Abstract
Xestospongia muta is among the most emblematic sponge species inhabiting coral reefs of the Caribbean Sea. Besides being the largest sponge species growing in the Caribbean, it is also known to produce secondary metabolites. This study aimed to assess the effect of depth and season on the symbiotic bacterial dynamics and major metabolite profiles of specimens of X. muta thriving in a tropical marine biome (Portobelo Bay, Panamá), which allow us to determine whether variability patterns are similar to those reported for subtropical latitudes. The bacterial assemblages were characterized using Illumina deep-sequencing and metabolomic profiles using UHPLC-DAD-ELSD from five depths (ranging 9–28 m) across two seasons (spring and autumn). Diverse symbiotic communities, representing 24 phyla with a predominance of Proteobacteria and Chloroflexi, were found. Although several thousands of OTUs were determined, most of them belong to the rare biosphere and only 23 to a core community. There was a significant difference between the structure of the microbial communities in respect to season (autumn to spring), with a further significant difference between depths only in autumn. This was partially mirrored in the metabolome profile, where the overall metabolite composition did not differ between seasons, but a significant depth gradient was observed in autumn. At the phyla level, Cyanobacteria, Firmicutes, Actinobacteria, and Spirochaete showed a mild-moderate correlation with the metabolome profile. The metabolomic profiles were mainly characterized by known brominated polyunsaturated fatty acids. This work presents findings about the composition and dynamics of the microbial assemblages of X. muta expanding and confirming current knowledge about its remarkable diversity and geographic variability as observed in this tropical marine biome.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Marine sponges (phylum Porifera) are the most ancient of the extant multicellular animals (metazoa) evolving some 800 million years ago [1,2,3,4]. They are sessile benthic organisms contributing to diversity and biomass flourishing in contrasting oceanic habitats such as shallow tropical, temperate, polar reefs, or abyssal plains [5, 6]. Sponges are pivotal in shaping ecosystems, due to their potential to influence benthic or pelagic processes [7]. Firstly, they are involved in structuring the seafloor and stabilization of substrate, offering refuge/habitat to a wide range of infauna species, and so they participate in complex biotic interactions with diverse macrobiotic taxa, influencing turbulence, spatial competition, and predation [7,8,9,10]. Secondly, sponges are key elements of biogeochemical cycles in coral reefs [11,12,13,14], participating in benthic-pelagic coupling of nutrient transfer. Indeed, they capture large amounts of food particles and other nutrients from the pelagic habitat via their capacity to pump thousands of liters per kilogram of sponge per day [7]. Like other animals, sponges have a well-developed innate immune system, with the ability to produce an arsenal of bioactive compounds including antimicrobials [6, 15, 16].
Marine sponges have formed close associations with a wide variety of microorganisms [15, 17,18,19,20], where bacteria can constitute up to 40% of the sponge tissue volume [21]. To date, 52 different microbial phyla have been associated with marine sponges [22]. A comprehensive review by Taylor et al. [15] suggested that approximately 15 bacterial phyla, two major archaeal lineages, and associated microbial eukaryotes can be found in sponges. However, in 2012, more than 25 bacterial phyla were reported [22] and the true number of microbial phyla actually associated with sponges is still not resolved.
Interest in sponge microbiology has two main justifications: ecological and biotechnological. First, the diverse assemblages of sponge-associated microbial communities are seen as models for early evolutionary microbiome interaction with higher organisms. Increasing knowledge on the diversity of sponge symbionts enables a better understanding of beneficial microbial consortia and the ecological and chemical functions of these microorganisms. Such associations contribute to the health and nutrition of sponges by (i) preserving the host from colonizers and grazers; (ii) producing antibiotics, repellent, or antagonism compounds; (iii) acquiring or synthesizing limiting nutrients; and (iv) processing metabolic waste [11,12,13, 23]. As sessile animals, marine sponges have evolved chemical defense strategies to thrive in competitive environmental conditions. One of the factors associated with adaptation is indeed the production of secondary metabolites [24]. Sponges continue to be the most productive sources of new bioactive compounds in the marine environment, accounting for approx. 50% of all marine natural products or secondary metabolites discovered between 1990 and 2009 [25]. Sponge holobionts are then considered a primary resource offering a wealth of novel chemical structures relevant to industry. While some of these natural products are of sponge origin, others have chemical features reminiscent of a microbial origin like polyketides or peptides [26]. Insights into the origin of these compounds will further aid towards the exploitation of the biotechnological potential of these holobionts [18, 26, 27].
Exploring the relationships between the structure of the symbiotic communities and the concentration of sponge-derived natural products provides insights into the real contribution of the microbiont in the production of important sponge metabolites. The giant barrel sponge Xestospongia muta is known for its rich source of diverse secondary metabolites (with over 40 metabolites identified so far) [28,29,30]. Being also a model “bacteriosponge” with clear biological fitness and the biggest sponge in the Caribbean Sea, X. muta is an excellent candidate to explore the sponge-associated microbial communities and their relationship with metabolite production [12, 31,32,33]. In this work, the stability of the symbiotic prokaryotic communities and the metabolomic profiles of X. muta were tested on different seasons and depths at Portobelo Bay marine biome, Tropical Western Caribbean Sea of Panamá.
Methods
Sponge Collection
Samples of the marine sponge Xestospongia muta (class: Demospongiae, order: Haplosclerida, family: Petrosiidae) were collected from the reef complex in the port of Portobelo (Panama) at five differing depths (9, 12, 18, 20, and 28 m) over two seasons, which correspond to the influences of autumn (September 2013, samples named as “S1”) and spring (May 2014, samples named as “S2”), periods corresponding to the annual Caribbean dry season [34]. Sampling sites are neotropical locations at the Northern Hemisphere and with less pronounced temperature differences. The average water temperature in September 2013 (S1) and May 2014 (S2) was 27 °C and 28 °C, respectively. Samples were collected from sponges estimated to be > 15 years of age [35] and showing no apparent signs of disease. For each depth and at each season, three independent sponge samples were collected via scuba diving at the same approximate location with 4 m between independent samples to obtain different asexual sponge clones (denoted as A, B, C for S1 and X, Y, Z for S2). A total of 30 samples were collected from the five different depths and two different seasons. A specimen corresponding to a volume of approximately 15 mL was collected from the sponge in the projecting edge of the side-wall osculum and placed in a 50-mL falcon tube underwater. Following the sampling, seawater was removed, and each specimen was placed into 30 mL of ethanol, transported in ice to the laboratory and stored at − 80 °C until further study.
Metagenomic DNA Extraction
Total DNA was extracted from ~ 0.5 g of sponge tissue using a combination of protocols from two established methods: one for the extraction of DNA from natural environments [36] and the other for marine sponge tissues [37]. In brief, the sponges were first ground and mixed with equal volume of sterile glass beads and 600 μL of extraction buffer (100 mM Tris-Cl, 100 mM EDTA, 1.5 M NaCl) were added. Six cycles of intermittent 30 s of vortexing and 30 s on ice were performed before centrifugation for 30 min (10 min at 2500×g and 20 min at 10,000×g at 4 °C). Three microliters of 10 mg/mL DNA-free RNAse were added to the total volume of supernatant and incubated at 37 °C for 1 h. After inactivation of the enzyme for 1 min (90 °C), an equal volume of phenol-chloroform-isoamyl alcohol (25:24:1, pH 8) was added. The aqueous phase was separated by centrifugation for 10 min (13,000 rpm at 4 °C) and then mixing with an equal volume of chloroform-isoamyl alcohol (24:1) followed by another 10 min centrifugation (13,000 rpm at 4 °C). The total DNA was precipitated from the extracted aqueous layer with 0.1× volume of 3 M sodium acetate and 0.6× volume of isopropanol. Subsequently, extracts were incubated overnight at − 20 °C followed by 30 min of centrifugation (13,000 rpm at 4 °C). The pelleted DNA was then washed in ice-cold 70% (v/v) ethanol, air dried, and re-suspended in 30 μL of MiliQ water.
16S rRNA Gene Library Construction and Sequencing
PCR amplicon libraries of the V5-V6 region of the 16S rRNA gene were prepared using the primers 807F [5′-GGATTAGATACCCBRGTAGTC-3′] and 1050R [5′-AGYTGDCGACRRCCRTGCA-3′] [38]. Each 25 μL reaction contained 1X buffer, 0.3 mM dNTPs, 0.5 μM forward primer, 0.5 μM reverse primer, 500 μg/mL BSA, 0.04 U/μL Accuzyme polymerase from Bioline, and 1 ng of template DNA. The PCR cycle conditions included initial denaturation at 95 °C for 2 min followed by 4 cycles of denaturation at 95 °C for 30 s, primer annealing at 60 °C for 30 s with a touchdown of 2 °C per cycle, and extension at 68 °C for 30 s. These cycles were followed by 26 cycles consisting of a denaturation at 95 °C for 30 s, primer annealing at 53 °C for 30 s, and extension at 68 °C for 30 s, and this was finally followed by a last extension at 68 °C for 7 min.
Libraries were constructed using a customized Illumina paired-end library sample preparation protocol [39]. Two subsequent overlapping PCRs were performed (6 cycles each at a constant annealing temperature of 55 °C) in fresh tubes and including all described components but changing the primers in order to attach the barcodes, adaptors, spacers, and indexes to the amplicons, and using as template 10−1 of the previous PCR reaction. A final pool with equal amounts of each sample (5 ng of final purified PCR product of each one) was prepared and sequenced on an Illumina-MiSeq platform [40], using ~ 250 nt paired-end sequencing chemistry.
Bioinformatic Analyses
All downstream sequence analyses were performed using QIIME 1.9.1 [41]. Approximately ~ 1.2 million reads were generated from the 30 independent sponge samples. As a first step, paired sequences were filtered (Q 25) and split according to the barcodes. Chimeric sequences were identified, extracted, and removed from the datasets using Usearch 6.1. A multi-step open-reference OTU picking workflow was performed within the QIIME system. Using this methodology, operational taxonomic units (OTUs) were picked assigning the reads to species groups based on 97% sequence similarity. This workflow used the PyNAST alignment [42] against the SILVA core set to assign OTUs. In the next steps, a single representative sequence for each OTU was again realigned using PyNAST to build a phylogenetic tree using FastTree. In total, 485,466 good-quality paired-end reads derived 24,417 OTUs, where the median sample size was 16,297 reads (min 7303 reads and max 32,682 reads) (Supplementary Figure 1a). Centroid sequences of core microbiome OTUs of X. muta found in this study had been deposited in GenBank database available under accession numbers KY274458–KY274480.
Metabolite Profiling
Each collected sponge specimen was also analyzed for its concentration of secondary metabolites. In brief, pre-frozen specimens were mechanically ground, then ~ 0.5 g of the dried sample was extracted three times with 10 mL a mixture of CH2Cl2/MeOH (1:1, v/v) under ultrasonication. The organic phases were combined and concentrated in vacuum on a bulk C18 solid phase added before evaporation (0.1 g). To focus on the secondary metabolites, a solid phase extraction (SPE) on C18 was performed as follows: the extract supported on the C18 phase was deposited on SPE cartridges (Strata-C18-E, Phenomenex, 500 mg) and compounds were eluted by a series of solutions (10 mL) of decreasing polarities: H2O (F1), H2O:MeOH (1:1) (F2), MeOH (F3), and CH2Cl2:MeOH (1:1) (F4). After evaporation of the solvents, fraction F3 was dissolved in methanol and ultra-high-performance liquid chromatography-diode array detector-evaporative light scattering detector (UHPLC-DAD-ELSD) analyses were performed [43].
The identification of the main metabolites of X. muta was performed beforehand with one injection in UHPLC-high-resolution mass spectrometry (UHPLC-HRMS) using the ultra-high resolution Qq-time-of-flight impact II (Bruker) in positive ion mode (Supplementary Figure 3). Due to the difficulty in the ionization of non-polar and halogenated compounds like unsaturated fatty acid derivatives by electrospray, metabolite profiling using DAD and ELSD analyses was preferred [44]. First, DAD allowed the detection of unsaturated fatty acids due to the presence of characteristic chromophores, then ELSD known as a universal detector revealed the presence of most types of metabolites. The resolution in the ELSD was found to be sufficient to relatively quantify the major specialized metabolites. Because major secondary metabolites from this sponge are known to be halogenated unsaturated fatty acids containing enyne or enediyne unsaturations, UV detection by DAD was appropriate for this study: one major band at around λmax 230 nm, as an indicative of enynes, and λmax 240, 250, 270, and 280 indicating enediynes [29]. UHPLC-DAD-ELSD analyses were therefore performed on an UltiMate 3000 UHPLC (Thermo Scientific) equipped with a DAD (Thermo Scientific) and an ELSD (Agilent). The column was a kinetex C18 (Phenomenex, 2.1 × 100 mm, 1.7 μm), with an injection volume of 5 μL and the flow rate 0.5 mL/min. The elution was set as follows: 0–3 min of isocratic H2O/CH3CN (90:10) then a linear gradient between 3 and 10 min until H2O/CH3CN (0:100), isocratic at this composition for 10–12 min and equilibration to the initial conditions for 12–15 min.
Statistical Analysis
Multivariate data analysis was used to explore and test for patterns in the global microbial and metabolite profiles across depth and season. The experimental design was set up as a two-way crossed design with season and depths being the main factors (effects) with the possibility to seek for an interaction effect (depth × season). First, each sample was compared to each other sample (using the different data matrices comprising either the global microbial OTUs or the secondary metabolites) via ordination using principal coordinate analysis (PCoA) following construction of sample-similarity matrix using either the weighted UniFrac distance for the global microbial profiles [45] or the Spearman rank correlation for the global metabolome profiles [46]. Significant differences between a priori predefined groups of samples (depth, season) were evaluated using the two-way permutational multivariate analysis of variance (PERMANOVA), allowing for type III (partial) sums of squares, fixed effects sum to zero for mixed terms, and exact p values generated using unrestricted permutation of raw data [47]. Groups of samples (depth, season) were considered significantly different if the p value falls < 0.05. These multivariate analyses were performed across a combination of software including PRIMER (v.6) PRIMER-E, Plymouth Marine Laboratory, UK, [46], QIIME, and the Phyloseq R package 1.5.21 [48]. Then, to test which of the 17 dominant phyla were significantly different across depths and seasons, a two-way ANOVA was performed using Prism v. 7.01 (Graphpad Software Inc.), while to test which of the nine metabolites were significantly different across depths and seasons, a two-way PERMANOVA was performed using PRIMER following the use of Euclidean distance to measure the difference between samples. Groups of samples (depth, season) were considered significantly different if the p value falls < 0.05.
To test for a correlation between global secondary metabolite profiles and microbiome composition, Mantel’s test was performed in PRIMER. Independent similarity matrices (as described above) were correlated using the Spearman rank correlation (9999 permutations). While the Mantel test highlights the significance of the relationships [49], it cannot provide information about any specific element of the data; thus, Spearman’s rank correlation was performed to determine the strength of each OTU on the concentration of each secondary metabolite [50]. In this case, the Spearman correlation coefficient was calculated using the stats and lineup packages of R. The scatterplot was constructed to show the dispersion of the coefficients using ggplot2 in the R package.
The microbial profiles were also compared using k-dominance plots and species diversity measures. Conventional measures of species diversity, richness, and evenness were calculated using algorithms for total OTUs (S), Shannon diversity (H′), Simpson (1-λ), and Pielou’s evenness (J′) [46]. Rarefaction analyses were performed to gauge for adequate sequencing depth per sample [40]. These univariate analyses were performed across a combination of software including PRIMER (v.6) PRIMER-E, Plymouth Marine Laboratory, UK [46], QIIME, and the Phyloseq R package 1.5.21 [48].
Availability of Data and Materials
Datasets of predominant sequence representatives of the main bacterial OTUs found in this study are available in the GenBank database under accession numbers KY274458–KY274480.
Results
Microbial Diversity of Xestospongia muta and Its Core Microbial Community
To explore the dynamics of the symbiotic community of marine sponge Xestospongia muta in the Port of Portobelo in Panama, microbial community structure and secondary metabolites were assessed across 30 sponge specimens collected in relation to depth and season. The microbial diversity was characterized by sequencing the V5-V6 region of the 16S rRNA gene using the Illumina platform. From a total of 485,466 good-quality paired-end sequences generated from the 30 sponge extracts, 24,417 microbial OTUs representing 24 microbial phyla were derived; 14 described bacteria phyla, 6 candidate phyla, 2 unclassified phyla (herein referred as Unclassified Candidates 1 and 2), and 2 archaea lineages. Between these, two highly dominant phyla were identified: Proteobacteria and Chloroflexi accounting for 61% of all reads (37.5% and 23.5% relative abundance, respectively). Other dominant phyla contributing to the global microbial assemblages were the uncultured sponge symbiont PAUC34f (6.6%), Actinobacteria (6.0%), Gemmatimonadetes (4.4%), Unclassified Candidate 1 (3.6%), Crenarchaeota (3.6), Cyanobacteria (3.4%), Nitrospirae (3.2%), Acidobacteria (2.2%), AncK6 (1.4%), Poribacteria (1.2%), and SBR1093 (1.2%) (Fig. 1, Table 1). The four most abundant phyla were also the four richest OTU phyla, where Proteobacteria, Chloroflexi, PAUC34f, and Actinobacteria were represented by 7935, 4517, 926, and 1395 OTUs, respectively. In contrast, two of the lowest predominant phyla LCP-89 and Tenericutes both contributing to ~ 34 reads are first reported as bacterial symbionts of X. muta [51,52,53]. Noteworthy, despite being described as a dominant sponge symbiont, the contribution of the phylum Poribacteria to the microbiome X. muta is only minor, accounting for 1.2% of the reads. Lastly, 119 OTUs were affiliated to an unclassified bacteria lineage here showed as Unclassified Candidate 1. Some authors have referred this taxonomic group as sponge-associated unidentified lineage (SAUL) [54, 55]. Although limited information is available for this lineage, it has also been reported in other marine environments, albeit at very low frequencies [56].
Although thousands of OTUs were derived (using 97% sequence identity to cluster reads), only 996 OTUs contributed to more than 30 sequence reads across all 30 samples, accounting for > 80% of the read counts, and there was a predominance of few OTUs. For example, OTU167 belonging to the class Gammaproteobacteria was present in all 30 samples and comprised 16,500 reads (or 3.4% of the total reads). Seven further OTUs representing the classes SAR202 (3), Nitrospira (2), Deltaproteobacteria (1), and Acidimicrobiia (1) were also predominant, each accounting for > 1% of the total reads and being present in most samples. With each of the 30 sponge specimens comprising between 150 and 250 microbial OTUs (as seen in the rarefaction curves, Supplementary Figure 1a) which indicates a high microbial richness for X. muta, most of the derived OTUs only contributed few sequences reads representing a sizable rare biosphere.
The core community of X. muta in the Port of Portobelo is composed of 23 OTUs (Fig. 2). The core microbial community is usually defined as the collection of OTUs present in at least 70% of collected sponge specimens; however, given that sponges were collected over differing seasons and depths, the defined core community of X. muta in this work are those OTUs present in all 30 sponge specimens. The cumulative relative abundance of these 23 OTUs varies between 10 and 35% within the different samples. Most of the OTUs belong to the phyla Proteobacteria and Chloroflexi (43.0% and 34.8% respectively) where Gammaproteobacteria and SAR202 were the most common represented classes. Other phyla represented in the core community were PAUC34f, Actinobacteria, Nitrospirae, Spirochaetes, and Poribacteria.
Microbial Community Changes Across Depth and Season
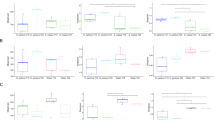
To assess whether the global microbial assemblages change with season and depth, ordination of the 30 samples via PCoA following the use of the weighted UniFrac distance metric was performed. As it was evident by the two-way crossed ANOVA results (Table 1), the most significant phyla variations were influenced by seasonal factors. PCoA revealed a clear shift with the season on the PCO1 axis (Fig. 3a) with all S1 (autumn) samples clustering independently to the right of the plot and all S2 (spring) samples to the left. Then when each season was ordinated separately, we observe a clear depth gradient in autumn (S1), with shallow and deeper depths separating on the PCO1 axis (Fig. 3b). While 9 and 12 m samples clustered together, they separated out from the 18 m samples on the PCO2 axis and were all independent from the 20 and 28 m samples. However, this depth pattern was not observed for spring (S2) (Fig. 3c). This observation was confirmed by the two-way PERMANOVA, which combined season with depth. Then there is a highly significant difference between autumn (S1) and spring (S2) (pseudo-F = 5.8976, p value = 0.0001) and a highly significant interaction effect with season/depth (pseudo-F = 2.1166, p value = 0.0001) (Table 2). This confirms the pattern observed in the PCoA plots where there was a strong depth gradient in autumn (S1) and a lack of this gradient in spring (S2), where there was a highly significant difference between depths during autumn (S1) (one-way PERMANOVA pseudo-F = 3.1975, p value = 0.0001), but no significant differences between depths during spring (S2) (one-way PERMANOVA pseudo-F = 1.1536, p value = 0.2491). Furthermore, out of the 17 dominant phyla, 8 were significantly different between seasons, comprising Cyanobacteria, Crenarchaeota, TM7, Gemmatimonadetes, Unclassified Candidate 1, SBR1093, Nitrospirae, and Acidobacteria, where PAUC34f, Actinobacteria, and Poribacteria showed a significant interaction effect with season/depth (two-way ANOVA p values < 0.05, Table 1).
Ordination of the global microbial assemblages and global metabolome profiles using principal coordinate analysis (PCoA). Plots on the left ordinate samples assessing the global microbial assemblages following the use of the weighted UniFrac distance algorithm, while plots on the right ordinate the global metabolome profiles following the use of Euclidean distance algorithm. Plots represent X. muta samples in respect to season (a, d) and depth (b, c, e, f)
The difference between spring and autumn can also be seen at the diversity level, where OTU richness (S), Shannon diversity (H′), Simpson diversity (1-lambda), and Pielou’s evenness (J′) were all higher for autumn (S1) (Fig. 4a–d). These diversity metrics can also be inferred from the k-dominance plots (Supplementary Figure 1b-d), where curves for the autumn (S1) samples fall below those for spring (S2) further indicating that these samples comprised less dominant taxa, having more even abundance profiles between taxa and overall greater species richness (Fig. 4d). When assessing k-dominance of the depths during autumn (S1), there appears to be a depth gradient with the shallower depths being less dominated by taxa, having more even abundance between taxa (Supplementary Figure 1b-c). Notably, from 18 m in depth, there was an evident decline in the mean Shannon diversity (H′) and Pielou’s evenness (J′) compared to the shallower depths (Fig. 4). In spring (S2), these patterns were not observed (Fig. 4, Supplementary Figure 1d).
Metabolome Patterns Across Season and Depth
To assess changes in the secondary metabolite production with season and depth, UV/ELSD metabolome profiles were generated for each sponge specimen (Supplementary Figure 2). Comparison of UV profiles (Supplementary Figure 3) of each derived compound with those previously isolated from X. muta [28, 29, 57] allowed the assignment of each peak to a known family of metabolite. Because ELSD is known as a universal detector with areas roughly proportional to the quantity of each compound, the focus here was on the major compounds of the profiles, corresponding to the peaks with RT 7.95 (1), 8.15 (2), and 8.35 (3) min. A strong and large band was observed at λmax 229 for compound 1 while four fine bands were observed at λmax 238, 253, 272, and 283 for 2 and 3, all these data being consistent with enyne- and enediyne-conjugated systems, respectively [29, 57]. On the basis of UV and HRMS data, the following identification was made: (7E,13E,15Z)-14,16-dibromohexadeca-7,13,15-triene-5-ynoic acid (1) (Supplementary Figure 2(b)) at m/z 420.0170, 422.0149, 424.0130 (ratio 1:2:1, [M + NH4]+) [29], (7E,13E,17E)-18-bromooctadeca-7,13,17-triene-5,15-diynoic acid (2) (Supplementary Figure 2(c)) at m/z 366.1064, 368.1045 (ratio 1:1, [M + NH4]+) [57], and a non-described dihomo derivative (3) of this compound at m/z 394.1378, 396.1358 (ratio 1:1, [M + NH4]+, retention time 8.35 min). Thus, the metabolomic profile comprised two distinct brominated polyunsaturated fatty acids.
The global metabolome profile comprising the nine metabolite bands was then compared across all 30 samples using the Spearman rank correlation. A two-way PERMANOVA crossed season with depth and showed that there were no significant differences between the main factors: spring and autumn, or depths. However, there was a significant interaction effect with season/depth (pseudo-F = 3.3703, p value = 0.0078) (Table 3). This lack of separation of points denoting autumn and spring is evident on the PCoA ordination (Fig. 3d). Furthermore, there was a significant difference in the metabolome profiles between depths during autumn (S1) (one-way PERMANOVA, pseudo-F = 2.9812, p value = 0.0097), but not between depths during spring (S2) (one-way PERMANOVA, pseudo-F = 2.3519, p value = 0.0749) (Table 3). However, this depth gradient is not as evident in the metabolome profile as it was in the microbiome profile (comparing Fig. 3c–e). Out of the nine measured metabolites, only RT_7.46 was significantly different between spring and autumn (Table 4), while RT_7.72, RT_7.95, and RT_8.15 showed a significant interaction effect with season/depth (two-way PERMANOVA p values < 0.05, Table 3).
Relationship Between Global Metabolome Profile and Microbial Community Composition
Because shifts in the microbial community were observed with season and depth, and a corresponding interaction effect was also deduced for the metabolome profile, a global relationship between the two variables was assessed. Using Mantel’s test, the pattern generated between the 30 samples derived from the global microbial community data was matched with the one generated by the metabolomic analysis. There is no clear correlation between both global datasets (Mantel’s Rho = 0.03874, p value < 0.05). However, when testing each of the phyla abundances independently, four phyla have a significant relationship with the global metabolome pattern: Cyanobacteria (Mantel’s Rho = 0.376, p value 0.0028), Firmicutes (Mantel’s Rho = 0.395, p value 0.026), Actinobacteria (Mantel’s Rho = 0.198, p value 0.020), and Spirochaete (Mantel’s Rho = 0.222, p value 0.039), all showing a moderate correlation with changes to the metabolome profile. Further tests were performed at each OTU-metabolite level, where most of the Spearman rank correlation coefficients fell between − 0.2 and 0.2 (Fig. 5). This suggests a lack of direct relationships between any particular OTU and metabolite combination in the conditions assayed and from the techniques used.
Discussion
Marine sponges are one of the most dominant sessile groups of the coral reefs and there are some evidences that in the mesophotic zone, their biomass and diversity increase with depth. This change occurs due in part to the increasing particulate organic carbon and differences in predation [58]. Despite these evidences, only few studies have analyzed whether their microbiome presents also depth-dependent changes [58, 59]. The giant barrel sponge Xestospongia muta is a good model for investigating these questions as it is (i) one of the most abundant sponge species in the Caribbean Sea, (ii) a high microbial abundance (HMA) bacteriosponge with a diverse symbiotic microbial community, and (iii) it is recognized as a rich source of secondary metabolites. Clarifying the shifts and stability of microbiome and metabolomic profile of marine sponges across a depth gradient and through different seasons is of importance for understanding the holobiont interaction and the involvement of symbionts in the secondary metabolite biosynthesis.
In the present study, besides the significant interactions of some taxonomic groups with depth/season, a core set of 23 OTUs was present in all 30 sampled X. muta specimens, suggesting that these taxa could be used to “ecotype” this sponge species in this ecoregion. HMA sponges, such as X. muta, are characterized by maintaining their complex microbial communities by a combination of mechanisms of vertical and horizontal transmission [60]; however, it has been found that only a few members of the specific adult microbial community are vertically transferred and only a small portion of them are maintained after the uptake of symbionts during filter-feeding periods of the host [60]; OTUs from the core set community found in this study could represent those vertically transferred symbiotic association of X. muta with more biological fitness.
Nevertheless, when assessing their taxonomic relatedness and their relative proportions of the global microbial assemblages (comprising thousands of OTUs per sponge specimen), some differences were noticed between the seasons spring (May 2014) and autumn (September 2013) which border the mean annual Caribbean dry season. This difference was evident when assessing changes with depths: in autumn, there was a clear depth gradient with shallow sponges’ microbiome being more similar and then being different to the deeper sponge specimens. This was not evident in spring some months later. Interestingly, such a trend was also observed in the global metabolome profile, although not as strong between the seasons. There are several possible explanations for such a loss of a depth gradient as, for example, a variability and/or mixing in abiotic factors as the temperature gradient, turbidity, light intensity, or nutrient availability.
As it has been previously reported, Proteobacteria and Chloroflexi were the predominant phyla in the X. muta’s microbiome [51, 61]. Other phyla such as PAUC34f, Actinobacteria, Gemmatimonadetes, Crenarchaeota, Cyanobacteria, Nitrospirae, Acidobacteria, AncK6, Poribacteria, and SBR1093 were also found associated with this species [53]. Eight of the dominating phyla were significantly different between seasons, comprising Cyanobacteria, Crenarchaeota, TM7, Gemmatimonadetes, Unclassified Candidate 1, SBR1093, Nitrospirae, and Anck6. Phyla PAUC34f, Actinobacteria, Gemmatimonadetes, Crenarchaeota, Cyanobacteria, TM7, and Poribacteria showed a significant interaction effect with season/depth (Table 1). Some of these taxa have been associated with functions of health and nutrition for sponges. For example, the ecological role of Cyanobacteria is well known, and it has been demonstrated that photosynthesized glycerol and organic phosphates are transferred from bacteria to the sponge host supplying up to 50% of its energy and 80% of its carbon budget [62]. Despite a low taxonomic resolution of the cyanobacterial sequences, it is likely that the cyanobacterial representatives encountered in X. muta in this work are the symbiont “Candidatus Synechococcus spongiarum,” which represents an independent cyanobacterial lineage that has become a sponge symbiont from past times and it is phylogenetically equidistant from the marine and freshwater strains of Synechococcus/Prochlorococcus subclade (CitamBio-2015). “Candidatus Synechococcus spongiarum” is the most widespread cyanobacterial symbiont found in marine sponges [63] and it has an extremely flexible photosynthetic apparatus that allows them to grow in a wide range of irradiances [64]. This may explain why, in autumn (S1), the proportion of cyanobacterial population increased in deeper communities.
Genomics studies focused on “Ca. Synechococcus spongiarum” have revealed a variety of clades that with high 16S rRNA sequence identity (~ 99%) may differ widely in genomic databases, due to physiological and functional variations [63]. As it has been identified in four different clades of this candidate symbiont, one of the detected modifications is the deletion of the psbP gene. This modification could lead to a less efficient photosynthetic system affecting their productivity and ability to assimilate carbon and transfer it to the host sponge [65], and potentially decreasing also the competitiveness under low light exposures [63]. This variety of clades that may differ in genomic repertoire and in physiology and biological function as symbionts may be the explanation for the differences observed in the patterns of this cyanobacterial class between the two seasons, where the decreasing number of Cyanobacteria during spring (S2) through the depth gradient was also statistically significant.
In the same way as in Cyanobacteria, Crenarchaeota and Gemmatimonadetes showed a significant interaction between the variables depth and season. Crenarchaeota, as the most abundant lineage of archaea, was first reported as sponge-associated microorganisms in 1996 [66] and they are now recognized as important components of the natural microbiota of many bacteriosponges [67]. Its ecological role has been related to ammonia-oxidizing activity which is one of the first reactions carried out to achieve nitrification [67]. This is an important biological process since sponges release ammonia as a metabolic waste product and its accumulation may have adverse effects on sponge health [67]. In the same way, the role of Gemmatimonadetes has been also associated with nutrition processes. The culturable representatives of this phylum have shown the ability to accumulate intracellular polyphosphates [68], a property absolutely beneficial for the sponges, as phosphorous (P) is one of the limiting elements in the marine ecosystems. Its accumulation by the symbionts could then enhance the recycling of P in the local environment of the marine sponge [13]. Trend in S2 to the increasing accumulation of these phyla in deeper areas could be related to the increasing production of metabolic wastes by the sponge. In both seasons, the samples collected in shallow waters were less diverse, showing a greater proportion of dominant clusters in their microbial communities. This may be related to the increasing availability of particulate food in deeper zones. The increased availability of particulate matter with depth has been considered as one of the reasons explaining why deep sponges grow significantly faster than their shallow congeners [69]. Therefore, mesophotic sponges may be less dependent of their associated bacterial communities for nutrition and digestion [59].
The metabolome profiles of X. muta revealed the dominant presence of brominated compounds [30]. The patterns and statistical analysis of the metabolomic data indicate that there is not a considerable influence of season or depth in the relative contribution of the metabolites to the global metabolome. However, the compounds (1) and (2) characterized here as the main compounds of the X. muta metabolome presented a significant interaction effect of the two variables, suggesting that these two compounds have a pattern influenced by depth in spring and autumn. Because the symbiotic community taxonomical composition shifts are not directly linked to the concentrations of these two main metabolites, the differential production detected is modulated either by changes in gene expression of activities encoded in the marine sponge genome or in the associated microbiome and/or due to environmental conditions, i.e., those enriching particulate matters available, captured and filtered with metabolite precursors.
To study in detail the involvement of symbiotic bacteria in the biosynthesis of secondary metabolites of marine sponges, it is necessary to go deeper into functional analyses such as (meta) genomics and (meta) transcriptomics of the holobiont. Although previous evidences suggest that the biosynthesis occurs in a collaborative process between the sponges and its microbial associate community [26], there are several bacterial phylotypes that could be involved as functional units of biosynthesis together with sponge cells, and this may be the explanation of the lack of direct correlation between any pair of OTU/metabolite in our results.
Notwithstanding, we found a moderate correlation between the phyla Cyanobacteria and Firmicutes with the global metabolome profile. Cyanobacteria isolated from marine sponges had been found able to produce brominated metabolites [70], a chemical feature of the main secondary metabolites found in the specimens of X. muta. Our results and the evidence from previous studies suggest that Cyanobacteria may be involved not only in the nutrition of the host, but also in the development of its chemical defense system. On the other hand, members of the phylum Firmicutes have been considered important for the health of the sponges, because they have been found to have the ability to produce substances with antimicrobial and antifungal properties [71]. These metabolic features suggest that members of these phyla might be involved in the chemical defense system of their host protecting them from both predators and microbial pathogens. In a recent transcriptomic holobiont profiling survey of X .muta specimens thriving in the northern Caribbean sea [72], it was detected the correspondence in actively transcribed genes ascribed to central metabolite production and degradation thus evidencing examples of interplay between host and microbiome functions, but the homology-based automatic annotation of genes involved in peripheral metabolism or in specialized secondary metabolite production or modification remains a challenging issue, as it is to define the origin and shared contribution of the sponge and the hosted microbiome in key abundant metabolites for a given sponge species.
The diversity of microbial symbionts may confer benefits in health and nutrition to their host, but they can also contribute to a larger metabolic repertoire. However, the mechanisms responsible for the emergence and maintenance of this diversity are still open questions including how related species from geographically isolated regions can acquire shared bacterial symbionts when they were apparently absent (below the detection limit of the technologies applied) from the seawater. These fundamental questions in sponge evolutionary biology may find some first clues on the shared OTUs between individuals of X. muta sampled at different depths and season (this study), and in different regions [51], together with future continuous spatio-temporal sequential tracking, which support the idea of coevolution of some of the sponge-associated bacteria in the course of the holobiont consolidation. Our results provide additional evidence to the hypothesis considering that there is not only horizontal transfer of symbiotic microorganisms from the surrounding aquatic environment, but also vertical transmission [6, 19, 56].
References
Li CW, Chen JY, Hua TE (1998) Precambrian sponges with cellular structures. Science 279:879–882. https://doi.org/10.1126/science.279.5352.879
Yin Z, Zhu M, Davidson EH, Bottjer DJ, Zhao F, Tafforeau P (2015) Sponge grade body fossil with cellular resolution dating 60 Myr before the Cambrian. Proc Natl Acad Sci U S A 112:E1453–E1460. https://doi.org/10.1073/pnas.1414577112
Van-Soest R, Boury-Esnault N, Vacelet J et al (2012) Global diversity of sponges (Porifera). PLoS One 7:e35105. https://doi.org/10.1371/journal.pone.0035105
Gold DA, Grabenstatter J, De Mendoza A et al (2016) Sterol and genomic analyses validate the sponge biomarker hypothesis. Proc Natl Acad Sci 1–6:2684–2689. https://doi.org/10.1073/pnas.1512614113
Dayton PK (1989) Interdecadal variation in an antarctic sponge and its predators from oceanographic climate shifts. Science 245:1484–1486. https://doi.org/10.1126/science.245.4925.1484
Hentschel U, Piel J, Degnan SM, Taylor MW (2012) Genomic insights into the marine sponge microbiome. Nat Rev Microbiol 10:641–654. https://doi.org/10.1038/nrmicro2839
Bell JJ (2008) The functional roles of marine sponges. Estuar Coast Shelf Sci 79:341–353. https://doi.org/10.1016/j.ecss.2008.05.002
Deignan LK, Pawlik JR (2014) Perilous proximity: does the Janzen-Connell hypothesis explain the distribution of giant barrel sponges on a Florida coral reef? Coral Reefs 34:561–567. https://doi.org/10.1007/s00338-014-1255-x
Rix L, Naumann MS, de Goeij JM et al (2015) Coral mucus fuels the sponge loop in warm- and cold-water coral reef ecosystems. Scientific 6:1–11. https://doi.org/10.1038/srep18715
Pawlik JR, McMurray SE, Erwin P, Zea S (2015) No evidence for food limitation of Caribbean reef sponges: reply to Slattery & Lesser (2015). Mar Ecol Prog Ser 527:281–284. https://doi.org/10.3354/meps11308
De Goeij JM, Van Oevelen D, Vermeij MJA et al (2013) Surviving in a marine desert: the sponge loop retains resources within coral reefs. Science 342:108–110. https://doi.org/10.1126/science.1241981
Loh T-L, Pawlik JR (2014) Chemical defenses and resource trade-offs structure sponge communities on Caribbean coral reefs. Proc Natl Acad Sci U S A 111:4151–4156. https://doi.org/10.1073/pnas.1321626111
Zhang F, Blasiak LC, Karolin JO, Powell RJ, Geddes CD, Hill RT (2015) Phosphorus sequestration in the form of polyphosphate by microbial symbionts in marine sponges. Proc Natl Acad Sci 201423768:4381–4386. https://doi.org/10.1073/pnas.1423768112
Gatti S (2002) The role of sponges in high-Antarctic carbon and silicon cycling—a modelling approach. Ber Polarforsch Meeresforsch 434:1–134
Taylor MW, Radax R, Steger D, Wagner M (2007) Sponge-associated microorganisms: evolution, ecology, and biotechnological potential. Microbiol Mol Biol Rev 71:295–347. https://doi.org/10.1128/MMBR.00040-06
Indraningrat AAG, Smidt H, Sipkema D (2016) Bioprospecting sponge-associated microbes for antimicrobial compounds. Mar Drugs 14:1–66. https://doi.org/10.3390/md14050087
Li Z-Y, He L-M, Wu J, Jiang Q (2006) Bacterial community diversity associated with four marine sponges from the South China Sea based on 16S rDNA-DGGE fingerprinting. J Exp Mar Biol Ecol 329:75–85. https://doi.org/10.1016/j.jembe.2005.08.014
Wang G (2006) Diversity and biotechnological potential of the sponge-associated microbial consortia. J Ind Microbiol Biotechnol 33:545–551. https://doi.org/10.1007/s10295-006-0123-2
Webster NS, Taylor MW (2012) Marine sponges and their microbial symbionts: love and other relationships. Environ Microbiol 14:335–346. https://doi.org/10.1111/j.1462-2920.2011.02460.x
Hentschel U, Usher KM, Taylor MW (2006) Marine sponges as microbial fermenters. FEMS Microbiol Ecol 55:167–177. https://doi.org/10.1111/j.1574-6941.2005.00046.x
Vacelet J, Donadey C (1975) Electron microscope study of the association between some sponges and bacteria. Bio Ecol 30:301–314
Webster NS, Thomas T (2016). Defining the Sponge Hologenome 7:1–14. https://doi.org/10.1128/mBio.00135-16.Invited
Ribes M, Jiménez E, Yahel G, López-Sendino P, Diez B, Massana R, Sharp JH, Coma R (2012) Functional convergence of microbes associated with temperate marine sponges. Environ Microbiol 14:1224–1239. https://doi.org/10.1111/j.1462-2920.2012.02701.x
Thoms C, Schupp P (2007) Chemical defense strategies in sponges: a review. Porifera Res Biodiversity Innov Sustain 627–637
Leal MC, Puga J, Serôdio J, Gomes NCM, Calado R (2012) Trends in the discovery of new marine natural products from invertebrates over the last two decades—where and what are we bioprospecting? PLoS One 7:e30580. https://doi.org/10.1371/journal.pone.0030580
Sacristán-Soriano O, Banaigs B, Casamayor EO, Becerro MA (2011) Exploring the links between natural products and bacterial assemblages in the sponge aplysina aerophoba. Appl Environ Microbiol 77:862–870. https://doi.org/10.1128/AEM.00100-10
Sipkema D, Franssen MCR, Osinga R, Tramper J, Wijffels RH (2005) Marine sponges as pharmacy. Mar Biotechnol 7:142–162. https://doi.org/10.1007/s10126-004-0405-5
Morinaka BI, Skepper CK, Molinski TF (2007) Ene-yne tetrahydrofurans from the sponge Xestospongia muta. Exploiting a Weak CD Effect for Assignment of Configuration 1:3–6
Schmitz FJ, Gopichand Y (1978) (7E,13E,15Z)-14,16-dibromo-7,13,15-hexadecatrien-5-ynoic acid. A novel dibromo acetylenic acid from the marine sponge Xestospongia muta. Tetrahedron Lett 3637–3640
Zhou X, Lu Y, Lin X, Yang B, Yang X, Liu Y (2011) Brominated aliphatic hydrocarbons and sterols from the sponge Xestospongia testudinaria with their bioactivities. Chem Phys Lipids 164:703–706. https://doi.org/10.1016/j.chemphyslip.2011.08.002
McMurray SE, Finelli CM, Pawlik JR (2015) Population dynamics of giant barrel sponges on Florida coral reefs. J Exp Mar Biol Ecol 473:73–80. https://doi.org/10.1016/j.jembe.2015.08.007
Gloeckner V, Wehrl M, Moitinho-Silva L, Gernert C, Schupp P, Pawlik JR, Lindquist NL, Erpenbeck D, Wörheide G, Hentschel U (2014) The HMA-LMA dichotomy revisited: an electron microscopical survey of 56 sponge species. Biol Bull 227:78–88
Zhou X, Xu T, Yang X-W, Huang R, Yang B, Tang L, Liu Y (2010) Chemical and biological aspects of marine sponges of the genus Xestospongia. Chem Biodivers 7:2201–2227. https://doi.org/10.1002/cbdv.201000024
Giannini A, Kushnir Y, Cane MA (2000) Interannual variability of Caribbean rainfall, ENSO, and the Atlantic Ocean. J Clim 13:297–311. https://doi.org/10.1175/1520-0442(2000)013<0297:IVOCRE>2.0.CO;2
McMurray SE, Blum JE, Pawlik JR (2008) Redwood of the reef: growth and age of the giant barrel sponge Xestospongia muta in the Florida keys. Mar Biol 155:159–171. https://doi.org/10.1007/s00227-008-1014-z
Griffiths R, Whiteley A (2000) Rapid method for coextraction of DNA and RNA from natural environments for analysis of ribosomal DNA-and rRNA-based microbial community composition. Appl Environ Microbiol 66:1–5. https://doi.org/10.1128/AEM.66.12.5488-5491.2000.Updated
Simister RL, Schmitt S, Taylor MW (2011) Evaluating methods for the preservation and extraction of DNA and RNA for analysis of microbial communities in marine sponges. J Exp Mar Biol Ecol 397:38–43. https://doi.org/10.1016/j.jembe.2010.11.004
Bohorquez LC, Delgado-Serrano L, López G, Osorio-Forero C, Klepac-Ceraj V, Kolter R, Junca H, Baena S, Zambrano MM (2012) In-depth characterization via complementing culture-independent approaches of the microbial community in an acidic hot spring of the Colombian Andes. Microb Ecol 63:103–115. https://doi.org/10.1007/s00248-011-9943-3
Galvez E, Junca H, Riaño D (2012) Assessment of microbial composition and degradation functions in PAH contaminated neotropical Caribbean marine sediments (Cartagena Bay, Colombia). Universidad de Los Andes
Camarinha-Silva A, Juregui R, Chaves-Moreno D et al (2014) Comparing the anterior nare bacterial community of two discrete human populations using Illumina amplicon sequencing. Environ Microbiol 16:2939–2952. https://doi.org/10.1111/1462-2920.12362
Kuczynski J, Stombaugh J, Walters WA et al (2011) Using QIIME to analyze 16S rRNA gene sequences from microbial communities. Curr Protoc Bioinformatics Chapter 10:Unit 10.7. https://doi.org/10.1002/0471250953.bi1007s36
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R (2010) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26:266–267. https://doi.org/10.1093/bioinformatics/btp636
Ivanišević J, Thomas OP, Lejeusne C, Chevaldonné P, Pérez T (2010) Metabolic fingerprinting as an indicator of biodiversity: towards understanding inter-specific relationships among Homoscleromorpha sponges. Metabolomics 7:289–304. https://doi.org/10.1007/s11306-010-0239-2
Greff S, Zubia M, Payri C, Thomas OP, Perez T (2017) Chemogeography of the red macroalgae Asparagopsis: metabolomics, bioactivity, and relation to invasiveness. Metabolomics 13:0. https://doi.org/10.1007/s11306-017-1169-z
Hamady M, Lozupone C, Knight R (2010) Fast UniFrac: facilitating high-throughput phylogenetic analyses of microbial communities including analysis of pyrosequencing and PhyloChip data. ISME J 4:17–27. https://doi.org/10.1038/ismej.2009.97
Clarke KR, Warwick RM (2001) Change in marine communities: an approach to statistical analysis and interpretation, 2nd edition. Prim Plymouth UK p. 172
Anderson MJ (2001) A new method for non parametric multivariate analysis of variance. Austral Ecol 26:32–46. https://doi.org/10.1111/j.1442-9993.2001.01070.pp.x
McMurdie PJ, Holmes S (2013) phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8:e61217. https://doi.org/10.1371/journal.pone.0061217
Hollister EB, Engledow AS, Hammett AJM, Provin TL, Wilkinson HH, Gentry TJ (2010) Shifts in microbial community structure along an ecological gradient of hypersaline soils and sediments. ISME J 4:829–838. https://doi.org/10.1038/ismej.2010.3
Barberan A, Bates ST, Casamayor EO, Fierer N (2012) Using network analysis to explore co-occurrence patterns in soil microbial communities. Isme J 6:343–351. https://doi.org/10.1038/ismej.2011.119
Fiore CL, Jarett JK, Lesser MP (2013) Symbiotic prokaryotic communities from different populations of the giant barrel sponge, Xestospongia muta. Microbiologyopen 2:938–952. https://doi.org/10.1002/mbo3.135
Montalvo NF, Hill RT (2011) Sponge-associated bacteria are strictly maintained in two closely related but geographically distant sponge hosts. Appl Environ Microbiol 77:7207–7216. https://doi.org/10.1128/AEM.05285-11
Thomas T, Moitinho-Silva L, Lurgi M, Björk JR, Easson C, Astudillo-García C, Olson JB, Erwin PM, López-Legentil S, Luter H, Chaves-Fonnegra A, Costa R, Schupp PJ, Steindler L, Erpenbeck D, Gilbert J, Knight R, Ackermann G, Victor Lopez J, Taylor MW, Thacker RW, Montoya JM, Hentschel U, Webster NS (2016) Diversity, structure and convergent evolution of the global sponge microbiome. Nat Commun 7:11870. https://doi.org/10.1038/ncomms11870
Webster NS, Taylor MW, Behnam F, Lücker S, Rattei T, Whalan S, Horn M, Wagner M (2010) Deep sequencing reveals exceptional diversity and modes of transmission for bacterial sponge symbionts. Environ Microbiol 12:2070–2082. https://doi.org/10.1111/j.1462-2920.2009.02065.x
Schmitt S, Tsai P, Bell J, Fromont J, Ilan M, Lindquist N, Perez T, Rodrigo A, Schupp PJ, Vacelet J, Webster N, Hentschel U, Taylor MW (2012) Assessing the complex sponge microbiota: core, variable and species-specific bacterial communities in marine sponges. ISME J 6:564–576. https://doi.org/10.1038/ismej.2011.116
Simister RL, Deines P, Botté ES, Webster NS, Taylor MW (2012) Sponge-specific clusters revisited: a comprehensive phylogeny of sponge-associated microorganisms. Environ Microbiol 14:517–524. https://doi.org/10.1111/j.1462-2920.2011.02664.x
Patil A, Kokke W, Cochran S et al (1992) Brominated polyacetylenic acids from the marine sponge Xestospongia muta: inhibitors of HIV protease. J Nat Prod 55:1170–1177
Morrow KM, Fiore C, Lesser M (2016) Environmental drivers of microbial community shifts in the giant barrel sponge, Xestospongia muta, over a shallow to mesophotic depth gradient. Environ Microbiol n/a-n/a. https://doi.org/10.1111/1462-2920.13226
Olson JB, Gao X (2013) Characterizing the bacterial associates of three Caribbean sponges along a gradient from shallow to mesophotic depths. FEMS Microbiol Ecol 85:74–84. https://doi.org/10.1111/1574-6941.12099
Schmitt S, Angermeier H, Schiller R, Lindquist N, Hentschel U (2008) Molecular microbial diversity survey of sponge reproductive stages and mechanistic insights into vertical transmission of microbial symbionts. Appl Environ Microbiol 74:7694–7708. https://doi.org/10.1128/AEM.00878-08
Lesser MP, Fiore C, Slattery M, Zaneveld J (2016) Climate change stressors destabilize the microbiome of the Caribbean barrel sponge, Xestospongia muta. J Exp Mar Biol Ecol 475:11–18. https://doi.org/10.1016/j.jembe.2015.11.004
Wilkinson C (1978) Microbial associations in sponges. III. Ultrastructure of the in situ associations in coral reef sponges. Mar Biol 49:177–185
Burgsdorf I, Slaby BM, Handley KM, Haber M, Blom J, Marshall CW, Gilbert JA, Hentschel U, Steindler L (2015) Lifestyle evolution in cyanobacterial symbionts of sponges. MBio 6:1–14. https://doi.org/10.1128/mBio.00391-15
Usher KM (2008) The ecology and phylogeny of cyanobacterial symbionts in sponges. Mar Ecol 29:178–192. https://doi.org/10.1111/j.1439-0485.2008.00245.x
Freeman CJ, Thacker RW, Baker DM, Fogel ML (2013) Quality or quantity: is nutrient transfer driven more by symbiont identity and productivity than by symbiont abundance? ISME J 7:1116–1125. https://doi.org/10.1038/ismej.2013.7
Preston CM, Wu KY, Molinskit TF, Delong EF (1996) A psychrophilic crenarchaeon inhabits a marine sponge. Proc Natl Acad Sci U S A 93:6241–6246
Steger D, Ettinger-Epstein P, Whalan S, Hentschel U, de Nys R, Wagner M, Taylor MW (2008) Diversity and mode of transmission of ammonia-oxidizing archaea in marine sponges. Environ Microbiol 10:1087–1094. https://doi.org/10.1111/j.1462-2920.2007.01515.x
Zhang H, Sekiguchi Y, Hanada S et al (2003) Gemmatimonas aurantiaca gen. nov., sp. nov., a Gram-negative, aerobic, polyphosphate-accumulating micro-organism, the first cultured representative of the new bacterial phylum Gemmatimonadetes phyl. nov. Int J Syst Evol Microbiol 53:1155–1163. https://doi.org/10.1099/ijs.0.02520-0
Lesser MP (2006) Benthic–pelagic coupling on coral reefs: feeding and growth of Caribbean sponges. J Exp Mar Biol Ecol 328:277–288. https://doi.org/10.1016/j.jembe.2005.07.010
Unson MD, Holland ND, Faulkner DJ (1994) A brominated secondary metabolite synthesized by the cyanobacterial symbiont of a marine sponge and accumulation of the crystalline metabolite in the sponge tissue. Mar Biol 119:1–11. https://doi.org/10.1007/BF00350100
Kennedy J, Baker P, Piper C, Cotter PD, Walsh M, Mooij MJ, Bourke MB, Rea MC, O’Connor PM, Ross RP, Hill C, O’Gara F, Marchesi JR, Dobson ADW (2009) Isolation and analysis of bacteria with antimicrobial activities from the marine sponge Haliclona simulans collected from Irish waters. Mar Biotechnol 11:384–396. https://doi.org/10.1007/s10126-008-9154-1
Fiore CL, Labrie M, Jarett JK, Lesser MP (2015) Transcriptional activity of the giant barrel sponge, Xestospongia muta Holobiont: molecular evidence for metabolic interchange. Front. Microbiol. https://doi.org/10.3389/fmicb.2015.00364, 6
Acknowledgments
We thank the additional support from Dr. Regis Guillaume, Embassy of France in Colombia, and Raúl de León from Dive Adventure Panama and Portobelo Dive Center for his guidance and collaboration during the dive sampling, to Iris Plumeier and Silke Kahl for excellent technical assistance and to the members of the BIOMMAR research group for their helpful technical advices.
Funding
The authors would like to thank financial support granted by the European Commission to EC/FP7 research project consortium MAGICPAH (KBBE-2009-245226), to the Colombian Administrative Department for Science, Technology and Innovation - Colciencias for financial support through grants EcosNord-Colciencias (Convocatoria 652-2014) “International cooperation and research mobility grants in marine research between Colombia and France 2015-2017” and “Joven Investigador” 2012 fellowship to M. Villegas-Plazas and to Universidad de los Andes for the MSc scholarship to M. Villegas-Plazas.
Author information
Authors and Affiliations
Contributions
MV analyzed and interpreted the microbial community data and wrote the manuscript. OT analyzed and interpreted the metabolomic data. MO supported statistical analyses. HJ conceptualized, analyzed the results, and supervised the study. JS contributed with the experimental design and DP with the sequencing and analyzing the results. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
Not applicable
Consent for Publication
Not applicable
Competing Interests
The authors declare that they have no competing interests.
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Villegas-Plazas, M., Wos-Oxley, M.L., Sanchez, J.A. et al. Variations in Microbial Diversity and Metabolite Profiles of the Tropical Marine Sponge Xestospongia muta with Season and Depth. Microb Ecol 78, 243–256 (2019). https://doi.org/10.1007/s00248-018-1285-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-018-1285-y