Abstract
The modification of flowering date is considered an important way to escape the current or future climatic constraints that affect wheat crops. A better understanding of its genetic bases would enable a more efficient and rapid modification through breeding. The objective of this study was to identify chromosomal regions associated with earliness in wheat. A 227-wheat core collection chosen to be highly contrasted for earliness was characterized for heading date. Experiments were conducted in controlled conditions and in the field for 3 years to break down earliness in the component traits: photoperiod sensitivity, vernalization requirement and narrow-sense earliness. Whole-genome association mapping was carried out using 760 molecular markers and taking into account the five ancestral group structure. We identified 62 markers individually associated to earliness components corresponding to 33 chromosomal regions. In addition, we identified 15 other significant markers and seven more regions by testing marker pair interactions. Co-localizations were observed with the Ppd-1, Vrn-1 and Rht-1 candidate genes. Using an independent set of lines to validate the model built for heading date, we were able to explain 34% of the variation using the structure and the significant markers. Results were compared with already published data using bi-parental populations giving an insight into the genetic architecture of flowering time in wheat.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Earliness, defined as the time needed to reach a specified development stage, plays a major role in plant adaptation to the environment. Its selection is considered to have been one of the main explanations for the gradual extension of wheat cultivation (Stelmakh 1990; Law and Worland 1997). In the context of global climate changes associated with increased drought and heat stresses, strategies to escape these constraints based on earliness adaptation are often advocated (e.g., Debaeke 2004; Gate et al. 2008). This predominant role of earliness in adaptation was highlighted using wheat experimental populations, where heading time was the fastest changing trait in response to local climatic conditions (Goldringer et al. 2006). The identification of the most determinant chromosomal regions would then be of interest to better control flowering time (Jung and Müller 2009) which is a key factor in wheat adaptation.
The development toward the reproductive stage is a complex phenomenon that implies different processes. Earliness for flowering time is generally decomposed into three processes: vernalization requirement, photoperiodic sensitivity and narrow-sense earliness (also named intrinsic earliness and earliness per se). Vernalization induces the shoot apical meristem to switch from a vegetative phase where it produces leaf primordia to a reproductive phase where it produces floral primordia. Optimal vernalization temperatures lie in the 5–10°C range, but higher and lower nonfreezing temperatures still have an effect (Brooking and Jamieson 2002). Provided that the vernalization requirement is satisfied, long days promote the switch from the vegetative to reproductive phase (Trevaskis 2010). However, vernalization and photoperiod interact in a complex way. The vernalization rate is still high above 10°C for short days, while it decreases rapidly for long days (Brooking and Jamieson 2002).
It is well known that cereals are characterized by a strong genetic variability for both vernalization requirement and photoperiod sensitivity. Winter-type cereals require an extended vernalization period, while spring-type cereals can reach anthesis after no or only a short exposure to vernalizing temperatures. Flowering in photoperiod-insensitive wheat genotypes is only marginally delayed in short days (10 h or less light) compared to long days (14 h or more light), while flowering of sensitive types is massively delayed and may not even occur (Beales et al. 2007). Genetic variability for narrow-sense earliness was also observed (Worland 1996). This refers to the remaining earliness variability when vernalization requirements and photoperiodic sensitivity are satisfied.
Our understanding of wheat flowering time determinism has largely improved in the past few years. Early genetic studies focused first on single chromosomes (e.g., Galiba et al. 1995; Worland 1996, Kato et al. 1998) and mainly concerned chromosome groups 2 and 5 that were known to be highly determinant. Then, thanks to the development of dense comprehensive genetic maps, approaches at the whole-genome level have been developed (e.g., Sourdille et al. 2000, 2003; Börner et al. 2002; Hanocq et al. 2003, 2004; Kuchel et al. 2006; Baga et al. 2009; Griffiths et al. 2009). Recently, some of the major genes involved in determining flowering time have been identified and characterized.
Photoperiod requirement for flowering involves mainly the Ppd-1 genes on group 2 chromosomes (Welsh et al. 1973; Law et al. 1978). Turner et al. (2005) cloned the barley Ppd-H1 gene using the colinearity with rice and Brachypodium. The gene was shown to be a member of the pseudo-response regulator (PRR) family most similar to Arabidopsis PRR7 (Turner et al. 2005). Beales et al. (2007) then cloned the wheat orthologs using the sequence of the Ppd-H1 gene. A 2-kb deletion upstream from the coding region was associated with dominant photoperiod insensitivity at the Ppd-D1 locus. This deletion caused a misexpression of Ppd-D1 with the peak of expression shifted from the light to the dark period associated with the induction of the FT gene (now called Vrn-3 in wheat). No candidate polymorphism was identified for the other homoeologous copies.
Vernalization requirement is determined by different genetic factors localized on several chromosomes. The Vrn-1 genes localized on chromosomes of group 5 are long known to be major determinants of the growth habit (Halloran 1967; Law et al. 1976; Maystrenko 1980). Yan et al. (2003) determined that in both diploid and hexaploid wheats, Vrn-1 was homologous to the Arabidopsis meristem identity gene Apetala 1. Insertions or deletions in the regulatory regions of the three homoeologous copies of bread wheat are associated with dominant homoeoalleles for the spring growth habit (Yan et al. 2004a; Fu et al. 2005; Santra et al. 2009). The Vrn-A2 locus was cloned using a positional cloning approach in T. monoccocum (Yan et al. 2004b). It acts as a dominant repressor of flowering down-regulated by vernalization. Distelfeld et al. (2009a) recently characterized the A and B loci in tetraploid wheat. The Vrn-2 locus includes two ZCCT genes (putative zinc finger and CCT domain) in tandem that code for highly similar proteins (Yan et al. 2004b). The Vrn-A2 was shown to be located on chromosome 5AL in a region translocated from the distal part of chromosome 4AL (Dubcovsky et al. 1998). Other homoeologous copies of the Vrn-2 locus are located on the 4B and 4D hexaploid wheat chromosomes. The Vrn-B3 gene was shown to be orthologous to the Arabidopsis FT gene and located on chromosome 7B (Yan et al. 2006). The dominant allele promotes the transcription of Vrn-1 and accelerates flowering. Bonnin et al. (2008) later reported that single nucleotide polymorphism (SNP) or insertions–deletions on A and D copies could also explain differences in heading date in a wheat core collection. Finally, the Vrn-D4 gene was also located in the centromeric region of chromosome 5D (Kato et al. 1993, 2003; Yoshida et al. 2010), but its function is not known yet. Interactions between these major genes are now better understood and models of the regulation of flowering have been proposed (Trevaskis et al. 2007; Distelfeld et al. 2009b; Trevaskis 2010).
Following a study conducted on maize (Chardon et al. 2004), a method called meta-analysis was used to integrate and summarize results from separate wheat studies conducted on flowering time (Hanocq et al. 2007). Combining 13 different detection studies, 84 QTL were projected on a reference map leading to the identification of 18 meta-QTL on chromosomes 2B, 2D, 4A, 4B, 5A, 5B, 5D, 6A, 7A, 7B and 7D. More recently, Griffiths et al. (2009) using four wheat populations identified 19 meta-QTL, some of them appearing unique to their survey. Baga et al. (2009) identified 11 genomic regions associated with floral transition, length of vegetative growth and flowering time. In all the studies, regions containing the major Ppd and Vrn genes are usually detected, but several other regions were also identified showing that several genetic factors were not yet determined. This is reinforced by the recent report involving a large complex population composed of 5,000 recombinant maize (Zea mays L.) inbred lines from 25 crosses, which showed that about 50–60 QTL regions affected flowering time in maize (Buckler et al. 2009).
High-throughput genotyping technologies and statistical methods based on linkage disequilibrium (LD) that take population structure into account (Kang et al. 2008) enabled association mapping studies to be developed for complex traits analysis. In wheat, genetic association studies have already proved successful to study complex traits at the scale of candidate genes using SNPs on worldwide genetic resources (Ravel et al. 2006; Bonnin et al. 2008; Charmet et al. 2009), and at the chromosome scale using SSR on modern cultivars (Breseghello and Sorrells 2006; Zheng et al. 2009). At the whole genome scale, DArTs were used to study associations with yield, disease resistance (Crossa et al. 2007) and bread quality (Bordes et al. 2011).
The objective of this study was to identify chromosomal regions associated with development using whole-genome association mapping on a specific 227-wheat core collection. Memberships of lines in this panel have previously been allocated to five ancestral groups using SSR markers (Rousset et al. 2011). Briefly, groups correspond to Central Asia, North-West Europe, worldwide CIMMYT bread lines, Eastern Asia and Eastern Europe. Then, experiments in controlled conditions and in the field for 3 years (2004, 2005 and 2006) were conducted to dissect earliness into its components: photoperiod sensitivity, vernalization requirement and narrow-sense earliness.
Materials and methods
The population, genotyping and structure
A sample of 227 bread wheat accessions was used for the present study. This sample is part of the INRA bread wheat core collection of 372 accessions (372CC) set up by Balfourier et al. (2007). To fit available phenotyping capacity while best conserving variability, the accessions were sub-sampled using passport data and previous field evaluation data to cover an expected wide range of variation for earliness and growth habit. This sub-sample of 227 accessions, all common to the sub-sample of 235 described by Rousset et al. (2011) captured more than 88% of the alleles observed at the 38 SSR loci among the 4,000 accessions used by Balfourier et al. (2007). Finally, this sub-sample includes landraces and cultivars from 56 different countries.
Genotyping available on the 372CC core collection was used for this study. Briefly, 578 DArT markers were generated by Triticarte Pty. Ltd. (Canberra, Australia; http://www.triticarte.com.au) as previously described (Wenzl et al. 2004; Akbari et al. 2006). All the accessions were also genotyped with 171 SSR markers assigned to the 21 chromosomes (Balfourier et al. 2007) and particularly to the 3B chromosome (Horvath et al. 2009). Based on data obtained from two previous genotyping studies (Balfourier et al. 2006; Bérard et al. 2009), 89 SNP markers within 44 genes were also genotyped. None of these genes were a priori related to flowering time, but were used here as polymorphic markers. For association mapping, we considered rare alleles (<5%) as missing data and then markers with too many missing values (>25%) were removed from the analysis. This resulted in 760 markers tested in the association as 572 DArT, 114 SSR and 74 SNP.
A consensus map using published data was built with the MetaQTL software (Veyrieras et al. 2007) and the map published by Somers et al. (2004) was used as the reference map (following Hanocq et al. (2007) chromosome 4B was reversed). Where markers had not been genetically mapped, they were placed close to the DArT markers with which the LD was the highest. Major growth habit genes (Ppd-1, Vrn-1, Vrn-2, Vrn-3, Vrn-4, Rht-1) were placed on the map using published and communicated data (Table S1). Linkage disequilibrium was calculated with the Tassel v2.1 software (Bradbury et al. 2007), which computes R 2 as proposed by Weir (1996) when only two alleles are observed, and R 2 for each allele pair and then the weighted mean when more than two alleles are present.
We used the structure of five ancestral groups proposed by Rousset et al. (2011) with 82 SSR markers. The first group gathered all Nepalian accessions together with accessions from Afghanistan, Tadjikistan and some from India and Pakistan. The second group was formed with the major part of accessions from North-West Europe. A third group included accessions possibly related to the worldwide CIMMYT breeding program based on common genetic resources and scattered in very diverse regions such as Mexico, Australia, Israel, South America and Africa. Most of the Japanese and Chinese accessions formed a fourth group with some Mediterranean accessions from Spain and Italy. A majority of European, East European and South-East European accessions together with Australian, Canadian and North American accessions were clustered in a fifth group also including some accessions from the Middle East. Trait values weighted by the contribution of each genotype to each group were used to compute group means.
A locus-by-locus analysis of variance was carried out using a fixed linear model written in R (R Development Core Team 2009) to identify DArT markers associated with the contribution to groups. To take into account multiple comparisons, significance was tested using a 0.05 false discovery rate as proposed by Storey and Tibshirani (2003) and implemented in the q value software.
Phenotyping
The measurements of phenotypic traits related to earliness assessed in field and nursery experiments at Le Moulon, France (48.4°N, 02.1°E) in 3 consecutive years (2004, 2005 and 2006) have been described in Rousset et al. (2011). Greenhouse experiments were conducted at Mons-Péronne, France (49.1°N, 03.3°E). Briefly, in the field experiment, the 227 genotypes were sown at the end of October in 2004 and the beginning of November in 2005 and 2006, in a two-replicate complete block design, where they received natural full vernalization during winter. Average heading dates were recorded for each accession and denoted as HdField. In the nursery experiment, the genotypes were spring planted in April (long days) after 0, 4 or 8 weeks in controlled conditions under cold temperature (4°C). Heading dates recorded after no, partial or full vernalization treatments were denoted HdNV, HdV4 and HdV8, respectively. In 2007, heading dates of vernalized plants (8 weeks at 4°C) were also measured in a photoperiodic glasshouse under long day (20 h of light per day) and short day conditions (10 h of light per day) and were denoted, respectively, as HdLD and HdSD. Temperature conditions were 18°C during the day and 14°C at night. For all experiments, the heading date of each plant was recorded when the first ear (main stem) was half out of the flag leaf sheath. Heading dates were transformed into thermal time, calculating temperature sums in degree-days (°Cd) from germination to heading (0°C basis).
Following Rousset et al. (2011), the measured traits and some calculated combinations of these traits were considered to be good proxies of earliness components. Narrow-sense earliness (NSE) was estimated using HdV8, HdLD and HdField. These three traits provide an estimate of wheat earliness when vernalization requirements and photoperiodic sensitivity are satisfied. Vernalization requirement (VR) was assessed using three different traits: HdNV, HdV4 and partial vernalization sensitivity (PVS) with PVS calculated as the quantitative response to extended exposition to cold temperature as PVS = HdV4 − HdV8 (Goldringer et al. 2006; Rousset et al. 2011).
As shown in Rousset et al. (2011), HdNV exhibited a bimodal distribution with accessions heading for the unvernalized May planting with less than 1,450°Cd classified in the spring subpopulation and the accessions heading with more than 1,450°Cd or not heading classified in the winter subpopulation. As observations were stopped while some accessions were still in a vegetative stage, an arbitrary HdNV value of 2,000°Cd (in 2004 and 2005) and 2,100°Cd (in 2006) was attributed to genotypes that did not head before the end of the experiment.
Photoperiod sensitivity (PS) was estimated using HdSD from the glasshouse experiment and day length sensitivity (DLS) calculated from the field experiment as DLS = HdField − HdV8, where HdField is supposed to express both PS and NSE and HdV8 only NSE. Moreover, the quantitative nature of photoperiodic sensitivity was also assessed using two other calculated variables: rapSDLD = HdSD/HdLD and diffSDLD = HdSD − HdLD.
Association
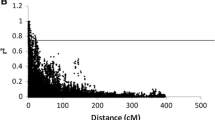
The association between phenotype and single markers was tested with a fixed linear model implemented in the Tassel v2.1 software (Bradbury et al. 2007). Population structure was incorporated into the model by using the covariates indicating the relative contribution of each genotype to each ancestral group. P values adjusted for multiple comparisons using a test derived from a step-down MinP procedure (Ge et al. 2003) of 0.05 or less were considered. Associations between phenotype and pairwise marker interactions were also tested using a fixed linear model written in R (R Development Core Team 2009). Again, population structure was incorporated into the model using covariates. First, pairwise interactions were tested between markers that were identified as being significant using the Tassel software. Then, all combinations of markers with more than ten individuals in each of the defined genotypic classes were tested. This resulted in 146,487 interactions tested. Significant associations were identified with a 0.10 false discovery rate (FDR) considering all traits in the first step and trait by trait in the second step (Storey and Tibshirani 2003). A fixed confidence interval was considered for each association. Horvath et al. (2009) showed that for the 372CC core collection, the most significant LD was limited to 5 cM, so we used twice this value as an estimate of the confidence interval. Maps and QTL were represented with the MapChart v2.1 software (Voorrips 2002).
Validation for field heading date
The whole core collection of 372 accessions was sown in Clermont-Ferrand on 2 November 2005. The heading date was scored on each genotype. These data were used to validate the model derived from the markers identified on the 227 lines. This concerned 130 lines for which the phenotype was predicted using their genotype at the significant loci. The structure of Horvath et al. (2009) based on DArT markers was used.
where HdField is the heading date scored in Clermont-Ferrand on genotype i, G ij the contribution of ancestor group j to genotype i, M ik the effect of the allele of genotype i at marker k, and R i the residual effect. Marker effects were based on a multiple regression analysis where all the significant markers were tested simultaneously.
Results
The structure explained about 30% of the phenotypic variation
Ten traits related to vernalization requirement (VR), photoperiod sensitivity (PS) and narrow-sense earliness were measured on 227 accessions of a core collection devised to represent the wheat worldwide diversity (Balfourier et al. 2007). The structure based on 82 SSR markers (Rousset et al. 2011) explained between 9 and 46% of the phenotypic variance with an average of 30%. Weighted means were computed for each trait and each group (Table 1). Only small differences were observed between groups 2 and 5, and between groups 1, 3 and 4. Except for DLS, groups 2 and 5 had higher values than the other three groups. For HdField, the difference between these two and the other three groups was 122°Cd. Groups 2 and 5 broadly correspond to accessions with a European origin while groups 1, 3 and 4 correspond to accessions with an Asian origin (Balfourier et al. 2007). To identify chromosomal regions possibly contributing to this pattern of mainly two larger groups differing in earliness, based on similarity in heading date groups 1, 3 and 4 were considered together as one large group and groups 2 and 5 as another. For each of the 572 DArT markers, an ANOVA was computed to test whether they participated in the partition into two groups. A total of 373 markers explained significantly the contribution to the two groups. These markers were spread all over the genome.
Single marker association identified 33 regions associated with earliness
For each of the ten traits, an association was tested with 760 markers including SSR, SNP and DArT. A total of 130 significant trait–marker associations were found (Table 1, Tables S2, S3) with 8–22 significant loci for trait-specific association depending on the trait. This represented 62 markers that were spread on all the chromosomes except chromosomes 5D. Chromosome 5D suffered from weak marker coverage as did all group D chromosomes. One to five significant associations per marker were observed (Table S2). The percentage of phenotypic variance explained by individual markers was on average 4.8% and ranged from 2.7 to 9.4% with the largest effect observed for wPt-9971 (6B) on HdV4 (Table S3). Despite some high phenotypic correlations between some traits (data not shown), in most cases at least one specific significant marker was identified for each trait (Table 1). Based on overlap of confidence intervals (fixed to 10 cM), 33 regions associated with earliness and growth habit were identified (Fig. 1).
Results of the association studies conducted on 227 wheat genotypes with 760 polymorphic markers for ten traits related to earliness. The Somers et al. (2004) map was used as a reference map on which markers were projected using MetaQTL (Veyrieras et al. 2007). Only markers used in the association study are represented. When two markers were less than 1 cM apart and not associated, only one marker was represented. Approximate centromere positions are represented in gray based on Somers et al. (2004). For interacting QTL, the name is composed of the trait(s) followed by the name (and chromosome assignation) of the other marker. A confidence interval of the QTL was fixed to 10 cM, based on the average LD in the core collection (Horvath et al. 2009). Distances are in cM. Traits are as follows: HdField heading date of plants sown in the field in autumn, HdV4 heading date of plants vernalized for 4 weeks and planted in the field in mid-April, HdV8 heading date of plants vernalized for 8 weeks and planted in the field in mid-April, HdNV heading date of nonvernalized plants sown in the field in mid-April, PVS partial vernalization requirement, HdV4 − HdV8, HdLD heading date of plants vernalized for 8 weeks and planted in the greenhouse in long day conditions (20 h photoperiod), HdSD heading date of plants vernalized for 8 weeks and planted in the greenhouse in short day conditions (8 h photoperiod), DLS day length sensitivity, HdField − HdV8, difSDLD partial photoperiod sensitivity, HdSD – HdLD, and rapSDLD partial photoperiod sensitivity, HdSD/HdLD. When QTL for a same earliness component overlapped, they were clustered together and their confidence intervals were combined. Regions (based on confidence interval overlap) are numbered as A.01–1.33 for additive regions and I.01–I.07 for interactive regions. Meta-QTLs from Hanocq et al. (2007) and Griffiths et al. (2009) are reported. Chromosome 5D was not represented, as no significant association was identified on it
For narrow-sense earliness, the five-group structure explained about 50% of the total phenotypic variance (Table 1). Twenty markers were associated with the three traits that we considered as representative of narrow-sense earliness (Fig. 1). They were mapped on all chromosomes except the three chromosomes of group 1, and chromosomes 2B, 3A, 3D and 5D. The significant markers carried by chromosomes 2A, 3B, 4A, 5A and 6D were not associated with other components of earliness (trait-specific markers), while for the other loci the markers were also associated with vernalization requirement and/or photoperiodic sensitivity. On average narrow-sense earliness-associated markers explained 20–25% of the phenotypic variance (Table 1).
For vernalization requirement, about 10–30% of the total phenotypic variance for vernalization requirements traits was explained by the structure (Table 1). Twenty-eight loci were associated with the three traits describing vernalization requirements (see “Materials and methods”) and were mapped on all chromosomes except 4D, 5D, 6D and 7A. Seven loci were specifically associated with vernalization requirements. As much as 30–45% of the phenotypic variance was explained by the associated loci (Table 1).
The five-group structure explained from 9 to 38% of the total phenotypic variance for photoperiod sensitivity (Table 1). Twenty-five markers were associated with the four traits used to characterize photoperiodic sensitivity (Fig. 1). They were mapped on all chromosomes except on chromosomes 1D, group 3 chromosomes, 4B, 5A, 5D and 6D. Three regions were associated specifically with photoperiod sensitivity, while the others were also associated with vernalization requirements and/or narrow-sense earliness. The markers explained about 10–30% of the phenotypic variance (Table 1).
Interactions between loci were detected
An analysis of variance was used to test for interaction between the 62 markers that were individually associated. Seven significant interactions were found involving marker wPt-0408 (2B) in interaction with wPt-4144 (2D), wPt-4077 (3A), wPt-1409 (5B), wPt-8283 (7B) for HdV4 and in interaction with wPt-4144 (2D), wPt-4077 (3A), wPt-8283 (7B) for PVS. In each case, the presence of the DArT allele “1” at the wPt-0408 locus gives a spring phenotype regardless of the allele present at the other locus as shown on Fig. 2 for the wPt-0408 × wPt-8283 interaction effect on the HdV4 trait. The presence of the DArT allele “0” at the wPt-0408 locus provides a spring or a winter phenotype depending on the allele present at the other locus.
Allelic effects on heading date of plants partially vernalized for 4 weeks planted in the field in mid-April (ResHdV4) for the four haplotypic classes at the wPt-0408 (2B) and wPt-8283 (7B) loci. Means were calculated with an ANOVA model taking into account the structure of the core collection. The boxes are drawn with widths proportional to the square roots of the number of observations in the groups
For interactions involving at least one marker not individually associated, 11 significant interactions were identified at the 0.10 FDR level (Fig. 1). Those interactions involved 17 different markers in which 2 were also individually associated (wPt-7330 and wPt-0408). This led to the identification of 7 new regions that differed from the 33 regions previously identified with single marker associations. The most significant interactions were identified for wPt-9268 (3A) × wPt-1269 (7D) for HdV4 (Fig. 3a) and wPt-2587 (6B) × wPt-2019 (1B) for the photoperiodic sensitivity-related traits difSDLD (Fig. 3b) and rapSDLD. In the first case, the interaction was similar to the one involving wPt-0408 described above: the presence of the DArT allele “1” at the wPt-9268 or wPt-1269 locus provides a spring phenotype regardless of the allele present at the other locus. A winter phenotype is observed for the combination of the “0” allele at both loci. In the second case, a cross-over interaction is observed and a photoperiod-insensitive phenotype is obtained with two different combinations (Fig. 3b).
Allelic effects for the four haplotypic classes on heading date of plants partially vernalized for 4 weeks planted in the field in mid-April (ResHdV4) at the wPt-9268 (3A) and wPt-1269 (7D) loci (3A) and on the difference in heading dates in short day and long day greenhouse grown plants (ResdifSDLD) at the wPt-2587 (6B) and wPt-2019 (1B) loci (3B). Means were calculated with an ANOVA model taking into account the structure of the core collection. The boxes are drawn with widths proportional to the square roots of the number of observations in the groups
Candidate genes colocalized for some associations
When possible, the map positions of the most obvious candidate genes (Ppd-1, Vrn-1, Vrn-2, Vrn-3, Vrn-4, Rht-1) were added to be compared with significant associations (Fig. 1). The three homoeologous copies of Ppd-1 on group 2 chromosomes were close to markers significantly associated with earliness components (NSE on 2A, VR on 2B, NSE and VR on 2D). Copies of the dwarfing gene Rht1 on chromosomes 4B and 4D and of vernalization gene Vrn-1 on 5A (NSE and VR) and Vrn-2 on 4B (VR) were close or in the confidence intervals of significant associations. Although not in the considered confidence interval, an association was identified near Vrn-D3 (7D) (~20 cM away) for the three components of earliness (NSE, VR, PS). No significant association colocalized near Vrn-B1 (5B), Vrn-A2 (5A), Vrn-D2 (4D), Vrn-A3 (7A), Vrn-B3 (7B) and Vrn-D4 (5D), but no polymorphic markers were tested close to Vrn-A2, Vrn-D2 and Vrn-B3. So 7 out of 11 candidate genes were close to the associated regions.
Assuming a 10-cM confidence interval for each associated marker and using the consensus map to estimate confidence intervals overlap, it is possible to know what proportion of the genome tested for association is covered by associated regions (P = 0.309). We then used a binomial distribution to calculate the probability that at least 7 out of 11 genes are by chance close to associated regions (P = 0.309, k = 7, n = 11). The low probability obtained (0.026) is in favor of a nonrandom distribution of associated regions and mapped major candidate genes.
The model was used to predict field heading date of an independent set of accessions observed at different environmental conditions (location and year)
Independent field data were used to validate the markers identified. The whole 372 core collection was sown at Clermont-Ferrand in autumn 2005. The correlation between HdField (3-year average on the Moulon) and 2005 heading date in Clermont was high for the 225 common accessions (r = 0.94). The structure and the 13 markers associated to HdField (Table 1) were used to predict heading date of 130 lines that were in the core collection CC372, but were not used in the present association study. These lines could be considered as a random sample of the whole core collection as far as heading date is concerned: the average heading date was 1,321°Cd (min = 1,127°Cd, max = 1,517°C) for the 130 genotypes compared to 1,317°Cd for the 225 common accessions (min = 1,138°Cd, max = 1,572°C). A highly significant correlation was found between heading dates measured in Clermont and the predicted heading dates (r = 0.58) (Fig. 4). The structure alone based on the five ancestral groups explained about 20% of the observed heading date in Clermont (r = 0.45). The total model with both the structure and the associated markers explained about 34% of the variation with a root mean square error of prediction of about 59°Cd, which corresponded to about 3.4 days considering that the mean daily temperature in Clermont in May–June 2005 was 17.4°C.
Discussion
A structured core collection was used to identify the main chromosomal regions determining photoperiod sensitivity, vernalization requirement and narrow-sense earliness in wheat. We identified 62 markers individually associated with earliness components and 15 additional markers with interactive effects leading to 40 regions involved. Co-localizations were observed with the Vrn-1, Ppd-1 and Rht-1 candidate genes. Finally, we were able to explain 34% of the variation for field heading date of an independent set of accessions using the structure and the significant markers. We first discuss the number and associated loci in relation to previously published results and known candidate genes, and then the interactions observed between the loci and the validation of the marker model using the independent set of accessions.
A whole-genome scan enabled detection of significant regions
The determinism of flowering date is a complex phenomenon that is likely to be controlled by a larger number of genes than the network composed by the few well-known major genes. The oldest studies focused on chromosomes of group 2 (Welsh et al. 1973; Law et al. 1978) and 5 (Halloran 1967; Law et al. 1976; Maystrenko 1980) that were soon identified as carrying major genes, Ppd-1 and Vrn-1, respectively. Thanks to the fast development of polymorphic molecular markers, whole-genome QTL detections were then conducted on bi-parental recombinant populations. These studies identified from 2 to 11 chromosomal regions (Table S4) associated with earliness-related traits depending on the genetic material used. The information contained in these studies was then compiled using meta-analysis approach that was adapted to QTL studies (Goffinet and Gerber 2000). This approach was first carried out on maize data (Chardon et al. 2004) and then on wheat (Hanocq et al. 2007; Griffiths et al. 2009). The two latter reported 18 and 19 earliness-related meta-QTL, respectively. QTL analysis has the advantages over whole-genome association mapping in that it requires fewer markers to identify rare alleles and is not hindered by population structure (Bergelson and Roux 2010). Combining data through a meta-analysis is then a powerful way to overcome some of its limitations, namely low mapping resolution and low genetic diversity. However, except if large homogenous public datasets are available, it requires more work than association studies as each QTL detection requires about the same quantity of phenotyping as an association study. Moreover, even with the large number of QTL studies published on flowering time in wheat (Table S4), out of the 40 regions that we reported 17 were not identified as metaQTL by either Hanocq et al. (2007) or Griffiths et al. (2009).
A survey of published mapping studies on wheat (Table S4) reveals that probably all chromosomes are involved in the determinism of earliness. This is reinforced by reports on other crop species such as maize for which Buckler et al. (2009) identified about 50–60 QTL regions affecting flowering time, rice with over 80 earliness-related genes and QTL reported (Uwatoko et al. 2008) and oil seed rape (Brassica napus L.) for which Shi et al. (2009) identified 57 meta-QTL associated with earliness traits. On Arabidopsis where the physiological determinants of flowering are better known, the network of interacting genes is very complex, so that quantitative variation may be explained by several loci (Jung and Müller 2009; Higgins et al. 2010). For example, the key integrator gene Flowering Locus C (FLC) is controlled by numerous regulators acting either positively or negatively. In the present study, using association genetics on traits related to earliness component, we identified 40 regions. Additional loci were possibly not identified due to the deficit of markers on group D chromosomes, which may be related to the already known lower level of genetic polymorphism for the D genome (Akbari et al. 2006). We used 760 markers in this study, of which 607 were projected onto the reference map. Considering the Somers et al. (2004) map as a reference, mapped markers covered approximately 61% of the wheat genome, excluding gaps more than 20 cM. This led to an average interval distance of 2.5 cM to be compared with the extent of the LD, which was estimated to be around 5 cM in the core collection used in this study (Horvath et al. 2009). If we consider that marker density should approximately match LD to conduct a comprehensive association study, we can estimate that about 1,000 well-spaced markers would be needed. Such a density is not yet available for bread wheat, but the current progress in sequencing and genotyping technologies will probably fill this gap in the upcoming years. Based on the same calculation, considering that the marker coverage was about 60% and that we detected 40 regions, we can extrapolate that about 65 regions are involved in flowering time on the whole genome. This will nicely correspond to the already cited figures for maize (Buckler et al. 2009) and oilseed rape (Shi et al. 2009).
The panel structured in five main groups based on markers is mostly separated into two groups when earliness is concerned (Table 1). The European accessions (groups 2 and 5, originating from the European wheat genepool) are more photoperiod sensitive and require more vernalization than the other three groups (originating from the same Asian wheat genepool). It is then probable that some associations were reduced or not detected by the structure correction performed in the analysis. However, we identified regions probably corresponding to the major genes on groups 2 (Ppd-1) and 5 (Vrn-A1). This was however not the case for chromosome 5D for which we did not find any association. As previously stated, this was probably caused by the low marker density on this chromosome. Rousset et al. (2011), using the same collection genotyped for the known polymorphism for the three Vrn-1 homoeologs, confirmed their association with vernalization requirements. However, the Vrn-D1 “spring” allele was shown to be rare in this collection, thus exhibiting a weak effect on vernalization requirement. Bonnin et al. (2008) and Rousset et al. (2011) also showed significant associations for Vrn-3 A and D copies on chromosomes of group 7. We found significant associations on chromosomes 7A and 7D, but with markers mapped through projection several cM away from the Vrn-3 genes (Fig. 1). Significant LD existed, however, between wPt-4515 and Vrn-3A on the 7A and between gpw4129 and Vrn-3D on the 7D (Table S5). A consensus map was built to compare the positions of the markers used in the association and candidate genes. Such an approach enabled us to combine in the same map a large number of markers that were mapped on populations that differed in their parents, structure and size. This may, however, lead to approximate order and distance between markers, in particular for SSR markers that amplify different loci on the same linkage group and for which polymorphic bands may differ between populations. We also identified associations on chromosomes 3B, 6A and 6B where the candidate genes Constans (CO), Gigantea (GI) and Luminidependens (LD) were previously localized (Nemoto et al. 2003; Dunford et al. 2005; Rhoné et al. 2010). Based on the LD with markers mapped on chromosome 3B, LD-B1 and GI-B1 probably corresponded to the A.12 region (Fig. 1; Table S5). CO-A1 is in LD with marker cfe273 on the 6A, so that region A.26 may correspond to this candidate gene. A nearby region was already reported by Hanocq et al. (2007) and Griffiths et al. (2009) during their meta-analysis. No significantly associated marker is in LD with CO-B1 on chromosome 6B, so that probably other genes were involved. Finally, we observed significant associations on the chromosomes of group 4. Hanocq et al. (2007) identified meta-QTL on chromosomes 4A and 4B that corresponded to regions A.16 and A.18 in our study. These regions encompassed the dwarfing Rht-1 genes (Fig. 1). Griffiths et al. 2009 identified metaQTL on 4B and 4D. Due to a nonoverlapping confidence interval in their populations, they concluded that it did not appear to be a pleiotropic effect at least for the Rht-D1 gene. Indeed, Youssefian et al. (1992) and Miralles et al. (1997) reported no differences between isogenic Rht1 lines in term of earliness. However, Singh et al. (2001) found on 10 isogenic couples (9 for Rht-B1, 1 for Rht-D1) grown in six environments that dwarf lines headed on average 1.3 days later. These discrepancies could be explained either by background or linked effects, or by the existence of different alleles with different pleiotropic effects. Addisu et al. (2009) reported no differences between the tall genotype (rht-B1a/rht-D1a) and semi-dwarf isogenic lines (Rht-B1b and Rht-D1b), but found a marked delay in flowering (+6 days) for dwarf isogenic lines carrying either the Rht-B1c or Rht-D1c allele. It is highly probable that different Rht-1 alleles are present in the core collection as the range of variation is very large for plant height (Bordes et al. 2008). Therefore, both Rht-B1 and Rht-D1 may be candidate genes for significant associations on chromosomes of group 4, but this still need to be validated. No candidate genes have been reported in additional regions identified in this study; however, analysis of gene networks may provide further candidates for test (Jung and Müller 2009; Higgins et al. 2010).
Two SNP were significantly associated, but none of them seemed to be a direct candidate gene for flowering time. The storage protein activator (SPA) was significantly associated with photoperiod sensitivity on chromosome 1B. However, this transcription factor is known to be specifically expressed in the grain as its maize orthologues opaque-2 (Schmidt et al. 1987). The psy2a gene was significantly associated on chromosome 5A to narrow-sense earliness. This gene is known to be involved in carotenoid biosynthesis and is not associated with endosperm carotenoids (Palaisa et al. 2003; Pozniak et al. 2007). Carotenoids are accessory pigments which play a role in photosynthesis. They are also precursors to abscisic acid (Hirschberg 2001). Thus, this gene may interfere with the photosynthetic complex and impact the photosensitivity in the plant, but the most probable hypothesis is that of a linkage disequilibrium with a flowering time gene.
Several significant interactions between loci were detected
We identified 18 significant interactions between markers. Significant interactions have already been reported between homoeologs for Ppd-1 and Vrn-1. Welsh et al. (1973) first reported that Ppd-D1 and Ppd-B1 showed epistatic interactions using monosomic lines to assign the genes and characterizing distributions for photoperiod response in the progenies of crosses. For both the Ppd-1 and Vrn-1 genes, the epistatic interaction is caused by loss-of-function mutations in regulatory regions that change the pattern of expression (Yan et al. 2004a; Beales et al. 2007). In the model of Loukoianov et al. (2005), a single Vrn-1 allele not repressed by Vrn-2 is sufficient to induce flowering, resulting in an epistatic dominant spring growth habit. Other interactions have been reported between Vrn-1 and Vrn-D4 (Yoshida et al. 2010), between Vrn-D1 and Vrn-3A (Rousset et al. 2011) and between QTL for heading date located on chromosomes 2B–6D and 7A–7D (Zhang et al. 2009). Buckler et al. (2009) reported only very few epistatic loci in maize, making the hypothesis that for outcrossing species the genetic architecture of flowering time is dominated by small additive QTLs with few genetic or environmental interactions. In our study, several of the interactions we detected involved the marker wPt-0408. This marker was mapped on chromosome 2B using mapping populations (Baga et al. 2009; Francki et al. 2009) and Chinese Spring deletion lines (Francki et al. 2009), but a homolog locus was also reported on chromosome 7B (Francki et al. 2009; Bogard et al. 2011). Both loci may be polymorphic in the core collection so that scores may represent a combination of the 2B and 7B loci creating the interaction. The wPt-0408 marker involved interactions with markers on chromosomes 2D, 3A, 5B and 7B. All these interactions dealt with vernalization requirements. Along with the interaction between wPt-9268 (3A) × wPt-2019 (7D) for HdV4, the spring phenotype is dominant with three out of four haplotypes showing a spring habit. This is consistent with interactions between homoeologous copies of the Vrn-1 gene. The crossover interaction reported for wPt-2587 (6B) × wPt-2019 (1B) for photoperiod sensitivity (Fig. 3) is less common. Uwatoko et al. (2008) reported such an interaction also for photoperiod sensitivity in rice between the gene Early Flowering 1 (Ef1 = Ehd1, Os10g32600) and its modifier (e1 = m-Ef1). Ehd1 is a B-type response regulator, whose role is to upregulate FT (Hd3a) in inductive conditions (short days). Ehd1 orthologs were, however, identified in neither Arabidopsis nor Brachypodium suggesting that it was an additional flowering component that evolved within the rice lineage or was deleted from the temperate grasses (Higgins et al. 2010). A crossover interaction suggests alternative routes for the regulation of photoperiod in wheat as was observed in rice.
The marker model explained about 34% of variation in heading date of an independent set of accessions
Thirteen makers were significantly associated with heading date scored for 3 years in the field. We used these markers and the structure developed on the whole core collection to predict heading date measured on an independent set of lines in a different site. Despite a high correlation coefficient of 0.94 between the field heading dates used for the association study and the ones used for the test, only 34% of the variation was explained by the markers and the genetic structure, while 65% was reached in the association study. Different hypotheses can explain the drop in the heading date prediction. The first one is a bad assignation to ancestral groups. Two different structures were used, based on either the 227 genotypes (Rousset et al. 2011) or the whole core collection (Horvath et al. 2009) and the structure alone explained about 20% of the heading date. The five groups identified in the two structures were very similar, but a wrong assignation could have an impact as the structure explained a large part of the variability for earliness components. About 0.1 point r 2 is lost (data not shown) when applying the structure used in the calibration process (Rousset et al. 2011) compared to the structure used in the validation on the same set of genotypes (Horvath et al. 2009). The second hypothesis is the occurrence of missing data. Markers with too many missing data were removed from the association analysis, after rare alleles were coded as missing for SSR. Even if their number was limited (8.3% considering all markers) with a 13-marker model, more than half of the lines had at least one missing data. As each marker explained about 5% of the phenotyping variance, a decrease in the prediction was observed. The third hypothesis is the presence in the validation set of new alleles or loci. This could not be ruled out even if there was no bias in terms of heading date range and geographic origin between the two sets. It is expected that additional studies using different association panels will bring more information on the possibility of constructing a prediction model based on markers.
Conclusions
Genetic studies of flowering time were up to now restricted to bi-parental populations, eventually combined together thanks to a meta-analysis. We carried out a genome-wide association study on a wheat core collection showing the efficiency of such an approach. We thus identified 40 potentially different regions influencing heading date and its components, most of them showing only additive effects. The identification of the most determinant chromosomal regions is of interest to better modulate flowering time, which is a key factor in wheat adaptation to biotic and abiotic stress. The number of regions involved and the small additive effects observed (a little less than 5% on average) show that although major regions have been known for a long time, flowering may still be considered a complex trait. Using a model that included significant markers and the structure showed that it was possible to predict heading date with a mean error of prediction of less than 4 days, which gives hopes for future marker-assisted selection when abundant and cheap markers will be available.
References
Addisu M, Snape JW, Simmonds JR, Gooding MJ (2009) Effects of reduced height (Rht) and photoperiod insensitivity (Ppd) alleles on yield of wheat in contrasting production systems. Euphytica 172:169–181
Akbari M, Wenzl P, Caig V, Carling J, Xia L, Yang S, Uszynski G, Mohler V, Lehmensiek A, Kuchel H, Hayden MJ, Howes N, Sharp P, Vaughan P, Rathmell B, Huttner E, Kilian A (2006) Diversity arrays technology (DArT) for high-throughput profiling of the hexaploid wheat genome. Theor Appl Genet 113:1409–1420
Baga M, Fowler BD, Chibbar RN (2009) Identification of genomic regions determining the phenological development leading to floral transition in wheat (Triticum aestivum L.). J Exp Bot 60:3575–3585
Balfourier F, Ravel C, Bochard AM, Exbrayat-Vinson F, Boutet G, Sourdille P, Dufour F, Charmet G (2006) Développement utilisation et comparaison de différents types de marqueurs pour étudier la diversité parmi une collection de blé tendre. Les Actes du BRG 6:129–144
Balfourier F, Roussel V, Strelchenko P, Exbrayat-Vinson F, Sourdille P, Boutet G, Koenig J, Ravel C, Mitrofanova O, Beckert M, Charmet G (2007) A worldwide bread wheat core collection arrayed in a 384-well plate. Theor Appl Genet 114:1265–1275
Beales J, Turner A, Griffiths S, Snape J, Laurie D (2007) A pseudo-response regulator is misexpressed in the photoperiod insensitive Ppd-D1a mutant of wheat (Triticum aestivum L.). Theor Appl Genet 115:721–733
Bérard A, Le Paslier M, Dardevet M, Exbrayat-Vinson F, Bonnin I, Cenci A, Haudry A, Brunel D, Ravel C (2009) High-throughput single nucleotide polymorphism genotyping in wheat (Triticum spp.). Plant Biotechnol J 7:364–374
Bergelson J, Roux F (2010) Towards identifying genes underlying ecologically relevant traits in Arabidopsis thaliana. Nat Rev Genetics 11:867–879
Bogard M, Jourdan M, Allard V, Martre P, Perretant M-R, Ravel C, Heumez E, Orford S, Snape J, Griffiths S, Gaju O, Foulkes J, Le Gouis J (2011) Anthesis date mainly explained correlations between post-anthesis leaf senescence, grain yield and grain protein concentration in a winter wheat population segregating for flowering time QTL. J Exp Bot 62:3621–3636
Bonnin I, Rousset M, Madur D, Sourdille P, Dupuits C, Brunel D, Goldringer I (2008) FT genome A and D polymorphisms are associated with the variation of earliness components in hexaploid wheat. Theor Appl Genet 116:383–394
Bordes J, Branlard G, Oury FX, Charmet G, Balfourier F (2008) Agronomic characteristics, grain quality and flour rheology of 372 bread wheats in a worldwide core collection. J Cereal Sci 48:569–579
Bordes J, Ravel C, Le Gouis J, Charmet G, Balfourier F (2011) Use of global wheat core collection for association analysis of flour and dough quality traits. J Cereal Sci. doi:10.1016/j.jcs.2011.03.004
Börner A, Schumann E, Fürste A, Cöster H, Leithold B, Röder MS, Weber WE (2002) Mapping of quantitative trait loci determining agronomic important characters in hexaploid wheat (Triticum aestivum L.). Theor Appl Genet 105:921–936
Bradbury PJ, Zhang ZC, Kroon D, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Breseghello F, Sorrells ME (2006) Association mapping of kernel size and milling quality in wheat (Triticum aestivum L.) cultivars. Genetics 172:1165–1177
Brooking IR, Jamieson PD (2002) Temperature and photoperiod response of vernalization in near-isogenic lines of wheat. Field Crop Res 79:21–38
Buckler ES, Holland JB, Bradbury PJ, Acharya CB, Brown PJ, Browne C, Ersoz E, Flint-Garcia S, Garcia A, Glaubitz JC, Goodman MM, Harjes C, Guill K, Kroon DE, Larsson S, Lepak NK, Li H, Mitchell SE, Pressoir G, Peiffer JA, Rosas MO, Rocheford TR, Romay MC, Romero S, Salvo S, Villeda HS, da Silva HS, Sun Q, Tian F, Upadyayula N, Ware D, Yates H, Yu J, Zhang Z, Kresovich S, McMullen MD (2009) The genetic architecture of maize flowering time. Science 235:714–718
Chardon F, Virlon B, Moreau L, Falque M, Joets J, Decousset L, Murigneux A, Charcosset A (2004) Genetic architecture of flowering time in maize as inferred from quantitative trait loci meta-analysis and synteny conservation with the rice genome. Genetics 168:2169–2185
Charmet G, Masood-Quraishi U, Ravel C, Romeuf I, Balfourier F, Perretant M-R, Joseph J-L, Rakszegi M, Guillon F, Sado PE, Bedo Z, Saulnier L (2009) Genetics of dietary fibre in bread wheat. Euphytica 170:155–168
Crossa J, Burgueno J, Dreisickacker S, Vargas M, Herrera-Foessel SA, Lillemo M, Singh RP, Trethowan R, Warburton M, Franco J, Reynolds M, Crouch JH, Ortiz R (2007) Association analysis of historical bread wheat germplasm using additive genetic covariance of relatives and population structure. Genetics 177:1889–1913
Debaeke P (2004) Scenario analysis for cereal management in water-limited conditions by the means of a crop simulation model (STICS). Agronomie 24:315–326
Distelfeld A, Tranquilli G, Li C, Yan L, Dubcovsky J (2009a) Genetic and molecular characterization of the VRN2 Loci in tetraploid wheat. Plant Physiol 149:245–257
Distelfeld A, Li C, Dubcovsky J (2009b) Regulation of flowering in temperate cereals. Curr Opin Plant Biol 12:178–184
Dubcovsky J, Lijavetzky D, Appendino L, Tranquilli G (1998) Comparative RFLP mapping of Triticum monococcum genes controlling vernalization requirement. Theor Appl Genet 97:968–975
Dunford RP, Griffiths S, Christodoulou V, Laurie D (2005) Characterisation of a barley (Hordeum vulgare L.) homologue of the Arabidopsis flowering time regulator GIGANTEA. Theor Appl Genet 110:925–931
Francki MG, Walker E, Crawford AC, Broughton S, Ohm HW, Barclay I, Wilson RE, McLean R (2009) Comparison of genetic and cytogenetic maps of hexaploid wheat (Triticum aestivum L.) using SSR and DArT markers. Mol Gen Genomics 281:181–191
Fu D, Szücs P, Yan L, Helguera M, Skinner J, von Zitzewitz J, Hayes P, Dubcovsky J (2005) Large deletions within the first intron in VRN-1 are associated with spring growth habit in barley and wheat. Mol Gen Genomics 273:54–65
Galiba G, Quarrie S, Sutka J, Morgunov A, Snape J (1995) RFLP mapping of the vernalization (Vrn1) and frost resistance (Fr1) genes on chromosome 5A of wheat. Theor Appl Genet 90:1174–1179
Gate P, Blondlot A, Gouache D, Deudon O, Vignier L (2008) Impacts of climate change on wheat growth in France. What solutions and actions to undertake? Oléagineux Corps gras Lipides 15:332–336
Ge Y, Dudoit S, Speed TP (2003) Resampling-based multiple testing for micro-array data analysis. Test 12:1–44
Goffinet B, Gerber S (2000) Quantitative trait loci: a meta-analysis. Genetics 155:463–473
Goldringer I, Prouin C, Rousset M, Galic N, Bonnin I (2006) Rapid differentiation of experimental populations of wheat for heading time in response to local climatic conditions. Ann Bot 98:805–817
Griffiths S, Simmonds J, Leverington M, Wang Y, Fish L, Sayers L, Alibert L, Orford S, Wingen L, Herry L, Faure S, Laurie D, Bilham L, Snape J (2009) Meta-QTL analysis of the genetic control of ear emergence in elite European winter wheat germplasm. Theor Appl Genet 119:383–395
Halloran GM (1967) Gene dosage and vernalization response in homoeologous group 5 of Triticum aestivum. Genetics 57:401–407
Hanocq E, Sayers EJ, Niarquin M, Le Gouis J, Charmet G, Gervais L, Dedryver F, Duranton N, Marty N, Dufour P, Rousset M, Worland AJ (2003) A QTL analysis for earliness under field and controlled environment conditions in a bread wheat doubled-haploid population. In: Börner A, Snape J (eds), 12th International European Wheat Aneuploid Cooperative, Norwich, UK, pp 57–59
Hanocq E, Niarquin M, Heumez E, Rousset M, Le Gouis J (2004) Detection and mapping of QTL for earliness components in a bread wheat recombinant inbred lines population. Theor Appl Genet 110:106–115
Hanocq E, Laperche A, Jaminon O, Lainé A-L, Le Gouis J (2007) Most significant genome regions involved in the control of earliness traits in Bread Wheat, as revealed by QTL meta-analysis. Theor Appl Genet 114:569–584
Higgins JA, Bailey PC, Laurie DA (2010) Comparative genomics of flowering time pathways using Brachypodium distachyon as a model for the temperate grasses. PLoS One 5:e10065
Hirschberg J (2001) Carotenoid biosynthesis in flowering plants. Curr Opin Plant Biol 4:210–218
Horvath A, Didier A, Koenig J, Exbrayat F, Charmet G, Balfourier F (2009) Analysis of diversity and linkage disequilibrium along chromosome 3B of bread wheat (Triticum aestivum L.). Theor Appl Genet 119:1523–1537
Jung C, Müller AE (2009) Flowering time control and applications in plant breeding. Trends Plant Sci 14:1360–1385
Kang HM, Zaitlen NA, Wade CM, Kirby A, Heckerman D, Daly MJ, Eskin E (2008) Efficient control of population structure in model organism association mapping. Genetics 178:1709–1723
Kato K, Nakagawa K, Kuno H (1993) Chromosomal location of the vernalization response, Vrn2 and Vrn4, in common wheat, Triticum aestivum L. Wheat Inf Serv 76:53
Kato K, Miura H, Akiyama M, Kuroshima M, Sawada S (1998) RFLP mapping of the three major genes, Vrn1, Q and B1, on the long arm of chromosome 5A of wheat. Euphytica 101:91–95
Kato K, Yamashita M, Ishimoto K, Yoshino H, Fujita M (2003) Genetic analysis of two genes for vernalization response, the former Vrn2 and Vrn4, using PCR based molecular markers. In: Pogna NE, Romano N, Pogna EA, Galterio G (eds), 10th international wheat genetics symposium, Paestum, Italy, pp 971–973
Kuchel H, Hollamby G, Langridge P, Williams K, Jefferies SP (2006) Identification of genetic loci associated with ear-emergence in bread wheat. Theor Appl Genet 113:1103–1112
Law CN, Worland AJ (1997) Genetic analyses of some flowering time and adaptative traits in wheat. New Phytol 137:19–28
Law CN, Worland AJ, Giorgi B (1976) Genetic-control of ear-emergence time by chromosomes-5a and chromosomes-5D of wheat. Heredity 36:49–58
Law CN, Sutka J, Worland AJ (1978) A genetic study of daylength response in wheat. Heredity 41:185–191
Loukoianov A, Yan L, Blechl A, Sanchez A, Dubcovsky J (2005) Regulation of VRN-1 vernalization genes in normal and transgenic polyploid wheat. Plant Physiol 138:2364–2373
Maystrenko OI (1980) Cytogenetic study of the growth habit and ear emergence time on chromosome 2B of wheat. In: Vartanian ME (ed) Well-being in mankind and genetics, 14th international congress of genetics, Vol 1, MIR Publishers, Moscow, pp 267–282
Miralles DJ, Slafer GA, Lynch V (1997) Rooting patterns in near-isogenic lines of spring wheat for dwarfism. Plant Soil 197:79–86
Nemoto Y, Kisakal M, Fuse T, Yano M, Ogihara Y (2003) Characterization and functional analysis of three wheat genes with homology to the CONSTANS flowering time gene in transgenic rice. Plant J 36:82–93
Palaisa KA, Morgante M, Williams M, Rafalski A (2003) Contrasting effects of selection on sequence diversity and linkage disequilibrium at two phytoene synthase loci. Plant Cell 15:1795–1806
Pozniak CJ, Knox RE, Clarke FR, Clarke JM (2007) Identification of QTL and association of a phytoene synthase gene with endosperm colour in durum wheat. Theor Appl Genet 114:525–537
Ravel C, Praud S, Murigneux A, Linossier L, Dardevet M, Balfourier F, Dufour P, Brunel D, Charmet G (2006) Identification of Glu-B1-1 as a candidate gene for the quantity of high-molecular-weight glutenin in bread wheat (Triticum aestivum L.) by means of an association study. Theor Appl Genet 112:738–743
R Development Core Team (2009). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. ISBN 3-900051-07-0, URL http://www.R-project.org
Rhoné B, Vitalis R, Goldringer I, Bonnin I (2010) Evolution of flowering time in experimental wheat populations: a comprehensive approach to detect genetic signatures of natural selection. Evolution 64:2110–2125
Rousset M, Bonnin I, Remoué C, Falque M, Rhoné B, Veyrieiras J-B, Madur D, Balfourier F, Le Gouis J, Santoni S, Goldringer I (2011) Deciphering the genetics of growth habit by an association study on candidate genes in bread wheat (Triticum aestivum L.). Theor Appl Genet. doi:10.1007/s00122-011-1636-2
Santra D, Santra M, Allan RE, Campbell K, Kidwell K (2009) Genetic and molecular characterization of vernalisation genes Vrn-A1, Vrn-B1, and Vrn-D1 in spring wheat germplasm from the Pacific Northwest Region of the USA. Plant Breed 128:576–584
Schmidt RJ, Burr A, Burr B (1987) Transposon tagging and molecular analysis of the maize regulatory locus opaque-2. Science 238:960–963
Shi J, Li R, Qiu D, Jiang C, Long Y, Morgan C, Bancroft I, Zhao J, Meng J (2009) Unraveling the complex trait of crop yield with quantitative trait loci mapping in Brassica napus. Genetics 182:851–861
Singh RP, Huerta-Espino J, Rajaram S, Crossa J (2001) Grain yield and other traits of tall and dwarf isolines of modern bread and durum wheats. Euphytica 119:241–244
Somers D, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Sourdille P, Snape JW, Cadalen T, Charmet G, Nakata N, Bernard S, Bernard M (2000) Detection of QTLs for heading time and photoperiod response in wheat using a doubled-haploid population. Genome 43:487–494
Sourdille P, Cadalen T, Guyomarc’h H, Snape JW, Perretant MR, Charmet G, Boeuf C, Bernard S, Bernard M (2003) An update of the Courtot × Chinese Spring intervarietal molecular marker linkage map for the QTL detection of agronomic traits in wheat. Theor Appl Genet 106:530–538
Stelmakh A (1990) Geographic distribution of Vrn genes in landraces and improved varieties of spring bread wheat. Euphytica 45:113–118
Storey JD, Tibshirani R (2003) Statistical significance for genomewide studies. Proc Natl Acad Sci USA 100:9440–9445
Trevaskis B (2010) The central role of the VERNALIZATION1 gene in the vernalization response of cereals. Funct Plant Biol 37:479–487
Trevaskis B, Hemming M, Dennis E, Peacock W (2007) The molecular basis of vernalization-induced flowering in cereals. Trends Plant Sci 12:352–357
Turner A, Beales J, Faure S, Dunford RP, Laurie DA (2005) The pseudo-response regulator Ppd-H1 provides adaptation to photoperiod in barley. Science 310:1031–1034
Uwatoko N, Onishi A, Ikeda Y, Kontani M, Sasaki A, Matsubara K, Itoh Y, Sano Y (2008) Epistasis among the three major flowering time genes in rice: coordinate changes of photoperiod sensitivity, basic vegetative growth and optimum photoperiod. Euphytica 163:167–175
Veyrieras J-B, Goffinet B, Charcosset A (2007) MetaQTL: a package of new computational methods for the meta-analysis of QTL mapping experiments. BMC Bioinformatics 8:1–16
Voorrips R (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Weir BS (1996) Genetic data analysis. II. Methods for discrete population genetic data. Sinauer Associates Inc, Sunderland, p 376
Welsh JR, Keim DL, Pirasteh B, Richards RD (1973) Genetic control of photoperiod response in wheat. In: 4th international wheat genetics symposium, Missouri, pp 879–884
Wenzl P, Carling J, Kudrna D, Jaccoud D, Huttner E, Kleinhofs A, Kilian A (2004) Diversity arrays technology (DArT) for whole-genome profiling of barley. Proc Natl Acad Sci USA 101:9915–9920
Worland A (1996) The influence of flowering time genes on environmental adaptability in European wheats. Euphytica 89:49–57
Yan L, Loukoianov A, Tranquilli G, Helguera M, Fahima T, Dubcovsky J (2003) Positional cloning of the wheat vernalization gene VRN1. Proc Natl Acad Sci USA 100:6263–6268
Yan L, Helguera M, Kato K, Fukuyama S, Sherman J, Dubcovsky J (2004a) Allelic variation at the VRN-1 promoter region in polyploid wheat. Theor Appl Genet 109:1677–1686
Yan L, Loukoianov A, Blechl A, Tranquilli G, Ramakrishna W, SanMiguel P, Bennetzen JL, Echenique V, Dubcovsky J (2004b) The wheat VRN2 gene is a flowering repressor down-regulated by vernalization. Science 12:1640–1644
Yan L, Fu D, Li C, Blechl A, Tranquilli G, Bonafede M, Sanchez A, Valarik M, Yasuda S, Dubcovsky J (2006) The wheat and barley vernalization gene VRN3 is an orthologue of FT. Proc Natl Acad Sci USA 103:19581–19586
Yoshida T, Nishida H, Zhu J, Nitcher R, Distelfeld A, Akashi Y, Kato K, Dubcovsky J (2010) Vrn-D4 is a vernalization gene located on the centromeric region of chromosome 5D in hexaploid wheat. Theor Appl Genet 120:543–552
Youssefian S, Kirby EJM, Gale MD (1992) Pleiotropic effects of the GA-insensitive Rht dwarfing genes in wheat. 1. Effects on development of the ear, stem and leaves. Field Crops Res 28:179–190
Zhang K, Tian J, Zhao L, Liu B, Chen G (2009) Detection of quantitative trait loci for heading date based on the doubled haploid progeny of two elite Chinese wheat cultivars. Genetica 135:257–265
Zheng S, Byrne P, Bai G, Shan X, Reid S, Haley S, Seabourn B (2009) Association analysis reveals effects of wheat glutenin alleles and rye translocations on dough-mixing properties. J Cereal Sci 50:283–290
Acknowledgments
The authors acknowledge the experimental work carried out by Jean-Pierre Noclerq (INRA, Estrées-Mons). This work was partly supported by the “Agence Nationale de la Recherche” (ANR) Wheat Performance Project (ANR07-GPLA023).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by I. Mackay.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Le Gouis, J., Bordes, J., Ravel, C. et al. Genome-wide association analysis to identify chromosomal regions determining components of earliness in wheat. Theor Appl Genet 124, 597–611 (2012). https://doi.org/10.1007/s00122-011-1732-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-011-1732-3