Abstract
In barley α-amylase 2, two and three tryptophans are protected against reaction with dimethyl(2-hydroxy-5-nitrobenzyl)sulphonium bromide by β-cyclodextrin and the pseudooligosaccharide inhibitor aplanin, respectively.
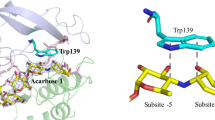
Fragments were generated from the enzyme derivatives by digestion with Armillaria mellea protease and trypsin, and isolated by RP-HPLC. The substituted tryptophans were identified by amino acid and sequence analyses of modified peptides. Aplanin and β-cyclodextrin both reduced the accessibility of Trp276 and −277. In addition, aplanin hindered modification of Trp206, and only this derivative retained activity. Trp206 probably belongs to the active site region, whereas Trp276 and −277 are located in a different binding site. This suggestion is supported by a comparison with the 3-D structure of Taka-amylase A guided by sequence homology between it and barley α-amylase.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Abbreviations
- HNB:
-
dimethyl(2-hydroxy-5-nitrobenzyl)sulphonium bromide
- NBS:
-
N-bromosuccinimide
- PTH:
-
phenylthiohydantoin
- RP-HPLC:
-
reverse phase high performance liquid chromatography
References
Barman, T. E. &D. E. Koshland, Jr.: A colorimetric procedure for the quantitative determination of tryptophan residues in proteins. J. Biol. Chem. 242, 5771–5776 (1967)
Brown, A. H. D. &J. V. Jacobsen: Genetic basis and natural variation of α-amylase isozymes in barley. Genet. Res. Camb. 40, 315–324 (1982)
Chandler, P. M., J. A. Zwar, J. V. Jacobsen, T. J. V. Higgins &A. S. Inglis: The effects of giberellic acid and abscisic acid on α-amylase mRNA levels in barley aleurone layers studies using an α-amylase cDNA clone. Plant Mol. Biol. 3, 407–418 (1984)
Clarke, A. J. &B. Svensson: The role of tryptophanyl residues in the function of Aspergillus niger glucoamylase G1 and G2. Carlsberg Res. Commun. 49, 111–122 (1984)
Clarke, A. J. &B. Svensson: Identification of an essential tryptophanyl residue in the primary structure of glucoamylase G2 from Aspergillus niger. Carlsberg Res. Commun. 49, 559–566 (1984)
Cohen, P., A. Sutter, G. Landreth, A. Zimmermann &E. M. Shooter: Oxidation of tryptophan-21 alters the biological activity and receptor binding characteristics of mouse nerve growth factor. J. Biol. Chem. 255, 2949–2954 (1980)
Dopheide, T. A. A. &W. M. Jones: Studies on the tryptophan residues in porcine pepsin. J. Biol. Chem. 243, 3906–3911 (1968)
Fletterick, R. J., J. Sygusch, H. Semple &N. B. Madsen: Structure of glycogen phosphorylase a at 3.0 Å resolution and its ligand binding sites at 6 Å. J. Biol. Chem. 251, 6142–6146 (1976)
Friedberg, F.: On the primary structure of amylases. FEBS Lett. 152, 139–140 (1983)
Gibson, R. M. &B. Svensson: Chemical modification of barley malt α-amylase 2: Involvement of tryptophan and tyrosine residues in enzyme activity. Carlsberg Res. Commun. 51, 295–308 (1986)
Horton, H. R. &W. P. Tucker: Dimethyl(2-hydroxy-5-nitrobenzyl)sulphonium salts. Water-soluble environmentally sensitive protein reagents. J. Biol. Chem. 245, 3397–3401 (1970)
Johansen, J. T., C. Overballe-Petersen, B. Martin, V. Hasemann &I. Svendsen: The complete amino acid sequence of copper,zinc superoxide dismutase from Saccharomyces cerevisiae. Carlsberg Res. Commun. 44, 201–217 (1979)
Kita, Y., M. Fukazawa, Y. Nitta &T. Watanabe: Kinetic study on chemical modification of Takaamylase A. I. Location and role of tryptophan residues. J. Biochem. 92, 653–659 (1982)
Kochhar, S. &R. D. Dua: An active centre tryptophan residue in liquifying α-amylase from Bacillus amyloliquefaciens. Biochem. Biophys. Res. Commun. 126, 966–973 (1985)
Lewis, W. G., J. M. Basford &P. L. Walton: Specificity and inhibition studies of Armillaria mellea protease. Biochim. Biophys. Acta 522, 551–560 (1978)
Loyter, A. &M. Schramm: Multimolecular complexes of α-amylase with glycogen limit dextrin. J. Biol. Chem. 241, 2611–2617 (1966)
Lundblad, R. L. &C. M. Noyes: Observations on the reaction of 2-hydroxy-5-nitrobenzyl bromide with a peptide-bound tryptophanyl residue. Anal. Biochem. 136, 93–100 (1984)
MacGregor, E. A. &A. W. MacGregor: The action of cereal α-amylases on solubilized starch and cereal starch granules. In: New Approaches to Research on Cereal Carbohydrates. R. D. Hill & L. Munck, ed., Elsevier, Amsterdam, 149–160 (1985)
Matsuura, Y., M. Kusunoki, W. Harada &M. Kakudo: Structure and possible catalytic residues of Taka-amylase A. J. Biochem. 95, 697–702 (1984)
Nakajima, R. T. Imanaka &S. Aiba: Comparison of amino acid sequences of eleven different α-amylases. Appl. Microbiol. Biotechnol. 23, 355–360 (1986)
Nitta, Y., T. Kunikata &T. Watanabe: Difference spectroscopic study of the interaction between soybean β-amylase and substrate or substrate analogues. J. Biochem. 93, 1195–1201 (1983)
Ohnishi, M. &K. Hiromi: Studies on the subsites of amylases. IV. Tryptophan residues of glucoamylase from Rhizopus niveus studied by chemical modification with N-bromosuccinimide. J. Biochem. 79, 11–16 (1976)
Payan, F., R. Haser, M. Pierrot, M. Frey, J. P. Astier, B. Abadie, E. Duee &G. Buisson: The three-dimensional structure of α-amylase from porcine pancreas at 5 Å resolution—the active site location. Acta. Cryst. B36, 416–421, (1980)
Quiocho, F. A.: Carbohydrate-binding proteins: Tertiary structures and protein sugar interactions. Ann. Rev. Biochem. 55, 287–315 (1986)
Robinson, G. W.: Reaction of a specific tryptophan residue in streptococcal proteinase with 2-hydroxy-5-nitrobenzyl bromide. J. Biol. Chem. 245, 4832–4841 (1970)
Rogers, J. C.: Two barley malt α-amylase gene families are regulated differently in aleurone cells. J. Biol. Chem. 260, 3731–3738 (1985)
Rogers, J. C.: Conserved amino acid sequence domains in alpha-amylases from plants, mammals, and bacteria. Biochem. Biophys. Res. Commun. 128, 470–476 (1985)
Sansom, M. S. P., D. I. Stuart, K. R. Acharya, J. Hajdu, P. J. McLaughlin &L. N. Johnson: Glycogen phosphorylase b.—The molecular anatomy of a large regulatory enzyme. J. Mol. Struc. 123, 3–25 (1985)
Silvanovich, M. P. &R. D. Hill: Affinity chromatography of cereal α-amylase. Anal. Biochem. 73, 430–433 (1976)
Spande, T. F. &B. Witkop: Determination of the tryptophan content of proteins with N-bromosuccinimide. Meth. Enzymol. XI, 489–506 (1967)
Svensson, B.: Regional distant sequence homology between amylases, α-glucosidases, and transglucanosylases. (Submitted)
Svensson, B., A. J. Clarke &I. Svendsen: Influence of acarbose and maltose on the reactivity of individual tryptophanyl residues in glucoamylase from A. niger. Carlsberg Res. Commun. 51, 61–73 (1986)
Svensson, B., R. M. Gibson, R. Haser &J. P. Astier: Crystallization of barley malt α-amylases and preliminary x-ray diffraction studies of the high-pI isoenzyme, α-amylase 2. J. Biol. Chem. 262, 13682–13684 (1987)
Svensson, B., K. Larsen &I. Svendsen: Amino acid sequence of tryptic fragments of glucoamylase G1 from Aspergillus niger. Carlsberg Res. Commun. 48, 517–527 (1983)
Svensson, B., J. Mundy, R. M. Gibson &I. Svendsen: Partial amino acid sequences of α-amylase isozymes from barley malt. Carlsberg Res. Commun. 50, 15–22 (1985)
Truscheit, E., W. Frommer, B. Junge, L. Müller, D. D. Schmidt &W. Wingender: Chemie und Biochemie mikrobieller α-Glucosidaseinhibitoren. Angew. Chem. 93, 738–755 (1981)
Ueda, S.: Fungal glucoamylases and raw starch digestion. Trends Biochem. Sci. 6, 89–90 (March, 1981)
Weselake, R. J. &R. D. Hill: Cycloheptaamylose as an affinity ligand of cereal alpha-amylases. Characteristics and a possible mechanism of interaction. Carbohydr. Res. 108, 153–161 (1982)
Weselake, R. J. &R. D. Hill: Inhibition of alpha-amylase-catalysed starch granule hydrolysis by cycloheptaamylose. Cereal Chem. 60, 98–101 (1983)
Author information
Authors and Affiliations
Additional information
Accepted byH. Klenow
Rights and permissions
About this article
Cite this article
Gibson, R.M., Svensson, B. Identification of tryptophanyl residues involved in binding of carbohydrate ligands to barley α-amylase 2. Carlsberg Res. Commun. 52, 373–379 (1987). https://doi.org/10.1007/BF02907525
Issue Date:
DOI: https://doi.org/10.1007/BF02907525