Abstract
Magnaporthe oryzae and Rhizoctonia solani, are among the most important pathogens of rice, severely limiting its productivity. Dm-AMP1, an antifungal plant defensin from Dahlia merckii, was expressed in rice (Oryza sativa L. sp. indica cv. Pusa basmati 1) using Agrobacterium tumefaciens-mediated transformation. Expression levels of Dm-AMP1 ranged from 0.43% to 0.57% of total soluble protein in transgenic plants. It was observed that constitutive expression of Dm-AMP1 suppresses the growth of M. oryzae and R. solani by 84% and 72%, respectively. Transgenic expression of Dm-AMP1 was not accompanied by an induction of pathogenesis-related (PR) gene expression, indicating that the expression of DmAMP1 directly inhibits the pathogen. The results of in vitro, in planta and microscopic analyses suggest that Dm-AMP1 expression has the potential to provide broad-spectrum disease resistance in rice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice is a host for more than 70 diseases caused by fungi, bacteria and viruses. Rice blast (M. oryzae) and sheath blight (R. solani) are among the most devastating fungal diseases affecting productivity of rice (Ou 1985). M. oryzae, an Ascomycete fungus, produces specialised infection structures known as appressoria that infect aerial tissues and hyphopodia that can infect root tissues (Sesma and Osbourn 2004). M. oryzae is highly variable and hence, durable resistance through breeding has attained limited success. Sheath blight is considered to be an important disease of rice next to the blast disease. It is caused by the fungus R. solani which is a Basidiomycete that occasionally produces sexual spores (basidiospores) but no asexual spores (conidia). In nature, R. solani reproduces asexually and exists primarily as vegetative mycelium and/or sclerotia and has a broad host range (Anderson 1982). Attempts to control R. solani with resistant rice cultivars have not been successful because effective source of resistance is not available. Currently, these diseases are controlled by the application of chemicals, use of resistant cultivars and agronomic management practices. However, plant genetic engineering might be an attractive method for improving resistance to fungal diseases in commercial rice varieties.
Plants possess an impressive arsenal of antimicrobial compounds that are either constitutively arrayed within certain tissues or synthesized in direct response to attack by pathogens (Gu et al. 1992; Terras et al. 1995). Plant defensins are small peptides (45–54 amino acids) with a characteristic three-dimensional folding pattern stabilized by disulfide-linked cysteines. They share common characters with those described in insects and mammals, suggesting that these defense molecules pre-date the evolutionary divergence of plants and animals (Broekaert et al. 1995). So far, over 80 different antimicrobial peptides from 38 plant species have been identified as belonging to a group of small, highly basic, and cysteine-rich peptides, the defensins (Thomma et al. 2002). Molecular mechanisms of plant defensins action are not completely understood. Neurospora crassa hyphae treated with either Rs-AFP2 or Dm-AMP1 defensins showed a rapid potassium (K+) efflux, calcium (Ca2+) uptake and alkalinization of the incubation medium. In addition, membrane potential changes, but not the formation of membrane pores, were observed (Thevissen et al. 1996). Defensins are able to bind to hyphal membranes at specific sites and indirect evidence indicates that this binding is required for antifungal activity (Thevissen et al. 1997; Thevissen et al. 2000). Recently, it was shown that different plant defensins are able to interact with different sphingolipids. Dm-AMP1 interacts with mannosylated sphingolipids occurring in the outer plasma membrane (Thevissen et al. 2000, 2003), and displaying a broad-spectrum in vitro antifungal activity (Osborn et al. 1995; Thomma et al. 2002). We hypothesized that expression of the Dm-AMP1 gene in rice may confer resistance to M. oryzae and R. solani. Thus, we introduced the gene for Dm-AMP1 in Pusa basmati 1 (highly susceptible to both blast and sheath blight), analysed its expression in different plant tissues, and studied the in planta interaction of the antimicrobial peptide with M. oryzae and R. solani.
Materials and methods
Plasmid construction and rice transformation
Dm-AMP1 gene was amplified from plasmid pFAJ3105 (Francois et al. 2002) and cloned at BamHI site of pAHC17 under the maize Ubiquitin (Ub1) promoter (Christensen and Quail 1996) to obtain pUb1-Dm. Plasmid pUb1-Dm was digested and cloned at HindIII site in pCAMBIA-1305.2 (Cambia, Australia) to obtain pUb1-Dm-T. The binary vector (pUb1-Dm-T) was mobilized into Agrobacterium tumefaciens strain LBA4404 harbouring pSB1 (a plasmid that carries virB, virG and virC from pTiBo542; Komari et al. 1996). The scutellum-derived calli of rice (Oryza sativa L. cv Pusa Basmati 1) were transformed using A. tumefaciens strain LBA4404 (pSB1, pUb1-Dm-T) as described earlier (Hiei et al. 1994). Total DNA from various independent transgenic lines was extracted (Dellaporta et al. 1983) and screened by PCR (using the same primers used for Dm-AMP1 isolation). The PCR profile was 94°C, 30 s; 52°C, 30 s; 72°C, 1 min, for 30 cycles in a Minicycler (MJ Research, Waltham, MA, USA).
Southern blot analysis
The genomic DNA (10 μg) from non-transformed and T2 transgenic plants was digested with restriction enzymes KpnI/SacI, electrophoresed on 0.8% agarose gel and blotted onto Hybond-N+ nylon membrane (Amersham, Buckinghamshire, England). DNA was fixed to the membranes using a UV cross-linker, 12 × 104 mJ/cm2, (Spectrolinker, Spectronics Corporation, USA). Hybridization probe (280 bp Dm-AMP1 gene) was labeled non-radioactively using Gene Images AlkPhos Direct Labelling and Detection System (Amersham Biosciences, Hong Kong). Hybridization was performed as recommended by supplier (Amersham Biosciences, Hong Kong).
Northern blot analysis
Total RNA was isolated from the leaves of transgenic and control plants as described by Logeman et al. (1987). About 20 μg/sample of total RNA were subjected to formaldehyde-containing agarose gel electrophoresis, and transferred onto nylon membrane (Hybond-N+, Amersham, Buckinghamshire, England). Hybridization probe was labeled non-radioactively using Gene Images AlkPhos Direct Labelling and Detection System (Amersham Biosciences, Hong Kong). Hybridization was performed as recommended by supplier (Amersham Biosciences, Hong Kong).
Raising antibodies against Dm-AMP1
Dm-AMP1 coding sequence was translationally fused to Glutathione-S-Transferase (GST) gene in pEG (KT) vector (Mitchell et al. 1993) and expressed in Saccharomyces cerevisiae strain, s288C by ‘one-step transformation’ method (Chen et al. 1992). Four percent galactose was used to induce expression of GST-Dm-AMP1 fusion protein in the yeast transformant. Purification of the fusion protein was carried out by affinity chromatography using GST purification module as per manufacturer’s instructions (Amersham Biosciences, Hong Kong). The purified fraction was treated with factor Xa to get purified Dm-AMP1. The purified fraction was used to raise polyclonal antibodies in rabbit as follows: 5 μg of purified DmAMP1 in 0.5 ml of 0.1 M phosphate buffered saline (PBS) was emulsified with an equal volume of Freund’s complete adjuvant and was used immediately for subcutaneous injection in rabbit. Two booster doses of antigen- adjuvant mixture were given after every 4 weeks. Fifteen millilitre of blood was drawn seven days after the second booster dose. The antibody (unfractionated serum) titer was estimated by indirect enzyme-linked immunosorbent assay (ELISA).
Western blot analysis
Rice leaf tissue was crushed into fine powder in liquid nitrogen and resuspended in 1× PBS containing 2 mM EDTA and 1 mM PMSF. Protein concentration was estimated by Bradford method (Bradford 1976). Total soluble proteins from four pUb1-Dm-T transgenic lines (Dm-T-14, Dm-T-16, Dm-T-21 and Dm-T-32; hereafter mentioned as “transgenic lines”) were extracted, resuspended in 0.1 M PBS) and quantified by Bradford method (1976). Around 20 μg of total soluble proteins from each sample were heated for 10 min at 85°C and heat-labile denatured proteins were removed by centrifugation (12,000g), and were separated on a 15% SDS-PAGE. Purified Dm-AMP1 (50 ng) was used as a positive control. Immunoblots were developed with antibodies raised against Dm-AMP1 using the alkaline phosphatase detection method (Bangalore Genei, Bangalore, India).
Estimation of Dm-AMP1 in leaf tissues of transgenic plants
Approximately 20 μg of protein extracted from the leaf tissues of transgenic lines was taken as the antigen for indirect ELISA using antibodies against Dm-AMP1. Known concentrations of purified Dm-AMP1 (2–100 ng) were used as a standard to plot absorbance (405 nm) versus concentration of purified Dm-AMP1 (ng per assay system), to estimate the levels of Dm-AMP1 in transgenic rice lines.
Immunohistochemical procedure
As measured by ELISA, Dm-T-32 had highest expression of Dm-AMP1. Thus, it was used for immunohistochemical localization studies. Leaves and roots from 21-day-old transformed (Dm-T-32) and control plants were fixed with 10% formaldehyde, 5% acetic acid, and 85% ethanol for 30 min at room temperature. The tissue was then dehydrated through a graded ethanol series (30 min each in 30%, 50%, 70%, 95%, and 100% EtOH). Dehydrated tissue was gradually embedded in paraffin wax. For sectioning, a rotary microtome (Weswox, India) and a freshly sharpened knife blade were used. Tissue ribbons with 8 μm transverse sections were fixed on clean glass slides. Slides were then deparaffinised in xylene for 10–15 min, and rehydrated through a graded ethanol series (10 min each in 100%, 95%, 70%, 50%, and 30% EtOH). In order to quench most of the tissue autofluorescence from chlorophyll and cell walls, sections were incubated in 0.1% Toluidine blue overnight. Sections of approximately 8 mm were prepared and processed for immunological detection of Dm-AMP1 protein (Patkar and Chattoo 2006). Anti-Dm-AMP1 antibody (1:100 dilution) and FITC-conjugated anti-rabbit IgG (Sigma Chemical Co, St Louis, MO, USA) diluted to 1:20, were used for immunological staining. The slides were observed under an epifluorescence microscope (Nikon, Eclipse 80i, Kanagawa, Japan) at ×200 and ×400 magnification. Control slides were treated with non-immune rabbit serum and FITC-conjugated goat anti-rabbit IgG.
Similarly, fungal mycelia were fixed and treated with purified Dm-AMP1. The samples were incubated for 90 min with anti-Dm-AMP1 antibody (diluted to 1:100) in blocking solution and then stained with FITC-conjugated anti-rabbit IgG raised in goat (Sigma Chemical Co, St Louis, MO, USA) diluted to 1:20 in PBS for 30 min. The slides were observed under an epifluorescence microscope at ×1,000 magnification.
Rice blast assay
Whole plant infection assays with M. oryzae (laboratory strain B157 corresponding to international race IC9) were carried out as described by Bonman and Mackill (1988). Briefly, 21-day-old transgenic plants and non-transformed control plants were inoculated with 1 × 105 spores/ml of M. oryzae using a manual sprayer. The spore suspension contained 0.05% Triton-X-100. The inoculated plants were incubated at 25°C with 90% relative humidity in the dark for 20 h and then maintained at 200 μmol/m2 s1 light intensity, and 14 h photoperiods at 28°C. Blast disease development was monitored as % diseased leaf area (% DLA), over a period of 10 days post inoculation (dpi).
Sheath blight assay
Non-transformed and transgenic lines were inoculated with R. solani as described by Jach et al. (1995). R. solani, was cultivated in 1 l sterile potato-dextrose broth (PDB) (Hi-Media, India) for 4–6 days at 28–30°C at 100 rpm. Mycelia were harvested and homogenised by vortexing in fresh sterile PDB medium; mixed with fresh soil and used as inoculated soil. About 25 seeds each from non-transformed plants and four transgenic lines were de-husked, surface sterilized by 70% ethanol and placed on solid Murashige and Skoog (MS) medium containing 3% sucrose (Jach et al. 1995). Seedlings were incubated on medium for 3–4 weeks and then transferred to un-inoculated soil for a week. About 4–5 week-old seedlings (20 seedlings each from non-transformed and 4 transgenic lines) were then transferred to inoculated soil and infection was monitored.
In vitro antifungal activity assay and microscopic studies
About 20 μg of total protein from leaf tissue of transgenic plants or 5 μg of purified Dm-AMP1 in 100 μl of 0.1 M PBS was used for all in vitro antifungal activity assay and microscopic studies unless otherwise stated. About 100 μl of M. oryzae (isolate B157; IC9) spore suspension (103 spores) were germinated on a glass slide at 28°C in the presence or absence of Dm-AMP1, mounted with appropriate concentration of glycerol to maintain constant relative humidity in sealed humidity chamber. Growth inhibition was assayed after 24 h, in terms of percent germination or altered morphology of hyphae as compared to the control samples. The treated and untreated samples were air dried, fixed in 10% formaldehyde, 5% acetic acid, and 85% ethanol for 30 min at room temperature, and washed thoroughly with water. The fixed samples were stained with 0.3 g/l calcofluor white (Sigma-Aldrich, USA) for 30 min at room temperature. Samples were washed with 0.1 M PBS (pH 7.0) before resuspending in the mounting medium. Glycerol was used as a mounting agent for fluorescence microscopy. All samples were examined at ×400 magnification under epifluorescence microscope (Eclipse 80i, 40X, Nikon, Japan), equipped with 100-W high-pressure mercury lamp (Osram HBO 103 W/2 N), and filter set for calcofluor white (UV-2A, Nikon, Japan).
Results
Characterization of Dm-AMP1 transformed plants
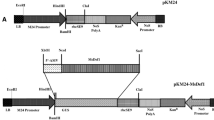
Scutellum-derived embryogenic Pusa Basmati 1 calli were used for Agrobacterium tumefaciens-mediated genetic transformation. The antibiotic resistance gene hpt was used as a selectable marker (Fig. 1b). About 78 out of 150 (52%) calli were found to be hygromycin resistant, of which 39 (50%) calli regenerated into whole plants and 25 (~65%) plants set seeds. About 25 hygromycin resistant plants were selected and analysed by PCR. A 280-bp PCR-amplified product corresponding to the size of Dm-AMP1 was observed in all the 25 putative transformed plants and the positive control (pUb1-Dm), but was absent in non-transformed plants (Fig. 1a; data of three representative lines are presented, data for other lines are not shown). Transformed lines were self-pollinated to obtain T1 plants. The T1 hygromycin-resistant plants were further self-pollinated to obtain T2 plants. Genomic DNA from each transgenic T2-line was digested with KpnI or SacI and hybridized with the Dm-AMP1 DNA probe to analyse the integration events (Fig. 1c). Single bands were detected in four lines (Dm-T-14, Dm-T-16, Dm-T-21 and Dm-T-32), suggesting that these lines have single site of integration for the transgene. Conversely, the multiple bands (2–3 bands) were detected in other lines (Dm-T-18, Dm-T-25) (Fig. 1c). The four independent transgenic lines with single transgene integration were used for further studies.
(a) Total genomic DNA from three representative transgenic plants was extracted and amplified using specific primers for Dm-AMP1 coding region. lane 1: 100 bp DNA ladder (Cat no: 105993: Bangalore Genei Pvt Ltd, Bangalore, India); lane 2: negative control (non-transformed Pusa basmati 1); lane 3–5: Dm-AMP1 transformed rice; lane 6: positive control (pUb1-Dm). (b) Plant expression vector p1305-Ub-Dm-T. pUb1: Maize ubiquitin promoter; SP: Signal peptide; tNos: 3′ untranslated terminator region of the Agrobacterium tumefaciens nopaline synthase gene; p35S: CaMV35S promoter; hpt: hygromycin phosphotransferase. (c) Southern blot analysis of KpnI-digested genomic DNA from non-transformed and representative Dm-AMP1- transformed transgenic plants in T2 generation. Genomic DNA (10 μg) was digested with restriction enzymes and blotted onto nitrocellulose membranes after electrophoresis in a 0.8% agarose gel. The membrane was hybridized with 280 bp Dm-AMP1 coding sequence; lane 1: Ladder (NEB 2-log ladder, Cat. No. N3200S); lane 2: Dm-T-18; lane 3: Dm-T-21; lane 4: Dm-T-16; lane 5; Dm-T-25; lane 6: Dm-T-14; lane 7: Dm-T-32; lane 8: negative control (non-transformed Pusa basmati 1)
Accumulation of Dm-AMP1 in rice plants
Leaves of T2 seedlings were analysed by western blot to detect the presence of Dm-AMP1. The transformed plants showed the expression of Dm-AMP1 protein, co-migrating with the purified protein, which was absent in the control (Fig. 2a). The Dm-AMP1 protein was also detected in roots (Fig. 2b) as well as in seeds (Fig. 2c) of the transgenic plants. The plants showing Dm-AMP1 expression were analysed by ELISA and the expression levels obtained in the leaf extracts of Dm-T-16, Dm-T-21, Dm-T-14 and Dm-T-32 were 4.33 ± 0.42, 4.54 ± 0.72, 4.82 ± 0.48 and 5.69 ± 0.61 μg/mg of total soluble proteins, respectively (Fig. 3). These lines were selected for analysis of disease resistance.
Immunodetection of Dm-AMP1 in transgenic rice lines. Total leaf protein extracts from T2-progeny plants were subjected to immunoblot analysis with polyclonal anti-Dm-AMP1. About 20 μg of total soluble proteins from each sample (T2-transformed and non-transformed rice plants) were separated on a 15% SDS-PAGE. GST-Dm-AMP1 was cleaved with factor Xa to get purified Dm-AMP1 and 50 ng of purified Dm-AMP1 was used as positive control. (a) Leaves; lane 1: Marker (PMWB, Bangalore genei Pvt Ltd, Banglore, India); lane 2: Dm-T-21; lane 3: Dm-T-16; lane 4: Dm-T-14; lane 5; Dm-T-32; lane 6: positive control; (b) Root; lane 1: positive control; lane 2: Dm-T-21; lane 3: Dm-T-16; lane 4: Dm-T-14; lane 5; Dm-T-32; lane 6: Marker (PMWB, Bangalore genei Pvt Ltd, Banglore, India); (c) Seed; lane 1: positive control; lane 2: Dm-T-21; lane 3: Dm-T-16; lane 4: Dm-T-14; lane 5; Dm-T-32; lane 6: negative control (non-transformed rice plants)
ELISA of T2-transgenic rice plants expressing Dm-AMP1. Levels of Dm-AMP1 in different T2-lines were estimated by indirect ELISA. Concentration of Dm-AMP1 in the protein extracts from transgenic leaf tissues was estimated by plotting the absorbance values on standard graph prepared using known concentrations of purified Dm-AMP1. The data are presented as means ± SEM from three independent estimations. The value obtained for the non-transformed plant was subtracted from the values for the Dm-AMP1-lines to eliminate the background levels
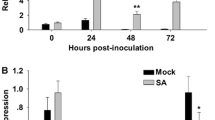
Transgenic plants constitutively expressing certain transgenes are known to activate pathogenesis-related proteins, which are only activated during pathogenesis in wild-type plants (Mittler et al. 1995). Thus, expression of PR-1 type genes has been widely used to indicate the induction of defense responses. To explore a possible activation of the plant defense system in the DmAMP1 transgenic rice lines, the expression of the endogenous PR-1a gene was analyzed at the RNA level, without challenging the plants with pathogen. The transcripts for PR-1a were not detected in DmAMP1 transgenic lines (Fig. 4). As expected, PR-1a transcripts were detected in M. oryzae infected control and transformed plants (Fig. 4, lane 1 and 6).
Northern-blot analysis of Dm-AMP1-expressing lines for the expression of rice PR-1a gene. About 20 μg/sample of total RNA were separated in a formaldehyde/agarose gel, transferred onto nylon membranes and hybridized to a non-radioactively labeled PR1a probe. (a) lane 1: M. oryzae-infected non-transformed plants; lane 2: control, uninfected non-transformed plants; lane 3: uninfected Dm-T-16; lane 4: uninfected Dm-T-14; lane 5; uninfected Dm-T-32; lane 6: M. oryzae-infected Dm-T-32 line. (b) rRNA bands (28 and 18S) from an ethidium bromide-stained gel of the corresponding RNA samples indicate equivalent loading of RNA
Localization of the Dm-AMP1 protein in rice and M. oryzae
Subcellular localization of Dm-AMP1 was done immunohistochemically. Dm-AMP1 was observed in the apoplast of leaves (Fig. 5a) and roots (Fig. 5b) of the transformed rice, while cytoplasmic content remained unlabelled.
(a) Immunolocalization of Dm-AMP1 in transgenic rice leaf. Transverse sections were prepared from leaf tissues of Dm-AMP1 transgenic lines and control non-transformed plants; A: Transverse sections of transgenic rice leaf; B: control non-transformed leaf tissue. Sections ‘A’ and ‘B’ were treated with anti-Dm-AMP1 and FITC-conjugated goat anti-rabbit IgG. Bar corresponds to 10 μm. (b) Localization of Dm-AMP1 in transgenic rice root. Transverse sections were prepared from root tissues of Dm-AMP1 transgenic lines and control non-transformed plants; A: Transverse sections of transgenic root; B: control (non-transformed root tissue). Sections ‘A’ and ‘B’ were treated with anti-DmAMP1 and FITC-conjugated goat anti-rabbit IgG. Bar corresponds to 10 μm
Further, tissue localization of Dm-AMP1 was studied by treating mycelia with Dm-AMP1 and anti-DmAMP1 antibody followed by FITC-conjugated goat anti-rabbit IgG. Distinct fluorescence was observed on the fungal membrane. Since, no additional proteins were tried for the binding assay possibility of non-specific binding could not be negated (data not shown).
Enhanced fungal disease tolerance of transgenic rice expressing Dm-AMP1
In vitro bioassays and in planta infection assays were performed by challenging four plants from each of four transgenic lines with M. oryzae and R. solani to evaluate the degree of resistance imparted by expression of Dm-AMP1. Development of necrotic areas on leaves and stem of control and transformed plants was monitored. When transgenic plants were challenged with M. oryzae, there was a remarkable difference in diseased leaf area (%DLA) between non-transformed (69.68 ± 2.52) and transgenic lines (11.12 ± 0.65 to 18.35 ± 1.74) at 99% confidence interval (CI) (Fig. 6a). Similarly, when plants were challenged with R. solani, the % infected plants ranged from 18.58 ± 4.82 to 28 ± 6.88, whereas it was 66.33 ± 3.55 in non-transformed plants at 99% CI (Fig. 6b).
(a) in planta bioassay of M. oryzae (105 spores/ml) on transgenic lines. A: Whole plant infection assay of non-transformed control (C) and transgenic plants; B: Blast infected leaves of Dm-AMP1 lines and non-transformed control rice plants. Infected leaf area was measured as % DLA (Diseased leaf area) at 10 dpi. The data are presented as means ± SEM from three independent estimations. (b) in planta bioassay of Dm-AMP1 transgenic lines with R. solani. The number of plants showing disease symptoms was used as disease scoring system. A: Whole plant infection assay of non-transformed control (C) and transgenic lines; B: Response of transgenic lines and non-transformed plants to sheath blight. Twenty seeds, each from non-transformed plants and the transgenic lines were used for sheath blight assay. The data are presented as means ± SEM from three independent estimations
Development of disease symptoms on inoculated leaves was monitored microscopically. Tissue sections of the inoculated leaves were cleaned with 6% KOH, stained with calcofluor white, and examined under epifluorescence microscope. In control (non-transformed) leaves infected with M. oryzae spores, extensive fungal development was observed. In contrast, under the same experimental conditions, reduced fungal colonization was observed in leaves of the Dm-AMP1 lines (Fig. 7). Inhibition of growth was observed, when M. oryzae was treated with total soluble protein from transgenic lines (data not shown). These results further support that constitutive expression of Dm-AMP1 suppresses the growth of M. oryzae.
Disease response of transformed and non-transformed rice leaves challenged with M. oryzae spores. Twenty-one day old transgenic lines and non-transformed control plants were inoculated with M. oryzae. Transverse sections of Dm-AMP1 transgenic lines and non-transformed control were prepared. The samples were stained with calcofluor white. A: Transverse sections of transgenic leaves with its corresponding bright field image; B: Non-transformed control with its corresponding bright field image. Bar corresponds to 10 μm
Discussion
The important role played by antimicrobial proteins in plant defense has been well documented (Broekaert et al. 1995; Terras et al. 1995; de Bolle et al. 1996; Francois et al. 2002). The expression of antimicrobial proteins has been used to engineer disease resistance in tobacco, scented geranium, potato and rice (Terras et al. 1995; de Bolle et al. 1996; Bi et al. 1999; Gao et al. 2000; Patkar and Chattoo 2006). Transgenic plants have the potential to provide a generic resistance against different pathogens, and are likely to reduce dependence on chemical pesticides.
Correct processing and higher expression of antimicrobial peptides in transgenic plants is important for in planta antimicrobial activity. Dm-AMP1 protein folds into a small and compact β-barrel, which is stabilized by four internal disulfide bridges (Yun et al. 1997). Secretory proteins have to pass through endoplasmic reticulum for correct folding and assembly of proteins (Vitale and Denecke 1999). Therefore, in the present work, the secretory signal of Dm-AMP1 was used to target Dm-AMP1 to the apoplastic region for its proper folding. Dm-AMP1 was produced constitutively since the maize ubiquitin promoter was used to drive the expression of transgene. This is an appropriate strategy for the control of pathogens, like M. oryzae, which can infect through the plant roots (Sesma and Osbourn 2004), stems or leaves.
The independent transgenic plants showed a range of Dm-AMP1 expression levels, which might be due to position effects, and/or the effects of transgene copy number (Meyer 1998). The expression level of Dm-AMP1 in transgenic lines ranged between 0.43% and 0.57% of the total soluble protein, which was four times higher than papaya and tobacco plants transformed with Dm-AMP1, Mj-AMP2 and Ac-AMP1 (Zhu et al. 2007; de Bolle et al. 1996). There was a significant correlation between the levels of Dm-AMP1 and the degree of disease resistance observed in a particular line. Among the four transgenic lines, Dm-T-32 showed highest level of transgene product of 5.69 μg/mg of total soluble protein, and a correspondingly higher level of resistance to M. oryzae and R. solani. In addition, all transgenic plants were healthy and showed no morphological or developmental abnormalities even in the lines with the highest Dm-AMP1 accumulation. Thus, the expression level of DmAMP1 in rice plants was sufficient to maintain its antifungal activity without any toxic effects to the host plant.
The results of in vitro inhibition assays suggest that Dm-AMP1 inhibits the growth of a broad range of fungal pathogens (Osborn et al. 1995; Thomma et al. 2002). However, it was not known whether the expression level in rice would be sufficient for the inhibition of M. oryzae and R. solani. Our study shows that the level of constitutive expression of the Dm-AMP1, as evaluated by in vitro growth inhibition and in planta assays, was sufficient to inhibit the growth of M. oryzae and R. solani. The plants expressing Dm-AMP1 protein showed significantly improved resistance to M. oryzae and R. solani, by 84% and 72%, respectively, as compared to non-transformed plants. We observed a consistent difference in the severity of tissue damage between control and Dm-AMP1 transgenic plants, when they were monitored for the development of disease symptoms. Constitutively-expressed Dm-AMP1 gene in rice appears to inhibit the proliferation of M. oryzae hyphae at the infection site. This was observed when plants were challenged with M. oryzae; small lesions were observed, which did not spread to form the large lesions as seen on the control plants. Further evidence for enhanced resistance comes from growth anomalies seen in the fungal hyphae in the Dm-AMP1 transgenic lines. Hyphae showing similar alterations in their morphology have been described in transgenic plants over-expressing Ace-AMP1 (ns-LTP from Allium cepa) and other defensins (Patkar and Chattoo 2006; Epple et al. 1997). Furthermore, transgene expression of the Dm-AMP1 gene was not accompanied by an induction of pathogenesis-related (PR) gene expression, indicating that the transgene product itself is directly active against the pathogen. Interestingly, the expression of Dm-AMP1 was localized to extracellular space of leaves and roots of the transgenics, as seen in Ace-AMP1 (Patkar and Chattoo 2006). Expression of AMPs in the apoplast of transgenic plants is preferred because most pathogenic microorganisms are present in the extracellular space during the early stages of infection.
Dm-AMP1 was found to target the fungal membrane as observed in the localization experiment (data not shown). It has been suggested that growth inhibition may be due to membrane destabilization (Thevissen et al. 1996). Interaction between Dm-AMP1 and the fungal membrane signifies that this binding might be essential for inhibitory activity against M. oryzae. Dm-AMP1 is active against the phytopathogenic fungus Botrytis cinerea, whereas it did not interfere with recognition responses and symbiosis establishment by the arbuscular mycorrhizal fungus Glomus mosseae, indicating that it may target the potential pathogenicity factors present in pathogenic fungi (Turrini et al. 2004).
The results of the current study indicate that Dm-AMP1 expressed in the apoplastic region of the plant tissue appears to interact with the fungal membrane leading to membrane destabilization, and reduced proliferation of fungal pathogen inside the plant tissue, thus imparting enhanced disease resistance to a broad-spectrum of fungal phytopathogens.
References
Anderson NA (1982) The genetics and pathology of Rhizoctonia solani. Annu Rev Phytopathol 20:329–347. doi:10.1146/annurev.py.20.090182.001553
Bi YM, Cammue BPA, Goodwin PH, KrishnaRaj S, Saxena PK (1999) Resistance to Botrytis cinerea in scented geranium transformed with a gene encoding the antimicrobial protein Ace-AMP1. Plant Cell Rep 18(10):835–840. doi:10.1007/s002990050670
de Bolle MFC, Osborn RW, Goderi IJ, Noe L, Acland D, Hart CA et al (1996) Antimicrobial peptides from Mirabilis jalapa and Amaranthus caudatus: expression, processing, localization and biological activity in transgenic tobacco. Plant Mol Biol 31:993–1008. doi:10.1007/BF00040718
Bonman JM, Mackill DJ (1988) Durable resistance to rice blast disease. Oryza 25:103–110
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. doi:10.1016/0003-2697(76)90527-3
Broekaert WF, Terras FR, Cammue BP, Osborn RW (1995) Plant defensins: novel antimicrobial peptides as components of the host defense system. Plant Physiol 108:1353–1358. doi:10.1104/pp.108.4.1353
Chen DC, Yang BC, Kuo TT (1992) One-step transformation of yeast in stationary phase. Curr Genet 21:83–84. doi:10.1007/BF00318659
Christensen AH, Quail P (1996) Ubiquitin promoter-based vectors for high-level expression of selectable and/or screenable marker genes in monocotyledonous plants. Transgenic Res 5:213–218. doi:10.1007/BF01969712
Dellaporta SL, Wood J, Hicks JB (1983) A plant DNA minipreparation: version 2. Plant Mol Biol Rep 1:19–22. doi:10.1007/BF02712670
Epple P, Apel K, Bohlmann H (1997) Overexpression of an endogenous thionin enhances resistance of arabidopsis against Fusarium oxysporum. Plant Cell 9:509–520
Francois IE, De Bolle MF, Dwyer G, Goderis IJ, Woutors PF, Verhaert PD et al (2002) Transgenic expression in Arabidopsis of a polyprotein construct leading to production of two different antimicrobial proteins. Plant Physiol 128:1346–1358. doi:10.1104/pp.010794
Gao AG, Haikimi SM, Mittanck CA, Wu Y, Woerner BM, Stark DM et al (2000) Fungal pathogen protection in potato by expression of a plant defensin peptide. Nat Biotechnol 18:1307–1310. doi:10.1038/82436
Gu Q, Kawata EE, Morse MJ, Wu HM, Cheung AY (1992) A flower-specific cDNA encoding a novel thionin in tobacco. Mol Gen Genet 234:89–96
Hiei Y, Ohta S, Komari T, Kumashiro T (1994) Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6(2):271–282. doi:10.1046/j.1365-313X.1994.6020271.x
Jach G, Gornhardt B, Mundy J, Logemann J, Pinsdorf E, Leah R et al (1995) Enhanced quantitative resistance against fungal disease by combinatorial expression of different barley antifungal proteins in transgenic tobacco. Plant J 8(1):97–109. doi:10.1046/j.1365-313X.1995.08010097.x
Komari T, Hiei Y, Saito Y, Murai N, Kumashiro T (1996) Vectors carrying two separate T-DNAs for co-transformation of higher plants mediated by Agrobacterium tumefaciens and segregation of transformants free from selection markers. Plant J 10:165–174. doi:10.1046/j.1365-313X.1996.10010165.x
Logeman J, Schell J, Willmitzer L (1987) Improved method for the isolation of RNA from plant tissues. Ann Biochem 163:16–20. doi:10.1016/0003-2697(87)90086-8
Mitchell DA, Marshall TK, Deschenes RJ (1993) Vectors for the inducible expression of glutathione S-transferase fusion proteins in yeast. Yeast 9:715–723. doi:10.1002/yea.320090705
Mittler R, Shulaev V, Lam E (1995) Coordinated activation of programmed cell death and defense mechanisms in transgenic tobacco plants expressing a bacterial proton pump. Plant Cell 7:29–42
Osborn RW, De Samblanx GW, Thevissen K, Goderis I, Torrekens S, Van Leuven F et al (1995) Isolation and characterisation of plant defensins from seeds of Asteraceae, Fabaceae, Hippocastanaceae and Saxifragaceae. FEBS Lett 368:257–262. doi:10.1016/0014-5793(95)00666-W
Ou SH (1985) Rice Diseases, 2nd edn. Commonwealth Mycological Institute Publication, Kew, Surrey, UK, pp 280–282
Patkar RN, Chattoo BB (2006) Transgenic indica rice expressing ns-LTP like protein shows enhanced resistance to both fungal and bacterial pathogens. Mol Breed 17:159–171. doi:10.1007/s11032-005-4736-3
Sesma A, Osbourn AE (2004) The rice leaf blast pathogen undergoes developmental processes typical of root-infecting fungi. Nature 431:582–586. doi:10.1038/nature02880
Terras FRG, Eggermont K, Kovaleva V, Raikhel NV, Osborn RW, Kester A et al (1995) Small cysteine-rich antifungal proteins from radish: their role in host defence. Plant Cell 7:573–588
Thevissen K, Ghazi A, de Samblanx GW, Brownlee C, Osborn RW, Broekaert WF (1996) Fungal membrane responses induced by plant defensins and thionins. J Biol Chem 271:15018–15025. doi:10.1074/jbc.271.25.15018
Thevissen K, François IEJA, Takemoto JY, Ferket KKA, Meert EMK (1997) Specific, high affinity binding sites for an antifungal plant defensin on Neurospora crassa hyphae and microsomal membranes. J Biol Chem 272:32176–32181
Thevissen K, Cammue BP, Lemaire K, Winderickx J, Dickson RC, Lester RL et al (2000) A gene encoding a sphingolipid biosynthesis enzyme determines the sensitivity of Saccharomyces cerevisiae to an antifungal plant defensin from dahlia (Dahlia merckii). Proc Natl Acad Sci USA 97:9531–9536. doi:10.1073/pnas.160077797
Thevissen K, Francois IE, Takemoto JY, Ferket KK, Meert EM, Cammue BP (2003) DmAMP1, an antifungal plant defensin from dahlia (Dahlia merckii), interacts with sphingolipids from Saccharomyces cerevisiae. FEMS Microbiol Lett 226:169–173. doi:10.1016/S0378-1097(03)00590-1
Thomma BPHJ, Cammue BPA, Thevissen K (2002) Plant defensins. Planta 216:193–202. doi:10.1007/s00425-002-0902-6
Turrini A, Sbrana C, Nuti MP, Pietrangeli BM, Giovannetti M (2004) Development of a model system to assess the impact of genetically modified corn and aubergine plants on abruscular mycorrhizal fungi. Plant Soil 266:69–75. doi:10.1007/s11104-005-4892-6
Vitale A, Denecke J (1999) The endoplasmic reticulum-gateway of the secretory pathway. Plant Cell 11:615–628
Yun DJ, Bressan RA, Hasegawa PM (1997) Plant antifungal proteins. Plant Breed Rev 14:39–88
Zhu YJ, Agbayani R, Moore PH (2007) Ectopic expression of Dahlia merckii defensin DmAMP1 improves papaya resistance to Phytophthora palmivora by reducing pathogen vigor. Planta 226(1):87. doi:10.1007/s00425-006-0471-1
Acknowledgments
We thank Dr. B. Cammue, University of Leuven, Belgium, for providing the plasmid pFAJ3105. We also extend our thanks to Prof. R. N. Pandey, Anand Agricultural University, Anand, India, for providing cultures of R. solani. This work was supported by the grants from Department of Biotechnology, Ministry of Science and Technology, Government of India.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jha, S., Tank, H.G., Prasad, B.D. et al. Expression of Dm-AMP1 in rice confers resistance to Magnaporthe oryzae and Rhizoctonia solani . Transgenic Res 18, 59–69 (2009). https://doi.org/10.1007/s11248-008-9196-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11248-008-9196-1