Abstract
Key message
Using a much higher number of SNP markers and larger sample sizes than all the previous studies, we characterized the genetic relationships among wild and cultivated plants of section Beta.
Abstract
We analyzed the genetic variation of Beta section Beta, which includes wild taxa (Beta macrocarpa, B. patula, B. vulgaris subsp. adanensis and B. vulgaris subsp. maritima) and cultivars (fodder beet, sugar beet, garden beet, leaf beet, and swiss chards), using 9724 single nucleotide polymorphism markers. The analyses conducted at the individual level without a priori groups confirmed the strong differentiation of B. macrocarpa and B. vulgaris subsp. adanensis from the other taxa. B. vulgaris subsp. maritima showed a complex genetic structure partly following a geographical pattern, which confounded the differences between this taxon and the cultivated varieties. Cultivated varieties were structured into three main groups: garden beets, fodder and sugar beets, and leaf beets and swiss chards. The genetic structure described here will be helpful to correctly estimate linkage disequilibrium and to test for statistical associations between genetic markers and environmental variables.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants closely related to crops (crop wild relatives) have the potential to contribute beneficial traits for crop improvement (Maxted et al. 2006) and are receiving increased attention in plant breeding (Warschefsky et al. 2014). Assessing genetic diversity and structure of crops and their wild relatives is useful for determining levels of genetic variability and estimating potential losses of genetic diversity associated with selection and domestication. A description of the genetic structure of wild and cultivated plants is also needed to identify markers under selection through genome–environment association methods (Schoville et al. 2012), which require a characterization of the neutral genetic structure of the data, and to correctly estimate the linkage disequilibrium between markers (Mangin et al. 2012), which can then be used to infer the selection history of genetic pools and to design genome-wide association studies for quantitative trait loci (e.g. Mangin et al. 2015). Germplasm collections represent the core resource to conduct genetic diversity analysis on crops and their wild relatives.
The sugar beet (Beta vulgaris subsp. vulgaris) is one of the most important crops in terms of metric tonnage (about 140 Mt in year 2000) and accounts for approximately one-fourth of the world’s sugar production (Draycott 2006). The domestication of sugar beet started late, in the eighteenth century, and is, therefore, relatively recent when compared to that of other major crop plants (Biancardi et al. 2012). Wild and cultivated relatives of the sugar beet are included in Beta section Beta (Kadereit et al. 2006; Frese 2010). The genetic diversity of Beta section Beta germplasm accessions has been studied with various genetic markers, including allozymes (Abe and Tsuda 1987; Letschert 1993), chloroplast, and mitochondrial markers (Mikami et al. 1984; Kishima et al. 1987; Fritzsche et al. 1987; Bonavent et al. 1989; Ecke and Michaelis 1990; Senda et al. 1998; Cheng et al. 2009, 2011; Yoshida et al. 2012), restriction fragment length polymorphisms (RFLPs; Mita et al. 1991; Santoni and Bervillé 1992a; Jung et al. 1993; Hjerdin et al. 1994; Desplanque et al. 1999), random amplification of polymorphic DNA (McGrath et al. 1999; Wang and Goldman 1999; Shun et al. 2000; Litwiniec et al. 2016), satellites and short-sequence repeats (SSRs; Santoni and Bervillé 1992b; Fénart et al. 2008; Stevanato et al. 2013), amplified fragment length polymorphisms (Saccomani et al. 2009), and diversity array technology (Andrello et al. 2016). These studies differed greatly in the sample size, taxonomic coverage, and geographical extent of the genotyped accessions. Microsatellites were also extensively used to study genetic structure and gene flow among sugar beets, ruderal and sea beets (B. vulgaris subsp. maritima), and weed (feral) beets in the field (e.g., Viard et al. 2004; Fievet et al. 2007; Fénart et al. 2008; Arnaud et al. 2009; Leys et al. 2014). The most recent investigations on genetic variation in Beta section Beta were performed using single nucleotide polymorphisms (SNP) markers (Adetunji et al. 2014; Mangin et al. 2015). In addition, the first genome of sugar beet was published (Dohm et al. 2014).
SNPs offer numerous advantages over other types of markers for studies on genetic diversity and varietal identification of crop and wild relatives (Kilian and Graner 2012). Compared to SSR markers, SNPs are often more densely and evenly distributed in the genome and their number is much larger (Morin et al. 2004). In addition, the bi-allelic nature of SNPs makes them easy to read and compare between different data sources. Despite some drawbacks, especially for SNPs genotyped by SNP arrays (ascertainment bias and low diversity), SNPs are now widely used for analysis of genetic diversity in crops (Lu et al. 2009; Winkler et al. 2016) and varietal identification by DNA fingerprinting (Curk et al. 2015; Tian et al. 2015). Species relationships and taxonomic studies on the basis of SNP markers have been reported for several crop–wild-relative systems such as Citrus ssp. (Curk et al. 2015), Brassica ssp. (Mason et al. 2015), carrots (Rong et al. 2014) and tomatoes (Bauchet et al. 2014), including the aforementioned studies on taxa of Beta section Beta (Adetunji et al. 2014; Mangin et al. 2015).
The taxonomy and genetic structure of section Beta has been extensively studied and today three species are recognized (Letschert 1993): Beta macrocarpa Gussone, Beta patula Aiton, and Beta vulgaris. The species B. vulgaris is further subdivided into three subspecies: B. vulgaris subsp. maritima (L.) Arcangeli, which is considered the ancestor of all cultivated beets (Biancardi et al. 2012); B. vulgaris subsp. adanensis (Pamukçuoglu) Ford-Lloyd and Williams; and B. vulgaris subsp. vulgaris, which contains all the cultivated forms: sugar beets (the most important source of sugar worldwide after sugar cane), fodder beets (used to feed livestock), garden beets (also known as table beets or red beets, utilized as vegetables for their roots), leaf beets, and swiss chards (utilized as vegetables for their leaves). Letschert (1993) discouraged any formal classification of the cultivated plants and advised referring them to categories such as the cultivar groups or cultivars.
While the genetic relationships among the wild forms of section Beta are substantially clear (Letschert 1993; Jung et al. 1993; Andrello et al. 2016), the genetic structure of the cultivated varieties and the relationships between them and the wild forms are still poorly understood. Slight genetic differences are shown when comparing cultivated varieties with allozymes (Abe and Tsuda 1987; Nagamine et al. 1989). Mitochondrial DNA is very effective to distinguish male sterile plants from fertile plants and to distinguish different sterile lines (Santoni et al. 1991; Weihe et al. 1991), because male sterility is a consequence of inactivation of mitochondrial functions (Mikami et al. 2011), but does not differentiate among varieties with fertile cytoplasm (Ecke and Michaelis 1990). RFLPs make it possible to distinguish between cultivated and wild plants (Mita et al. 1991; Santoni and Bervillé 1992a), but the genetic relationships between varieties are not consistent across different studies. While some analyses showed a clear genetic separation between swiss chards and the other varieties (Santoni and Bervillé 1992a; Jung et al. 1993) and between fodder and sugar beets (Hjerdin et al. 1994), other analyses did not lead to distinct genetic clusters separating fodder beets, sugar beets, garden beets, and swiss chards (Jung et al. 1993; Wang and Goldman 1999). The most recent studies using SNP markers did not detect a clear differentiation between B. vulgaris subsp. maritima and cultivated beets (Adetunji et al. 2014) or among varieties (Mangin et al. 2015), most likely because of the limited number of accessions and SNP markers (91 accessions genotyped with 418 SNP markers in Adetunji et al. 2014; 2035 accessions with 320 markers in Mangin et al. 2015).
The first objective of the present study is to describe the genetic structure of wild and cultivated accessions of Beta section Beta using a larger sample size, a wider geographical coverage and a higher number of SNP markers than all previous studies. The second objective is to compare genetic diversity among the taxa and cultivars. Our accessions came from the entire distribution area of the wild varieties and the main areas of cultivation of the cultivars (Andrello et al. 2016). This large collection effort allowed us to describe the genetic diversity and structure within taxa and to account for the effects of within-taxon diversity on genetic differentiation between taxa.
Materials and methods
Plant materials and genetic markers
The dataset analyzed in this study comprises 1512 individuals of beets belonging to 1080 accessions (for 36 accessions, multiple individuals were genotyped) recovered from 12 germplasm banks worldwide (Mangin et al. 2015; Andrello et al. 2016). Information on the individuals (mainly country of origin, species and subspecies) were found in web databases such as GRIN-ARS (USDA, US, http://www.ars-grin.gov/npgs/) and EURISCO (ECPGR, Europe, http://www.ecpgr.cgiar.org/germplasm_databases.html). This set included individuals of all taxa and cultivated varieties of Beta section Beta (Table 1) and covers the entire native range of wild beets and the main growing regions of the cultivated varieties. Since different ploidies are found in section Beta germplasm (Letschert 1993; Draycott 2006), only diploid accessions were used after ploidy determination using flow cytometry. Polyploids were removed because crossing them with the diploid elite line would generate triploid F1s, which most likely would be sterile. Returning to a diploid level would have required much labor and time through back-crossing. Of the 25 tetraploid accessions found in our initial dataset, there were 1 fodder beet, 1 sugar beet, and 23 B. macrocarpa.
Seeds were germinated in the greenhouse and leaf material was collected prior to DNA extraction and genotyping on a 35 k Axiom® beet array. The 35 k Axiom® beet genotyping array was developed based on structural differences between an elite line and 16 accessions for which 2 × 100 paired-end NGS reads were mapped on the reference draft genome of the sugar beet (Dohm et al. 2014). The 16 accessions consisted of 13 B. vulgaris subsp. maritima, 2 sugar beet, and 1 fodder beet; therefore, this design was suitable to detect polymorphism among cultigroups and wild beets. The method used to choose the 13 B. vulgaris subsp. maritima accessions to develop the Affymetrix array was based on genetic diversity and allele richness (determined with SNP and DArT markers) and spatial extent. Of these 13 accessions, 10 were included in subsequent analyses of genetic structure.
Using a bioinformatics pipeline, a subset of 33,000 high-quality SNPs was selected to design the Axiom® array. Then, the array was employed to generate 18,333 high-density SNP markers for the individuals analyzed in this study. The analyses presented in this study were performed on a subset of 9724 SNPs corresponding to the “Poly High Resolution” “Mono High Resolution” and “No Minor Homozygote” categories of the Axiom® genotyping array. These categories comprise the most accurate and polymorphic SNPs according to the Axiom® best practices genotyping workflow (Thermo Fisher Scientific Inc. 2017).
Analysis of genetic structure
To assess genetic variation at the individual level, we first applied a principal component analysis on individual genotypes; values were centered but not reduced and the analysis was performed in R with the function “glPca” of adegenet. The resulting principal components were used to cluster individuals into distinct genetic groups, using a k-means algorithm as implemented in the “find.clusters” function of adegenet. We determined the optimal number of genetic groups by visually inspecting the curve of the Bayesian information criterion (BIC) against the number of groups. We then applied a discriminant analysis of principal components (DAPC) to describe the genetic clusters (Jombart et al. 2010) using the clusters identified with the k-means algorithm, as implemented in the “dapc” function of adegenet. All the principal components of genetic data (1512) were kept for the k-means algorithm, while only 701 principal components (which explained 90% of the genetic variance) were retained in the discriminant analysis to avoid overfitting.
Estimation of genetic diversity
For each wild taxon and each cultivar, genetic diversity was estimated using the expected heterozygosity (H e) and the proportion of polymorphic loci (P). H e was estimated using the functions “summary” of the R package adegenet 2.0.0 (Jombart and Ahmed 2011). P was calculated as A r − 1, where A r is the mean allelic richness after rarefaction (El Mousadik and Petit 1996) estimated using the function “allel.rich” of the R package PopGenReport; the rarefaction method allows for comparisons between groups with different sample sizes, as it estimates A r using the smallest sample size of the dataset. B. patula was excluded from the calculation of A r as it is represented by only one individual; thus, the smallest sample size was represented by the 26 individuals (hence, 52 alleles) of B. vulgaris subsp. adanensis. Differences in P between every pair of taxa or cultivars were assessed using one-tailed permutation tests (999 permutations).
Results
Analysis of genetic structure
The visual inspection of the curve of BIC of the k-means algorithm suggested between 9 and 20 genetic groups. For simplicity, we chose nine groups for the DAPC (Fig. 1). Table 2 contains the number of individuals for each taxon and cultivar in each genetic group. All posterior membership probabilities were > 0.99.
Discriminant analysis of principal components (DAPC). a The first and second discriminant functions, b the first and third discriminant function. Each individual is represented by a dot colored according to its taxonomic identification. The nine genetic groups identified with the k-means algorithm are identified by dotted and numbered ellipses
The DAPC showed several patterns. First, most genetic groups comprised several taxa and/or cultivars, thus there was no exact correspondence between genetic and taxonomic groups. Second, some groups were not well separated by the three first discriminant functions: for example, group #9 overlapped with group #3, group #1 overlapped with group #8. Since the DAPC calculates the discriminant axes to maximize the separation of genetic groups, the overlap demonstrates that these groups are very similar. Both these observations suggest that Beta section Beta has a complex genetic structure that differs from the taxonomic structure.
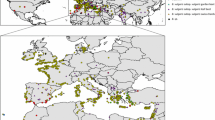
The first discriminant axis described genetic differentiation between group #4 and the other groups. Group #4 contained almost all the individuals of B. vulgaris subsp. adanensis and 110 individuals of B. vulgaris subsp. maritima, plus two individuals belonging to the cultivars. This group can therefore be associated with wild accessions from the Eastern Mediterranean basin and the Middle East (Fig. 2). The 110 B. vulgaris subsp. maritima individuals are probably misclassified and are actually B. vulgaris subsp. adanensis. Misclassifications are common given the difficulty of morphological discrimination for wild beets, which relies on floral traits and is thus very difficult without flowers (Letschert 1993; Leys et al. 2014; Andrello et al. 2016). The second discriminant axis described genetic differentiation between group #5 and the others. Group #5 comprised almost exclusively individuals of B. macrocarpa, with one individual of B. vulgaris subsp. maritima (probably misclassified) and the only accession of B. patula. The geographic distribution of this group; therefore, corresponds to the native range of these two species (Mediterranean basin and Canary Islands for B. macrocarpa and Madeira for B. patula; Fig. 2). In summary, the first and second discriminant axes separated three clusters of individuals, the first one consisting of B. macrocarpa and B. patula, the second one formed by the subspecies B. vulgaris subsp. adanensis and the third one composed by the other accessions. The structure within this third cluster was resolved by the third discriminant axis.
Geographical distribution of the nine genetic groups identified with the k-means algorithm. Each individual is represented by a dot colored according to its taxonomic group as in Fig. 1
The third discriminant axis described the genetic structure of B. vulgaris subsp. maritima and the cultivars. Four large groups, composed almost exclusively by accessions of B. vulgaris subsp. maritima, represented the genetic differentiation within this taxon (groups #2, #3, #6 and #7). Group #2 had its geographical center on the Atlantic coasts and extended from Denmark to Western Mediterranean (Fig. 2). Group #7 consisted exclusively of accessions from Morocco. Group #3 was centered on the Mediterranean basin, but extended from the British Isles to the Red Sea. Group #6 had an even more Eastern distribution, centered on India. The third discriminant axis thus corresponded to a geographical gradient going from North–West (negative values) to South–East (positive values).
On the third discriminant axis, cultivars were located within the geographical gradient of B. vulgaris subsp. maritima. Three groups could be recognized. Group # 1 included fodder and sugar beets and individuals of B. vulgaris subsp. maritima. Geographically, the individuals of B. vulgaris subsp. maritima of this group were scattered over a large area. Group #8 comprised most garden beet accessions, but included individuals of other cultivars and of B. vulgaris subsp. maritima. The individuals of B. vulgaris subsp. maritima in this group were located in the Mediterranean basin. Group #9 included most leaf beet and swiss chards accessions and individuals of B. vulgaris subsp. maritima. The individuals of B. vulgaris subsp. maritima of this group were scattered over a large geographical area. In summary, cultivars were distributed in three groups, the first one corresponding to fodder and sugar beets, the second one to garden beets and the third one to leaf beets and swiss chards, but all these groups also contained wild individuals.
The location of the ten accessions of B. vulgaris subsp. maritima used to develop the Axiom® genotyping array in the DAPC plane is shown in Supplementary Fig. 1. This shows that this SNP array is probably more appropriate to study genetic diversity of groups #2 and #9 than other groups.
Estimation of genetic diversity of wild and cultivated taxa
The lowest levels of genetic diversity, in terms of proportion of polymorphic markers P and expected heterozygosity H e, were observed in B. macrocarpa (Table 1), while the other taxa and cultivars exhibited significantly higher levels of genetic diversity (permutation tests, p < 0.05). The proportion of polymorphic markers did not significantly differ among the other taxa and cultivars (permutation tests, p > 0.25).
Discussion
Genetic differentiation between species and within B. vulgaris subsp. maritima
The first result of our study is the differentiation of B. macrocarpa and B. vulgaris subsp. adanensis relatively to the other species. One hundred and ten individuals of B. vulgaris subsp. maritima (10%) clustered with the accessions of B. vulgaris subsp. adanensis. This is likely the result of misclassifications of some accessions. In a previous study on a similar set of accessions using DArTs (Andrello et al. 2016), we observed the same patterns of genetic differentiation, which confirms the results of crossings (Abe et al. 1986) and the patterns of morphological (Buttler 1977; Letschert 1993) and genetic differentiation obtained with other types of markers: allozymes (Letschert 1993), RFLPs (Jung et al. 1993), RAPDs (Shen et al. 1998), nuclear sequences (Kadereit et al. 2006; Villain 2007), mitochondrial, and nuclear SSRs (Leys et al. 2014). In the present study, B. patula clustered with B. macrocarpa, while DArTs led us to cluster B. patula with B. vulgaris subsp. maritima. However, both analyses were based on a single accession, and a clearer picture of the position of B. patula among the species of section Beta would benefit from the collection of further data on this microendemic taxon (Frese et al. 2012).
Because B. macrocarpa and B. vulgaris subsp. adanensis are the most genetically divergent taxa, crossings with cultivated varieties are expected to be less successful (van Geyt et al. 1990). However, B. macrocarpa was used as a pollen donor to obtain highly productive germplasm lines (Campbell 2010). B. macrocarpa has its distributional range in the Southern Mediterranean where the climate is dry and warm, and has been suggested as a source of genetic material for the introgression of drought-adaptive traits in sugar beet (Monteiro et al. 2013). Improving drought tolerance in beets is useful for the development of rain-fed beets (Ober et al. 2004), which may suffer important productivity reduction from the effects of changing climate (Jones et al. 2003).
Genetic structure within B. vulgaris subsp. maritima was driven by geography, with four large groups corresponding to as many geographical regions. However, two of these four groups (#3 and #6, Fig. 2) had a very large geographical spread and group plants with very distant spatial locations. The Strait of Gibraltar does not seem to constitute a barrier to gene flow, given the presence of Mediterranean individuals in cluster #2, which is composed mainly by Northern Atlantic individuals. The patterns of genetic variation of wild taxa thus resemble much the patterns obtained with DArTs markers (Andrello et al. 2016). In those analyses, we showed that the genetic clusters are disposed onto a spatial gradient going from North–West to South–East, which is part of a genetic continuum showing isolation by distance and no clear genetic discontinuities.
Differentiation between cultivars and B. vulgaris subsp. maritima
Past studies suggested a scenario of both strong genetic similarities among cultivars (Ecke and Michaelis 1990; Jung et al. 1993) and clear genetic separation (Hjerdin et al. 1994; McGrath et al. 1999) between cultivars and B. vulgaris subsp. maritima, but were based on smaller numbers of accessions, fewer genetic markers, and smaller geographical regions than the ones of the present study. One of the most recent studies, using 418 SNP markers on 324 individuals coming from the entire distribution area of the taxon (Adetunji et al. 2014), showed no clear separation between B. vulgaris subsp. maritima and cultivated taxa. Adetunji et al. (2014) attributed the lack of structure to small sample sizes, incorrect annotation of the germplasm material, and weak resolution power of their SNP panel. Another explanation is the complex genetic structure of wild taxa, especially B. vulgaris subsp. maritima, which is far from being a homogeneous taxon. The large within-taxon diversity of B. vulgaris subsp. maritima was also evidenced by past morphological (Letschert 1993) and genetic studies (Andrello et al. 2016). It is difficult to observe clear patterns of genetic differentiation between the accessions of B. vulgaris subsp. maritima taken as a whole and those of the cultivated beets, because the complex structure of B. vulgaris subsp. maritima blurs the differences that may exist between single accessions. If the B. vulgaris subsp. maritima taxon is considered by separating it into genetic groups as those evidenced with the k-means algorithm, clearer genetic differences may emerge between wild and cultivated taxa. For example, cultivated accessions will likely be more differentiated from wild accessions of cluster #2 than from wild accessions of clusters #8 (which includes many garden beets) and #9 (which includes many swiss chards).
A specific pattern uncovered by our analysis is the presence of cultivated and wild individuals in the same genetic cluster (group #1, #8 and #9). The wild individuals may be related to the original individuals used for the domestication of the crops, but the genetic clusters do not allow us to identify the original geographic regions of crop domestication: the B. vulgaris subsp. maritima accessions of group #1 (which includes the accessions of sugar and fodder beet) are scattered over the entire distribution area of the species, and the same can be said of group #9 (swiss chards and leaf beets). Even if group #8 (including garden beet) shows a more circumscribed distribution, centered on the Mediterranean basin, it comprises a lower number of wild individuals than the others, and its small spatial range may be the result of small sample size. In addition, the genetic signature of domestication in present samples may be confounded by the repeated crossings of cultivated lines with wild material (e.g. Bonavent et al. 1989). Advanced methods for demographic reconstruction based on SNPs (Excoffier et al. 2013; Schiffels and Durbin 2014) could be employed to assess the likelihood of several demographic scenarios and give deeper insights on the history of domestication in beets.
Genetic diversity is expected to be lower in cultivated than in wild varieties due to bottlenecks and selection of adaptive traits associated with the domestication process (Hyten et al. 2006). Past studies on taxa of section Beta have evidenced generalized reductions of genetic diversity in cultivars relative to B. vulgaris subsp. maritima or feral beets using a variety of molecular markers (Desplanque et al. 1999; Arnaud et al. 2003; Viard et al. 2004; Fénart et al. 2008; Litwiniec et al. 2016). In our study, the proportion of polymorphic SNP markers in cultivars was not significantly smaller than in B. vulgaris subsp. maritima. This is most likely due to the limited number of accessions (16) used to develop the genotyping array, which likely led to an ascertainment bias towards SNPs with medium frequencies and against SNPs with low frequency in B. vulgaris subsp. maritima. The ascertainment bias may have limited the power to detect a genetic bottleneck associated with domestication, because the effects of bottlenecks are stronger for rare alleles (Hyten et al. 2006).
Differentiation among cultivars
Past studies did not fully clarify the genetic structure between cultivars (Ecke and Michaelis 1990; Mita et al. 1991; Jung et al. 1993; Wang and Goldman 1999). Jung et al. (1993) analyzed sugar beets, fodder beets, garden beets, and swiss chards. In their dendrogram, the sugar beet accessions were closer to the fodder beet than to the swiss chards accessions, which suggests that fodder and sugar beets should be genetically similar. The position of garden beets was uncertain because some accessions were closer to the fodder and sugar beets and some others were closer to swiss chards. Recently, Mangin et al. (2015) used the same analytical methods used in our work (k-means on principal components on SNP markers) and found three large groups for cultivated beets: a first group containing sugar and fodder beets (and some garden beets), a second group including fodder and garden beets (and some sugar beets), and a third group composed of sugar beets, fodder beets, garden beets, and swiss chards in similar proportions. Therefore, the data reported by Mangin et al. (2015) did not allow them to fully separate the cultigroups on a genetic basis.
Using a much higher number of SNP markers, we could deepen the pattern of genetic structure among cultivated groups. In particular, our analyses confirmed the pattern of separation between fodder and sugar beets on one side and leaf beets and swiss chards on the other side. In addition, we showed that garden beets are separated from the other cultivars. The distinct origin of garden beets from leaf and sugar beets confirms the results obtained with mitochondrial minisatellites and the RNA of beet cryptic virus (Hammer et al. 1990; Cheng et al. 2011). It is thought that garden beets were already domesticated in the twelfth century in Northern Italy and/or Southern Germany (Hammer et al. 1990), thus their domestication history would be distinct from that of modern sugar beets, which is thought to have started in the eighteenth century with the work of Franz Carl Achard (Fischer 1989). However, mitochondrial DNA restriction profiles indicated that the Owen cytoplasm found in all modern sugar beet lines was identical to that found in the garden beet Crapaudine variety, which suggests that garden beets have been unintentionally involved in the breeding of sugar beets by Owen (Bonavent et al. 1989). This can explain the small genetic distance between garden and sugar beets calculated from nuclear markers. The use of mitochondrial markers would be useful to clarify the relationships between the samples used in this study.
Concluding remarks
Beta section Beta contains the closest wild relatives of the sugar beet and it is the primary source of wild material for introgression of genetic variation into cultivated beets (Frese 2010). The analyses presented in this study clarified the relationships between the different taxa and cultivars forming the section, in particular the relationships between cultivated varieties and wild taxa, and the genetic differences among cultivars. Determining the genetic structure of the accessions is needed for detecting markers under selection in genomic-environmental association studies (Hoban et al. 2016). The case of section Beta is particularly complex due to the simultaneous effects of taxonomic and spatial structures on neutral genetic variation (Andrello et al. 2016). Limiting the search for genomic–environmental associations to a single taxon (such as B. vulgaris subsp. maritima, Manel et al. submitted) facilitates the analysis by simplifying the structure of neutral variation and decreases the rate of false positives. However, the geographical structure of B. vulgaris subsp. maritima shows complex patterns even after removing the taxonomic signal, because it is shaped by multiple factors, such as spatial distance, seed dispersal through marine currents, postglacial range expansion, and hybridization with crop plants (Santoni and Bervillé 1992c; Fievet et al. 2007; Arnaud et al. 2009; Leys et al. 2014; Andrello et al. 2016). Such a complex genetic structure will cause much difficulty in interpreting the results of genomic–environmental association studies in Beta section Beta.
A correct description of neutral genetic structure is also necessary to correctly estimate linkage disequilibrium patterns (Mangin et al. 2012), which are useful to reveal the selection history of genetic pools and to design genome-wide association studies for quantitative trait loci (QTL) (Würschum 2012; Adetunji et al. 2014; Mangin et al. 2015). Although the characterization of QTL is normally performed on elite lines of cultivated taxa (Würschum et al. 2011), the existence of untapped genetic variation in crop wild relatives (Warschefsky et al. 2014) may drive association studies towards wild accessions. In this case, the complex structure identified within B. vulgaris subsp. maritima is likely to affect estimates of linkage disequilibrium when all the accessions are analyzed together.
Data availability statement
All accessions including those of the present investigation have been obtained from germplasm banks worldwide under Material Transfer Agreement (MTA). They have been listed, described including from where they originated at: https://urgi.versailles.inra.fr/siregal/siregal/accessionAction.do?collectionId=146.
Author contribution statement
KH, PD, DV, and BD collected the accessions, grew the plants and performed the molecular and bioinformatic work, MA conceived, designed and performed the statistical analyses and led the writing, all authors reviewed the manuscript.
References
Abe J, Tsuda C (1987) Genetic analysis for isozyme variation in the section Vulgares, genus Beta. Jpn J Breed 37:253–261
Abe J, Yoshikawa H, Tsuda C (1986) Reproductive barriers in sugar beet (Beta vulgaris) and its wild relatives of the section Vulgares, the genus Beta. J Fac Agric Hokkaido Univ Jpn 63:40–48
Adetunji I, Willems G, Tschoep H et al (2014) Genetic diversity and linkage disequilibrium analysis in elite sugar beet breeding lines and wild beet accessions. Theor Appl Genet 127:559–571. doi:10.1007/s00122-013-2239-x
Andrello M, Henry K, Devaux P et al (2016) Taxonomic, spatial and adaptive genetic variation of Beta section Beta. Theor Appl Genet 129:257–271. doi:10.1007/s00122-015-2625-7
Arnaud J-F, Viard F, Delescluse M, Cuguen J (2003) Evidence for gene flow via seed dispersal from crop to wild relatives in Beta vulgaris (Chenopodiaceae): consequences for the release of genetically modified crop species with weedy lineages. Proc R Soc Lond B Biol Sci 270:1565–1571
Arnaud J-F, Fénart S, Godé C et al (2009) Fine-scale geographical structure of genetic diversity in inland wild beet populations. Mol Ecol 18:3201–3215. doi:10.1111/j.1365-294X.2009.04279.x
Bauchet G, Munos S, Sauvage C et al (2014) Genes involved in floral meristem in tomato exhibit drastically reduced genetic diversity and signature of selection. BMC Plant Biol 14:1
Biancardi E, Panella LW, Lewellen RT (2012) Beta maritima. The origin of beets. Springer, New York
Bonavent J-F, Bessone L, Geny A et al (1989) A possible origin for the sugar beet cytoplasmic male sterility source Owen. Genome 32:322–327. doi:10.1139/g89-448
Buttler KP (1977) Variation in wild populations of annual beet (Beta, Chenopodiaceae). Plant Syst Evol 128:123–136
Campbell LG (2010) Registration of seven sugarbeet germplasms selected from crosses between cultivated sugarbeet and wild species. J Plant Regist 4:149. doi:10.3198/jpr2009.11.0673crg
Cheng D, Kitazaki K, Xu D et al (2009) The distribution of normal and male-sterile cytoplasms in Chinese sugar-beet germplasm. Euphytica 165:345–351. doi:10.1007/s10681-008-9796-0
Cheng D, Yoshida Y, Kitazaki K et al (2011) Mitochondrial genome diversity in Beta vulgaris L. ssp. vulgaris (leaf and garden beet groups) and its implications concerning the dissemination of the crop. Genet Resour Crop Evol 58:553–560. doi:10.1007/s10722-010-9598-9
Curk F, Ancillo G, Ollitrault F et al (2015) Nuclear species-diagnostic SNP markers mined from 454 amplicon sequencing reveal admixture genomic structure of modern citrus varieties. PLoS One 10:e0125628
Desplanque B, Boudry P, Broomberg K et al (1999) Genetic diversity and gene flow between wild, cultivated and weedy forms of Beta vulgaris L. (Chenopodiaceae), assessed by RFLP and microsatellite markers. Theor Appl Genet 98:1194–1201
Dohm JC, Minoche AE, Holtgräwe D et al (2014) The genome of the recently domesticated crop plant sugar beet (Beta vulgaris). Nature 505:546–549. doi:10.1038/nature12817
Draycott AP (ed) (2006) Sugar beet. Blackwell, Oxford
Ecke W, Michaelis G (1990) Comparison of chloroplast and mitochondrial DNA from five morphologically distinct Beta vulgaris cultivars: sugar beet, fodder beet, beet root, foliage beet, and Swiss chard. TAG Theor Appl Genet Theor Angew Genet 79:440–442. doi:10.1007/BF00226149
El Mousadik A, Petit RJ (1996) High level of genetic differentiation for allelic richness among populations of the argan tree (Argania spinosa (L.) Skeels) endemic to Morocco. Theor Appl Genet 92:832–839. doi:10.1007/BF00221895
Excoffier L, Dupanloup I, Huerta-Sánchez E et al (2013) Robust demographic inference from genomic and SNP data. PLoS Genet 9:e1003905. doi:10.1371/journal.pgen.1003905
Fénart S, Arnaud J-F, De Cauwer I, Cuguen J (2008) Nuclear and cytoplasmic genetic diversity in weed beet and sugar beet accessions compared to wild relatives: new insights into the genetic relationships within the Beta vulgaris complex species. Theor Appl Genet 116:1063–1077. doi:10.1007/s00122-008-0735-1
Fievet V, Touzet P, Arnaud J-F, Cuguen J (2007) Spatial analysis of nuclear and cytoplasmic DNA diversity in wild sea beet (Beta vulgaris ssp. maritima) populations: do marine currents shape the genetic structure? Mol Ecol 16:1847–1864. doi:10.1111/j.1365-294X.2006.03208.x
Fischer HE (1989) Origin of the “Weisse Schlesische Rübe” (white Silesian beet) and resynthesis of sugar beet. Euphytica 41:75–80. doi:10.1007/BF00022414
Frese L (2010) Conservation and Access to Sugarbeet Germplasm. Sugar Tech 12:207–219. doi:10.1007/s12355-010-0054-0
Frese L, Nachtigall M, Enders M, Pinheiro De Carvalho MAA (2012) Beta patula (Aiton): Genetic diversity analysis. In: Maxted N (ed) \Agrobiodiversity conservation: securing the diversity of crop wild relatives and landraces. CABI, Wallingford
Fritzsche K, Metzlaff M, Melzer R, Hagemann R (1987) Comparative restriction endonuclease analysis and molecular cloning of plastid DNAs from wild species and cultivated varieties of the genus Beta (L.). Theor Appl Genet 74:589–594. doi:10.1007/BF00288857
Hammer K, Stanarius A, Kühne T (1990) Differential occurrence of beet cryptic viruses—a new tool for germplasm characterization and evolutionary studies in beets? Euphytica 45:23–27. doi:10.1007/BF00032145
Hjerdin A, Sall T, Nilsson NO et al (1994) Genetic variation among wild and cultivated beets of the section Beta as revealed by RFLP analysis. J Sugar Beet Res 31:59–67
Hoban S, Kelley JL, Lotterhos KE et al (2016) Finding the genomic basis of local adaptation: pitfalls, practical solutions, and future directions. Am Nat 188:379–397. doi:10.1086/688018
Hyten DL, Song Q, Zhu Y et al (2006) Impacts of genetic bottlenecks on soybean genome diversity. Proc Natl Acad Sci 103:16666–16671. doi:10.1073/pnas.0604379103
Jombart T, Ahmed I (2011) Adegenet 1.3-1: new tools for the analysis of genome-wide SNP data. Bioinformatics. doi:10.1093/bioinformatics/btr521
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11:94
Jones PD, Lister DH, Jaggard KW, Pidgeon JD (2003) Future climate impact on the productivity of sugar beet (Beta vulgaris L.) in Europe. Clim Change 58:93–108
Jung C, Pillen K, Frese L et al (1993) Phylogenetic relationships between cultivated and wild species of the genus Beta revealed by DNA “fingerprinting”. Theor Appl Genet 86:449–457. doi:10.1007/BF00838560
Kadereit G, Hohmann S, Kadereit JW (2006) A synopsis of Chenopodiaceae subfam. Betoideae and notes on the taxonomy of Beta. Willdenowia 36:9–19. doi:10.3372/wi.36.36101
Kilian B, Graner A (2012) NGS technologies for analyzing germplasm diversity in genebanks. Brief Funct Genom 11:38–50. doi:10.1093/bfgp/elr046
Kishima Y, Mikami T, Hirai A et al (1987) Beta chloroplast genomes: analysis of fraction I protein and chloroplast DNA variation. Theor Appl Genet. doi:10.1007/BF00262497
Letschert JPW (1993) Beta section Beta: biogeographical patterns of variation, and taxonomy. Wagening Agric Univ Pap 93(1):1–155
Leys M, Petit EJ, El-Bahloul Y et al (2014) Spatial genetic structure in Beta vulgaris subsp. maritima and Beta macrocarpa reveals the effect of contrasting mating system, influence of marine currents, and footprints of postglacial recolonization routes. Ecol Evol 4:1828–1852. doi:10.1002/ece3.1061
Litwiniec A, Gośka M, Choińska B et al (2016) Evaluation of rhizomania-resistance segregating sequences and overall genetic diversity pattern among selected accessions of Beta and Patellifolia. Potential implications of breeding for genetic bottlenecks in terms of rhizomania resistance. Euphytica 207:685–706. doi:10.1007/s10681-015-1570-5
Lu Y, Yan J, Guimarães CT et al (2009) Molecular characterization of global maize breeding germplasm based on genome-wide single nucleotide polymorphisms. Theor Appl Genet 120:93–115. doi:10.1007/s00122-009-1162-7
Mangin B, Siberchicot A, Nicolas S et al (2012) Novel measures of linkage disequilibrium that correct the bias due to population structure and relatedness. Heredity 108:285–291. doi:10.1038/hdy.2011.73
Mangin B, Sandron F, Henry K et al (2015) Breeding patterns and cultivated beets origins by genetic diversity and linkage disequilibrium analyses. Theor Appl Genet 128:2255–2271. doi:10.1007/s00122-015-2582-1
Mason AS, Zhang J, Tollenaere R et al (2015) High-throughput genotyping for species identification and diversity assessment in germplasm collections. Mol Ecol Resour 15:1091–1101. doi:10.1111/1755-0998.12379
Maxted N, Ford-Lloyd BV, Jury S et al (2006) Towards a definition of a crop wild relative. Biodivers Conserv 15:2673–2685. doi:10.1007/s10531-005-5409-6
McGrath JM, Derrico CA, Yu Y (1999) Genetic diversity in selected, historical US sugarbeet germplasm and Beta vulgaris ssp. maritima. Theor Appl Genet 98:968–976
Mikami T, Kishima Y, Sugiura M, Kinoshita T (1984) Chloroplast DNA diversity in the cytoplasms of sugar beet and its related species. Plant Sci Lett 36:231–235. doi:10.1016/0304-4211(84)90174-3
Mikami T, Yamamoto MP, Matsuhira H et al (2011) Molecular basis of cytoplasmic male sterility in beets: an overview. Plant Genet Resour 9:284–287. doi:10.1017/S1479262111000177
Mita G, Dani M, Casciari P et al (1991) Assessment of the degree of genetic variation in beet based on RFLP analysis and the taxonomy of Beta. Euphytica 55:1–6. doi:10.1007/BF00022552
Monteiro F, Romeiras MM, Batista D, Duarte MC (2013) Biodiversity assessment of sugar beet species and its wild relatives: linking ecological data with new genetic approaches. Am J Plant Sci 04:21–34. doi:10.4236/ajps.2013.48A003
Morin PA, Luikart G, Wayne RK, The SNP workshop group (2004) SNPs in ecology, evolution and conservation. Trends Ecol Evol 19:208–216. doi:10.1016/j.tree.2004.01.009
Nagamine T, Catty JP, Ford-Lloyd BV (1989) Phenotypic polymorphism and allele differentiation of isozymes in fodder beet, multigerm sugar beet and monogerm sugar beet. Theor Appl Genet 77:711–720
Ober ES, Clark CJA, Bloa ML et al (2004) Assessing the genetic resources to improve drought tolerance in sugar beet: agronomic traits of diverse genotypes under droughted and irrigated conditions. Field Crops Res 90:213–234. doi:10.1016/j.fcr.2004.03.004
Rong J, Lammers Y, Strasburg JL et al (2014) New insights into domestication of carrot from root transcriptome analyses. BMC Genom 15:1
Saccomani M, Stevanato P, Trebbi D et al (2009) Molecular and morpho-physiological characterization of sea, ruderal and cultivated beets. Euphytica 169:19–29. doi:10.1007/s10681-009-9888-5
Santoni S, Bervillé A (1992a) Characterization of the nuclear ribosomal DNA units and phylogeny of Beta L. wild forms and cultivated beets. Theor Appl Genet 83:533–542
Santoni S, Bervillé A (1992b) Two different satellite DNAs in Beta vulgaris L.: evolution, quantification and distribution in the genus. Theor Appl Genet 84:1009–1016
Santoni S, Bervillé A (1992c) Evidence for gene exchanges between sugar beet (Beta vulgaris L.) and wild beets: consequences for transgenic sugar beets. Plant Mol Biol 20:578–580
Santoni S, Faivre-Rampant P, Moreau E, Bervillé A (1991) Rapid control of purity for the cytoplasm of male-sterile seed stocks by means of a dot hybridization assay. Mol Cell Probes 5:1–9
Schiffels S, Durbin R (2014) Inferring human population size and separation history from multiple genome sequences. Nat Genet 46:919–925. doi:10.1038/ng.3015
Schoville SD, Bonin A, François O et al (2012) Adaptive genetic variation on the landscape: methods and cases. Annu Rev Ecol Evol Syst 43:23–43. doi:10.1146/annurev-ecolsys-110411-160248
Senda M, Onodera Y, Mikami T (1998) Cytoplasmic diversity in leaf beet cultivars as revealed by mitochondrial DNA analysis. Hereditas 128:127–132
Shen Y, Ford-Lloyd BV, Newbury J (1998) Genetic relationships within the genus Beta determined using both PCR-based marker and DNA sequencing techniques. Heredity 80:624–632
Shun ZF, Chu SY, Frese L (2000) Study on the relationship between Chinese and East Mediterranean Beta vulgaris L. subsp. vulgaris (leaf beet group) accessions. In: Maggioni L, Frese L, Germeier C, Lipman E (eds) Report of a working group on Beta. International Plant Genetic Resources Institute, Rome, pp 65–69
Stevanato P, Trebbi D, Biancardi E et al (2013) Evaluation of genetic diversity and root traits of sea beet accessions of the Adriatic Sea coast. Euphytica 189:135–146. doi:10.1007/s10681-012-0775-0
Thermo Fisher Scientific Inc. (2017) AxiomTM Genotyping Solution Data Analysis Guide. http://tools.thermofisher.com/content/sfs/manuals/axiom_genotyping_solution_analysis_guide.pdf. Accessed 23 March 2017
Tian H-L, Wang F-G, Zhao J-R et al (2015) Development of maizeSNP3072, a high-throughput compatible SNP array, for DNA fingerprinting identification of Chinese maize varieties. Mol Breed. doi:10.1007/s11032-015-0335-0
van Geyt JPC, Lange W, Oleo M, De Bock TSM (1990) Natural variation within the genus Beta and its possible use for breeding sugar beet : a review. Euphytica 49:57–76
Viard F, Arnaud J-F, Delescluse M, Cuguen J (2004) Tracing back seed and pollen flow within the crop-wild Beta vulgaris complex: genetic distinctiveness vs. hot spots of hybridization over a regional scale. Mol Ecol 13:1357–1364. doi:10.1111/j.1365-294X.2004.02150.x
Villain S (2007) Histoire évolutive de la section Beta: mise en évidence des phénomènes d’hybridation et de spéciation au sein de la Section dans le bassin Méditerranéen. Ph.D., Université Lille 1-Sciences et Technologies
Wang M, Goldman IL (1999) Genetic distance and diversity in table beet and sugar beet accessions measured by randomly amplified polymorphic DNA. J Am Soc Hortic Sci 124:630–635
Warschefsky E, Penmetsa RV, Cook DR, von Wettberg EJB (2014) Back to the wilds: tapping evolutionary adaptations for resilient crops through systematic hybridization with crop wild relatives. Am J Bot 101:1791–1800. doi:10.3732/ajb.1400116
Weihe A, Dudareva NA, Veprev SG et al (1991) Molecular characterization of mitochondrial DNA of different subtypes of male-sterile cytoplasms of the sugar beet Beta vulgaris L. Theor Appl Genet 82:11–16. doi:10.1007/BF00231271
Winkler LR, Michael Bonman J, Chao S et al (2016) Population structure and genotype–phenotype associations in a collection of oat landraces and historic cultivars. Front Plant Sci. doi:10.3389/fpls.2016.01077
Würschum T (2012) Mapping QTL for agronomic traits in breeding populations. Theor Appl Genet 125:201–210. doi:10.1007/s00122-012-1887-6
Würschum T, Maurer HP, Kraft T et al (2011) Genome-wide association mapping of agronomic traits in sugar beet. Theor Appl Genet 123:1121–1131. doi:10.1007/s00122-011-1653-1
Yoshida Y, Matsunaga M, Cheng D et al (2012) Mitochondrial minisatellite polymorphisms in fodder and sugar beets reveal genetic bottlenecks associated with domestication. Biol Plant 56:369–372. doi:10.1007/s10535-012-0101-7
Acknowledgements
This study was funded by the French Government, under the management of the Research National Agency (ANR-11-BTBR-0007) through the AKER program (included in the “Investissements d’avenir”).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Compliance with ethical standards
Not applicable.
Additional information
Communicated by André J. Bervillé.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Andrello, M., Henry, K., Devaux, P. et al. Insights into the genetic relationships among plants of Beta section Beta using SNP markers. Theor Appl Genet 130, 1857–1866 (2017). https://doi.org/10.1007/s00122-017-2929-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-017-2929-x