Abstract
Key message
We identified 21 new and stable QTL, and 11 QTL clusters for yield-related traits in three bread wheat populations using the wheat 90 K SNP assay.
Abstract
Identification of quantitative trait loci (QTL) for yield-related traits and closely linked molecular markers is important in order to identify gene/QTL for marker-assisted selection (MAS) in wheat breeding. The objectives of the present study were to identify QTL for yield-related traits and dissect the relationships among different traits in three wheat recombinant inbred line (RIL) populations derived from crosses Doumai × Shi 4185 (D × S), Gaocheng 8901 × Zhoumai 16 (G × Z) and Linmai 2 × Zhong 892 (L × Z). Using the available high-density linkage maps previously constructed with the wheat 90 K iSelect single nucleotide polymorphism (SNP) array, 65, 46 and 53 QTL for 12 traits were identified in the three RIL populations, respectively. Among them, 34, 23 and 27 were likely to be new QTL. Eighteen common QTL were detected across two or three populations. Eleven QTL clusters harboring multiple QTL were detected in different populations, and the interval 15.5–32.3 cM around the Rht-B1 locus on chromosome 4BS harboring 20 QTL is an important region determining grain yield (GY). Thousand-kernel weight (TKW) is significantly affected by kernel width and plant height (PH), whereas flag leaf width can be used to select lines with large kernel number per spike. Eleven candidate genes were identified, including eight cloned genes for kernel, heading date (HD) and PH-related traits as well as predicted genes for TKW, spike length and HD. The closest SNP markers of stable QTL or QTL clusters can be used for MAS in wheat breeding using kompetitive allele-specific PCR or semi-thermal asymmetric reverse PCR assays for improvement of GY.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bread wheat is one of the most important staple crops worldwide. Genetic improvement of wheat yield potential has been made in many countries in Europe (Austin et al. 1980; Brancourt-Hulmel et al. 2003), North America (McCaig and DePauw 1995; Donmez et al. 2001), South America (Ortiz-Monasterio et al. 1997) and Oceania (Siddique et al. 1989) over the past few decades. Significant genetic progress has also been made in improving yield potential of Chinese wheat cultivars during the past 50 years (Zhou et al. 2007; Zheng et al. 2011; Xiao et al. 2012; Gao et al. 2017). However, the gradual decrease in farmland, rapid increase in global population, decline in available water and climate change are important factors necessitating yield improvement. Thus, increased grain yield (GY) is the main goal in wheat breeding. Considering the complex, polygenic inheritance, low heritability, and significant influence of environment, yield improvement continues to be a huge challenge.
Wheat GY comprises three main components, viz. spike number per unit area (SN), kernel number per spike (KNS) and thousand-kernel weight (TKW). Increases in grain yield were mainly associated with increased components as well as harvest index (HI) (Donmez et al. 2001; Brancourt-Hulmel et al. 2003). Kernel, spike, plant height (PH) and flag leaf-related traits can also affect GY through impacting yield components. Among the three yield components, SN and KNS are more easily influenced by environment. Numerous quantitative trait loci (QTL) for these two traits have been identified, and one gene TaTEF-7A related to KNS was cloned (Cuthbert et al. 2008; Deng et al. 2011; Lee et al. 2014; Zheng et al. 2014; Gao et al. 2015; Sun et al. 2017). Previous studies showed higher heritabilities of TKW than SN and KNS, varying from 0.59 to 0.80 (Xiao and He 2003). TKW is determined by kernel dimensions, such as kernel length (KL) and kernel width (KW) (Zhang et al. 2014a; Huang et al. 2015). More than 100 QTL for TKW have been mapped on all 21 wheat chromosomes (Gegas et al. 2010; Ramya et al. 2010; Gao et al. 2015; Sukumaran et al. 2015; Wu et al. 2015; Sun et al. 2017), and over 20 kernel weight-related genes were cloned (Ma et al. 2010; Jiang et al. 2011; Su et al. 2011; Yang et al. 2012; Zhang et al. 2012b; Chang et al. 2013; Guo et al. 2013; Chang et al. 2014; Dong et al. 2014; Qin et al. 2014; Zhang et al. 2014b; Hanif et al. 2015; Wang et al. 2015; Ma et al. 2016; Wang et al. 2016b; Zhang et al. 2017c). KNS and TKW can be defined by spike length (SL) and dry weight (SDW) that are easier to select in small plots in wheat breeding. Genetic studies on these two traits were not given enough attention previously, particularly SDW. Heading date (HD), which is critical for optimal crop adaptation, yield potential and stability, is controlled by genes for vernalization (Vrn), photoperiod (Ppd) and earliness per se (Eps). Previous studies indicated that HD associated genes were distributed on almost all 21 chromosomes and expressed at different developmental stages (Griffiths et al. 2009; Huang et al. 2011; Zhang et al. 2012a; Takenaka and Kawahara 2013; Zikhali et al. 2014; Muterko et al. 2016).
Following deployment of the Green Revolution genes (Rht-B1b and Rht-D1b) much success was achieved in breeding wheat varieties with increased lodging resistance, higher HI, and higher yield potential (Peng et al. 1999; Ellis et al. 2005). To date, 24 Rht genes have been identified; Rht-B1b, Rht-D1b and Rht8 are the three dwarfing genes that are widely used in wheat cultivars (Peng et al. 1999; Hedden 2003; Ellis et al. 2005; Chen et al. 2015; Tian et al. 2017). Recently, Rht24 was identified on chromosome 6AL. This gene was widely distributed in elite wheat varieties, and reduced PH by 6.0–7.9 cm (Tian et al. 2017; Würschum et al. 2017). Varieties with long uppermost internode length (UIL) are preferred in wheat breeding, and related genes were mapped (Yu et al. 2014; Zhang et al. 2017b). Previous studies and production practice indicated that PH had a significant influence on yield-related traits (Peng et al. 1999; McCartney et al. 2005; Cuthbert et al. 2008; Gao et al. 2015).
Flag leaf area, including flag leaf length (FLL) and width (FLW), does not only affect photosynthetic performance (Xu and Zhao 1995; Sharma et al. 2003), but also determine population structure and impact plant stress response (Sourdille et al. 2002; Perez–Perez et al. 2010). Thus, flag leaf area is significantly associated with GY. The TaFLW1 gene associated with FLW was finely mapped close to Fhb5 on chromosome 5A (Xue et al. 2013). Although several QTL for FLL and FLW have been identified, it is necessary to identify more common and stable QTL for marker-assisted selection (MAS) in wheat breeding (Jia et al. 2013; Xue et al. 2013; Wu et al. 2016).
The International Wheat Genome Sequencing Consortium (IWGSC 2014) identified more than 124,000 genes in bread wheat. However, only about 65 important genes have been cloned and 150 functional markers were developed for these genes (Liu et al. 2012; Rasheed et al. 2016; Nadolska-Orczyk et al. 2017). Although numerous yield-related QTL have been identified (Azadi et al. 2015; Cui et al. 2014; Gao et al. 2015; Sun et al. 2017), there is still a large gap between the discovery of QTL, gene cloning and MAS due to the low heritability of related traits, limited number of genetic markers and the narrow genetic background of single mapping populations in previous studies (Nadolska-Orczyk et al. 2017).
Single nucleotide polymorphisms (SNP) are abundant in plant and animal genomes (Wang et al. 2014). SNPs are now widely used in QTL mapping and genome-wide association studies (GWAS) for economically important crops and animals (Wiedmann et al. 2008; Ganal et al. 2011; Song et al. 2013). The rice (44 K) and maize (50 and 55 K) SNP arrays are widely used in detecting new genes (Tung et al. 2010; Zhao et al. 2011; Cook et al. 2012; Xu et al. 2017a, b). Recently, the wheat 90 K iSelect SNP genotyping array was used to construct high-density linkage maps for QTL analysis in bread and durum wheats (Wang et al. 2014; Gao et al. 2015; Jin et al. 2016; Zhai et al. 2016; Wen et al. 2017). In addition, once genes and QTL have been cloned, gene-based SNP haplotypes can be identified to improve selection of target alleles in breeding programs (Rasheed et al. 2016).
The aims of the present study were to: (1) identify QTL and tightly linked SNP markers for yield-related traits using three RIL populations, (2) detect QTL clusters or pleiotropic loci associated with those traits, and (3) investigate genetic relationships among different yield-related traits.
Materials and methods
Plant materials and field trials
Three RIL populations derived from crosses Doumai × Shi 4185 (D × S, 275 F2:6 RILs), Gaocheng 8901 × Zhoumai 16 (G × Z, 176 F2:6 RILs) and Linmai 2 × Zhong 892 (L × Z, 273 F2:6 RILs) were used for constructing high-density linkage maps and QTL mapping. Doumai is a winter wheat line with a larger spike, TKW and flag leaf area and a lower tillering capacity; Shi 4185 (pedigree: Zhi 8094/Baofeng 7228//Shi 84-7120) is a winter wheat cultivar released in Hebei province, which has a smaller spike and TKW, and a higher tillering capacity. Gaocheng 8901 (pedigree: 77546-2/Linzhangmai) was cultivated in Hebei province with a taller plant height (above 90 cm), a smaller flag leaf width and good processing quality, while Zhoumai 16 (pedigree: Zhoumai 9/Zhou 8425B) is a widely grown facultative wheat cultivar in Henan province, which has a short stature (about 70 cm) and a larger spike and flag leaf width. Linmai 2 (pedigree: Lumai 23/Lin 90-15) is a semi-winter wheat cultivar in Shandong province with a larger spike and TKW and a lower tillering capacity, whereas Zhong 892 is a semi-winter wheat line from Henan province with a fewer KNS, a shorter stature (about 73 cm), and a narrow flag leaf.
The D × S population was grown at Shunyi in Beijing and Shijiazhuang in Hebei province during the 2012–2013, 2013–2014 and 2014–2015 cropping seasons; the G × Z and L × Z populations were grown at Anyang in Henan province and Suixi in Anhui province during the 2012–2013 and 2013–2014 cropping seasons. A randomized complete block design with three replicates was employed. Each plot comprised three 1.5 m rows spaced 20 cm apart, with 50 seeds in each row. Agronomic management was performed according to local practices at each location.
Phenotyping
Twelve phenotypic traits, SN, KNS, TKW, KL, KW, SL, SDW, HD, PH, UIL, FLL and FLW, were assessed in each population (Tables S1, S2 and S3).
Among yield components, SN was estimated by counting spikes at physiological maturity in one meter row section of each plot and expressed as m−2; KNS was measured by the mean kernel number of 20 randomly selected spikes in each plot at physiological maturity; and TKW was determined by the weight of 500 kernels from each plot after harvest. For kernel-related traits, 20 kernels were randomly selected from each plot after harvest for measuring KL and KW, represented by the mean length and width of 20 kernels, respectively. For spike-related traits, 20 spikes from 20 single plants in each plot were randomly selected at physiological maturity for measuring SL and SDW, the mean distance between the base and the top excluding the awns and the mean weight of 20 spikes after drying at 75 °C for 48 h, respectively. HD was recorded at half-spike emergence on a plot basis and represented as days from sowing to heading. For PH-related traits, 10 single plants in each plot were randomly selected at physiological maturity for measuring PH and UIL as the mean distance between the stem base and the top of spikes excluding awns and the mean length of the uppermost internode, respectively. For flag leaf area-related traits, 10 random flag leaves in each plot at the mid grain-fill stage were used to measure FLL and FLW, represented by the distance between the base and the tip, and width at the widest point, respectively.
Statistical analysis
For each trait, best linear unbiased estimation (BLUE) were calculated across environments using the ANOVA function in IciMapping V4.0 software (Li et al. 2007; http://www.isbreeding.net) assuming fixed effects for the genotype. Analysis of variance (ANOVA) was computed using the PROC GLM procedure in SAS 9.4 software (SAS Institute, http://www.sas.com). Broad-sense heritabilities (h2) were calculated following Nyquist and Baker (1991). PROC CORR procedure was used to calculate Pearson’s correlation coefficients (r) among the traits based on BLUE values.
Genotyping and high-density linkage map construction
The populations were genotyped using the wheat 90 K iSelect SNP array outsourced from Capital Bio Corporation, Beijing, China (http://www.capitalbiotech.com/). To reduce the influence of low-quality SNPs on the mapping results, three approaches were used to control the SNP data: (1) the heterozygous loci were treated as missing data, (2) the SNPs with missing data > 10% were filtered, and (3) allelic frequencies at a locus should be between 0.3 and 0.7.
The construction of high-density linkage maps was previously described by Wen et al. (2017) and the main steps are as follows: firstly, the BIN function of the IciMapping V4.0 software (Li et al. 2007; http://www.isbreeding.net) was used to place the SNP markers with no recombination into one bin and identify markers with minimum percentage of missing data as frame markers; secondly, the “Grouping” function in JoinMap 4.0 (Stam, 1993) was employed in sorting the frame markers into different groups with LOD thresholds ≥ 7; thirdly, the “AutoMap” command in MapDisto 1.7 (Lorieux, 2012) was used to order the frame markers and calculate genetic distances; finally, based on the CSS database (IWGSC RefSeq v1.0, https://urgi.versailles.inra.fr/blast_iwgsc/blast.php), linkage groups were assigned and oriented on chromosomes.
QTL analysis
The frame markers identified in each bin were used for QTL mapping in IciMapping V4.0 software (Li et al. 2007; http://www.isbreeding.net). Composite interval mapping (ICIM) was chosen to search for QTL of phenotypic traits of all lines from each environment and BLUE values. LOD scores to declare significant QTL for all traits ranged from 2.0 to 2.5 across environments following 2000 permutations at P = 0.01; thus a LOD threshold of 2.5 was chosen for declaration of putative QTL. QTL detected in different environments for one trait were considered to be the same if the distance between the LOD contour peaks is less than 20 cM. QTL for same traits identified in more than half of the environments tested were considered to be stable.
QTL position comparison and candidate gene discovery
QTL identified in each population were integrated on the consensus map constructed in our previous study (Wen et al. 2017). The positions of QTL for the same traits in different populations were compared based on consensus maps (Wang et al. 2014; Wen et al. 2017). Flanking sequences of SNPs linked to QTL within a confidence interval (Darvasi and Soller 1997) were also used to blast against the CSS database (IWGSC RefSeq v1.0, https://urgi.versailles.inra.fr/blast_iwgsc/blast.php) to identify the positions of QTL on the physical map. QTL for same traits mapped in different populations were considered to be the same if their positions were close in all three maps.
Candidate genes for QTL were determined using the following two approaches. Firstly, sequences of cloned genes on the same chromosome as QTL for each trait were used to blast against the CSS database. The gene was considered to be a candidate gene if its location was the same as SNPs mapped within a confidence interval to the LOD contour peak of the QTL on the physical map. Secondly, the flanking sequences of SNPs mapped within a confidence interval to the LOD contour peak of QTL were used to blast against the NCBI database (http://www.ncbi.nlm.nih.gov/) to identify putative gene functions. BLAST hits were filtered to an e-value threshold of 10−5 with sequence similarities higher than 75%.
Results
Phenotypic evaluation
The distributions of all 12 traits revealed continuous variation and transgressive segregation in the three populations, indicating polygenic inheritance. Among these, SN, KNS, TKW, SL, SDW, PH, UIL, FLL and FLW exhibited larger ranges in the populations, whereas KL, KW and HD showed less variation (Tables S1, S2 and S3).
ANOVA showed highly significant effects (P < 0.01) of lines and environments on all traits (Tables S8, S9 and S10). KNS, TKW, KL, KW, SL, PH and FLW had high h2 (above 0.80) in all three populations, whereas SN, SDW, HD, UIL and FLL showed moderate h2 (0.60–0.80), indicating that most of these traits were stable and mainly determined by genetic factors (Tables S8, S9 and S10).
SN exhibited significant (P < 0.01) and negative correlations with KNS, TKW, KW, SDW and FLW in both the D × S and L × Z populations (Tables S11, S12 and S13). KNS was significantly and positively correlated with SDW and FLW in all three populations (Tables S11, S12 and S13). TKW had significant and positive correlations with KL, KW and PH in all three populations, and the highest correlation was between TKW and KW (r = 0.71–0.82) in all three populations (Tables S11, S12 and S13).
High-density linkage maps
Based on the criteria mentioned above, 11,012 (D × S), 11,979 (G × Z) and 10,443 (L × Z) markers were used for the construction of genetic linkage maps (Wen et al. 2017). For the D × S population, all polymorphic markers were assigned to 2851 recombinant bins, and the total length of the linkage map constructed by the frame markers is 2030.0 cM, with an average frame marker density of 1.40 markers per cM (Tables S4 and S7). For the G × Z population, all polymorphic markers were placed into 3284 recombinant bins, covering a total length of 3130.3 cM, with an average frame marker density of 1.04 markers per cM (Tables S5 and S7). For the L × Z population, all polymorphic markers were sorted into 3489 recombinant bins, and the total length of the linkage map constructed by the frame markers is 2954.8 cM, with an average frame marker density of 1.08 markers per cM (Tables S6 and S7).
QTL for yield-related traits
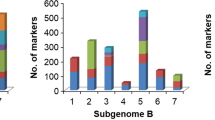
Sixty five, 46 and 53 QTL for yield-related traits were identified in the D × S, G × Z and L × Z populations, respectively (Table 1 and Fig. 1).
QTL for yield-related traits on the wheat consensus map (Wen et al. 2017). Markers are indicated on the right side, and corresponding centimorgan (cM) distances are shown on the left. QTL confidence intervals with a LOD score ≥ 2.5 are indicated by vertical bars. Different shapes and colors represent QTL identified in different populations and traits, respectively
QTL for yield components
Nine QTL for SN were mapped on chromosomes 1AL, 1BL, 2BS, 3AL, 4BS (2), 5DL, 6AL and 7BS, with 4, 3 and 2 in the D × S, G × Z and L × Z populations, respectively (Table 1). QSN.caas-4BS was stably identified in the D × S and L × Z populations, explaining 22.0–59.0 and 16.0–23.3% of the phenotypic variances across environments, respectively (Tables 1, 3). QSN.caas-1AL (closest marker IWB40317) identified in two environments and BLUE value in the G × Z population explained 9.1–16.2% of the phenotypic variance.
Sixteen QTL for KNS were identified on chromosomes 1BS, 1BL (2), 3DL, 4AL, 4BS (3), 5BL, 5DL, 7AS, 7AL (2), 7BS, 7BL and 7DS, with 5, 7 and 4 in the D × S, G × Z and L × Z populations, respectively (Table 1). QKNS.caas-4BS was stably detected in the D × S and L × Z populations, accounting for 9.9–43.1 and 17.8–36.6% of the phenotypic variances, respectively (Tables 1, 3). QKNS.caas-7AL was also stably identified in the D × S and L × Z populations, explaining 3.5–4.7 and 2.7–5.2% of the phenotypic variances, respectively. QKNS.caas-3DL (closest marker IWB66510) in the G × Z population was significant in two environments, accounting for 8.0–10.8% of the phenotypic variance.
Seventeen QTL for TKW were detected on chromosomes 2AS, 2BS, 3AL, 3B, 3DL, 4AL, 4BS (2), 4DS (2), 5AL (2), 5DL, 6AL, 6BL, 7AL and 7BL, with 8, 3 and 6 in the D × S, G × Z and L × Z populations, respectively (Table 1). QTKW.caas-4BS was stably identified in the D × S and G × Z populations, explaining 12.1–45.6 and 6.6–9.2% of the phenotypic variances, respectively (Tables 1, 3). QTKW.caas-5AL, QTKW.caas-5DL and QTKW.caas-6BL in the D × S population and QTKW.caas-3B and QTKW.caas-7AL.1 in the L × Z population were detected in most environments and BLUE values. QTKW.caas-6AL (closest marker IWA428) in the L × Z population was found in two environments and BLUE value, accounting for 10.3–18.8% of the phenotypic variance.
QTL for kernel-related traits
Eleven QTL for KL were mapped on chromosomes 1AL, 1BS, 1BL, 4BL, 4DS, 5BS, 5DL, 6AL, 6DL, 7AS and 7DS, with 6, 3 and 2 in the D × S, G × Z and L × Z populations, respectively (Table 1). QKL.caas-5BS and QKL.caas-5DL in the D × S population and QKL.caas-1BL and QKL.caas-6DL in the L × Z population were stably identified in all environments and BLUE values. QKL.caas-1BS (closest marker IWB31844) in the G × Z population was significant in three environments and BLUE value, explaining 8.2–17.1% of the phenotypic variance.
Thirteen QTL for KW were identified on chromosomes 2BL, 3AL, 3B (2), 4AS, 4AL, 4BS, 4DS, 5AL, 5BL, 5DL, 6AL and 7AL, with 4, 3 and 6 in the D × S, G × Z and L × Z populations, respectively (Table 1). QKW.caas-4AL and QKW.caas-4BS in the D × S population, QKW.caas-3AL and QKW.caas-5AL in the G × Z population and QKW.caas-2BL, QKW.caas-3B.2 and QKW.caas-4AS in the L × Z population were stably identified in most of the environments and BLUE values. QKW.caas-3B.1 explained 8.4–13.9% of the phenotypic variance in the G × Z population.
QTL for spike-related traits
Seventeen QTL for SL were detected on chromosomes 1AL, 2DS (3), 2DL, 3B, 3DL, 4AL, 4BS, 5AL, 6AL (2), 6BL (4) and 6DL, with 6, 6 and 5 in the D × S, G × Z and L × Z populations, respectively (Table 1). QSL.caas-2DS stably detected in the G × Z and L × Z populations accounting for 21.1–29.9 and 3.2–8.3% of the phenotypic variances, respectively (Tables 1, 3); QSL.caas-6AL was stably identified in the D × S and L × Z populations, explaining 1.7–3.6 and 2.7–4.8% of the phenotypic variances, respectively; QSL.caas-6BL.1 was stably detected in all three populations, accounting for 9.8–28.5, 4.8–9.2 and 10.4–19.1% of the phenotypic variances, respectively. QSL.caas-2DS.1, QSL.caas-4AL, QSL.caas-5AL.2 in the G × Z population, QSL.caas-2DL in the D × S population and QSL.caas-4BS in the L × Z population were stably identified in four or five environments and BLUE values.
Ten QTL for SDW were identified on chromosomes 1BS, 4BS (2), 5DL (2), 6AL (2), 6BL, 7AS and 7BL, with 4, 3 and 3 in the D × S, G × Z and L × Z populations, respectively. QSDW.caas-4BS was stably detected in the D × S and L × Z populations, explaining 37.2–42.9 and 6.4–19.3% of the phenotypic variances, respectively (Tables 1, 3). QSDW.caas-6BL (closest marker IWB73837) in the D × S population was significant in two environments and BLUE value, accounting for 15.2–28.3% of the phenotypic variance; QSDW.caas-5DL.2 (closest marker IWB12546) in the D × S population was stably detected in all four environments and BLUE value.
QTL for HD
Nineteen QTL for HD were mapped on chromosomes 1BL, 1DL (2), 2AL, 2BS (2), 2DS, 3AS, 3AL (2), 4BS, 5AL, 5DL (2), 6BL (2), 7BS, 7BL and 7DL, with 6, 6 and 7 in the D × S, G × Z and L × Z populations, respectively (Table 1). Three common QTL were detected on chromosomes 2BS, 5DL and 6BL in the D × S and L × Z populations, accounting for 5.1–16.0, 3.0–16.5 and 2.7–11.0% of the phenotypic variances, respectively (Tables 1, 3). QHD.caas-1DL.2 (closest marker IWB49684) was a major QTL and stably identified in all environments and BLUE value in the G × Z population.
QTL for PH-related traits
Eighteen QTL for PH were identified on chromosomes 1BL, 2AL, 2BL (2), 3AS, 3AL, 3B, 4BS (2), 4BL, 4DS (2), 5AL (2), 5BL, 6BL (2) and 6DS, with 9, 5 and 4 in the D × S, G × Z and L × Z populations, respectively (Table 1). QPH.caas-4BS was stably identified in the G × Z and L × Z populations, accounting for 13.2–21.1 and 7.4–23.7% of the phenotypic variances, respectively; QPH.caas-4DS was also stably identified in the G × Z and L × Z populations, explaining 9.5–15.0 and 6.7–18.5% of the phenotypic variances, respectively; QPH.caas-6BL was stably detected in the D × S and L × Z populations, accounting for 2.3–4.8 and 3.4–11.9% of the phenotypic variances, respectively (Tables 1, 3). QPH.caas-1BL, QPH.caas-5AL.1 and QPH.caas-6DS in the D × S population and QPH.caas-2BL.2 in the L × Z population were stably identified in all environments and BLUE values.
Fifteen QTL for UIL were detected on chromosomes 2AS (3), 2AL, 2BL, 3AL, 3DL, 4AS, 4BS (3), 4DS, 5AL, 6BL and 7DS, with 6, 4 and 5 in the D × S, G × Z and L × Z populations, respectively (Table 1). QUIL.caas-2AS was stably identified in the D × S and L × Z populations, accounting for 8.5–10.8 and 2.0–3.2% of the phenotypic variances, respectively; QUIL.caas-4BS stably detected in all three populations, explained 4.0–7.0, 7.1–17.2 and 7.0–10.7% of the phenotypic variances, respectively (Tables 1, 3). QUIL.caas-4DS and QUIL.caas-6BL were significant and stable across environments, explaining 7.9–18.6 and 10.4–13.8% of the phenotypic variances in the G × Z and L × Z populations, respectively. QUIL.caas-3AL and QUIL.caas-4AS in the L × Z population, and QUIL.caas-3DL in the D × S population were stably identified in all environments and BLUE values.
QTL for flag leaf-related traits
Six QTL for FLL were mapped on chromosomes 2AL, 2BL, 3B, 4AS, 4BS and 5DL, with 3 and 3 in the D × S and L × Z populations, respectively (Table 1). QFLL.caas-3B (closest marker IWB54417) was significant in two environments and BLUE value in the L × Z population, explaining 10.1–14.1% of the phenotypic variances.
Thirteen QTL for FLW were identified on chromosomes 1BL, 2BS (2), 4AL (2), 4BS (2), 5AL (2), 5BL, 5DL, 6AS and 6BL, with 4, 3 and 6 in the D × S, G × Z and L × Z populations, respectively (Table 1). QFLW.caas-2BS was stably detected in the G × Z and L × Z populations, accounting for 10.5–14.3 and 1.9–4.8% of the phenotypic variances, respectively; QFLW.caas-4BS, stably identified in the D × S and L × Z populations, explained 2.8–15.5 and 11.5–22.1% of the phenotypic variances, respectively (Tables 1, 3). QFLW.caas-4AL.2 (closest marker IWB38036) in the G × Z population was significant in two environments and BLUE value. QFLW.caas-5BL and QFLW.caas-5DL in the D × S population, and QFLW.caas-5AL.1, QFLW.caas-6AS and QFLW.caas-6BL in the L × Z population were stably identified in all environments and BLUE values.
QTL clusters and pleiotropic loci
Eleven QTL clusters were identified on chromosomes 1BL, 2AS, 3DL, 4BS, 4DS, 5AL (2), 5DL, 6AL (2) and 6BL (Table 2). Clusters on chromosomes 4BS, 4DS and 6BL were observed in all three populations, whereas clusters on chromosomes 5DL and 6AL (interval 74.3–75.9 cM) were identified in the D × S and L × Z populations. Among these QTL clusters, 10 were associated with kernel-related traits, except for that on chromosome 3DL. The interval 15.5–32.3 cM on chromosome 4BS harboring 20 QTL for all traits except for KL, with 7, 4 and 9 QTL in the D × S, G × Z and L × Z populations, respectively; intervals 32.7–45.1 cM and 17.0–30.3 cM on chromosomes 2AS and 4DS, respectively, were associated with both kernel and PH-related traits; intervals 12.7–50.4 cM and 86.6–105.7 cM on chromosomes 5DL and 6BL harboring 11 and 9 QTL, respectively, were important in determining yield-related traits.
Discussion
Comparison with previous studies
The Green Revolution genes Rht-B1b and Rht-D1b greatly contributed to wheat yield potential through improving lodging resistance and modifying dry matter distribution (Peng et al. 1999; Ellis et al. 2005). Blasting results indicated that the QTL for PH on chromosomes 4BS and 4DS in the G × Z and L × Z populations were Rht-B1b and Rht-D1b, respectively (Peng et al. 1999). These two genes were also detected by Gao et al. (2015) and Sun et al. (2017) using the wheat 90 K iSelect SNP array. Interestingly, we identified QTL for SN, KNS, TKW, KW, SDW, UIL and FLW around the Rht-B1 locus on chromosome 4BS with large contributions in the D × S population, whereas no QTL for PH was detected. Sequencing of this locus indicated that there is a large deletion in Doumai, whereas the allele in Shi 4185 is Rht-B1b (data not shown). Four QTL for KNS, TKW, PH and UIL in the G × Z population and nine QTL for SN, KNS, SL, SDW, HD, PH, UIL, FLL and FLW in the L × Z population were detected around the Rht-B1 locus. These were attributed to the effect of Rht-B1, in agreement with previous reports (Cuthbert et al. 2008; Deng et al. 2011; Zhang et al. 2017b). Gao et al. (2015) identified QTL for TKW and KW at the Rht-B1 locus, and QTL were similarly reported by Gegas et al. (2010) and Ramya et al. (2010).
Six QTL for PH and kernel-related traits around the Rht-D1 locus were detected in the G × Z and L × Z populations, indicating a lesser effect on yield-related traits than Rht-B1. Gao et al. (2015) identified four QTL for GY, chlorophyll content and normalized difference in vegetation index around Rht-D1. Cuthbert et al. (2008) and Zhang et al. (2017b) both reported a QTL-rich region for yield-related traits around the Rht-B1 locus, whereas no QTL-rich regions around Rht-D1 were identified in these studies.
Many studies to date have implied that SN is controlled by polygenes (Cuthbert et al. 2008; Cui et al. 2014; Lee et al. 2014; Gao et al. 2015). A minor QTL on chromosome 6AL identified in the present study was also detected by Gao et al. (2015). Another QTL on chromosome 3AL is at a similar position with that in Lee et al. (2014). The major QTL on chromosome 1AL detected in two environments and BLUE value is likely to be new.
QKNS.caas-7AL, stably identified in the D × S and L × Z populations, is at the same position as SNP wsnp_Ex_c11047_17915103 detected by Sun et al. (2017). Another minor QTL on chromosome 7BS was previously observed by Gao et al. (2015). QKNS.caas-1BL.2 and QKNS.caas-7BL are close to those in Xu et al. (2017a, b) and Wang et al. (2011a), respectively. The major QTL on chromosome 3DL is likely to be new.
TKW-associated QTL were mapped on all 21 chromosomes in previous studies (Gegas et al. 2010; Ramya et al. 2010; Gao et al. 2015; Wu et al. 2015; Sukumaran et al. 2015; Cheng et al. 2017; Sun et al. 2017). A gene for TKW on chromosome 7D was finely mapped by Röder et al. (2008) with the SSR marker Xgwm1002-7D. Blasting results revealed that QTKW.caas-3B and QTKW.caas-6AL are probably TaGS5 and TaTPP-6AL1, respectively (Wang et al. 2015; Ma et al. 2016; Wang et al. 2016b; Zhang et al. 2017c), whereas QTKW.caas-2BS, QTKW.caas-3DL.1 and QTKW.caas-7AL.1 are different from the cloned genes TaSus2-2B (2BS), TaCKX6-D1 (3D) and TaGASR7-A1 (7AL) (Jiang et al. 2011; Zhang et al. 2012b; Dong et al. 2014), respectively. The 3AL QTL is at the same position as SNP Excalibur_c39508_88 associated with TKW (Sun et al. 2017). In addition, QTKW.caas-2AS, QTKW.caas-4AL.1, QTKW.caas-4DS.1 and QTKW.caas-6BL are located at similar positions to those in Cui et al. (2014), Wu et al. (2015), Mohler et al. (2016) and Mir et al. (2012), respectively. QTKW.caas-5AL, QTKW.caas-5DL and QTKW.caas-7AL.1 are likely to be new QTL detected in most environments and BLUE values.
Blasting results indicated that QKL.caas-6DL is probably TaGS1a (Guo et al. 2013); QKL.caas-1BL and QKL.caas-5BS are at the same positions as those in Wu et al. (2015) and Zhai et al. (2018), respectively. QKL.caas-4DS, co-localized with QTKW.caas-4DS, is at similar position to that in Gegas et al. (2010). In addition, QKL.caas-1AL, QKL.caas-1BS, QKL.caas-6AL, QKL.caas-7AS and QKL.caas-7DS are at similar positions to those in Cui et al. (2014), Mohler et al. (2016), Li et al. (2015), Tyagi et al. (2014) and Gegas et al. (2010), respectively. QKL.caas-5DL, co-localized with QTKW.caas-5DL and stably identified in all six environments and BLUE value, is likely to be a new QTL.
QKW.caas-6AL, co-localized with QTKW.caas-6AL, is likely to be TaTPP-6AL1 (Zhang et al. 2017c); QKW.caas-3B.2, identified in all four environments and BLUE value, is at the same position as that in Zhai et al. (2018). QKW.caas-3AL, co-localized with QTKW.caas-3AL, is at a similar position to SNP Excalibur_c39508_88 for TKW (Sun et al. 2017). QKW.caas-5BL and QKW.caas-5DL are at similar positions to those in Cui et al. (2014) and Mohler et al. (2016), respectively. QKW.caas-5AL, co-localized with QTKW.caas-5AL.3, is likely to be a new QTL. QKW.caas-2BL, QKW.caas-3B.1, QKW.caas-3B.2, QKW.caas-4AS and QKW.caas-4AL are also new QTL.
QSL.caas-6BL.1, identified in all three populations, is at the same position as those in Gao et al. (2015) and Sun et al. (2017); QSL.caas-2DS, detected in the G × Z and L × Z populations, is at a similar position to that in Azadi et al. (2015); QSL.caas-6AL, identified in the D × S and L × Z populations, is at a similar position to QTL reported previously (Lee et al. 2014; Liu et al. 2014). QSL.caas-3B, QSL.caas-3DL, QSL.caas-4AL and QSL.caas-5AL.2 are also in similar positions to those reported in Azadi et al. (2015), Liu et al. (2014), Gao et al. (2015) and Wang et al. (2011a), respectively. QSL.caas-2DS.1 and QSL.caas-2DL identified in four or five environments and BLUE values are likely to be new.
QSDW.caas-6BL and QSDW.caas-7BL are at similar positions as SNPs RAC875_c31299_1302 and BS00055584_51 identified by Valluru et al. (2017), respectively. QSDW.caas-4BS stably identified in the D × S and L × Z populations is likely to be a new QTL. QSDW.caas-5DL detected in all four environments and BLUE value is also likely to be new.
Blasting results showed that QHD.caas-5DL in the D × S and L × Z populations should be contributed by Vrn-D1 (Fu et al. 2005); QHD.caas-7BS is about 6 Mb from Vrn-B3 locus (Yan et al. 2006) on the physical map (IWGSC RefSeq v1.0, https://urgi.versailles.inra.fr/blast_iwgsc/blast.php); QHD.caas-2BS, identified in the D × S and L × Z populations and close to the Eps QTL detected by Le Gouis et al. (2012), is about 56 Mb from Ppd-B1 locus (Nishida et al. 2013); QHD.caas-2DS is about 46 Mb from Ppd-D1 (Beales et al. 2007). Because there are no variations at Vrn-B3 and Ppd-D1 loci in the parents of three populations, QHD.caas-7BS and QHD.caas-2DS should be different from Vrn-B3 and Ppd-D1, respectively. Comparison of the physical locations of QHD.caas-2BS and Ppd-B1 also shows that the two loci are different. Previously, Wang et al. (2016a) cloned gene TaELF3-1DL associated with early flowering on chromosome 1DL using the G × Z population. This locus is consistent with QHD.caas-1DL.2 in the present study and at the same position as the major Eps QTL flanked by Xgdm111 and Xbarc62 (Griffiths et al. 2009; Zikhali et al. 2014). Another QTL on chromosome 1DL (QHD.caas-1DL.1) about 60 Mb from QHD.caas-1DL.2 should be the flowering time-related gene TaFT3-D1, which is a homolog of the barley gene HvFT3 (Faure et al. 2007; Halliwell et al. 2016). QHD.caas-1BL, QHD.caas-3AL.1, QHD.caas-5AL, QHD.caas-7BL and QHD.caas-7DL are at similar positions to those in Zhang et al. (2009), Edae et al. (2014), Reif et al. (2011), Bordes et al. (2013) and Cuthbert et al. (2008). QHD.caas-6BL identified in the D × S and L × Z populations is probably a new QTL.
Rht4 and Rht12 were identified as induced semi-dwarf mutants in wheat and mapped on chromosomes 2BL and 5AL, respectively (Konzak 1988; Ellis et al. 2005). Blasting results indicated that QPH.caas-2BL.1 is about 7 Mb from marker Xwmc317 linked to Rht4 on the physical map (IWGSC RefSeq v1.0, https://urgi.versailles.inra.fr/blast_iwgsc/blast.php). Nevertheless, no sufficient evidence can prove that they are the same. Two QTL for PH on chromosome 5AL are different from Rht12, whereas QPH.caas-5AL.1 is at a similar position to the QTL detected by Zhang et al. (2017b). QPH.caas-1BL, QPH.caas-2BL.2, QPH.caas-6BL and QPH.caas-6DS are likely to be new.
UIL plays an important role in determining PH and GY, but relatively little attention was given to this trait in previous studies. Our results indicated that Rht-B1 and Rht-D1 affect UIL based on the co-localization of PH and UIL QTL at two loci. QUIL.caas-2AS, identified in the D × S and L × Z populations, was located at the same position as a gene Rht_NM9 that was finely mapped within a region of 8.86 Mb on chromosome 2AS (Lu et al. 2015), and close to the QTL in Yu et al. (2014). Zhang et al. (2017b) reported a QTL associated with PH and related traits close to QUIL.caas-3DL. The major QTL for UIL on chromosome 6BL, with no correlation with PH, is likely to be new.
Genetic studies on flag leaf-related traits (FLL and FLW) in wheat have always been a big challenge due to instability across environments. In rice, two genes (Nal1 and Nal7) for FLW have been cloned, and two QTL (qFL1 and qFLW4) associated with FLL and FLW, respectively, were finely mapped (Fujino et al. 2008; Qi et al. 2008; Wang et al. 2011b; Chen et al. 2012). However, to date, no genes for FLL and FLW have been cloned in wheat, and one gene (TaFLW1) next to Fhb5 on chromosome 5A was finely mapped (Xue et al. 2013). QFLL.caas-2BL, previously identified by Wu et al. (2016), was close to a QTL reported in Edae et al. (2014). QFLL.caas-2AL and QFLL.caas-3B detected in two or three environments and BLUE values are probably new.
Several previous studies indicated that chromosome 5A is important in determining FLW in wheat (Jia et al. 2013; Xue et al. 2013; Wu et al. 2016). Two QTL on chromosome 5AL were detected in the present study, and QFLW.caas-5AL.2 is at the same position as that in Wu et al. (2016). QFLW.caas-1BL and QFLW.caas-6BL are also at the same positions with those in Wu et al. (2016). QFLW.caas-2BS, QFLW.caas-4AL.2, QFLW.caas-5AL.1, QFLW.caas-5BL, QFLW.caas-5DL and QFLW.caas-6AS are likely to be new QTL.
Genetic relationships among grain yield-related traits
GY is a complex quantitative trait controlled by multiple genes and highly influenced by environment. As expected, yield-related traits are significantly correlated. In the present study, 11 QTL clusters or QTL-rich regions on chromosomes 1BL, 2AS, 3DL, 4BS, 4DS, 5AL (2), 5DL, 6AL (2) and 6BL were identified. The pleiotropic loci on chromosomes 4BS and 5DL were associated with 11 and 9 traits, respectively, showing crucial roles in affecting GY. The interval 15.5–32.3 cM around Rht-B1 locus on chromosome 4BS, harboring 20 QTL for all traits except KL, is a pleiotropic locus. This represented a strong validation that Rht-B1 has a significant influence on yield-related traits. Several studies have reported this QTL-rich region for yield (McCartney et al. 2005; Cuthbert et al. 2008; Zhang et al. 2017b) and physiological-related traits (Gao et al. 2015), and our study revealed more detail on the effect of this locus. Another QTL-rich region is around the Rht-D1 locus on chromosome 4DS, including QTL for TKW, KL, KW, PH and UIL. The large effect of Rht-D1 on TKW is consistent with the significantly positive correlation between TKW and PH-related traits. The QTL-rich region around this locus for leaf area (Wu et al. 2016) and physiological-related traits (Gao et al. 2015) were also identified previously.
Among the 11 QTL clusters, 10 were associated with kernel-related traits, indicating that the variation of kernel-related traits is easily caused by the altered agronomic traits. Six QTL for KW were co-localized with those for TKW, whereas only two for KL were co-localized with the TKW QTL, which was reflected by the higher correlations between KW and TKW (r = 0.71–0.82). Three QTL for SN detected among the pleiotropic regions were co-localized with TKW and KW QTL. As SN was significantly and negatively correlated with TKW in the three populations, the effects of these pleiotropic regions on SN and TKW are likely to be contrasting. In addition, QTL for SN and KNS on chromosomes 4BS and 5DL identified in the D × S population were co-localized, in agreement with the significant and negative correlation between two traits (r = − 0.66). Three KNS QTL were co-localized with PH or UIL QTL, and three with kernel-related traits. Due to the negative correlations, increased PH, UIL and kernel-related traits may have adverse effects on KNS. Moreover, four QTL for SDW on chromosomes 1BS, 4BS, 5DL and 7AS and four for FLW on chromosomes 1BL, 4AL, 4BS and 5DL were co-localized with the QTL for KNS, in agreement with the significant correlations among these traits. The HD QTL were co-localized with those for TKW, SL, SDW and PH, respectively. Based on the correlations between HD and each trait, it was considered that taller plants are likely to be heading earlier, whereas the earlier heading date is favorable for spike development and grain-filling. Le Gouis et al. (2012) identified two HD loci close to Rht-B1 and Rht-D1, but the conclusion on effects of two genes for HD remains controversial.
Molecular mechanisms underlying yield-related traits
With availability of the CSS database (IWGSC RefSeq v1.0, https://urgi.versailles.inra.fr/blast_iwgsc/blast.php) QTL mapped with high-density SNPs can be in comparison with cloned genes. Compared to SSR markers, the 90 K iSelect SNP assay was developed from the transcriptome (Wang et al. 2014), providing a better approach in searching for candidate genes. In the present study, 11 candidate genes were identified, including eight cloned genes for kernel traits, HD and PH as well as three predicted genes for TKW, SL and HD (Table S14).
GA-insensitive plants carrying the Rht-B1b, Rht-B1e and Rht-D1b alleles were characterized by reduced height, because of decreased sensitivity of their vegetative tissues to endogenous gibberellin (GA) (Keyes et al. 1989). This effect is manifested by decreased cell length in almost all vegetative organs, followed not only by reduced plant height, but also by decreased coleoptile, internode lengths, and leaf area. Chebotar et al. (2016) reported that the GA-sensitive (Rht8) and GA-insensitive (Rht-B1 and Rht-D1) dwarfing genes had pleiotropic effects on all traits studied except for the number of fertile spikelets. In the present study, Rht-B1 and Rht-D1 as well as several QTL for PH were identified. As expected, Rht-B1 and Rht-D1 showed large effects on PH and yield-related traits, consistent with the previous reports (Peng et al. 1999; Ellis et al. 2005; Gao et al. 2015; Sun et al. 2017). Moreover, other QTL for PH were always co-localized with the QTL for yield traits, especially for kernel-related traits, indicating similar but lesser effects compared to Rht-B1 and Rht-D1.
Kernel-related genes are involved in the regulation of cell division and expansion, presumably via three major pathways, including ubiquitination-mediated proteasomal degradation, G-protein signaling, and phytohormones, and other unknown pathways (Huang et al. 2009; Li et al. 2011; Ma et al. 2016; Zhang et al. 2017c). Three QTL for kernel-related traits in the present study were associated with cloned genes TaGS5 (3BS), TaTPP-6AL1 (6AL) and TaGS1a (6DL) (Guo et al. 2013; Ma et al. 2016; Zhang et al. 2017c), respectively. Moreover, the co-localized QTL (QTKW.caas-5AL.3 and QKW.caas-5AL) putatively correspond to the isoamylase 3 gene EMS47868.1 involved in biosynthesis of amylopectin in wheat and rice. The function of this gene is similar to Trehalose 6-phosphate phosphatase (TPP), which is associated with starch accumulation and grain yield in several crops (Zhang et al. 2017c).
The effect of Ppd and Vrn genes on HD or flowering time is relatively well understood; however, much less is known about the influence of genetic factors that modify flowering time once photoperiod and vernalization requirements have been met. Three QTL for HD in the present study are associated with cloned genes Vrn-D1, TaFT3-D1 and TaELF3-1DL (Fu et al. 2005; Faure et al. 2007; Halliwell et al. 2016; Wang et al. 2016a), respectively. Another QTL for HD on chromosome 4BS corresponds to Tubulin/FtsZ family protein gene NP_849388.1 involved in plant growth rhythm. In addition, a putative candidate gene corresponding to the bidirectional sugar transporter SWEET6b (EMS56832.1) is associated with SL on chromosome 2DS. The molecular mechanism for many yield-related traits is still not well understood, and more experimental analysis should be undertaken to confirm the roles of these genes on yield-related traits.
Potential implications in wheat breeding
Wheat GY is significantly influenced by environment, which creates major challenges in selection of high-yielding lines in small plots at the early stages of breeding programs. In contrast, GY components, as well as kernel, spike, PH and flag leaf-related traits are less influenced by environment. Consequently, more attention has focused on yield-related traits so as to improve GY. It has been proven previously, and in the present study, that TKW is significantly affected by KW (Huang et al. 2015; Wu et al. 2015). Considering the problems of kernel color preference and possible existence of black point, TKW can be selected directly in wheat breeding.
The present study showed that KNS was significantly and positively correlated with SDW and FLW in wheat. This was supported by the co-localization of QTL for these three traits. Due to its simplicity FLW can be used to select lines with large KNS. More physiological traits should be considered in future studies to dissect factors associated with SN.
The presence of common QTL identified in two or three populations reveals that the QTL have consistently stable effects in different genetic backgrounds. Therefore, the common QTL listed in Table 3 might be valuable to be employed in improving GY. In addition, stable QTL with large contributions identified in a single population, such as QTKW.caas-5DL, QKL.caas-5BS, QKL.caas-5DL, QKL.caas-6DL, QKW.caas-3B.2, QSL.caas-4AL, QPH.caas-5AL.1 and QFLW.caas-6AS, are also useful for MAS in breeding varieties adapted to specific environments. Moreover, flanking SNP markers of pleiotropic loci or QTL-rich regions for yield-related traits are suitable to select varieties with good performance. Our study proved that QTL mapping using three RIL populations together with high-density linkage maps offered an opportunity to identify common and stable QTL for MAS.
Kompetitive allele-specific PCR (KASP) is a homogenous, fluorescence-based genotyping variation of PCR (Rasheed et al. 2016). Currently, KASP assays are widely used in genotyping important traits due to high-throughput and low cost. Semi-thermal asymmetric reverse PCR (STARP) is a newly developed genotyping method adapting to multiple platforms and throughputs (Long et al. 2017). The development of these two assays offers better choices in detecting InDels or SNPs. SNP markers tightly linked with yield-related traits detected by QTL mapping or GWAS can be successfully transformed into KASP or STARP assays for MAS in wheat breeding (Rasheed et al. 2016; Liu et al. 2017). With the use of high-density linkage maps in gene mining and high-throughput platforms in genotyping, wheat breeding by molecular design is not a distant goal.
Abbreviations
- BLUE:
-
Best linear unbiased estimation
- FLL:
-
Flag leaf length
- FLW:
-
Flag leaf width
- GY:
-
Grain yield
- GWAS:
-
Genome-wide association study
- h 2 :
-
Broad-sense heritability
- HD:
-
Heading date
- KASP:
-
Kompetitive allele-specific PCR
- KL:
-
Kernel length
- KNS:
-
Kernel number per spike
- KW:
-
Kernel width
- LOD:
-
Logarithm of odds
- MAS:
-
Marker-assisted selection
- PH:
-
Plant height
- QTL:
-
Quantitative trait loci
- R 2 :
-
Phenotypic variance explained
- RIL:
-
Recombinant inbred line
- SDW:
-
Spike dry weight
- SL:
-
Spike length
- SN:
-
Spike number per unit area
- SNP:
-
Single nucleotide polymorphism
- STARP:
-
Semi-thermal asymmetric reverse PCR
- TKW:
-
Thousand-kernel weight
- UIL:
-
Uppermost internode length
References
Austin RB, Bingham J, Blackwell RD, Evans LT, Ford MA, Morgan CL, Taylor M (1980) Genetic improvement in winter wheat yields since 1900 and associated physiological changes. J Agric Sci (Camb) 94:675–689
Azadi A, Mardi M, Hervan EM, Mohammadi SA, Moradi F, Tabatabaee MT, Pirseyedi SM, Ebrahimi M, Fayaz F, Kazemi M, Ashkani S, Nakhoda B, Mohammadi-Nejad G (2015) QTL mapping of yield and yield components under normal and salt-stress conditions in bread wheat (Triticum aestivum L.). Plant Mol Biol Rep 33:102–120
Beales J, Turner A, Griffiths S, Snape JW, Laurie DA (2007) A pseudo-response regulator is misexpressed in the photoperiod insensitive Ppd-D1a mutant of wheat (Triticum aestivum L.). Theor Appl Genet 115:721–733
Bordes J, Goudemand E, Duchalais L, Chevarin L, Oury FX, Heumez E, Lapierre A, Perretant MR, Rolland B, Beghin D, Laurent V, Le Gouis J, Storlie E, Robert O, Charmet G (2013) Genome-wide association mapping of three important traits using bread wheat elite breeding populations. Mol Breed 33:755–768
Brancourt-Hulmel Doussinault MG, Leconte C, Berard P, Le Buanec B, Trottet M (2003) Genetic improvement of agronomic traits of winter wheat cultivars released in France from 1946 to 1992. Crop Sci 43:37–45
Chang JZ, Zhang JA, Mao XG, Li A, Jia JZ, Jing RL (2013) Polymorphism of TaSAP1-A1 and its association with agronomic traits in wheat. Planta 237:1495–1508
Chang JZ, Hao CY, Chang XP, Zhang XY, Jing RL (2014) HapIII of TaSAP1-A1, a positively selected haplotype in wheat breeding. J Integr Agric 13:1462–1468
Chebotar GA, Chebotar SV, Motsnyy II (2016) Pleiotropic effects of gibberellin-sensitive and gibberellin-insensitive dwarfing genes in bread wheat of the southern step region of the Black Sea. Cytol Genet 50:20–27
Chen ML, Luo J, Shao GN, Wei XJ, Tang SQ, Sheng ZH, Song J, Hu PS (2012) Fine mapping of a major QTL for flag leaf width in rice, qFLW4, which might be caused by alternative splicing of NAL1. Plant Cell Rep 31:863–872
Chen SL, Gao RH, Wang HY, Wen MX, Xiao J, Bian NF, Zhang RQ, Hu WJ, Cheng SH, Bie TD, Wang XE (2015) Characterization of a novel reduced height gene (Rht23) regulating panicle morphology and plant architecture in bread wheat. Euphytica 203:583–594
Cheng RR, Kong ZX, Zhang LW, Xie Q, Jia HY, Yu D, Huang YL, Ma ZQ (2017) Mapping QTLs controlling kernel dimensions in a wheat inter-varietal RIL mapping population. Theor Appl Genet 130:1405–1414
Cook JP, McMullen MD, Holland JB, Tian F, Bradbury P, Ross-Ibarra J, Buckler ES, Flint-Garcia SA (2012) Genetic architecture of maize kernel composition in the nested association mapping and inbred association panels. Plant Physiol 158:824–834
Cui F, Zhao CH, Ding AM, Li J, Wang L, Li XF, Bao YG, Li JM, Wang HG (2014) Construction of an integrative linkage map and QTL mapping of grain yield-related traits using three related wheat RIL populations. Theor Appl Genet 127:659–675
Cuthbert JL, Somers DJ, Brûlé-Babel AL, Brown PD, Crow GH (2008) Molecular mapping of quantitative trait loci for yield and yield components in spring wheat (Triticum aestivum L.). Theor Appl Genet 117:595–608
Darvasi A, Soller M (1997) A simple method to calculate resolving power and confidence interval of QTL map location. Behav Genet 27:125–132
Deng SM, Wu XR, Wu YY, Zhou RH, Wang HG, Jia JZ, Liu SB (2011) Characterization and precise mapping of a QTL increasing spike number with pleiotropic effects in wheat. Theor Appl Genet 122:281–289
Dong LL, Wang FM, Liu T, Dong ZY, Li AL, Jing RL, Mao L, Li YW, Liu X, Zhang KP, Wang DW (2014) Natural variation of TaGASR7-A1 affects grain length in common wheat under multiple cultivation conditions. Mol Breed 34:937–947
Donmez E, Sears RG, Shroyer JP, Paulsen GM (2001) Genetic gain in yield attributes of winter wheat in the Great Plains. Crop Sci 41:1412–1419
Edae EA, Byrne PF, Haley SD, Lopes MS, Reynolds MP (2014) Genome-wide association mapping of yield and yield components of spring wheat under contrasting moisture regimes. Theor Appl Genet 127:791–807
Ellis MH, Rebetzke GJ, Azanza F, Richards RA, Spielmeyer W (2005) Molecular mapping of gibberellin-responsive dwarfing genes in bread wheat. Theor Appl Genet 111:423–430
Faure S, Higgins J, Turner A, Laurie DA (2007) The FLOWERING LOCUS T-like gene family in barley (Hordeum vulgare). Genetics 176:599–609
Fu DL, Szucs P, Yan LL, Helguera M, Skinner JS, Zitzewitz J, Hayes PM, Dubcovsky J (2005) Large deletions within the first intron in VRN-1 are associated with spring growth habit in barley and wheat. Mol Gen Genomics 273:54–65
Fujino K, Matsuda Y, Ozawa K, Nishimura T, Koshiba T, Fraaije MW, Sekiguchi H (2008) NARROW LEAF 7 controls leaf shape mediated by auxin in rice. Mol Genet Genomics 279:499–507
Ganal MW, Durstewitz G, Polley A, Berard A, Buckler ES, Charcosset A, Clarke JD, Graner EM, Hansen M, Joets J, Le Paslier MC, McMullen MD, Montalent P, Rose M, Schon CC, Sun Q, Walter H, Martin OC, Falque M (2011) A large maize (Zea mays L.) SNP genotyping array: development and germplasm genotyping, and genetic mapping to compare with the B73 reference genome. PLoS ONE 6:e28334
Gao FM, Wen WE, Liu JD, Rasheed A, Yin GH, Xia XC, Wu XX, He ZH (2015) Genome-wide linkage mapping of QTL for yield components, plant height and yield-related physiological traits in the Chinese wheat cross Zhou 8425B/Chinese Spring. Front Plant Sci 6:1099
Gao FM, Ma DY, Yin GH, Rasheed A, Dong Y, Xiao YG, Xia XC, Wu XX, He ZH (2017) Genetic progress in grain yield and physiological traits in Chinese wheat cultivars of Southern Yellow and Huai Valley since 1950. Crop Sci 57:760–773
Gegas VC, Nazari A, Griffiths S, Simmonds J, Fish L, Orford S, Sayers L, Doonan JH, Snape JW (2010) A genetic framework for grain size and shape variation in wheat. Plant Cell 22:1046–1056
Griffiths S, Simmonds J, Leverington M, Wang Y, Fish L, Sayers L, Alibert L, Orford S, Wingen L, Herry L, Faure S, Laurie D, Bilham L, Snape J (2009) Meta-QTL analysis of the genetic control of ear emergence in elite European winter wheat germplasm. Theor Appl Genet 119:383–395
Guo Y, Sun JJ, Zhang GZ, Wang YY, Kong FM, Zhao Y, Li SS (2013) Haplotype, molecular marker and phenotype effects associated with mineral nutrient and grain size traits of TaGS1a in wheat. Field Crops Res 154:119–125
Halliwell J, Borrill P, Gordon A, Kowalczyk R, Pagano ML, Saccomanno B, Bentley AR, Uauy C, Cockram J (2016) Systematic investigation of FLOWERING LOCUS T-like poaceae gene families identifies the short-day expressed flowering pathway gene, TaFT3 in wheat (Triticum aestivum L.). Front. Plant Sci 7:857
Hanif M, Gao FM, Liu JD, Wen WE, Zhang YJ, Rasheed A, Xia XC, He ZH, Cao SH (2015) TaTGW6-A1, an ortholog of rice TGW6, is associated with grain weight and yield in bread wheat. Mol Breed 36:1–8
Hedden P (2003) The genes of the Green Revolution. Trends Genet 19:5–9
Huang XZ, Qian Q, Liu ZB, Sun HY, He SY, Luo D, Xia GM, Chu CC, Li JY (2009) Natural variation at the DEP1 locus enhances grain yield in rice. Nat Genet 41:494–497
Huang L, Wang Q, Zhang LQ, Yuan ZW, Wang JR, Zhang HG, Zheng YL, Liu DC (2011) Haplotype variations of gene Ppd-D1 in Aegilops tauschii and their implications on wheat origin. Genet Resour Crop Evol 59:1027–1032
Huang YL, Kong ZX, Wu XY, Cheng RR, Yu D, Ma ZQ (2015) Characterization of three wheat grain weight QTLs that differentially affect kernel dimensions. Theor Appl Genet 128:2437–2445
Jia HY, Wan HS, Yang SH, Zhang ZZ, Kong ZX, Xue SL, Zhang LX, Ma ZQ (2013) Genetic dissection of yield-related traits in a recombinant inbred line population created using a key breeding parent in China’s wheat breeding. Theor Appl Genet 126:2123–2139
Jiang QY, Hou J, Hao CY, Wang LF, Ge HM, Dong YS, Zhang XY (2011) The wheat (T. aestivum) sucrose synthase 2 gene (TaSus2) active in endosperm development is associated with yield traits. Funct Integr Genomics 11:49–61
Jin H, Wen WE, Liu JD, Zhai SN, Zhang Y, Yan J, Liu ZY, Xia XC, He ZH (2016) Genome-wide QTL mapping for wheat processing quality parameters in a Gaocheng 8901/Zhoumai 16 recombinant inbred line population. Front Plant Sci 7:1032
Keyes GJ, Paolillo DJ, Sorrells ME (1989) The effects of dwarfing genes Rht1 and Rht2 on cellular dimensions and rate of leaf elongation in wheat. Ann Bot 64:683–690
Konzak CF (1988) Genetic analysis, genetic improvement and evaluation of induced semi-dwarf mutants in wheat. Semi-dwarf cereal mutants and their use in cross-breeding III research coordination meeting, 16–20 December 1985. International Atomic Energy Agency, Vienna, Austria 39–50
Le Gouis J, Bordes J, Ravel C, Heumez E, Faure S, Praud S, Galic N, Remoue C, Balfourier F, Allard V, Rousset M (2012) Genome-wide association analysis to identify chromosomal regions determining components of earliness in wheat. Theor Appl Genet 124:597–611
Lee HS, Jung JU, Kang CS, Heo HY, Park CS (2014) Mapping of QTL for yield and its related traits in a doubled haploid population of Korean wheat. Plant Biotechnol Rep 8:443–454
Li HH, Ye GY, Wang JK (2007) A modified algorithm for the improvement of composite interval mapping. Genetics 175:361–374
Li YB, Fan CC, Xing YZ, Jiang YH, Luo LJ, Sun L, Shao D, Xu CJ, Li XH, Xiao JH, He YQ, Zhang QF (2011) Natural variation in GS5 plays an important role in regulating grain size and yield in rice. Nat Genet 43:1266–1269
Li QF, Zhang Y, Liu TT, Wang FF, Liu K, Chen JS, Tian JC (2015) Genetic analysis of kernel weight and kernel size in wheat (Triticum aestivum L.) using unconditional and conditional QTL mapping. Mol Breed 35:194
Liu YN, He ZH, Appels R, Xia XC (2012) Functional markers in wheat: current status and future prospects. Theor Appl Genet 125:1–10
Liu G, Jia LJ, Lu LH, Qin DD, Zhang JP, Guan PF, Ni ZF, Yao YY, Sun QX, Peng HR (2014) Mapping QTLs of yield-related traits using RIL population derived from common wheat and Tibetan semi-wild wheat. Theor Appl Genet 127:2415–2432
Liu JD, He ZH, Rasheed A, Wen WE, Yan J, Zhang PZ, Wan YX, Zhang Y, Xie CJ, Xia XC (2017) Genome-wide association mapping of black point reaction in common wheat (Triticum aestivum L.). BMC Plant Biol 17:220
Long YM, Chao WS, Ma GJ, Xu SS, Qi LL (2017) An innovative SNP genotyping method adapting to multiple platforms and throughputs. Theor Appl Genet 130:597–607
Lorieux M (2012) MapDisto: fast and efficient computation of genetic linkage maps. Mol Breed 30:1231–1235
Lu Y, Xing LP, Xing SJ, Hu P, Cui CF, Zhang MY, Xiao J, Wang HY, Zhang RQ, Wang XE, Chen PD, Cao AZ (2015) Characterization of a putative new semi-dominant reduced height gene, Rht_NM9, in wheat (Triticum aestivum L.). J Genet Genomics 42:685–698
Ma DY, Yan J, He ZH, Wu L, Xia XC (2010) Characterization of a cell wall invertase gene TaCwi-A1 on common wheat chromosome 2A and development of functional markers. Mol Breed 29:43–52
Ma L, Li T, Hao CY, Wang YQ, Chen XH, Zhang XY (2016) TaGS5-3A, a grain size gene selected during wheat improvement for larger kernel and yield. Plant Biotecnol J 14:1269–1280
McCaig TN, DePauw RM (1995) Breeding hard red spring wheat in western Canada: historical trends in yield and related variables. Can J Plant Sci 75:387–393
McCartney CA, Somers DJ, Humphreys DJ, Lukow O (2005) Mapping quantitative trait loci controlling agronomic traits in the spring wheat cross RL 4452 × AC ‘Domain’. Genome 48:870–883
Mir RR, Kumar N, Jaiswal V, Girdharwal N, Prasad M, Balyan HS, Gupta PK (2012) Genetic dissection of grain weight in bread wheat through quantitative trait locus interval and association mapping. Mol Breed 29:963–972
Mohler V, Albrecht T, Castell A, Diethelm M, Schweizer G, Hartl L (2016) Considering causal genes in the genetic dissection of kernel traits in common wheat. J Appl Genet 57:467–476
Muterko A, Kalendar R, Salina E (2016) Allelic variation at the VERNALIZATION-A1, VRN-B1, VRN-B3, and PHOTOPERIOD-A1 genes in cultivars of Triticum durum Desf. Planta 244:1253–1263
Nadolska-Orczyk A, Rajchel IK, Orczyk W, Gasparis S (2017) Major genes determining yield-related traits in wheat and barley. Theor Appl Genet 130:1081–1098
Nishida H, Yoshida T, Kawakami K, Fujita M, Long B, Akashi Y, Laurie DA, Kato K (2013) Structural variation in the 5′ upstream region of photoperiod-insensitive alleles Ppd-A1a and Ppd-B1a identified in hexaploid wheat (Triticum aestivum L.), and their effect on heading time. Mol Breed 31:27–37
Nyquist W, Baker R (1991) Estimation of heritability and prediction of selection response in plant populations. Crit Rev Plant Sci 10:235–322
Ortiz-Monasterio R, Sayre KD, Rajaram S, McMahon M (1997) Genetic progress in wheat yield and nitrogen use efficiency under four nitrogen rates. Crop Sci 37:898–904
Peng J, Richards D-E, Hartley N-M, Murphy GP, Devos KM, Flintham JE, Beales J, Fish LJ, Worland AJ, Pelica F, Sudhakar D, Christou P, Snape JW, Gale MD, Harberd NP (1999) ‘Green Revolution’ genes encode mutant gibberellin response modulators. Nature 400:256–261
Perez-Perez JM, Esteve-Bruna D, Micol JL (2010) QTL analysis of leaf architecture. J Plant Res 123:15–23
Qi J, Qian Q, Bu QY, Li SY, Chen Q, Sun JQ, Liang WX, Zhou YH, Chu CC, Li XG, Ren FG, Palme K, Zhao BR, Chen JF, Chen MS, Li CY (2008) Mutation of the rice narrow leaf1 gene, which encodes a novel protein, affects vein patterning and polar auxin transport. Plant Physiol 147:1947–1959
Qin L, Hao CY, Hou J, Wang YQ, Li T, Wang LF, Ma ZQ, Zhang XY (2014) Homologous haplotypes, expression, genetic effects and geographic distribution of the wheat yield gene TaGW2. BMC Plant Biol 14:107
Ramya P, Chaubal A, Kulkarni K, Gupta L, Kadoo N, Dhaliwal HS, Chhuneja P, Lagu M, Gupta V (2010) QTL mapping of 1000-kernel weight, kernel length, and kernel width in bread wheat (Triticum aestivum L.). J Appl Genet 51:421–429
Rasheed A, Wen WE, Gao FM, Zhai SN, Jin H, Liu JD, Guo Q, Zhang Y, Dreisigacker S, Xia XC, He ZH (2016) Development and validation of KASP assays for genes underpinning key economic traits in bread wheat. Theor Appl Genet 129:1843–1860
Reif JC, Maurer HP, Korzun V, Ebmeyer E, Miedaner T, Würschum T (2011) Mapping QTLs with main and epistatic effects underlying grain yield and heading time in soft winter wheat. Theor Appl Genet 123:283–292
Röder MS, Huang XQ, Rörner A (2008) Fine mapping of the region on wheat chromosome 7D controlling grain weight. Funct Integr Genomics 8:79–86
Sharma SN, Sain RS, Sharma RK (2003) The genetic control of flag leaf length in normal and late sown durum wheat. J Agric Sci 141:323–331
Siddique KHM, Belford RK, Perry MW, Tennant D (1989) Growth, development and light interception of old and modern wheat cultivars in a Mediterranean-type environment. Aust J Agric Res 40:473–487
Song Q, Hyten DL, Jia G, Quigley CV, Fickus EW, Nelson RL, Cregan PB (2013) Development and evaluation of SoySNP50 K, a high-density genotyping array for soybean. PLoS ONE 8:e54985
Sourdille P, Cadalen T, Gay G, Gill B, Bernard M (2002) Molecular and physical mapping of genes affecting awning in wheat. Plant Breed 121:320–324
Stam P (1993) Construction of integrated genetic linkage maps by means of a new computer package: JOINMAP. Plant J 3:739–744
Su Z, Hao C, Wang L, Dong Y, Zhang X (2011) Identification and development of a functional marker of TaGW2 associated with grain weight in bread wheat (Triticum aestivum L.). Theor Appl Genet 122:211–223
Sukumaran S, Dreisigacker S, Lopes M, Chavez P, Reynolds MP (2015) Genome-wide association study for grain yield and related traits in an elite spring wheat population grown in temperate irrigated environments. Theor Appl Genet 128:353–363
Sun CW, Zhang FY, Yan XF, Zhang XF, Dong ZD, Cui DQ, Chen F (2017) Genome-wide association study for 13 agronomic traits reveals distribution of superior alleles in bread wheat from the Yellow and Huai Valley of China. Plant Biotechnol J 15:953–969
Takenaka S, Kawahara T (2013) Evolution of tetraploid wheat based on variations in 5′UTR regions of Ppd-A1: evidence of gene flow between emmer and timopheevi wheat. Genet Resour Crop Evol 60:2143–2155
The International Wheat Genome Sequencing Consortium (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:1251788
Tian XL, Wen WE, Xie L, Fu LP, Xu DA, Fu C, Wang DS, Chen XM, Xia XC, Chen QJ, He ZH, Cao SH (2017) Molecular mapping of reduced plant height gene Rht24 in bread wheat. Front Plant Sci 8:1379
Tung CW, Zhao K, Wright MH, Ali ML, Jung J, Kimball J, Tyagi W, Thomson MJ, McNally K, Leung H, Kim H, Ahn S-N, Reynolds A, Scheffler B, Eizenga G, McClung A, Bustamante C, McCouch SR (2010) Development of a research platform for dissecting phenotype–genotype associations in rice (Oryza spp.). Rice 3:205–217
Tyagi S, Mir RR, Balyan HS, Gupta PK (2014) Interval mapping and meta-QTL analysis of grain traits in common wheat (Triticum aestivum L.). Euphytica 201:367–380
Valluru R, Reynolds MP, Davies WJ, Sukumaran S (2017) Phenotypic and genome-wide association analysis of spike ethylene in diverse wheat genotypes under heat stress. New Phytol 214:271–283
Wang JS, Liu WH, Wang H, Li LH, Wu J, Yang XM, Li XQ, Gao AN (2011a) QTL mapping of yield-related traits in the wheat germplasm 3228. Euphytica 177:277–292
Wang P, Zhou GL, Yu HH, Yu SB (2011b) Fine mapping a major QTL for flag leaf size and yield-related traits in rice. Theor Appl Genet 123:1319–1330
Wang SC, Wong D, Forrest K, Allen A, Chao S, Huang BE, Maccaferri M, Salvi S, Milner SG, Cattivelli L, Mastrangelo AM, Whan A, Stephen S, Barker G, Wieseke R, Plieske J, Lillemo M, Mather D, Appels R, Dolferus R, Guedira GB, Korol A, Akhunova AR, Feuillet C, Salse J, Morgante M, Pozniak C, Luo MC, Dvorak J, Morell M, Dubcovsky J, Ganal M, Tuberosa R, Lawley C, Mikoulitch I, Cavanagh C, Edwards KJ, Hayden M, Akhunov E (2014) Characterization of polyploid wheat genomic diversity using a high-density 90000 single nucleotide polymorphism array. Plant Biotechnol J 12:787–796
Wang SS, Zhang XF, Chen F, Cui DQ (2015) A single-nucleotide polymorphism of TaGS5 gene revealed its association with kernel weight in Chinese bread wheat. Front Plant Sci 6:1166
Wang JP, Wen WE, Hanif M, Xia XC, Wang HG, Liu SB, Liu JD, Yang L, Cao SH, He ZH (2016a) TaELF3-1DL, a homolog of ELF3, is associated with heading date in bread wheat. Mol Breed 36:161
Wang SS, Yan XF, Wang YY, Liu HM, Cui DQ, Chen F (2016b) Haplotypes of the TaGS5-A1 gene are associated with thousand-kernel weight in Chinese bread wheat. Front Plant Sci 7:783
Wen WE, He ZH, Gao FM, Liu JD, Jin H, Zhai SN, Qu YY, Xia XC (2017) A high-density consensus map of common wheat integrating four mapping populations scanned by the 90 K SNP array. Front Plant Sci 8:1389
Wiedmann RT, Smith TPL, Nonneman DJ (2008) SNP discovery in swine by reduced representation and high throughput pyrosequencing. BMC Genet 9:81
Wu QH, Chen YX, Zhou SH, Fu L, Chen JJ, Xiao Y, Zhang D, Ouyang SH, Zhao XJ, Cui Y, Zhang DY, Liang Y, Wang ZZ, Xie JZ, Qin JX, Wang GX, Li DL, Huang YL, Yu MH, Lu P, Wang LL, Wang L, Wang H, Dang C, Li J, Zhang Y, Peng HR, Yuan CG, You MS, Sun QX, Wang JR, Wang LX, Luo MC, Han J, Liu ZY (2015) High-density genetic linkage map construction and QTL mapping of grain shape and size in the wheat population Yanda 1817 × Beinong6. PLoS ONE 10:e0118144
Wu QH, Chen YX, Fu L, Zhou SH, Chen JJ, Zhao XJ, Zhang D, Ouyang SH, Wang ZH, Li D, Wang GX, Zhang DY, Yuan CG, Wang LX, You MS, Han J, Liu ZY (2016) QTL mapping of flag leaf traits in common wheat using an integrated high-density SSR and SNP genetic linkage map. Euphytica 208:337–351
Würschum T, Langer SM, Longin CFH, Tucker MR, Leiser WL (2017) A modern Green Revolution gene for reduced height in wheat. Plant J 92:892–903
Xiao SH, He ZH (2003) Wheat yield and end use quality improvement in China (Chapter 13). In: Zhuang QS (ed) Chinese wheat improvement and pedigree analysis. China Agricultural Publish Press, Beijing
Xiao YG, Qian ZG, Wu K, Liu JJ, Xia XC, Ji WQ, He ZH (2012) Genetic gains in grain yield and physiological traits of winter wheat in Shandong province, China, from 1969 to 2006. Crop Sci 52:44
Xu HY, Zhao JS (1995) Canopy photosynthesis capacity and the contribution from different organs in high-yielding winter wheat. Acta Agron Sin 21:204–209
Xu C, Ren Y, Jian Y, Guo Z, Zhang Y, Xie C, Fu J, Wang H, Wang G, Xu Y, Li P, Zou C (2017a) Development of a maize 55 K SNP array with improved genome coverage for molecular breeding. Mol Breed 37:20
Xu YF, Li SS, Li LH, Ma FF, Fu XY, Shi ZL, Xu HX, Ma PT, An DG (2017b) QTL mapping for yield and photosynthetic related traits under different water regimes in wheat. Mol Breed 37:34
Xue SL, Xu F, Li GQ, Zhou Y, Lin MS, Gao ZX, Su XH, Xu XW, Jiang G, Zhang S, Jia HY, Kong ZX, Zhang LX, Ma ZQ (2013) Fine mapping TaFLW1, a major QTL controlling flag leaf width in bread wheat (Triticum aestivum L.). Theor Appl Genet 126:1941–1949
Yan L, Fu D, Li C, Blechl A, Tranquilli G, Bonafede M, Sanchez A, Valarik M, Yasuda S, Dubcovsky J (2006) The wheat and barley vernalization gene VRN3 is an orthologue of FT. Proc Natl Acad Sci USA 103:19581–19586
Yang ZB, Bai ZY, Li XL, Wang P, Wu QX, Yang L, Li LQ, Li XJ (2012) SNP identification and allelic-specific PCR markers development for TaGW2, a gene linked to wheat kernel weight. Theor Appl Genet 125:1057–1068
Yu M, Mao SL, Chen GY, Pu ZE, Wei YM, Zheng YL (2014) QTLs for uppermost internode and spike length in two wheat RIL populations and their affect upon plant height at an individual QTL level. Euphytica 200:95–108
Zhai SN, He ZH, Wen WE, Jin H, Liu JD, Zhang Y, Liu ZY, Xia XC (2016) Genome-wide linkage mapping of flour color-related traits and polyphenol oxidase activity in common wheat. Theor Appl Genet 129:377–394
Zhai HJ, Feng ZY, Du XF, Song YE, Liu XY, Qi ZQ, Song L, Li J, Li LH, Peng HR, Hu ZR, Yao YY, Xin MM, Xiao SH, Sun QX, Ni ZF (2018) A novel allele of TaGW2-A1 is located in a finely mapped QTL that increases grain weight but decreases grain number in wheat (Triticum aestivum L.). Theor Appl Genet 131:539–553
Zhang KP, Tian JC, Zhao L, Liu B, Chen GF (2009) Detection of quantitative trait loci for heading date based on the doubled haploid progeny of two elite Chinese wheat cultivars. Genetica 135:257–265
Zhang J, Wang YY, Wu SW, Yang JP, Liu HW, Zhou Y (2012a) A single nucleotide polymorphism at the Vrn-D1 promoter region in common wheat is associated with vernalization response. Theor Appl Genet 125:1697–1704
Zhang L, Zhao YL, Gao LF, Zhao GY, Zhou RH, Zhang BS, Jia JZ (2012b) TaCKX6-D1, the ortholog of rice OsCKX2, is associated with grain weight in hexaploid wheat. New Phytol 195:574–584
Zhang XY, Deng ZY, Wang YR, Li JF, Tian JC (2014a) Unconditional and conditional QTL analysis of kernel weight related traits in wheat (Triticum aestivum L.) in multiple genetic backgrounds. Genetica 142:371–379
Zhang YJ, Liu JD, Xia XC, He ZH (2014b) TaGS-D1, an ortholog of rice OsGS3, is associated with grain weight and grain length in common wheat. Mol Breed 34:1097–1107
Zhang HX, Zhang FN, Li GD, Zhang SN, Zhang ZG, Ma LJ (2017a) Genetic diversity and association mapping of agronomic yield traits in eighty six synthetic hexaploid wheat. Euphytica 213:211
Zhang N, Fan XL, Cui F, Zhao CH, Zhang W, Zhao XQ, Yang LJ, Pan RQ, Chen M, Han J, Ji J, Liu DC, Zhao ZW, Tong YP, Zhang AM, Wang T, Li JM (2017b) Characterization of the temporal and spatial expression of wheat (Triticum aestivum L.) plant height at the QTL level and their influence on yield-related traits. Theor Appl Genet 130:1235–1252
Zhang PF, He ZH, Tian XL, Gao FM, Xu DA, Liu JD, Wen WE, Fu LP, Li GY, Sui XX, Xia XC, Wang CP, Cao SH (2017c) Cloning of TaTPP-6AL1 associated with grain weight in bread wheat and development of functional marker. Mol Breed 37:78
Zhao K, Tung C-W, Eizenga GC, Wright MH, Ali ML, Price AH, Norton GJ, Islam MR, Reynolds A, Mezey J, McClung AM, Bustamante CD, McCouch SR (2011) Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat Commun 2:467
Zheng TC, Zhang XK, Yin GH, Wang LN, Han YL, Chen L, Huang F, Tang JW, Xia XC, He ZH (2011) Genetic gains in grain yield, net photosynthesis and stomatal conductance achieved in Henan Province of China between 1981 and 2008. Field Crops Res 122:225–233
Zheng J, Liu H, Wang YQ, Wang LF, Chang XP, Jing RL, Hao CY, Zhang XY (2014) TEF-7A, a transcript elongation factor gene, influences yield-related traits in bread wheat (Triticum aestivum L.). J Exp Bot 65:5351–5365
Zhou Y, Zhu HZ, Cai SB, He ZH, Zhang XK, Xia XC, Zhang GS (2007) Genetic improvement of grain yield and associated traits in the southern China winter wheat region: 1949 to 2000. Euphytica 157:465–473
Zikhali M, Leverington-Waite M, Fish L, Simmonds J, Orford S, Wingen LU, Goram R, Gosman N, Bentley A, Griffiths S (2014) Validation of a 1DL earliness per se (eps) flowering QTL in bread wheat (Triticum aestivum). Mol Breed 34:1023–1033
Acknowledgements
The authors are grateful to Prof. R. A. McIntosh, Plant Breeding Institute, University of Sydney, for critical review of this manuscript. This work was funded by the National Natural Science Foundation of China (31461143021), National Key Research and Development Programs of China (2016YFD0101802, 2016YFD0100502, 2016YFE0108600), and CAAS Science and Technology Innovation Program.
Author information
Authors and Affiliations
Contributions
FL performed the experiment and wrote the paper; WW participated in the field trials and constructed the linkage maps; JL, HJ, JY, PZ and YW participated in the field trials; SC and HG assisted in writing the paper; ZH and XX designed the experiment and wrote the paper. All authors read the final version of the manuscript and approved for publication.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interests.
Ethical standards
We declare that these experiments complied with the ethical standards in China.
Additional information
Communicated by Susanne Dreisigacker.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, F., Wen, W., He, Z. et al. Genome-wide linkage mapping of yield-related traits in three Chinese bread wheat populations using high-density SNP markers. Theor Appl Genet 131, 1903–1924 (2018). https://doi.org/10.1007/s00122-018-3122-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-018-3122-6