Abstract
The ever-increasing demand for food production proportionate to exponential growth of global population evoked the need for applying innovative techniques for developing disease-resistant crop varieties, as the pest and pathogen attack causes considerable yield loss which in turn putts the agriculture sector and crop production in crisis. This is highly significant when the resistance conferred by conventional measures, like artificial hybridization, mutation breeding, marker-assisted selection, etc. appears to be inadequate in many cases, especially due to the evolution of new/more virulent strains/pathovars/isolates of pathogens and their unexpected host range expansion. Development of recombinant DNA technology, transgenic expression and RNA silencing strategies lead to a new era of transgenically engineered resistance in several crop species, many of which succeeded in field trials and got commercialized. Elucidation of genetic and molecular mechanisms underlying host-pathogen interactions, resistance, susceptibility and different levels of plant immune responses (PTI, ETI, HR, etc.) revealed the key genes in the host as well as pathogens that can be manipulated by transgenic approaches/techniques for conferring effective resistance. Reprogramming of phytohormonal regulatory pathways determining defence response and remodeling of molecular receptors/transcription factors involved in resistance or disease development can be brought about by transgenic methods to enhance the resistance in host plants. Targeting of conserved sequences or molecular components could provide broad-spectrum resistance in certain cases. Mining of R genes, transgenic expression of foreign R gene, etc. are other useful strategies. Potential of genome editing based on engineered nucleases like ZFNs, TALENs and CRISPR/Cas9 to precisely mutate the genomic sequence of interest can be exploited for specific targeting of the host/pathogenic genes associated with the biotic stress response. Application of all these approaches for the management of plant pathogens is discussed in this chapter.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
23.1 Introduction
The ever increasing demand for food production proportionate to exponential growth of global population evoked the need for applying innovative techniques for developing disease-resistant crop varieties, as the pest and pathogen attack causes considerable yield loss which in turn putting the agriculture sector and crop production in crisis. This is highly significant when the resistance conferred by conventional measures, like artificial hybridization, mutation breeding, marker-assisted selection, etc. appears to be inadequate in many cases, especially due to the evolution of new/more virulent strains/ pathovars/ isolates of pathogens and their unexpected host range expansion. Development of recombinant DNA technology, transgenic expression, and RNA silencing strategies lead to a new era of transgenically engineered resistance in several crop species, many of which succeeded in field trials and got commercialized. Elucidation of genetic and molecular mechanisms underlying host-pathogen interactions, resistance, susceptibility, and different levels of plant immune responses (PTI, ETI, HR, etc.) revealed the key genes in the host as well as pathogens that can be manipulated by transgenic approaches/ techniques for conferring effective resistance. Reprogramming of phytohormonal regulatory pathways determining defense response and remodeling of molecular receptors/ transcription factors involved in resistance or disease development can be brought about by transgenic methods to enhance the resistance in host plants. Targeting of conserved sequences or molecular components could provide broad-spectrum resistance in certain cases. Mining of R genes, transgenic expression of foreign R gene, etc. is another useful strategy. Potential of genome editing based on engineered nucleases like ZFNs, TALENs, and CRISPR/ Cas9 to precisely mutate the genomic sequence of interest can be exploited for specific targeting of the host/ pathogenic genes associated with the biotic stress response. Application of all these approaches for the management of plant pathogens were discussed in this chapter.
During the earlier times, resistant varieties were created by artificial hybridization with wild-type resistant varieties, which was purely depended on phenotypic selection. Diminishing genetic variability and lack of resistant germplasm limited the possibilities of resistant gene introgression via conventional breeding. Moreover, it is time-consuming and associated linkage drag may result in introgression of undesirable traits too. Mutation breeding was an alternative to this, where chemical mutagens are used for inducing mutation in target genes to create desirable phenotype. Resistance to powdery mildew has been successfully engineered in this way by mutating Mlo locus. As mutagenesis is random, the chances of off-target mutations is high. QTL mapping and marker-associated selection and breeding are other ways to create disease-resistant varieties but require large mapping population and are time-consuming and labour-intensive as in the case of hybridization. Even though these conventional strategies enabled crop protection to a great extent, over the centuries, rapid genetic variation and adaptive evolution of pathogens engendered for novel, robust and sustainable methods for controlling pathogens and pest management.
The development of genetic engineering (GE) technology for designing tailor-made plants by transforming plant with the desired gene from other organism was a major breakthrough in crop improvement programs. Initially, transgenic experiments were confined to model plants merely for demonstration and proof-of-concept purpose; later, species wide dispersal of technology was made possible by the development of optimum transformation and regeneration protocols, and this widened the applicability of GE for many economically important crops. Since disease resistance is an important yield determinant, transgenic technologies have given considerable focus on developing/reinforcing resistance in economically important crop species. Crop improvement program for disease resistance gained a significant progress by deploying various GE techniques. Rapid introgression of promising candidate genes, even into phylogenetically unrelated species, precise alteration or modification of host factors involved in the perception of pathogenic effectors and deployment of defence signals, metabolic pathways and biomolecules involved in direct defence responses as well as targeted by pathogen effectors have been successfully implemented using transgenic strategies. Similarly, multitude of pathogenic genes and effectors have also been manipulated transgenically, in order to reduce virulence and pathogenicity. Furthermore, transgenic strategies permit precise spatio-temporal regulation of trait of interest. The emergence of new breeding techniques (NBT), including targeted mutagenesis and precision breeding, further revolutionized crop improvement strategies. Genome editing tools, like ZFNs, TALENs and CRISPR/Cas9, based on site-directed nucleases enabled precise editing of targeted genomic loci to confer/modify resistance traits in many of the crop plant species. Altogether transgenic strategies facilitated the fine-tuning of the plant immune system for ensuring sustainable production. In this chapter, we will describe about the strategies adopted for creating resistance in transgenic plants against virus, bacteria and fungi and at then the upcoming strategies based on the information obtain from plant–pathogen interactions.
23.2 Transgenic Technology for Virus Resistance
Viruses are notorious plant pathogens causing devastating damage to crop yield. These obligate parasites invade, replicate and proliferate in host cell and result in infection, which reduces fitness and productivity of crop plants and decreases market and aesthetic values of the products. Attributes like great evolvability, large population sizes, error-prone replication and efficient host-dependence render the control of plant viruses extremely difficult. Development of sustained resistance against pathogenic virus, broad-spectrum as well as durable, throughout the productive stage while infection pressure persists is always a challenge in crop improvement. Farmers rely on combining traditional cultural management practices, such as field sanitation, crop rotation, planting of trap plants, spraying for vector, rouging and manual removal of infected plants upon detection of disease symptoms, use of certified virus-free seeds or planting materials (Kamala and Makeshkumar 2015; Deepthi and Makeshkumar 2016), as there is no specific direct control strategy; even chemical pesticides are not available. All these measures are laborious and time-consuming. Chemical control of vectors, in addition, causes health and environmental hazards. The most efficient practical solution available is the use of resistant varieties. But elite cultivars might not always be endowed with resistance traits, which further make the problem complicated and uneconomical. Even though resistance genes can be introgressed from wild varieties, either by conventional breeding or marker-assisted selection, linkage drag and other inherent shortcomings of these techniques render it inadequate for successful crop improvement. Resistance genes in unrelated species or sometimes related species cannot be introgressed by hybridization due to barriers, like sexual incompatibility, male sterility, etc. Similarly, attaining broad-spectrum resistance requires stacking of resistance genes from different sources, which is again a laborious and less efficient strategy. Moreover, virus resistance conferred by R genes is less durable due to the suppression of R–Avr interaction by creating variant Avr protein, which might be unrecognizable by the host R protein, by quickly evolving viral genome, rendering the host susceptible for compatible host-pathogen interaction. However, transgenic approaches have succeeded to a great extend in engineering efficient virus resistance in crop species.
Recessive resistance, conferred by mutating host factors required for infection cycle completion and thereby making the host non-permissive to virus, is more efficient and durable strategy. Resistance thus conferred is passive as there is no active involvement of plant immune system. Resistance mechanism, to be adopted, and respective targets vary with stage of infection (Johnson 1981).
Transgenic antiviral approaches used so far were based on the expression of various viral proteins; RNAs; nonviral genes like nucleases, antiviral inhibitors and plantibodies; plant defence response elicitors; host-derived resistance genes (dominant resistance genes and recessive resistance genes); and various factors involved in host defence responses. Viral proteins usually used for engineering resistance are capsid protein, replicase proteins and movement proteins. Several RNA molecules, viz. sense RNAs, antisense RNAs, satellite RNAs, defective interfering RNAs, hairpin RNAs and artificial microRNAs, noncoding RNAs, antisense RNAs, ribozymes, double-stranded RNAs (dsRNAs) and inverted repeat RNAs (irRNAs), have been employed for conferring virus resistance by post-transcriptional or transcriptional gene silencing. Transgenic virus resistance strategies are deployed principally by three mechanisms: pathogen-derived resistance, pathogen-targeted resistance and RNA interference.
23.2.1 Pathogen-Derived Resistance
Pathogen-derived resistance (PDR) is protein-mediated resistance conferred by viral protein expressed in host cells (Sanford and Johnston 1985). PDR is accomplished by different a mode of action, which varies with strains and stages of infection cycle, like whether it is in movement/transport/replication phase. It can be inhibition of replication and viral particle accumulation in the early stages of infection while limiting the spread via apoplastic/symplastic/phloem stream during the movement stage (Galvez et al. 2014). The first type of PDR is based on the silencing of pathogenic gene by expression of a part of that gene in host.
23.2.2 Coat Protein–Mediated Resistance (CPMR)
Since coat protein (CP) has implications in almost all stages of infection, like uncoating, systemic movement, long-distance transport, replication, symptom development, etc., it can be the ideal candidate for engineering PDR, even in non-host plants (Galvez et al. 2014). CPMR is manifested by expression of coat protein gene in the host cell and the subsequent interaction between transgenic CP and viral CP (Koo et al. 2004). The very first time application of CPMR was showed in tobacco (Nicotiana tabacum) plants, expressing the capsid protein-encoding sequences of Tobacco mosaic virus (TMV), and this resulted in partial resistance to TMV (Abel et al. 1986). Later it was expanded to tomato, using the same CP construct, with resistance to Tomato mosaic virus (ToMV) (Nelson et al. 1988). Similarly, capsid protein gene of a potyvirus was expressed in a non-host plant, tobacco, conferring resistance to other potyviruses. CPMR has been used to confer resistance to at least 35 viruses, representing more than 15 different taxonomic groups (Table 23.1). Several virus-resistant transgenic crop plants were developed by using a suitable coat protein gene (Mundembe et al. 2009; Nomura et al. 2004). CPMR has been successfully established in host plants, including potato, tomato, tobacco and papaya, exhibiting resistance to Potato virus Y (PVY) and Potato leaf roll virus (PLRV), Cucumber mosaic virus (CMV) and Papaya ring spot virus (PRSV) (Makeshkumar et al. 2002). The efficiency of CPMR varies for each virus with different stages of infection cycle (Bendahmane et al. 2007). The underlying molecular mechanism of resistance manifestation is either through recoating of invading viral particles or by blocking of receptors in transgenic plants (Saharan et al. 2016).
23.2.3 Replicase- or Rep-Associated Protein-Mediated Resistance
Replicase is another potential candidate to engineer resistance against viral genome. Expression of intact or truncated or mutant virus-encoded replicase in host can confer resistance to that virus. It was reported for the first time when a 54 kDa truncated protein of TMV replicase protein was expressed in N. benthamiana, where it conferred high level of resistance against TMV infection (Golemboski et al. 1990). This strategy has been extended to many food crops, including rice, bean, potato etc., and mostly resulted in narrow-spectrum resistance towards a particular race of the pathogen (Saharan et al. 2016) (Table 23.2). Rep-associated protein, which interacts with host DNA polymerase during replication of ssDNA virus, can also be manipulated in a similar manner. Resistance to two ssDNA viruses, Tomato yellow leaf curl virus-Israel (TYLCV-Is [Mild]) and Bean golden mosaic virus were conferred by transgenic expression of a truncated replication-associated protein gene and rep gene, respectively, in tomato and P. vulgaris (Brunetti et al. 1997; Faria et al. 2006). As in the case of CPMR, the active entity conferring resistance can either be protein or RNA or both.
23.2.4 Movement Protein–Mediated Resistance
Movement proteins (MP) facilitate the intracellular or cellular movement of viral particles through plasmodesmata, by modifying the gating channels of plasmodesmata. Resistance conferred by dysfunctional or mutated MP is mostly broad-spectrum in nature when compared to CP or replicase-mediated resistance (Prins et al. 2008). The first MP-mediated resistance was shown in transgenic tobacco plants expressing a 30 kDA mutant defective MP (dMP), which competed with the wild-type virus-encoded MP for the binding sites in the plasmodesmata and conferred resistance against eponymous virus infection. dMP conferred broad-spectrum resistance by preventing the systemic spread of distantly related and unrelated viruses (Lapidot et al. 1993; Cooper et al. 1995). Similar broad-spectrum resistance was observed in transgenic potato expressing wild-type Potato leafroll virus (PLRV) movement protein against PLRV, PVY and PVX (Tacke et al. 1996), while narrow-spectrum resistance was shown in transgenic N. benthamiana expressing wild-type movement proteins of Cowpea mosaic virus (CPMV) and transgenic tobacco expressing PVX movement protein (Sijen et al. 1995).
23.2.5 Other Viral Protein-Mediated Resistance
Transgenic expression of viral proteins other than those discussed previously, such as replication-associated protein, NIa protease, P1 protein and HC- Pro, has been tried out in order to achieve resistance against viruses (Cillo and Palukaitis 2014). Transgenic expression of partial or complete Rep gene was found to confer resistance against Tomato golden mosaic virus (TGMV), Tomato yellow leaf curl virus (TYLCV) (Yang et al. 2004; Antignus et al. 2004; Lucioli et al. 2003), African cassava mosaic virus (ACMV) (Chellappan et al. 2004), Bean golden mosaic virus (BGMV) (Faria et al. 2006), Maize streak virus (Shepherd et al. 2007), Cotton leaf curl virus (CLCuV) (Hashmi et al. 2011) and Tomato leaf curl Taiwan virus (ToLCTWV) (Lin et al. 2012) in transgenic plants.
Transgenic tobacco plants expressing the NIa protein of Tobacco vein mottling virus (TVMV) exhibited resistance against TVMV, whereas it failed to confer resistance to two other potyviruses, Tobacco etch virus (TEV) and PVY (Maiti et al. 1999). Transgenic tobacco lines expressing paired NIa protease coding sequences for two viruses, like TEV–PVY, TEV–TVMV and TVMV–PVY, were assessed for virus resistance and mostly resulted in the recovery-type resistance (Fellers et al. 1998). However, expression of multiple genes (NIa/NIb/capsid protein) from a single potyvirus using a single construct failed to confer any enhanced resistance in most cases (Maiti et al. 1999).
Transgenic expression of complete or partial sequence encoding P1 protein conferred resistance, of varying degree, against viruses like PVY, PPV, TVMV and PVA. A recovery-type resistance was obtained in most cases rather than complete resistance (Germundsson and Valkonen 2006). Expression P1 coding sequence of PVY-O and PPV, respectively, showed either complete resistance or a recovery-type resistance against PVY-O infection in potato and PPV infection in N. benthamiana (Maki-Valkama et al. 2001; Tavert-Roudet et al. 1998).
Expression of viral HC-Pro protein resulted in recovery phenotypes instead of conferring resistance to PVA and PPV in transgenic N. benthamiana and SMV in transgenic soybean (Savenkov and Valkonen 2002; Barajas et al. 2004; Lim et al. 2007). Expression level of transgene considerably influenced the resistance mediated by HC-Pro in most of the experiments, wherein low expression levels are mostly favoured for resistant or recovery phenotype rather than high expression levels. Deletion of central domain of HC-Pro protein has found to confer recover disease symptoms of Cowpea aphid-borne mosaic virus (CABMV) in transgenic N. benthamiana compared to intact protein (Mlotshwa et al. 2002).
Other viral genes, such as VPg-protease coding region of Tomato ringspot virus (ToRSV), capsid protein domain of BNYVV (Andika et al. 2005) and p23 silencing suppressor protein of Citrus tristeza virus (CTV) (Fagoaga et al. 2006), were also used as potential transgene candidates for providing resistance against ToRSV infection in N. benthamiana, olymyxa betae infection in transgenic N. benthamiana plants and CTV infection in transgenic Mexican lime plants, respectively (Table 23.3).
23.2.6 Viral RNA-Mediated Resistance
Although virus resistance has been successfully manifested in several crop species by transgenic expression of intact or dysfunctional or mutated structural proteins of virus, resistance in many plants has been found to be mediated by corresponding mRNAs rather than the encoded protein moieties (Saharan et al. 2016). The active role of RNA in conferring resistance was first revealed when an untranslatable coat protein gene in transgenic tobacco plants provided resistance against Tobacco etch virus (TEV) (Lindbo and Dougherty 1992a, b). Many cases of transgenic expression of untranslatable viral proteins further substantiated the direct participation of RNA in developing resistance. Later, it has been shown that the virus resistance can be RNA-mediated, and viral protein expression is not necessary for the same (Dougherty et al. 1994). Viral RNA-mediated resistance is carried out by transcriptional gene silencing (PTGS), RNA interference or RNAi or RNA silencing, where a transgenic viral RNA introduced into the host plant drives sequence-specific homology-dependent degradation of viral genomic RNA or viral mRNAs by employing the small interfering RNA (siRNA) pathway of host cell machinery (Voinnet 2008; Waterhouse et al. 2001; Lindbo and Dougherty 2005). This homology-dependent gene silencing mechanism is conserved among higher eukaryotes and operates for gene expression regulation and in host defence against transposable elements and viruses (Hannon 2002; Mawassi and Gera 2012).
23.2.7 RNA Interference: Mechanism
RNA interference is widely accepted as a potential gene silencing strategy that can be easily manipulated to obtain desirable trait in organism of interest. RNA silencing process requires a double-stranded RNA precursor to trigger the silencing process. This dsRNA precursor is derived from viral RNA intermediates or viral RNA secondary structures. After recognition of this dsRNA by the RNase III-like enzyme, namely, DICER, it is cleaved into 21–25 nucleotide duplexes, called as small interfering RNAs (siRNAs). The siRNAs thus formed are incorporated and converted to ssRNAs by Argonaut (Ago) containing multisubunit ribonuclease named as RNA-induced silencing complex (RISC). Subsequently RISC targets the specific mRNAs that share sequence similarity with siRNA through degradation of transcript (Waterhouse et al. 1998; Hamilton and Baulcombe 1999; Voinnet 2008).
RNA-mediated resistance engineering approaches include expression of non-coding regions of viral genome, viral CP mRNA, satellite (sat) RNA, defective interfering (DI) RNA and viral sequences in sense or antisense orientation or in double-stranded forms in host plants (Cillo and Palukaitis 2014).
Non-coding ssRNA
Viral non-coding RNAs like 5′ and 3′ NTRs, intergenomic regions and non-coding RNAs, either in sense or antisense orientation, have been used for engineering resistance in various crop species. Transgenic expression of 5′ and 3′ NTRs and intergenomic regions could successfully confer resistance in crop plants. Sometimes simultaneous expression of non-coding NTR regions with mRNAs also confer resistance. For instance, transgenic tobacco expressing 3′-NTR of Andean potato mottle virus (APMoV), which was expressed with a smaller portion of capsid protein-coding region, showed excellent resistance in several lines (Vaskin et al. 2001). Transgenic oilseed rape plant expressing 3′-NTR of Turnip yellow mosaic virus (Zaccomer et al. 1993) and intergenic region of PLRV expressing transgenic potato (Dong et al. 1999) showed resistance to respective pathogens.
Non-translatable Sense RNAs
Non-translatable sense RNAs of coat protein created by frameshift mutation have been widely used to confer resistance to a large number of crops, including tobacco, rice, papaya, peanut, grape, sugar cane, etc. Expression of frameshifted, capsid protein coding sequences of PVY-NTN in transgenic potato was shown to be resistant to PVYNTN infection (Rachman et al. 2001). There are some other viral proteins which can be suitable candidates to be manipulated in similar manner to engineer resistance. Potential of non-translatable coding sequences of movement protein, nucleoprotein, Vpg, NSm, etc. to mediate viral gene silencing has been proved through different experiments. Expression of non-translatable form of the capsid protein gene has been conferred resistance to Papaya leaf-distortion mosaic virus (PLDMV) (Kung et al. 2010), Zucchini yellow mosaic virus (ZYMV) and PRSV-W (Wu et al. 2010; Yu et al. 2011) and Sugarcane yellow leaf virus (Zhu et al. 2011). Transgenic tobacco expressing a non-translatable form of the NSM coding region of TSWV and a non-translatable form of the TEV 6-kDa/VPg encoding region exhibited resistance towards TSWV and TEV, respectively (Prins et al. 1996; 1997; Swaney et al. 1995). Moreover, hybrid constructs, designed by a fusion of non-translatable coding sequence of coat protein of different viruses, have found to be efficient for developing broad-spectrum resistance.
Antisense RNAs
Transformation of host plant with antisense RNAs of certain viral genes is a powerful strategy to silence invading viral genes. Transgenic plants expressing sense RNAs for viral replicase, transcription activator protein (TrAP), replication enhancer protein (Ren), AV1, etc. had been developed. Antisense RNA-mediated silencing mostly resulted in attenuated or delayed symptoms and complete resistance in some cases. Trials to engineer resistance to Tomato leaf curl virus (Praveen et al. 2005), African cassava mosaic virus (Zhang et al. 2005) and Mung bean yellow mosaic India virus (Singh et al. 2013) have been performed.
Satellite RNA
Some viruses exhibit a supernumerary RNA component that does not show any apparent homology to the viral RNA genome but depends on its helper virus for replication, encapsidation and transmission and is defined as a satellite RNA (Simon et al. 2004). These sequences have found to have a highly variable range of effects on various components like viral replication, pathogenesis and symptom expression in certain host-pathogen interactions. Sat variants with attenuating effects are regarded as potential biocontrol agents in transgenic plants. Transgenic expression of an attenuating CMV-associated satRNA suppressed viral replication and symptom development and thereby conferred tolerance to CMV in tobacco and other solanaceous plants including tomato (Baulcombe et al. 1986; Harrison et al. 1987; Kim et al. 1997; Kim et al. 1995)). Transgenic N. benthamiana and A. thaliana plants expressing Bamboo mosaic virus (BaMV) satRNA showed high resistance to helper virus (Lin et al. 2013).
Defective Interfering DNAs and RNAs
Defective DNAs and RNAs are produced during the replication of certain viral species, which can reduce the full-length genome accumulation and results in the denomination of DI nucleic acids, associated with the modulation of symptom expression, and thus serves as a source for transgenic resistance strategies. ssDNA geminiviruses, like ACMV, and ssRNA Tombusvirus, like Cymbidium ringspot virus (CymRSV), are known to form DIs. Interfered viral replication and milder symptoms were observed in transgenic N. benthamiana expressing naturally occurring subgenomic ACMV DNA B upon ACMV infection (Stanley et al. 1990). Tolerance to other viral species, including Beet curly top virus (BCTV) (Frischmuth and Stanley 1998; Stenger 1994), Cucumber necrosis virus (CNV), Carnation Italian ringspot virus and CymRSV (Kollàr et al. 1993; Rubio et al. 1999), has been obtained by employing DI DNA and DI RNA.
Silencing by Ds Inverted Repeat Sequence/hpRNA
Combined expression of sense and antisense transcripts in the same transgenic plants is advantageous for obtaining more transgenic lines exhibiting resistance. It is more efficient than the expression of either strand alone (Waterhouse et al. 1998; Smith et al. 2000; Waterhouse and Helliwell 2003). This was made possible by designing a single transcript of both polarities, with sense and antisense sequences separated by introns or other spacer sequences, to generate dsRNAs with loops, known as hpRNAs, intron hpRNAs (ihpRNAs) or irRNAs, which stabilize the inverted repeat DNA sequences in Escherichia coli (Smith et al. 2000; Wesley et al. 2001). dsRNA thus produced is then acted upon by siRNA pathway, resulting in virus resistance (Castel and Martienssen 2013; Csorba et al. 2009; Ding 2010; Eames et al. 2008; Pumplin and Voinnet 2013). Broad-spectrum resistance can be achieved by creating a chimeric, fused hpRNAs expressing sequences of several viruses from the same vector (Cillo and Palukaitis 2014). The application of hpRNAs as a transgenic resistance strategy was first demonstrated in transgenic tobacco and barley expressing a hpRNA specifically targeting NIa protease coding sequences of PVY and polymerase coding sequences of BYDV, respectively (Smith et al. 2000, Abbott et al. 2000). Development of viral resistance in crop plants was tremendously advantaged from the establishment of simple and efficient technique for stable integration of self-complementary hairpin construct, designed specifically for the cognate RNA target. dsRNA and siRNA are generated in host cell and silence the pathogen gene expression by cleaving target RNA. The major crop plants, like rice, maize, citrus, cassava, legumes, etc., have been conferred durable and efficient resistance using this method (Cillo and Palukaitis 2014). Although RNAi confers efficient and durable resistance, plant viruses have evolved smart measures, like suppressors of RNA-induced gene silencing, to escape RNAi which is a challenge before the advancements of transgenic resistance (Table 23.3).
Silencing by Artificial MicroRNAs
MicroRNAs are small (20–250 nt), non-coding RNAs present in eukaryotic cells and facilitate gene regulation at post-transcriptional or translational level (Bartel 2004). The precursor miRNA transcripts are processed into mature miRNA and directed to target mRNA by the same mechanism involving DICER, Ago and RISC, as that of siRNAs (Jones-Rhoades et al. 2006). Customized miRNA precursors producing target-specific siRNAs, which are having no similarity with mature endogenous miRNAs, known as artificial microRNAs (amiRNAs), can be transgenically expressed to target invading viral sequences. amiRNA-mediated virus resistance was first reported in transgenic Arabidopsis showing resistance against Turnip yellow mosaic virus (TYMV) and Turnip mosaic virus (TuMV) by stable expression of amiRNAs targeting the RNA sequences encoding silencing suppressors P69 and HC-Pro of the virus (Niu et al. 2006). amiRNAs targeting RNA silencing suppressors of virus have found to be more efficient in conferring resistance either complete resistance or delayed infection/susceptibility, when compared to those targeting CP or other structural proteins (van Vu et al. 2013). Silencing suppressors Hc-Pro and TGB1/p25 of PVY and PVX, respectively, were targeted efficiently by amiRNAs (Ai et al. 2011). amiRNAs have been demonstrated to confer resistance against a vast range of viruses, including positive-sense ssRNA genome such as CMV, PVY and PVX; negative-sense ssRNA viruses like WSMoV (genus Tospovirus); and ssDNA viruses of the genus Begomovirus like ToLCV New Delhi variant (van Vu et al. 2013) and Cotton leaf curl Burewala virus (Ali et al. 2013). Efficiency of amiRNA-induced viral gene silencing is determined by several factors, like expression levels of pre-miRNA backbone and sequence complementarity of amiRNA with target and structural features of target RNA and accessibility of amiRNA to target RNA (Ali et al. 2013; Cillo and Palukaitis 2014; Simón-Mateo and García 2011; Duan et al. 2008). Viruses may quickly evolve new strains with mutations in amiRNA target site and escape silencing, which can be circumvented by simultaneous targeting of multiple regions of highly conserved RNA motifs of viral genome with multiple amiRNAs. A polycistronic amiRNA designed from a modified rice miRNA395, targeting different conserved regions of the WSMV, conferred resistance to WSMV in wheat (Fahim et al. 2012).
Silencing by Co-suppression
Resistance to virus can be achieved by activating co-suppression by introducing surplus amount of viral transcripts in the host, where overabundance of sense strand induces downregulation or suppression of transgene and invaded viral gene. Such overabundance of sense strand leads to the removal of all homologous transcripts, beyond a critical threshold, by respective cellular machineries (Stam et al. 1997). Overexpressed transcripts, when recognized by plant RNA-dependent RNA polymerase, act as primers for dsRNA synthesis, which then subjected to cleavage and degradation by DICER and RISC complex (Dougherty and Parks 1995).
23.2.8 Transgenic Technology to Engineer Pathogen-Targeted Virus Resistance
PDR and R gene often do not confer complete resistance to invading viruses, and mostly resistance is not stable over several generations. Pathogen-targeted resistance is manifested by introducing silencing constructs into host plants, where it targets viral genome. Synthetic constructs designed to target viral sequences are neither of plant origin nor pathogen-derived. Such non-viral-mediated resistance is manifested by transgenic expression of synthetic nucleases like zinc finger nucleases, transcription activator like effector nucleases, CRISPR/Cas9, antiviral inhibitor proteins like ribosome-inhibiting proteins, peptide aptamers and plantibodies (Bastet et al. 2018).
Nucleases
Resistance against Fijivirus RBSDV and TMV was conferred by transgenic expression of an E. coli dsRNase gene (Cao et al. 2013), bovine pancreatic RNase (Trifonova et al. 2007) and an inducible extracellular RNase from Zinnia elegans (Trifonova et al. 2012). E. coli dsRNase (RNase III) conferred high-level resistance to a tomato isolate of TSWV also (Langenberg et al. 1997). Feasibility of transgenic expression of antiviral pathways or invading nucleic acid targeting pathways from heterologous systems in host plant for targeting infectious viruses has been demonstrated using OAS system 2,5A-oligoadenylate synthetase (OAS)/RNase L system, which is also called as 2,5A oligoadenylate pathway, an animal antiviral pathway induced by interferons in mammalian cells. The OAS catalyses the polymerization of ATP producing 2-5-linked oligoadenylates, pppA (2_p5_A) nor 2,5A, when it recognizes a dsRNA, which can be replicative intermediates of single-stranded RNA viruses or viral dsRNA genomes. The subsequent activation of the latent ribonuclease, RNAse L, by the 2,5A oligonucleotides produced by OAS, carries out the degradation of both viral and cellular RNAs, thereby hampering the viral replication and infection (Floyd-Smith et al. 1981). The efficacy of this RNA targeting mechanism to confer broad-spectrum resistance has been demonstrated in tobacco plants (Mitra et al. 1996). Resistance to several DNA and RNA viruses has been engineered employing ZFNs, TALENs and CRISPR/Cas9. Artificial TALE proteins and FokI nuclease expressing Nicotiana benthamiana plants showed resistance to different Begomoviruses. Transgenic expression of yeast-derived dsRNase, Pac1, confer broad-spectrum resistance to phytopathogenic viruses. Pac1 has been successfully used to confer resistance to CMV, Tomato mosaic virus (ToMV) and PVY in tobacco (Watanabe et al. 1995) and PSTVd in potato (Sano et al. 1997).
Ribosome-Inactivating Proteins
Ribosome-inhibiting proteins or ribosome-inactivating proteins found in certain plant species possess anti-viral activities. Transgenic expression of RIP from the pokeweed (Phytolacca americana) conferred broad-spectrum resistance to virus infection in several plant species, including tobacco, potato and N. benthamiana and Brassica napus against PVX, PVY and TuMV (Lodge et al. 1993; Zhang et al. 1999), respectively. Transgenic expression of RIP from Dianthus caryophyllus, Trichosanthes kirilowii and Iris hollandica also was observed to provide varying degrees of resistance to ACMV, TMV and CMV and TEV and TMV, respectively. However, any of the RIPs evaluated so far has not found to have an efficient broad-spectrum anti-viral activity.
Peptide Aptamers
There are some short-peptide molecules, which can confer low-level broad-spectrum resistance and are advantageous in that the need for producing high level of transgene to induce PTGS can be avoided. Such short-peptide-mediated resistance sometimes provides better resistance than RNAi- and protein-mediated PDR as in the case of transgenic N. benthamiana with an introduced target-specific peptide aptamer showing broad-spectrum resistance to Tomato spotted wilt virus (TSWV), Groundnut ringspot virus (GRSV) and Chrysanthemum stem necrosis virus (CSNV) (Rudolph et al. 2003). Similarly, a conformationally constrained peptide aptamer was found to interfere the replication of Tomato golden mosaic virus (TGMV) and Cabbage leaf curl virus (CaLCuV) (Lopez-Ochoa et al. 2006). Further investigation of such peptides and understanding of their mode of action enable their application for engineering broad-spectrum resistance.
Plantibodies
Antibody-mediated resistance to a plant virus was first demonstrated in transgenic N. benthamiana plants expressing an scFv against Artichoke mottled crinkle virus (Tavladoraki et al. 1993). Later, transgenic expression of scFv in different crop plants, like Chinese cabbage, citrus, gladiolus, potato and tomato, conferred them with virus resistance. Initial experiments resulted in delayed or reduced disease symptoms only, and complete resistance was not observed. Improvements in plantibody targeting and stabilization and development of nonstructural viral proteins targeting plantibodies enhanced the level of resistance provided by them (Gargouri-Bouzid et al. 2006; Nickel et al. 2008, Boonrod et al. 2004; Gil et al. 2011). Transgenic N. benthamiana expressing scFvs targeted against the TBSV replicase showed broad-spectrum resistance to TBSV, two members of family Tombusviridae (CNV and TCV) and Dianthovirus Red clover necrotic mosaic virus (Boonrod et al. 2004).
23.3 Engineering Resistance to Vectors
Many pathogens, particularly viruses, spread by different insect vectors, mostly hemipterans, aphids, dipterans, etc. Efficient vector control strategies significantly reduce the infection and crop damage. In certain cases, a single vector may harbour multiple pathogens, which are infecting same or different species. Transgenic strategies for vector control can be implemented by taking cues from vector-pathogen–host interactions and underlying genetic, molecular and biochemical processes. Persistent, semi-persistent and non-persistent vectors differ highly in their feeding nature, subcellular localization of virus in them and the status of virus in vector, whether replicating or non-replicative, circulating (Jones 2014; Brault et al. 2010). Viruses themselves influences host interaction with vectors and environment and modulate host immune response accordingly to regulate the feeding pattern and preference in order to manage the acquisition as well as injection of pathogens from viruliferous and non-viruliferous plants (Ingwell et al. 2012; Stafford et al. 2011). All these are accomplished by regulating host defence signalling hubs, manipulation of plant biochemistry for enhanced vector performance and virus-mediated suppression of plant immunity. Suppression of JA- and SA-mediated immune response to reduce insect repellents signals, i.e. different volatile compounds is a typical example for how pests reprogramme plant immunity for enhancing infection (Westwood et al. 2013). Interestingly, such negative regulation of defence-related signalling can be manipulated transgenically to reverse the effects so that resistance to both pests and pathogens can be achieved.
Understanding of involvement of semiochemicals and subsequent and signal generation and transduction can give an insight into potential semiochemical targets that can be modified transgenically to control vectors. Transgenic plants constitutively emitting the semiochemical (E)-b-farnesene, an aphid alarm pheromone and predator attractant, showed repulsion to aphid species and attraction to aphid-parasitizing wasps under controlled conditions (Bruce et al. 2015). However, the metabolically engineered plants showed no differences in aphid infestation or presence of natural enemies in field. Stimulated expression or mixed expression of different semiochemicals can be an alternative strategy to effectively control vectors.
Introgression of R genes conferring vector resistance, like the melon Vat (‘virus aphid transmission’) gene and the tomato Mi-1.2 gene, is useful for controlling vectors. Vat confers effective resistance to multiple viruses but only when transmitted by specific aphid species (Goggin et al. 2006), whereas Mi-1.2 provide resistance to the nematode Meloidogyne incognita and many phloem-feeding insects, including B. tabaci (Guo et al. 2016; Peng et al. 2016). As the vectors that are identified by NB-LRR protein domains of plants are limited, modifying these receptors in order to expand the range of defendable vectors using transgenic methods can be a promising strategy (Harris et al. 2013; Kim et al. 2016). The vector genes/transcripts encoding effectors can be targeted by transgenic expression of cognate RNA molecules in host, which is taken up by feeding invertebrates and silences the respective target. Fusion protein consisting of luteoviral coat protein–spider toxin was expressed in host, where viral coat protein moiety facilitates the uptake of toxin by vector (Whitfield and Rotenberg 2015; Bonning et al. 2014). Disruption of transmission of three groups of vectors – whiteflies, aphids and mealy bugs – was obtained by phloem-specific expression of a spider-derived toxin and a lectin (Javaid et al. 2016). Engineering plant as vectors of viruses, which are pathogenic on aphids and leafhoppers, including Densoviruses (DNA) and Dicistroviruses (positive-sense RNA), is a potential strategy, where host plants act as biocontrol agents. It was tested using cricket paralysing virus, and it disrupted normal aphid responses to olfactory cues from other aphids, causing them to scatter and predisposing them to attack by predators and parasitoids or promote transition to winged morphs, enhancing virus dissemination (Kerr et al. 2015; Ban et al. 2008; Ryabov et al. 2009).
23.4 Transgenic Approach for Resistance Against Viroids
Viroids are small (~250 to 400 nt) non-protein-coding, circular, single-stranded RNAs, which autonomously replicate through an RNA–RNA rolling-circle mechanism that is catalysed by host enzymes (RNA polymerases, RNases and RNA ligases) in the nucleus (Pospiviroidae), or in plastids, primarily chloroplasts (Avsunviroidae). These small infectious nucleic acids can infect economically important higher plants potato, tomato, citrus, hop and temperate fruits, like peach, apple and pear. Systemic infection of viroids involves entry into specific subcellular organelles, replication, exit of the organelles and cell-to-cell trafficking, access into and long-distance trafficking within the vasculature and exit and invasion of nonvascular tissue to restart the cycle. Interaction of the viroid RNA with cellular proteins involved in its replication/trafficking or from defensive responses is triggered by the host results in infection and disease development (Navarro et al. 2012; Flores et al. 2005). As conventional control measures sound insufficient to offer complete protection against these pathogens, transgenic approaches, like targeting the viroid RNA for degradation or manipulating the host defensive response in order to disrupt host–viroid interactions, may efficiently alleviate the disease symptoms. A peculiar resistance mechanism, namely, cross-protection, wherein a previous infection from mild strain induces development of resistance against further infection from a severely virulent strain, operates against viroids. The underlying molecular mechanism of cross-protection can be sequence specific, which may be related to RNA silencing (Niblett et al. 1978; Khoury et al. 1988; Flores et al. 2005).
As viroid replicates via dsRNA intermediates, yeast-derived dsRNA-specific RNase pac1 has been engineered to target this dsRNA of PSTVd in potato, which resulted in reduced infection and symptoms. Such RNAi approach can confer additional resistance to some RNA viruses as well. These pac1 expressing transgenic potato lines showed resistance to Tomato spotted wilt virus also (Flores et al. 2017). Expression of a catalytic single-chain variable antibody (3D8 scFv) with intrinsic RNase (and DNase) activities, in chrysanthemum plants, resulted in resistance to CSVd infection. This strategy is advantageous in that antibody accumulates to very low levels and antibody expression does not induce any phenotypic alteration, even though it acts in sequence-independent manner, and protects plants not only against CSVd but also confers resistance to DNA and RNA viruses (Lee et al. 2013; Tran et al. 2016). Different trials for antisense RNA-mediated silencing of viroid genomic RNA couldn’t establish a stable resistance strategy based on RNAi. Even so, CSVd resistance was conferred by Agrobacterium-mediated transformation of a commercial chrysanthemum cultivar with four different constructs carrying sense or antisense CSVd-specific RNAs of 75–82 nt (Matoušek et al. 1994; Jo et al. 2015). Although initial attempts to engineer hammerhead ribozymes to target the plus and minus strands of CEVd did not confer viroid resistance in vivo, hammerhead ribozyme with shorter recognition sequences (9–11 bases) engineered to target PSTVd minus RNA suppressed viroid accumulation in transgenic potato lines (Yang et al. 1997). Tertiary stabilizing motifs (TSMs) of hammerhead ribozymes is suspected to play a critical role in their potential to cleave specific RNA in trans, and this was substantiated when hammerhead ribozymes, with their tertiary stabilizing motifs (TSMs) preserved, engineered to target PSTVd minus RNA successfully interfered with systemic PSTVd infection in in transgenic N. benthamiana. Target accessibility of the substrate and the subcellular co-localization of ribozyme and substrate are major hurdles to extract the complete potential of these small RNAs for crop protection (Flores et al. 2017). RNA interference strategy for targeting viral genome can be applied to viroids also. Transgenic tomato and N. benthamiana expressing hpRNA construct with near full-length (340-nt) and specific truncated sequences of PSTVD, respectively, showed resistance to this viroid. As RISC is active in cytoplasm, it may not interfere with viroid replication and accumulation host in nucleus but can be targeted while they move to cytoplasm for invasion into neighbouring cells (Schwind et al. 2009). Overexpression of AGO also has found to attenuate viroid accumulation (Minoia et al. 2014). Artificial small RNAs, artificial miRNAs (amiR-NAs) and synthetic trans-acting siRNAs (syn-tasiRNAs) are another efficient transgenic strategy for controlling viroids, when stably expressed in transgenic plants (Carbonell et al. 2014). Concurrent application of more than one strategy might have synergistic effects, resulting in enhanced resistance to viroids.
23.5 Transgenic Technology for Resistance Against Fungi
Fungal pathogens cause several diseases in plants resulting in catastrophic effects on crop yield (10% of global crop loss). Conventional control strategies include the application of chemical fungicides and biocontrol agents, phytosanitation, crop rotation, destruction of intermediate hosts, etc., which are inadequate to provide complete protection from infection. Transgenic strategies can be adopted to confer/enhance the resistance to fungal pathogens so that yield loss and disease management expenses can be considerably reduced. Several plants naturally produce antifungal compounds, and genes encoding these compounds or regulating their biosynthesis can be manipulated to confer fungal resistance. Prime plant-derived antifungal compounds are chitinase and glucanases, enzymes that degrade major components of fungal cell wall, chitin and glucan, thereby imparting resistance (Silva et al. 2018, 2019). Stacking of these two genes can yield enhanced resistance against fungal pathogens. Transgenic carrot expressing chitinase and β1,3-glucanase, together with AP24 gene, exhibited broad-spectrum resistance against fungal pathogens (Ram and Mohandas 2003). Transgenic grape and cotton plants constitutively expressing chitinase from Trichoderma species conferred resistance to several fungal diseases (Rubio et al. 2015; Emani et al. 2003). Plant-derived inhibitors of microbial cell wall degrading enzymes are other potential candidates for engineering resistance. Transgenic expression of inhibitors of cell wall degrading enzymes has conferred resistance against many dreaded fungal pathogens in crop plants. Reduced susceptibility to necrotrophic fungus Botrytis cinerea has been observed in PGIP overexpressing transgenic tomato, grapevine, tobacco and Arabidopsis thaliana (Ferrari et al. 2012; Liu et al. 2017; Manfredini et al. 2005). PGIP degrades microbial polygalacturonases and thereby delays plant cell pectin hydrolysis, which in turn restricts fungal infection (Ferrari et al. 2013). Xylanase inhibitor protein controls the xylanases, which degrades the main component of cell wall, xylan, as in the case of effective inhibition and counteraction of the F. graminearum necrotic xylanase activity by constitutive expression of Triticum aestivum xylanase inhibitor III (TAXI-III) in GM wheat (Tundo et al. 2016). Expression of pectin methyl esterase (PME) inhibitors in Arabidopsis could prevent damage to the plant cell wall during Botrytis cinerea infection (Lionetti et al. 2017). Transgenic strategies to control insect pests, including cry genes from Bacillus thuringiensis, provide dual protection by reducing the chances of fungal infection and mycotoxin (fumonisin, Fusarium spp.; aflatoxin ergotoxine, Claviceps spp.; aflatoxins, Aspergillus spp.) contamination by reducing the rate of insect wounding. This would reduce management costs and yield loss considerably. Overexpression of PnAMP-h2 gene from Pharbitis nil and barley chitinase (chi-2) genes were found to provide enhanced resistance to fungal pathogens. Antimicrobial peptides derived from non-plant, non-phytopathogenic microbes can also be used for imparting disease resistance as in the case of enhanced resistance to R. solani and soil-borne pathogen A. alternata observed in transgenic cotton and tobacco plants expressing endochitinase gene from a mycoparasitic fungus Trichoderma virens. Phytoalexin capsidiol provided resistance to the potato blight pathogen, Phytophthora infestans (P. infestans) in pepper (Capsicum spp.) (Lee et al. 2017) (Table 23.4).
Defensins are cysteine-rich antimicrobial peptide of plant origin, which is a potential candidate to protect crop plants with phytopathogenic fungal infection by their transgenic expression in plants. Constitutive expression of an alfalfa seed defensin MsDef1 and NmDef02 defensin in potato provided strong resistance to Verticillium dahliae and Phytophthora infestans, respectively (Gao et al. 2000). Constitutive expression of defensins mostly causes undesirable side effects, including reduced growth and yield performance (Chen and Chen 2002). Regulation of defensin expression by using (native or heterologous) pathogen-induced promoter like barley GER4c promoter or tissue-specific promoter, which mainly depends on the infection biology and tissues affected, would be advantageous to avoid the negative fitness effects associated with their constitutive expression (Himmelbach et al. 2010; Rushton et al. 2002). Expression of antifungal defensins under root-specific promoter confers resistance to sudden death syndrome in soybean, because it is caused by a root-colonizing pathogen Fusarium virguliforme. Similarly, lifestyle-specific expression of defensins by fine-tuning their subcellular localization, which in turn depends upon whether pathogen is biotrophic or hemibiotrophic, is also effective to confer durable resistance without compromising yield. Extracellularly targeted defensins enable protection from biotrophic fungi, whereas extra- and intracellularly targeted defensins should be coexpressed in order to confer resistance to hemibiotroph. However, the effect of transgene encoded antimicrobial compound on animal/mammalian system should be assessed before employing this strategy for transforming edible crops. A detailed understanding of the underlying molecular mechanisms of their mode of action would permit the development of much sophisticated strategies for the regulated expression of antifungal defensins to confer broad-spectrum durable resistance against many notorious fungal pathogens (Kaur et al. 2011). Synergistic effects by coexpression of different plant defensins with different mode of action or plant defensin with antifungal pathogenesis-related protein (PR) or HIGS with plant defensins enhance resistance to a wide range of fungal infection (Chen et al. 2009; Ntui et al. 2011).
Silencing of pathogen genes by RNAi-based host-induced gene silencing (HIGS) is another efficient transgenic strategy to develop resistance. Resistance to Blumeria graminis and F. verticillioides was achieved by HIGs of their endogenous gene and fungal transgene, respectively (Nowara et al. 2010; Tinoco et al. 2010).
Plant resistance genes have extensively used for engineering resistance against phytopathogenic fungi. Introgression of R genes from related or unrelated species is facilitated by transgenic technologies. Overexpression of NPR1, a key regulator of SAR, conferred resistance to blight and blast disease causing fungal and bacterial pathogens, respectively (Chern et al. 2005; Yuan et al. 2007). Transgenic rice plants overexpressing AtNPR1 gene and translational suppressor uORFs from the TBF1 gene displayed resistance towards bacterial and fungal pathogens and reduced negative fitness costs associated with constitutive expression of NPR (Xu et al. 2017). In another case, introgression of the gene involved in non-host resistance, Phytophthora soaje susceptible 1 (AtPSS1), from Arabidopsis into soybean by transformation enhanced resistance to the soybean host fungal pathogen Fusarium virguliforme, by a suspected autophagy mechanism without yield penalty (Wang et al. 2018). Transgenic expression of R gene RPI-BLB2 from Solanum bulbocastanum in cultivated potato showed resistance to Phytophthora infestans (van der Vossen et al. 2005). Resistance conferred by single R gene is mostly not/less durable as pathogen is smart enough to circumvent the resistance. Stacking of multiple R genes, either targeting single or different pathogens, in single plant is an efficient alternative to impart durable and/or broad-spectrum resistance against destructive fungal pathogens. Robust transgenic strategies for gene pyramiding are required for executing this. Host genes manipulated by fungus for apressoria formation and haustoria development can be a potential target to be modified in order to confer pre-haustorial resistance. Arabidopsis PENETRATION genes involved in haustoria establishment is a suitable candidate in that regard (Fonseca and Mysore 2019). Integrated transgenic approach combining resistance genes, defensins and antifungal proteins is expected to confer effective durable resistance.
23.6 Transgenic Technology for Bacterial Resistance in Crop Plants
Bacteria are causing many diseases in economically important plants, where it severely impairs the growth, development and yield potential of plants. Plants defend bacterial pathogens with common components of immunity, viz. R genes, ETI-associated HR, antimicrobial compounds, inhabitants of bacterial enzymes, etc. Transgenic approaches for engineering resistance to bacteria in plants can be executed in the following ways: introduction of host R genes and bacterial avirulence genes, incorporation of pathogen-derived genes for resistance to bacterial phytotoxins and expression of antibacterial proteins from plants, insects or bacteriophages as bactericidal or bacteriolytic agents (Panopoulos et al. 1996). Transfer of maize R gene RESISTANCE TO XANTHOMONAS ORYZAE 1 (RXO1) to rice conferred enhanced resistance to the rice host pathogen Xanthomonas oryzae pv. oryzicola causing bacterial streak disease. Similarly, single R gene obtained from pepper was introgressed to tomato in order to control highly destructive bacterial leaf spot (Zhao et al. 2004, 2005; Tai et al. 1999). Unravelling novel candidate receptors or enzymes or structural proteins from bacteria that can be targeted by transgenic methods will have promising results in controlling bacterial diseases (Table 23.5).
Antimicrobial peptides of plant origin as well as those from heterologous non-plant systems are efficient candidates for engineering disease resistance. Plant-derived antimicrobial peptides contain cysteine-rich short amino acid sequence and target the outer membrane structures. Overexpression of pepper antimicrobial peptide-encoding gene CaAMP1 in Arabidopsis conferred broad-spectrum resistance to bacterial, fungal and oomycete pathogens (Lee et al. 2008).
A significant reduction of fire blight symptoms in pear and partial resistance to bacterial blight in tomato was achieved by expression of lactoferrin, a mammalian glycoprotein (Malnoy et al. 2003). Attacin is another heterologous protein that has been used for conferring resistance against Erwinia amylovora, causing fire blight in transgenic apple (Aldwinckle et al. 2003). Transgenic expression of another protein, pectate lyase 3 (PL3), in potato resulted in increased resistance to Erwinia soft rot (Wegener 2002).
Cecropins are antimicrobial amphipathic peptides of 31–39 aa long and interact with bacterial membranes or induce pore formation in membrane. Transgenic tomato expressing high level of cecropin displayed enhanced resistance against bacterial speck disease. An increased resistance to bacterial blight in rice was engineered by targeting cecropin towards intercellular species (Alan et al. 2004; Jan et al. 2010).
Increased expression and accumulation of ROS, hydrolytic enzymes, antimicrobial peptides and proteins in transgenic host plant can enhance resistance, including HR, to bacterial pathogens and avoid the energy and resource loss in inducing immune responses, which in turn enable plant to channelize the saved energy and nutrients/metabolites into growth and development process so that yield is not compromised. Resistance to several bacterial and fungal pathogens has been achieved by transgenic expression of genes encoding cell wall degrading hydrolytic enzymes, defence-related proteins with protease inhibitor activity, etc. (Shin et al. 2008; Senthilkumar et al. 2010).
23.7 Genome Editing Techniques for Engineering Disease Resistance
Genome editing technology, which emerged in the 1990s, enabled targeted mutagenesis of genomic loci of interest by exploiting cell’s intrinsic DNA repair pathways. Site-directed nuclease fused with sequence-specific DNA-binding protein domains or RNAs creates double-stranded breaks in target genomic site. DSBs thus formed are subsequently repaired either by error-prone DNA repair pathway non-homologous end joining (NHEJ) or high-fidelity homology-directed repair pathway (HDR). NHEJ usually results in point mutations at the site of DSB, and HDR enables replacement or introduction of a DNA fragment, while a repair template is provided. Mutations thus induced are stably inherited over generations. Genome editing techniques are useful for functional genomic studies as well as for creating or modifying desired phenotype or trait in organism of interest. Meganucleases, zinc finger nucleases (ZFN), TALENs and CRISPR/Cas are the commonly used genome editing tools. ZFNs and TALENS are fusion proteins in which specific DNA-binding proteins domains are fused with endonuclease domain of FokI nuclease, so that protein guides FokI to respective DNA targets. CRISPR/Cas9, in contrast, is an RNA-guided engineered nuclease, where a single guide RNA with 5′ target-specific 20 nt spacer directs Cas nuclease to target DNA. CRISPR/Cas is a part of bacterial adaptive immune system, providing resistance to invading viruses or nucleic acids. There are different classes of CRISPR/Cas systems, of which dsDNA targeting class 2, type II system seen in Streptococcus pyogenes, popular as CRISPR/Cas9 is the most common CRISPR system exploited for genome editing. Several variants of CRISPR systems and Cas protein have been discovered, each of which varies in their target recognition and cleavage properties.
Genome editing techniques have been successfully employed to develop resistance against bacterial, fungal and viral pathogens. ZFNs conferred broad-spectrum resistance to various Begomoviruses, including Tomato yellow leaf curl China virus (TYLCCNV) and Tobacco curly shoot virus (TbCSV) by targeting a single site in viral genome (Chen et al. 2014). Modified ZFN called as artificial zinc finger protein (AZP), which lacks the cleavage domain compared to ZFN, has been used to confer resistance to Beet severe curly top virus (BSCTV, family Geminiviridae) and Rice tungro bacilliform virus (RTBV) in Arabidopsis by targeting the intergenic region (IR) of BSCTV and blocking viral promoter sequences of RTBV (Ordiz et al. 2010). TALEs have been engineered to confer resistance to TbCSV and TYLCCNV in tobacco (Cheng et al. 2015). Complexity of protein engineering and off-target effects limited the use of these modular protein toolboxes. CRISPR/Cas9, whose genome editing potential was revealed recently only, quickly superseded and emerged as a promising tool for crop improvement with high efficiency, precision, robustness and simple designing strategy.
CRISPR/Cas9 has been widely used for engineering disease resistance in many crop plants by targeting pathogenic as well as host factors. Pathogenic genes that are inevitable for survival and infection cycle are appropriate candidate targets for CRISPR/Cas9-mediated knockout. Adaptive potentials enable pathogens to overcome the dominant R gene-mediated resistance. Thus, susceptibility genes are considered as potential candidates to be targeted in order to develop durable resistance. Disruption of susceptibility genes or S proteins by CRISPR/Cas9-mediated targeted mutagenesis has been successfully demonstrated. The first report of the application of CRISPR/Cas9 for virus resistance came when double-stranded replicative from geminiviral DNA was targeted to confer resistance. Later, this strategy was used to provide resistance to several DNA viruses. Apart from the demonstration for proof of concept of the CRISPR/Cas9 in model plants, highly efficient resistance against different viral, bacterial and fungal pathogens has been obtained in various crops, including major food crops like wheat and rice (Table 23.6, 23.7 and 23.8). For instance, mutagenesis of ERF transcription factor (OsERF922) gene using CRISPR/Cas9 imparted resistance in rice against fungal pathogen Magnaporthe oryzae (Wang et al. 2016). Similarly, CRISPR/Cas9-induced mutation in homeoalleles of mildew resistance locus (MLO), encoding a transmembrane protein, and enhanced disease resistance 1 (EDR1), encoding a Raf-like mitogen-activated protein, conferred resistance to powdery mildew in hexaploid wheat (Wang et al. 2014). CRISPR/Cas9-mediated loss-of-function mutation in eukaryotic translation initiation factor eIF4e, which is essential for infection of Potyviridae family viruses, conferred resistance against potyvirus (Zucchini yellow mosaic virus and Papaya ringspot mosaic virus-W) and Ipomovirus in Arabidopsis and cucumber (Cucumber vein yellowing virus) (Chandrasekaran et al. 2016). CRISPR/Cas9-mediated targeted mutagenesis of promoter sequence of susceptibility gene CsLOB1 conferred resistance to bacterial pathogen Xanthomonas citri subsp. citri in citrus (Peng et al. 2017). This tool has also been shown to be effective to control dsDNA viruses, as in the case of resistance to Cauliflower mosaic virus (CaMV) in Arabidopsis (Liu et al. 2018).
Loss-of-function mutations created by CRISPR/Cas9 is useful for identifying resistance and susceptibility-determining genes and corresponding regulatory elements like promoters or enhancers. As the NHEJ-induced mutations are stably inherited and the CRISPR/Cas expression cassettes can be eliminated in successive generations, the resulting plant remains transgene-free and thus overcomes the regulatory issues associated with Agrobacterium transformed conventional transgenic plants. Delivery of CRISPR/Cas9 constructs as ribonucleoprotein (RNP) complex composed of sgRNA and Cas9 protein into host plant also yields non-transgenic edited plant with disease resistance. Non-transgenic tomato, resistant to powdery mildew, has been created in this manner (Nekrasov et al. 2017). Multiple DNA sequences can be targeted simultaneously by designing gRNA cassettes using golden gate cloning/tRNA: gRNA/gRNA: ribozyme assembly strategy. The ease of multiplexing is highly beneficial for conferring host with sound resistance against single pathogen as well as with broad-spectrum resistance. Multiple regions of pathogen genome can be targeted to alleviate/reduce the possibilities for the evolution of cleavage or recognition resistant variants by overcoming mutations at single site. Although incorporation of resistance gene from foreign species can also be performed by inducing HDR repair pathway of DSB created, efficiency of such events is very less and still needs to be optimized for obtaining high rate of HDR. Cisgenic applications of genome editing can be useful for enhancing disease resistance in species, which are difficult to hybridize by introducing a desirable gene fragment from its natural gene pool using CRISPR/Cas9-induced HDR-based pathway. Genetic engineering approaches manipulating endogenous genetic variations related to specific traits are preferred to exogenous genomic targeting recombinant DNA technology.
Apart from dsDNA targeting Cas9, there are certain CRISPR systems with RNA targeting properties, which can be engineered to confer resistance to RNA viruses or silencing of viral transcripts in host. Class II, type VI Cas effector proteins, Cas13a (C2c2), Cas13b (C2c6), Cas13c (C2c7), Cas13d and Cas9 from Francisella novicida (FnCas9), RNA targeting SpCas9 (RCas9), are the RNA-guided RNA targeting Cas effector variants identified in different microbial genomes so far (Abudayyeh et al. 2017). Expression of Fncas9 and sgRNA targeting conserved 3′ UTR of Cucumber mosaic virus (CMV) and Tobacco mosaic virus (TMV) developed stable immunity in N. benthamiana against CMV and/or TMV (Zhang et al. 2018). CRISPR-Cas13a was used to confer resistance to Turnip mosaic virus (TuMV) in N. benthamiana by targeting Hc-Pro and GFR regions of genome (Aman et al. 2018).
A class II, type V CRISPR/Cpf1 system that efficiently target DNA was discovered from Prevotella and Francisella species and named as Cas12a (Zetsche et al. 2015; Makarova et al. 2015). Cpf1 from Acidaminococcus sp. BV3L6 Cpf1 (AsCpf1) and Lachnospiraceae bacterium Cpf1 (LbCpf1) are other two promising nucleases that can be used for genome editing purposes (Zetsche et al. 2015).
Gene expression regulation and epigenetic modification is also facilitated by mutant Cas9, which lacks cleavage activity. Transcriptional activators or repressors can be recruited to a particular gene of interest by fusing them with dCas9, which then is directed by specific sgRNA to the target site. Similar strategy can be adopted for editing epigenetic marks using appropriate epigenetic modifiers (Dominguez et al. 2016). This enable fine-tuning of defence regulatory pathways and metabolic pathways involved in immune response. Transcriptional activation of positive regulators of immunity and biomolecules like pathogenesis-related proteins or hydrolytic enzymes, which are directly involved in defence manifestation, repression of negative regulators of immunity, etc., can be achieved using CRISPR/dCas9 system. Resistance traits that are unexpressed due to epigenetic suppression can be activated by reversing the methylation and similarly epigenetically controlled susceptibility also modified. Tissue and developmental stage or stress status specific activity of CRISPR components and target editing events can be achieved by employing tissue-specific/inducible promoters for the expression of Cas effector and guide RNA. This is particularly beneficial for reducing unwanted or off-target editing.
Genome-wide mining of S genes may unravel novel candidate S genes that can confer resistance, durable as well as broad-spectrum, without having any fitness costs. Several trace elements, like Fe, Ca, Cu, K, Mn and Zn, are required by pathogens to meet their nutritional requirements, whereas the same are required in host plants for nutrition as well as to activate/drive certain critical defence responses like Cu-dependent binding of SA binding to its NPR1 receptor (Yuan et al. 2010). Coordinated activities of plant immune signalling pathways to inhibit/suppress bacterial iron acquisition mechanisms substantiated the role of Fe in establishing infection (Nobori et al. 2018). Pathogens compete with host to acquire these mineral nutrients, which is counteracted by plant, by limiting their availability to invading pathogens by a mechanism called as nutritional immunity. Pathogenic mineral transporters strive to acquire Fe, Mn, Zn, K and Cu, which are essential for their virulence/survival, from the host, leaving host deprived of these trace elements and thereby inhibiting immune responses that require these minerals (Ren et al. 2016; Hood and Skaar 2012). Thus, such trace element transporters or their acquisition mechanisms can be potential nutritional immunity-related S gene targets for genome editing (Zaidi et al. 2018).
Complete knockout of S genes mostly has negative fitness effects, like reduced growth, yield and fertility; early senescence; and reduced tolerance to abiotic stress, as most of them being primary in function. Even though fitness cost is not lethal, it can cause phenotypic abnormalities. Alternative strategies, like introduction of S gene variant, creation of intermediate alleles by promoter targeting, transient knockout of S genes by using pathogen-inducible promoter, etc., using CRISPR/Cas9 would efficiently confer resistance, avoiding any cost of fitness. Identification of S gene allelic variants, which can confer resistance, leads to create resistant S gene variant by specifically editing the respective SNP using CRISPR/Cas9 base editors (Rodríguez-Leal et al. 2017; Yan et al. 2018).
Even though the characteristics, like specificity, efficiency, simplicity, flexibility, etc., of this genome editing tool are highlighted, several shortcomings and complications, including off-target effects, low rate of successful transformation events and recalcitrance for regeneration and establishment of edited plants, are associated with CRISPR/Cas. Each stage of experiment, from the selection of target sequence to stable integration of mutations, needs to be optimized for each and every species for utilizing maximum potential of this genome editing tool for disease resistance.
23.8 Strategies Based on the Mechanism of Plant–Pathogen Interaction
Plants are constantly subjected to stress from various biotic agents/pathogens, like virus, bacteria, fungi, oomycetes, nematodes, insects and parasitic plants, that are causing serious crop loss. Plant–pathogen interaction studies have revealed various components involved in immunity and their mechanism of action during defence response. Interaction is specific characteristic of each group of pathogens. A vast array of well-programmed defensive mechanisms driven by multiple biomolecules which are involved in each stage, viz. invasion of pathogen into host, recognition and initiation of immune response, active deployment of defensive strategies and establishment of resistance, together constitute plant immunity. Pathogen-derived signals during infection process is perceived by specific receptors, mostly located in the cell membrane, which then trigger a cascade of defence activities/pathways making use of a large spectrum of biomolecules and molecular mechanisms (Silva et al. 2018).
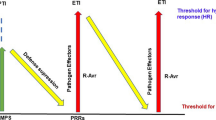
Plant innate immunity has organized as a three-layered system with pathogen-triggered immunity (PTI), effector-triggered susceptibility (ETS) and effector-triggered immunity (ETI). The first active line of plant immunity is PTI, which is triggered when pathogen-associated molecular patterns (PAMP), together with this damage-associated molecular patterns (DAMP) released during pathogen invasion and damage, are recognized by transmembrane pattern recognition receptors (PRRs) (Boutrot and Zipfel 2017; Uma et al. 2011). PTI is suppressed by effector-triggered susceptibility (ETS) induced by pathogen-derived susceptibility proteins/effector and results in infection (Jones and Dangl 2006; Chisholm et al. 2006). This activates the second line of defence, effector-triggered immunity induced by recognition of specific effectors/cognate factors, pathogen avirulence (Avr) proteins by another group of receptors encoded by resistance genes (R). A co-evolutionary gene-to-gene molecular arm race occurs between pathogen effectors and host R genes, while pathogen evolves a new effector to restore the compatible interaction to facilitate infection, parallel to which host evolves a new R protein, to strengthen immunity. PTI is conserved over a range of organisms, whereas ETI specific to Avr protein is produced by each organism. ETI mostly/usually activates localized cell death pathway, otherwise known as hypersensitive response (HR), to restrain infection by transmitting defence signals to neighbouring non-infected cells via plasmodesmata and to other systemic organs via phloem, resulting in distal resistance responses, namely, local acquired resistance (LAR) and systemic acquired resistance (SAR), respectively (Coll et al. 2011; Dangl and Jones 2001). Like HR, ETI responses also involve production of salicylic acid (SA), reactive oxygen species (ROS), necrosis and also structural changes, such as lignification and callose deposition. Various biomolecules, like pathogenesis-related proteins (PR), antimicrobial peptides (AMP), ribosome-inhibiting proteins (RIPs), defensive secondary metabolites, etc., are produced by plants as a part of defence response (Kachroo and Robin 2013; Dempsey and Klessig 2012). Increasing the cytosolic calcium levels in response to exogeneous signals, like H2O2/PAMP/DAMPs, etc., produced during infection is sensed by cellular and membrane calcium receptors, which in turn transduce signals to elicit defence responses, like HR, ROs production, transcriptional regulation of stress responsive genes, etc. (Seybold et al. 2014; Silva et al. 2018).
Incompatible interaction between non-host species and pathogen results in immunity against non-host pathogen in non-host plant. Preinvasive NHR is mostly passive mainly based on different physical and chemical barriers to interfere proliferation and accumulation of pathogen. Sometimes, NHR involves active defence response mediated by incompatible R–Avr gene-to-gene interaction result in ETI, and this shows that common mechanisms converge at some points in NHR and HR (Flor 1971; Glazebrook 2005; Gill et al. 2015). Prolonged interaction between host and pathogen occurs, wherein pathogens are under constant dual selection pressures, for increased resistance from host and on pathogen for increased performance of pathogen, which results in co-evolution of pathogen and host with virulence specificities on specific hosts (Allen et al. 2004). Such co-evolved pathogens become unable to infect phylogenetically unrelated hosts, resulting in non-host resistance, which is highly beneficial to control pathogens. NHR is a quantitative trait with multiple genes and pathways, whereas HR has specific R gene (Senthil-Kumar and Mysore 2013). Based on plant response, NHR is considered to be of two types. Type 1 NHR involves no visible symptom as pathogen fails to penetrate tissues. It is manifested by different physical, chemical and metabolical barriers that block pathogen penetration without activating ETI and PTI. Type II NHR involves some degree of pathogen penetration or entry in plant tissue, not as strong as a susceptible host pathogen infection but in a dose sufficient to trigger HR (cell death), and involves ETI. During post-invasive NHR, some pathogens penetrate host by overcoming PTI responses and involve activation of defence responses, like HR, ROS, cell death or the formation of cell wall appositions in infected cells (Collins et al. 2003; Rojas et al. 2012).
23.9 Enhancing Immunity: Transgenic Approaches to Manipulate Plant Innate Immunity
23.9.1 Upregulation of Defence Pathways
Upregulation of molecules involved in defence regulation, signalling and other allied cellular processes can boost hosts’ general immune responses, including ROs production, callose deposition, PR proteins and activation of SAR, etc. Resistance to several microbial (bacterial and fungal) pathogens has been achieved by employing this strategy without introducing new metabolic pathway or new gene but exploiting plant’s own immune system. Resistance to Rhizoctonia solani and Magnaporthe oryzae (rice blast causative fungi) has been achieved by expressing a native rice gene under the control of a constitutive promotor from maize (Bundó and Coca 2016; Chen et al. 2016; Vincelli 2016). Genome editing techniques, like CRISPR/cas9-mediated transcriptional regulation, can be a potential strategy to serve this purpose.
23.9.2 Production of Antimicrobial Compounds
Microbial pathogens secrete different cell wall degrading enzymes (CWDE) like cellulases, polygalacturonases, xylanases, xyloglucan endoglucanase, chitinases and protease inhibitors to damage cell wall for making their way into host cells. In order to combat CWDEs, plants initiate cell wall strengthening/damage preventing mechanisms, like production of polygalacturonase-inhibiting proteins (PGIPs), xylanase-inhibiting proteins (XIPs) and xyloglucan-specific endoglucanase-inhibiting proteins (XEGIPs) (Schüttelkopf et al. 2010; Xu et al. 2011). As these mechanisms are mostly evolutionarily conserved, their manipulation could confer durable and broad-spectrum resistance in many crop species. Reduced susceptibility to Phytophthora capsici by expressing a pepper PGIP in GM tobacco; resistance to Verticillium and Fusarium wilts by expressing protein GhPGIP1 in Arabidopsis and cotton; resistance to Phytophthora sojae in soybean by constitutive expression of elevated levels of a Glycine max XEGIP (GmGIP1), an inhibitor of a Phytophthora sojae, xyloglucan-specific endoglucanase (PsXEG1); etc. have proved the efficacy of enhancing plant’s own immune mechanisms by transgenic strategies to confer resistance (Silva et al. 2013; Liu et al. 2017; Wang et al. 2013; Ma et al. 2015).
23.9.3 Enhancing Plant Recognition of Infection
PTI results from perception of self-derived DAMPs and non-self PAMPs signals by PRR comprising receptor like kinases and receptor like proteases, whose active domains include leucine-rich repeats (LRR), lysine M domain (LysM), epidermal growth factor (EGF) like domain and lectin motif (Wu and Zhou 2013; Silva et al. 2018). Transfer and expression of PRRs in heterologous species by transgenic methods broaden the range of pathogens that can be recognized by the host while plant’s own defence system or metabolic pathways are left unmodified. Expansion of pathogen recognition window is of prime importance in field conditions, where multiple infections prevail. Even though certain microbes are highly adapted with species-/race-/strain-specific PAMPs and have specific residues/motifs to block perception by PRRs, most of the PAMPs are conserved across microbial species, and thus individual PRR genes can confer broad-spectrum resistance (Zipfel and Felix 2005). Manipulation of PRRs and other receptors or downstream components involved, from a non-host species or unrelated family, is useful for boosting plant immunity as it will be difficult for pathogen to overcome the resulting new PRR/PAMP recognition system and the downstream signalling pathways. Resistance to Xanthomonas citri in transgenic Hamlin sweet orange expressing NbFLS2 and resistance to Pseudomonas syringae in Arabidopsis expressing Nicotiana benthamiana receptor-like protein are required for Csp22 responsiveness (NbCSPR); NbCSPR exemplifies the effectiveness of transgenic expression of PRRs, even in heterologous system (Hao et al. 2016). Similarly, the transgenic expression of AtEFR in tomato (against P. syringae pv. syringae, Ralstonia solanacearum and Xanthomonas perforans), tomato PRR Ve1 in Arabidopsis and the Arabidopsis PRR AtEFR-Tu in wheat and potato against bacterial wilt (BW) causing soil-borne pathogen Ralstonia solanacearum is also expected to confer durable and effective resistance to microbial pathogens (Fradin et al. 2011; Lacombe et al. 2010; Schoonbeek et al. 2015). As most of the PAMPs are conserved and metabolically vital molecules of pathogen, it may not evolve quickly along with transferred PRRs in order to widen the host range.
23.9.4 Targeting Susceptible Gene/Protein
PTI is modulated or suppressed by the binding of pathogen effector proteins to host S proteins and thereby triggering ETS, which facilitate pathogen’s invasion and establishment. Plant S genes are responsible for susceptibility to pathogen, ETS establishment and supporting compatibility. Molecular mechanisms associated with S genes are exercised in three levels: basic compatibility or early establishment in preinvasive stage when recognition and invasion occurs, sustained compatibility during which proliferation and spread of pathogen occurs and, finally, negative regulation of immune signals via respective signalling during post-penetration stage (van Schie and Takken 2014; Pavan et al. 2010). S proteins can either be positive regulators of immunity by promoting infection and disease development or negative regulators of immunity, and S gene products either facilitate pathogen’s growth in host system or negatively regulate the plant immunity (Silva et al. 2018). Disarming/dysfunctioning, such susceptibility genes, by knockout or loss-of-function mutation is an efficient strategy to develop immunity, and the resulting resistance is mostly recessive and may be associated with pleiotropic effects and negative fitness costs, which may be due to constitutive activation of defence response. Durable and broad-spectrum resistance to powdery mildew in different plant species has been conferred in the same manner, by mutating mildew resistance locus O (MLO). Effector targets in host are activated by pathogenic effectors, which then negatively regulate plant immunity. S gene-mediated resistance can be pathogen specific, if the impaired pathway is inevitable for pathogen in any of the pre-penetration/penetration/post-penetration events, and it can be broad-spectrum, if the target S gene has implications in constitutive defence response. Another point to be noted while S proteins are encoded by multiple S genes is that all of them need to be targeted simultaneously; otherwise, the feasibility of same also should be checked. S gene LOF confers plants with broad-spectrum durable resistance than dominant R gene-mediated resistance as this is race/pathovar/strain non-specific. In order to overcome a modified or silenced susceptibility gene conferring host-pathogen compatibility, pathogen should acquire a new function to replace the host factor it was exploiting, which is a tough task.
Eukaryotic translation initiation factors eIF4e and eIF4g usually interact with the cap structure of transcripts. Upon Potyviridae family virus attack, this interaction is disrupted and initiates interaction with Vpg proteins from virus and facilitates translation of viral proteins in host cell, thereby eIF4e and eIF4g act as negative regulators of plant immunity (Zhang et al. 2006; Callot and Gallois 2014). Similar factor rwm1 has been identified in watermelon under Cucumber mosaic virus infection (Ouibrahim et al. 2014). Loss-of-function mutation of such factors confers recessive resistance towards respective pathogens and is advantageous as far as it does not cause any serious pleiotropic effect. Sugar efflux from the host cell is activated by TAL activator from Xanthomonas oryzae, by binding to promoter region of SWEET14 gene in order to nourish bacterial pathogen (Yuan and Wang 2013; Boch et al. 2009; Chen et al. 2012; Li et al. 2012). Since this may have an adverse impact on plant health and agronomic performance due to altered sugar flux, modification of binding sites or specific residues of SWEET14 gene render is non-accessible or useless for pathogen. This can be accomplished by CRISPR/Cas9 genome editing tool.
23.9.5 Mining of R Genes
Overexpression or transfer of resistance genes from other related/unrelated species can confer durable broad-spectrum resistance. This is particularly advantageous in some crop species, where natural R gene is lacking or endogenous R genes fail to confer effective broad-spectrum resistance to most dreaded pathogens. In such cases, resistance genes can be introgressed from distantly related or even from unrelated species by genetic engineering, which in any way is impossible with conventional breeding. Conventional breeding itself is a hurdle in the case of polyploid crops, like potato, grape, banana, apple, etc. In such cases, the useful Cis genes also can be introgressed by genetic engineering, by avoiding major drawbacks of breeding techniques, viz, time consumption and linkage drag (Jones et al. 2014). Resistance to potato late blight has been achieved by this strategy (Jo et al. 2014). Though resistance genes efficiently confer resistance, their durability in heterologous hosts, over the course of time in field conditions, is limited. Since R gene action is strain-/race-specific, it confers species/subspecies level durable, broad-spectrum resistance, particularly in phylogenetically related hosts by heterologous expression of R genes (Eg: Bs2 gene). GM tomato plants expressing Bs2, an R gene from pepper that recognizes AvrBs2 present in some Xanthomonas campestris pathovars, were conferred with resistance against Xanthomonas perforans (Horvath et al. 2012). The efficacy of R genes in heterologous systems in conferring disease resistance has been demonstrated by similar transgenic plants. Bs2 gene functions as a solid source of resistance to different Xanthomonas species and X. campestris pathovars (e.g. pv. vesicatoria (Xcv)) in multiple solanaceous species. Co-transformation of R genes with cognate Avr gene can be followed by stacking of heterologous R genes, once the R gene becomes able to overcome phytopathogen. Such interspecific R gene transfer confers host with broad-spectrum resistance in unrelated host as well (Gururani et al. 2012; Horvath et al. 2012). Overexpression of certain post-invasion- induced non-host resistance genes (PINGs) and bright trichomes (BRT1) genes that encode UDP glycosyltransferase 84A2, both of which have been found to be involved in NHR, conferred enhanced resistance against P. pachyrhizi in soybean (Langenbach et al. 2016). It has been found that expression of translational repressor domain from the TBF1 gene (uORFs), along with the defence-related gene, can circumvent the negative fitness costs associated with expression of defence-related genes in plants. For example, transgenic rice plants overexpressing the AtNPR1 gene and translational suppressor uORFs from the TBF1 gene displayed the expected resistance phenotype towards bacterial and fungal pathogens and reduced negative fitness costs associated with constitutive expression of NPR (Xu et al. 2017).
The principal functional domains of R genes are Toll/interleukin-1 receptor (TIR), nucleotide-binding site (NBS), leucine-rich repeat (LRR), LRR transmembrane domain (TrD), coiled coil (CC), nuclear localization signal (NLS), mitogen-activated protein kinase (MAPK) and MAPK kinase (MAPKK) domains. R proteins are classified into eight groups based on the arrangement of functional elements, and the major subset among them is LRR-NBS protein (LRR interact to Avr proteins with their highly variable, specificity-determining C- terminus. Mostly, R–Avr interactions occur via intermediate molecules regarded as effector targets. Mutation of host receptors mostly by altering or modifying functional domains by transgenic methods would enhance resistance and can be used in agriculture even though it is quite difficult to achieve broad-spectrum resistance by this strategy due to high R–Avr specificities. Mutation of functional domains will alter binding specificities of R proteins, which in turn strengthen the defence response either by blocking perception of specific pathogen by native R gene or by broadening the range of pathogens that can be recognized by R protein. This strategy handles pathogen recognition mechanisms only, while the rest of the defence response is part of the innate immunity only. Solanum tuberosum R3a gene (from the CNL group) confers resistance to the P. infestans Avr3aKI isolate but not to the Avr3aEM isolate. N. benthamiana expressing R3a* mutant variants showed significantly improved recognition of AVR3aEM (Chapman et al. 2014). Such mutated plant R genes enhance possibilities of recognizing pathogens/races that usually escape original R gene perception and thus confer durable resistance in field.
23.9.6 Modifying Pathogen Effector Targets in Host and Targeting Pathogen Effectors
Pathogenic virulence factors involved in virulence usually bind to certain host targets during their activity. Genetic modification of such host molecules targeted by pathogen can efficiently inhibit infection process. Modification/deletion of binding site residues in host target without compromising its other biological functions can be an alternative strategy to obstruct infection. Pathogen-derived proteins either sustain pathogen growth in susceptible host or induce HR in resistant host, and some of them have proved to be effective in enhancing plant defence response while expressed in host systems and thus can serve as functional genes for creating resistant GM plants (Silva et al. 2018). Expression of such defence promoting effectors from pathogens may confer resistance in host plants. N. benthamiana expressing P. sojae CRN effector PsCRN115 was conferred with upregulated defence responses, such as ABC transporters, cytochrome P450 and PRRs, and increased the level of plant resistance to P. capsici and P. parasitica oomycetes (Zhang et al. 2015).
23.9.7 Silencing Pathogen Genes
RNA interference is a potential strategy to control microbial pathogens of all classes – viral, bacterial, fungal (bio-necrotrophic and oomycetes) and nematode – wherein expression of essential genes of pathogen required for proliferation and establishment of infection is silenced post-transcriptionally by siRNAs complementary to pathogen-derived mRNAs. A prerequisite for RNAi is a double-stranded DNA precursor derived from a replicating viral genome or viral RNA secondary structure, which will be cleaved by DICER-like proteins (ribonuclease III – DCL), and the resulting ssRNA is loaded into silencing complex (RISC complex). siRNA-guided activated complex further recognizes and degrades the target mRNA by Argonaut (Ago) protein in cytosol (Seo et al. 2013; Weiberg et al. 2014). Conserved domains of vital genes related to cellular processes, morphogenesis or pathogenesis can be potential target sites for host-induced gene silencing, which may confer broad-spectrum durable resistance. Similarly, host genes, which are inevitable for the infection process, whose silencing does not have any/have minimal negative fitness costs also, can be targeted. Transgenic plants expressing target-specific siRNAs successfully confer resistance by sequence-specific degradation of target mRNA.
23.9.8 Transgenic Expression of Detoxifying Pathogenic Toxins
Several pathogens produce toxins, which adversely affect various metabolic pathways/processes of host system. The expression of detoxifying enzymes, either native or transgenic, specific to each pathogen under native or heterologous promoter would considerably reduce/alleviate the detrimental effects of such pathogenic toxins. For instance, detoxification of phytotoxin oxalic acid from Cryphonectria parasitica causing chestnut blight disease was achieved in American chestnut transformed with wheat gene coding for the production of the degradative enzyme, oxalate oxidase, and the resultant GM chestnut plant showed considerable reduction in chestnut blight disease development (Zhang et al. 2013). Similarly, transgenic wheat expressing a toxin-degrading enzyme from barley showed resistance to Fusarium head blight (Li et al. 2015).
23.9.9 Expression of Antimicrobial Compounds
Introduction of genes encoding antimicrobial compounds into host plant can confer efficient resistance to specific pathogens. Defensins, an antimicrobial peptide of plant origin, chitin-degrading enzyme from other microbial species, is a well-established example for this strategy. Resulting transgenic crops must display pathovar-/strain-/race-specific resistance to a respective pathogen in the field, while exhibiting normal growth and development pattern without compromising the yield and their responses to other biotic as well as abiotic stress stimuli. Spatio-temporally regulated expression of these genes using tissue-/development-specific promoters or pathogen-inducible promoter would be advantageous to minimize the negative fitness effects, if any. The effect of transgene encoded antimicrobial compound on animal/mammalian system should be assessed before employing this strategy for transforming edible crops. The manipulation of bacterial cry genes is an efficient strategy for controlling insect pests and thereby prevents vector-borne diseases (Mayee et al. 2003). A detailed understanding of the underlying molecular mechanisms of their mode of action would permit the development of much sophisticated strategies for the regulated expression of appropriate antimicrobial compounds to confer broad-spectrum durable resistance to crop plants.
23.10 Cisgenesis: Alternative to Transgenics
Cisgenesis is emerging as an efficient alternative strategy of crop improvement, which combines the beneficial aspects of classical breeding and modern biotechnology. In cisgenesis, desirable genes are transferred among crossable species/species capable of sexual hybridization (Schouten et al. 2006). In this method, the cisgene to be transferred is same as that in breeding procedure, and the resulting plant also is the same as a classically bred plant. Thus, cisgenesis eliminates biosafety concerns regarding transgenic plants, thereby overcoming the regulatory issues for field cultivation and marketing. As cisgenesis facilitates precise gene transfer, negative effects of linkage drag that associated with conventional breeding can be avoided. Additionally, cisgenesis is not time-consuming as the conventional breeding. Thus, cisgenesis can be adopted for specifically stacking resistance genes from crossable gene pools.
23.11 Conclusion
Genetic engineering technology has made tremendous contributions in developing disease-resistant crop plants. Transgenic technology has evolved multitude of options for precise alteration of crop genome, even at nucleotide level over a very short time span, and crop improvement programmes for disease resistance have taken advantage of these methods in order to circumvent the prevailing as well as expanding challenges/vulnerabilities posed by pathogenic organisms. From the incorporation of resistance genes or pathogen targeting genes and manipulation of plant’s biochemistry for harming pathogens to editing of single bases to disrupt the entire plant–pathogen interaction and thereby disease development, genetic engineering techniques have opened up every possible way for crops to combat pathogens. Sequence databases and functional annotation information are expanding on a daily basis with the establishment of high-throughput sequencing and genotyping platforms. This further unravels novel molecular and genomic targets that can be manipulated for conferring robust – broad-spectrum – durable resistance to all crop plant species.
Despite the availability of transgenic technology for developing crops with desirable resistance traits, field-level cultivation and commercialization of produce are still a major challenge in the case of these plants due to stringent regulations. Although some transgenic crops have been commercialized, many are not going for field trials. Stability of resistant phenotypes without any negative fitness effects under field conditions needs to be ensured. Protocols for transformation and regeneration of farmer-preferred cultivars/local cultivars of crop of interest need to be optimized. Stigma associated with transgenic plants is another major issue that restricts the GE crops in laboratories or experimental glasshouses. Even in the case of engineering resistance, GE strategies, which modify or alter the endogenous immune components rather than those adding/introducing novel/foreign gene or sequence or metabolic pathways, are preferred, when such issues are concerned. Ecological and health concerns regarding possibilities of horizontal gene flow, especially the bacterial marker genes used in the transformation process and the loss of endogenous agrobiodiversity, are the prominent underlying causes of such regulations. The evaluation of transgenic crop products to ensure biosafety should be followed before releasing. For transgene-free editing options opened up by third-generation genome editing tool, CRISPR/Cas is a promising strategy to overcome regulatory barriers associated with transgenic crops. Recently, the definitions of genetically modified plants are getting revised, enabling field-level cultivation of many transgenic disease-resistant crops. However, the innovations that render transgenic technology free of any risks need to be emerged for better public acceptance of genetically modified crops. But lastly, the conventional breeding techniques, which created the large pool of disease-resistant cultivars we had all this time, cannot be replaced completely and are powerful to sustain crop production, despite its shortcomings. Genetic engineering can supplement and renovate at points where breeding techniques are struggling to prevail over and that will lead us to the prime goal, sustainable crop/food production.
References
Abel PP, Nelson RS, De B, Hoffmann N, Rogers SG, Fraley RT, Beachy RN (1986) Delay of disease development in transgenic plants that express the tobacco mosaic virus coat protein gene. Science 232:738–743
Abudayyeh OO, Gootenberg JS, Essletzbichler P, Han S, Joung J, Belanto JJ, Lander ES (2017) RNA targeting with CRISPR–Cas13. Nature 550:280
Ai T, Zhang L, Gao Z, Zhu CX, Guo X (2011) Highly efficient virus resistance mediated by artificial microRNAs that target the suppressor of PVX and PVY in plants. Plant Biol 13:304–316
Ali I, Amin I, Briddon RW, Mansoor S (2013) Artificial microRNA-mediated resistance against the monopartite begomovirus cotton leaf curl Burewala virus. Virol J 10(1):231
Ali Z, Abulfaraj A, Idris A, Ali S, Tashkandi M, Mahfouz MM (2015) CRISPR/Cas9-mediated viral interference in plants. Genome Biol 16:238
Ali Z, Ali S, Tashkandi M, Zaidi SSEA, Mahfouz MM (2016) CRISPR/Cas9-mediated immunity to geminiviruses: differential interference and evasion. Sci Rep 6:1–13
Alan AR, Blowers A, Earle ED (2004) Expression of a magainin-type antimicrobial peptide gene (MSI-99) in tomato enhances resistance to bacterial speck disease. Plant Cell Rep 22(6):388–396
Allen RL, Bittner-Eddy PD, Grenville-Briggs LJ, Meitz JC, Rehmany AP, Rose LE, Beynon JL (2004) Host-parasite coevolutionary conflict between Arabidopsis and downy mildew. Science 306:1957–1960
Aldwinckle HS, Borejsza-Wysocka EE, Malnoy M, Brown SK, Norelli JL, Beer SV, Jin QL (2003) Development of fire blight resistant apple cultivars by genetic engineering. Acta Hortic 622:105–111
Antignus Y, Vunsh R, Lachman O, Pearlsman M, Maslenin L, Hananya U, Rosner A (2004) Truncated Rep gene originated from Tomato yellow leaf curl virus-Israel [Mild] confers strainspecific resistance in transgenic tomato. Ann Appl Biol 144:39–44
Andika IB, Kondo H, Tamada T (2005) Evidence that RNA silencing-mediated resistance to Beet necrotic yellow vein virus is less effective in roots than in leaves. Mol Plant Microbe Interact 18:194–204
Arce P, Moreno M, Gutierrez M, Gebauer M, Dell’Orto P, Torres H, Acuna I, Oliger P, Venegas A, Jordana X, Kalazich J, Holuigue L (1999) Enhanced resistance to bacterial infection by Erwinia carotovora subsp. atroseptica in transgenic potato plants expressing the attacin or the cecropin SB-37 genes. Am J Potato Res 76:169–177
Arif M, Azhar U, Arshad M, Zafar Y, Mansoor S, Asad S (2012) Engineering broad-spectrum resistance against RNA viruses in potato. Transgenic Res 21:303–311
Aman R, Ali Z, Butt H, Mahas A, Aljedaani F, Khan MZ, Mahfouz M (2018) RNA virus interference via CRISPR/Cas13a system in plants. Genome Biol 19:1
Bai Y, Guo Z, Wang X, Bai D, Zhang W (2009) Generation of double-virus-resistant marker-free transgenic potato plants. Prog Nat Sci 19:543–548
Balaji V, Smart CD (2012) Over-expression of snakin-2 and extensin-like protein genes restricts pathogen invasiveness and enhances tolerance to Clavibacter michiganensis subsp. michiganensis in transgenic tomato (Solanum lycopersicum). Transgenic Res 21:23–37
Baltes NJ, Hummel AW, Konecna E, Cegan R, Bruns AN, Bisaro DM, Voytas DF (2015) Conferring resistance to geminiviruses with the CRISPR–Cas prokaryotic immune system. Nat Plants 1:1–4
Baulcombe DC, Saunders GR, Bevan MW, Mayo MA, Harrison BD (1986) Expression of biologically active viral satellite RNA from the nuclear genome of transformed plants. Nature 321:446–449
Ban L, Ahmed E, Ninkovic V, Delp G, Glinwood R (2008) Infection with an insect virus affects olfactory behaviour and interactions with host plant and natural enemies in an aphid. Entomol Exp Appl 127:108–117
Barajas D, Tenllado F, González-Jara P, Martínez-García B, Atencio FA, Díaz-Ruíz JR (2004) Resistance to plum pox virus (PPV) in Nicotiana benthamiana plants transformed with the PPV HC-Pro silencing suppressor gene. Plant Pathol 88:239–248
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Bastet A, Lederer B, Giovinazzo N, Arnoux X, Retana S, Reinbold C, Lemaire O (2018) Trans-species synthetic gene design allows resistance pyramiding and broad-spectrum engineering of virus resistance in plants. Plant Biotechnol J 16:1569–1581
Bendahmane M, Chen I, Asurmendi S, Bazzini AA, Szecsi J, Beachy RN (2007) Coat protein-mediated resistance to TMV infection of Nicotiana tabacum involves multiple modes of interference by coat protein. Virology 366:107–116
Boonrod K, Galetzka D, Nagy PD, Conrad U, Krczal G (2004) Single-chain antibodies against a plant viral RNA-dependent RNA polymerase confer virus resistance. Nat Biotechnol 22:856–862
Boch J, Scholze H, Schornack S, Landgraf A, Hahn S, Kay S, Lahaye T, Nickstadt A, Bonas U (2009) Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326:1509–1512
Bonning BC, Pal N, Liu S, Wang Z, Sivakumar S, Dixon PM, Miller WA (2014) Toxin delivery by the coat protein of an aphid-vectored plant virus provides plant resistance to aphids. Nat Biotechnol 32:102
Boutrot F, Zipfel C (2017) Function, discovery, and exploitation of plant pattern recognition receptors for broad-spectrum disease resistance. Annu Rev Phytopathol 55:257–286
Bonfim K, Faria JC, Nogueira EO, Mendes ÉA, Aragão FJ (2007) RNAi-mediated resistance to bean golden mosaic virus in genetically engineered common bean (Phaseolus vulgaris). Mol Plant Microbe Interact 20:717–726
Brault V, Uzest M, Monsion B, Jacquot E, Blanc S (2010) Aphids as transport devices for plant viruses. C R Biol 333:524–538
Bruce TJ, Aradottir GI, Smart LE, Martin JL, Caulfield JC, Doherty A, Jones HD (2015) The first crop plant genetically engineered to release an insect pheromone for defence. Sci Rep 5:11183
Brunetti A, Tavazza M, Noris E, Tavazza R, Caciagli P, Ancora G, Accotto GP (1997) High expression of truncated viral Rep protein confers resistance to tomato yellow leaf curl virus in transgenic tomato plants. Mol Plant-Microbe Interact 10:571–579
Bundó M, Coca M (2016) Enhancing blast disease resistance by overexpression of the calcium-dependent protein kinase Os CPK 4 in rice. Plant Biotechnol J 14:1357–1367
Carbonell A, Takeda A, Fahlgren N, Johnson SC, Cuperus JT, Carrington JC (2014) New generation of artificial microRNA and synthetic trans-acting small interfering RNA vectors for efficient gene silencing in Arabidopsis. J Plant Physiol 165:15–29
Cao X, Lu Y, Di D, Zhang Z, Liu H, Tian L (2013) Enhanced virus resistance in transgenic maize expressing a dsRNA-specific endoribonuclease gene from E. coli. PLoS One 8:e60829
Castel SE, Martienssen RA (2013) RNA interference in the nucleus: roles for small RNAs in transcription, epigenetics and beyond. Nat Rev Genet 14:100–112
Callot C, Gallois JL (2014) Pyramiding resistances based on translation initiation factors in Arabidopsis is impaired by male gametophyte lethality. Plant Signal Behav 9:e27940
Chang C, Chen YC, Hsu YH, Wu JT, Hu CC, Chang WC, Lin NS (2005) Transgenic resistance to Cymbidium mosaic virus in Dendrobium expressing the viral capsid protein gene. Transgenic Res 14:41–46
Chapman S, Stevens LJ, Boevink PC, Engelhardt S, Alexander CJ, Harrower B, Champouret N, McGeachy K, Van Weymers PS, Chen X, Birch PR (2014) Detection of the viru lent form of AVR3a from Phytophthora infestans following artificial evolution of potato resistance gene R3a. PLoS One 9:e110158
Chandrasekaran J, Brumin M, Wolf D, Leibman D, Klap C, Pearlsman M, Gal-On A (2016) Development of broad virus resistance in non-transgenic cucumber using CRISPR/Cas9 technology. Mol Plant Pathol 17:1140–1153
Chauhan RD, Beyene G, Kalyaeva M, Fauquet CM, Taylor N (2015) Improvements in agrobacterium-mediated transformation of cassava (Manihot esculenta Crantz) for large-scale production of transgenic plants. Plant Cell Tissue Organ Cult 121:591–603
Chen C, Chen Z (2002) Potentiation of developmentally regulated plant defense response by AtWRKY18, a pathogen-induced Arabidopsis transcription factor. Plant Physiol 129:706–716
Chen YK, Lohuis D, Goldbach R, Prins M (2004) High frequency induction of RNA-mediated resistance against cucumber mosaic virus using inverted repeat constructs. Mol Breed 14:215–226
Chen SC, Liu AR, Zou ZR (2006) Overexpression of glucanase gene and defensin gene in transgenic tomato enhances resistance to Ralstonia solanacearum. Russ J Plant Physiol 53:671–677
Chen SC, Liu AR, Wang FH, Ahammed GJ (2009) Combined overexpression of chitinase and defensin genesin transgenic tomato enhances resistance to Botrytis cinerea. Afr J Biotechnol 8:5182–5188
Chen LQ, Qu XQ, Hou BH, Sosso D, Osorio S, Fernie AR, Frommer WB (2012) Sucrose efflux mediated by SWEET proteins as a key step for phloem transport. Science 335:207–211
Chen W, Qian Y, Wu X, Sun Y, Wu X, Cheng X (2014) Inhibiting replication of begomoviruses using artificial zinc finger nucleases that target viral-conserved nucleotide motif. Virus Genes 48:494–501
Cheng X, Li F, Cai J, Chen W, Zhao N, Sun Y, Guo Y, Yang X, Wu X (2015) Artificial TALE as a convenient protein platform for engineering broad-spectrum resistance to begomoviruses. Viruses 7:4772–4782
Chern M, Fitzgerald HA, Canlas PE, Navarre DA, Ronald PC (2005) Overexpression of a rice NPR1 homolog leads to constitutive activation of defense response and hypersensitivity to light. Mol Plant Microbe Interact 18:511–520
Chung BN, Yoon J, Palukaitis P (2013) Engineered resistance in potato against potato leaf roll virus, potato virus A and potato virus Y. Virus Genes 47:86–92
Chen XJ, Chen Y, Zhang LN, Xu B, Zhang JH, Chen ZX, Xu JY (2016) Overexpression of OsPGIP1 enhances rice resistance to sheath blight. Plant Dis 100:388–395
Chellappan P, Masona MV, Vanitharani R, Taylor NJ, Fauquet CM (2004) Broad spectrum resistance to ssDNA viruses associated with transgene-induced gene silencing in cassava. Plant Mol Biol 56:601–611
Chisholm ST, Coaker G, Day B, Staskawicz BJ (2006) Host-microbe interactions: shaping the evolution of the plant immune response. Cell 124:803–814
Chye ML, Zhao KJ, He ZM, Ramalingam S, Fung KL (2005) An agglutinating chitinase with two chitin-binding domains confers fungal protection in transgenic potato. Planta 220:717–730
Cillo F, Palukaitis P (2014) Transgenic resistance. Adv Virus Res 90:35–146
Cruz ARR, Aragão FJL (2014) RNAi-based enhanced resistance to Cowpea severe mosaic virus and Cowpea aphid-borne mosaic virus in transgenic cowpea. Plant Pathol 63:831–837
Coca M, Penas G, Gómez J, Campo S, Bortolotti C, Messeguer J, San Segundo B (2006) Enhanced resistance to the rice blast fungus Magnaporthe grisea conferred by expression of a cecropin A gene in transgenic rice. Planta 223:392–406
Coll NS, Epple P, Dangl JL (2011) Programmed cell death in the plant immune system. Cell Death Differ 18:1247
Collins NC, Thordal-Christensen H, Lipka V, Bau S, Kombrink E, Qiu JL, Schulze-Lefert P (2003) SNARE-protein-mediated disease resistance at the plant cell wall. Nature 425:973
Cooper B, Lapidot M, Heick JA, Dodds JA, Beachy RN (1995) A defective movement protein of TMV in transgenic plants confers resistance to multipleviruses whereas the functional analog increases susceptibility. Virology 206:307–313
Csorba T, Pantaleo V, Burgyán J (2009) RNA silencing: an antiviral mechanism. Adv Virus Res 75:35–230
Dangl JL, Jones JD (2001) Plant pathogens and integrated defence responses to infection. Nature 411:826
De Gray G, Rajasekaran K, Smith F, Sanford J, Daniell H (2001) Expression of an antimicrobial peptide via the chloroplast genome to control phytopathogenic bacteria and fungi. Plant Physiol 127:852–862
Dempsey DMA, Klessig DF (2012) SOS–too many signals for systemic acquired resistance. Trends Plant Sci 17:538–545
Deepthi DC, Makeshkumar T (2016) Elimination of cassava mosaic disease through meristem culture and field evaluation for yield loss assessment in cassava genotypes. J Root Crops 42(1):45–52
Di Nicola-Negri E, Brunetti A, Tavazza M, Ilardi V (2005) Hairpin RNA-mediated silencing of plum pox virus P1 and HC-Pro genes for efficient and predictable resistance to the virus. Transgenic Res 14:989–994
Ding SW (2010) RNA-based antiviral immunity. Nat Rev Immunol 10:632–644
Dong J, Li T, Hasi A, Zhang H (1999) Resistance of transgenic potato expressing intergenic sequence of potato leafroll virus. Virol Sin 14:66–72
Domínguez A, de Mendoza AH, Guerri J, Cambra M, Navarro L, Moreno P, Peña L (2002) Pathogen-derived resistance to Citrus tristeza virus (CTV) in transgenic Mexican lime (Citrus aurantifolia (Christ.) Swing.) plants expressing its p25 coat protein gene. Mol Plant Breed 10:1–10
Dominguez AA, Lim WA, Qi LS (2016) Beyond editing: repurposing CRISPR–Cas9 for precision genome regulation and interrogation. Nat Rev Mol Cell Biol 17:5
Dougherty WG, Lindbo JA, Smith HA, Parks TD, Swaney S, Proebsting WM (1994) RNA-mediated virus resistance in transgenic plants: exploitation of a cellular pathway possibly involved in RNA degradation. Mol Plant Microbe Interact 7:544–552
Dougherty WG, Parks TD (1995) Transgene and gene suppression: telling us something new. Curr Opin Cell Biol 7:399–405
Duan CG, Wang CH, Fang RX, Guo HS (2008) Artificial microRNAs highly accessible to targets confer efficient virus resistance in plants. J Virol 82:11084–11095
Dubey VK, Chandrasekhar K, Srivastava A, Aminuddin Singh VP, Dhar K, Arora PK (2015) Expression of coat protein gene of cucumber mosaic virus (CMV-subgroup IA) Gladiolus isolate in Nicotiana tabacum. J Plant Interact 10:296–304
Eames A, Wang MB, Smith NA, Waterhouse PM (2008) RNA silencing in plants: yesterday, today and tomorrow. Plant Physiol 147:456–468
Elayabalan S, Kalaiponmani K, Sreeramanan S, Selvarajan R, Radha P, Ramlatha M, Kumar KK, Balasubramanian P (2013) Development of Agrobacterium-mediated transformation of highly valued hill bananacultivar Virupakshi (AAB) for resistance to BBTV disease. World J Microb Biotech 29:589–596
Emani C, Garcia JM, Lopata-Finch E, Pozo MJ, Uribe P, Kim DJ, Rathore KS (2003) Enhanced fungal resistance in transgenic cotton expressing an endochitinase gene from Trichoderma virens. Plant Biotechnol J 1:321–336
Fagoaga C, López C, de Mendoza AH, Moreno P, Navarro L, Flores R, Pena L (2006) Post-transcriptional gene silencing of the p23 silencing suppressor of Citrus tristeza virus confers resistance to the virus in transgenic Mexican lime. Plant Mol Biol 60:153–165
Fahim M, Ayala-Navarrete L, Millar AA, Larkin PJ (2010) Hairpin RNA derived from viral NIa gene confers immunity to wheat streak mosaic virus infection in transgenic wheat plants. Plant Biotechnol J 8(7):821–834
Fahim M, Millar AA, Wood CC, Larkin PJ (2012) Resistance to Wheat streak mosaic virus generated by expression of an artificial polycistronic microRNA in wheat. Plant Biotechnol J 10:150–163
Faria JC, Albino MM, Dias BB, Cançado LJ, da Cunha NB, Silva LDM, Aragao FJ (2006) Partial resistance to Bean golden mosaic virus in a transgenic common bean (Phaseolus vulgaris L.) line expressing a mutated rep gene. Plant Sci 171:565–571
Febres VJ, Lee RF, Moore GA (2008) Transgenic resistance to Citrus tristeza virus in grapefruit. Plant Cell Rep 27:93–104
Fellers JP, Collins GB, Hunt AG (1998) The NIa-proteinase of different plant potyviruses provides specific resistance to viral infection. Crop Sci 38:1309–1319
Fermin G, Inglessis V, Garboza C, Rangel S, Dagert M, Gonsalves D (2004) Technology transfer for engineered resistance against PRSV in Venezuelan transgenic papayas. Plant Dis 88:516–522
Ferreira SA, Pitz KY, Manshardt R, Zee F, Fitch M, Gonsalves D (2002) Virus coat protein transgenic papaya provides practical control of papaya ringspot virus in Hawaii. Plant Dis 86:101–105
Ferrari S, Sella L, Janni M, De Lorenzo G, Favaron F, D’ovidio R (2012) Transgenic expression of polygalacturonase-inhibiting proteins in Arabidopsis and wheat increases resistance to the flower pathogen Fusarium graminearum. Plant Biol 14:31–38
Ferrari S, Savatin DV, Sicilia F, Gramegna G, Cervone F, De Lorenzo G (2013) Oligogalacturonides: plant damage-associated molecular patterns and regulators of growth and development. Front Plant Sci 4:49
Fister AS, Landherr L, Maximova SN, Guiltinan MJ (2018) Transient expression of CRISPR/Cas9 machinery targeting TcNPR3 enhances defense response in Theobroma cacao. Front Plant Sci 9:268
Flor HH (1971) Current status of the gene-for-gene concept. Annu Rev Phytopathol 9:275–296
Flores R, Hernández C, Alba AEMD, Daròs JA, Serio FD (2005) Viroids and viroid-host interactions. Annu Rev Phytopathol 43:117–139
Flores R, Navarro B, Kovalskaya N, Hammond RW, Di Serio F (2017) Engineering resistance against viroids. Curr Opin Virol 26:1–7
Floyd-Smith G, Slattery E, Lengyel P (1981) Interferon action: RNA cleavage pattern of a (2′-5′) oligoadenylate-dependent endonuclease. Science 212:1030–1032
Fonseca JP, Mysore KS (2019) Genes involved in nonhost disease resistance as a key to engineer durable resistance in crops. Plant Sci 279:108–116
Fradin EF, Abd-El-Haliem A, Masini L, van den Berg GC, Joosten MH, Thomma BP (2011) Interfamily transfer of tomato Ve1 mediates Verticillium resistance in Arabidopsis. Plant Physiol 156:2255–2265
Frischmuth T, Stanley J (1998) Recombination between viral DNA and the transgenic coat protein gene of African cassava mosaic geminivirus. J Gen Virol 79:1265–1271
Furutani N, Hidaka S, Kosaka Y, Shizukawa Y, Kanematsu S (2006) Coat protein gene-mediated resistance to soybean mosaic virus in transgenic soybean. Breed Sci 56:119–124
Fuentes A, Ramos PL, Fiallo E, Callard D, Sánchez Y, Peral R (2006) Intron-hairpin RNA derived from replication association protein C1 gene confers immunity to tomato yellow leaf curl virus infection in transgenic tomato plants. Transgenic Res 15:291–304
Ganesan U, Suri SS, Rajasubramaniam S, Rajam MV, Dasgupta I (2009) Transgenic expression of coat protein gene of Rice tungro bacilliform virus in rice reduces the accumulation of viral DNA in inoculated plants. Virus Genes 39:113–119
Gargouri-Bouzid R, Jaoua L, Rouis S, Saïdi MN, Bouaziz D, Ellouz R (2006) PVY-resistant transgenic potato plants expressing an anti-NIa protein scFv antibody. Mol Biotechnol 33:133–140
Germundsson A, Valkonen JP (2006) P1-and VPg-transgenic plants show similar resistance to Potato virus A and may compromise long distance movement of the virus in plant sections expressing RNA silencing-based resistance. Virus Res 116:208–213
Galvez LC, Banerjee J, Pinar H, Mitra A (2014) Engineered plant virus resistance. Plant Sci 228:11–25
Gao AG, Hakimi SM, Mittanck CA, Wu WBM, Stark DM, Shah DM, Liang J, Rommens CM (2000) Fungal pathogen protection in potato by expression of a plant defensin peptide. Nat Biotechnol 18:1307–1310
Gao L, Ding X, Li K, Liao W, Zhong Y, Ren R, Liu Z, Adhimoolam K, Zhi H (2015) Characterization of soybean mosaic virus resistance derived from inverted repeat-SMV-HC-Pro genes in multiple soybean cultivars. Theor Appl Genet 128:1489–1505
Gil M, Esteban O, García JA, Pena L, Cambra M (2011) Resistance to plum pox virus in plants expressing cytosolic and nuclear single-chain antibodies against the viral RNA NIb replicase. Plant Pathol 60:967–976
Gill US, Lee S, Mysore KS (2015) Host versus nonhost resistance: distinct wars with similar arsenals. Phytopathology 105:580–587
Glazebrook J (2005) Contrasting mechanisms of defense against biotrophic and necrotrophic pathogens. Annu Rev Phytopathol 43:205–227
Goggin FL, Jia L, Shah G, Hebert S, Williamson VM, Ullman DE (2006) Heterologous expression of the Mi-1.2 gene from tomato confers resistance against nematodes but not aphids in eggplant. Mol Plant-Microbe Interact 19:383–388
Golemboski DB, Lomonossoff GP, Zaitlin M (1990) Plants transformed with a tobacco mosaic virus nonstructural gene sequence are resistant to the virus. Proc Natl Acad Sci 87:6311–6315
Guo Y, Ruan MH, Yao W, Chen L, Chen RK, Zhang MQ (2008) Difference of coat protein mediated resistance to sugarcane mosaic virus between Badila and Funong 91–4621. J Fujian Agric For Univ (Natural Science Edition) 37:7–12
Guo HJ, Huang LC, Sun YC, Guo H, Ge F (2016) The contrasting effects of elevated CO2 on TYLCV infection of tomato genotypes with and without the resistance gene, Mi-1.2. Front Plant Sci 7:1680
Gubba A, Gonsalves C, Stevens MR, Tricoli DM, Gonsalves D (2002) Combining transgenic and natural resistance to obtain broad resistance to tospovirus infection in tomato (Lycopersicon esculentum Mill). Mol Breed 9:13–23
Gururani MA, Venkatesh J, Upadhyaya CP, Nookaraju A, Pandey SK, Park SW (2012) Plant disease resistance genes: current status and future directions. Physiol Mol Plant Pathol 78:51–65
Hannon GJ (2002) RNA interference. Nature 418:244–251
Hashmi JA, Zafar Y, Arshad M, Mansoor S, Asad S (2011) Engineering cotton (Gossypium hirsutum L.) for resistance to cotton leaf curl disease using viral truncated AC1 DNA sequences. Virus Genes 42:286–296
Hao G, Pitino M, Duan Y, Stover E (2016) Reduced susceptibility to Xanthomonas citri in transgenic citrus expressing the FLS2 receptor from Nicotiana benthamiana. Mol Plant-Microbe Interact 29:132–142
Harris CJ, Slootweg EJ, Goverse A, Baulcombe DC (2013) Stepwise artificial evolution of a plant disease resistance gene. Proc Natl Acad Sci 110:21189–21194
Harrison BD, Mayo MA, Baulcombe DC (1987) Virus resistance in transgenic plants that express cucumber mosaic virus satellite RNA. Nature 328:799–802
Hamilton AJ, Baulcombe DC (1999) A species of small antisense RNA in posttranscriptional gene silencing in plants. Science 286:950–952
He X, Miyasaka SC, Fitch MM, Moore PH, Zhu YJ (2008) Agrobacterium tumefaciens-mediated transformation of taro (Colocasia esculenta (L.) Schott) with a rice chitinase gene for improved tolerance to a fungal pathogen Sclerotium rolfsii. Plant Cell Rep 27:903–909
Himmelbach A, Liu L, Zierold U, Altschmied L, Maucher H, Beier F, Muller D, Hensel G, Heise A, Schutzendubel A, Kumlehn J, Schweizer P (2010) Promoters of the barley germin-like GER4 gene cluster enable strong transgene expression in response to pathogen attack. Plant Cell 22(3):937–952
Hood MI, Skaar EP (2012) Nutritional immunity: transition metals at the pathogen–host interface. Nat Rev Microbiol 10:525
Horvath DM, Stall RE, Jones JB, Pauly MH, Vallad GE, Dahlbeck D, Scott JW (2012) Transgenic resistance confers effective field level control of bacterial spot disease in tomato. PLoS One 7:e42036
Huang Y, Nordeen RO, Di M, Owens LD, Mc Beth JH (1997) Expression of an engineered cecropin gene cassette in transgenic tobacco plants confers disease resistance to Pseudomonas syringae pv. tabaci. Phytopathology 87:494–499
Kamala S, Makeshkumar T (2015) Transcriptome profiling of in vitro lines propagated from corm bud tip of elephant foot yam (Amorphophallus paeoniifolius) approves complete potyviruses elimination. Eur J Plant Pathology 143(2):363–371
Ingwell LL, Eigenbrode SD, Bosque-Pérez NA (2012) Plant viruses alter insect behavior to enhance their spread. Sci Rep 2:578
Ilardi V, Tavazza M (2015) Biotechnological strategies and tools for Plum pox virus resistance: trans-, intra-, cis-genesis, and beyond. Front Plant Sci 6:379
Iwanami T, Shimizu T, Ito T, Hirabayashi T (2004) Tolerance to citrus mosaic virus in transgenic trifoliate orange lines harboring capsid polyprotein gene. Plant Dis 88(8):865–868
Jan PS, Huang HY, Chen HM (2010) Expression of a synthesized gene encoding cationic peptide cecropin B in transgenic tomato plants protects against bacterial diseases. Appl Environ Microbiol 76:769–775
Javaid S, Amin I, Jander G, Mukhtar Z, Saeed NA, Mansoor S (2016) A transgenic approach to control hemipteran insects by expressing insecticidal genes under phloem-specific promoters. Sci Rep 6:34706
Jha S, Tank HG, Prasad BD, Chattoo BB (2009) Expression of Dm-AMP1 in rice confers resistance to Magnaporthe oryzae and Rhizoctonia solani. Transgenic Res 18:59–69
Ji X, Zhang H, Zhang Y, Wang Y, Gao C (2015) Establishing a CRISPR–Cas-like immune system conferring DNA virus resistance in plants. Nat Plants 1:1–4
Jia H, Orbovic V, Jones JB, Wang N (2016) Modification of the PthA4 effector binding elements in type I Cs LOB 1 promoter using Cas9/sgRNA to produce transgenic Duncan grapefruit alleviating XccΔpthA4: dCs LOB 1.3 infection. Plant Biotechnol J 14:1291–1301
Jo KR, Kim CJ, Kim SJ, Kim TY, Bergervoet M, Jongsma MA, Vossen JH (2014) Development of late blight resistant potatoes by cisgene stacking. BMC Biotechnol 14:50
Jo KM, Jo Y, Choi H, Chu H, Lian S, Yoon JY, Choi SK, Kim KH, Cho WK (2015) Development of genetically modified chrysanthemums resistant to Chrysanthemum stunt viroid using sense and antisense RNAs. Sci Hortic 195:17–24
Johnson R (1981) Durable resistance: definition of, genetic control, and attainment in plant breeding. Phytopathology 71:567–568
Jones RAC (2014) Plant virus ecology and epidemiology: historical perspectives, recent progress and future prospects. Ann Appl Biol 164:320–347
Jones JD, Dangl JL (2006) The plant immune system. Nature 444:323
Jones JD, Witek K, Verweij W, Jupe F, Cooke D, Dorling S, Foster S (2014) Elevating crop disease resistance with cloned genes. Philos Trans R Soc B 369:0087
Jones-Rhoades MW, Bartel DP, Bartel B (2006) MicroRNAs and their regulatory roles in plants. Annu Rev Plant Biol 57:19–53
Joubert DA, Kars I, Wagemakers L, Bergmann C, Kemp G, Vivier MA, van Kan JA (2007) A polygalacturonase-inhibiting protein from grapevine reduces the symptoms of the endopolygalacturonase BcPG2 from Botrytis cinerea in Nicotiana benthamiana leaves without any evidence for in vitro interaction. Mol Plant Microbe Interact 20:392–402
Kamachi S, Mochizuki A, Nishiguchi M, Tabei Y (2007) Transgenic Nicotiana benthamiana plants resistant to cucumber green mottle mosaic virus based on RNA silencing. Plant Cell Rep 26:1283–1288
Kawazu Y, Fujiyama R, Sugiyama K, Sasaya T (2006) A transgenic lettuce line with resistance to both lettuce big-vein associated virus and mirafiori lettuce virus. J Am Soc Horticult Sci 131:760–763
Kachroo A, Robin GP (2013) Systemic signaling during plant defense. Curr Opin Plant Biol 16:527–533
Kaur J, Sagaram US, Shah D (2011) Can plant defensins be used to engineer durable commercially useful fungal resistance in crop plants. Fungal Biol Rev 25:128–135
Kertbundit S, Pongtanom N, Ruanjan P, Chantasingh D, Tanwanchai A, Panyim S, Juříček M (2007) Resistance of transgenic papaya plants to papaya ringspot virus. Biol Plant 51:333–339
Kerr CH, Wang QS, Keatings K, Khong A, Allan D, Yip CK, Jan E (2015) The 5′ untranslated region of a novel infectious molecular clone of the dicistrovirus cricket paralysis virus modulates infection. J Virol 89:5919–5934
Khoury J, Singh R, Boucher A, Coombs D (1988) Concentration and distribution of mild and severe strains of potato spindle tuber viroid in cross-protected tomato plants. Phytopathology 78:1331–1336
Khan RS, Sjahril R, Nakamur I (2008) Production of transgenic potato exhibiting enhanced resistance to fungal infections and herbicide applications. Plant Biotechnol Rep 2:13–20
Kim SJ, Paek KH, Kim BD (1995) Delay of disease development in transgenic petunia plants expressing cucumber mosaic virus I17N-satellite RNA. J Am Soc Hortic Sci 120:353–359
Kim SJ, Lee SJ, Kim BD, Paek KH (1997) Satellite-RNA-mediated resistance to cucumber mosaic virus in transgenic plants of hot pepper (Capsicum annuum cv. Golden Tower). Plant Cell Rep 16:825–830
Kim JK, Jang IC, Wu R, Zuo WN, Boston RS, Lee YH, Ahn IP, Nahm BH (2003) Co-expression of a modified maize ribosome-inactivating protein and a rice basic chitinase gene in transgenic rice plants confers enhanced resistance to sheath blight. Transgenic Res 12:475–484
Kim SH, Qi D, Ashfield T, Helm M, Innes RW (2016) Using decoys to expand the recognition specificity of a plant disease resistance protein. Science 351:684–687
Kishimoto K, Nishizawa Y, Tabei Y, Hibi T, Nakajima M, Akutsu K (2002) Detailed analysis of rice chitinase gene expression in transgenic cucumber plants showing different levels of disease resistance to gray mold (Botrytis cinerea). Plant Sci 162:655–662
Koo JC, Asurmendi S, Bick J, Woodford-Thomas T, Beachy RN (2004) Ecdysone agonist inducible expression of a coat protein gene from tobacco mosaic virus confers viral resistance in transgenic Arabidopsis. Plant J 37:439–448
Kollàr À, Dalmay T, Burgyàn J (1993) Defective interfering RNA-mediated resistance against cymbidium ringspot tombusvirus in transgenic plants. Virology 193:313–318
Koseoglou E (2017) The study of SlPMR4 CRISPR/Cas9-mediated tomato allelic series for resistance against powdery mildew (Doctoral dissertation, Wageningen University and Research, Wageningen)
Kouassi NK, Chen L, Siré C, Bangratz-Reyser M, Beachy RN, Fauquet CM, Brugidou C (2006) Expression of rice yellow mottle virus coat protein enhances virus infection in transgenic plants. Arch Virol 151:2111–2122
Krubphachaya P, Juricek M, Kertbundit S (2007) Induction of RNA-mediated resistance to papaya ringspot virus type W. J Biochem Mol Biol 40:404–411
Kung YJ, Bau HJ, Wu YL, Huang CH, Chen TM, Yeh SD (2009) Generation of transgenic papaya with double resistance to papaya ringspot virus and papaya leaf-distortion mosaic virus. Phytopathology 99:1312–1320
Kung YJ, Yu TA, Huang CH, Wang HC, Wang SL, Yeh SD (2010) Generation of hermaphrodite transgenic papaya lines with virus resistance via transformation of somatic embryos derived from adventitious roots of in vitro shoots. Transgenic Res 19:621–635
Langenberg WG, Zhang L, Giunchedi L, Mitra A (1997) Transgenic tobacco plants expressing the bacterial mc gene resist virus infection. Mol Breed 3:391–399
Lacombe S, Rougon-Cardoso A, Sherwood E, Peeters N, Dahlbeck D, Van Esse HP, Jones JD (2010) Interfamily transfer of a plant pattern-recognition receptor confers broad-spectrum bacterial resistance. Nat Biotechnol 28:365
Langenbach C, Schultheiss H, Rosendahl M, Tresch N, Conrath U, Goellner K (2016) Interspecies gene transfer provides soybean resistance to a fungal pathogen. Plant Biotechnol J 14:699–708
Lapidot M, Gafny R, Ding B, Wolf S, Lucas WJ, Beachy RN (1993) A dysfunctional movement protein of tobacco mosaic virus that partially modifies the plasmodesmata and limits virus spread in transgenic plants. Plant J 4:959–970
Lee SC, Hwang IS, Choi HW, Hwang BK (2008) Involvement of the pepper antimicrobial protein CaAmp1 gene in broad spectrum disease resistance. Plant Physiol 148:1004–1020
Lee YH, Jung M, Shin SH, Lee JH, Choi SH, Her NH, Lee JH, Ryu KH, Paek KY, Harn CH (2009) Transgenic peppers that are highly tolerant to a new CMV pathotype. Plant Cell Rep 28:223–232
Lee G, Shim HK, Kwon MH, Son SH, Kim KY, Park EY, Yang JK, Lee TK, Auh CK, Kim D Kim YS (2013) RNA virus accumulation is inhibited by ribonuclease activity of 3D8 scFv in transgenic Nicotiana tabacum. Plant Cell Tissue Organ Cult 115:189–197
Lee HA, Kim S, Kim S, Choi D (2017) Expansion of sesquiterpene biosynthetic gene clusters in pepper confers nonhost resistance to the Irish potato famine pathogen. New Phytol 215:1132–1143
Leibman D, Wolf D, Saharan V, Zelcer A, Arazi T, Yoel S, Gaba V, Gal-On A (2011) A high level of transgenic viral small RNA is associated with broad potyvirus resistance in cucurbits. Mol Plant Microbe Interact 24:1220–1238
Li T, Liu B, Spalding MH, Weeks DP, Yang B (2012) High-efficiency TALEN-based gene editing produces disease-resistant rice. Nat Biotechnol 30:390
Li X, Shin S, Heinen S, Dill-Macky R, Berthiller F, Nersesian N, Clemente T, McCormick S Muehlbauer GJ (2015) Transgenic wheat expressing a barley UDP-glucosyltransferase detoxifies deoxynivalenol and provides high levels of resistance to Fusarium graminearum. Mol Plant Microbe Interact 28:1237–1246
Lionetti V, Fabri E, De Caroli M, Hansen AR, Willats WG, Piro G, Bellincampi D (2017) Three pectin methylesterase inhibitors protect cell wall integrity for Arabidopsis immunity to Botrytis. Plant Physiol 173:1844–1863
Liu XH, Zhang HW, Liu X, Liu XJ, Tan ZB, Rong TZ (2005) Isolation of the capsid protein gene of maize dwarf mosaic virus and its transformation in maize. Sheng wu gong cheng xue bao Chin J Biotechnol 21:144–148
Liu X, Tan Z, Li W, Zhang H, He D (2009) Cloning and transformation of SCMV CP gene and regeneration of transgenic maize plants showing resistance to SCMV strain MDB. Afr J Biotechnol 8:3747–3753
Liu N, Zhang X, Sun Y, Wang P, Li X, Pei Y, Hou Y (2017) Molecular evidence for the involvement of a polygalacturonase-inhibiting protein, GhPGIP1, in enhanced resistance to Verticillium and Fusarium wilts in cotton. Sci Rep 7:39840
Liu H, Soyars CL, Li J, Fei Q, He G, Peterson BA, Wang X (2018) CRISPR/Cas9-mediated resistance to cauliflower mosaic virus. Plant Direct 2:e00047
Lin CY, Ku HM, Chiang YH, Ho HY, Yu TA, Jan FJ (2012) Development of transgenic watermelon resistant to cucumber mosaic virus and Watermelon mosaic virus by using a single chimeric transgene construct. Transgenic Res 21:983–993
Lin KY, Hsu YH, Chen HC, Lin NS (2013) Transgenic resistance to Bamboo mosaic virus by expression of interfering satellite RNA. Mol Plant Pathol 14:693–707
Lindbo JA, Dougherty WG (1992a) Pathogen-derived resistance to a potyvirus: immune and resistant phenotypes in transgenic tobacco expressing altered forms of a potyvirus coat protein nucleotide sequence. Mol Plant Microbe Interact 5(2):144–153
Lindbo JA, Dougherty WG (1992b) Untranslatable transcripts of the tobacco etch virus coat protein gene sequence can interfere with tobacco etch virus replication in transgenic plants and protoplasts. Virology 189(2):725–733
Lindbo JA, Dougherty WG (2005) Plant pathology and RNAi: A brief history. Ann Rev Phytopath 43:191–204
Li Q, Lawrence CB, Xing HY, Babbitt RA, Bass WT, Maiti IB, Everett NP (2001) Enhanced disease resistance conferred by expression of an antimicrobial magainin analogue in transgenic tobacco. Planta 212:635–639
Li H, Conner RL, Chen Q, Graf RJ, Laroche A, Ahmad F, Kuzyk AD (2005) Promising genetic resources for resistance to wheat streak mosaic virus and the wheat curl mite in wheat-Thinopyrum partial amphiploids and their derivatives. Genet Resour Crop Evol 51:827–835
Liao LJ, Pan IC, Chan YL, Hsu YH, Chen WH, Chan MT (2004) Transgene silencing in Phalaenopsis expressing the coat protein of Cymbidium mosaic virus is a manifestation of RNAmediated resistance. Mol Breed 13:229–242
Lim HS, Ko TS, Hobbs HA, Lambert KN, Yu JM, McCoppin NK, Korban SS, Hartman GL, Domier LL (2007) Soybean mosaic virus helper component-protease alters leaf morphology and reduces seed production in transgenic soybean plants. Phytopathology 97:366–372
Lopez-Ochoa L, Ramirez-Prado J, Hanley-Bowdoin L (2006) Peptide aptamers that bind to a geminivirus replication protein interfere with viral replication in plant cells. J Virol 80:5841–5853
Lopez C, Cervera M, Fagoaga C, Moreno P, Navarro L, Flores R, Pena L (2010) Accumulation of transgene-derived siRNAs is not sufficient for RNAi-mediated protection against Citrus tristeza virus in transgenic Mexican lime. Mol Plant Pathol 11:33–41
Lodge JK, Kaniewski WK, Tumer NE (1993) Broad-spectrum virus resistance in transgenic plants expressing pokeweed antiviral protein. Proc Natl Acad Sci 90:7089–7093
Loeza-Kuk E, Gutiérrez-Espinosa MA, Ochoa-Martínez DL, Villegas-Monter Á, Mora-Aguilera G, Palacios-Torres EC, Pérez-Molphe-Balch E (2011) Análisis de la resistencia en pomelo y limón mexicano transformados con el gen p25 del Citrus tristeza virus. Agrociencia 45(1):55–65
Lucioli A, Noris E, Brunetti A, Tavazza R, Ruzza V, Castillo AG, Bejarano ER, Accotto GP Tavazza M (2003) Tomato yellow leaf curl Sardinia virus rep-derived resistance to homologous and heterologous geminiviruses occurs by different mechanisms and is overcome if virus-mediated transgene silencing is activated. J Virol 77:6785–6798
Ludlow EJ, Mouradva A, Spangenberg G (2009) Post transcriptional gene silencing as an efficient tool for engineering resistance to white clover mosaic virus in white clover. J Pl Physiol 166:1557–1567
Ma J, Song Y, Wu B, Jiang M, Li K, Zhu C, Wen F (2011) Production of transgenic rice new germplasm with strong resistance against two isolations of Rice stripe virus by RNA interference. Transgenic Res 20:1367–1377
Ma Z, Song T, Zhu L, Ye W, Wang Y, Shao Y, Tyler BM (2015) A Phytophthora sojae glycoside hydrolase 12 protein is a major virulence factor during soybean infection and is recognized as a PAMP. Plant Cell 27:2057–2072
Ma J, Chen J, Wang M, Ren Y, Wang S, Lei C, Cheng Z (2018) Disruption of OsSEC3A increases the content of salicylic acid and induces plant defense responses in rice. J Exp Bot 69:1051–1064
Maiti IB, Von Lanken C, Hong Y, Dey N, Hunt AG (1999) Expression of multiple virus-derived resistance determinants in transgenic plants does not lead to additive resistance properties. J Plant Biochem Biotechnol 8(2):67–73
Malnoy M, Venisse JS, Brisset MN, Chevreau E (2003) Expression of bovine lactoferrin cDNA confers resistance to Erwinia amylovora in transgenic pear. Mol Breed 12:231–244
Makeshkumar T, Varma A, Singh KK, Malathi VG, Duttagupta M (2002) Coat protein mediated resistance against potato virus Y in transgenic tobacco. Ind Phytopath 55(2):187–194
Makarova KS, Wolf YI, Alkhnbashi OS, Costa F, Shah SA, Saunders SJ, Barrangou R, Brouns SJ, Charpentier E, Haft DH, Horvath P (2015) An updated evolutionary classification of CRISPR–Cas systems. Nat Rev Microbiol 13:722–736
Mäki-Valkama T, Valkonen JP, Lehtinen A, Pehu E (2001) Protection against potato virus Y (PVY) in the field in potatoes transformed with the PVY P1 gene. Am J Potato Res 78:209–214
Matoušek J, Trněná L, Rakouský S, Riesner D (1994) Inhibition of potato spindle tuber viroid (PSTVd) infectivity with antisense RNA transcripts. J Phytopathol 141:10–24
Malnoy M, Viola R, Jung MH, Koo OJ, Kim S, Kim JS, Velasco R, Nagamangala Kanchiswamy C (2016) DNA-free genetically edited grapevine and apple protoplast using CRISPR/Cas9 ribonucleoproteins. Front Plant Sci 7:1904
Manfredini C, Sicilia F, Ferrari S, Pontiggia D, Salvi G, Caprari C, De Lorenzo G (2005) Polygalacturonase-inhibiting protein 2 of Phaseolus vulgaris inhibits BcPG1, a polygalacturonase of Botrytis cinerea important for pathogenicity, and protects transgenic plants from infection. Physiol Mol Plant Pathol 67:108–115
Mayee CD, Singh P, Dongre AB, Rao MRK, Raj S (2003) Transgenic Bt cotton. CICR Technical Bulletin No.22. Nagpur. Central Institute for Cotton Research
Mawassi M, Gera A (2012) Controlling plant response to the environment: viral diseases. In: Altman A, Hasegawa PM (eds) Plant biotechnology and agriculture: prospects for the 21st century, pp 343–352
Macovei A, Sevilla NR, Cantos C, Jonson GB, Slamet-Loedin I, Čermák T, Voytas DF, Choi IR, Chadha-Mohanty P (2018) Novel alleles of rice eIF4G generated by CRISPR/Cas9-targeted mutagenesis confer resistance to Rice tungro spherical virus. Plant Biotechnol J 16(11):1918–1927
McCue KF, Ponciano G, Rockhold DR, Whitworth JL, Gray SM, Fofanov Y, Belknap WR (2012) Generation of PVY coat protein siRNAs in transgenic potatoes resistant to PVY. Am J Potato Res 89:374–383
Mehta R, Radhakrishnan T, Kumar A, Yadav R, Dobaria JR, Thirumalaisamy PP, Jain RK, Chigurupati P (2013) Coat protein-mediated transgenic resistance of peanut (Arachis hypogaea L.) to peanut stem necrosis disease through agrobacterium-mediated genetic transformation. Indian J Virol 24:205–213
Mentag R, Lukevich M, Morency MJ, Seguin A (2003) Bacterial disease resistance of transgenic hybrid poplar expressing the synthetic antimicrobial peptide D4E1. Tree Physiol 23:405–411
Minoia S, Carbonell A, Di Serio F, Gisel A, Carrington JC, Navarro B, Flores R (2014) Specific argonautes selectively bind small RNAs derived from potato spindle tuber viroid and attenuate viroid accumulation in vivo. J Virol 88:11933–11945
Missiou A, Kalantidis K, Boutla A, Tzortzakaki S, Tabler M, Tsagris M (2004) Generation of transgenic potato plants highly resistant to potato virus Y (PVY) through RNA silencing. Mol Breed 14:185–197
Mitra A, Higgins DW, Langenberg WG, Nie H, Sengupta DN, Silverman RH (1996) A mammalian 2-5A system functions as an antiviral pathway in transgenic plants. Proc Natl Acad Sci 93:6780–6785
Moravčíková J, Matušíková I, Libantova J, Bauer M, Mlynárová LU (2004) Expression of a cucumber class III chitinase and Nicotiana plumbaginifolia class I glucanase genes in transgenic potato plants. Plant Cell Tissue Organ Cult 79:161–168
Moravčíková J, Libantová J, Heldák J, Salaj J, Bauer M, Matušíková I, Gálová Z, Mlynárová Ľ (2007) Stress-induced expression of cucumber chitinase and Nicotiana plumbaginifolia β-1,3-glucanase genes in transgenic potato plants. Acta Physiol Plant 29:133–141
Mlotshwa S, Verver J, Sithole-Niang I, Prins M, Van Kammen AB, Wellink J (2002) Transgenic plants expressing HC-Pro show enhanced virus sensitivity while silencing of the transgene results in resistance. Virus Genes 25:45–57
Mundembe R, Matibiri A, Sithole-Niang I (2009) Transgenic plants expressing the coat protein gene of cowpea aphid-borne mosaic potyvirus predominantly convey the delayed symptom development phenotype. Afr J Biotechnol 8:2682–2690
Muniz FR, De Souza AJ, Stipp LCL, Schinor E, Freitas W, Harakava R, Stach-Machado DR, Rezende JAM, Mourão Filho FAA, Mendes BMJ (2012) Genetic transformation of Citrus sinensis with Citrus tristeza virus (CTV) derived sequences and reaction of transgenic lines to CTV infection. Biol Plant 56:162–166
Namukwaya B, Tripathi L, Tripathi JN, Arinaitwe G, Mukasa SB, Tushemereirwe WK (2012) Transgenic banana expressing Pflp gene confers enhanced resistance to Xanthomonas wilt disease. Transgenic Res 21:855–865
Navarro B, Gisel A, Rodio ME, Delgado S, Flores R, Di Serio F (2012) Viroids: how to infect a host and cause disease without encoding proteins. Biochimie 94:1474–1480
Nekrasov V, Wang C, Win J, Lanz C, Weigel D, Kamoun S (2017) Rapid generation of a transgene-free powdery mildew resistant tomato by genome deletion. Sci Rep 7:482
Nelson RS, McCormick SM, Delannay X, Dubé P, Layton J, Anderson EJ, Fraley RT (1988) Virus tolerance, plant performance of transgenic tomato plants expressing coat protein from tobacco mosaic virus. Biotechnology 6:403
Niblett CL, Dickson E, Fernow KH, Horst RK, Zaitlin M (1978) Cross protection among four viroids. Virology 91:198–203
Nickel H, Kawchuk L, Twyman RM, Zimmermann S, Junghans H, Winter S, Fischer R, Prüfer D (2008) Plantibody-mediated inhibition of the potato leafroll virus P1 protein reduces virus accumulation. Virus Res 136:140–145
Nishizawa Y, Saruta M, Nakazono K, Nishio Z, Soma M, Yoshida T, Nakajima E, Hibi T (2003) Characterization of transgenic rice plants over-expressing the stress-inducible β-glucanase gene Gns1. Plant Mol Biol 51:143–152
Niu QW, Lin SS, Reyes JL, Chen KC, Wu HW, Yeh SD, Chua NH (2006) Expression of artificial microRNAs in transgenic Arabidopsis thaliana confers virus resistance. Nat Biotechnol 24:1420
Nobori T, Velásquez AC, Wu J, Kvitko BH, Kremer JM, Wang Y, Tsuda K (2018) Transcriptome landscape of a bacterial pathogen under plant immunity. Proc Natl Acad Sci 115:E3055–E3064
Norelli JL, Mills JZ, Momol MT, Aldwinkle HS (1998) Effect of cercropintype transgenes on fire blight resistanc of apple. Acta Hortic 489:273–278
Nowara D, Gay A, Lacomme C, Shaw J, Ridout C, Douchkov D, Schweizer P (2010) HIGS: host-induced gene silencing in the obligate biotrophic fungal pathogen Blumeria graminis. Plant Cell 22:3130–3141
Nomura K, Ohshima K, Anai T, Uekusa H, Kita N (2004) RNA silencing of the introduced coat protein gene of turnip mosaic virus confers broad-spectrum resistance in transgenic Arabidopsis. Phytopathology 94:730–736
Ntui VO, Azadi P, Thirukkumaran R, Chin DP, Nakamura I, Mii M (2011) Increased resistance to Fusarium wilt in transgenic tobacco co-expressing chitinase and wasabi defensin genes. Plant Pathol 60:221e231
Ntui VO, Kong K, Khan RS, Igawa T, Janavi GJ, Rabindran R, Nakamura I, Mii M (2015) Resistance to Sri Lankan cassava mosaic virus (SLCMV) in genetically engineered cassava cv. KU50 through RNA silencing. PLoS One 10:e0120551
Nyaboga NE, Ateka ME, Bulimo DW (2008) Serological detection of virus diseases of sweet potato in Kenya. J Appl Biosci 7:222–229
Ogwok E, Odipio J, Halsey M, Gaitán-Solís E, Bua A, Taylor NJ, Fauquet CM, Alicai T (2012) Transgenic RNA interference (RNAi)-derived field resistance to cassava brown streak disease. Mol Plant Pathol 13:1019–1031
Okada Y, Saito A, Nishiguchi M, Kimura T, Mori M, Hanada K, Sakai J, Miyazaki C, Matsuda Y, Murata T (2001) Virus resistance in transgenic sweetpotato [Ipomoea batatas L.(lam)] expressing the coat protein gene of sweet potato feathery mottle virus. Theor Appl Genet 103:743–751
Okada Y, Saito A (2008) Evaluation of resistance to complex infection of SPFMVs in transgenic sweet potato. Breed Sci 58:243–250
Okada Y, Yoshinaga M (2010) Advanced resistance to Sweetpotato feathery mottle virus (SPFMV) in transgenic sweetpotato. Sweetpotato Res Front 23:4
Okubara P, Blechl A, McCormick S, Alexander N, Dill-Macky R, Hohn T (2002) Engineering deoxynivalenol metabolism in wheat through the expression of a fungal trichothecene acetyltransferase gene. Theor Appl Genet 106:74–83
Ordiz MI, Magnenat L, Barbas CF, Beachy RN (2010) Negative regulation of the RTBV promoter by designed zinc finger proteins. Plant Mol Biol 72:621–630
Ortigosa A, Gimenez-Ibanez S, Leonhardt N, Solano R (2019) Design of a bacterial speck resistant tomato by CRISPR/Cas9-mediated editing of SlJAZ2. Plant Biotechnol J 17(3):665–673
Ortigosa A, Gimenez-Ibanez S, Leonhardt N, Solano R (2018) Design of a bacterial speck resistant tomato by CRISPR/Cas9-mediated editing of SlJAZ2. Plant Biotechnol J 17:665–673
Osusky M, Osuska L, Hancock RE, Kay WW, Misra S (2004) Transgenic Potatoes Expressing a Novel Cationic Peptide are Resistant to Late Blight and Pink Rot. Transgenic Res 13:181–190
Ouibrahim L, Mazier M, Estevan J, Pagny G, Decroocq V, Desbiez C, Caranta C (2014) Cloning of the Arabidopsis rwm1 gene for resistance to Watermelon mosaic virus points to a new function for natural virus resistance genes. Plant J 79:705–716
Pack IS, Kim YJ, Youk ES, Lee WK, Yoon WK, Park KW, Kim CG, Harn CH, Kim HM, Jeong SC (2012) A molecular framework for risk assessment of a virus-tolerant transgenic pepper line. J Crop Sci Biotechnol 15:107–115
Park SM, Lee JS, Jegal S, Jeon BY, Jung M, Park YS, Han SL, Shin YS, Her NH, Lee JH, Lee MY (2005) Transgenic watermelon rootstock resistant to CGMMV (cucumber green mottle mosaic virus) infection. Plant Cell Rep 24:350–356
Park HM, Choi MS, Kwak DY, Lee BC, Lee JH, Kim MK, Kim YG, Shin DB, Park SK, Kim YH (2012) Suppression of NS3 and MP is important for the stable inheritance of RNAimediated Rice stripe virus (RSV) resistance obtained by targeting the fully complementary RSV-CP gene. Mol Cells 33:43–51
Panopoulos NJ, Hatziloukas E, Afendra AS (1996) Transgenic crop resistance to bacteria. Field Crops Res 45:85–97
Pang SZ, Jan FJ, Tricoli DM, Russell PF, Carney KJ, Hu JS, Fuchs M, Quemada HD, Gonsalves D (2000) Resistance to squash mosaic comovirus in transgenic squash plants expressing its coat protein genes. Mol Breed 6:87–93
Pavan S, Jacobsen E, Visser RG, Bai Y (2010) Loss of susceptibility as a novel breeding strategy for durable and broad-spectrum resistance. Mol Breed 25:1
Peng HC, Mantelin S, Hicks GR, Takken FL, Kaloshian I (2016) The conformation of a plasma membrane-localized somatic embryogenesis receptor kinase complex is altered by a potato aphid-derived effector. Plant Physiol 171:2211–2222
Peng A, Chen S, Lei T, Xu L, He Y, Wu L, Zou X (2017) Engineering canker-resistant plants through CRISPR/Cas9 targeted editing of the susceptibility gene Cs LOB 1 promoter in citrus. Plant Biotechnol J 15:1509–1519
Prasad K, Bhatnagar-Mathur P, Waliyar F, Sharma KK (2013) Overexpression of a chitinase gene in transgenic peanut confers enhanced resistance to major soil borne and foliar fungal pathogens. J Plant Biochem Biotechnol 22:222–233
Pratap D, Kumar S, Raj SK, Sharma AK (2011) Agrobacterium-mediated transformation of eggplant (Solanum melongena L.) using cotyledon explants and coat protein gene of cucumber mosaic virus. Indian J Biotechnol 10:19–24
Pratap D, Raj SK, Kumar S, Snehi SK, Gautam KK, Sharma AK (2012) Coat protein-mediated transgenic resistance in tomato against a IB subgroup cucumber mosaic virus strain. Phytoparasitica 40:375–382
Praveen S, Kushwaha CM, Mishra AK, Singh V, Jain RK, Varma A (2005) Engineering tomato for resistance to tomato leaf curl disease using viral rep gene sequences. Plant Cell Tissue Org Cult 83:311–318
Prins M, Laimer M, Noris E, Schubert J, Wassenegger M, Tepfer M (2008) Strategies for antiviral resistance in transgenic plants. Mol Plant Pathol 9:73–83
Prins M, Resende RD, Anker C, van Schepen A, de Haan P, Goldbach R (1996) Engineered RNA-mediated resistance to tomato spotted wilt virus is sequence specific. Mol Plant Microbe Interact 9:416–418
Prins M, Kikkert M, Ismayadi C, De Graauw W, De Haan P, Goldbach R (1997) Characterization of RNA-mediated resistance to tomato spotted wilt virus in transgenic tobacco plants expressing NSM gene sequences. Plant Mol Biol 33:235–243
Pumplin N, Voinnet O (2013) RNA silencing suppression by plant pathogens: defence, counter-defence and counter-counter-defence. Nat Rev Microbiol 11:745–760
Pyott DE, Sheehan E, Molnar A (2016) Engineering of CRISPR/Cas9-mediated potyvirus resistance in transgene-free Arabidopsis plants. Mol Plant Pathol 17:1276–1288
Rachman DS, McGeachy K, Reavy B, Štrukelj B, Žel J, Barker H (2001) Strong resistance to potato tuber necrotic ringspot disease in potato induced by transformation with the coat protein gene sequences from an NTN isolate of potato virus Y. Ann Appl Biol 139:269–275
Ram MN, Mohandas S (2003) Transformation of African violet (Saintpaulia ionantha) with glucanase-chitinase genes using Agrobacterium tumefaciens. Acta Hort 624:471–478
Raj SK, Singh R, Pandey SK, Singh B (2005) Agrobacterium-mediated tomato transformation and regeneration of transgenic lines expressing tomato leaf curl virus coat protein gene for resistance against TLCV infection. Curr Sci 88:1674–1679
Reyes CA, De Francesco A, Peña EJ, Costa N, Plata MI, Sendin L, Castagnaro AP, García ML (2011) Resistance to citrus psorosis virus in transgenic sweet orange plants is triggered by coat protein–RNA silencing. J Biotechnol 151:151–158
Ren W, Wang K, Liao P, Yang G, Zhao Y, Zhou Y (2016) The regulation of innate immunity by nutritional factors. Biomed Res Int 2016:5138706
Reyes CA, Peña EJ, Zanek MC, Sanchez DV, Grau O, García ML (2009) Differential resistance to Citrus psorosis virus in transgenic Nicotiana benthamiana plants expressing hairpin RNA derived from the coat protein and 54K protein genes. Plant Cell Rep 28:1817–1825
Rodriguez-Hernabdez AM, Gosalvez B, Sempere RN, Burgos L, Aranda MA, Truniger V (2012) Melon RNA interference (RNAi) lines silenced for cm-eIF4E show broad virus resistance. Mol Plant Pathol 13:755–763
Rodríguez-Leal D, Lemmon ZH, Man J, Bartlett M, Lippman ZB (2017) Engineering quantitative trait variation for crop improvement by genome editing. Cell 171:470–480
Rojas CM, Senthil-Kumar M, Wang K, Ryu CM, Kaundal A, Mysore KS (2012) Glycolate oxidase modulates reactive oxygen species–mediated signal transduction during nonhost resistance in Nicotiana benthamiana and Arabidopsis. Plant Cell 24:336–352
Rubio T, Borja M, Scholthof HB, Feldstein PA, Morris TJ, Jackson AO (1999) Broad-spectrum protection against tombusviruses elicited by defective interfering RNAs in transgenic plants. J Virol 73:5070–5078
Rubio J, Montes C, Castro Á, Alvarez C, Olmedo B, Munoz M, Miccono M (2015) Genetically engineered Thompson Seedless grapevine plants designed for fungal tolerance: selection and characterization of the best performing individuals in a field trial. Transgenic Res 24:43–60
Rudolph C, Schreier PH, Uhrig JF (2003) Peptide-mediated broad-spectrum plant resistance to tospoviruses. Proc Natl Acad Sci 100:4429–4434
Rushton PJ, Reinstädler A, Lipka V, Lippok B, Somssich IE (2002) Synthetic plant promoters containing defined regulatory elements provide novel insights into pathogen-and wound-induced signaling. Plant Cell 14:749–762
Ryabov EV, Keane G, Naish N, Evered C, Winstanley D (2009) Densovirus induces winged morphs in asexual clones of the rosy apple aphid, Dysaphis plantaginea. Proc Natl Acad Sci U S A 106:8465–8470
Sano T, Nagayama A, Ogawa T, Ishida I, Okada Y (1997) Transgenic potato expressing a double-stranded RNA-specific ribonuclease is resistant to potato spindle tuber viroid. Nat Biotechnol 15:1290–1294
Savenkov EI, Valkonen JP (2002) Silencing of a viral RNA silencing suppressor in transgenic plants. J Gen Virol 83:2325–2335
Sarowar S, Kim YJ, Kim KD, Hwang BK, Ok SH, Shin JS (2009) Overexpression of lipid transfer protein (LTP) genes enhances resistance to plant pathogens and LTP functions in longdistance systemic signaling in tobacco. Plant Cell Rep 28:419–427
Saur IM, Kadota Y, Sklenar J, Holton NJ, Smakowska E, Belkhadir Y, Zipfel C, Rathjen JP (2016) NbCSPR underlies age-dependent immune responses to bacterial cold shock protein in Nicotiana benthamiana. Proc Natl Acad Sci 113:3389–3394
Saharan V, Jain D, Pareek S, Pal A, Kumaraswamy RV, Jakhar SK, Singh M (2016) Viral, fungal and bacterial disease resistance in transgenic plants. In: Advances in plant breeding strategies: agronomic, abiotic and biotic stress traits. Springer, Cham, pp 627–656
Sanford JC, Johnston SA (1985) The concept of parasite-derived resistance—deriving resistance genes from the parasite’s own genome. J Theor Biol 113:395–405
Schouten HJ, Krens FA, Jacobsen E (2006) Cisgenic plants are similar to traditionally bred plants: international regulations for genetically modified organisms should be altered to exempt cisgenesis. EMBO Rep 7:750–753
Schwind N, Zwiebel M, Itaya A, Ding B, Wang MB, Krczal G, Wassenegger M (2009) RNAi-mediated resistance to potato spindle tuber viroid in transgenic tomato expressing a viroid hairpin RNA construct. Mol Plant Pathol 10:459–469
Schoonbeek HJ, Wang HH, Stefanato FL, Craze M, Bowden S, Wallington E, Ridout CJ (2015) Arabidopsis EF-Tu receptor enhances bacterial disease resistance in transgenic wheat. New Phytol 206:606–613
Schüttelkopf AW, Gros L, Blair DE, Frearson JA, van Aalten DM, Gilbert IH (2010) Acetazolamide-based fungal chitinase inhibitors. Bioorgan Med Chem 18:8334–8340
Scorza R, Callahan AM, Damsteegt V, Levy L, Ravelonandro M (1997) Transferring potyvirus coat protein genes through hybridization of transgenic plants to produce plum pox virus resistant plums (Prunus domestica L.). In: In XVII international symposium virus and virus-like diseases of temperate fruit crops, vol 472, pp 421–428
Senthil-Kumar M, Mysore KS (2013) Nonhost resistance against bacterial pathogens: retrospectives and prospects. Annu Rev Phytopathol 51:407–427
Senthilkumar R, Cheng CP, Yeh KW (2010) Genetically pyramiding protease-inhibitor genes for dual broad-spectrum resistance against insect and phytopathogens in transgenic tobacco. Plant Biotechnol J 8:65–75
Sengoda VG, Tsai WS, De La Peña RC, Green SK, Kenyon L, Hughes J (2012) Expression of the full-length coat protein gene of tomato leaf curl Taiwan virus is not necessary for recovery phenotype in transgenic tomato. J Phytopathol 160:213–219
Seybold H, Trempel F, Ranf S, Scheel D, Romeis T, Lee J (2014) Ca2+ signalling in plant immune response: from pattern recognition receptors to Ca2+ decoding mechanisms. New Phytol 204:782–790
Seo GJ, Kincaid RP, Phanaksri T, Burke JM, Pare JM, Cox JE, Hsiang TY, Krug RM, Sullivan CS (2013) Reciprocal inhibition between intracellular antiviral signaling and the RNAi machinery in mammalian cells. Cell Host Microbe 14:435–445
Shepherd DN, Mangwende T, Martin DP, Bezuidenhout M, Kloppers FJ, Carolissen CH, Monjane AL, Rybicki EP, Thomson JA (2007) Maize streak virus-resistant transgenic maize: a first for Africa. Plant Biotechnol J 5:759–767
Shekhawat UK, Ganapathi TR, Hadapad AB (2012) Transgenic banana plants expressing small interfering RNAs targeted against viral replication initiation gene display high-level resistance to banana bunchy top virus infection. J Gen Virol 93:1804–1813
Shin R, Han JH, Lee GJ, Peak KH (2002) The potential use of a viral coat protein gene as a transgene screening marker and multiple virus resistance in pepper plants coexpressing coat proteins of cucumber mosaic virus and tomato mosaic virus. Transgenic Res 11:215–219
Shin S, Mackintosh CA, Lewis J, Heinen SJ, Radmer L, Dill-Macky R, Baldridge GD, Zeyen RJ, Muehlbauer GJ (2008) Transgenic wheat expressing a barley class II chitinase gene has enhanced resistance against Fusarium graminearum. J Exp Bot 59:2371–2378
Shimizu T, Nakazono-Nagaoka E, Uehara-Ichiki T, Sasaya T, Omura T (2011) Targeting specific genes for RNA interference is crucial to the development of strong resistance to Rice stripe virus. Plant Biotechnol J 9:503–512
Shimizu T, Nagaoka EK, Akita F, Wei T, Sasaya T, Omura T, Ichiki TU (2012) Haripin RNA derived from the gene for Pns9, a viroplasm matrix protein of Rice gall dwarf virus, confers strong resistance to virus infection in transgenic rice plants. J of Biotech 157:421–427
Shimizu T, Yoshii M, Wei T, Hirochika H, Omura T (2009) Silencing by RNAi of the gene for Pns12, a viroplasm matrix protein of Rice dwarf virus, results in strong resistance of transgenic rice plants to the virus. Plant Biotechnol J 7:24–32
Shimizu T, Ogamino T, Hiraguri A, Nakazono-Nagaoka E, Uehara-Ichiki T, Nakajima M, Akutsu K, Omura T, Sasaya T (2013) Strong resistance against Rice grassy stunt virus is induced in transgenic rice plants expressing double-stranded RNA of the viral genes for nucleocapsid or movement proteins as targets for RNA interference. Phytopathology 103:513–519
Sijen T, Wellink J, Hendriks J, Verver J, van Kammen A (1995) Replication of cowpea mosaic virus RNA1 and RNA2 is specifically blocked in transgenic Nicotiana benthamiana plants expressing the full-length replicase or movement protein genes. Mol Plant-Microbe Interact 8:340–347
Silva Y, Portieles R, Pujol M, Terauchi R, Matsumura H, Serrano M, Borrás-Hidalgo O (2013) Expression of a microbial serine proteinase inhibitor gene enhances the tobacco defense against oomycete pathogens. Physiol Mol Plant Pathol 84:99–106
Silva MS, Arraes FBM, de Araújo CM, Grossi-de-Sa M, Fernandez D, de Souza CE, Grossi-de-Sa MF (2018) Potential biotechnological assets related to plant immunity modulation applicable in engineering disease-resistant crops. Plant Sci 270:72–84
Simon AE, Roossinck MJ, Havelda Z (2004) Plant virus satellite and defective interfering RNAs: new paradigms for a new century. Annu Rev Phytopathol 42:415–437
Simón-Mateo C, García JA (2011) Antiviral strategies in plants based on RNA silencing. BBA-Gene Regul Mech 1809:722–731
Singh A, Chaudhary S, Shankar A, Prasad V (2019) Enhanced resistance to fungal pathogens through selective utilization of useful microbial genes. In: New and future developments in microbial biotechnology and bioengineering. Elsevier, Amsterdam, pp 87–93
Singh VB, Haq QMR, Malathi VG (2013) Antisense RNA approach targeting Rep gene of Mungbean yellow mosaic India virus to develop resistance in soybean. Arch Phytopathol Plant Protect 46:2191–2207
Sivamani E, Brey CW, Talbert LE, Young MA, Dyer WE, Kaniewski WK, Qu R (2002) Resistance to wheat streak mosaic virus in transgenic wheat engineered with the viral coat protein gene. Transgenic Res 11:31–41
Smith NA, Singh SP, Wang MB, Stoutjesdijk PA, Green AG, Waterhouse PM (2000) Gene expression: total silencing by intron-spliced hairpin RNAs. Nature 407:319
Stafford CA, Walker GP, Ullman DE (2011) Infection with a plant virus modifies vector feeding behavior. Proc Natl Acad Sci 108:9350–9355
Stanley J, Frischmuth T, Ellwood S (1990) Defective viral DNA ameliorates symptoms of geminivirus infection in transgenic plants. Proc Natl Acad Sci 87:6291–6295
Stenger DC (1994) Strain-specific mobilization and amplification of a transgenic defective-interfering DNA of the geminivirus beet curly top virus. Virology 203:397–402
Stam M, Mol JN, Kooter JM (1997) The silence of genes in transgenic plants. Ann Bot 79:3–12
Sridevi G, Parameswari C, Sabapathi N, Raghupathy V, Veluthambi K (2008) Combined expression of chitinase and β-1, 3-glucanase genes in indica rice (Oryza sativa L.) enhances resistance against Rhizoctonia solani. Plant Sci 175:283–290
Soler N, Plomer M, Fagoaga C, Moreno P, Navarro L, Flores R, Peña L (2012) Transformation of Mexican lime with an intron-hairpin construct expressing untranslatable versions of the genes coding for the three silencing suppressors of Citrus tristeza virus confers complete resistance to the virus. Plant Biotechnol J 10:597–608
Souza JMT, Nickel O, Gonsalves D (2005) Development of virus resistant transgenic papayas expressing the coat protein gene from a Brazilian isolate of papaya ringspot virus. Fitopatol Bras 30:357–365
Swaney S, Powers H, Goodwin J, Rosales LS, Dougherty WG (1995) RNA-mediated resistance with nonstructural genes from the tobacco etch virus genome. Mol Plant Microbe Interact 8:1004–1011
Tavert-Roudet G, Ravelonandro M, Bachelier JC, Dunez J (1998) Transgenic Nicotiana benthamiana plants containing the P1 gene of plum pox virus are resistant to virus challenge. Eur J Plant Pathol 104:103–107
Tavladoraki P, Benvenuto E, Trinca S, De Martinis D, Cattaneo A, Galeffi P (1993) Transgenic plants expressing a functional single-chain Fv antibody are specifically protected from virus attack. Nature 366:469–472
Tacke E, Salamini F, Rohde W (1996) Genetic engineering of potato for broad-spectrum protection against virus infection. Nat Biotechnol 14:1597
Tai TH, Dahlbeck D, Clark ET, Gajiwala P, Pasion R, Whalen M, Staskawicz BJ (1999) Expression of the Bs2 pepper gene confers resistance to bacterial spot disease in tomato. Proc Natl Acad Sci 96:14153–14158
Tang X, Xie M, Kim YJ, Zhou J, Klessig DF, Martin GB (1999) Overexpression of Pto activates defense responses and confers broad resistance. Plant Cell 11:15–29
Tamarzizt HB, Chouchane SG, Lengliz R, Maxwell DP, Marrakchi M, Fakhfakh H, Gorsane F (2009) Use of tomato leaf curl virus (TYLCV) truncated Rep gene sequence to engineer TYLCV resistance in tomato plants. Acta Virol 53:99–104
Tinoco MLP, Dias BB, Dall’Astta RC, Pamphile J, Aragão FJ (2010) In vivo trans-specific gene silencing in fungal cells by in planta expression of a double-stranded RNA. BMC Biol 8:27
Tran DT, Cho S, Hoang PM, Kim J, Kil EJ, Lee TK, Rhee Y, Lee S (2016) A codon-optimized nucleic acid hydrolyzing single-chain antibody confers resistance to Chrysanthemums against Chrysanthemum stunt viroid infection. Plant Mol Biol Report 34:221–232
Trifonova EA, Sapotsky MV, Komarova ML, Scherban AB, Shumny VK, Polyakova AM, Lapshina LA, Kochetov AV, Malinovsky VI (2007) Protection of transgenic tobacco plants expressing bovine pancreatic ribonuclease against tobacco mosaic virus. Plant Cell Rep 26:1121–1126
Trifonova EA, Romanova AV, Sangaev SS, Sapotsky MV, Malinovsky VI, Kochetov AV (2012) Inducible expression of the gene of Zinnia elegans coding for extracellular ribonuclease in Nicotiana tabacum plants. Biol Plant 56:571–574
Tougou M, Furutani N, Yamagishi N, Shizukawa Y, Takahata Y, Hidaka S (2006) Development of resistant transgenic soybeans with inverted repeat-coat protein genes of soybean dwarf virus. Plant Cell Rep 25:1213–1218
Tripathi L, Mwaka H, Tripathi JN, Tushemereirwe WK (2010) Expression of sweet pepper Hrap gene in banana enhances resistance to Xanthomonas campestris pv. musacearum. Mol Plant Pathol 11:721–731
Tripathi L, Tripathi JN, Shah T, Muiruri KS, Katari M (2019) Molecular basis of disease resistance in banana progenitor Musa balbisiana against Xanthomonas campestris pv. musacearum. Sci Rep 9:1–17
Tripathi L, Tripathi JN, Kiggundu A, Korie S, Shotkoski F, Tushemereirwe WK (2014) Field trial of Xanthomonas wilt disease-resistant bananas in East Africa. Nat Biotechnol 32:868–870
Tougou M, Yamagishi N, Furutani N, Shizukawa Y, Takahata Y, Hidaka S (2007) Soybean dwarf virus-resistant transgenic soybeans with the sense coat protein gene. Plant Cell Rep 26:1967–1975
Tundo S, Kalunke R, Janni M, Volpi C, Lionetti V, Bellincampi D, D’Ovidio R (2016) Pyramiding PvPGIP2 and TAXI-III but not PvPGIP2 and PMEI enhances resistance against Fusarium graminearum. Mol Plant-Microbe Interact 29:629–639
Uma B, Rani TS, Podile AR (2011) Warriors at the gate that never sleep: non-host resistance in plants. J Plant Physiol 168:2141–2152
van der Vossen EA, Gros J, Sikkema A, Muskens M, Wouters D, Wolters P, Pereira A, Allefs S (2005) The Rpi-blb2 gene from Solanum bulbocastanum is an Mi-1 gene homolog conferring broad-spectrum late blight resistance in potato. Plant J 44:208–222
Vanderschuren H, Moreno I, Anjanappa RB, Zainuddin IM, Gruissem W (2012) Exploiting the combination of natural and genetically engineered resistance to cassava mosaic and cassava brown streak viruses impacting cassava production in Africa. PLoS One 7:e45277
van Vu T, Choudhury NR, Mukherjee SK (2013) Transgenic tomato plants expressing artificial microRNAs for silencing the pre-coat and coat proteins of a begomovirus, tomato leaf curl New Delhi virus, show tolerance to virus infection. Virus Res 172:35–45
van Schie C, Takken FL (2014) Susceptibility genes 101: how to be a good host. Annu Rev Phytopathol 52:551–581
Vaskin MFS, Vidal MS, Alves ED, Farinelli L, de Oliveira DE (2001) Co-suppression mediated virus resistance in transgenic tobacco plants harboring the 3-untranslated region of Andean potato mottle virus. Transgenic Res 10:489–499
Verma V, Sharma S, Devi SV, Rajasubramaniam S, Dasgupta I (2012) Delay in virus accumulation and low virus transmission from transgenic rice plants expressing Rice tungro spherical virus RNA. Virus Genes 45:350–359
Vidya CS, Manoharan M, Kumar CR, Savtthri HS, Sita GL (2000) Agrobacterium-mediated transformation of tomato (Lycopersicon esculentum var. Pusa Ruby) with coat-protein gene of Physalis mottle tymovirus. J Plant Physiol 156:106–110
Vincelli P (2016) Genetic engineering and sustainable crop disease management: opportunities for case-by-case decision-making. Sustainability 8:495
Voinnet O (2008) Use, tolerance and avoidance of amplified RNA silencing by plants. Trends Plant Sci 13:317–328
Wally O, Jayaraj J, Punja Z (2009) Comparative resistance to foliar fungal pathogens in transgenic carrot plants expressing genes encoding for chitinase, β-1, 3-glucanase and peroxidise. Eur J Plant Pathol 123:331–342
Wang M-B, Abbott DC, Waterhouse PM (2000) A single copy of a virus-derived transgene encoding hairpin RNA gives immunity to barley yellow dwarf virus. Mol Plant Pathol 1(6):347–356
Wang X, Eggenberger AL, Nutter FW Jr, Hill JH (2001) Pathogen-derived transgenic resistance to soybean mosaic virus in soybean. Mol Breed 8:119–127
Wang HZ, Zhao PJ, Xu JC, Zhao H, Zhang HS (2003) Virus resistance in transgenic watermelon plants containing a WMV-2 coat protein gene. Acta Genet Sin 30:70–75
Wang X, Zhu X, Tooley P, Zhang X (2013) Cloning and functional analysis of three genes encoding polygalacturonase-inhibiting proteins from Capsicum annuum and transgenic CaPGIP1 in tobacco in relation to increased resistance to two fungal pathogens. Plant Mol Biol 81:379–400
Wang Y, Cheng X, Shan Q, Zhang Y, Liu J, Gao C, Qiu JL (2014) Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat Biotechnol 32:947
Wang F, Wang C, Liu P, Lei C, Hao W, Gao Y, Zhao K (2016) Enhanced rice blast resistance by CRISPR/Cas9-targeted mutagenesis of the ERF transcription factor gene OsERF922. PLoS One 11:e0154027
Wagaba H, Beyene G, Aleu J, Odipio J, Okao-Okuja G, Chauhan RD, Munga T, Obiero H, Halsey ME, Ilyas M, Raymond P, Bua A, Taylor NJ, Miano D Alicai T (2017) Field level RNAimediated resistance to cassava brown streak disease across multiple cropping cycles and diverse east African agro-ecological locations. Front Plant Sci 7:2060
Wang B, Sumit R, Sahu BB, Ngaki MN, Srivastava SK, Yang Y, Bhattacharyya MK (2018) Arabidopsis novel glycine-rich plasma membrane PSS1 protein enhances disease resistance in transgenic soybean plants. Plant Physiol 176:865–878
Watanabe Y, Ogawa T, Takahashi H, Ishida I, Takeuchi Y, Yamamoto M, Okada Y (1995) Resistance against multiple plant viruses in plants mediated by a double stranded-RNA specific ribonuclease. FEBS Lett 372:165–168
Waterhouse PM, Graham MW, Wang MB (1998) Virus resistance and gene silencing in plants can be induced by simultaneous expression of sense and antisense RNA. Proc Natl Acad Sci 95:13959–13964
Waterhouse PM, Helliwell CA (2003) Exploring plant genomes by RNA-induced gene silencing. Nat Rev Genet 4:29
Waterhouse PM, Wang MB, Lough T (2001) Gene silencing as an adaptive defence against viruses. Nature 411:834–842
Wako T, Terami F, Hanada K, Tabei Y (2001) Resistance to Zucchini yellow mosaic virus (ZYMV) in transgenic cucumber plants (Cucumis sativus L.) harboring the coat protein gene of ZYMV. Bull Natl Res Inst Veg Ornament Plant Tea (Japan) 16:175–186
Wegener CB (2002) Induction of defence responses against Erwinia soft rot by an endogenous pectate lyase in potatoes. Physiol Mol Plant Pathol 60(2):91–100
Westwood JH, Groen SC, Du Z, Murphy AM, Anggoro DT, Tungadi T, Smith AG (2013) A trio of viral proteins tunes aphid-plant interactions in Arabidopsis thaliana. PLoS One 8:e83066
Weiberg A, Wang M, Bellinger M, Jin H (2014) Small RNAs: a new paradigm in plant-microbe interactions. Annu Rev Phytopathol 52:495–516
Wesley SV, Helliwell CA, Smith NA, Wang M, Rouse DT, Liu Q, Gooding PS, Singh SP, Abbott D, Stoutjesdijk PA, Robinson SP (2001) Construct design for efficient, effective and high throughput gene silencing in plants. Plant J 27(6):581–590
Whitfield AE, Rotenberg D (2015) Disruption of insect transmission of plant viruses. Curr Opin Insect Sci 8:79–87
Wu HW, Yu TA, Raja JA, Wang HC, Yeh SD (2009) Generation of transgenic oriental melon resistant to Zucchini yellow mosaic virus by an improved cotyledon-cutting method. Plant Cell Rep 28(7):1053–1064
Wu HW, Yu TA, Raja J, Christopher SJ, Wang SL, Yeh SD (2010) Double-virus resistance of transgenic oriental melon conferred by untranslatable chimeric construct carrying partial coat protein genes of two viruses. Plant Dis 94(11):1341–1347
Wu Y, Zhou JM (2013) Receptor-Like kinases in plant innate immunity. J Integr Plant Biol 55(12):1271–1286
Xu BL, Shi GY, Xue YY (2005) Plant regeneration of transgenic Huanghemi (Cucumis melo) and identification of its resistance to virus disease. J Fruit Sci 22:734–736
Xu W, Yang S, Bhadury P, He J, He M, Gao L, Song B (2011) Synthesis and bioactivity of novel sulfone derivatives containing 2, 4-dichlorophenyl substituted 1, 3, 4-oxadiazole/thiadiazole moiety as chitinase inhibitors. Pestic Biochem Physiol 101:6–15
Xu G, Yuan M, Ai C, Liu L, Zhuang E, Karapetyan S, Dong X (2017) uORF-mediated translation allows engineered plant disease resistance without fitness costs. Nature 545:491
Yu TA, Chiang CH, Wu HW, Li CM, Yang CF, Chen JH, Chen YW, Yeh SD (2011) Generation of transgenic watermelon resistant to Zucchini yellow mosaic virus and papaya ringspot virus type W. Plant Cell Rep 30(3):359–371
Yan F, Kuang Y, Ren B, Wang J, Zhang D, Lin H, Zhou H (2018) Highly efficient AT to GC base editing by Cas9n-guided tRNA adenosine deaminase in rice. Mol Plant 11:631–634
Yadav JS, Ogwok E, Wagaba H, Patil BL, Bagewadi B, Alicai T, Gaitan-Solis E, Taylor NJ, Fauquet CM (2011) RNAi-mediated resistance to cassava brown streak Uganda virus in transgenic cassava. Mol Plant Pathol 12(7):677–687
Yang Y, Sherwood TA, Patte CP, Hiebert E, Polston JE (2004) Use of tomato yellow leaf curl virus (TYLCV) Rep gene sequences to engineer TYLCV resistance in tomato. Phytopathology 94(5):490–496
Yang X, Wang X, Li X, Zhang B, Xiao Y, Li D, Xie C, Pei Y (2008) Characterization and expression of an nsLTPs-like antimicrobial protein gene from motherwort (Leonurus japonicus). Plant Cell Rep 27(4):759–766
Yang X, Yie Y, Zhu F, Liu Y, Kang L, Wang X, Tien P (1997) Ribozyme-mediated high resistance against potato spindle tuber viroid in transgenic potatoes. Proc Natl Acad Sci 94(10):4861–4865
Yang X, Niu L, Zhang W, Yang J, Xing G, He H (2018) RNAi-mediated SMV P3 cistron silencing confers significantly enhanced resistance to multiple Potyvirus strains and isolates in transgenic soybean. Plant Cell Rep 37:103–114
Yassaie M, Afsharifar AR, Niazi A, Salehzadeh S, Izadpanah K (2011) Induction of resistance to barley yellow dwarf virus (PAV) in bread wheat using post-transcriptional gene silencing (PTGS). Iran J Plant Pathol 47:67–82
Yuan Y, Zhong S, Li Q, Zhu Z, Lou Y, Wang L, Wang J, Wang M, Li Q, Yang D, He Z (2007) Functional analysis of rice NPR1-like genes reveals that OsNPR1/NH1 is the rice orthologue conferring disease resistance with enhanced herbivore susceptibility. Plant Biotechnol J 5(2):313–324
Yuan M, Wang S (2013) Rice MtN3/saliva/SWEET family genes and their homologs in cellular organisms. Mol Plant 6:665–674
Yuan M, Chu Z, Li X, Xu C, Wang S (2010) The bacterial pathogen Xanthomonas oryzae overcomes rice defenses by regulating host copper redistribution. Plant Cell 22:3164–3176
Zaccomer B, Cellier F, Boyer JC, Haenni AL, Tepfer M (1993) Transgenic plants that express genes including the 3′ untranslated region of the turnip yellow mosaic virus (TYMV) genome are partially protected against TYMV infection. Gene 136(1-2): 87–94
Zaidi SSEA, Mukhtar MS, Mansoor S (2018) Genome editing: targeting susceptibility genes for plant disease resistance. Trends Biotechnol 36:898–906
Zetsche B, Gootenberg JS, Abudayyeh OO, Slaymaker IM, Makarova KS, Essletzbichler P, Koonin EV (2015) Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163:759–771
Zhang H, Tian Y, Zhou Y, Dang B, Lan H, Song G (1999) Introduction of pokeweed antiviral protein cDNA into Brassica napus and acquisition of transgenic plants resistant to viruses. Chin Sci Bull 4:701–704
Zhang P, Vanderschuren H, Fütterer J, Gruissem W (2005) Resistance to cassava mosaic disease in transgenic cassava expressing antisense RNAs targeting virus replication genes. Plant Biotechnol J 3(4):385–397
Zhang YY, Li HX, Ouyang B, Ye ZB (2006) Regulation of eukaryotic initiation factor 4E and its isoform: implications for antiviral strategy in plants. J Integr Plant Biol 48:1129–1139
Zhang ZY, Fu FL, Gou L, Wang HG, Li WC (2010) RNA interference-based transgenic maize resistant to maize dwarf mosaic virus. J Plant Biol 53:297–305
Zhang ZY, Yang L, Zhou SF, Wang HG, Li WC, Fu FL (2011) Improvement of resistance to maize dwarf mosaic virus mediated by transgenic RNA interference. J Biotechnol 153:181–187
Zhang B, Oakes AD, Newhouse AE, Baier KM, Maynard CA, Powell WA (2013) A threshold level of oxalate oxidase transgene expression reduces Cryphonectria parasitica-induced necrosis in a transgenic American chestnut (Castanea dentata) leaf bioassay. Transgenic Res 22(5):973–982
Zhang M, Li Q, Liu T, Liu L, Shen D, Zhu Y (2015) Two cytoplasmic effectors of Phytophthora sojae regulate plant cell death via interactions with plant catalases. Plant Physiol 167(1):164–175
Zhang T, Zheng Q, Yi X, An H, Zhao Y, Ma S, Zhou G (2018) Establishing RNA virus resistance in plants by harnessing CRISPR immune system. Plant Biotechnol J 16:1415–1423
Zhao B, Ardales EY, Raymundo A, Bai J, Trick HN, Leach JE, Hulbert SH (2004) The avrRxo1 gene from the rice pathogen Xanthomonas oryzae pv. oryzicola confers a nonhost defense reaction on maize with resistance gene Rxo1. Mol Plant-Microbe Interact 17:771–779
Zhao B, Lin X, Poland J, Trick H, Leach J, Hulbert S (2005) A maize resistance gene functions against bacterial streak disease in rice. Proc Natl Acad Sci 102:15383–15388
Zhao Y, Yang X, Zhou G, Zhang T (2019) Engineering plants virus resistance: from RNA silencing to genome editing strategies. Plant Biotechnol J 18(2):328–336
Zhou Y, Yuan Y, Yuan F, Wang M, Zhong H, Gu M, Liang G (2012) RNAi-directed down-regulation of RSV results in increased resistance in rice (Oryza sativa L.). Biotechnol Lett 34:965–972
Zhou J, Peng Z, Long J, Sosso D, Liu B, Eom JS, Huang S, Liu S, Vera Cruz C, Frommer WB, White FF (2015) Gene targeting by the TAL effector PthXo2 reveals cryptic resistance gene for bacterial blight of rice. Plant J 82:632–643
Zhu YX, Ou-Yang WJ, Zhang YF, Chen ZL (1996) Transgenic sweet pepper plants from agrobacterium mediated transformation. Plant Cell Rep 16:71–75
Zhu H, Xu X, Xiao G, Yuan L, Li B (2007) Enhancing disease resistances of super hybrid rice with four antifungal genes. Sci China Life Sci 50:31–39
Zhu YJ, McCafferty H, Osterman G, Lim S, Agbayani R, Lehrer A, Schenck S, Komor E (2011) Genetic transformation with untranslatable coat protein gene of sugarcane yellow leaf virus reduces virus titers in sugarcane. Transgenic Res 20(3):503–512
Zipfel C, Felix G (2005) Plants and animals: a different taste for microbes. Curr Opin Plant Biol 8:353–360
Zrachya A, Kumar PP, Ramakrishnan U, Levy Y, Loyter A, Arazi T, Lapidot M, Gafni Y (2007) Production of siRNA targeted against TYLCV coat protein transcripts leads to silencing of its expression and resistance to the virus. Transgenic Res 16:385–398
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Makeshkumar, T., Divya, K., Asha, S. (2021). Transgenic Technology for Disease Resistance in Crop Plants. In: Singh, K.P., Jahagirdar, S., Sarma, B.K. (eds) Emerging Trends in Plant Pathology . Springer, Singapore. https://doi.org/10.1007/978-981-15-6275-4_23
Download citation
DOI: https://doi.org/10.1007/978-981-15-6275-4_23
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-6274-7
Online ISBN: 978-981-15-6275-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)