Abstract
3-Phenyllactic acid (PLA), which is an organic acid widely existing in honey and lactic acid bacteria fermented food, can be produced by many microorganisms, especially lactic acid bacteria. It was proved as an ideal antimicrobial compound with broad and effective antimicrobial activity against both bacteria and fungi. In addition, it could be used as feed additives to replace antibiotics in livestock feeds. This article presented a review of recent studies on the existing resource, antimicrobial activity, and measurement of PLA. In addition, microorganism strains and dehydrogenases producing PLA were reviewed in detail, the metabolic pathway and regulation of PLA synthesis in LAB strains were discussed, and high-level bioproduction of PLA by microorganism fermentation was also summarized.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Food safety is generally recognized as a primary public safety issue in the world. Food microbial contamination is an important factor to result in food safety problems, which possibly bring about public health problems and great economic losses. Bacterial contamination, especially by pathogenic species including Listeria, Staphylococcus, Escherichia coli, and Salmonella, can cause foodborne illnesses (Kaneko et al. 1999). Fungal contamination by yeasts and moulds, which easily grow and reproduce in food system with feeble oxygen, can cause food spoilage (Schnurer et al. 1999) and even can produce toxic secondary metabolites, namely, mycotoxins, which are capable of causing disease and death in humans and animals (Riley et al. 1993). Application of antimicrobial preservatives, which inhibit the growth of bacteria or fungi, is an effective approach to bring down the food safety hazards by microbial contaminations.
3-Phenyllactic acid (2-hydroxy-3-phenylpropanoic acid or β-phenyllactic acid, PLA), a kind of an organic acid, has been reported as an antimicrobial compound with broad-spectrum activity against bacteria including Listeria monocytogenes (Dieuleveux et al. 1998b), Staphylococcus aureus, and Escherichia coli O157:H7 (Ohhira et al. 2004), and fungi including yeasts (Schwenninger et al. 2008) and a wide range of moulds, such as Aspergillus ochraceus, Penicillium roqueforti, and Penicillium citrinu (Lavermicocca et al. 2003). The present article is a review of recent studies on the properties, measurement, antimicrobial activity, and biosynthesis pathway, as well as its biological production by microbial fermentation and the possible enzymes producing PLA.
PLA
Existing sources
PLA was found widely existing in honey, and its content was commonly much higher than other phenolic acids in honey. PLA was suggested as chemical marker for thistle (Galactites tomentosa Moench) unifloral honeys, in which PLA content reached 100–800 mg/kg (Tuberoso et al. 2011). It was also found in high concentration in heather, ling heather, and manuka honeys (820, 875, and 243 mg/kg) (Tan et al. 1988; Dimitrova et al. 2007). Most of other honeys had PLA concentration level of less than 100 mg/kg (Wilkins et al. 1993 and 1995; Dimitrova et al. 2007), while it could not be detected in sunflower honey (Dimitrova et al. 2007). Recently, PLA was reported as metabolite of food microorganism, especially lactic acid bacteria (LAB). So, PLA was found existing in fermentation foods using LAB as starter, such as sourdoughs (Van der Meulen et al. 2007; Ryan et al. 2009).
Chemical structure
The molecular formula and molecular weight of PLA are C9H10O3 and 166 g/mol, respectively. PLA has an asymmetric carbon atom and thus has two chiral isomers: D- and L-PLA (Fig. 1).
Measurement
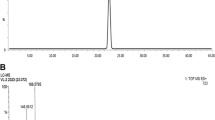
Reverse-phase HPLC is a simple and widely applicable method for measuring PLA. It was adopted in most of quantitative PLA measurement of microbial fermentation broth (Armaforte et al. 2006). In addition, it was developed to quantify the PLA content in rumen fluid (Khan et al. 1998) and honey (Tuberoso et al. 2011) (shown in Table 1). Regular gas chromatography–mass spectrometry (GC/MS) and capillary electrophoresis could also be used to measure PLA (Wilkins et al. 1995; Sarkissian et al. 2000). However, the aforementioned methods are usually non-stereospecific and cannot separate and detect the PLA enantiomers, L- and D-PLA.
Tekewe et al. (2008) reported the stereospecific determination of PLA using chiral HPLC method, in which Chiralcel OJ-H column based on cellulose tris-(4-methyl benzoate) chiral stationary phase was used to separate PLA enantiomers. In addition, using modified cyclodextrins as chiral additive during chromatography or capillary electrophoresis process could achieve the separation of PLA enantiomers (Nardi and Eliseev 1993; Heil et al. 1998).
Antibacterial activity
Dieuleveux et al. (1998b) firstly reported the antibacterial activity of PLA. They purified and identified the novel anti-Listeria compound, PLA, produced and excreted from Geotrichum candidum, and found that D-PLA showed slightly higher anti-Listeria activity than L-PLA. Then, PLA was proved to be able to inhibit Listeria monocytogenes growth in culture medium, milk, and cheese. It could reduce the bacteria population by 4.5 log in ultra-high-temperature treatment whole milk, to give fewer cells than in the control after 5 days of culture (Dieuleveux and Gueguen 1998).
In addition, PLA could inhibit a range of Gram-positive bacteria, such as Staphylococcus aureus, Enterococcuss faecalis, and Bacillus cereus, and Gram-negative bacteria, such as Salmonella enterica, Escherichia coli, Providencia stuartii, and Klebsiella oxytoca (Dieuleveux et al. 1998a; Ohhira et al. 2004). PLA could show higher inhibitory effect in acidic pH (Ohhira et al. 2004). The mechanism of antibacterial action is not clear yet, but it was suggested that the bacterial cell wall should be an action site of PLA. Scanning electron microscope studies showed that the bacteria exposed to PLA had damaged, even broken cell wall structure. The bacteria formed aggregates and secreted polysaccharides; then, the cell wall lost rigidity, causing the cells to swell, even collapse (Dieuleveux et al. 1998a).
Antifungal activity
In addition to antibacterial activity, the inhibitory properties of PLA have also been demonstrated against yeasts, such as Candida pulcherrima, Candida parapsilosis, and Rhodotorula mucilaginosa (Schwenninger et al. 2008) and a wide range of mould species isolated from bakery products, flour, and cereals, including some mycotoxigenic species, namely, Aspergillus ochraceus, Penicillium roqueforti, Penicillium citrinu, etc. (Lavermicocca et al. 2000 and 2003). PLA has relatively high MIC value for antiyeast activity (50 to more than 500 mM at pH 4.0 to 6.0), and the value of PLA decreased with decreasing pH (Schwenninger et al. 2008). And MIC value against moulds at pH 4.0 is 45 mM (Strom et al. 2002).
The fungal inhibitory activity of PLA was firstly characterized by Lavermicocca et al. (2000), who purified the antifungal compounds from Lactobacillus plantarum Strain 21B, a lactic acid bacterium (LAB) with high antifungal activity. Using the PLA-producing strain, L. plantarum 21B, as a starter during sourdough bread fermentation process, the fungal growth could be delayed for 7 days (Lavermicocca et al. 2000). PLA has been considered as antifungal compound marker (Schnürer and Magnusson 2005) and been widely purified and characterized from various LABs, such as L. plantarum 21B (Lavermicocca et al. 2000), L. plantarum MiLAB 393 (Strom et al. 2002), L. plantarum IMAU10014 (Wang et al. 2012), L. plantarum (Prema et al. 2010), and Weissella cibaria FMF4B16 (Ndagano et al. 2011). And it was presumed that the behavior of the antifungal activity of LAB strains was positively related to the metabolic content of PLA (Valerio et al. 2004).
Other applications
Like other organic acids, PLA can be used as feed additives to replace antibiotics in livestock feeds. It may exert some positive effects to the immune system of laying hens and then effectively improve production performance and egg quality (Wang et al. 2009b). When supplemented in long-term diet of chick feeds, PLA improves the growth performance, has antipathogen effect in large intestine, and reduces yellowness of meat (Wang et al. 2010). Also, it was reported that PLA may increase immune-related blood cell counts and potentially reduce E. coli numbers in weanling and growing pigs (Wang et al. 2009a).
In addition, PLA has potential as a pharmaceutical agent to treat coronary disease since its analogue “Danshensu” from Chinese medicine is applied presently (Wang et al. 1991). And it also has been patented to be used as a skin-protecting ingredient to reduce skin wrinkles (Yu and Van Scott 1997).
Biological PLA production
Microorganisms producing PLA
Lavermicocca et al. (2000) reported the production of PLA from L. plantarum 21B, which was the first report showing the production of PLA by lactic acid bacteria (LAB) (2000). Subsequently, it was found that PLA could be generally produced by a wide range of LAB species, such as Lactobacillus, Enterococcus, Weissella, and Leuconostoc, but the production varied greatly among strains and species. When grown in DeMan–Rogosa–Sharpe (MRS) medium, most of LAB strains produced less than 1-mM PLA; however, L. plantarum 1081, L. plantarum 778, L. plantarum 1073, L. acidophilus 1063 (Gerez et al. 2010), and L. plantarum CECT-221 (Rodriguez et al. 2012) could produce 5.2-, 4.1-, 2.6-, 1.1-, and 1.38-mM PLA, respectively (shown in Table 2).
PLA could also be produced by a series of dairy propionic acid bacteria (PAB) strains, such as Propionibacterium jensenii DSMZ 20535, P. thoenii DSMZ 20276, P. acidipropionici DSMZ 4900, and P. freudenreichii ssp. freudenreichii DSMZ 2027, with the production of 0.01–0.1 mM (Lind et al. 2007). And cofermentation with P. jensenii SM11 and Lactobacillus paracasei subsp. paracasei SM20 produced 1-mM PLA (Schwenninger et al. 2008).
In addition, PLA could be produced by other microorganism strains, such as bacillaceae (Bacillus coagulans) (Zheng et al. 2011), fungus (Geotrichum candidum) (Dieuleveux et al. 1998b), and Brevibacteriaceae (Brevibacterium lactofermentum) (Kamata et al. 1986). And both G. candidum and B. lactofermentum could produce much higher PLA than LAB and PAB strains (shown in Table 2).
Metabolic pathway and regulation of PLA synthesis in LAB strains
The synthesis of PLA in LAB strains results from the catabolism of phenylalanine, in which phenylalanine is transaminated to phenylpyruvic acid (PPA) and PPA further reduced to PLA (shown in Fig. 2) (McSweeney and Sousa 2000; Yvon and Rijnen 2001; Li et al. 2007; Vermeulen et al. 2006). The amino acid phenylalanine had remarkable effect on PLA production in LAB strains. When cultured in synthetic medium without phenylalanine, L. plantarum ITM21B did not produce PLA at all; however, the PLA fermentation amounts could be observed and gradually increased if phenylalanine was supplemented with concentrations ranging from 0.1 to 0.4 g/L in initial medium (Valerio et al. 2004). The direct correlation between PLA production and phenylalanine content in medium was also reported in many other references (Li et al. 2007; Dallagnol et al. 2011; Rodriguez et al. 2012).
The transamination reaction is the first catabolic step of phenylalanine and is initiated by an aromatic aminotransferase (AAT) which has broad substrate spectrum, including leucine, tyrosine, tryptophan, and methionine (Yvon et al. 1997). AAT catalyzes transferring the amino group from the amino acid to a suitable α-keto acid acceptor, which commonly is α-ketoglutarate in most LAB strains (Fig. 2) (Yvon et al. 1998; Rijnen et al. 2000). Therefore, α-ketoglutarate has important effect phenylalanine catabolism and impacts on the regulation of PLA biosynthesis (Vermeulen et al. 2006; Dallagnol et al. 2011). Vermeulen et al. reported that the addition of α-ketoglutarate strongly increased PLA formation in L. plantarum TMW1.468 by 5 to >30 % (2006). On the other hand, α-ketoglutarate is produced from glutamate by glutamate dehydrogenase, and the glutamate dehydrogenase activity is influenced by the redox state of the cell, so PLA formation can be up-regulated indirectly by adding those compounds, which act as alternative electron acceptors to increase NAD(P)+ level, such as citrate, fructose, and glucose (Vermeulen et al. 2006; Li et al. 2007; Dallagnol et al. 2011).
Actually, phenylpyruvic acid (PPA) is the direct precursor of PLA in PLA biosynthesis of LAB strains (Fig. 2) and shows much higher effect on PLA production than phenylalanine. When PPA was used to replace phenylalanine as supplemented substrate at the same concentration, PLA production increased 14-fold during Lactobacillus sp. SK007 fermentation (Li et al. 2007). It was suggested that phenylalanine transamination was the limiting factor in PLA production in LAB strains, and the bottleneck could be overcome using PPA as substrate (Li et al. 2007; Mu et al. 2009a and 2009b; Zheng et al. 2011).
Dehydrogenases converting PPA to PLA
There have been several different kinds of dehydrogenases that were characterized to convert PPA to PLA, and lactate dehydrogenase (LDH) is the main kind. In nature, there are two forms of LDH with different catalytic stereospecificity, L-LDH (EC 1.1.1.27) and D-LDH (EC 1.1.1.28). They have the highest catalytic activity for pyruvic acid and wide substrate specificity for α-ketonic acids, such as 2-ketobutyrate, α-ketoglutaric acid, etc. (el Hawrani et al. 1996). So far, it was found that LDHs from many LAB strains have the substrate specificity for PPA, including L-LDH from Pediococcus acidilactici DG302 (Garmyn et al. 1995), L. plantarum SK002 (Jia et al. 2010), and Lactobacillus helveticus 53/7 (Savijoki and Palva 1997), and D-LDH from Pediococcus pentosaceus ATCC 25745 (Yu et al. 2012), P. acidilactici DSM 20284 (Mu et al. 2012), L. plantarum SK002 (Jia et al. 2010), Lactobacillus pentosus JCM1558 (Tokuda et al. 2003) (previously called L. plantarum ATCC 8041 (Taguchi and Ohta 1991)), and Lactobacillus confusus 20196 (Hummel et al. 1983). Also, it was reported that some LDHs from non-LAB organisms could also transform PPA into PLA, such as Thermoanaerobacter ethanolicus JW200 (Zhou and Shao 2010), B. coagulans SDM (Zheng et al. 2011), and Clonorchis sinensis (Yang et al. 2006).
The optimum pH and temperature for LDH from LAB strains are in the range of 5.5–7.0 and 30–45 °C, respectively, probably because most of LAB strains are mesophile and acidophile (Table 3). The LDH from LAB strains generally showed weak thermostability, especially under more than 45 °C (Li et al. 2008; Jia et al. 2010; Yu et al. 2012; Mu et al. 2012). Some LDHs were characterized from thermophilic non-LAB strains, which showed higher optimum and thermostability, such as T. ethanolicus JW200 (Zhou and Shao 2010) and B. coagulans SDM (Zheng et al. 2011). In all of the LDHs reported, P. acidilactici D-LDH showed the highest catalytic efficiency (k cat/K m) value of 105 mM−1 s−1 in D-LDHs, and the k cat/K m of L-LDH was only reported in B. coagulans L-LDH with 110 mM−1 s−1.
Although natural LDH showed less substrate specificity to PPA than pyruvic acid, the substrate specificity could be improved by modification of enzyme. Tokuda et al. (2003) constructed a mutant L. pentosus D-LDH in which the single amino acid of Tyr52 was replaced by Leu, and this mutant D-LDH (Y52L) showed much higher substrate specificity and catalytic efficiency to PPA than the wild-type D-LDH.
In addition to LDH, some other dehydrogenases were also reported exhibiting catalytic hydrogenation activity to phenylpyruvate, in which D-form dehydrogenases included D-hydroxyisocaproate dehydrogenase (D-HicDH) from Lactobacillus casei (Hummel et al. 1985) and the D-mandelate dehydrogenase (D-ManDH) from Enterococcus faecalis (Tamura et al. 2002; Wada et al. 2008) and Lactobacillus curvatus (Hummel et al. 1988). These D-form dehydrogenases have different optimum substrate; however, interestingly, all of them show broad substrate specificity to 2-ketoacids including phenylpyruvate (Table 4). Especially, L. casei D-HicDH (Hummel et al. 1985) and L. curvatus D-ManDH (Hummel et al. 1988) show relatively high specificity to phenylpyruvate with K m of 0.15 mM. In L-form dehydrogenases, L. confuses L-HicDH was characterized to have substrate specificity to PPA with k cat/K m of 2.81 × 108 s−1 M−1 (Feil et al. 1997).
High-level bioproduction of PLA by microorganism fermentation
PLA production could be remarkably improved when adding PPA in initial medium or during fermentation process. Commercially, PPA can be obtained easily at relatively low price through organic synthesis from hydantoin (Christidis and Schouteeten 1985) and has been used to produce phenylalanine which is in high demand for the production of an artificial sweetener “aspartame” (Matsunaga et al. 1987; Leng et al. 2006). Therefore, it is an effective approach to produce PLA in large scale using PPA as material. It was reported that PLA content increased 14-fold in Lactobacillus sp. SK007 fermentation, which reached 1.12 g L−1, when PPA was added in MRS broth (Li et al. 2007). Using response surface methodology, the medium components containing PPA were optimized for PLA production of Lactobacillus sp. SK007, and the PLA fermentation yield increased to 2.30 g L−1 (Mu et al. 2009a).
Further, the fed-batch fermentation of Lactobacillus sp. SK007 with substrate PPA feeding and pH-control was reported to be able to produce 17.38-g L−1 PLA, with the conversion ratio of PPA to PLA of 51.1 % and PLA production rate of 0.241 g L−1 h−1 (Mu et al. 2009b). Recently, Zheng et al. isolated a thermophilic bacteria, B. coagulans SDM, having PLA producing ability at a high temperature, which was helpful to improve the solubility and dissolution rate of substrate PPA. Using whole cells of B. coagulans SDM converting PPA, PLA was produced in a high concentration of 37.3 g L−1 and high productivity of 2.3 g L−1 h−1 (Zheng et al. 2011).
Future
So far, the scientific researches for the antimicrobial activity of PLA focused on the PLA producing strains in the form of protective cultures. There were not enough application researches of PLA in food system as a pure antimicrobial agent. More researches are needed to verify the effectiveness of PLA in food system and to compare it with other typical antimicrobial agents in detail.
Although PLA exists in many ordinary foods such as honey and LAB-fermented foods, PLA is still not allowed by legislation to be used as an additive. It is necessary to implement more human trials to study the metabolism pathway, health effects, and possible toxicity effects of PLA. And these data would give a guide to whether it can be approved as a legal additive.
To our best knowledge, there is no reference reporting the downstream process of PLA preparation. Therefore, in addition to optimizing the high-level production of PLA, the downstream process researches should be strengthened in the future.
References
Arai K, Kamata T, Uchikoba H, Fushinobu S, Matsuzawa H, Taguchi H (2001) Some Lactobacillus L-lactate dehydrogenases exhibit comparable catalytic activities for pyruvate and oxaloacetate. J Bacteriol 183:397–400
Armaforte E, Carri S, Ferri G, Caboni MF (2006) High-performance liquid chromatography determination of phenyllactic acid in MRS broth. J Chromatogr A 1131:281–284
Christidis Y, Schouteeten A (1985) Process for preparation of crystallized monohydrated sodium phenylpyruvate. US Patent 4,518,800
Dallagnol AM, Catalan CAN, Mercado MI, de Valdez GF, Rollan GC (2011) Effect of biosynthetic intermediates and citrate on the phenyllactic and hydroxyphenyllactic acids production by Lactobacillus plantarum CRL 778. J Appl Microbiol 111:1447–1455
Dieuleveux V, Gueguen M (1998) Antimicrobial effects of D-3-phenyllactic acid on Listeria monocytogenes in TSB-YE medium, milk, and cheese. J Food Prot 61:1281–1285
Dieuleveux V, Lemarinier S, Gueguen M (1998a) Antimicrobial spectrum and target site of D-3-phenyllactic acid. Int J Food Microbiol 40:177–183
Dieuleveux V, Van Der Pyl D, Chataud J, Gueguen M (1998b) Purification and characterization of anti-Listeria compounds produced by Geotrichum candidum. Appl Environ Microbiol 64:800–803
Dimitrova B, Gevrenova R, Anklam E (2007) Analysis of phenolic acids in honeys of different floral origin by solid-phase extraction and high-performance liquid chromatography. Phytochem Anal 18:24–32
el Hawrani AS, Sessions RB, Moreton KM, Holbrook JJ (1996) Guided evolution of enzymes with new substrate specificities. J Mol Biol 264:97–110
Feil IK, Hendle J, Schomburg D (1997) Modified substrate specificity of L-hydroxyisocaproate dehydrogenase derived from structure-based protein engineering. Protein Eng 10:255–262
Garmyn D, Ferain T, Bernard N, Hols P, Delcour J (1995) Cloning, nucleotide sequence, and transcriptional analysis of the Pediococcus acidilactici L-(+)-lactate dehydrogenase gene. Appl Environ Microbiol 61:266–272
Gerez CL, Carbajo MS, Rollan G, Torres Leal G, Font de Valdez G (2010) Inhibition of citrus fungal pathogens by using lactic acid bacteria. J Food Sci 75:354–359
Heil M, Podebrad F, Beck T, Mosandl A, Sewell AC, Bohles H (1998) Enantioselective multidimensional gas chromatography–mass spectrometry in the analysis of urinary organic acids. J Chromatogr B Biomed Sci Appl 714:119–126
Hummel W, Schutte H, Kula MR (1983) Large-scale production of D-lactate dehydrogenase for the stereospecific reduction of pyruvate and phenylpyruvate. Eur J Appl Microbiol Biotechnol 18:75–85
Hummel W, Schutte H, Kula MR (1985) D-2-hydroxyisocaproate dehydrogenase from Lactobacillus casei—a new enzyme suitable for stereospecific reduction of 2-ketocarboxylic acids. Appl Microbiol Biotechnol 21:7–15
Hummel W, Schutte H, Kula MR (1988) D-(−)-mandelic acid dehydrogenase from Labtobacillus curvatus. Appl Microbiol Biotechnol 28:433–439
Jia J, Mu W, Zhang T, Jiang B (2010) Bioconversion of phenylpyruvate to phenyllactate: gene cloning, expression, and enzymatic characterization of D- and L1-lactate dehydrogenases from Lactobacillus plantarum SK002. Appl Biochem Biotechnol 162:242–251
Kamata M, Toyomasu R, Suzuki D, Tanaka T (1986) D-phenyllactic acid production by Brevibacterium or Corynebacterium. Patent JP 86108396
Kaneko KI, Hayashidani H, Ohtomo Y, Kosuge J, Kato M, Takahashi K, Shiraki Y, Ogawa M (1999) Bacterial contamination of ready-to-eat foods and fresh products in retail shops and food factories. J Food Prot 62:644–649
Khan RI, Amin MR, Mohammed N, Onodera R (1998) Quantitative determination of aromatic amino acids and related compounds in rumen fluid by high-performance liquid chromatography. J Chromatogr B Biomed Sci Appl 710:17–25
Lavermicocca P, Valerio F, Evidente A, Lazzaroni S, Corsetti A, Gobbetti M (2000) Purification and characterization of novel antifungal compounds from the sourdough Lactobacillus plantarum strain 21B. Appl Environ Microbiol 66:4084–4090
Lavermicocca P, Valerio F, Visconti A (2003) Antifungal activity of phenyllactic acid against molds isolated from bakery products. Appl Environ Microbiol 69:634–640
Leng L, Zheng P, Sun Z (2006) Continuous production of l-phenylalanine from phenylpyruvic acid and l-aspartic acid by immobilized recombinant Escherichia coli SW0209-52. Process Biochem 41:1669–1672
Li XF, Jiang B, Pan BL (2007) Biotransformation of phenylpyruvic acid to phenyllactic acid by growing and resting cells of a Lactobacillus sp. Biotechnol Lett 29:593–597
Li XF, Jiang B, Pan BL, Mu WM, Zhang T (2008) Purification and partial characterization of Lactobacillus species SK007 lactate dehydrogenase (LDH) catalyzing phenylpyruvic acid (PPA) conversion into phenyllactic acid (PLA). J Agric Food Chem 56:2392–2399
Lind H, Sjogren J, Gohil S, Kenne L, Schnurer J, Broberg A (2007) Antifungal compounds from cultures of dairy propionibacteria type strains. FEMS Microbiol Lett 271:310–315
Matsunaga T, Higashijima M, Sulaswatty A, Nishimura S, Kitamura T, Tsuji M, Kawaguchi T (1987) Repeated batch production of L-phenylalanine from phenylpyruvate and NH4Cl by immobilized cells of Nocardia opaca under hydrogen high pressure. Biotechnol Bioeng 31:834–840
McSweeney PLH, Sousa MJ (2000) Biochemical pathways for the production of flavour compounds in cheeses during ripening: a review. Lait 80:293–324
Mu WM, Chen C, Li XF, Zhang T, Jiang B (2009a) Optimization of culture medium for the production of phenyllactic acid by Lactobacillus sp SK007. Bioresour Technol 100:1366–1370
Mu WM, Liu FL, Jia JH, Chen C, Zhang T, Jiang B (2009b) 3-Phenyllactic acid production by substrate feeding and pH-control in fed-batch fermentation of Lactobacillus sp SK007. Bioresour Technol 100:5226–5229
Mu W, Yu S, Jiang B, Li X (2012) Characterization of D-lactate dehydrogenase from Pediococcus acidilactici that converts phenylpyruvic acid into phenyllactic acid. Biotechnol Lett. doi:10.1007/s10529-012-0847-1
Nardi A, Eliseev A (1993) Use of charged and neutral cyclodextrins in capillary zone electrophoresis: enantiomeric resolution of some 2-hydroxy acids. J Chromatogr 638:247–253
Ndagano D, Lamoureux T, Dortu C, Vandermoten S, Thonart P (2011) Antifungal activity of 2 lactic acid bacteria of the Weissella genus isolated from food. J Food Sci 76:305–311
Ohhira I, Kuwaki S, Morita H, Suzuki T, Tomita S, Hisamatsu S, Sonoki S, Shinoda S (2004) Identification of 3-phenyllactic acid as a possible antibacterial substance produced by Enterococcus faecalis TH10. Biocontrol Sci 9:77–81
Prema P, Smila D, Palavesam A, Immanuel G (2010) Production and characterization of an antifungal compound (3-phenyllactic acid) produced by Lactobacillus plantarum strain. Food Bioprocess Tech 3:379–386
Rijnen L, Courtin P, Gripon JC, Yvon M (2000) Expression of a heterologous glutamate dehydrogenase gene in Lactococcus lactis highly improves the conversion of amino acids to aroma compounds. Appl Environ Microbiol 66:1354–1359
Riley RT, Norred WP, Bacon CW (1993) Fungal toxins in foods: recent concerns. Annu Rev Nutr 13:167–189
Rodriguez N, Salgado JM, Cortes S, Dominguez JM (2012) Antimicrobial activity of D-3-phenyllactic acid produced by fed-batch process against Salmonella enterica. Food Control 25:274–284
Ryan LA, Dal Bello F, Czerny M, Koehler P, Arendt EK (2009) Quantification of phenyllactic acid in wheat sourdough using high resolution gas chromatography–mass spectrometry. J Agric Food Chem 57:1060–1064
Sarkissian CN, Scriver CR, Mamer OA (2000) Measurement of phenyllactate, phenylacetate, and phenylpyruvate by negative ion chemical ionization–gas chromatography/mass spectrometry in brain of mouse genetic models of phenylketonuria and non-phenylketonuria hyperphenylalaninemia. Anal Biochem 280:242–249
Savijoki K, Palva A (1997) Molecular genetic characterization of the L-lactate dehydrogenase gene (ldhL) of Lactobacillus helveticus and biochemical characterization of the enzyme. Appl Environ Microbiol 63:2850–2856
Schnürer J, Magnusson J (2005) Antifungal lactic acid bacteria as biopreservatives. Trends Food Sci Tech 16:70–78
Schnurer J, Olsson J, Borjesson T (1999) Fungal volatiles as indicators of food and feeds spoilage. Fungal Gen Biol 27:209–217
Schwenninger SM, Lacroix C, Truttmann S, Jans C, Sporndli C, Bigler L, Meile L (2008) Characterization of low-molecular-weight antiyeast metabolites produced by a food-protective Lactobacillus-Propionibacterium coculture. J Food Prot 71:2481–2487
Strom K, Sjogren J, Broberg A, Schnurer J (2002) Lactobacillus plantarum MiLAB 393 produces the antifungal cyclic dipeptides cyclo(L-Phe-L-Pro) and cyclo(L-Phe-trans-4-OH-L-Pro) and 3-phenyllactic acid. Appl Environ Microbiol 68:4322–4327
Taguchi H, Ohta T (1991) D-lactate dehydrogenase is a member of the D-isomer-specific 2-hydroxyacid dehydrogenase family. Cloning, sequencing, and expression in Escherichia coli of the D-lactate dehydrogenase gene of Lactobacillus plantarum. J Biol Chem 266:12588–12594
Tamura Y, Ohkubo A, Iwai S, Wada Y, Shinoda T, Arai K, Mineki S, Iida M, Taguchi H (2002) Two forms of NAD-dependent D-mandelate dehydrogenase in Enterococcus faecalis IAM 10071. Appl Environ Microbiol 68:947–951
Tan S, Wilkins A, Molan P, Holland P, Reid M (1988) A chemical approach to the determination of floral sources of New Zealand honeys. J Apicult Res 28:212–222
Tekewe A, Singh S, Singh M, Mohan U, Banerjee UC (2008) Development and validation of HPLC method for the resolution of drug intermediates: DL-3-phenyllactic acid, DL-O-acetyl-3-phenyllactic acid and (+/−)-mexiletine acetamide enantiomers. Talanta 75:239–245
Tokuda C, Ishikura Y, Shigematsu M, Mutoh H, Tsuzuki S, Nakahira Y, Tamura Y, Shinoda T, Arai K, Takahashi O, Taguchi H (2003) Conversion of Lactobacillus pentosus D-lactate dehydrogenase to a D-hydroxyisocaproate dehydrogenase through a single amino acid replacement. J Bacteriol 185:5023–5026
Tuberoso CI, Bifulco E, Caboni P, Sarais G, Cottiglia F, Floris I (2011) Lumichrome and phenyllactic acid as chemical markers of thistle (Galactites tomentosa Moench) honey. J Agri Food Chem 59:364–369
Valerio F, Lavermicocca P, Pascale M, Visconti A (2004) Production of phenyllactic acid by lactic acid bacteria: an approach to the selection of strains contributing to food quality and preservation. FEMS Microbiol Lett 233:289–295
Van der Meulen R, Scheirlinck I, Van Schoor A, Huys G, Vancanneyt M, Vandamme P, De Vuyst L (2007) Population dynamics and metabolite target analysis of lactic acid bacteria during laboratory fermentations of wheat and spelt sourdoughs. Appl Environ Microbiol 73:4741–4750
Vermeulen N, Ganzle MG, Vogel RF (2006) Influence of peptide supply and cosubstrates on phenylalanine metabolism of Lactobacillus sanfranciscensis DSM20451(T) and Lactobacillus plantarum TMW1.468. J Agric Food Chem 54:3832–3839
Wada Y, Iwai S, Tamura Y, Ando T, Shinoda T, Arai K, Taguchi H (2008) A new family of D-2-hydroxyacid dehydrogenases that comprises D-mandelate dehydrogenases and 2-ketopantoate reductases. Biosci Biotechnol Biochem 72:1087–1094
Wang J, Shao Y, Zhang Y, Jiao B, Dai H, Xue F (1991) Experimental studies of β-phenyllactic acid on the coronary system. J Shanghai Med Univ 18:295–297
Wang JP, Yoo JS, Lee JH, Jang HD, Kim HJ, Shin SO, Seong SI, Kim IH (2009a) Effects of phenyllactic acid on growth performance, nutrient digestibility, microbial shedding, and blood profile in pigs. J Anim Sci 87:3235–3243
Wang JP, Yoo JS, Lee JH, Zhou TX, Jang HD, Kim HJ, Kim IH (2009b) Effects of phenyllactic acid on production performance, egg quality parameters, and blood characteristics in laying hens. J Appl Poultry Res 18:203–209
Wang JP, Lee JH, Yoo JS, Cho JH, Kim HJ, Kim IH (2010) Effects of phenyllactic acid on growth performance, intestinal microbiota, relative organ weight, blood characteristics, and meat quality of broiler chicks. Poultry Sci 89:1549–1555
Wang H, Yan Y, Wang J, Zhang H, Qi W (2012) Production and characterization of antifungal compounds produced by Lactobacillus plantarum IMAU10014. PloS one 7:e29452
Wilkins A, Lu Y, Tan S (1993) Extractives from New Zealand honeys. 4. Linalool derivatives and other components from nodding thistle (Carduus nutans) honey. J Agric Food Chem 41:873–878
Wilkins A, Lu Y, Tan S (1995) Extractives from New Zealand honeys. 5. Aliphatic dicarboxylic acids in New Zealand rewarewa (Knightea excelsa) honey. J Agric Food Chem 43:3021–3025
Yang G, Jing C, Zhu P, Hu X, Xu J, Wu Z, Yu X (2006) Molecular cloning and characterization of a novel lactate dehydrogenase gene from Clonorchis sinensis. Parasitol Res 99:55–64
Yu R, Van Scott E (1997) Method of using 3-phenyllactic acid for treating wrinkles. Patent US 5643953
Yu S, Jiang H, Jiang B, Mu W (2012) Characterization of D-lactate dehydrogenase producing D-3-phenyllactic acid from Pediococcus pentosaceus. Biosci Biotechnol Biochem 76:853–855
Yvon M, Rijnen L (2001) Cheese flavour formation by amino acid catabolism. Int Dairy J 11:185–201
Yvon M, Thirouin S, Rijnen L, Fromentier D, Gripon JC (1997) An aminotransferase from Lactococcus lactis initiates conversion of amino acids to cheese flavor compounds. Appl Environ Microbiol 63:414–419
Yvon M, Berthelot S, Gripon JC (1998) Adding alpha-ketoglutarate to semi-hard cheese curd highly enhances the conversion of amino acids to aroma compounds. Int Dairy J 8:889–898
Zhang DL, Li WL, Zhang JB, Tang WR, Chen XF, Cao KW, Chu QC, Ye JN (2010) Determination of unconjugated aromatic acids in urine by capillary electrophoresis with dual electrochemical detection—potential application in fast diagnosis of phenylketonuria. Electrophoresis 31:2989–2996
Zheng ZJ, Ma CQ, Gao C, Li FS, Qin JY, Zhang HW, Wang K, Xu P (2011) Efficient conversion of phenylpyruvic acid to phenyllactic acid by using whole cells of Bacillus coagulans SDM. PloS one 6:e19030
Zhou Q, Shao WL (2010) Molecular genetic characterization of the thermostable L-lactate dehydrogenase gene (ldhL) of Thermoanaerobacter ethanolicus JW200 and biochemical characterization of the enzyme. Biochemistry (Mosc) 75:526–530
Acknowledgements
This work was supported by the 973 Project (No. 2012CB720802), the 863 Project (No. 2011AA100904), the Natural Science Foundation of China Project (No. 31171705), and the Support Project of Jiangsu Province (No. BE2011622, BE2011766, BE2010678, and BE2010626).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mu, W., Yu, S., Zhu, L. et al. Recent research on 3-phenyllactic acid, a broad-spectrum antimicrobial compound. Appl Microbiol Biotechnol 95, 1155–1163 (2012). https://doi.org/10.1007/s00253-012-4269-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-012-4269-8