Abstract
A novel bacterium, Cr-10, was isolated from a chromium-contaminated site and capable of removing toxic chromium species from solution by reducing hexavalent chromium to an insoluble precipitate. Sequence analysis of 16S rRNA gene of strain Cr-10 showed that it was most closely related to Serratia rubidaea JCM 1240T (97.68%). Physiological and chemotaxonomic data also supported that strain Cr-10 was identified as Serratia sp., a genus which was never specially reported chromate-resistant before. Serratia sp., Cr-10 was tolerant to a concentration of 1,500 mg Cr(VI) L−1, which was the highest level reported until now. The optimum pH and temperature for reduction of Cr(VI) by Serratia sp. Cr-10 were found to be 7.0 and 37 °C, respectively. The Cr(VI) reduction was significantly influenced by additional carbon sources, and among them fructose and lactose offered maximum reduction, with a rate of 0.28 and 0.25 mg Cr(VI) L−1 h−1, respectively. The cell-free extracts and filtrate of the culture were able to reduce Cr(VI) while concentration of total chromium remained stable in the process, indicating that the enzyme-catalyzed mechanism was applied in Cr(VI) reduction by the isolate. Additionally, it was found that there was hardly any chromium on the cell surface of the strain, further supporting that reduction, rather than bioadsorption, plays a major role in the Cr(VI) removal.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Chromium is essential for life in trace amounts, but is highly toxic to most organisms at low concentrations. Extensive use of chromium in industries such as leather tanning, metallurgical, electroplating, etc. resulting in industrial wastes containing hexavalent chromium [Cr(VI)], has led to the contamination of soil, sediment, surface, and ground waters (Szulczewski et al. 2001; Malik 2004). Compared to Cr(VI), trivalent chromium [Cr(III)] is nontoxic, and due to its lower environmental mobility, thus exhibiting limited environmental disruption.
Certain species have mechanisms of chromate tolerance and resistance, such as Enterobacter cloacae, Desulfovibrio vulgaris, Pseudomonas aeruginosa, Cupriavidus metallidurans, Ralstonia metallidurans, P. putida, Escherichia coli, Caulobacter crescentus, Shewanella oneidensis, Bacillus firmus, and Burkholderia cepacia. The mechanisms by which these microorganisms reduce Cr(VI) are variable and are species dependent. Several species contain membrane transporters responsible for mediating efflux of chromate ions from cell cytoplasm, such as P. aeruginosa (Alvarez et al. 1999) and She. oneidensis (Brown et al. 2006). Some species utilize a soluble enzyme that catalyses the reduction of Cr(VI)) into the much less toxic and less mobile Cr(III), leading to decreased uptake of chromium such as E. coli (Ackerley et al. 2004), Cau. crescentus (Hu et al. 2005), Cup. metallidurans (Juhnke et al. 2002), B. firmus (Sau et al. 2010), and Acidiphilium cryptum (Magnuson et al. 2010) even could reduce Cr(VI) by periplasmic c-type cytochrome in the way of obtaining electrons from the respiratory chain and divert them to Cr(VI). In addition, E. coli (Llagostera et al. 1986), P. aeruginosa (Miranda et al. 2005), and She. oneidensis (Chourey et al. 2006) also use DNA repair system to resist toxicity caused by chromium compounds.
Currently, chemical remediation process to immobilize toxic heavy metals such as chromium can be expensive and complicated. The capacity of microbial biomass removing chromium from effluent represents a potential and more economic strategy. In this study, we described the capacity for chromate reduction by a newly isolated bacterium from a chromate-contaminated site, designated as Cr-10, which may be applied in the simple, efficient, and economic bioremediation strategy. We characterized the physiological and biochemical properties, chromate-reducing capacity of this isolate, and the effect factors such as temperature, initial pH, amount of inocula, initial concentration of Cr(VI), and carbon sources were also investigated.
Materials and methods
Isolation of Cr-resistant bacteria
Cr-resistant bacteria were isolated from soil at four sites (see supplementary Table S1) near a chromate-producing chemical factory in Qingdao, China. The heavy metal content of the soil sample was evaluated by modified colorimetric (542 nm) diphenylcarbazide method (Pattanapipitpaisal et al. 2001). The soil sample was diluted with sterile water and the dilutions were plated onto nutrient agar (beef extract 0.3%; peptone 0.5%; NaCl 0.5%; agar 1.5%; w/v) amended with 40 mg Cr(VI) L−1. A filter-sterilized solution of K2Cr2O7 (Guangdong Guanghua Chemical Factory Co. Ltd, AR Grade) was used as the source of Cr(VI), which was added to the sterile molten nutrient agar to avoid reduction during autoclaving. The plates were incubated at 30 °C for a week. Totally, 16 distinct colonies were selected as Cr-resistant isolates and one isolate Cr-10 (CCTCC AB 209242, China Center for Type Culture Collection) which was identified as Serratia sp. was used in the subsequent study.
Identification of strains
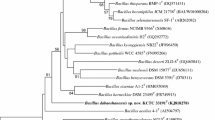
The 16S rRNA gene was amplified by PCR using bacterial universal primers 27F and 1540R (Rainey et al. 1996) and the PCR products were sequenced by Invitrogen Biotechnology Co. Ltd. (Shanghai, China). Sequence similarity was searched by the National Center for Biotechnology Information blast and calculated by pairwise alignment obtained from the EzTaxon database (Chun et al. 2007). Analysis of the 16S rRNA gene sequence data was performed by using the software package MEGA version 4.0 (Tamura et al. 2007) after multiple alignment of the data by CLUSTAL_X (Thompson et al. 1997). The phylogenetic tree was inferred using the neighbor-joining (Saitou and Nei 1987) methods. Bootstrap analysis based on 1,000 replications was undertaken to test the robustness of the phylogenetic tree (Felsenstein 1985).
Cell morphology was examined by phase-contrast (Olympus BX51, Tokyo, Japan) and transmission electron microscopy (Hitachi H-7650, Tokyo, Japan) using cells negatively stained with 1% (w/v) phosphotungstic acid after air drying. Motility was examined according to the method of Bernardet et al. (2002). Growth under anaerobic conditions was tested on nutrient agar in a GasPak (BBL) jar at 30 °C for 24 h. Catalase activity was determined by assessing bubble production in 3% (v/v) H2O2 and oxidase activity was determined using 1% (w/v) tetramethyl-p-phenylenediamine. Carbon-source utilization and enzyme activities were tested with Ser. ficaria DSM 4569T under the same condition, using the API 20NE, API 20E, and API ZYM test kits (bioMérieux, Marcy, France) at 30 °C for 24 h to 48 h.
To determine its whole-cell fatty acid profile, strain Cr-10 and Ser. ficaria DSM 4569T were cultured at 30 °C for 4 days on TSB agar. Analysis of the fatty acid methyl esters was carried out according to the standard protocol of the Sherlock Microbial Identification System (MIDI, Newark, USA).
Determination of inhibitory concentration of Cr(VI)
The minimal inhibitory concentration (MIC) of Cr(VI) at which no colony growth occurred was determined by broth agar dilution method (Luli et al. 1983). One percent (v/v) of the log-phase culture was inoculated into nutrient broth containing different concentrations of Cr(VI) (40–1500 mg L−1) at pH 7.0 and incubated at 30 °C at 180 rpm for 24 h. Afterwards, the liquid culture was diluted with sterile water and plated onto nutrient agar. These plates were incubated at 30 °C for 48 h, the MIC was considered to be the lowest concentration of Cr(VI) at which no growth occurred (McLean et al. 2000).
Batch reduction of Cr(VI) with free cells
Complete reduction of 20–23 mg L−1 chromate at different temperatures and pH were observed in cultures of strain Cr-10 aerobically grown in nutrient broth with 1% (w/v) glucose as the additional carbon source. The batch reduction test was also performed with initial Cr(VI) concentrations ranging from 10 to 50 mg L−1 in the same medium, as well as various additional 1% (w/v) carbon sources including glucose, fructose, sucrose, lactose, sodium acetate, and succinic acid in nutrient broth containing 20 mg L−1 chromate. The samples were withdrawn at 6-h intervals and subjected to Cr(VI) estimation according to the method mentioned above. The medium containing appropriate concentration of Cr(VI) without Serratia sp. was used as control.
To estimate the influence of temperature on Cr(VI) reduction, experiments were carried out with free cells incubated in nutrient broth containing 23 mg L−1 Cr(VI) at 25, 30, 37, and 40 °C. Samples were withdrawn at 6-h intervals and analyzed for Cr(VI) reduction. The effect of initial pH on Cr(VI) reduction was studied by varying pH values as 5.0, 6.0, 7.0, 8.0, and 9.0 and the pH was adjusted using 1 N NaOH and/or 1 N HCl. A series of amount of inocula as 0%, 0.5%, 1%, 2%, and 5% and growth at initial concentration of Cr(VI) varied as 0, 10, 20, 40, and 50 mg L−1 were performed to investigate their effect on Cr(VI) reduction. The influence of various carbon sources was studied by adding glucose, fructose, sucrose, lactose, sodium acetate, and succinic acid.
Bioremediation mechanisms experiments
The free cells, cell-free extracts, and filtrate of culture were used to reduce Cr(VI) in order to investigate the mechanisms of Cr(VI) reduction by the isolate Cr-10. The nutrient broth without additional carbon source was inoculated with overnight grown bacterial culture and incubated for 24 h and then was centrifuged at 6,000 rpm for 15 min. The supernatant was filter-sterilized at 0.45 μm and supplemented with K2Cr2O7 to a final concentration of 20 mg L−1 Cr(VI), while the cells were resuspended with phosphate buffer solution (PBS, pH 7.0) and were disrupted by sonication on ice with output setting of 30% for 10 min with 30 s bursts and 3 min breaks. Following centrifugation, the supernatant, so is called cell-free extracts, was supplemented with 20 mg L−1 Cr(VI) and tested for Cr(VI) reduction together with the filtrate of culture. The PBS and nutrient broth containing the same concentration of Cr(VI) were used as control. In addition, the cells in PBS suspension was carried out Cr(total) test with the PBS as control, both of which were added to 20 mg L−1 Cr(VI). Concentration of the total Cr [Cr(VI) plus Cr(III)] was obtained by oxidation of Cr(III) to Cr(VI) using a 5-ml system, 250-μl sample, 250 μl H2SO4 (2 M), 1 ml (NH4)2S2O8 (10 g/l), and 3 ml H2O were mixed to boil for 20 min; after cooling down, 500 μl DPC (0.80 g/l) was added to incubate for 5 min and then resorted to Cr(VI) measurement method. To evaluate the heavy metal adsorption ability of Serratia sp., cells incubated in nutrient broth containing 1,500 mg L−1 Cr(VI) for 24 h were prepared for visual observation and elemental analysis by scanning electron microscopy (Hitachi S-4800, Toyko, Japan) and energy dispersive X-ray spectroscopy (EDS, Horiba EMAX, Kyoto, Japan).
The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequence of strain Cr-10 is HM475151.
Results
Isolation and identification of chromium-resistant microorganisms
Totally, 16 strains were isolated and identified as eight different species (see supplementary Table S2). Strain Cr-10 was studied further because of its low 16S rRNA gene sequence similarity to described species and high Cr(VI) concentration resistibility. The 16S rRNA gene sequence of strain Cr-10 was a continuous stretch of 1,423 bp. Sequence similarity calculations indicated strain Cr-10 showed the highest degree of identity to Ser. rubidaea DSM 4480T (97.68%), Ser. entomophila DSM 12358T (97.26%), and Ser. ficaria DSM 4569T (97.12%). The phylogenetic analysis based on 16S rRNA gene sequences demonstrated that the isolate formed a distinct branch within the genus Serratia (Fig. 1). The strain was Gram-negative, facultatively aerobic, oxidase-negative, and catalase-positive, gave a nitrate reduction test, an indole production test, and a Voges–Proskauer test positive. Colonies on nutrient agar were smooth, convex, circular, and red-colored. Other phenotypic characteristics that differentiated it from Ser. ficaria DSM 4569T were shown in Table 1. The isolate showed a minimum inhibitory concentration of 1,500 mg L−1 Cr(VI), with the cell deformation seriously due to the high concentration of toxic Cr(VI) but still keep viable, which indicated that strain Cr-10 was a novel chromium-resistant species (Fig. 2).
Factors affecting reduction of Cr(VI)
Temperature
After incubation in nutrient broth containing 23 mg L−1 Cr(VI) over temperatures between 25 and 40 °C, reduction was observed in all cultures (Fig. 3a). Complete reduction was achieved at 37 °C after 36 h, which appears to be the optimal temperature for chromate reduction under these conditions.
Factors affecting reduction of Cr(VI). Error bars indicate standard deviations from mean activities of triplicate assays. a Percentage of initial Cr(VI) (23 or 22 mg L−1) reduced at various temperatures and pH values. b Effect of initial concentration on % Cr(VI) removal and cell growth. Square, 10 mg L−1; diamond, 20 mg L−1; triangle, 40 mg L−1; inverted triangle, 50 mg L−1; hollow, concentration of Cr(VI) (mg L−1); solid, cell growth (A600). c Effect of carbon sources on Cr(VI) reduction
pH
The effect of initial solution pH on abiotic and biotic chromate reduction was investigated in flasks at an initial Cr (VI) concentration of 22 mg L−1 (Fig. 3b). The results clearly showed that pH value has more correlation with reduction than cell growth, as the optimum pH value for growth is 5.0 (Table 1) while it has the lowest Cr(VI) removal rate at pH 5.0.
Initial concentration of Cr(VI)
The effect of initial Cr(VI) concentration on the overall rate of reduction and cell growth was investigated at concentrations from 10 to 50 mg L−1. Complete reduction occurred at the initial concentrations of 10 and 20 mg L−1 after 12 h, while the Cr(VI) removal percent only reached 26% and 23% after 60 h at the initial concentrations of 40 and 50 mg L−1, respectively. At the same time, the influence of initial concentration on cell growth was studied and higher amount of biomass was obtained at an initial concentration of 10 mg L−1, while the cell growth in the initial concentration of 40 and 50 mg L−1 were strongly inhibited (Fig. 3c).
Carbon sources
Microbial reduction of toxic Cr(VI) to nontoxic Cr(III) is an electron-requiring process and effect of carbon sources, identically called electron donors such as glucose, fructose, sucrose, lactose, sodium acetate, and succinic acid on chromium reduction was studied. It was shown that fructose had the maximum reduction rate, lactose secondly, and sucrose thirdly, with a rate of 0.28, 0.25, and 0.23 mg Cr L−1 h−1, respectively, higher than the rate of glucose (0.16 mg Cr L−1 h−1) (Fig. 3d).
Bioremediation mechanism
To determine whether bioreduction or bioadsorption plays a major role in the Cr(VI) removal, the cell-free extracts, filtrate of culture, and whole cells of strain Cr-10 suspended in PBS were used to measure the Cr(VI) reduction capacity and the results were shown in Fig. 4. The cell-free extracts derived from cells incubated in nutrient broth and the filtrate of culture showed a reduction rate of 0.054 and 0.078 mg Cr L−1 h−1, respectively (Fig. 4a), and the overall reduction rate was 0.132 mg Cr L−1 h−1 and similar to the rate of culture without additional carbon source (0.125 mg Cr L−1 h−1, Fig. 3d). Moreover, the rate of filtrate was a little higher than that of cell-free extracts, which meant that the Cr(VI) reductase existed both extracellular and intracellular and more reductase were secreted to extracellular.
The whole cell reduction test results (Fig. 4b) showed that there was no significant decline but a little increase for the concentration of Cr(total) in the incubation process, which indicated that bioadsorption plays hardly any role in the Cr(VI) removal action. The EDS spectrum confirmed the conclusion as the Cr precipitates bound to the cell surface accounted for 0.33% only (see supplementary Table S3).
Discussion
Cr(VI) is mobile in the environment because of its high solubility in water. Thus, polluted soils with low concentrations of chromium are widespread in the environment. Bacteria with the ability to resist and reduce Cr(VI) can be used for detoxification of soils contaminated with Cr(VI). In this study, we isolated a strain Cr-10 from a high concentration of Cr(VI)-contaminated soil site (approximately 0.4 g Kg−1), identified as Serratia sp. The species in the genus had been investigated for their ability to produce cohesive biofilm supporting nanocrystalline palladium catalyst, which was then used to reduce Cr(VI) to less dangerous Cr(III) (Beadregard et al. 2010); additionally, several Ser. mercascens strains were isolated from tannery effluent containing Cr(VI) (Srivastava and Thakur 2007), no Cr-resistant Serratia sp. reported expressly.
Serratia sp. Cr-10 showed Cr(VI) reduction activity with the highest reduction rate of 0.28 mg Cr L−1 h−1 (0.028 mg Cr h−1), lower than Brucella sp. (0.92 mg Cr L−1 h−1) (Thacker et al. 2007) and Pseudomonas sp. (0.24 mg Cr h−1) (McLean et al. 2000). However, strain Cr-10 has a MIC of 1,500 mg Cr(VI) L−1, much higher than that of Brucella sp. (1,000 mg L−1) (Thacker et al. 2007) and Pseudomonas sp. (520 mg L−1) (McLean et al. 2000), indicating it has a strong defense system for Cr(VI) tolerance. In addition, it appears that among the carbon sources detected under our experiment conditions, fermentable carbon sources were fairly suited to enrich for Cr(VI) reducers, as the rate of without additional carbon source was 0.125 mg Cr L−1 h−1, less than half of which with fructose as additional carbon source. It is well known, Cr(VI) was transformed to Cr(III) via a reduction reaction, during which electrons transmitted from electron donor to Cr(VI). The carbon source just acted as electron donor in the process, thus it had a higher reduction rate with the additional carbon source. Therefore, it may be possible to stimulate Cr(VI)-reducing microorganisms at the natural site using relatively economical sources of fermentable, such as molasses which was the by-product in sucrose production.
There are two main mechanisms by which chromate could be reduced (Ramirez-Diaz et al. 2008). First, Cr(VI) could be reduced to the less toxic Cr(III) by an enzymatic reaction in strain Cr-10. Results described here showed an obvious correlation among Cr(VI) reduction and temperature and pH. The optimum temperature for the reduction activity of strain Cr-10 was 37 °C and the reaction was inhibited at pH 5.0 and 9.0. Since Cr(VI) reduction is enzyme-mediated, temperature and pH changes may affect the enzyme ionization rate and the protein folding, consequently affect enzyme activity. The initial Cr(VI) concentration effect test results showed a higher rate of growth and Cr(VI) reduction during the exponential growth phase because a great deal of active Cr(VI) reductase was produced during that phase. Bioadsorption is the second mechanism by which the Cr(VI) concentration could be reduced. In the bioremediation mechanism experiments, it showed that there was rarely chromium precipitates on the cell surface and that concentration of Cr(total) in the PBS kept almost invariable, indicating the weak adsorption strength of the cells which may be due to lack of viscous secretions such as exopolysaccharides. The bioinformation analysis revealed that there was chromate reductase in Ser. odorifera DSM 4582T (http://www.ncbi.nlm.nih.gov/genome) which was 37% identical to ChrR and 82% identical to YieF, respectively, two of the best studied chromate reductase (Ramirez-Diaz et al. 2008). Therefore, it is most likely that there is chromate reductase in strain Cr-10 which is homologous to ChrR and YieF to some extent and further studies are needed.
Besides, there were also several Bacillus sp., Alcaligenes sp. Cr-14, and Morganella sp. Cr-16 isolated from the chromate-contaminated site, which Cr-resistant ability was variable, consequently maybe forming a bacterial consortia to remove chromate more efficiently in situ. In conclusion, the study demonstrated the potential to stimulate naturally surviving microorganisms community at the polluting site to reduce Cr(VI) to Cr(III).
References
Ackerley DF, Gonzalez CF, Park CH, Blake R, Keyhan M, Matin A (2004) Chromate-reducing properties of soluble flavoproteins from Pseudomonas putida and Escherichia coli. Appl Environ Microbiol 70:873–882
Alvarez AH, Moreno-Sanchez R, Cervantes C (1999) Chromate efflux by means of the ChrA chromate resistance protein from Pseudomonas aeruginosa. J Bacteriol 181:7398–7400
Beadregard DA, Yong P, Macaskie LF, Johns ML (2010) Using non-invasive magnetic resonance imaging (MRI) to assess the reduction of Cr(VI) using a biofilm-palladium catalyst. Biotechnol Bioeng 107:11–20
Bernardet JF, Nakagawa Y, Holmes B (2002) Proposed minimal standards for describing new taxa of the family Flavobacteriaceae and emended description of the family. Int J Syst Evol Microbiol 52:1049–1070
Brown SD, Thormpson MR, Verberkmoes NC, Chourey K, Shah M, Zhou J, Hettich RL, Thompson DK (2006) Molecular dynamics of the Shewanella oneidensis response to chromate stress. Mol Cell Proteomics 5:1054–1071
Chourey K, Thormpson MR, Morrell-Falvey J, Verberkmoes NC, Brown SD, Shah M, Zhou J, Doktycz M, Hettich RL, Thompson DK (2006) Global molecular and morphological effects of 24-hour chromium(VI) exposure on Shewanella oneidensis MR-1. Appl Environ Microbiol 72:6331–6344
Chun J, Lee JH, Jung Y, Kim M, Kim S, Kim BK, Lim YW (2007) EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequences. Int J Syst Evol Microbiol 57:2259–2261
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Grimont PAD, Grimont F, Starr MP (1979) Serratia ficaria sp. nov., a bacterial species associated with Smyrna figs and the fig wasp Blastophaga psenes. Curr Microbiol 2:277–282
Hu P, Brodie EL, Suzuki Y, McAdams HH, Andersen GL (2005) Whole-genome transcriptional analysis of heavy metal stresses in Caulobacter crescentus. J Bacteriol 187:8437–8449
Juhnke S, Peitzsch N, Hubener N, Grosse C, Nies DH (2002) New genes involved in chromate resistance in Ralstonia metallidurans strain CH34. Arch Microbiol 179:15–25
Llagostera M, Garrido S, Guerrero R, Barbe J (1986) Induction of SOS genes of Escherichia coli by chromium compounds. Environ Mutagen 8:571–577
Luli GW, Talnagi JW, Strohl WR, Pfister RM (1983) Hexavalent chromium-resistant bacteria isolated from river sediments. Appl Environ Microbiol 46:846–854
Magnuson TS, Swenson MW, Paszczynski AJ, Deobald LA, Kerk D, Cummings DE (2010) Proteogenomic and functional analysis of chromate reduction in Acidiphilium cryptum JF-5, an Fe(III)-respiring acidophile. Biometal 23:1129–1138
Malik A (2004) Metal bioremediation through growing cells. Environ Int 30:261–278
McLean JS, Beveridge TJ, Phipps D (2000) Isolation and characterization of a chromium-reducing bacterium from a chromated copper arsenate-contaminated site. Environ Microbiol 2:611–619
Miranda AT, Gonzalez MV, Gonzalez G, Vargas E, Campos-Garcia J, Cervantes C (2005) Involvement of DNA helicases in chromate resistance by Pseudomonas aeruginosa PAO1. Mutat Res 578:202–209
Pattanapipitpaisal P, Brown NL, Macaskie LE (2001) Chromate reduction and 16S rRNA identification of bacteria isolated from a Cr(VI)-contaminated site. Appl Microbiol Biotechnol 57:257–261
Rainey FA, Ward-Rainey N, Kroppenstedt RM, Stackebrandt E (1996) The genus Nocardiopsis represents a phylogenetically coherent taxon and a distinct actinomycete lineage: proposal of Nocardiopsaceae fam. nov. Int J Syst Bacteriol 46:1088–1092
Ramirez-Diaz MI, Diaz-Perez C, Vargas E, Riveros-Rosas H, Campos-Garcia J, Cervantes C (2008) Mechanisms of bacterial resistance to chromium compounds. Biometals 21:321–332
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sau GB, Chatterjee S, Mukherjee SK (2010) Chromate reduction by cell-free extract of Bacillus firmus KUCr1. Pol J Microbiol 59:185–190
Srivastava S, Thakur IS (2007) Evaluation of biosorption potency of Acinetobacter sp. for removal of hexavalent chromium from tannery effluent. Biodegradation 18:637–646
Szulczewski MD, Helmke PA, Bleam WF (2001) XANES spectroscopy studies of Cr(VI) reduction by thiols in organosulfur compounds and humic substances. Environ Sci Technol 35:1134–1141
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thacker U, Parikh R, Shouche Y, Madamwar D (2007) Reduction of chromate by cell-free extract of Brucella sp. isolated from Cr(VI) contaminated sites. Bioresour Technol 98:1541–1547
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Acknowledgments
We thank Dr. Shu Xia for providing chromate-contaminated soil samples.
This work was supported by the Research Network for Applied Microbiology grant from Chinese Academy of Sciences.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
Soil samples used in this study were collected from chromate-contaminated sites (DOC 31 kb)
Table S2
Identification of the isolates based on 16S rRNA gene sequences and their Cr(VI)-resistant ability (DOC 35 kb)
Table S3
The EDS result of the cell surface of strain Cr-10 (DOC 32 kb)
Rights and permissions
About this article
Cite this article
Zhang, K., Li, F. Isolation and characterization of a chromium-resistant bacterium Serratia sp. Cr-10 from a chromate-contaminated site. Appl Microbiol Biotechnol 90, 1163–1169 (2011). https://doi.org/10.1007/s00253-011-3120-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-011-3120-y