauer M, Gormley N, Gilbert JA, Smith G, Knight R (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624
Dopson M, Baker-Austin C, Koppineedi PR, Bond PL (2003) Growth in sulfidic mineral environments: metal resistance mechanisms in acidophilic micro-organisms. Microbiology 149:1959–1970
Druschel GK, Baker BJ, Gihring TM, Banfield JF (2004) Acid mine drainage biogeochemistry at Iron Mountain, California. Geochem Trans 5:13–32
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200
Gansauge M-T, Meyer M (2013) Single-stranded DNA library preparation for the sequencing of ancient or damaged DNA. Nat Protoc 8:737–748
Garcia-Moyano A, Gonzalez-Toril E, Aguilera A, Amils R (2007) Prokaryotic community composition and ecology of floating macroscopic filaments from an extreme acidic environment, Rio Tinto (SW, Spain). Syst Appl Microbiol 30:601–614
Goebel BM, Stackebrandt E (1994) Cultural and phylogenetic analysis of mixed microbial populations found in natural and commercial bioleaching environments. Appl Environ Microbiol 60:1614–1621
Gołębiewski M, Deja-Sikora E, Cichosz M, Tretyn A, Wróbel B (2014) 16S rDNA pyrosequencing analysis of bacterial community in heavy metals polluted soils. Microb Ecol 67:635–647
Gonzalez-Toril E, Llobet-Brossa E, Casamayor EO, Amann R, Amils R (2003) Microbial ecology of an extreme acidic environment, the Tinto River. Appl Environ Microbiol 69:4853–4865
Gough HL, Stahl DA (2011) Microbial community structures in anoxic freshwater lake sediment along a metal contamination gradient. ISME J 5:543–558
He Z, Xie X, Xiao S, Li J, Qiu G (2007) Microbial diversity of mine water at Zhong Tiaoshan copper mine, China. J Basic Microbiol 47:485–495
Hennecke H, Kjelleberg S, Brussaard C (2013) Molecular insights into environmental microbes. Fems Microbiol Rev 37:285
Hugenholtz P, Goebel BM, Pace NR (1998) Impact of culture independent studies on the emerging phylogenetic view of bacterial diversity. J Bacteriol 180:4765–4774
Imarla T, Hector SB, Deane SM, Rawlings DE (2006) Resistance determinants of a highly arsenic-resistant strain of Leptospirillum ferriphilum isolated from a commercial biooxidation tank. Appl Environ Microbiol 72:2247–2253
Johnson DB, Hallberg KB (2003) The microbiology of acidic mine waters. Res Microbiol 154:466–473
Kamika I, Momba MNB (2013) Microbial diversity of Emalahleni mine water in South Africa and tolerance ability of the predominant organism to vanadium and nickel. Plos One 9:e86189
Kuang J-L, Huang L-N, Chen L-X, Hua Z-S, Li S-J, Hu M, Li J-T, Shu W-S (2013) Contemporary environmental variation determines microbial diversity patterns in acid mine drainage. ISME J 7:1038–1050
Kuczynski J, Lauber CL, Walters WA, Parfrey LW, Clemente JC, Gevers D, Knight R (2012) Experimental and analytical tools for studying the human microbiome. Nat Rev Genet 13:47–58
Kyrpides NC, Hugenholtz P, Eisen JA, Woyke T, Göker M, Parker CT, Amann R, Beck BJ, Chain PS, Chun J, Colwell RR, Danchin A, Dawyndt P, Dedeurwaerdere T, DeLong EF, Detter JC, De Vos P, Donohue TJ, Dong XZ, Ehrlich DS, Fraser C, Gibbs R, Gilbert J, Gilna P, Glöckner FO, Jansson JK, Keasling JD, Knight R, Labeda D, Lapidus A, Lee JS, Li WJ, Ma J, Markowitz V, Moore ER, Morrison M, Meyer F, Nelson KE, Ohkuma M, Ouzounis CA, Pace N, Parkhill J, Qin N, Rossello-Mora R, Sikorski J, Smith D, Sogin M, Stevens R, Stingl U, Suzuki K, Taylor D, Tiedje JM, Tindall B, Wagner M, Weinstock G, Weissenbach J, White O, Wang J, Zhang L, Zhou YG, Field D, Whitman WB, Garrity GM, Klenk HP (2014) Genomic encyclopedia of bacteria and archaea: sequencing a myriad of type strains. PLoS Biol 12:e1001920
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, Chichester, pp 115–175
Liu W-T, Marsh TL, Cheng H, Forney LJ (1997) Characterization of microbial diversity by determining terminal restriction fragment length polymorphisms of genes encoding 16S rRNA. Appl Environ Microbiol 63:4516–4522
Liu H, He H, Cheng C, Liu J, Shu M, Jiao Y, Tao F, Zhong W (2014a) Diversity analysis of the bacterial community in tobacco waste extract during reconstituted tobacco process. Appl Microbiol Biotechnol. doi:10.1007/s00253-014-5960-8
Liu J, Hua Z-S, Chen L-X, Kuang J-L, Li S-J, Shu W-S, Huang L-N (2014b) Correlating microbial diversity patterns with geochemistry in an extreme and heterogeneous environment of mine tailings. Appl Environ Microbiol 80:3677–3686
Loman NJ, Constantinidou C, Christner M, Rohde H, Chan JZ, Quick J, Weir JC, Quince C, Smith GP, Betley JR, Aepfelbacher M, Pallen MJ (2013) A culture-independent sequence-based metagenomics approach to the investigation of an outbreak of shiga-toxigenic Escherichia coli O104:H4. JAMA 309:1502–1510
Lu X-M, Lu P-Z (2014) Diversity, abundance, and spatial distribution of riverine microbial communities response to effluents from swine farm versus farmhouse restaurant. Appl Microbiol Biotechnol 98:7597–7608
Markle JG, Frank DN, Mortin-Toth S, Robertson CE, Feazel LM, Rolle-Kampczyk U, Von Bergen M, McCoy KD, Macpherson AJ, Danska JS (2013) Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science 1(339):1084–1088
Maynaud G, Brunel B, Mornico D, Durot M, Severac D, Dubois E, Navarro E, Cleyet-Marel J-C, Quéré AL (2013) Genome-wide transcriptional responses of two metal-tolerant symbiotic Mesorhizobium isolates to zinc and cadmium exposure. BMC Genomics 14:292
Meyer F, Paarmann D, D’Souza M, Olson R, Glass EM, Kubal M, Paczian T, Rodriguez A, Stevens R, Wilkie A, Wilkening J, Edwards RA (2008) The metagenomics RAST server—a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinforma 9:386–394
Mohapatra BR, Gould WD, Dinardo O, Koren DW (2011) Tracking the prokaryotic diversity in acid mine drainage-contaminated environments: a review of molecular methods. Miner Eng 24:709–718
Muyzer G, De Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Nies DH (2004) Metals and their compounds in the environment. Part II. In: Anke K, Ihnat M, Stoeppler M (eds) The elements: essential and toxic effects on microorganisms. Wiley-VCH, Weinheim, Germany
Oelofse SHH (2009) Mine water pollution—acid mine decant, effluent and treatment: a consideration of key emerging issues that may impact the state of the environment. In: Krishna CS (ed) Mining: environment and health concerns, 1st edn. The Icfai University Press, India, pp 84–91
Portune KJ, Pérez MC, Álvarez-Hornos FJ, Gabaldón C (2014) Investigating bacterial populations in styrene-degrading biofilters by 16S rDNA tag pyrosequencing. Appl Microbiol Biotechnol. doi:10.1007/s00253-014-5868-3
Quince C, Lanzén A, Curtis TP, Davenport RJ, Hall N, Head IM, Read LF, Sloan WT (2009) Accurate determination of microbial diversity from 454 pyrosequencing data. Nat Methods 6:639–641
Raji AI, Moller C, Litthauer D, van Heerden E, Piater LA (2008) Bacterial diversity of biofilm samples from deep mines in South Africa. Biokemistri 20:53–62
Ren G, Zhang H, Lin X, Zhu J, Jia Z (2014) Response of phyllosphere bacterial communities to elevated CO2 during rice growing season. Appl Microbiol Biotechnol 98:9459–9471
Reuter S, Ellington MJ, Cartwright EP, Köser CU, Török ME, Gouliouris T, Harris SR, Brown NM, Holden MT, Quail M, Parkhill J, Smith GP, Bentley SD, Peacock SJ (2013) Rapid bacterial whole-genome sequencing to enhance diagnostic and public health microbiology. JAMA Intern Med 173:1397–1404
Röske I, Sabra W, Nacke H, Daniel R, Zeng A-P, Antranikian G, Sahm K (2014) Microbial community composition and dynamics in high-temperature biogas reactors using industrial bioethanol waste as substrate. Appl Microbiol Biotechnol 98:9095–9106
Shi Y, Yang H, Zhang T, Sun J, Lou K (2014) Illumina-based analysis of endophytic bacterial diversity and space-time dynamics in sugar beet on the north slope of Tianshan mountain. Appl Microbiol Biotechnol 98:6375–6385
Singh B, Crippen TL, Zheng L, Fields AT, Yu Z, Ma Q, Wood TK, Dowd SE, Flores M, Tomberlin JK, Tarone AT (2014) A metagenomic assessment of the bacteria associated with Lucilia sericata and Lucilia cuprina (Diptera: Calliphoridae). Appl Microbiol Biotechnol. doi:10.1007/s00253-014-6115-7
Slonczewski JL, Fujisawa M, Dopson M, Krulwich TA, Robert KP (2009) Cytoplasmic pH measurement and homeostasis in bacteria and archaea. Adv Microb Physiol 55:1–79
Staley C, Unno T, Gould TJ, Jarvis B, Phillips J, Cotner JB, Sadowsky MJ (2013) Application of Illumina next-generation sequencing to characterize the bacterial community of the Upper Mississippi River. J Appl Microbiol 115:1147–1158
Stroud JL, Low A, Collins RN, Manefield M (2014) Metal(loid) bioaccessibility dictates microbial community composition in acid sulfate soil horizons and sulfidic drain sediments. Environ Sci Technol 48:8514–8521
Su X, Zhang Q, Hu J, Hashmi MZ, Ding L, Shen C (2014) Enhanced degradation of biphenyl from PCB-contaminated sediments: the impact of extracellular organic matter from Micrococcus luteus. Microbiol Biotechnol. doi:10.1007/s00253-014-6108-6
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Wang L-Y, Ke W-J, Sun X-B, Liu J-F, Gu J-D, Mu B-Z (2014) Comparison of bacterial community in aqueous and oil phases of water-flooded petroleum reservoirs using pyrosequencing and clone library approaches. Appl Microbiol Biotechnol 98:4209–4221
Whitman WB, Coleman DC, Wiebe WJ (1998) Prokaryotes: the unseen majority. Proc Natl Acad Sci U S A 95:6578–6583
Wu H, Zhang J, Mi Z, Xie S, Chen C, Zhang X (2014) Biofilm bacterial communities in urban drinking water distribution systems transporting waters with different purification strategies. Appl Microbiol Biotechnol. doi:10.1007/s00253-014-6095-7
Ye L, Zhang T (2013) Bacterial communities in different sections of a municipal wastewater treatment plant revealed by 16S rDNA 454 pyrosequencing. Appl Microbiol Biotechnol 97:2681–2690
Zhang Y, Wang X, Hu M, Li P (2014) Effect of hydraulic retention time (HRT) on the biodegradation of trichloroethylene wastewater and anaerobic bacterial community in the UASB reactor. Appl Microbiol Biotechnol. doi:10.1007/s00253-014-6096-6
Zhao D, Huang R, Zeng J, Yu Z, Liu P, Cheng S, Wu QL (2014) Pyrosequencing analysis of bacterial community and assembly in activated sludge samples from different geographic regions in China. Appl Microbiol Biotechnol 98:9119–9128
Acknowledgments
The authors are grateful to the two South African mining companies for allowing the researchers to sample their evaporation dams. The financial support of National Research Foundation (NRF) and the Department of Science and Technology, South Africa for carrying out this project is thankfully acknowledged. JK gratefully acknowledges the Postdoctoral Fellowship from Tshwane University of Technology, Pretoria, South Africa.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 13.8 kb)
Rights and permissions
About this article
Cite this article
Keshri, J., Mankazana, B.B.J. & Momba, M.N.B. Profile of bacterial communities in South African mine-water samples using Illumina next-generation sequencing platform. Appl Microbiol Biotechnol 99, 3233–3242 (2015). https://doi.org/10.1007/s00253-014-6213-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-6213-6