Abstract
An agar-degrading bacterium, Catenovulum sp. X3, was isolated from the seawater of Shantou, China. A novel β-agarase gene agaXa was cloned from the strain Catenovulum sp. X3. The gene agaXa consists of 1,590 bp and encodes a protein of 529 amino acids, with only 40 % amino acid sequence identity with known agarases. AgaXa should belong to the glycoside hydrolase family GH118 based on the amino acid sequence similarity. The molecular mass of the recombinant AgaXa (rAgaXa) was estimated to be 52 kDa by sodium dodecyl sulfate–polyacrylamide gel electrophoresis. It had a maximal agarase activity at 52 °C and pH 7.4 and was stable over pH 5.0 ~ 9.0 and at temperatures below 42 °C. The K m and V max for agarose were 10.5 mg/ml and 588.2 U/mg, respectively. The purified rAgaXa showed endolytic activity on agarose degradation, yielding neoagarohexaose, neoagarooctaose, neoagarodecaose, and neoagarododecaose as the end products. The results showed that AgaXa has potential applications in agar degradation for the production of oligosaccharides with various bioactivities.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
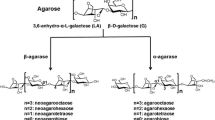
Agar, which is composed of agarose and agaropectin, is the main component of the cell walls of red alga. Agarose is composed alternatively of 1–4-linked 3,6-anhydro-α-l-galactose and 1–3-linked β-d-galactose (Duckworth and Yaphe 1971). Agarase is a kind of glycoside hydrolase that catalyzes the hydrolysis of agarose. According to the cleavage pattern, agarases are classified into α-agarase (EC 3.2.1.158) and β-agarase (EC 3.2.1.81). α-Agarases cleave to α-1,3 linkages of agarose to produce agarooligosaccharides, and β-agarases cleave to β-1,4 linkages to produce neoagarooligosaccharides. Most of the reported agarases are β-agarases, which are classified into four glycoside hydrolase (GH) families, GH16, GH50, GH86, and GH118, based on the amino acid sequence similarity on the Carbohydrate-Active EnZyme database (CAZy) (Cantarel et al. 2009; Chi et al. 2012).
Agarase has potential applications in the food, cosmetic, and medical industries for the production of neoagarooligosaccharides (NAOS) from agar. It has been proven that NAOS have various special biological activities, such as high antioxidative activity (Wu et al. 2005), moisturizing effects on the skin, and whitening effects on the melanoma cells (Kobayashi et al. 1997; Lee et al. 2008), as well as inhibiting the growth of bacteria and slowing down the degradation of starch (Giordano et al. 2006). Moreover, agarases have been used to recover DNA from the agarose gel (Finkelstein and Rownd 1978; Zhang and Sun 2007) and to prepare seaweed protoplasts (Chen et al. 1994; Dipakkore et al. 2005).
In this study, a new β-agarase gene agaXa was cloned from Catenovulum sp. X3. It is the first reported agarase gene in genus Catenovulum. According to the amino acid sequence similarity analysis, AgaXa should belong to the novel glycoside hydrolase family GH118. The gene agaXa was expressed in Escherichia coli BL21 (DE3), and the recombinant protein (rAgaXa) was purified and characterized. Based on its high activity and thermal and pH stability, agarase AgaXa has potential applications in the industry.
Materials and methods
Chemicals and reagents
All chemicals were reagent grade or higher and were purchased from commercial sources, unless otherwise stated. Enzymes were purchased from TaKaRa (Dalian, China). Oligonucleotides were synthesized by BGI (Beijing, China).
Bacterial strains, plasmids and culture conditions
E. coli DH5α and BL21 (DE3) were used as the host for cloning and expression, respectively. Plasmid pBluescript SK(+) and pET-32a(+) were used as vectors for cloning and expression, respectively. The E. coli strain was grown in Luria–Bertani (LB) medium at 37 °C, supplemented with ampicillin (100 μg/ml) when required. The agarase-producing bacterium was cultured using marine agar or marine broth 2216E (Oppenheimer and ZoBell 1952) supplemented with 0.1 % agar.
Isolation and identification of strain X3
Strain X3 was isolated from seawater samples collected from the coastal sea of Shantou, China. The colony that produces an obvious degradation zone on marine agar 2216E at 37 °C was selected for identification. The 16S rRNA gene was amplified using the universal primers 27 F and 1492R (Lane 1991). The 16S rRNA gene sequence was identified using the EzTaxon-e server (Kim et al. 2012). Strain X3 had been deposited at the NITE Biological Resource Center of Japan (NBRC) under the accession number NBRC 107874.
Cloning and analysis of the agarase gene
To construct a partial genomic library, the genomic DNA of strain X3 was isolated using the MiniBEST Bacterial Genomic DNA Extraction Kit version 2.0 (TaKaRa, Dalian). The genomic DNA was digested by restriction endonuclease EcoRΙ and HindIII and separated on 0.7 % agarose gels, and the DNA fragments with sizes from 2 to 7 kb were purified using the TaKaRa Agarose Gel DNA Purification Kit version 2.0 and ligated into pBluescript SK(+) containing the same endonuclease site. The ligated DNA was transformed into E. coli DH5α cells. The transformants were plated on LB agar plates with ampicillin (100 μg/ml) and 1 mM IPTG. The clone showing agarase activity was screened and sequenced. Open reading frame was analyzed using the DNASTAR software. The signal peptide was predicted using the SignalP 4.0 server (Petersen et al. 2011). The searches for amino sequence similarities and conserved domains were performed with the BLAST programs from the National Center for Biotechnology Information. Sequence alignment was conducted using the BioEdit software. A phylogenetic tree was generated using the neighbor-joining method as described by Lin et al. (2012a).
Expression and purification of the recombinant AgaXa
The agarase gene agaXa including its signal sequence was amplified from the chromosomal DNA of strain X3, using the forward primer, 5′-GCGCGGATCCATCAAACTATTGAAT-3′, and reverse primer, 5′-GGCCTCGAGAATTACAACCCAT-3′, containing BamHΙ and XhoΙ sites, respectively. The terminator of agaXa was removed, and a poly-6-histidine residue was added to the C terminus of the recombinant protein in order to purify the protein by affinity chromatography. The amplified product was purified and digested by BamHΙ and XhoΙ and ligated to the corresponding vector pET-32a(+). Finally, the ligated plasmid pET-32a(+)–agaXa was transformed into E. coli BL21 (DE3) competent cells. The recombinant strain E. coli BL21 (DE3)–pET-32a(+)–agaXa was cultured in a 500-ml LB medium supplemented with 100 μg/ml ampicillin at 37 °C. When the culture turbidity at 600 nm reached 0.6, IPTG and sodium deoxycholate were added to a final concentration of 0.2 mM and 0.01 % (w/v), respectively. The cells were grown for an additional 20 h at 25 °C. Cells and medium were separated by centrifugation at 10,000 × g for 10 min. Proteins in the supernatant were precipitated by adding ammonium sulfate to a final concentration of 75 % (w/v). After centrifugation, the protein pellet was dissolved in a 20-mM Tris–HCl (pH 7.4) buffer at 4 °C. Following dialysis, the protein was loaded onto a Ni–NTA agarose column (10 ml of set volumes) (QIAGEN China Co., Ltd, Shanghai). The column was washed with buffer A (20 mM Tris–HCl with 0.5 M NaCl, pH 7.4) and then with buffer B (buffer A with 20 mM imidazole). The recombinant protein was eluted with buffer C (buffer A with 100 mM imidazole) and analyzed by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). Protein concentrations were measured using the Enhanced BCA Protein Assay Kit (Beyotime Institute of Biotechnology, Jiangsu, China).
Enzyme assay
The agarase activity was measured by the release of the reducing sugar equivalent using 3,5-dinitrosalicylic acid method (Miller 1959). The reaction mixture (1 ml) consisted of 5 μl of agarase solution and 995 μl 50 mM Tris–HCl (pH 7.4) containing 0.25 % melted agarose. The mixture was incubated at 52 °C for 10 min, and the reaction was terminated by boiling for 10 min. Optical density was read at 520 nm, and values for reducing sugars were expressed as d-galactose equivalents. One unit was defined as the amounts of enzyme required to release 1 μmol of the reducing sugars per minute at the above conditions.
Effects of temperature and pH on enzyme activity and stability
The optimal temperature of enzyme activity was determined over a temperature range of 27–62 °C and at pH 7.4 for 10 min. The thermostability assay was performed by incubating the enzyme for 1 h at different temperatures and measuring the residual enzyme activity. The effect of pH on agarase activity was assayed at a pH range of 5.3–8.3 in 50 mM PBS buffer. The pH stability of the agarase was determined by pre-incubating the enzyme solution at each pH (4.0–11.0) at 4 °C for 12 h, and residual enzyme activity was measured under standard conditions.
Effects of various reagents on enzyme activity
The effects of various reagents on enzyme activity were determined by adding various reagents to the reaction mixture with final concentrations of 2 and 10 mM. The relative activity was defined as the percentage of the activity determined under the standard condition described previously without any reagent addition.
Determination of kinetic parameters
Kinetic parameters of AgaXa were determined in 50-mM Tris–HCl buffer (pH 7.4) with 1–5 mg/ml substrate (agarose, molecular mass of 10 kDa). The samples were incubated at 52 °C for 10 min. K m and V max for the substrate were obtained using the Lineweaver–Burke equation. Values of K cat (turnover number) and K cat / K m (catalytic efficiency) were calculated based on the K m, V max, and [E] (concentration of agarose) values.
Enzymatic product analysis
The purified agarase (5 U/ml) was incubated with the agarose (10 mg/ml) or a NAOS mixture at 42 °C for 24 h. The NAOS mixture (named 814) containing neoagarooctaose (DP8), neoagarodecaose (DP10), neoagarododecaose (DP12), and neoagarotetradecaose (DP14) was prepared by enzymatic hydrolysis of agarose using other β-agarases studied in our lab (Lin et al. 2012b). For the agarose degradation, the reaction mixture was centrifuged at 12,000 × g for 15 min; the supernatant was analyzed by thin layer chromatography (TLC) using silica gel 60 TLC plates (Merck, Darmstadt, Germany) and lyophilized. The plates were developed with a solvent system composed of n-butanol/acetic acid/H2O (2:1:1, by volume). The spots were visualized by spraying with 10 % (v/v) H2SO4 and heating (80 °C). The molecular mass of the enzymatic product was determined using a matrix-assisted laser desorption ionization time-of-flight (MALDI-TOF) mass spectrometer (QSTAR Pulsar I, Applied Biosystems). Carbon-13 nuclear magnetic resonance (13C NMR) spectrum of the lyophilized products (dissolved in D2O, Sigma, USA) was recorded on AVANCE 400 MHz (Brucker Biospin, Swiss). For the NAOS degradation, the reaction mixture was directly analyzed by TLC after centrifugation.
Nucleotide sequence accession number
The nucleotide sequence of 16S rRNA gene of strain X3 and agaXa reported here had been submitted to the GenBank database under the accession numbers GU975828 and GU975829, respectively.
Results
Identification of the agar-degrading bacterium
One strain (named X3) showing high agarolytic activity on the marine agar plate at 37 °C was picked out and purified. The nearly complete 16S rRNA gene sequence (1,494 bp) of strain X3 was determined and aligned with the 16S rRNA gene sequences of all known type strains in the EzTaxon-e server. The sequences having the highest similarities with the 16S rRNA gene sequence of strain X3 were as follows: Catenovulum agarivorans DSM 23111T (97.6 %), Bowmanella pacifica LMG 24568T (91.1 %), and Bowmanella denitrificans BD1T (90.6 %). The results showed that the 16S rRNA gene sequence of strain X3 had very low similarities to other known bacteria except to that of the type strain of the newly established genus Catenovulum (Yan et al. 2011). Therefore, strain X3 was identified as a member of Catenovulum.
Cloning and sequence analysis of the agarase gene
A clone, showing agarase activity, was screened and named pBSK-Xa. A 3,505-bp inserted DNA fragment was sequenced and analyzed; the result showed that it harbored an open reading frame of 1,590 bp which was named agaXa. The gene agaXa encoded a protein of 429 amino acids with an isoelectric point of pH 7.8 and a predicted molecular weight of 56.6 kDa. SignalP 4.0 server predicted that the encoded protein (AgaXa) contained a 28-amino acid signal peptide. According to the analysis in the BLAST search, AgaXa showed the highest amino acid sequence identity of only 40 % with two reported agarases with identical amino acid sequences: they are AgaB from Pseudoalteromonas sp. CY24 (Ma et al. 2007) and AgaC from Vibrio sp. PO-303 (Dong et al. 2006). Non-identified conserved domain was found in AgaXa. In order to verify the relationship between AgaXa and the β-agarases from known glycoside hydrolase families, a phylogenetic tree based on the amino acid sequences was constructed (Fig. 1). It showed that the AgaXa formed a clade with agarase AgaB which had been proposed for the representation of a novel glycoside hydrolase family GH118 (Ma et al. 2007; Chi et al. 2012). The amino acid sequence comparison between AgaXa and AgaB (Fig. 2) showed that both of them have an N-terminal signal peptide and some conservative amino acid residues.
Alignment of amino acid sequences of AgaXa and AgaB (BAF3590) from Pseudoalteromonas sp. CY24. The alignment was performed using the BioEdit software. The signal peptide was underlined. The conserved Glu and Asp residues were marked with triangles and the conserved Gly, Ala, and Arg were marked with asterisks
Expression and purification of AgaXa
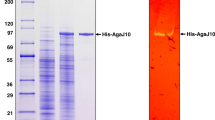
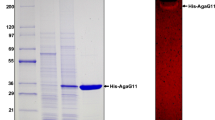
The agarase AgaXa with the signal peptide and a polyhistidine (6 × His) sequence on the C terminus was overexpressed in the E. coli BL21 (DE3) system. The culture supernatant showed high extracellular agarase activity. AgaXa purified from the culture supernatant formed a single band in the SDS-PAGE with an apparent molecular mass of 52 kDa (Fig. 3).
Effects of temperature and pH on enzyme activity and stability
The AgaXa showed high activity when the temperature was between 37 and 52 °C (the optimum temperature was 52 °C); it was stable at temperatures below 42 °C, and more than 95 % activity of the maximal activity was retained (Fig. 4a). The optimum pH of AgaXa was 7.4. It retained more than 85 % activity after being preincubated at a wide range of pH 5.0–9.0 at 4 °C for 12 h (Fig. 4b).
Effects of various reagents on enzyme activity
The activity of AgaXa was strongly inhibited by Cu2+, Fe3+, Mn2+, Al3+, and SDS (Table 1). Oppositely, AgaXa's activity was increased by 127, 54, and 44 % by adding 10 mM of DDT, β-met, and metal ions Mg2+, respectively.
Kinetic parameters
The kinetic parameters of AgaXa for agarose were calculated based on Lineweaver–Burke plots. The K m, V max, K cat, and K cat / K m of AgaXa were 10.5 mg/ml, 588.2 U/mg, 5.15 × 102 s−1, and 4.9 × 106 s−1 M−1, respectively.
Enzymatic product analysis
The agarose degradation products by AgaXa were analyzed by TLC (Fig. 5a). At the initial stage, the enzyme hydrolyzed agarose to generate oligosaccharides with various polymerization degrees, suggesting that AgaXa is an endo-type agarase. The enzymatic products at 2 h were identified by MALDI-TOF mass spectrometer (Fig. 5c). As shown in the mass spectrum, the main peaks were at 975, 1,281, 1,587, 1,893, and 2,199 m/z which were close to the calculated molecular mass of neoagarohexaose (DP6), DP8, DP10, DP12, and DP14, respectively. Therefore, after 24 h of agarose degradation, the end enzymatic products were DP6, DP8, DP10, and DP12 (Fig. 5a). Also, AgaXa could hydrolyze the NAOS mixture containing DP8, DP10, DP12, and DP14 into DP4, DP6, and DP8 (Fig. 5b). Therefore, AgaXa cannot hydrolyze DP8 or smaller NAOS. The end products of agarose degradation by AgaXa were further identified by 13C NMR. The resonances at about 97 and 93 ppm, which were assigned to α- and β-anomer carbon, respectively, at the reducing end of NAOS, were found in the 13C NMR spectrum (Fig. 6). No signal at 90.72 ppm was detected, which was typically found at the reducing end of agarooligosaccharides. These results confirmed that AgaXa is a β-agarase.
Analysis of enzymatic products. a TLC analysis of agarose degradation by rAgaXa. b The NAOS degradation by rAgaXa: lane 1, self-made DP4 and DP6; lane 2, self-made NAOS mixture 814; lane 3, the products of the NAOS degradation by rAgaXa. c MALDI-TOF mass spectrum of the 2-h enzymatic products of agarose by rAgaXa
Discussion
Most of the agar-degrading bacteria were isolated at 25 °C. In order to obtain some new agar-degrading bacteria with thermostable agarase, the culture temperature for strain isolation was increased to 37 °C in this study. Finally, the agarolytic strain X3 showing good growth at 37 °C was obtained, indicating the presence of thermostable agarase in the bacterium. New bacteria suggest the presence of novel agarase genes. Agarase AgaXa is the first reported agarase of Catenovulum.
CAZy has classified AgaXa, AgaB from Pseudoalteromonas sp. CY24 (Ma et al. 2007), and AgaC from Vibrio sp. PO-303 (Dong et al. 2006) into a novel glycoside hydrolase family GH118. This classification is supported by the phylogenetic tree constructed in this study based on the amino acid sequences of AgaXa and the β-agarases from known glycoside hydrolase families (Fig. 1) in which AgaXa and agarase AgaB formed an independent clade. According to the reported properties of AgaB and AgaC, there is one common characteristic of this new glycoside hydrolase family GH118—the members cannot hydrolyze DP8 or smaller NAOS. A structural analysis revealed that AgaB has a large substrate-binding cleft that accommodates 12 sugar units, with eight sugar units toward the reducing end, spanning subsites +1 to +8; with four sugar units toward the non-reducing end, spanning subsites −4 to −1; and with enzymatic cleavage taking place between subsites −1 and +1. This structural evidence helps explain the inability of AgaB to hydrolyze neoagarooctaose or smaller neoagarooligosaccharides and its ability to produce larger agarooligosaccharides, predominantly including neoagarooctaose, than other β-agarases (Ma et al. 2007; Ren et al. 2010; Chi et al. 2012). AgaXa may have a similar substrate-binding cleft as AgaB.
Polysaccharide-degrading enzymes generally contain a catalytic module and one or more carbohydrate-binding modules (CBM). The amino acids Gly, Ala, and/or Arg are the key residues in the CBM of glycoside hydrolase (Ma et al. 2007), while the key residues in the catalytic module of glycoside hydrolase are Glu and/or Asp (Ohta et al. 2004a). Therefore, some of these amino acid residues may play an important role in the agarose degradation by AgaXa or AgaB (Fig. 2). The crystallization of AgaB had been reported (Ren et al. 2010), but the detailed structural properties especially the substrate-binding cleft and its key residues are still unknown. The crystallization of AgaXa is now undertaken for the determination of its 3D structure. It would be helpful to understand the structural properties of members in the family GH118.
AgaXa also has different properties from AgaB and AgaC as follows. AgaXa has the maximum activity at 52 °C while AgaB and AgaC at 40 and 35 °C, respectively. Therefore, AgaXa is more thermostable than AgaB and AgaC. Also, the optimum pH of AgaXa is 7.4, while those of AgaB and AgaC are 6.0 (Table 2). Besides, based on the sequence analysis results of InterProScan, AgaB has a transmembrane motif located at the region of amino acid 21–41, while there is no such motif in the N terminus of AgaXa. Also, AgaB owns more predicted parallel β-helixes (nine helixes) than AgaXa (five helixes).
Industrial application requires agarase with high activity and stability at temperatures higher than the gelling temperature of agar (about 40 °C) (Ohta et al. 2004a). Nowadays, there are only a few β-agarases that can meet that condition. For example, Ohta et al. (2004b) reported a β-agarase, AgaA, from Microbulbifer thermotolerans JAMB-A94; the maximal activity was observed at 55 °C. The enzyme was stable up to 60 °C. The thermophilic bacteria, Thermoanaerobacter wiegelii and Caldoanaerobacter sp. (Bannikova et al. 2008), were isolated from hot springs and can produce agarase with optimum temperature over 55 °C, but the agarase genes have not been reported. In this study, the stable and optimum temperatures of AgaXa are higher than the gelling temperature of agar. Therefore, agarase AgaXa has potential applications in the industry.
References
Bannikova GE, Lopatin SA, Varlamov VP, Kuznetsov BB, Kozina IV, Miroshnichenko ML, Chernykh NA, Turova TP, Bonch-Osmolovskaya EA (2008) The thermophilic bacteria hydrolyzing agar: characterization of thermostable agarase. Appl Biochem Microbiol 45:366–371
Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, Henrissat B (2009) The Carbohydrate-Active EnZyme database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res 37:D233–238
Chen LCM, Craigiel JS, Xie ZK (1994) Protoplast production from Porphyra linearis using a simplified agarase procedure capable of commercial application. J Appl Phycol 6:35–39
Chi WJ, Chang YK, Hong SK (2012) Agar degradation by microorganisms and agar-degrading enzymes. Appl Microbiol Biotechnol 94:917–930
Dipakkore S, Reddy CRK, Jha B (2005) Production and seeding of protoplasts of Porphyra okhaensis (Bangiales, Rhodophyta) in laboratory culture. J Appl Phycol 17:331–337
Dong J, Hashikawa S, Konishi T, Tamaru Y, Araki T (2006) Cloning of the novel gene-encoding beta-agarase C from a marine bacterium, Vibrio sp. strain PO-303, and characterization of the gene product. Appl Environ Microbiol 72:6399–6401
Duckworth M, Yaphe W (1971) Structure of agar: part I. Fractionation of a complex mixture of polysaccharides. Carbohydr Res 16:189–197
Giordano A, Andreotti G, Tramice A, Trincone A (2006) Marine glycosyl hydrolases in the hydrolysis and synthesis of oligosaccharides. J Biotechnol 1:511–530
Finkelstein M, Rownd RH (1978) A rapid method for extracting DNA from agarose gels. Plasmid 1:557–562
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kobayashi R, Takimasa M, Suzuki T, Kirimura K, Usami S (1997) Neoagarobiose as a novel moisturizer with whitening effect. Biosci Biotechnol Biochem 61:162–163
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. John Wiley & Sons, Ltd., Chichester, pp 115–175
Lee DG, Jang MK, Lee OH, Kim NY, Ju SA, Lee SH (2008) Overproduction of a glycoside hydrolase family 50 beta-agarase from Agarivorans sp. JA-1 in Bacillus subtilis and the whitening effect of its product. Biotechnol Lett 30:911–918
Lin B, Lu G, Zheng Y, Xie W, Li S, Hu Z (2012a) Aquimarina agarilytica sp. nov., a novel agarolytic species isolated from red alga. Int J Syst Evol Microbiol 62:869–873
Lin B, Lu G, Zheng Y, Xie W, Li S, Hu Z (2012b) Gene cloning, expression and characterization of a neoagarotetraose-producing β-agarase from a marine bacterium Agarivorans sp. HZ105. World J Microb Biot 28:1691–1697
Ma CP, Lu XZ, Shi C, Li J, Gu Y, Ma Y, Chu Y, Han F, Gong Q, Yu W (2007) Molecular cloning and characterization of a novel β-agarase, AgaB, from marine Pseudoalteromonas sp. CY24. J Biol Chem 282:3747–3754
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31:426–428
Ohta Y, Hatada Y, Nogi Y, Miyazaki M, Li Z, Akita M, Hidaka Y, Goda S, Ito S, Horikoshi K (2004a) Enzymatic properties and nucleotide and amino acid sequences of a thermostable β-agarase from a novel species of deep-sea microbulbifer. Appl Microbiol Biotechnol 64:505–514
Ohta Y, Nogi Y, Miyazaki M, Li Z, Hatada Y, Ito S, Horikoshi K (2004b) Enzymatic properties and nucleotide and amino acid sequences of a thermostable β-agarase from the novel marine isolate JAMB-A94. Biosci Biotechnol Biochem 68:1073–1081
Oppenheimer CH, ZoBell CE (1952) The growth and viability of sixty-three species of marine bacteria as influenced by hydrostatic pressure. J Mar Res 11:10–18
Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786
Ren A, Xia ZX, Yu W, Zhou J (2010) Expression, crystallization and preliminary X-ray analysis of an anomeric inverting agarase from Pseudoalteromonas sp. CY24. Acta Cryst F66:1635–1639
Wu SC, Wen TN, Pan CL (2005) Algal–oligosaccharide–lysates prepared by two bacterial agarases stepwise hydrolyzed and their antioxidative properties. Fish Sci 71:1149–1159
Yan S, Yu M, Wang Y, Shen C, Zhang XH (2011) Catenovulum agarivorans gen. nov., sp. nov., a peritrichously flagellated, chain-forming, agar-hydrolysing gammaproteobacterium from seawater. Int J Syst Evol Microbiol 61:2866–2873
Zhang WW, Sun L (2007) Cloning, characterization, and molecular application of a beta-agarase gene from Vibrio sp. strain V134. Appl Environ Microbiol 73:2825–2831
Acknowledgements
This work was supported by the National Natural Science Foundation of China (nos. 41076106 and 31200077), Guangdong Natural Science Foundation (no. S2011030005257), and the Key Science and Technology Innovation Project for University by the Department of Education of Guangdong Province (no. CXZD1124) and the Science & Technology Project of Guangdong Province (no. 2012A031100009).
Author information
Authors and Affiliations
Corresponding author
Additional information
Wei Xie, Bokun Lin, and Zhengrong Zhou contributed equally to this work.
Rights and permissions
About this article
Cite this article
Xie, W., Lin, B., Zhou, Z. et al. Characterization of a novel β-agarase from an agar-degrading bacterium Catenovulum sp. X3. Appl Microbiol Biotechnol 97, 4907–4915 (2013). https://doi.org/10.1007/s00253-012-4385-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-012-4385-5