Abstract
Grapevines are challenged by a range of diseases and pests, causing economic losses and requiring often costly approaches to mitigate damage. Public interest in reducing the use of chemicals is a related challenge, along with climate change. Yet, the Vitis gene pool provides vast resources for the development of genetic resistance in rootstock and scion cultivars. Traditional breeding approaches have made great strides in the development of adaptive traits, and recent access to ‘omic technologies has further facilitated the identification of useful loci along with rapid trait introgression from wild species. Moreover, marker technologies are now used to stack multiple genes for the same trait into a single genotype, a heretofore barely accessible technology. Genomic technologies are also impacting germplasm characterization, and thereby facilitating “Breeding by Design” approaches. Genetic transformation and gene-editing technologies are also applicable for both cultivar improvement as well as functional studies of genes. The landscape for acceptance of new resistant cultivars is complex and with wine grapes, subject to high degrees of regulation especially in the European Union. With rootstocks, as well as table/raisin grapes, gaining acceptance in the marketplace for new cultivars developed through either traditional or marker-assisted approaches is routine. Yet even in the highly regulated EU environment, the adoption of new wine cultivars of interspecific origins is beginning to take place in both traditional wine growing regions as well as non-traditional regions nearby.
Reisch and Hausmann: Co-last authors, equally contributed
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
4.1 Introduction
Viticulture and winemaking are largely recognized worldwide, having a strong socio-economic role for many countries. These techniques and processes developed by man are based on the cultivation of the vine. Most cultivated grapevines belong to the Eurasian species Vitis vinifera L. Although 12,000 names are reported, the actual number of vine varieties for the V. vinifera species in the world is estimated at 6,000. ‘Kyoho’, ‘Cabernet-Sauvignon’, and ‘Sultanina’ are the most abundant varieties grown worldwide (OIV 2017). In 2019 the global surface area planted with vines for all purposes (wine, table grapes, juice and raisins), including young vines not yet in production, is estimated at 7.4 mha (EUROSTAT 2019). In 2019, world wine production, excluding juices and musts, is estimated at 260 mhl and the world wine export market—considered as the sum of the exports of all countries—has expanded with respect to 2018 both in volume and in value. Strong increases can be observed in exports from Italy, Spain, Canada and Chile; France was still the most important world exporter in terms of value. Bottled wines (<2 L) represented 53% of trade volumes globally, a share in line with 2018. Sparkling wine once again saw a significant growth in 2019, in terms of both volume and value. This can be partially explained by the ongoing trend for the Italian Prosecco wine throughout the world (OIV 2020). The European Union (EU) is the world-leading producer of wine. Between 2014 and 2018, the average annual production was 167 mhl. It accounts for 45% of world wine growing areas, 65% of production, 60% of global consumption and 70% of exports (EUROSTAT 2019).
Besides spirits, distillates, and liquors, in recent years, there has been a great deal of interest in non-alcoholic products of the viti-vinicultural sector intended for human consumption. The discovery that both the edible flesh of the grape and its by-products (such as the seeds and skin) contain components that are beneficial to human health (called “nutraceuticals”) has led to rapidly expanding markets for grapes and their by-products. For example, grape seed extracts have been used as nutritional supplements in fruit flavoured beverages, cereals, snack bars, and dairy desserts such as yogurt. Grape leaves have been used to stop bleeding, inflammation and pain. Unripe grapes used to treat sore throats and dried grapes (raisins) to cure constipation and thirst. Round, ripe, and sweet grapes have been used to treat a range of health problems including cancer, cholera, smallpox, nausea, eye infections, skin, kidney, and liver diseases. Grape phenolics have been proved to possess several health promoting properties playing an important role in the inhibition of carcinogenesis, mutagenesis, and cardiovascular diseases. These activities have been associated with their in vivo and in vitro antioxidative activities. Flavonoids in grape seeds have also been reported to exhibit activities against peptic ulcer and several dermal disorders (reviewed by Ali et al. 2010). In addition, grape berry and seed extracts are used in cosmetics and personal care products (FAO-OIV 2016). The vine-trend has landed in these sectors also thanks to “Vinotherapy” (wine therapy), based on the discovery of new ingredients extracted from various organs. Approximately 75% of produced grapes is intended for wine production, out of which 20–30% represents waste products. This waste is also called “grape pomace” and consists of skins, remaining pulp, seeds, and stalks. These by-products represent waste disposal or they are used for wine alcohol production, serve as fertilizer or as animal feed (Antonić et al. 2020). Noble alternatives to the use of whole grapes and seeds are represented by the production of (balsamic) vinegar and virgin oil. Processing grapes into juice is definitely a minor reality compared to winemaking. In fact, FAOSTAT (2014) reports 73.7% world production concentrated in the Americas, with only 21% and 5.1% in Europe and Asia, respectively. In addition, the top world grape juice producers are not the same as the top wine producers, with the USA the leading one, followed by Spain, Argentina, Chile, and Brazil. Especially in the latter, new juice grape cultivars, with shorter phenological cycles, higher glucometric potential and adapted to tropical climates point to a progressive and increasing substitution of traditional cultivars for juice production (Spigno et al. 2017).The impact of the Covid-19 crisis on wine producers varied depending on their sales focus. Smaller wineries were particularly affected by the closures of restaurants and hotels and the lack of tourists. The simultaneous global impact of the pandemic also led to a global decline in wine exports, especially to countries with a high proportion of wine consumption at social events and in restaurants. For the majority of wine producers in Spain, France and Italy, several of their strongest distribution channels, in terms of value and volume, have been negatively affected at the same time. These effects, by far, could not be compensated by increases in online sales (ProWein Business report, Loose 2020).
Concomitantly, there is another ongoing crisis due to climate change. Considering all threats and challenges, 73% of wine businesses expect a very likely or likely effect of climate change on their business (ProWein Business report, Loose and Pabst 2019). Tackling climate change will be much cheaper than the disruption that global heating will cause, as well as bringing benefits to health. During the twentieth century, significant changes in temperatures were recorded, including increases from 2 to 5 °C in Europe, which is home to world-renowned wine regions. Moreover, decreases in the precipitations over Southern Europe were also observed. According to the latest report of the intergovernmental panel on climate change (IPCC 2013), following different representative concentration pathways, global temperature is expected to rise between 1 °C (least severe scenario) and 5° (most severe scenario) over the twenty-first century. Very recently, proxy-based reconstructions demonstrate that the modern global temperature has exceeded annual levels over the past 12,000 years and probably approaches the warmth of the last interglacial period (Bova et al. 2021). Climate is an important factor impacting grapevine physiological development, vegetative growth, phenology, production, and consequently, wine quality. Climatic factors also determine the geographical location of vineyards. In fact, over the past years the land area devoted to viticulture at the cooler end of the suitability spectrum has increased globally, reflecting the trend towards a warmer climate (Vigl et al. 2018). Moreover, the variability in the weather parameters, such as air temperatures, precipitation, and solar radiation, leads to annual changes in productivity. Weather extremes are also known to have detrimental impacts on grape yield and quality, namely hail, late frost spells, and excessive rainfall (Fraga et al. 2019). Besides having already tremendously impacted grape yield, dramatic changes in climate are expected to influence the incidence of biotic threats. In fact, temperature, humidity, and wind speed may also affect the different steps of the reproductive cycle of pathogens and pests, destabilizing even further the precarious equilibrium with the host plant (Caffarra et al. 2012).
Diseases caused by biotic factors (fungi, oomycetes, bacteria, phytoplasmas, viruses and nematodes) as well as pests (insects and mites) producing disease-like symptoms (Agrios 2004; Wilcox et al. 2015) cause a high expenditure of plant protection measurements in order to ensure the quantity and quality of the harvest. In almost all grape growing regions, either powdery or downy mildew (PM or DM) is the most serious disease that has to be controlled regularly at great expense. In a recent study for the state of California, Sambucci et al. (2019) calculated total pecuniary costs of $239 million for PM management (fungicides and application) in 2015 (without non-monetary costs for environmental pollution and risks to human health). They expect significant cost savings if new grapevine varieties resistant to PM are cultivated. Similar results can be expected for other diseases, i. e. the conclusions are generally true and promising. In addition to these economic reasons of the grape growers, the concerns of society and consumers about the use of chemicals are playing an increasingly important role and are being taken into account by legislators. Some varieties with resistance to PM and DM (also sometimes called “PIWI”, a German abbreviation for pilzwiderstandsfähig/ fungus-resistant; https://piwi-international.de/en/) already exist and allow reduced chemical plant protection applications. However, other fungal diseases can appear under these circumstances and infect susceptible PIWIs. For instance, black rot (BR) can lead to heavy yield losses, even to total failure in extreme cases, and has to be mitigated to avoid economic damage (Molitor and Beyer 2014). Grapevine trunk diseases (esca complex) became a limiting factor in viticulture in many countries in the recent past and the expenses for the replacement of dead vines are estimated at over $1.5 billion per year. Prolonged droughts as an effect of global warming will lead to more incidences, higher severities and ultimately increase economic damages (Fischer and Ashnaei 2019).
Our modern life also includes international trade and travel that favors the spread of alien species. The bacterium Xylella fastidiosa, causing Pierce's disease (PD), is native to southern areas of North and Central America and transmitted by leafhoppers. In California, it was rather a minor problem for viticulture until a non-native insect vector emerged, leading to devastating PD outbreaks in the late 1990s. For 2010, the direct economic losses were calculated to at least $56 million (Tumber et al. 2014). Xylella fastidiosa is now present in Spain, Turkey, Iran and Taiwan and is therefore of global concern (Godefroid et al. 2019). Another example involving an invaded insect vector is Flavescence dorée (FD), a disease mainly present in Europe. Causative agent is a phytoplasma transmitted by the leafhopper Scaphoideus titanus that originated in North America and emerged in Europe in the middle of the last century. Although FD is a quarantined organism, it is still spreading and causes yield losses and lower grape quality. For compensating the economic damages of FD, Italian grape growers were given €34 million in 2005 (Chuche and Thiéry 2014). Rootstocks were originally developed because of phylloxera, a pest that attacks the root. Grafted vines have proven to be effective and practical to tackle phylloxera, but mean otherwise additional costs for the grower. If fungus-resistant scions can be bred, it should also be possible to add resistances to phylloxera in the future. In the long run, this would make it possible to return to own-rooted grapevines and save the costs for rootstocks and grafting. Considering the urgent and challenging necessity of the implementation of sustainable viticulture with fewer fungicides and pesticides, new varieties resistant to pathogens and pests are highly desirable and should be tested in the long term for possible erosion of resistance, design of best cropping systems and adaptation to climate change (Delrot et al. 2020).
In this overall frame, it is therefore even more important to develop ad hoc grapevine ideotypes, capable of also leading to more competitive (niche) products and by-products. To achieve this, traditional genetic improvement–which has always operated at the service of the germplasm enhancement–is no longer sufficient and it has to make use of genetics and genomics tools, integrated with information from other ‘omic disciplines. Genetics is already supporting the classical genetic improvement through marker-assisted breeding (MAB). Indeed, genomics-assisted breeding (GAB) is still in its infancy but is a promising direction. Traditional breeding will be supported by re-sequenced germplasm repositories and numerous fully haplophased genome assemblies due to improved sequencing technologies and data mining (bioinformatics) approaches. The concept of “Breeding by Design” (Peleman et al. 2005) will become increasingly important as the full range of loci impacting important traits are identified, and as variability at each locus is better understood. In parallel, to overcome the genetic transformation limiting factors and issues, the novel technologies of genome or tailored gene editing offer new opportunities to be scouted and exploited towards the establishment of resistances to grapevine biotic stresses.
4.2 Description on Main and Emerging Diseases
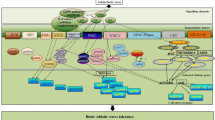
A wide range of diseases afflicts the Eurasian grape species (V. vinifera L.) affecting production and fruit quality, processing and exportation. Grapevine is known to host the widest variety of pathogens of any woody agricultural plant (Martelli 1997), including fungi, oomycetes, bacteria, phytoplasmas, viruses, and plant-parasitic nematodes with different infection mechanisms, life cycles and survival strategies (Fig. 4.1). All parts of the grapevine plant are subjected to attack by these organisms including the roots, trunk, arms, cordons, canes, shoots, leaves, rachis and berries. The importance of disease management in grapevine is critical, and derives from the relatively high value of the crop, the substantial annual production costs and its exposure to emerging pathogens, particularly since the mid nineteenth century, because of human activity and travel. One reason for the susceptibility of V. vinifera cultivars to some of the major pathogens is that these organisms are not indigenous to Eurasia, and as such, there has been no selection pressure to evolve resistance.
Examples of fungal, complex and viral disease symptoms on grapevine organs: a Black rot (Phyllosticta ampelicida), b Phomopsis cane and leaf spot (Diaporthe spp.), c Esca, d Grapevine yellows (Ca. Phytoplasma), e Fanleaf yellowing (fanleaf grapevine Nepovirus), f Petri disease (Phaeomoniella chlamydospora and other fungi), and root galls caused by nematode parasites, g Meloidogyne javanica, and h Xiphinema index
4.2.1 Caused by Fungi and Oomycetes
Among the wide range of pathogens affecting grapevine, fungal pathogens are of significant importance since V. vinifera is susceptible to 29 fungal diseases (Wilcox et al. 2015), including the emergent ascomycetes associated with anthracnose, BR and PM (Pirrello et al. 2019) and the re-emergence of fungal grapevine trunk diseases (Gramaje et al. 2018). The oomycete causal agent of DM is also one the most damaging threats affecting viticulture worldwide. Both fungi and oomycetes can differentially uncover diverse climatic conditions, from temperate-humid to warmer-dry climate regions (Bois et al. 2017).
4.2.1.1 Powdery Mildew
The causal agent of PM is the biotroph fungus Erysiphe necator Schw. (Gadoury et al. 2012). All above-ground parts of grapevine can be infected by the growing of this pathogen upon their epidermis. The fungus is easily recognized by the appearance of a whitish-gray powdery-appearing growth on the green tissues due to the presence of its mycelium and conidia (Pirrello et al. 2019). The infection usually begins on the lower leaf surface and often produces chlorotic spots on the upper side. Severely affected leaves usually undergo senescence, develop necrotic blotches, and fall prematurely. On stems, the infection produces similar symptoms to those on leaves, with affected areas turning black, as epidermal cells are killed (Gadoury et al. 2019). On berries, prebloom infection of the cluster or within one week after their formation is the most severe manifestation of the disease, causing the cease of berries growing and often the skin to split making them susceptible to infection by other pathogens (Gadoury et al. 2019). Field assays for the assessment of disease tolerant grapevine material upon natural infections are carried out by visual observations (Li 1993; Wang et al. 1995; Pap et al. 2016). These observations are made mostly in mid-summer, when the symptoms become more evident and fully developed and it is possible to estimate the occurrence of the pathogen in vineyards treated and not-treated with fungicide (Li 1993; Wang et al. 1995).
Powdery mildew is a polycyclic disease that evolves in two distinct phases, (i) the primary infection, caused by ascospores (sexual spores), and (ii) secondary infections, caused by conidia (asexual spores). Its epidemiology is well-studied (Gadoury et al. 2012). Pathogenic specialization in E. necator has been demonstrated on wild vines and Parthenocissus spp. (Gadoury and Pearson 1991; Frenkel et al. 2010; Gur et al. 2021). Direct evidence for race specificity resistance has been available for E. necator. Ramming et al. (2012) demonstrated the presence of fungal races differentially interacting with race-specific resistance genes with susceptible and resistant progeny artificially infected with several E. necator isolates under controlled conditions. Race specificity of “resistance to Uncinula necator” (Run)1/Run2, and “resistance to E. necator” (Ren)2 alleles, conferring resistance to E. necator isolates that were collected from introgression lines containing the Run1 locus, was also recently confirmed (Feechan et al. 2015). Although most E. necator isolates collected from Vitis appear to be equally aggressive on highly susceptible Vitis hosts (Gadoury et al. 2012), differences in aggressiveness among isolates were found in various wild and cultivated Vitis species and may reflect their differential responses to substantial host resistance (Frenkel et al. 2010; Gur et al. 2021).
Traditionally, two genetic groups or biotypes denoted A and B were demonstrated and have been considered within E. necator populations worldwide (Gadoury et al. 2012). Recently, by using multilocus sequencing, microsatellites and amplicon sequencing, Gur et al. (2021) identified a new group IL within E. necator populations in Israel, which was genetically differentiated from any known group in Eastern US and Europe. Groups A and B are genetically distinct, with very little variability within each group, however, the two groups correspond to different epidemiological parameters (Gadoury et al. 2012). Group IL was dominant on wild and traditional vines, and was more aggressive than A and B isolates on both wild and domesticated ones (Gur et al. 2021). Erysiphe necator genome is among the largest sequenced ascomycete genomes (size of ~125 Mb) (Jones et al. 2014). It is also exceptionally repetitive compared to other fungal plant pathogens, and this structure resulted in a broad range of copy numbers variation (CNV) in the demethylase inhibitor (DMI) fungicide target gene CYP51. Jones et al. (2014) concluded that CNV could be adaptive in the development of resistance to fungicides by providing increasing quantitative protection in a gene-dosage dependent manner.
Management of E. necator usually requires a significant use of fungicides, although cultural practices (e.g. pruning and training of the grapevine canopy, selective removal of leaves around the clusters) are important components of an integrated management program and can improve the success and the number of chemical applications. The timing of the fungicide applications may be influenced by the used fungicide, the stage of crop development and the potential for disease infection. There have been several attempts to link spray application timing and fungicide selection to climate and environmental information (Gubler et al. 1999; Calonnec et al. 2008; Carisse et al. 2009; Caffi et al. 2011; Lu et al. 2020). Application of fungicide spray when the first conidia infection occurs is crucial to stop the potential disease outbreak before it can establish itself. This spray timing can also reduce the total number of applications required over a growing season and therefore costs for spraying. Alternating fungicides with different modes of action is essential to prevent E. necator populations from developing resistance. Biological control of PM has met with limited experimental and/or commercial success (Gadoury et al. 2019). Regarding host resistance, all V. vinifera cultivars are susceptible to E. necator, except for the Ren1 and Ren1.2 carriers. All other PM resistances are found in North American Vitis spp., but demand for them is limited outside a few regions and markets (Gadoury et al. 2019). Up to 14 different Ren and Run loci have been identified and described (see Sect. 4.5.2.3).
4.2.1.2 Downy Mildew
The causal agent of grapevine DM is the biotrophic oomycete Plasmopara viticola (Berk. & Curt.) Berlese & de Toni (Gessler et al. 2011). This fungus can attack all green parts of the vine; however leaves, terminal part of the shoots, inflorescences, and young clusters are especially susceptible to infection. Disease symptoms on the upper leaf surface appear as slightly darker and shiny lesions, which rapidly become circular yellowish spots taking an oily appearance sometimes referred to as “oil spots”. Veins often limit leaf lesions and severe infection can result in defoliation (Kassemeyer et al. 2019). Infected shoot tips are thicken, flatten, and curl becoming white due to sporulation and eventually turn brown and necrotic (Gubler et al. 2015). A severe epinasty or “shepherd’s crook” appearance could be shown on shoot tips and rachises which become infected as they are rapidly elongating (Kassemeyer et al. 2019). Young berries are very susceptible turning grayish when infected. Berries become less susceptible as they age but pedicels remain susceptible for some time. Therefore, older berries can become infected if the pathogen infects the pedicels, leading to the so-called “brown rot” (Kassemeyer et al. 2019). Resistance and/or susceptibility of grapevine to DM infection has been evaluated in untreated fields following natural infection based mostly on visual observations focused on both foliage and cluster level (Brown et al. 1999; Cadle-Davidson 2008; Pavloušek 2012; Pacifico et al. 2013; Boso et al. 2005, 2011, 2014; Wan et al. 2007a; Prajongjai et al. 2014).
Plasmopara viticola overwinters as oospores embedded in fallen leaves or other grape tissues infected during the previous season. In spring, oospores germinate and produce macrosporangia, which release zoospores that are dispersed onto grapevine tissues by rain or wind. Secondary disease cycles can take place subsequently under appropriate infection conditions. Numerous clonal cycles may ocurre in one season (Gessler et al. 2011). Population genetics studies indicated that P. viticola populations from most temperate regions contain widespread footprints of recombination, thus suggesting the occurrence of recurrent sexual reproduction (Gobbin et al. 2003; Koopman et al. 2007; Li et al. 2016; Delmas et al. 2017; Zhang et al. 2017; Maddalena et al. 2020). Several studies carried out with isolates collected from a small number of countries and using different markers concluded that North American populations of P. viticola are much more diverse than European populations (Gobbin et al. 2006; Fontaine et al. 2013). Recently, Fontaine et al. (2021) analyzed almost 2,000 P. viticola samples, collected from cultivated and wild grapes in Northeast America, and from the main grape growing regions worldwide. High genetic differentiation occurred in the different regions, with little admixture between them and a low genetic diversity in invaded areas, the European population presenting the highest level of genetic diversity among all invaded populations (Fontaine et al. 2021).
Species boundaries in P. viticola were suggested by Schröder et al. (2011), when assessing the genetic diversity of 14 P. viticola isolates in North America. A cryptic speciation in P. viticola was later confirmed by Rouxel et al. (2013, 2014), with at least five P. viticola cryptic species (also called “formae speciales”), each with a unique degree of pathogenic specialization within the family Vitaceae. Two cryptic species showed a complete host plant specialization toward Parthenocissus quinquefolia and V. riparia, whereas P. viticola species found on V. aestivalis, V. cinerea, V. labrusca, V. vinifera and V. vulpina could infect a broad range of hosts under controlled conditions (Rouxel et al. 2014). Phenotypic variation in aggressiveness has been observed in P. viticola isolates collected from susceptible and partially resistant grapevine cultivars in France, Switzerland and Germany (Delmas et al. 2016). Cross-inoculation experiments demonstrated partial host resistance genotype selection for greater aggressiveness in P. viticola, despite the lack of neutral genetic differentiation among isolates, and the pathogen specificity for a particular grapevine cultivar (Delmas et al. 2016).
The sequenced genome of P. viticola obtained a ~95 Mb assembly size with high level of gene completeness, recovering a large number of genes encoding secreted proteins (Dussert et al. 2019). These included a large proportion of candidate pathogenicity-related genes involved in plant-pathogen interactions. Secretomes were enriched in functions linked to plant cell wall modifications, protease inhibition, reactive oxygen species metabolism, and proteolysis, which are involved in plant defenses (Dussert et al. 2019).
Chemical applications are usually necessary for P. viticola control. Fungicides commonly used for disease prevention are usually fungitoxic at multiple cellular sites, in contrast with penetrating fungicides that act at a single or very limited number of target sites in the P. viticola metabolism. Therefore, the risk of resistance development to specific classes of these latter materials is higher. Decision Support System (DSS) based on epidemiological models have been successfully developed to help viticulturists make informed decisions about fungicide treatments against DM (Caffi et al. 2012, 2013; Cola et al. 2014; Rossi et al. 2014). Cultural practices can also help to limit the potential for fungal infection and disease spread unless environmental conditions are favorable for disease development. Canopy management practices that promote rapid drying of inflorescences, leaves and clusters, site selection and mulches that impede the movement of primary zoospores from the soil to the grapevines are practical recommendations to reduce disease incidence (Kassemeyer et al. 2019). The use of grapevine cultivars showing partial resistance to DM represents an important and sustainable strategy for disease control (Töpfer et al. 2011). Wild grapevine species from Asia and North America, belonging to the Vitis and Muscadinia genera, developed different mechanisms of resistance against P. viticola (Gessler et al. 2011). To date, several quantitative trait loci (QTLs) named “resistance to P. viticola” (Rpv) have been identified conferring resistance to the pathogen (Delrot et al. 2020; Dry et al. 2019; Hausmann et al. 2019) (see Sect. 4.5.2).
4.2.1.3 Esca and Petri Disease
Esca and Petri disease are primarily caused several fungus co-occurring in the plant: Phaeomoniella chlamydospora (W. Gams, Crous, Wingf. & Mugnai) Crous & W. Gams; several species of the genus Phaeoacremonium W. Gams, Crous & M.J. Wingf., being P. minimum (Tul. & C. Tul.) D. Gramaje, L. Mostert & Crous the most prevalent (Gramaje et al. 2015); and several Cadophora spp., being C. luteo-olivacea (J. F. H. Beyma) Harr. & McNew the most prevalent species (Gramaje et al. 2011; Travadon et al. 2015). Esca diseased vines can be further colonized by several basidiomycetous taxa belonging to the genera Inocutis Fiasson & Niemelä, Inonotus P. Karst, Fomitiporella Murrill, Fomitiporia Murrill, Phellinus Quél, and Stereum Hill ex Pers (Cloete et al. 2015; Brown et al. 2020). The most characteristic foliar symptoms of the chronic esca comprise multiple banding discolourations on leaves known as ‘tiger-stripe’ pattern (Mugnai et al. 1999). Internal wood symptoms involve black spots in the xylem vessels, longitudinal brown to black vascular streaking, and white to light yellow soft rot that frequently develops in wood of older vines (Mugnai et al. 1999). Apoplectic esca form is characterized by a sudden and unexpected wilting of the whole vine or one/several arms or shoots (Lecomte et al. 2012). External symptoms of Petri disease include general stunting growth, delayed budbreak, retarded or absent sprouting, shortened internodes, chlorotic and sparse foliage with necrotic margins, leaves or entire shoots, wilting and dieback (Gramaje and Armengol 2011). Internal symptoms of affected vines include the presence of dark-coloured phenolic compounds formed inside xylem vessels of the trunks in response to the fungus growing in and around them, which exude out when cut in cross-sections and dark streaks in longitudinal section (Rooney-Latham et al. 2005). Esca symptoms in the vineyard have been reported with varying incidence between cultivars, rootstocks, and clones based on visual assessment of external symptoms, mainly examining foliar symptomatology (Marchi 2001; Fussler et al. 2008; Bruez et al. 2013; Murolo and Romanazzi 2014; Borgo et al. 2016). This kind of assessment has the limitation that the grapevine trunk diseases (GTD) pathogens often occur in mixed infections within the same vine (Gramaje et al. 2018) and thus there may be some uncertainty that the symptoms observed are due to the effects of a single GTD pathogen.
Esca and Petri disease pathogens are present in vineyards but also in nurseries. In fact, numerous investigations have shown that planting material used in young vineyards is already infected, either systemically from infected mother vines or by contamination during the propagation process (Gramaje and Armengol 2011). In mature vineyards, fruiting bodies (pycnidia or perithecia) containing the spores (conidia or ascorpores) of fungi are primarily developed in dead or infected tissues of spurs, cordons and trunks. Spores are disseminated from fruiting bodies by wind, rain or arthropods until they land on susceptible pruning wounds to germinate and start colonizing xylem vessels and pith parenchyma cells (Gramaje et al. 2018). No matter which GTD fungi are involved, spore release has generally been shown to correlate with rain events and moderate temperatures.
The genetic diversity of Pa. chlamydospora has been well studied in many grape growing areas around the world and by using different molecular techniques: molecular markers or multigene sequence analysis; however, most of these studies have reported a low level of genetic diversity (Comont et al. 2010). Considerable genetic variation, suggestive of ongoing recombination, was found in studies done with Pm. minimum strains collected from grapevines in Australia (Cottral et al. 2001), France (Borie et al. 2002), Italy (Tegli et al. 2000) and Spain (Gramaje et al. 2013; Martín et al. 2014). Contrast analysis among groups defined by molecular marker analyses showed no significant differences in the virulence of Pm. minimum isolates (Gramaje et al. 2013). The genetic study of a collection of C. luteo-olivacea isolates obtained from symptomatic vines in Spain and South Africa identified two highly differentiated genetic clusters in fungal populations with no intermediate genotypes between these clusters (Gramaje et al. 2014). All isolates were able to induce typical Petri disease symptoms in xylem vessels of 110 R rootstock; however, no association was found between virulence phenotype and genetic cluster. The genomes of the esca and Petri disease pathogens Pm. minimum (syn. aleophilum) with 47.5 Mb assembly size (Blanco-Ulate et al. 2013c), and Pa. chlamydospora with ~27 Mb assembly size (Antonielli et al. 2014; Morales-Cruz et al. 2015) have been sequenced in their entirety. In general, a low number of genes putatively coding for plant cell wall-degrading enzymes and secondary metabolite genes were found compared to other wood-degrading fungi.
Presently, no curative measures are known for control of both diseases in nurseries and young vineyards. These diseases would be best managed by an integrated disease management strategy that combines the use of preventive measures, control options throughout all the stages of the propagation process and newly planted nursery vineyards (Gramaje and Armengol 2011). Removing and destroying all diseased wood from the vineyard still remains the best practice to reduce the number of new infections for all GTD pathogens affecting mature plants. Pruning in wet weather should be avoided and conducted during periods when inoculum is less prominent and wound healing is more rapid (Gramaje et al. 2018). Wound protection is the most effective strategy for controlling GTD, and especially if adopted early in the life of the vineyard (Kaplan et al. 2016). Of the fungicides evaluated, based on frequency of reports from literature, thiophanate methyl alone (Rolshausen et al. 2010; Díaz and Latorre 2013) or mixed with myclobutanil (Brown et al. 2021), pyraclostrobin and boscalid alone (Brown et al. 2021) or mixed with a liquid polymer (Martínez-Diz et al. 2021a) are most effective against esca pathogens. Fosetyl-Al applications reduced both expression of esca leaf symptoms and vine mortality under field conditions (Di Marco et al. 2011). Regarding the use of biological control agents (BCAs), most research has been performed under controlled conditions. Biological control agent treatments and natural compounds have shown variable results for preventing infection by GTD under field conditions (Di Marco et al. 2004; Ayres et al. 2017; Cobos et al. 2015; Martínez-Diz et al. 2021a).
There have been reports of varying susceptibility of V. vinifera cultivars and rootstocks to GTD; however, no evidence of qualitative resistance to esca pathogens has been found among several commercial and wild Vitis spp. (Gramaje et al. 2010; Travadon et al. 2013; Murolo and Romanazzi 2014; Sofia et al. 2018; Martínez-Diz et al. 2019; Chacón-Vozmediano et al. 2021). Little is known about the mechanisms of grapevine resistance to esca and Petri disease. Recent reports suggested that V. vinifera susceptibility is positively correlated to xylem vessel diameter for Pa. chlamydospora (Pouzoulet et al. 2017, 2020). This finding warrants further research as such traits may be useful markers when selecting for tolerant or new genotypes.
In nurseries, the application of fungicides to control Petri disease pathogens is difficult since most of them have been phased out from the market (Gramaje and Di Marco 2015). Treating propagation material with hot water at 50 °C or 53 °C for 30 min is the most effective method to disinfect dormant canes during the propagation process (Halleen and Fourie 2016; Eichmeier et al. 2018). However, some anecdotal reports on unacceptably high losses when long duration hot water treatment (50 °C for 30 or 45 min) is applied to commercial batches of cuttings and rootlings have been published (Waite and Morton 2007). Regarding the biological control on esca and Petri disease pathogens, most studies have examined the application of Trichoderma spp. in nurseries. Dipping planting material in T. atroviride strain SC1 during the hydration stages resulted in a decreased incidence of these fungi in Italian (Pertot et al. 2016) and Spanish nurseries (Berbegal et al. 2020). However, BCA treatments have shown variable results for preventing infection by esca and Petri disease pathogens as a pre-planting strategy under field conditions (Martínez-Diz et al. 2021b).
4.2.1.4 Botrytis Bunch Rot
Botrytis cinerea Pers. Fr. is the necrotrophic fungal pathogen that causes Botrytis bunch rot (BBR) of grapevines (Elad et al. 2016). This pathogen can grow on any herbaceous plant tissue, including shoots, flowers and young leaves, developing patches of soft brown tissue, which result in the death of infected parts. On young expanded leaves, the infection can produce areas of brown necrotic tissue (Bettiga and Gubler 2015). Infection of young clusters causes the appearance of brown to black spots on rachises, calyptra and pedicels of the inflorescence. On the fruit, small water-soaked lesions or a symptom called “slip skin” in which the berry epidermis easily slips off are often observed (Wilcox et al. 2019a). Poorly hardened shoots may become infected late in the growing season showing patches of bleaching bark where sclerotia or grayish sporulating mycelia can be seen habitually (Bettiga and Gubler 2015). Regarding the screening of grapevine for resistance in fields upon natural infection, the damage caused by BBR was assessed at harvest in Italian vineyards for 2 years evaluating leaves and bunches separately using visual inspections, and ranked according to a scale (Pacifico et al. 2013). In Australian and New Zealand vineyards, Hill et al. (2014) used grape bunches naturally infected with B. cinerea to investigate the accuracy of visual estimation in comparison with alternative methods, such as NIR and mid-IR spectroscopy, digital image analysis and qPCR. It turned out that digital image analysis was the most suitable and practical alternative to visual estimation, with no specialised equipment required.
Host range of B. cinerea is wide and the potential for an alternative host plant to become an inoculum source is greater. Despite this broad host range, the most consistently available inoculum source within grapevine comes from the crop itself. B. cinerea typically overwinters as mycelia, sclerotia, or chlamydospores in infected grape tissues on the vineyard floor or on the vine (Wilcox et al. 2019a). Fungal infection pathways occur in two periods, from flowering to young cluster development, and after veraison. In the early season, B. cinerea infects inflorescences and young berries, resulting in (i) inflorescence and blossom blight, (ii) latent infections of berries, and (iii) saprophytic colonization of grape bunch trash (Ciliberti et al. 2015). After veraison, latent infections may become visible as rotted berries, and the colonized bunch trash may serve as a source of inoculum for spread inside the grapes. Ripening berries can also be infected through contact with the aerial mycelium produced on adjacent infected berries (berry-to-berry infection) (González-Domínguez et al. 2015).
Recent studies investigating the population structure of B. cinerea in grapevine and other hosts proposed that host plant should be considered as the crucial factor structuring pathogen populations, based on observed associations between patterns of population subdivision and the host of origin of isolates (Walker 2016; Mercier et al. 2019). Cross-pathogenicity experiments with isolates collected from several hosts were consistent with analyses of population subdivision, with patterns of quantitative pathogenicity consistent with specialization of grapevine isolates to grapevine hosts (Mercier et al. 2019). Botrytis cinerea collected from grapevine has a genome of approximately 42 Mb spread across 16 chromosomes and contains a large set of candidates of secreted proteins that are involved in plant tissue penetration and decomposition (Blanco-Ulate et al. 2013a, b).
Management of BBR is best achieved through an integrated approach including sanitation, canopy management, irrigation, reducing berry damage, fungicide or BCA applications and plant resistance (Wilcox et al. 2019a). Removal of clusters left on the vineyard or on vine from the previous season is important to eliminate potential sources of inoculum in the following spring. Control through canopy management is critical and can be obtained by creating a microclimate that is less conductive to fungal development. This alteration of the berry microclimate can be achieved through the choice of training system and the subsequent imposition of shoot positioning, hedging, and leaf removal. The objective is to expose the grape clusters to wind and light so that they dry out faster after a wetting (Bettiga and Gubler 2015). Choosing the right timing type or level of irrigation can also help control BBR. Irrigation should be managed to balance canopy vigour and avoid excessively succulent berries with tightly compacted clusters.
In grapevine regions where economic loss from BBR is important, appropriate cultural practices must be complemented with fungicide applications. Efficacy of chemicals is a function of the application timing, the product efficacy and of good spray coverage. Several fungicide classes have been developed for in-season control of BBR, including the anilinopyrimidines, the dicarboxamides, the hydroxyanilide, and the novel strobilurin and carboxin fungicides. B. cinerea is a pathogen at high risk for fungicide resistance development, so alternating fungicides that have different modes of action is essential to prevent it. A better method would be to apply sprays only when environmental conditions conducive to the fungal growth have been forecasted. In this sense, the mechanistic model developed by González-Domínguez et al. (2015) can predict the risk of B. cinerea development and the severity at harvest. The model has been validated (Fedele et al. 2020) and is currently integrated in a DSS (Caffi et al. 2017) to help growers schedule fungicide treatments.
Applications of BCA products based on antagonistic microorganisms (e.g. species of the fungal genera Trichoderma, Aureobasidium and Ulocladium, as well as fermentation products of Bacillus spp. bacteria) have provided a measure of control in some regions but have been ineffective in others, particularly when disease pressure was high (Bettiga and Gubler 2015). Moderate resistance or tolerance to BBR has been identified in closely related Vitis spp., including V. lincecumii, V. labrusca and V. rotundifolia, but they usually have poor fruit quality or flavors that are not commercially desirable. In particular, researchers evaluated indirect traits (see Sect. 4.6.2) such as physical, morphological and chemical components contributing to BBR in Vitis spp. For example, aromatic volatiles produced by V. labrusca reduced pathogenicity and Botrytis spore production (Naegele 2018). These species are important sources of resistance to several biotic and abiotic stresses, but lack desirable fruit quality characteristics found in V. vinifera.
Studies have evaluated chemical, morphological, physical, and genetic components contributing to B. cinerea resistance in Vitis spp. (Deytieux-Belleau et al. 2009; Herzog et al. 2015; Trotel-Aziz et al. 2006). The wax content and cuticle, as well as the number and thickness of epi- and hypodermal cell layers have been weakly positively correlated with resistance (Gabler et al. 2003; Deytieux-Belleau et al. 2009; Herzog et al. 2015; Rossmann et al. 2020). Compact clusters favours microclimates for diseases spread and infection and have more crackings or physical damage as berries expand against each other (Bettiga and Gubler 2015). Other studies suggested that aromatic volatiles produced by V. labrusca accessions were able to reduce spore production and pathogenicity of B. cinerea (Kulakiotu et al. 2004).
4.2.1.5 Anthracnose
Grapevine anthracnose is caused by the hemibiotrophic fungus Elsinoë ampelina Shear (Mirica 1988). The pathogen attacks mostly young aerial green tissues and succulent parts of the vine throughout the entire crop cycle, including leaves, tendrils, shoots, fruit stems, petioles, and clusters; however, lesions are prevalent and distinctive on shoots and berries (Thind 2019). Disease symptoms on shoots, petioles and tendrils appear as isolated, small, reddish, circular spots that enlarge to become brownish, sunken lesions with grayish centers and dark margins (Thind 2019). On leaves, the fungus causes numerous small, circular brown spots, which turn gray in the center with black round or angular margins, and as the lesions mature, the necrotic center often drops out, producing a “shot-hole” appearance (Carisse and Lefebvre 2011a). Infected clusters and berries initially show small, reddish-brown, circular spots which may become slightly sunken and whose centers turn whitish gray with black margins, sometimes resembling a bird’s eye (Pirrello et al. 2019). Screening of grapevines for anthracnose tolerance in fields upon natural infection have been assessed based on visual inspections of leaves and clusters (Wang et al. 1998; Yun et al. 2006; Li et al. 2008; Louime et al. 2011; Poolsawat et al. 2012). However, due to the perennial nature of grapevine, this approach is time-consuming, laborious and expensive and under field conditions. Anthracnose is highly influenced by the climatic conditions, requiring some years of evaluation to produce robust data (Pirrello et al. 2019).A recent study reports about a disease test system of artificially inoculated vines in the greenhouse (Modesto et al. 2020).
Elsinöe ampelina overwinters as sclerotia or mycelium in cane lesions formed during the previous growing season and in fallen mummified berries (Carisse and Lefebvre 2011b). Conidia are then released from these structures in spring and splashed by rain causing primary infections. Spores infect new tendrils, shoots, leaves and young berries, producing lesions with acervuli from which conidia are released during periods of humid conditions, thus serving as secondary inoculum for disease spread during the growing season (Carisse and Morissette-Thomas 2013). High genetic variability was observed among E. ampelina isolates collected from different regions in Thailand by random amplified polymorphic DNA (RAPD) analysis (Poolsawat et al. 2010). Recently, multilocus DNA analyses of E. ampelina populations collected from grapevine in Australia and Brazil (Santos et al. 2018a), and in Brazil (Santos et al. 2018b), resulted in high genetic diversity with the identification of four and five genetic haplotypes, respectively. High degree of variability in pathogen virulence to grapevine was observed in all studies (Poolsawat et al. 2010; Santos et al. 2018a, b). Elsinöe ampelina collected from grapevine has a genome of approximately 28 Mb and contain a large set of candidate secreted proteins as coding for CAZymes and 20 secondary metabolite clusters that may contain genes involved in secondary metabolite biosynthesis and pathogenesis (Li et al. 2020).
Sanitation measures to reduce inoculum sources of E. ampelina in the vineyard are very influential and important. Diseased pruning debris should be removed from the vineyard or destroyed. Mulching can also be used to cover infected berries on the vineyard floor (Thind 2019). Fungicide applications are recommended when the disease pressure is high in established vineyards. In general, several fungicides used in the management program against other diseases are also effective against anthracnose. However, the pathogen can develop resistance following repeated use due to their specificity in mode of action. Grapevine cultivars vary considerably in their susceptibility to anthracnose. In general, V. vinifera species are highly susceptible (Hopkins and Harris 2000; Yun et al. 2006), and table grape cultivars are more susceptible to anthracnose than wine grape cultivars (Hart et al. 1993; Kono et al. 2013). Several cold-tender hybrid cultivars derived from V. riparia are also highly susceptible to the disease (Carisse and Lefebvre 2011a).
4.2.1.6 Black Rot
Phyllosticta ampelicida (Engelm.) Aa (syn. Guignardia bidwellii, following the recommendation of the International Commission on the Taxonomy of Fungi, Rossman et al. 2015) is the causal agent of BR of grapevine. All herbaceous tissues of the vine are susceptible to infection by the pathogen, including leaves, shoots, tendrils, petioles and berries, with young leaves and fruit being extremely sensitive (Pirrello et al. 2019). Leaf lesions are initially small circular to irregularly cream-coloured dots delineated by narrow dark-brown margins that evolve into reddish brown lesions as they mature. Pycnidia develop as small black pimples within the leaf lesions often forming a ring (Wilcox et al. 2019b). The first symptom on berries is a small whitish dot surrounded by a chocolate-brown necrotic tissue. This necrosis rapidly expands, and the berry becomes rotted and shrivelled into a blue-black mummy that remains attached to the pedicel covered with pycnidia (Onesti et al. 2016). Shoot infections cause elongated black necrosis often containing abundant pycnidia. On the petioles and pedicels, lesions appear as small, darkened depressions, which turn black quickly. The tolerance of grapevine varieties to BR have been tested under field conditions via natural infection in Romania by the analysis of the severity symptoms based on observations of frequency and intensity of the attack of P. ampelicida fungus according to a scale (Tomoiaga and Chedea 2020). In Italian vineyards, the damage caused by BR was also visually assessed at harvest for two years evaluated leaves and bunches separately and ranked according to a scale (Pacifico et al. 2013) (for host genetic resistance studies see Sect. 4.5.2.1).
Phyllosticta ampelicida overwinters in mummified berries on vines and on the soil surface, and in cane lesions as pycnidia (Ramsdell and Milholland 1988). During the following grape growing season, the pathogen produces both conidia and ascospores on the inoculum sources (Hoffman et al. 2004). These spore types are repeatedly dispersed and cause primary infections in all green tissues (Ferrin 1977). Conidia developed from pycnicia in light-brown necrotic lesions on green tissues are dispersed by wind or droplets and cause secondary infections (Ferrin 1977). Black rot is a polycyclic disease with repeated cycles of primary and secondary infections. Three formae speciales (f. sp.) of “G. bidwellii” have been described (Luttrell 1946, 1948): (i) “G. bidwellii” f. sp. euvitis is pathogenic to V. vinifera and to the American bunch grape species of the section Vitis, (ii) “G. bidwellii” f. sp. muscadinii is pathogenic to V. rotundifolia and V. vinifera, and (iii) “G. bidwellii” f. sp. parthenocissi is pathogenic to Parthenocissus spp. Zhang et al. (2013) identified different species in the P. ampelicida complex, thus proposing P. parthenocissi as a distinct species based on morphological and phylogenetic data. The high degree of genetic variability of P. ampelicida was later confirmed by using microsatellite markers in broad collections of isolates collected worldwide (Narduzzi-Wicht et al. 2014; Rinaldi et al. 2017). The genome sequence of P. ampelicida is not available yet.
Black rot is best managed by integrating chemical applications and cultural practices. Removal of mummies from the canopy to the ground during winter pruning is a primary sanitation practice. In vineyards where the inoculum levels are high, additional cultural practices such as mulching the vineyard floor to cover mummies or physical removal of mummies from the vineyard are recommended. Diseased pruning debris should be removed from the vineyard or destroyed (Wilcox and Hoffman 2019). Several fungicide classes such as quinone outside inhibitors (QoIs), demethylation inhibitors (DMIs) and dithiocarbamates provide good control of BR (Molitor and Beyer 2014). In organic production systems, copper and sulfur are only moderately effective compared to the synthetic active compounds (Wilcox and Hoffman 2019).
4.2.1.7 Phomopsis Cane and Leaf Spot and Phomopsis Dieback
The causal agents of Phomopsis cane and leaf spot (PCLS) and Phomopsis dieback are up to 18 Diaporthe species, D. ampelina being the most common species (van Niekerk et al. 2005; Baumgartner et al. 2013; Úrbez-Torres et al. 2013; Guarnaccia et al. 2018). Phomopsis dieback is characterized by V- or irregular-shaped wood cankers in canes, spurs, cordons and/or trunk as well as general canopy symptoms of shoot dieback and dead spurs. During the development of PCLS, D. ampelina directly attacks all green tissues of the vine, causing necrotic lesions on the leaves, green stems, and fruit (Baumgartner et al. 2013). To date, no comparative studies are available about disease measurement on grapevine cultivars upon natural infections.
The ascogenous stage of Diaporthe spp. is rarely encountered in nature. Diaporthe species have septate mycelium, and reproduction occurs asexually by means of conidia produced in pycnidia. Two types of conidia are produced: fusiform 1-celled alpha conidia and filiform 1-celled beta conidia. The primary inoculum consists of alpha conidia that are responsible for infection of shoots and leaves primarily in spring, during cool and wet weather (Anco et al. 2013; Erincik et al. 2003). The role of beta conidia in the epidemiology of PCLS is still unclear, but their capacity to germinate and infect grapevine tissue has been demonstrated by artificial inoculation (Sergeeva et al. 2003). Both alpha and beta conidia are assumed to be dispersed by rain, although research on this type of dispersal is limited (Wilcox and Hoffman 2019).
Known hosts of the grapevine specialist D. ampelina include the Eurasian grapevine V. vinifera, the North American grapevine V. rupestris, and V. aestivalis, V. lambrusca and V. rotundifolia (Hewitt and Pearson 1988; Uecker 1988) (for host genetic resistance studies see Sect. 4.5.2.1). Other plants such as Ampelopsis quinquefolia, Hydrangea macrophylla and Olea europaea (Hewitt and Pearson 1988; Uecker 1988; Santos et al. 2010; Úrbez-Torres et al. 2013) can also be hosts of D. ampelina, inoculum may therefore originate from source other than grapevine. Research also demonstrated that Diaporthe spp. isolated from wood cankers of fruit and nut crops in Northern California are pathogenic on grapevines (Lawrence et al. 2015). Even though Diaporthe spp. are characterized under PCLS and Diaporthe dieback diseases, aggressiveness varies according to the species. Diaporthe ampelina has a long history as the most common and virulent species together with D. amygdali (Mostert et al. 2001; Van Niekerk et al. 2005). Lesuthu et al. (2019) reported D. ampelina, D. novem and D. nebulae as the most virulent species of Diaporthe associated with grapevines in South Africa. Diaporthe eres was found to be a weak to moderate pathogen in several studies carried out in Croatia (Kaliterna et al. 2012) and US (Baumgartner et al. 2013). Pathogenicity tests with 9 species collected in a broad survey of trunk diseases in Europe revealed D. bacchae, D. celeris, D. hispaniae and D. hungariae as pathogens of grapevine, while D. bohemiae did not induce necrotic lesions on the inoculated grapevine shoots (Guarnaccia et al. 2018). In China, D. gulyae was the most aggressive taxon, whereas D. hubeiensis was the least aggressive among eight Diaporthe spp. (Manawasinghe et al. 2019).
Few population genetic studies have been performed with Diaporthe spp. The reproductive biology of Diaporthe on grapevines is important when examining the genetic diversity within populations of each Diaporthe taxa. Genetic clones and microsatellite sequences used as probes, as well as mycelial incompatibility tests, indicated high genetic diversity in natural populations of four Diaporthe taxa affecting grapevine in Australia (Scheper 2001). High genetic nucleotide and haplotype diversity was observed for D. eres in Chinese vineyards (Manawasinghe et al. 2019). Haplotype networks including Chinese and European isolates suggested a close relationship between the two populations and evidence for recombination (Manawasinghe et al. 2019). The draft genome sequence of D. ampelina isolated from grapevine has been recently released with ~47.4 Mb assembly size (Morales-Cruz et al. 2015). The analysis of functional annotations of their predicted protein-coding genes indicated a complex repertoire of potential virulence functions and the identification of many genes associated with nutrient uptake, toxin production, and lignocellulose degradation (Morales-Cruz et al. 2015).
Management strategies to control PCLS still rely on cultural practices and fungicide application (Wilcox and Hoffman 2019). The introduction of D. ampelina into the vineyard should be avoided by using pathogen-free planting material from nurseries. Once the disease is present, dead wood and pruning debris should be destroyed preferably by burning. Broad-spectrum protectant fungicides such as captan, chlorothalonil, dithianon, folpet, mancozeb and ziram are very effective against PCLS and are frequently used for its control throughout the world. Other fungicide groups, such as QoI and DMI materials, have provided disease control in some grape regions but not in others. Proper timing of fungicide sprays is crucial for PCLS control in vineyards. Applications should be focused during the early shoot growth stages (Wilcox and Hoffman 2019). Contact fungicides are usually recommended before cool, wet weather to prevent infection from conidia dispersed by rain splashes, and additional applications are recommended if wet conditions continue in order to protect new growth (Wilcox and Hoffman 2019). A simple warning system based on the prediction of PCLS infection periods was developed to better time fungicide application (Nita et al. 2006). However, Anco et al. (2012) demonstrated that fungal inoculum is not always present in sufficient amounts in the vineyard and a temporal pattern exists in the production of inoculum by pycnidia. In this sense, González-Dominguez et al. (2021) recently studied the dynamics of conidial production during the season and the dispersal of this primary inoculum in vineyards in Italy and Montenegro in order to incorporate this information into an improved warning system for a proper fungicide timing. Management of Phomopsis dieback should be focused on pruning wound protection. Spray applications of thiophanate methyl mixed with myclobutanil, or pyraclostrobin and boscalid minimized D. ampelina infection in California table grapes (Brown et al. 2021).
4.2.2 Caused by Bacteria and Phytoplasmas
Various bacteria and numerous viruses are detectable in grapevine phloem and/or xylems. Pierce’s disease, a threatening disease of grapevine especially in the Americas, is caused by the bacterium X. fastidiosa subsp. fastidiosa, listed as a quarantine pathogen worldwide. Other bacteria are associated with diseases currently considered less severe, such as Allorizhobium vitis and Agrobacterium tumefaciens, causing crown gall in grapevines all over the world. Xylophilus ampelinus, also listed among the quarantine organisms in many countries, causes grapevine bacterial blight, which shows an apparently limited distribution in the world, probably due to the fact that it is often latent. Moreover, several species of phytoplasmas, the smallest known plant bacteria, have been reported in grapevines. However, only a few of them are associated with very serious diseases; among those FD, a European grapevine yellow, is the most destructive and is listed as a quarantine pathogen in all grape growing countries. Among bacteria, only the most common or most damaging are described (causing PD and crown gall), as well as the phytoplasmas associated only with the European grapevine yellows (FD and Bois noir, BN).
4.2.2.1 Pierce’s Disease
Pierce’s disease, caused by X. fastidiosa, is one of the most devastating bacterial diseases of grapevine. The bacterium grows in the xylem of the plant, where it actively multiplies and forms a biofilm. The leaves dry up starting from the margins in spring–summer, can show yellowing, and then fall down. Grapes wither before harvest, and canes do not mature or mature unevenly. Plants can die in 1–5 years. The disease is spread especially in North America, but recently was identified also in Europe, in the Canarian islands (Moralejo et al. 2019). It is transmitted by insects that feed on grapevine xylem sap, in particular several sharpshooters, leafhoppers and splittebugs. The pathogen occurs in many herbaceous and woody plants, cultivated and wild. Some of those host plants do not show any symptoms, and some others are hosts of the vectors. Currently, four subspecies of X. fastidiosa have been identified so far, and other two are proposed, but only one of them is present in grapevine, X. fastidiosa subsp. fastidiosa. Most of the strains are aggressive and cause severe damage to grapevine, however asymptomatic and weakly virulent strains have been identified in nature (Hopkins 2005). Strong population genetic structure was found. A high level of genetic diversity was detected among the bacterium populations isolated from grapevine in different viticulture locations using variable markers, and interestingly the diversity was linked to the geographic areas and not to the variety (Lin et al. 2013). The genome of X. fastidiosa subsp. fastidiosa from orange was sequenced in 2000, and showed a 2,679,305 bp circular chromosome and two plasmids of 51,158 bp and 1,285 bp (Simpson et al. 2000). Later on, another two genomes, originated from almond and from grapevine, were sequenced and showed to be very similar. Due to its complex epidemiology, the field control of the disease is not easy, and include agronomic strategies, such as control of insect vectors, removal of host plants and roguing of infected grapevines. Hot water treatment is used to control PD disease in nurseries and to prevent spread of the bacterium through grapevine multiplication material. Due to the difficulty in containing the disease, a number of other control strategies, ranging from biological agents to transgenic approaches, have been carried out in the last decades. Bacterial and fungal biological control agents have been identified. They can help in limiting PD damages in grapevines, such as Paraburkholderia phytofirmans (Baccari et al. 2019), but most of them were not able to provide long-term protection. Bacterial phage therapy looks to be a promising strategy (Das et al. 2015). Many progresses have been made in the individuation of resistance sources, occurring only outside of the V. vinifera species. A first resistance locus, PdR1, was discovered in V. arizonica/candicans on chromosome 14; a second one, PdR2, on chromosome 8 in V. arizonica/girdiana, and work is in progress on other Vitis species that showed resistance (Walker and Tenscher 2017; Lin 2017) (see Sect. 4.5.2.1). Five new varieties resistant to PD, obtained from V. arizonica after four to five traditional crossing with V. vinifera, were recently released in America (see Sect. 4.10.3). Several trials have been carried out to obtain transgenic resistant grapevines transformed with different strategies (reviewed in Kyrkou et al. 2018), however studies and field experiments are still ongoing to ascertain the long-term and effectiveness of the protection against the disease in the field (see also Sect. 4.12.2).
4.2.2.2 Crown Gall
Crown gall in grapevine is generally caused by the species Allorizhobium vitis (formerly named “Agrobacterium vitis”) (Ormeño-Orrillo et al. 2015), but can also be associated with Agrobacterium tumefaciens. These gram-negative bacteria are generally latent in grapevine for long time; uncontrolled proliferation of tissues, leading to the formation of galls and tumours, is triggered by mechanical damages in the wood tissues, due to freezing, to very hot climatic conditions, or to any other injury that can cause cracking of the tissues, included pruning. The disease is widespread in all the grape growing areas around the world. The tumours are located on the graft union or on trunk wounds, though can be also present in the canes. They can reduce or block the water and nutrient uptake, especially in young grapevines, causing plant decline. The bacterium inhabits the xylem of the plant, where it forms a biofilm. Virulent strains of All. vitis and A. tumefaciens are able to release to the plant cells a transfer-DNA (T-DNA), that finally leads to the uncontrolled production of the galls. Moreover, these bacteria can survive in the soil or in dead plant tissues for a long time (Bini et al. 2008).The genome of All. vitis is more than 5 Mb in length and includes two circular chromosomes and a variable number of plasmids (Slater et al. 2009). Virulent strains contain a tumour-inducing plasmid (Ti), able to deliver the T-DNA, which is not present in avirulent strains. High genetic diversity among strains has been found and revealed four genetic clusters, leading to consider All. vitis a species complex (Kuzmanovic et al. 2020). Management of crown gall is very hard and is mainly carried out by cultural control. Prevention is the first strategy, planting All. vitis-free grapevines, not replanting immediately after removal of an old symptomatic vineyard, and avoiding locations with frequent freezing. When removing an infected plant, it is helpful to eliminate the roots as much as possible. Hot water treatment of multiplication vine material can reduce the pathogen, but not eliminate it. The application of rameic compounds in the wounds and after the removal of the galls can have a bacteriostatic effect, but it is not a definitive solution. Application of bacterial biocontrol agents has been assayed in many studies with some success (Bazzi et al. 1999; Asghari et al. 2020), but no commercial effective product is available yet. Natural resistance occurs in different Vitis species, such as V. amurensis and V. labrusca, but not in V. vinifera. Resistance traits from V. amurensis were introgressed into V. vinifera and showed to be controlled by a single quantitative trait locus (QTL) (Kuczmong et al. 2012) (see Sect. 4.5.2.2). Transgenic silencing approaches using genes in the Ti-plasmid also proved to be promising in achieving crown gall resistance (Galambos et al. 2013).
4.2.2.3 Grapevine Yellows
Grapevine yellows (GYs) are diseases occurring worldwide. Leaves affected by GYs are crispy, brittle, downwards rolling and show reddening in red varieties and yellowing in white varieties. Discoloration occurs always also in the main veins. Flowers and bunches wither, shrivel and fall down. Canes exhibit short internodes, necrosis of terminal buds and sometimes black/brown pustules on the basis; moreover, they appear rubbery and weeping and do not lignify in autumn, or only partially. GYs are associated with phytoplasmas, which are wall-less gram-positive bacteria living in the phloem sieve tubes. Phytoplasmas belong to the genus Candidatus Phytoplasma (Ca. Phytoplasma), in the class Mollicutes. They cannot be maintained and propagate in laboratory culture media, thus are commonly classified in phylogenetic groups based on their 16S ribosomal nucleotide sequence (Firrao et al. 2004). Only some groups occur in grapevine, and the different ‘Ca. Phytoplasma’ species are generally typical of each continent. Though the symptoms are identical all over the world, not all the GYs show the same epidemiology, which is strictly depending on the specific insect vector. The most serious GYs occur in Europe, where two main diseases are present: BN, spread in all viticultural regions in Europe and the Mediterranean basin, and FD, a quarantine pest in the European Community, occurring in Central and Southern Europe. Symptoms of GYs were discovered in South Africa only a decade ago, and the disease is luckily confined to a few grape growing areas. In Australia and New Zealand several GY phytoplasmas are present in grapevines, while in the Americas and Asia GYs are only sporadically reported. In general, phytoplasmas infecting grapevines are present also in other plants, wild or cultivated, thus they are not strictly specialized pathogens regarding the plant host. The ecological cycle of ‘Ca. Phytoplasma solani’, the agent of BN in Europe, includes a few main wild plant hosts (among them Convolvolus arvensis, Urtica dioica, Cirsius spp.), which constitute a wild reservoir of the pathogen and also a host for the insect vectors, which are Hyalesthes obsoletus, Reptalus panzeri and R. quinquecostatus. Phytoplasmas causing FD, transmitted by the American leafhopper S. titanus, were supposed to be present only in grapevines at the beginning of the century; however, they occur at least in black alder and clematis, which are now thought to be the ancestral hosts in Europe. Specialization of phytoplasma colonization in plants is indeed determined by the vectors, which have unique and very specialized relationships with the pathogen. The genome of phytoplasmas is the smallest among the bacteria. Only a few phytoplasma genomes were fully sequenced, among them ‘Ca. Phytoplasma solani’. It includes a unique circular chromosome of about 1,000 kbp (Šeruga Musić et al. 2019), however the genome length can vary between different strains even in the same species. The genome of FD phytoplasma was not yet completely sequenced and it was estimated to be approximately 650 kbp in strain FD92 (Carle et al. 2011). Gene duplication and redundancy are very common, but phytoplasmas lack genes coding for many important metabolic pathways, such as ATP synthase genes (Namba 2019), according to their high level of specialization and parasitism. Management of GYs includes only preventative strategies that mainly rely on management of insect vectors by chemical treatments (see Sect. 4.3.1.4). Besides that, common and suggested strategies are uprooting of diseased plants and planting of healthy vine material. Specific field strategies can be settled down according to the ecology of the respective disease. Now, neither biological control agents nor resistance traits have been identified. However, multiyear field observations revealed that most American rootstocks and very few European varieties possess resistance or tolerance to GYs or to their vectors. Molecular studies and breeding activities to identify the genetic traits associated with resistance and susceptibility to GYs are ongoing (Bertazzon et al. 2019; Jarausch et al. 2013; Jollard et al. 2019).
4.2.3 Caused by Viruses
Grapevine can host more than 70 different viruses (Martelli 2017). Most of them fortunately do not cause evident damages to the plant or the production. However, a few diseases associated with viruses are very common, can cause important losses in vine production and therefore have been the object of specific legislation to avoid their spreading through world trade. Fanleaf and leafroll complexes are doubtless the most important viral diseases in all grape growing areas. Other minor viruses are listed for sanitary purposes in clonal selections or in other national sanitary protocols, such as those associated with rugose wood (Grapevine Virus A, GVA; Grapevine Virus B, GVB; Grapevine Rupestris Stem Pitting associated Virus, GRSPaV), or the newly discovered Grapevine Red Blotch Virus (GRBV) that emerged in the USA in the last few years, and Grapevine Pinot gris Virus (GPGV), which nowadays seems to be widespread in Europe and threatens all other viticultural countries in the world. Concerning viral diseases, only the most damaging and widespread (fanleaf and leafroll complex) are reported in detail.
4.2.3.1 Fanleaf
Fanleaf or infectious degeneration complex is the most destructive viral disease. Symptoms are very variable, but generally include typical deformations of leaves and canes and leaf yellowing. Leaves are asymmetrical with acute denticulation, may show enlarged petiolar sinuses and chlorotic mottles or bright yellow discoloration of veins. Canes show abnormal branching, short internodes, double nodes and fasciations. Moreover, the infected plants show dropping off flowers and berries, which are smaller and ripen irregularly. In more severe cases, grapevine plants can slowly decline. The etiological agents belong to Nepoviruses, and the most known is Grapevine Fanleaf Virus (GFLV). All Nepoviruses are transmitted specifically by nematodes living in the soil and feeding on roots, X. index in the case of GFLV. Grapevine Fanleaf Virus is generally restricted to grapevine, though a few weeds have been found to host it, but not having any epidemiological relevance. Also, the transmission by nematode species was demonstrated to be species-specific. While GFLV is spread almost all over the world, due to human trade, the other grapevine Nepoviruses and their vectors are confined only to the Old World or to North America (Martelli and Taylor 1990). Several Nepoviruses were completely sequenced, including GFLV. They are constituted by an isometric particle formed by a bipartite single-strand positive-sense RNA genome. Both RNAs have poly(A) tale at the 3’ end and a capped 5’ end. Both RNA1 and RNA2 have single open reading frame (ORF) coding for a single polypeptide that is subsequently cleaved in separate proteins. A satellite RNA is present in some GFLV isolates (Schmitt-Keichinger et al. 2017). Genetic variability of GFLV is wide and consistent with the notion of quasi species, and varies between different ORFs in the genome (Elbeaino et al. 2014). Management of fanleaf, such as for all grapevine viruses, relies on prevention of the infection, and is based on planting of healthy and certificated grapevine plants in nematode-free soils. Once a plant or a vineyard is infected, removal of infected grapevines is recommended, and crop rotation in order to sanitize the soil. Indeed, due to the specificity of the plant-pathogen-vector system, planting crops different from grapevine for a few years is the most effective and sustainable control strategy (see for more details on nematode control Sect. 4.2.4). It seems that there is no useful source of resistance to GFLV in Vitis species. However, some American species showed resistance to X. index. In M. rotundifolia the resistance is against X. index feeding and has been used in hybrids and crossing (Oliver and Fuchs 2011).
4.2.3.2 Leafroll
Leafroll complex disease is spread all over the world, and its typical symptom is rolling of the leaves, which become thick and brittle in summer. Moreover, leaves of infected plants show reddening in red varieties or yellowing in white varieties, though usually the primary and secondary veins remain green. Bunches from infected plants ripen later and can be smaller in number and size. The intensity of the leaf symptoms depends on the virus type and viral strain. Leafroll viruses are classified according to their genetic features in the family Closteroviridae. They are named as Grapevine Leafroll associated Virus (GLRaV), followed by a number. Most GLRaVs belong to the Ampelovirus genus (GLRaV-1, GLRaV-3, GLRaV-4), GLRaV-2 is a Closterovirus, and GLRaV-7 belongs to the Velavirus genus. Grapevine Leafroll associated Virus-2 is the only GLRaV that can be also associated with graft incompatibility, besides typical leafroll symptoms, and the symptomatology is linked to the viral strain (Bertazzon et al. 2010). GLRaV-7 is asymptomatic, whereas GLRaV-3, the most common GLRaV, causes the most typical leafroll symptoms. All the GLRaVs have been identified only in grapevines so far. They are phloem limited and non-mechanically transmitted. Ampeloviruses are transmitted by a plethora of mealybugs and scale insects. Among them, the most important is Planococcus ficus, but at least other 10 mealybugs and scale insects were demonstrated to be able to transmit one or another GLRaVs. On the contrary, the vectors of the leafroll viruses classified in the other two genera (GLRaV-2 and GLRaV-7) have not yet been discovered. The intra- and interspecific molecular diversity of GLRaVs is well known. For example, GLRaV-4, 5, 6 and 9 are now grouped under GLRaV-4, but they were considered different viruses until a few years ago, when a complete revision of GLRaVs’ phylogeny was carried out (Martelli et al. 2012). It is interesting to note that in general the genetic diversity is not linked to the geographic area of study, due to the wide grapevine trade in the past that allowed these viruses to circulate everywhere in the world grape growing countries. Leafroll is indeed one of the most widespread and common viral diseases of grapevine. The genome of Closteroviridae is constituted by a unique RNA, positive-sense single-strand filament, spanning from 13,700 to 18,500 nucleotides and including 6–12 typical ORFs (Martelli 2014). In the Ampelovirus genus, GLRaV-1 and GLRaV-3 have the larger genome (9–12 ORFs), while GLRaV-4 shows 7 ORFs. GLRaV-2 contains 9 ORFs, GLRaV-7 is formed by 8–10 ORFs, depending on the isolate.
As in the case of other grapevine viruses, the main control strategy is prevention, achieved by certification and sanitary selection regulative schemes, which aim to ensure the absence of dangerous viruses in the multiplication trade material. All grapevine species and varieties can be infected by GLRaVs, but not all of them show the symptoms at the same extent, especially in the leaves: indeed, evidence of leafroll in white varieties is difficult to identify. However, the damages to the grape and wine production occur in all varieties, thus no natural resistance or tolerance has been detected in grapevine so far. In the field, the only possible control strategy is targeted to the vector, and involves monitoring, chemical treatments, mating disruption (MD), and biological control by using parasitoids and predators (see Sect. 4.3.1.3).
4.2.4 Nematode Parasites
Two major plant-parasitic nematode (PPN) species can seriously damage grapevines, root-knot (Meloidogyne spp.) and dagger (Xiphinema index) nematodes. Other have been recognised to cause significant damage to grapevines under more specific conditions, for instance root-lesion (Pratylenchus spp.), citrus (Tylenchulus semipenetrans) and ring (Criconemoides xenoplax) nematodes. However, many PPNs could be detected in the rhizosphere feeding on grapes or alternative hosts as weeds (Téliz et al. 2007). Plants affected by PPNs show an unspecific symptomatology, in some cases confused with other problems such as root asphyxia or nutritional deficiencies, such as yellowing, poor growth, early ripening of grapes and stunting. Usually these affected plants form patches in the field that may follow or not the vine rows. Nematode damage is related to their population density, but differences in climate, soil characteristics and grape cultivar/rootstock could change its susceptibility (Nicol et al. 1999). Nematodes damage plants by direct feeding on the roots or, in the case of some species, by vectoring viruses (e.g. X. index as vector of GFLV) (Brown et al. 1993). In this sense, a nematological soil and root analysis is needed in order to detect and quantify them properly.
Meloidogyne spp. are sedentary endoparasites, establishing a permanent feeding site by inducing the formation of a root gall and giant cells inside the gall. In additionally to parasitism, the galling reduces the uptake of nutrients and water of the root that affects the growth during stressing periods of the year. The most important species are the tropical M. arenaria, M. incognita, M. javanica and the temperate M. hapla (Nicol et al. 1999). Other species (e.g. M. ethiopica, M. nataliei, M. hispanica) have been found to affect grapevines in some regions (Bird et al. 1994; Carneiro et al. 2004; Castillo et al. 2009). Recently, a new species, M. vitis, has been described as severely affecting grapevines in China (Yang et al. 2021). The genome of the most important species has been sequenced showing that the structure and function of their genomes are linked to differences in reproduction (Blanc-Mathieu et al. 2017; Szitenberg et al. 2017).
Xiphinema index has a worldwide distribution. This nematode feeds ectoparasitically on root tips (Nicol et al. 1999) resulting in retarded root extension, swelling and the formation of root tip galls. However, this kind of gall is usually more terminal compared to that produced by Meloidogyne spp. Xiphinema index is the vector of GFLV. Other species from Longidoridae could be virus vectors for grapevine (Brown et al. 1993). This nematode may persist in the soil for long periods (up to 10 years) feeding on root fragments or up to 4 years if the host is absent (Raski et al. 1965; Demangeat et al. 2005). Xiphinema index also remains viruliferous for up to 9 months in moist soil in the absence of host plants (Taylor and Raski 1964), and the depth where they are frequently found (40–110 cm) hinders field sampling and management (Villate et al. 2012). The phylogeography of this nematode has been studied using mitochondrial and microsatellite markers in samples from most regions of its worldwide distribution and the highest polymorphism level was found in the samples of the Middle and Near East, strongly suggesting that this region contained the native area of the nematode (Nguyen et al. 2019). The diversity of this nematodes appears to follow the routes of the domesticated grapevine during Antiquity, mainly dispersed by the Greeks and the Romans (Nguyen et al. 2019).
The best strategy for regulation of nematodes is to avoid the entry of these pathogens in the vineyard with soil and plant material. Once established in the field, they are very difficult to eradicate and measures can only be taken to reduce the quantity, damage and expansion in the field. Usually, replanting is the most vulnerable time for nematode infection and damage, when plantlets are smaller and more sensible to stresses. During this period, plants are young and the amount of inoculum may be high if the previous crop was also grapevine. For this reason, a reduction of the initial plant-parasitic nematode level is necessary after a nematological soil analysis. One option to achieve this goal could be crop rotations with non-host or weak hosts, depending on the target nematode species present in the soil. Cereals or corn could be an interesting option in grapevine for the reduction of Meloidogyne spp., while for X. index this crop rotation is less useful because it can survive for a long time without appropriate hosts. Biofumigation may be an alternative control method that can also increase soil biodiversity, while solarisation is not suitable for the control of X. index because they move to deeper horizons (Bello et al. 2004). Cover crops are currently an active field of research, mainly the use of species from the Brassicaceae family. In some cases, there are promising results, but they are highly dependent on the environmental conditions and their competence with the crop (Baginsky et al. 2013; Kruger et al. 2015). Weeds must be properly managed during fallow or crop rotations in the case of root-knot nematodes because many of them could be reservoirs and could increase the initial soil inoculum in some cases (Castillo et al. 2008). In certain countries, some nematicides are available for application before planting or during the season, so farmers need to check if the possibility of using this tool is allowed in their country. However, nematicides have a risk to the environment and in some cases are not fully effective because some nematodes (X. index) could reach important depths in soil. Additionally, in some countries the small returning profit in grapevine made this strategy unaffordable. Plant-parasitic nematodes have a high number of potential biocontrol agents that affect nematodes either by parasitizing them or by inducing a defense response in the attacked plants (Poveda et al. 2020; Topalović et al. 2020). However, specific studies about the application of these biocontrol agents on grapes are scarce. In some cases, it was looked for bacterial biocontrol species isolated from grapevine rhizosphere (Aballay et al. 2011, 2013, 2017). It has been found that the mycorrhizal species Rhizophagus intraradices BEG141 induces local and systemic protection against X. index by inducing a priming of defence responses in grapevine (Hao et al. 2012). A similar observation is described for Diversipora versiforme against M. incognita in V. amurensis (Li et al. 2006). Trichoderma spp. and other endophytic fungi have also a great potential to control specific problems of PPNs (Poveda et al. 2020). However, many of these strategies must be proven in an integrated pest management (IPM) approach, since a single control measure usually did not achieve good final results and many of them are at the experimental laboratory level.
In addition to other agronomical measures, the most efficient and economic method to control PPNs is the use of resistant rootstocks. In some cases, specific virulent nematode populations have been overcome the resistance, as it is the case for Harmony and Freedom rootstocks by virulent populations of M. incognita and M. arenaria (Esmenjaud and Bouquet 2009). In this sense, resistance against some specific nematodes has been found in several Vitis spp. and in other related genera (Esmenjaud and Bouquet 2009). Ferris et al. (2012) developed several rootstock lines in California with a different degree of resistance to M. arenaria, M. incognita, X. index, Pratylenchus vulnus, C. xenoplax and T. semipenetrans from diverse resistance sources (V. rupestris, M. rotundifolia, V. rufotomentosa, V. champinii, V. riparia). These rootstocks maintain their resistance at high temperature and with different combinations and population levels of nematodes (Ferris et al. 2012, 2013). The resistance against X. index and GFLV is challenging because the nematode can acquire the viral particles within 1–10 min while feeding (Wyss 2014). The Institut Nacional de la Recherche Agronomique (France) has developed the rootstock Nemadex Alain Bouquet, which delays the appearance of GFLV in infested vineyards (Ollat et al. 2011). However, the poor performance of some of these Muscadinia-based hybrids in calcareous or dry soil conditions could reduce their utility in specific Mediterranean dry conditions (Ollat et al. 2016). Hypersensitive resistance reaction preventing the feeding and reproduction of nematodes is the most common resistance mechanism (Staudt and Weischer 1992; Esmenjaud and Bouquet 2009). Several major genes and/or regions have been found to confer resistance to Meloidogyne (N, Mur1, MJR1) (reviewed by Saucet et al. 2016; Smith et al. 2018b) and to X. index (locus XiR1) from V. arizonica (XiR1 locus: Xu et al. 2008; Hwang et al. 2010) and from M. rotundifolia (XiR2, XiR3 and XiR4 loci: Rubio et al. 2020) (see Sect. 4.5.2.1). Alternatively, approaches with transgenic plants showed good and promising results using different strategies: expressing the coat protein (Vigne et al. 2004), artificial miRNAs targeting the coat protein gene (Jelly et al. 2012) and nanobody-mediated resistance (Hemmer et al. 2018).
4.3 Description on Main and Emerging Pests
Among arthropods threatening grapevines, phylloxera (Daktulosphaira vitifoliae) is probably the most well known and documented pest, being responsible for the European phylloxera crisis which brought the rootstock era initiation. Nowadays, grape berry moths (Lobesia botrana, Eupoecilia ambiguella) are of economic importance in most European areas. In addition to grapevine moths, a number of leafhoppers (e.g. S. titanus, Empoasca vitis and Jacobiasca lybica), scale insects (e.g. Parthenolecanium corni, Pulvinaria vitis, Planococcus ficus and Margarodes spp.) and mites (e.g. Panonychus ulmi, Eotetranychus carpini, Colomerus vitis, Calepitrimerus vitis) are locally important and, due to climate change, some of them are steadily expanding their range. In this scenario, new invasive alien species are becoming increasingly important. Their arrival and impact on viticulture is facilitated by the increase in globalisation processes, such as the exchange of goods and the movement of people (Cini et al. 2014). Some recent examples in Europe with consequences on grapevine cultivation are Drosophila suzukii and Halyomorpha halys (Fig. 4.2).
4.3.1 Insects
Grapevines are grown in a wide variety of climates and situations, ranging from extremely hot and dry environments (i.e. Israel, Greece, Southern Spain, California, Arizona) to cold and humid areas (Canada, New Zealand, Moldova, Brazil). For this reason, the threats posed by insect pests and other phytosanitary problems can be very different. Moreover, unlike other tree crops, the grapevine is characterised by indeterminate growth, which means availability of growing and tender tissues throughout the production season, favourable to the trophic activity of many different insects. As a fact, insects can damage all the plant organs: roots, trunk, shoots, buds, leaves and berries. Worldwide, approximately 150 species of arthropods are considered harmful to grapevines, with either reversible damage (most of them manageable with appropriate integrated control strategies; see Pertot et al. 2017) or irreversible damage (i.e. insect vectors of incurable diseases such as phytoplasmosis; see Chuche and Thiéry 2014).
In the following paragraphs, besides phylloxera (D. vitifoliae) as a major insect pest worldwide and L. botrana as the most widespread key species in the Mediterranean areas, the main characteristics of two other species, S. titanus and P. ficus, were summarized, since they are becoming increasingly important for vines, partly because of their ability to transmit serious incurable diseases. Among the exotic species, we dedicated a focus on D. suzukii, whose harmfulness to grapevines has been the subject of numerous evaluations and hypotheses in recent years and which nevertheless constitutes a new and considerable risk. In particular, the current knowledge about the intraspecific (pheromones and vibrations) and interspecific communication with the host plant (chemical and physical factors that are responsible for the susceptibility of grapevines or their varieties to attacks by these insects) of these species will be summarized. The main control methods applied or under study that are based on interference with these languages will be also overviewed.
4.3.1.1 Daktulosphaira vitifoliae
Grape phylloxera, D. vitifoliae Fitch (Hemiptera, Phylloxeridae) is a notorious and well documented pest in viticulture. Especially in the European phylloxera crisis of the nineteenth century, when the Eurasian grapevine (V. vinifera L.) and wine sector first encountered the insect, a great effort was put into its understanding and management (Gale et al. 2002). Grape phylloxera is an obligate biotroph of Vitis spp. and feeds by sucking plant sap from former parenchyma cells that are transformed by the insect saliva into starch and amino acid-laden nutritional cells (Griesser et al. 2015). In doing so, it creates the typical pocket-like leaf galls on leaves, hook-like nodosities on root tips or crater-like tuberosities on mature roots. The latter especially can be lethal for susceptible grapevines, which is a main reason why V. vinifera vines are globally grafted onto resistant rootstock hybrids (Riaz et al. 2019). General reviews are already written elsewhere on grape phylloxera’s biology (Granett et al. 2001), life cycle (Forneck and Huber 2009) and host plant interaction (Powell et al. 2013). This section focuses on novel insights that build onto the gathered knowledge of these reviews and is a selection of genomic related approaches.
In commercial vineyards planted with grafted grapevines, oviparous grape phylloxera reproduces through cyclical parthenogenesis on the rootstock roots (of American hybrids) and sometimes on the scion leaves (on V. vinifera or hybrids thereof) (Forneck and Huber 2009). The success of infestation and damage to the grapevine are thereby influenced by temperature, water availability and rootstock hybrid (Savi et al. 2019). Grape phylloxera strains moreover also show an innate variability to infest grapevine species, and are therefore controversially grouped into biotypes (c.f. Downie 2010; Granett et al. 1985). Using controlled infestation assays, these biotypes are defined based on the ability to create leaf galls, nodosities or tuberosities on Vitis spp. (Forneck et al. 2016). Such biotype-specific infestation traits can be suppressed by elevated temperatures, illustrating the synergy of pest, host plant and environmental factors (Wilmink et al. 2021a).
Grape phylloxera population dynamic studies were initially conducted with RAPD assessments (Fong et al. 1995), but over the past decade have made way for microsatellite-based markers (Tello and Forneck 2019). Using such markers, studies in Australia showed that some phylloxera strains (called “super-clones”) are well adapted to different environmental circumstances and are abundant in a wide geographic range (Corrie et al. 2002; Umina et al. 2007). Such dominant clones were not found in Austria (Forneck et al. 2015). Indeed, grape phylloxera populations show a high genetic diversity within vineyard populations, with some stratification between root and leaf populations (Arancibia et al. 2018; Bao et al. 2015; Corrie et al. 2002; Riaz et al. 2017; Wilmink et al. 2021b). Besides differentiating between root and leaf feeding, these studies focused on population structure based on host plant and climatic adaptation. Studies on phylloxera’s native habitat e.g. by Downie (1999) and Lund et al. (2017), enable studying the deviation of population structure and life cycle variations in introduced habitats from that of its native habitat, east of the Rocky Mountains. In its native range, population structure is visible based on geography and host plant, and sexual recombination is much more common (Lund et al. 2017). The effort to use a standardized set of microsatellite markers to compare studies (Forneck et al. 2017) enabled a historical backtracking of two migration events from Europe to the northeastern coast of North America (Tello et al. 2019), which was also confirmed by genome sequencing (Rispe et al. 2020).
The genome of grape phylloxera covers 294 Mb and shows a unique and extraordinary expansion in putative effector genes (>2700) (Rispe et al. 2020) which may indicate novel adaptations of the insect feeding strategy. The significance of effectors as key elements of the interaction between the grape phylloxera and host plant was further demonstrated by the work of Zhao et al. (2019a) who provided functional evidence for SPRING genes in phylloxera. Rispe et al. (2016) in a transcriptomic de novo sequencing and Eitle et al. (2019a) in a proteomic approach further confirmed the relevance of effectors for the host plant interaction and beyond. With the progress in the ‘omic technology more information will be available to understand the complex biology of grape phylloxera. The feeding site (leaf gall versus root gall) greatly affects the transcriptomic profile of adult root versus adult leaf feeding phylloxera as shown in a de novo transcriptome assembly by Rispe et al. (2016). The host plant and the larval stages also greatly affect the transcriptomic profile as shown in a comparative transcriptome analysis of two grape phylloxera lineages (host-adapted to V. vinifera and a rootstock) that were studied in the mobile stage and in the sessile pre-adult stage (Savoi et al. 2020).
Understanding the host resistance against grape phylloxera, challenged in the past and present, calls for deeper knowledge on the susceptibility and compatibility between host and insect. Although histoid leaf and organoid root galls have captured interest of scientists since phylloxera discovery in the past, only recently the complexity and whole plant effects were shown on a molecular/transcriptomic level. These studies on both leaf gall (Body et al. 2019; Nabity et al. 2013; Schultz et al. 2019) and root gall (Du et al. 2011; Eitle et al. 2019b, c; Griesser et al. 2015; Kellow 2004) show the massive manipulation and reprogramming of the host plant. Effects like the increase of sink activity (of the galls) and actively compensating for the negative effects of herbivory through carbon loss have been shown for leaf galls (Nabity et al. 2013) and root galls (Griesser et al. 2015). This shows that the physiological and genomic regulation of sink strength, plant defense mechanisms, and carbon and nitrogen allocation play a key role in the plant–insect interaction.
A general review on different grape phylloxera management options was written by Powell (2012). Focusing on host plant susceptibility, most of the globally used rootstocks are resistant against tuberosity formation, yet allow populations to endure on non-lignified roots (forming nodosities) (Riaz et al. 2019). During the annual growing season, these populations may migrate to infest the host plant’s leaves. Though V. vinifera scions are less susceptible to leaf galls than American Vitis spp. such as V. riparia or V. rupestris,—which are broadly used in rootstock hybridization—an increased leaf infestation rate in commercial vineyards is evident. Albeit the reasons are not yet clear, biotypes with altered leaf-feeding performance have been confirmed (Wilmink et al. 2021b). Several pesticides are available to combat such leaf infestations, reviewed by Yin et al. (2019). For countries that still grow own-rooted V. vinifera vines and rely on local quarantine measures (like Australia), the early detection of belowground phylloxera infestations is crucial. This can be based on classical phylloxera trapping systems that collect emerging larvae, and/or on high-throughput hyperspectral scanning to predict (phylloxera-related) grapevine health decline (Vanegas et al. 2018). Using portable loop-mediated isothermal amplification (LAMP) assays, phylloxera mitochondrial DNA can furthermore be identified genetically, without false-positives from non-target species contamination, allowing phylloxera identification on species level in both lab and field (Agarwal et al. 2020). To prevent phylloxera cross-contamination between vineyards, viticulture material can be treated with dry air, steam or hot water to eradicate phylloxera larvae (Clarke et al. 2018, 2019).
Host resistance mechanisms against phylloxera on roots (rootstock hybrids) and scion (V. vinifera) are mostly not complete, allowing the insect to continue feeding on its host, often without visual damage or decay of the host plant (Powell et al. 2013). The known defense mechanisms in rootstocks are both structural and biochemical, such as elevated levels of phenolic metabolites in parenchymal cells (Du et al. 2011; Eitle et al. 2019c; Lawo et al. 2011) as well as cell wall properties in root tips or lignified cambium cells. Defense mechanisms in roots and leaves seem to be involved, but studies are still scarce. Wang et al. (2019) showed that transcription factor WRKY46 is phylloxera-responsive and involved in the salicylic acid (SA)-mediated defense regulatory response in roots. The jasmonic acid (JA)/SA crosstalk is activated by phylloxera feeding and SA seems to play a role in root galls (Du et al. 2011; Eitle et al. 2019b). Because of the limited resistance of most commercial rootstock hybrids, which moreover only possess a narrow genetic base (Riaz et al. 2019), there is a pressing need to develop new rootstocks with defined resistance genes (Clark et al. 2018; Rubio et al. 2020). Rapid necrotic response and generally no gall induction (= no phylloxera feeding) are pursued resistant traits. Genetic sources for phylloxera resistance have been reported as in accessions of V. cinerea (‘Arnold’) and M. rotundifolia (Blank et al. 2009; Rubio et al. 2020; Smith et al. 2018a; Zhang et al. 2009) and first studies showed that variable numbers of loci control the resistant traits (tuberosities vs. nodosities vs. leaf galls) (Clark et al. 2018; Roush et al. 2007; Rubio et al. 2020). When these rootstock resistance traits are incorporated into broadband rootstock hybrids, work is still to be done for phylloxera leaf resistance (see Sects. 4.5.2.1 and 4.10).
4.3.1.2 Lobesia botrana
The European grapevine moth, L. botrana (Denis & Shiffermüller) (Lepidoptera Tortricidae) is the key pest of grapevine in Europe where it is endemic and widespread in all wine growing areas, but it is economically most important in Southern Europe. In Southern France, Central and Southern Spain, Portugal, Greece, Italy and the islands of the Mediterranean basin, it is the only moth that has a significant impact on grapevine production. Recently, L. botrana was found in the important South and North American wine growing regions of Argentina, Chile and California (Gilligan et al. 2011; Gonzalez 2010). However, in California it has been declared eradicated through the fast application and integration of different control methods on a territorial scale (Simmons et al. 2021).
Concerning pest genetics and genomics, a novel set of microsatellite markers for L. botrana were isolated using next-generation sequencing (NGS) and employed for genetic characterization of populations from Europe and the Middle East (Reineke et al. 2015). In addition, a complete mitochondrial genome of the European grapevine moth have been sequenced (Piper et al. 2016), followed by the analysis of the antennal transcriptome and expression of odorant-binding and chemosensory proteins (Rojas et al. 2018).
The damage is caused by larval feeding on grape clusters that renders them susceptible to B. cinerea in mid-season, leading to the development of primary and secondary roots at harvest. Due to the direct and indirect damage it causes, appropriate control measures are required, often represented by the use of broad-spectrum insecticides, which may cause outbreaks of spider mites and other minor pests (Thiéry and Moreau 2005). These issues have promoted the search for effective alternatives with a reduced risk, such as semiochemical-based control methods, biological control, microbial and botanical pesticides. One of the alternative biological control methods that has demonstrated the greatest effectiveness over large areas (area wide pest management, AWPM) and in the long term is the pheromone MD (Ioriatti et al. 2011; Ioriatti and Lucchi 2016). Mating disruption with hand-applied multipurpose reservoir dispensers is indeed the most effective and widely applied pheromone-based control technique used against grapevine moths worldwide. This strategy essentially relies on permeating the crop with relatively low amounts of synthetic sex pheromone that disrupt intraspecific communication of the target species and thus prevent mating. The case of the province of Trento in Northern Italy can be cited as an example and model for the effective and large-scale application of this control strategy (Ioriatti and Lucchi 2016). This territory indeed is recognised as a pioneer in Italy in the application of MD against grapevine moths, L. botrana, Eupoecilia ambiguella (Hb.) and in some areas also another tortricid moth, Argyrotaenia ljungiana (Thunb.), for which multiple purpose dispenses have been developed. Currently, the vine growing area in the Province of Trento involved in pheromone MD represents about 95% of the total (about 9,700 ha), and together with the area involving the apple tree, represents the largest area treated with pheromones in Italy. However, further research is still needed to improve the efficacy of MD technology, and to manage invasive and secondary pests without compromising the value of the pheromone-based approach for control of the target tortricid pests. For example, it remains essential to develop and commercialize novel and/or better formulations that are more effective, cheaper, and easier to deploy. In this sense, aerosol technologies may provide a cost effective alternative to hand applied dispensers (Benelli and Thomson 2019).
However, it would be very important to supplement the techniques based on sex pheromones with new technologies that can also interfere with the behaviour of the insect's females, through the use of volatile compounds emitted by the host plant that are attractive to the egg-laying females (kairomones). To this end, grapevine volatile organic compounds (VOCs) involved in host-plant interactions were extensively studied for L. botrana, and it was found that a specific blend of the terpenoids (E)-β-caryophyllene, (E)-β-farnesene and the homoterpene (E)-4,8-dimethyl-1,3,7-nonatriene was attractive in laboratory and field conditions (Anfora et al. 2009). The attractiveness was shown to be dependent on the kairomone ratio, decreasing significantly when deviating from the ratio found in the grapevine headspace collection (Tasin et al. 2006, 2011). Based on this knowledge, stable transgenic lines of grapevine have been generated recently with altered (E)-β-caryophyllene and (E)-β-farnesene emission compared to original unmodified plants (Salvagnin et al. 2016, 2018). It has been shown that modification of the kairomone ratio within the grape bouquet is sufficient to interfere with the host-finding behaviour of L. botrana making the transgenic plants less attractive to the female moths in wind tunnel experiments. This finding could form the basis for development of new environmentally friendly approaches for pest and pathogen control, also exploiting the new biotechnologies for plant genome editing such as the clustered regularly interspaced short palindromic repeats (CRISPR)-associated protein 9 (Cas9) system and its subsequent developments (Anzalone et al. 2019).
4.3.1.3 Planococcus f icus
The vine mealybug, Planococcus ficus (Signoret) (Hemiptera: Pseudococcidae), native to Mediterranean countries, is now widespread in all five continents where it is a key grape pest in the most important grape growing regions (Daane et al. 2018; Cocco et al. 2021). Planococcus ficus is a polyphagous species that can grow on herbaceous, shrubby and tree-like plants, with a preference for figs (hence the specific name) and grapevines. It infests all green organs. In Mediterranean areas, P. ficus can go through 3–4 generations. Overwintering is mainly carried out by fertilised females, although in many cases females with ovisacchus and juvenile forms may also be present under the rhytidome in winter.
Based on molecular genetic analysis, the geographic origin of invasive populations of the mealybug P. ficus in North America has been determined (Daane et al. 2018). Moreover, the complete genome of Planococcus spp. is now available (Jung et al. 2018).
The damage is due to the abundant emission of honeydew on which fungus settle, with leaf yellowing and phylloptosis (Walton et al. 2004). Furthermore, sooty mold and mealybug colonies severely affect the aesthetic quality of table grapes, thereby reducing its market value. Moreover, this pest is an effective vector of the GLRaV-1, 3, 4, 5 and 9, Kober stem grooving (GVA) and corky bark disease (GVB) (Bertin et al. 2016; Tsai et al. 2010). In particular, the spread of GLRaV-3 appears to be very rapid and can infect both white and red grapes. The symptoms of leafroll viruses are multiple and range from a reduction in photosynthetic capacity to effects on the growth of all plant organs and the sugar content of the berries (Daane et al. 2018).
Conventional control programs for P. ficus in vineyards still rely on repeated applications of synthetic insecticides throughout the grape growing season, including organophosphates and neonicotinoids (Walton et al. 2004, 2006; Daane et al. 2012), which showed both severe non-target effects on beneficial insects (Mansour et al. 2018). Moreover, the efficacy of non-systemic insecticides is often insufficient because normal treatments cannot adequately reach the individuals that often live under the bark, at root level and in the shelter of the clusters. Agronomic practices that increase sunlight and reduce humidity within the vineyard (i.e. rational irrigation, pruning, fertilization and grassland management) can help to at least partially reduce damage from P. ficus by making the environment less suitable for the explosion of its populations (Cocco et al. 2021).
In recent years, as part of the development of IPM strategies against P. ficus, augmentative and conservative biological control is playing an increasingly important role. In most cases, inoculation with one of the two species of koinobiont endoparasitoids Anagyrus pseudococci (Girault) and Anagyrus vladimiri Triapitsyn (Hymenoptera: Encyrtidae), and the generalist predator Cryptolaemus montrouzieri Mulsant (Coleoptera: Coccinellidae) is used for this type of biological control (Cocco et al. 2021).
The potential application of MD for controlling the mealybug infestations was also investigated by testing different dispensers and formulations of the synthetic sex pheromone (Walton et al. 2006; Daane et al. 2020). Results were promising as MD consistently altered male orientation and often reduced the pest density and/or the crop damage, leading to the development of commercial MD products. The use of this environmentally sustainable and preventive control technique against P. ficus is increasing in all wine growing areas of the world and new commercial products are being registered, thus ensuring further development. The recent development of double dispensers for the simultaneous control of L. botrana and P. ficus (i.e. Isonet® LPF) is another key factor in maintaining effective IPM in the vineyard (Cocco et al. 2021). As the integration of biological control and MD has proven to be very effective, the timely use of these two biological tools is the most sustainable and short-term perspective for the control of P. ficus (Shapira et al. 2018).
4.3.1.4 Scaphoideus t itanus
The American grapevine leafhopper, S. titanus Ball (Hemiptera Cicadellidae), is a leafhopper native to North America, which is currently widespread in most of the European wine growing areas (Chuche and Thiery 2014). Introduced at the end of the 1950s in France, it slowly spread to the rest of Europe and today has been recorded from the main grape growing EU Member States (Austria, Croatia, France, Hungary, Italy, Portugal, Slovenia and Spain) as well as in Switzerland and in Serbia (EPPO 2021). Although it can be collected from several arboreal plants (i.e. Crataegus, Juniperus, Ulmus and Fraxinus) (Barnett 1976), and weeds (i.e. Taraxacum, Trifolium) (Trivellone et al. 2013), S. titanus requires grapevine plants for the completion of its life cycle. Notably, while in North America S. titanus is mostly associated with wild grapevines (i.e. V. labrusca, V. riparia), in Europe it can be easily found also on cultivated V. vinifera (Jeger et al. 2016).
Microsatellite and mitochondrial data provided evidence for a single major introduction for S. titanus in Europe (Papura et al. 2012). Mitochondrial genomic variation was also exploited to detect phylogenetic relationships of three groups in the genus Scaphoideus (Du et al. 2017).
This leafhopper is univoltine and overwinters as eggs under the grapevine bark, especially on 2 years old shoots (Bagnoli and Gargani 2011). Most hatchings occur between May and June and the peak of nymphs’ presence in the field is in June while adults appear from early July and can be found in the vineyard until leaf fall (Chuche and Thiery 2014). Although the feeding activity does not cause any visible direct damage to the grapevine vegetation, S. titanus can transmit the phytoplasma causal agent of FD, a EU quarantine disease (listed under Commission Implementing Regulation 2019/2072 in Annex II, Part B), which is part of the elm yellows group (16Sr-V). The association of S. titanus to the FD is the reason for the adoption of mandatory insecticide treatments against the species in most regions by national decrees. The control strategy aims at preventing the spread of the disease, which is acquired by nymphs and requires an approximate latency time of 4–5 weeks before being transmittable to the grapevines. In adults, however, the latency period is much shorter and individuals can transmit the pathogen as early as one week after acquisition (Alma et al. 2018). Scaphoideus titanus is a persistent propagative vector, therefore once the pathogen has been acquired, it remains infective lifelong.
The main target of the insecticide treatments are nymphs of 3rd–5th instars, present in the vineyards around mid-June, with local variability due to the specific microclimate. However, in case the presence of adults is revealed by the monitoring through colored sticky panels, a second or even a third insecticide treatment will be performed later in the summer. Several chemical classes are used against S. titanus: neonicotinoids, organophosphates, pyrethrins, pyrethroids, growth regulators (thiadiazin, buprofezin). The use of pyrethrins by organic growers seems to be scarcely effective primarily because of the short persistence (Chuche and Thiéry 2014).
Measurements of FD control, beside the vector control through insecticides, also include uprooting of infected plants and the prophylaxis through the production of healthy propagation material. In case of infections exceeding 20–30% of the plants in a plot, the entire vineyard must be rogued. In this regard, phytosanitary services can impose sanctions to farmers that do not comply with the law obligations.
Biological control now does not appear to be a feasible strategy, since no efficient agents (parasitoids and predators) are known. Principal parasitoids of S. titanus nymphal stages are Dryinid wasps and Pipunculidae flies, but neither of them are species-specific and the few attempts resulted in poor results (Chuche and Thiéry 2014). As for behavioral manipulation, a technique of MD based on the transmission of disruptive noise (i.e. vibrations) is currently experimented in several vineyards of Northern Italy. In fact, S. titanus uses vision and olfaction for host detection (Mazzoni et al. 2009a) whereas the mating behavior is mediated exclusively by vibrational signals. The method of vibrational MD aims at interfering with the mating communication of S. titanus thus preventing mating (Mazzoni et al. 2009b). In practice, by exchanging species-specific vibrational signals, a male and a female establish a mating duet that allows a reciprocal identification and the location of the female by the male on the host plant. Both laboratory and field tests confirmed that the release, of disruptive vibrations by electromagnetic transducers, specifically designed to mask the species mating signals, compromises the process of pair formation similarly to what happens in the classic pheromone MD (Eriksson et al. 2012; Mazzoni et al. 2019; Nieri et al. 2021).
4.3.1.5 Drosophila s uzukii
Drosophila suzukii (Matsumura) (Diptera Drosophilidae) is native to the South East Asia and was found for the first time outside its native habitat in the Hawaiian Islands in 1980, then in North America and Europe in 2008 (Walsh et al. 2011; Cini et al. 2012; Asplen et al. 2015), and more recently in South America and Africa in 2013 (Deprá et al. 2014; Ouantar et al. 2020). The species has caused extensive damage in all regions where it became established and has demonstrated a very rapid expansion. Drosophila suzukii is a highly polyphagous insect but most of the damages were recorded on fruits of blueberry, strawberry, raspberry, cherry, and on some varieties of grapes (EPPO 2020a). In natural ecosystems, D. suzukii reproduces on many wild fruits such as blackberry, elder, buckthorn, etc. Unlike other Drosophilids, D. suzukii shows the peculiar characteristic of females being able to lay their eggs inside intact ripening fruits, starting from veraison and preferring fruits with a thin epicarp. As a fact, females have a well-developed saw-like ovipositor, which can penetrate beneath the skin of host fruits (Crava et al. 2020).
As mentioned before, grapevine is also counted among the host plants of this pest. However, it has been shown that only some varieties, due to their chemical-physical characteristics, are susceptible to oviposition during the ripening phase (e.g. the ‘Schiava’ variety) (Ioriatti et al. 2015; Baser et al. 2018). The key factor that makes some varieties more susceptible to D. suzukii attacks during ripening before harvest is the hardness of the skin, which the insect must be able to pierce with its ovipositor. Drosophila suzukii in fact prefers above all varieties with red berries, late harvest and less firm skin. Although the penetration force is the main element influencing the susceptibility of grapevine, host attractiveness will likely depend upon additional factors, such as soluble sugar content (Burrack et al. 2013; Ioriatti et al. 2015). The role of D. suzukii as a vector of spoilage bacteria responsible for sour rot of grape has been also demonstrated (Ioriatti et al. 2018).
Regarding pest genetics and genomics, studies about genetic variability of specific D. suzukii populations have been reported (e.g. Tait et al. 2017). Moreover, two draft genome assemblies already exist for this species. The first one was framed within the linking between genomics and ecology to investigate the invasive Drosophila complex evolution (Ometto et al. 2013). To enable basic and applied research of this important pest, basic properties of a second sequenced genome and transcriptome were reported, describing patterns of genome evolution in D. suzukii and its close relatives (Chiu et al. 2013). Both draft genomes contain pervasive assembly errors and are highly fragmented, which limits their values. To improve the assembly of the D. suzukii genome and to annotate it in a way that facilitates comparisons with D. melanogaster, long-read sequencing data were produce and assembled, resulting into a novel, high-quality D. suzukii genome assembly (Paris et al. 2020).
Although various control approaches have been implemented to suppress D. suzukii populations and reduce crop damage, current programs still rely primarily on insecticides that target adult flies. Insecticides can be effective, but there is a restricted list of permitted active ingredients and increasing problems with fruit residues (Haye et al. 2016). Insecticide applications may also lead to the development of resistance, which is of major concern for this insect over the longer term. Therefore, given also the high polyphagy and mobility of the pest, only area-wide IPM strategies aimed at reducing population densities at the landscape level would be able to achieve sufficient levels of effectiveness. New IPM tools are hence under development including those based on semiochemicals, cultural methods, exclusion nets, sterile and/or incompatible insect techniques possibly in combination with transgenic approaches (EPPO 2020b). However, none of these methods alone or together has shown yet to significantly control D. suzukii populations on a large scale. In the light of these results, the method that seems most promising, suitable and durable over large areas is biological control. In this sense, many studies have been carried out and are at an advanced stage of application using both indigenous and exotic arthropods, predators and microorganisms (Lee et al. 2019). Biological control in area-wide programs should be able to reduce pest populations in natural habitats, thereby reducing the number of flies that migrate into susceptible crops that, in turn, will improve the effectiveness of other integrated control tools and lower damage (Rossi Stacconi et al. 2019; Nieri et al. 2021). In several countries, classical biological control projects are in different stages of preparation and evaluation. For most of these projects the plan is to release the effective and highly specific wasp, Ganaspis brasiliensis (Ihering) (Hymenoptera: Figitidae), the main larval parasitoid of D. suzukii in the native areas.
4.3.2 Mites
Main mite pests of grapevine belong to the Tetranychidae and Eriophyidae families of the Acari Prostigmata. Spider mites of importance in European vineyards are Tetranychus urticae Koch, Panonychus ulmi (Koch) and Eotetranychus carpini (Oudemans). Among eriophyoid mites, Colomerus vitis (Pagenstecher) and Calepitrimerus vitis (Nalepa) are important for viticulture (Duso et al. 2012).
The tetranychid mite P. ulmi is widespread in temperate areas while less common in the warmer Mediterranean climate. Overwintering occurs as eggs laid from late summer onwards, usually at the brancing point of the shoot along one or two-year branches. The eggs hatch between April and May, and the first juveniles are present during sprouting when they complete their development in about 20 days. Afterward, P. ulmi can develop from four to seven generations per year (Bovey et al. 1979; Girolami 1981). Typically, P. ulmi populations remain at low levels in the first part of the season and increase in mid or late summer (Wermelinger et al. 1992; Duso and Pasqualetto 1993). Population increases are favored by high temperature and high leaf nitrogen. Infestations by P. ulmiare affected by grapevine variety, and in particular, leaf hairiness promotes high spider mite density (Schreiner 1984; Rilling 1989). Panonychus ulmi showed some differences in host preference between V. vinifera and V. labrusca, being the latter less preferred host for oviposition; however, when different varieties of V. vinifera are considered, no differences emerged (Johann et al. 2019; Silva et al. 2021). Eotetranychus carpini is another important tetranychid mite pest in Southern Europe. It overwinters as females under bark, and in spring it moves to the developing canopy. This spider mite can complete 4–8 generations per year. Optimal conditions for population increase are mild temperature, low relative humidity, and pubescent leaf undersurfaces (Duso and Vettorazzo 1999). The two-spotted spider mite T. urticae is a polyphagous species that has been observed to infest grapevines as well as several weeds occurring in vineyards (Schruft et al. 1979; Arias and Nieto 1981; Boller et al. 1985). It typically overwinters as females under bark and on weeds present in the vineyards (Schruft et al. 1979). In warm climate conditions, young stages can also be found during winter (Arias and Nieto 1980). Overwintered stages colonize the newly developed grape vegetation during spring and up to 15 generations develop afterward, depending on climate conditions. A grapevine cultivar effect has been observed related to biological parameter of these mites (Amala et al. 2016). Other spider mites infesting grapevine are Tetranychus mcdanieli (McGregor), important in North America and in France, Tetranychus turkestani (Ugarov and Nikolskii) observed to infest grapevine in France, Spain and Portugal, Tetranychus pacificus (McGregor) and Eotetranychus willametei (Ewing) recorded in vineyards in North America (Duso et al. 2012; Wilson and Daane 2017).
Spider mites damage grape leaves by puncturing spongy mesophyll and palisade cells and sucking out the contents. The symptoms associated with spider mite infestations can differ among mite species, but generally, they cause leaf discoloration, yellow (on some cultivars reddish) leaf spots, and changes in gas exchange rates, chlorophyll and other metabolites content. Severe infestation with high-density can disrupt leaf functionality and lead to defoliation, causing economic damages in some cases (Duso et al. 2012). Transcriptome analysis of grapevine responses to T. urticae infestation revealed a host-resistance response difference between grapevine-adapted or non-adapted mite strains. Spider mites infestation induced the down-regulation of photosynthesis, cell division and growth, and an up-regulation of genes involved in biosynthesis, signaling, and responses to JA, ethylene (ET), and SA. The level of responses was proportional to the damage inflicted by the different strains, with a weak response associated to the non-adapted strain and a strong but ineffective defense triggered by feeding of the grapevine-adapted strain (Díaz-Riquelme et al. 2016).
Eriophyoid mites are tiny organisms that entrain a close relationship with their host plants (de Lillo et al. 2018). Among them, two species are considered of importance for grapevines. The grape rust mite Cal. vitis is a severe pest in different viticultural regions in the world (Bernard et al. 2005; Duso et al. 2010). Females are the overwintering stages, under the bud scales or bark at the insertion between one- and two-year-old branches. In spring, overwintered females lay their eggs at the base of the shoot. Juveniles colonize the shoots during spring, colonizing leaf undersurfaces and new buds at the leaf axils. Generally, population densities increase afterward, with peaks reached in mid-summer. Two to four generations are reported, but a higher number of generations are possible in areas characterized by optimal conditions for population development (Duso et al. 2010; Walton et al. 2010). Symptoms related to feeding activity of the rust mites are scar tissue and bronzed leaves in summer, death of the growth point of buds, stunted shoot growth, shortened shoot internodes, development of lateral shoots, leaf deformation, reduced cluster size and flower drop and has been associated to the so-called “short shoot syndrome” (Walton et al. 2007; Duso et al. 2010). In Europe, severe infestations were recorded from the 1980s especially in young vineyards (Duso and de Lillo 1996).
The grape erineum mite Col. vitis is present in main viticultural areas of the world and based on the symptoms produced on grapevine three strains have been distinguished (Smith and Stafford 1948): bud gall, leaf erineum (with the acronym GEM, i.e. grape erineum mite) and leaf curl. The erineum strain is more widely distributed than the other strains (Duso and de Lillo 1996; Valenzano et al. 2020). This strain is characterized by overwintering as female under the external bud scales or under the bark. Overwintered females start being active in spring at bud break by inducing the first felty patches (named “erinea”) on newly formed leaves and then start reproducing. Basal leaves are usually infested early in the season, while in-season infestations are observed on apical leaves, where a large number of erinea are produced. Erinea can also be produced on inflorescences. Up to seven generations of this mite can develop yearly. Thus, long-time dispersal can play an important role in the distribution of this species (Valenzano et al. 2019). For a long time considered a minor pest of grapevine, recent studies highlight that Co. vitis infestation can negatively affect plant growth and physiology (Javadi Khederi et al. 2014a; Javadi Khederi et al. 2018a). Additionally, the erineum strain of Co. vitis was associated with the transmission GPGV and GINV (Malagnini et al. 2016; Morán et al. 2018).
Spider mites and rust mites are considered pesticide-induced pests. Chemical control using acaricides can be a possible option, but restrictions in the use of pesticides and risks of development of pesticide-resistance mite’s strains represent a limitation for its long-term adoption (van Leeuwen et al. 2010; Bajda et al. 2015; Van Leeuwen et al. 2015; Rameshgar et al. 2019a, b; Badieinia et al. 2020). In this context, it should be considered that sulfur-based fungicides could have some effect in reducing mite infestations, particularly eriophyids (Cooper et al. 2020). However, conservation and enhancement of the populations of their natural enemies, particularly predatory mites, should represent the first option for mite pest management (Duso et al. 2012). This can be obtained by reducing pesticides impact on beneficial mites and by adopting habitat management tactics (Pozzebon et al. 2015; Pennington et al. 2017; Sáenz-Romo et al. 2019; Tixier 2018; Duso et al. 2020; Zanettin et al. 2021). Leaf morphology strongly affects the presence and abundance of predatory mites (e.g., Duso et al. 1992; Camporese and Duso 1996; Schmidt 2014; Pozzebon et al. 2015), and genetic factors that control the grapevine foliar traits can be exploited in indirect forms for breeding of host-plant resistance to pests by promoting the abundance of predators in new cultivars (Barba et al. 2019) (see Sect. 4.6.2). Genetic information related to plant responses to pest mite infestation can be used in breeding programs aimed at direct host-plant resistance. This type of information is available for the interaction between grapevine and the eriophyid Col. vitis (Javadi Khederi et al. 2014b, 2018a, b, c, d).
4.4 Brief on the Host Phenotypic Characterization
4.4.1 Milestones from Phenotyping to Phenomics
Phenotypes are the qualitative and quantitative characteristics (traits) of organisms that are of the most interest and phenotyping is the process of their assessment. By studying phenotypic traits, useful explanations of important outcomes such as disease can be obtained. Phenotypic variation is produced through a complex web of interactions between genotype and environment, and such a genotype–phenotype map is inaccessible without the detailed phenotypic data that allow these interactions to be investigated. Despite this need, our ability to characterize phenomes—high-dimensional phenotypic data on an organism-wide scale—still lags behind our ability to characterize genomes. Given the high dimensionality of phenomes, analyses of phenomic data call for new concepts and techniques and require collaborations between scientists with diverse expertise (Houle et al. 2010). The phenomic tools and techniques are paving the way in harnessing the potentiality of genomic resources in genetic improvement. These techniques have become much more advanced and have now entered the era of high-throughput integrated phenotyping platforms to provide a solution to genomics-enabled improvement and address our need for precise and efficient phenotyping of crop plants (Zhao et al. 2019b).
With the rapid development in high-throughput phenotyping technologies, also this area of grapevine research is entering a new era called “phenomics”. Besides increased throughput, phenomics brings opportunities and challenges in objectivity, precision, dynamic measures, and integration that demand new approaches for standardization, data management, and analysis (Kumar et al. 2015). Reliable, (semi-)automated, multifunctional, and high-throughput phenotypic technologies are increasingly considered as important tools for rapid advancement in genetics as well as in grapevine breeding programs. Traditionally, phenotyping techniques relied on measurement of visual, chemical, physiological, or other characteristics by experts, often at low-throughput. The use of standardized descriptors and scales to phenotype grapevine traits has provided a good foundation for international adoption of phenotyping standards and cross-comparison of results. However, many of these descriptors are subjective and fail to capture complete trait variation (Cadle-Davidson et al. 2019).
4.4.2 Phenotype-Based Diversity Analysis: Possibilities and Constraints
Although the perennial grapevine entails a series of developmental features that makes phenotype studies monotonous and costly, in the last years the advent of new phenotyping techniques and technologies allowed biotic resistance traits to be characterized in large germplasm collections, segregating populations, series of varieties or specific clones. Some traits are more amenable to high-throughput methods by exploiting associations between the whole vine and samples of the vine, such as disease and pest resistance. While the evaluation of foliar resistance is straightforward, there are limitations in berry phenotyping, such as the long time required for fruiting and the protracted juvenile stage. Large areas are required for studies at the whole-plant level when comparisons require multiple genotypes and a sufficient number of replicates. Unlike model plants, grapevine is a typical outdoor crop that is exposed to fluctuating environmental conditions in the field. Therefore, the plasticity of the phenotype should be tested over a number of years at different locations before a new variety is released (reviewed by Delrot et al. 2020).
There is a great challenge for the grapevine community regarding the study of the genotype by environment interactions (GxE). Addressing the challenges of GxE requires standardization and careful documentation of phenotyping protocols, an effort that is gaining international relevance Several international collaborative phenotyping projects and networks (TransPLANT, regional and International Plant Phenotyping Networks and ELIXIR-EXCELERATE) have developed resources for standardized phenotyping. One notable effort is minimum information about a plant phenotyping experiment (MIAPPE; www.miappe.org), which outlines the suggested and required attributes for describing the metadata of the experiments. Standardization and careful documentation promote better data management and make data reusable for purposes beyond what was originally intended or beyond current resources. To this end, a set of findability, accessibility, interoperability and reusability (FAIR) principles has been developed. FAIR's vision in relation to the grape community has started to be organized through a global grape information system (GrapeIS) organized by the international grapevine genome program (IGGP; www.vitaceae.org), and its success will depend on active participation of those who are part of the grape community that generates, analyzes and publishes data (Adam-Blondon et al. 2016; Cadle-Davidson et al. 2019) (see Sect. 4.11.5).
While studies under field conditions are required to reproduce real vineyard environments, an initial assessment of the phenotype under controlled conditions is advisable in many instances to directly and efficiently evaluate biotic stress resistance—as most climate- and environment-smart traits—in large-scale screenings. In addition, bioassays for the assessment of symptoms are fundamental to shed light onto host–pathogen/pest interactions; conveniently, these can be carried out by observations in greenhouses or under laboratory conditions (ex vivo and in vitro).
In a greenhouse setting, different strategies can be applied according to the goal. Explants of potted grapevine are a valuable system to assess the response to some pathogens at leaf level (e.g. Boso et al. 2014), whereas plants in hydroponic media proved useful to evaluate responses to herbivorous pests (e.g. Díaz-Riquelme et al. 2016). Since it is known that some diseases have dual epidemics, it is also important to be able to evaluate the response of the flower organ and/or the fruit. A smart system that shortens time lapses for flower/fruit phenotyping is the use of fruitbearing (or fruiting) cuttings. To do so, after cane rooting induction and sprouting, only one inflorescence per cane is left, enabling fruit set by limiting competition for nutrients in the cutting (Mullins et Rajasekaran 1981; Antolín et al. 2010). Recently, the use of fruiting cuttings allowed the development of a new lab phenotyping method to assess the disease extent on inflorescences at the phenological stage of flower button (Buonassisi et al. 2018). Under laboratory conditions, ex vivo methods or in vitro protocols can be performed: they are only feasible for inoculation of some pathogens and challenging for infestation with some pests.
Ex vivo pathogenesis assays are conducted using detached leaves or leaf discs, and different inoculation strategies have been used. For instance, inoculation of detached leaves or leaf discs as well as detached berries in Petri dishes enables reliable phenotyping for tolerance/resistance of grapevine genotypes to major fungal diseases such as PM, DM, BBR and leaf rust (e.g. Sargolzaei et al. 2020; Bierman et al. 2019; Lovato et al. 2019; Gomes et al. 2019; Zendler et al. 2021a). Although in vitro methods are usually less representative of the whole-plant ecophysiology, they are still convenient for a rapid screening of specific features for ad hoc objectives (e.g. Algarra Alarcon et al. 2015).
4.4.3 Approaches for Biotic Stress Symptom Assessment
4.4.3.1 Visual Evaluation
Visual rating is a starting point using a comprehensive set of traits described by the Organisation Internationale de la Vigne et du Vin (OIV) phenotypic scales, which were developed primarily for the standardized description of grapevine varieties and species. Nowadays, the OIV code (OIV 2009) includes twelve resistance descriptors for five biotic stresses (such as DM, PM, BBR, Eutypa dieback and phylloxera) at various organs; some have recently been proposed for P. ampelicida at foliage (Rex et al. 2014) and for P. viticola at inflorescence (Buonassisi et al. 2018). The descriptors consist of a 9-point Likert rating scale (Likert 1932) with five discrete categories that describe a continuous variable (the resistance). Thus, for users keen on biotic stresses it functionally becomes a 5-point or even a 3-point scale that determines a limited resolution of knowledge gained. However, this symptom assessment method has been largely deployed in genetic analysis such as genotype–phenotype associations. Besides OIV, there are other discrete scales developed by unofficial or official agencies such as the International Union for the Protection of New Varieties of Plants (UPOV). Indeed, the European and Mediterranean Plant Protection Organization (EPPO) proposes the evaluation of symptoms both qualitative—counting the ratio of organs infected over the healthy ones (disease incidence)—and quantitative—by measuring the percentage of organ surface affected by symptoms with respect to the total surface (disease severity).
The degree of care in designing, executing, and describing resistance phenotyping experiments are highly variable but recent studies have improved attention to detail. Most studies rated disease on a categorical scale following natural infection, often attempting to relate their ratings to the OIV scale. Several of these studies rated disease progression over time and found that the significance of QTL changed over time (e.g. Bellin et al. 2009; Zendler et al. 2017), with some QTL being undetectable if the wrong time point was selected (Barba et al. 2014) or a too early time point was chosen (e.g. Vezzulli et al. 2019a). Time series ratings provide the added opportunity for area under the disease progress curve (AUDPC) analysis (e.g. Bove et al. 2019; Possamai et al. 2021; Clark et al. 2018; Yin et al. 2021). Recently, efforts to develop and standardize controlled infection ex vivo have allowed for detection of moderate or minor QTL that may not be detected in vineyard evaluations (e.g. Sapkota et al. 2019a; Ochssner et al. 2016; Rubio et al. 2020). For loci that have been genetically mapped using multiple phenotypic parameters, the degree of QTL significance may provide some insights into which phenotyping methods best explain the genetics of resistance. If feasible, single isolate inoculation of detached leaves or discs appears to be the current best method for detecting minor or moderate QTL and may be the most relevant for fine mapping and characterization of candidate genes (Cadle-Davidson et al. 2019). Improved accuracy in visual symptom assessment can be achieved through histochemical staining and microscopic analysis. Light microscopy can distinguish hyphae, appressoria, conidia and conidiophore, while haustoria cannot be monitored. Moreover, confocal scanning electron microscopy and low-temperature scanning electron microscopy have been used (Nascimento-Gavioli et al. 2020; Hu et al. 2019; Rahman et al. 2020; Ullrich et al. 2009). Studies in other pathosystems have emphasized the subjectivity and imprecision of visual ratings as well as the importance of rater subjectivity. Fortunately, QTL were consistent across raters, even if inconsistency in ratings affected the magnitude of these QTL effects (Poland and Nelson 2011).
4.4.3.2 Molecular Tools for Causal Agent Detection and Identification
While many phenotypes of interest represent the endpoints of gene expression, the molecular and genetic signals themselves could be considered valuable phenotypes to be measured and mapped. Unlike conventional methods that rely on visual symptoms, isolation, and/or culturing, pathogens can be detected using molecular techniques, such as enzyme-linked immunosorbent assays (ELISA), DNA/RNA probes, or polymerase chain reaction (PCR) amplification of nucleic acids including quantitative PCR or reverse-transcription PCR. Enzyme-linked immunosorbent assays, which confirm the presence of pathogens upon fluorescence or other visible chemical reactions, are standard techniques for detecting viruses, fungi, and other microbes, and are affordable and easy to perform. However, immunological procedures rely on antibody-based recognition of antigens produced by the pathogen, which may not be available for all pathogens of interest. The availability of extensive DNA and RNA sequence information greatly benefits most techniques for molecular detection and diagnostics of plant pathogens. PCR-based assays target sequences from the pathogen for amplification and detection, and species-specific primers have provided a powerful tool for pathogen identification. These primers usually target regions of ribosomal RNA genes exhibiting sufficient diversity among taxa, such as the internal transcribed spacer regions, ITS1 and/or ITS2, in fungi. The next step is characterization of the phytobiome via metagenomics to study the composition and expression profiles of microbial communities in and around the grapevines. While most phytobiomes change drastically in response to the environment, there appears to be a genetic basis to the phytobiome as a phenotype. Such studies can generate details on biological and metabolic processes in grapevine-microbe interactions (reviewed by Cadle-Davidson et al. 2019).
4.4.3.3 Biomarkers of Grapevine Resistance to Diseases and Pests
An upcoming non-invasive approach to phenotype biotic stress response is represented by the identification of biomarkers. These are biologically relevant compounds produced or emitted during the interactions between pathogen/pest and the host. Such compounds may be first sought among all the major groups of metabolites in grapevine leaves—phenolics, organic acids, terpenoids and lipids—upon pathogen infection or pest infestation. By far the most examined pathosystem is the grapevine-P. viticola one. Initial metabolite profiling has been replaced over time by metabolomics approaches. Stilbenes were among the first candidates as biomarkers for disease resistance (Gindro et al. 2012; Viret et al. 2018) and recently the large class of stilbenoids was confirmed to play a fundamental role at various post-pathogen inoculation times (Billet et al. 2020). Besides these phenolic compounds, some lipids have been identified as being associated with resistance to DM (Chitarrini et al. 2017; Cavaco et al. 2018; Negrel et al. 2018). Moreover, among metabolites analyzed from butanol extract, hexadecanoic and the monohydroxycarboxylic acids were related to resistance, while the more polar compounds were related to sensitivity (Batovska et al. 2009). The metabolite profiles of resistant grapevine species compared with representatives of the more susceptible cultivars gave some hints to unknown resistance biomarkers (or elicitors) as VOCs (Elfert et al. 2013). Emission of volatile sesquiterpenes and monoterpenes were also detected in grapevine genotypes upon P. viticola inoculation in vitro (Algarra Alarcon et al. 2015). Regarding pest response, chemical fingerprinting allowed the identification of flavonoids as biomarkers of early infestation of grape phylloxera (Benheim et al. 2011). In general, the field of pests is less investigated than that of pathogens, probably due to the limitations of managing experiments under controlled conditions. In addition to metabolic biomarkers, upstream biochemical levels have recently been investigated which have resulted in proteomics and transcriptomics studies. For instance, proteomic analysis of induced grapevine resistance reveals specific defence pathways activated as a response to DM (e.g. Palmieri et al. 2012; Perazzolli et al. 2016). The plethora of resistance-related transcriptomics studies started in 2010 (Polesani et al. 2010; Wu et al. 2010; see Sect. 4.11) and culminated with the characterization of tolerance to FD (Bertazzon et al. 2019). The entirety of these single ‘omic researches was prodromal to the development of an integrated approach today known as system biology.
Recently, systems biology has provided information about the interaction of genes, proteins, and metabolites through integration of ‘omic data (see Sect. 4.11). As technologies for generating ‘omic data improve in sensitivity, resolution, accuracy, depth, and speed, databases and data analysis pipelines must keep pace (Mosa et al. 2017). For instance, NGS technologies have simplified the simultaneously acquisition of transcriptome-wide expression profiles and genome-wide marker data. These achievements paved the way for the emergence of expression QTL (eQTL) studies, also applied in the dissection of biotic stress resistance traits such as Phomopsis resistance (Barba et al. 2018). In the era of system biology, biomarkers have been identified through double ‘omic approaches. In particular, the coupling of proteomic and transcriptomic data revealed physiological changes in response to esca proper and apoplexy (Spagnolo et al. 2012). Lately, the integration of transcriptomics and metabolomics data has been most widely exploited to uncover commonalities and differences between diverse R-gene-mediated resistance to P. viticola (Chitarrini et al. 2020) as well as potential biomarkers of fungal/oomycetes-associated disease susceptibility (Maia et al. 2020). However, new research frontiers are always present. By combining multiple ‘omic approaches (genomics, transcriptomics and degradomics), the same pathosystem was characterized and a specific effector was identified that triggers an immune response in a wild species. This effector is a potential biomarker to screen novel grape resistant varieties (Brilli et al. 2018).
4.5 Genetic Resources of Resistance Genes
4.5.1 Gene Pool and Gene Center
The grapevine belongs to the Vitaceae family, which is divided into different genera (19), including the Vitis genus (Aruani et al. 2015). Vitis species are distributed in three major geographic habitats between approximately 10° and 50° latitude in the northern hemisphere (Töpfer et al. 2011). The number of species is not exactly known and estimated to be 60–70 or even more (Alleweldt and Possingham 1988; Galet 1988). The two largest gene centers are North America and East Asia with about 30 and 40 species, respectively. In Eurasia, only one single species, V. vinifera subsp. sylvestris, is native from which the cultivated grapevine—V. vinifera subsp. vinifera (syn. sativa)—was domesticated. All Vitis species share the same number of chromosomes (2n = 38) and are interfertile, i. e. there is per se no sexual reproductive barrier within this genus. Natural sympatric hybridization occurs, which explains the difficulty in clearly distinguishing and botanically defining the number of Vitis species (Walker et al. 2019; Zecca et al. 2020).
For a more application-oriented characterization such as breeding, the gene pool is actually a more important concept than the species. A gene pool includes all wild species that can exchange genetic material. However, since this ability often differs between the wild species of a gene pool, Harlan and Wet (1971) refined this general concept and formed three subgroups. In the primary gene pool, genes can be easily transferred from wild species to cultivated forms by crosses; species in the secondary gene pool are more distinct but hybridizations are still possible with at least some fertile offspring; in the tertiary gene pool, crosses can be conducted only with special techniques and the progeny are often sterile. Almost all genes can be introgressed from non-related species independent of any sexual barrier with modern genetic engineering and tissue culture methods (see Sects. 4.12 and 4.13). For grapevine, a definition of primary, secondary and tertiary gene pool has not been established, as no relevant differentiation is possible. Within the Vitis genus there are no apparent mating obstacles; with species of more far related taxa no sexual hybridization is possible. The only but very important exception is M. rotundifolia (syn. V. rotundifolia), a species distributed in the southeast of North America that has 40 chromosomes (2n). Although the different number of chromosomes is a genetic barrier that prevents a natural hybridization, successful crossings were shown in the past enabling the transmission of several resistances to the Vitis gene pool (Pauquet et al. 2001; Rubio et al. 2020).
Within the gene centers of Vitis and M. rotundifolia, there are very diverse abiotic (temperature, precipitation, humidity, soil, etc.) and biotic conditions (pathogens and pests) to which the respective regionally occurring species are well adapted. The ability to cope with these stresses is based on the high genetic diversity of the grapevine wild species. In contrast, a reduced genetic diversity is found in the domesticated cultivated form.
The value of the wild species as genetic resources for grapevine breeding became evident twice in history. First, when European settlers introduced grapevine varieties to North America, where they died after encountering previously unknown pathogens and pests. Secondly, the reverse process in the middle of the eighteenth century, when these disease causal agents and pests were accidentally introduced from North America to Europe, ended with the same result of grapevine decline. Botanists and breeders recognized quickly the presence of resistances in grapevine wild species and exploited them for the improvement of cultivated varieties. For these historical reasons and the more comprehensive knowledge of North American wild grapevine species many biotic resistances from this gene pool are known today (Table 4.1).
About one and a half centuries later, mildew resistances were also described for East Asian Vitis species (e.g. Wan et al. 2007a, b; Eibach et al. 2010) and several were genetically mapped in V. amurensis, the best known and evaluated East Asian wild species (Blasi et al. 2011; Schwander et al. 2012; Venuti et al. 2013; Lin et al. 2019; Fu et al. 2020). This finding was surprising, since the grapevine mildew pathogens are native to North America and most scientists followed the concept of co-evolution of host and pathogens for the development of resistance genes.
Germplasm repositories have been regularly used to evaluate genetic resources ex situ for their breeding value. However, the success to find new resistances depends on the degree of genetic diversity and representativeness of such grapevine collections. Not all Vitis species are present in international repositories and therefore usable for breeding purposes, especially those of the East Asian gene center. Reasons for this include the less developed cultural ties to East Asia in the past compared to North America and limited accessibility today. For grapevine breeding, it is evident that some species have been more exploited than others have simply because they are easier to maintain and propagate, due to their rooting performance, their shared flowering time with crossing partners or further advantageous agronomic traits.
The assessment of grapevine wild species in their natural habitat (in situ) is very laborious and time consuming. Travelling through the natural distribution areas and collecting extensive genetic material is a promising approach to enhance and evaluate the high diversity in repositories (ex situ) (e.g. Schmid et al. 2009; Péros et al. 2021; Sargolzaei et al. 2021). In China, collection trips and systematic surveys of Vitis wild species began in the 1950s and intensified in the late 1970s. In the course of these activities, even new species were discovered (Wan et al. 2008a; Lu and Liu 2015). To save space and capacities in the germplasm repositories, core collections can be generated that consist of a minimal number of individuals with the highest degree of genetic diversity. These smaller sets of vines are easier to maintain ex situ and allow evaluation under different environmental conditions. Due to the broader genetic diversity these genetic resources are more representative and of high value for identifying new biotic stress resistances in the future.
4.5.2 Known Disease and Pest Resistances
Knowledge about genetic resistances is crucial for the development of new cultivars with a strong and durable field resistance to various diseases and pests. Breeding efforts are driven by climate change and the endeavour to reduce plant protection measures. The most effective approach to prevent the breakdown of resistance by an individual pathogen is the combination of multiple resistance (R)-loci from different genetic sources. Comprehensive research to identify R-loci against E. necator (formerly U. necator) and P. viticola over the past decades has resulted in a sufficient number of R-loci to follow a stacking (“pyramiding”) strategy in breeding (Töpfer et al. 2011; Dry et al. 2019). These loci differ in strength and probable mechanisms of the mediated resistance and in availability of adequate genetic markers for application in marker-assisted selection (MAS) (Eibach et al. 2007; Delrot et al. 2020). Moreover, efforts have been made to study resistances against biotic factors including fungi, bacteria, phytoplasmas, viruses, nematodes, insects, and mites. For some of these, varying levels of susceptibility or resistance has been described in varieties from different genetic backgrounds but also within the same species, however no R-loci have been mapped so far. For instance concerning grapevine anthracnose (E. ampelina) (Mortensen 1981; Wang et al. 1998; Kim et al. 2008; Kono et al. 2012), leaf rust (Phakopsora euvitis) (Clayton and Ridings 1970; Patil et al. 1998; Hennessy et al. 2007; Gomes et al. 2019), Eutypa dieback (e.g. Eutypa lata) (Ridgman 1991; Highet and Wicks 1998; Loschiavo et al. 2007; Sosnowski et al. 2007), Botryosphaeria dieback (Neofusicoccum spp., Diploida seriata) (Billones-Baaijens et al. 2014; Guan et al. 2016; Sosnowski et al. 2016), esca and Petri disease (P. chlamydospora, Phaeoacremonium spp.) (Marchi 2001; Feliciano et al. 2004; Fussler et al. 2008; Landi et al. 2012; Travadon et al. 2013; Murolo and Romanazzi 2014), as well as FD (Eveillard et al. 2016).
4.5.2.1 North American Gene Center
The majority of identified R-loci against diverse pathogens and pests originate from North American wild species. This is due to the good and early accessibility of plant material in large parts of the world based on strong trading activities in the nineteenth and twentieth century and subsequent breeding of interspecific hybrids. In addition, a widespread, comprehensive description and awareness of resistance behaviour of these species and the assumption that most of the pathogens and pests are native to North America combined with the theory of coevolution may have made them a popular research object.
To date, twelve R-loci conferring resistance to E. necator have been identified, eight of them from grapevine wild species native to North America. The locus Run1 was identified on chr12 of M. rotundifolia (Pauquet et al. 2001; Barker et al. 2005). The gene MrRUN1, coding for a Toll/interleukin-1 receptor (TIR)-NB-LRR protein, conferring a strong resistance via a hypersensitive response (HR) was cloned and functionally characterized (Feechan et al. 2013). Further R-loci identified in M. rotundifolia background are Run2 with two allelic variants (Run2.1 and Run2.2) on chr18 (Riaz et al. 2011) and Ren5 on chr14 (Blanc et al. 2012). Ren2 is also located on chr14 and could be traced back to V. cinerea (Dalbó et al. 2001). The Ren3 locus was identified on chr15 in the new bred variety ‘Regent’ (Welter et al. 2007). This German cultivar has a complex genetic background so that the original North American resistance donor species has not yet been identified. Later Zendler et al. (2017, 2021b) found out that the resistance in this region of chr15 is not conferred by one but actually by two adjacent loci: Ren3 and Ren9. Both loci are present in many French hybrids and have often been transmitted together in crosses due to their tight linkage. The very recently identified strong resistance Ren11 is derived from V. aestivalis and was also mapped on the same chromosome 15, but does not overlap with Ren3 and Ren9 (Karn et al. 2021). The North American wild species from which the R-loci Ren8 on chr18 (Zyprian et al. 2016) and Ren10 on chr10 (Teh et al. 2017) originate are not yet known.
Until today 31 Rpv loci have been identified, 18 of them originate from North American wild Vitis species. The Rpv1 locus from M. rotundifolia was found co-located within the same region of chr12 as Run1 (Merdinoglu et al. 2003; Anderson et al. 2011) and due to the lack of recombinant progeny between the two loci they can be considered as one Run1/Rpv1 locus (Dry et al. 2019). This locus contains seven genes coding for TIR-NB-LRR proteins like the gene MrRUN1 but also MrRPV1 conferring resistance to P. viticola (Feechan et al. 2013). A second locus from M. rotundifolia is Rpv2 on chr18 (Wiedemann-Merdinoglu et al. 2006). The loci Rpv5 on chr9, Rpv6 on chr12 (Marguerit et al. 2009), Rpv9 on chr7 and Rpv13 on chr12 (Moreira et al. 2011) originate from V. riparia. Rpv14 on chr5 was identified from the background of V. cinerea (Ochssner et al. 2016), Rpv19 on chr14 (Divilov et al. 2018) and Rpv28 on chr10 (Bhattarai et al. 2021) from V. rupestris and Rpv27 on chr18 (Sapkota et al. 2019a) from V. aestivalis. Similar to some Ren loci, there are several Rpv loci originating from North American wild species with unclear identity, namely Rpv4 on chr4 (Welter et al. 2007), Rpv7 on chr7 (Bellin et al. 2009), Rpv11 on chr5 (Fischer et al. 2004), Rpv17 on chr8, Rpv18 on chr11, Rpv20 on chr6 and Rpv21 on chr7 (Divilov et al. 2018). For the locus Rpv3 on chr18 (Welter et al. 2007; Bellin et al. 2009; van Heerden et al. 2014) seven different haplotypes were described by Di Gaspero et al. (2012) with at least four different North American wild species as sources (V. labrusca, V. lincecumii, V. riparia and V. rupestris). Three of these were validated in segregating populations and named Rpv3.1, Rpv3.2 and Rpv3.3 (Di Gaspero et al. 2012; Zyprian et al. 2016; Vezzulli et al. 2019a).
Further fungal pathogens have been the subject of resistance research like the ascomycete B. cinerea causing BBR. Sapkota et al. (2019b) were the first to report the successful mapping of a resistance locus against B. cinerea on chr2 of the V. aestivalis cultivar ‘Norton’. However, structure of the locus and resistance mechanisms remains to be elucidated. Recent studies have aimed at the identification of genetic loci underlying morphological traits for an indirect resistance to BBR like a loose bunch architecture (Correa et al. 2014; Tello et al. 2016; Richter et al. 2019). Apart from this, genetic resistances were identified for the grapevine BR disease caused by P. ampelicida. This hemibiotrophic ascomycete is native to North America like the two mildews. Dalbó et al. (2000) were the first who described the mapping of a resistance locus for BR in the breeding line Illinois 547–1, which is an interspecific offspring of the North American wild species V. rupestris and V. cinerea. Later, Rex et al. (2014) identified the loci Rgb1 on chr14 (Resistance to G. bidwellii, now P. ampelicida) and Rgb2 on chr16 in the interspecific rootstock cultivar ‘Börner’ (V. riparia ‘Gm183’ x V. cinerea ‘Arnold’). The exact origin of these R-loci is not yet clear and remains to be specified. The resistance locus of Illinois 547–1 and Rgb1 of ‘Börner’ mapped to the same genomic region, they are presumably not identical but allelic forms (Rex et al. 2014). Further wild species, almost exclusively from North America, have been described as resistant to BR, so identification and mapping of further resistance can be expected in future (Barrett 1953; Hausmann et al. 2017).
Grapevine trunk diseases are caused by several fungi from different families and comprise esca, Petri and black-foot disease as well as Eutypa, Botryosphaeria and Phomopsis dieback (Gramaje et al. 2018; Claverie et al. 2020). Phomopsis cane and leaf spot of grapevine is caused by D. ampelina. Recently, the two R-loci Rda1 and Rda2 have been described that reduce symptoms on berries and canes to a large degree. Rda2 was identified on chr7 of the complex interspecific hybrid ‘Horizon’ (V. vinifera, V. labrusca, V. aestivalis, V. rupestris). Rda1 is located on chr15 and originates from V. cinerea, it spans a cluster of five NB-LRR genes, and via candidates were narrowed down to two of these genes via transcriptome screening (Barba et al. 2018). No further R-loci against fungi within the grapevine trunk diseases complex have been reported to date.
Besides from fungal pathogens, bacteria, viruses and phytoplasmas constitute, to varying degrees, a threat to productivity and durability of vineyards (Wilcox et al. 2015). Pierce’s disease is caused by the xylem-inhabiting bacterium X. fastidiosa transferred to grapevines by various leafhoppers such as the large Homalodisca vitripennis. Xylem vessel occlusions by bacterial aggregation and formed gums and tyloses cause desiccation and death of vines within a few years (Fritschi et al. 2007; Riaz et al. 2008a). Whereas V. vinifera varieties are susceptible to X. fastidiosa, several North American wild species accessions were described as resistant (Fritschi et al. 2007; Riaz et al. 2018a). The resistance locus PdR1 was inherited from V. arizonica and identified in a mapping population based on bacterial number in stem tissue and disease symptoms of leaves and canes (Krivanek et al. 2006; Riaz et al. 2006). The resistance mechanism of PdR1 is still unknown, however a large cluster of putative LRR receptor kinase genes is located within the locus and two candidates are currently being studied further (Riaz et al. unpublished; Dry et al. 2019).
To date no Vitis species with resistances to viruses and phytoplasmas were identified (Belli et al. 2010; Oliver and Fuchs 2011; Eveillard et al. 2016; Meng et al. 2017). In addition, data on genetic resistances against nematode parasites and insects acting as virus-vectors are scarce. Resistance against the dagger nematode X. index, vectoring GFLV, has been described in several North American wild grapevine species (Kunde et al. 1968; Ollat et al. 2016). Resistance derived from V. arizonica, indicated by reduced number of root galls, was shown to be controlled by one locus on chr19 named XiR1 containing putative NB-LRR genes (Xu et al. 2008; Hwang et al. 2010). Rubio et al. (2020) recently identified the loci XiR2 (chr9), XiR3 (chr10) and XiR4 (chr18) in M. rotundifolia genetic background also using a root gall index and a nematode reproduction factor to determine the resistance behavior. So far, specification of the mediated resistance by the described loci remains undetermined. Rootstocks resistant to X. index could not be found to prevent replication of GFLV or translocation of the virus to the susceptible scion (Laimer et al. 2009; Oliver and Fuchs 2011). Resistance mediated by the XiR-loci is hypothesized to act via significant delay of GFLV transmission by the nematode to the grapevine, possibly through a reduction of nematode population and thus feeding events, allowing for sufficient production providing an economic benefit to the grower (Rubio et al. 2020; Oliver and Fuchs 2011). Moreover, several available rootstock varieties showing resistance to endoparasitic root knot nematode species descend from North American grapevine wild species (Ollat et al. 2016; Smith et al. 2018a). Recently the locus MJR1 was mapped on chr18 of a V. cinerea accession conferring resistance against M. javanica by induction of an HR in cells of the root meristem probably impairing nematode migration and giant cell formation (Smith et al. 2018b).
A major insect pest of grapevine is grape phylloxera (D. vitifoliae). To mitigate its negative impact on viticulture rootstocks, hybrid varieties with North American wild species in their background, showing certain levels of resistance are available. Rdv1 a genetic locus conferring root resistance against phylloxera via an HR was mapped on chr13 of ‘Börner’ and is assumed to originate from V. cinerea (Zhang et al. 2009; Hausmann et al. 2011). Clustered NBS-LRR-like genes in the central region of the locus were proposed as candidate genes for the Rdv1-mediated resistance (Hausmann et al. 2014). A second root resistance locus of V. cinerea was identified on chr14 and named Rdv2 (Smith et al. 2018a). A mapping approach in a cross of two breeding lines with complex pedigree (V. riparia, V. labrusca, V. rupestris, V. aestivalis, V. berlandieri, V. lincecumii) revealed three additional Rdv-loci, with the donor species remaining undetermined. Rdv3 on chr14, overlapping with the location of Rdv2, is the first locus reported to confer foliar resistance to phylloxera, Rdv4 (chr10) and Rdv5 (chr5) mediate root resistance (Clark et al. 2018). Rubio et al. (2020) recently identified the root R-loci Rdv6 (chr7), Rdv7 (chr3) and Rdv8 (chr10) from M. rotundifolia background.
4.5.2.2 East Asian Gene Center
To date, two wild East Asian Vitis species were identified as sources of R-loci against PM. Resistance mediated by the single dominant locus Ren4 on chr18 of V. romanetii was described to be associated with a reduction of the E. necator penetration success and no HR (Ramming et al. 2011; Mahanil et al. 2012). However Dry et al. (2019) report a Ren4-mediated post-penetration resistance via programmed cell death of epidermal cells or callose encasement of haustoria. The loci Ren6 on chr9 and Ren7 on chr19 originate from V. piasezkii and confer a resistance based on HR (Pap et al. 2016).
The wild species V. amurensis is the best-exploited East Asian source of resistance against DM (Table 4.1). To date eight Rpv-loci have been identified tracing back to this source. Rpv8 and Rpv12 were mapped to chr14 in approximately the same position, requiring clarification about whether two distinct genes are responsible for the strong resistance conferred by induction of an HR in both cases (Blasi et al. 2011; Venuti et al. 2013). Schwander et al. (2012) identified the Rpv10 locus on chr9 of the V. amurensis-derived cultivar ‘Solaris’. The loci Rpv23 (Fu et al. 2020), Rpv25 and Rpv26 (Lin et al. 2019) were mapped on chr15 of the half-sib individuals ‘ShuangHong’ and ‘Shuangyou’. Also described on chr2 and chr18 of ‘ShuangHong’ were the loci Rpv22 and Rpv24 (Fu et al. 2020). Apart from PM resistance, V. piasezkii was found to be a source of resistance against DM as well, with the identification of Rpv15 on chr18 and Rpv16 on chr9 (Pap et al. in preparation).
Another serious fungal disease of grapevine is ripe rot caused by Colletotrichum gloeosporioides or Colletotrichum acutatum. Few reports of resistance behaviour in varieties derived from North American wild species and even V. vinifera exist (Daykin 1984; Jang et al. 2011, 2020), however, the first resistance locus against ripe rot disease was identified in V. amurensis background. Cgr1 is located on chr14 and contains several putative resistance genes of the NBS-LRR-type (Fu et al. 2019).
The bacteria Agrobacterium/All. vitis and sporadically A. tumefaciens are known to cause crown gall disease of grapevine. Infected vines remain without symptoms until gall formation is triggered by freezing or mechanical wounding. The galls affect the vascular system and decline of water and nutrient supply can lead to reduced yield and even plant death (Kuczmog et al. 2012). Cultivars of V. vinifera are described as highly susceptible, V. labrusca and V. amurensis as resistant (De Cleene and De Ley 1976). A single dominant crown gall resistance locus Rcg1 was identified on chr15 of V. amurensis (Kuczmog et al. 2012); the mechanism conferring the resistance remains to be elucidated.
The reason that most of the known R-loci from East Asian sources were identified in varieties with V. amurensis background is the historical abundance of this material. Being native to the cold regions of the Far East V. amurensis can withstand temperatures as low as −40 °C (Wan et al. 2008b). This extreme cold-hardiness, resistance behavior against some fungal pathogens and the absence of a ‘foxy’ flavor made this species a valuable crossing partner with V. vinifera (Koleda 1975; Korbuly 2000; Wan et al. 2007b). Numerous hybrids emerged from extensive breeding efforts in the former Soviet Union in the beginning and middle of the twentieth century and in China after the end of World War II (Luo and Zang 1990; Korbuly 2000; Wan et al. 2008a). With the start of breeding programs in countries of East and Middle Europe (e.g. Hungary in the 1950s), V. amurensis and its hybrids also found their way to more western countries (Koleda 1975; Korbuly 2000).
4.5.2.3 Europe/Central Asian Gene Center
The Eurasian grapevine wild species V. v. subsp. sylvestris and its cultivated form V. v. subsp. sativa are generally described to be susceptible to numerous non-native pests and pathogens. In the nineteenth century, the introduction of E. necator and P. viticola from North America into Europe led to massive damages of local vineyards (Töpfer et al. 2011). Due to the relatively short time span that has been available for co-evolution between grapevine and pathogens since then as well as to clonal propagation, the probability to find resistances to these pathogens in V. vinifera were assumed to be small. Therefore the mapping of Ren1 (Hoffmann et al. 2008) and the putative allelic locus Ren1.2 (Possamai et al. 2021) as well as Rpv29-31 (Sargolzaei et al. 2020) in the genome of V. vinifera accessions was both very interesting and surprising.
The partial investigation of the Georgian grapevine germplasm regarding the level of resistance to P. viticola led to a small group of varieties with low disease severity values after infection with the pathogen (Bitsadze et al. 2015; Toffolatti et al. 2016). The variety displaying the most constant resistance behaviour ‘Mgaloblishvili’ was studied in detail regarding its resistance mechanism. No HR could be observed, resistance is rather associated with callose deposition, degeneration of the mycelium, altered morphology of sporangiophores and reduced production of sporangia. Transcriptome analysis showed an overexpression of genes related to pathogen recognition, the ET signalling pathway, synthesis of antimicrobial compounds and enzymes and development of structural barriers upon infection with P. viticola (Toffolatti et al. 2018, 2020). A genome-wide association (GWA) approach using 84 seedlings of an ‘Mgaloblishvili’ S1 population and 48 genotypes of the Georgian germplasm led to the identification of three R-loci against DM from the V. vinifera background: Rpv29 (chr14), Rpv30 (chr3) and Rpv31 (chr16) (Sargolzaei et al. 2020).
The Ren1 locus was identified on chr13 of the two Central Asian V. v. subsp. sativa cultivars ‘Kishmish vatkana’ and ‘Dzhandzhal kara’. Powdery mildew resistance is mediated by restriction of pathogen growth and sporulation as well as programmed cell death. The homologous region on chr13 of the ‘PN40024’ reference genome contains several NB-LRR-genes, however no specific candidate gene for Ren1 has been proposed (Hoffmann et al. 2008; Coleman et al. 2009). Riaz et al. (2013) identified a Ren1-like locus in six additional Central Asian V. vinifera but also in two V. v. subsp. sylvestris accessions displaying PM resistance. In a subsequent study, they were able to map Ren1 in the genome of V. v. subsp. sylvestris and to identify several additional resistant subsp. sylvestris accessions from five different Caucasian and Central Asian countries. Results of further genetic marker and sequence analysis of the locus suggest that Ren1-mediated resistance against E. necator of V. v. subsp. sylvestris is the progenitor of the resistance in the Central Asian V. vinifera cultivars (Riaz et al. 2020a). Very recently, a strong partial resistance in the Caucasian variety ‘Shavtsitska’ was described that co-localize with Ren1 of the Central Asian cultivar ‘Kishmish vatkana’ (Possamai et al. 2021). The resistance of ‘Shavtsitska’ seems to be widespread in Caucasian varieties, since it was also found in ‘Tskhvedianis tetra’ and 24 other varieties. Genetic analysis showed a different haplotype compared to Ren1 of ‘Kishmish vatkana’ and named it Ren1.2.
The identification of R-loci against E. necator and P. viticola in grapevine accessions with East Asian or V. vinifera background is contrary to the current theory of the North American pathogen origin and introduction to Europe and the rest of the world in the middle of the nineteenth century. Such a short period would not be sufficient for the local wild Vitis species to develop resistances against the pathogens by co-evolution. In the case of DM, some authors explain this unexpected observation with the presence of related species of P. viticola in the East Asian Vitis gene center (Jürges et al. 2009). Another attempt of explanation is the hypothesis that the pathogens were indeed present in Asia for longer than thought, possibly since before the continental split separated it from America, disregarding the total absence of historical records of PM and DM infections in Europe and Asia prior to the nineteenth century. The lack of resistance in the majority of grapevine cultivars could be explained if the focus in early breeding efforts was not on these traits and by long-standing tradition of clonal propagation (Riaz et al. 2020a).
Nevertheless, these findings suggest that present grapevine germplasms based on East Asian wild species or V. vinifera could be valuable sources for resistances to be used in breeding programs. Especially V. vinifera as a resistance source opens up the possibility to receive breeding lines without introducing unpleasant quality characteristics often passed on by North American (e.g. foxy flavor) wild species.
4.6 Glimpses on Traditional Breeding Towards Molecular Breeding
4.6.1 Traditional Breeding Objectives and Achievements
The disease resistance traditional breeding history in grapevine began in the late nineteenth century, when the import of grapevine material from America into Europe (primarily France) enabled the introduction of serious, previously absent grapevine pests and pathogens for which the native Eurasian vines (V. vinifera L.) had no resistance. In particular, the North American genotypes were used for the reason of being resistant to phylloxera and the first interspecific hybrids, mainly rootstocks, were bred to overcome the insect threat. Starting in the second half of the nineteenth century, several attempts to combine different resistance traits of American grapevines (V. riparia, V. labrusca, V. aestivalis, and V. berlandieri) with qualitative characteristics of European species were made, leading to the creation of interspecific resistant varieties. Besides the phylloxera plague, a serious invasion of fungal diseases contributed to the massive destruction of European vineyards and led to the increase in hybrid plantings. Relevant in the history of interspecific breeding are the genotypes coming from France called “first-generation hybrids”, usually meaning crossings between American species and cultivated French varieties, also called “direct producers”, namely vines grown on their own roots and used for wine making. The second-generation hybrids, basically crossings between first generation hybrids among themselves or with cultivated European varieties, came out later in the century and they contain a higher percentage of the V. vinifera genome, thereby increasing the quality of the wine (reviewed by Zini et al. 2019) (see Sect. 4.7.5).
Nowadays, around the world a myriad of expanded and minute cross breeding programs are run with traditional (or classical) methods by governmental institutions, universities, private companies as well as private growers. They mainly address three purposes: wine grapes, table grapes, and rootstocks. Minor although important and increasing uses are raisins and juice grapes (OIV 2020). Given all by-products known and emergent of grape growing and winemaking, we cannot exclude that, in the future, ad hoc breeding programs for the production of novel (niche) products will be undertaken. The breeding goals and selection achievements in agreement with the different final purposes intended for the grapevine have been recently summarized by Delrot et al. (2020). Among them, it is relevant to underline that the goals of (mainly) pathogen resistance coupled with yield and fair quality in wine grape varieties as well as (mainly) pest resistance coupled with relevant agronomical trait and/or abiotic stress resistance for rootstocks have been reached. These objectives have been achieved through both positive (e.g. high sugar, balanced acidity, flavour etc.) and negative (e.g. off-flavours) selection of traits (Töpfer et al. 2011).
Given the level of knowledge now achieved, it is important to underline that some common morphological and/or organoleptic characteristics may have a different, and sometimes new, interpretation as (integrative) indirect traits associated with resistance to biotic stress. For instance, part of leaf or berry resistance to pathogens may also be linked to leaf morphology or bunch architecture. Leaf hairs can influence pathogen infections, acting as a physical barrier or affecting the leaf micro-environmental conditions (Niks and Rubiales 2002). While a role of trichome density was often proposed in repelling water and thus favouring resistance to P. viticola (Staudt and Kassemeyer 1995; Kortekamp et al. 1998; Kono et al. 2018), lately a possible effected of prostrate hairs on the foliar resistance also to E. necator has been assumed (Possamai et al. 2021). The abundance of predatory phytoseiid mites, biological agents that control both spider mite pests as well as PM, can be influenced by breeding for tuft-form domatia in leaf vein axils (Barba et al. 2019). Moreover, loose clusters promote air exchanges, modifying the bunch microclimate and reducing incidence and severity of fungal diseases such as gray mold. Actually, the traditional breeding program carried out at FEM (Italy) has recently released cultivars bearing looser clusters with higher tolerance to gray mould (CIVIT, https://www.civit.tn.it/). Besides cluster looseness, berry skin thickness has long been suspected as relevant in promoting resistance to biotic stress; to date its role is controversial and unconfirmed in the literature. By contrast, berry skin and leaf composition seem to be more promising, as suggested by a very recent study providing for the basis for using cell wall traits in crop breeding programs (Molina et al. 2021). Finally, biomarker traits such as stilbenoid (and other phenolic compound) production as well as VOC emission are interesting candidates in the context of innovative resistance breeding (see Sect. 4.4). Some of these cited indirect traits can first be pursued in a cross breeding program and subsequently can be refined and stabilized during the clonal selection process, usually performed within existing cultivars in order to keep it phytosanitarily healthy and morphologically stable.
4.6.2 Limitations of Traditional Breeding and Rationale for Molecular Breeding
In spite of their achievements through the centuries, traditional cross breeding has some limitations. Crop peculiarities such as long reproductive cycle, large plant size, and expanded evaluation period do not facilitate these processes. At the beginning of the twenty-first century, as new tools were developed to genetically dissect traits, molecular approaches were introduced into breeding research starting the era of molecular breeding. In particular, the advent of molecular markers, genetic mapping and genotype–phenotype association analysis (see Sects. 4.7 and 4.8) initiated a paradigm change in grapevine genetic improvement from a pure empirical work to the strictly goal oriented design of crosses and of gene management (Töpfer et al. 2011).
Some tools and advantages of molecular breeding have already become popular and are no longer dispensable in grapevines. Markers linked to disease-resistance genes are currently utilised for discarding susceptible seedlings at the initial stage of development in large-scale breeding programs. Grapevine molecular breeding is now facing and undertaking the next step forward, from MAB to GAB (see Sect. 4.9). A genomics approach to crop breeding requires full knowledge of the association between DNA variation in gene sequence and phenotypic variation in the trait across the extant germplasm (the discovery of natural allelic variation), not just linking a gene to a segregating phenotype (reviewed by Di Gaspero and Cattonaro 2010). In the last fifteen years, genome sequencing projects expedited and enhanced this process, giving insights about trait/gene localization and molecular organization. After the first deciphering of the grapevine genome (‘PN40024’, Jaillon et al. 2007; ‘Pinot noir’, Velasco et al. 2007), nowadays, an increasing number of V. vinifera whole genome sequences were generated and freely available. A few hybrid and non-vinifera species genomes have been also decoded (see Sect. 4.11.1). Advances in automated technology enable a new approach in MAB: “Breeding by Design” (Peleman et al. 2005). The advances in applied genomics and the possibility of generating large-scale marker datasets provide us with the tools to determine the genetic basis for all traits of agronomical importance. Methods for assessing the allelic variation at these agronomically important loci are also now available. This combined knowledge will eventually allow the breeder to combine favorable alleles at all these loci in a controlled manner, leading to superior varieties.
Both based on crossing, traditional breeding and MAB lie under the hat of the well-established conventional breeding technologies (CBTs), aiming at the creation of new cultivars. This goal has been pursued in the last 20 years also through the adoption of argued methods of genetic transformation (ETGMs, see Sect. 4.12). Changing concepts and technological approaches have recently provided opportunities to develop rational and refined molecular breeding strategies. Today, the last frontier of grapevine genetic improvement consists of the debated new breeding technologies, allowing the development of new clones by editing already existing varieties (NBTs, see Sect. 4.13) (EU Explanatory Note 2017).
4.7 Brief on the Host Genetic Diversity
4.7.1 Genetic Diversity Analysis in Grapevine Based on Molecular Markers
Genetic diversity of grapevine germplasm has been extensively investigated first with morphologic descriptors (i.e. “classical” ampelography) and then with different molecular markers, of which microsatellites (or simple sequence repeats, SSRs) and single nucleotide polymorphisms (SNPs) are the most common ones. Since the milestone research papers published in the 1990’s (Thomas and Scott 1993; Bowers et al. 1996; Sefc et al. 1999), more than 400 nuclear SSR (nSSR) markers have been developed and tested by the scientific community on grapevine. SSRs are suitable for many purposes, among them identifying grapevine varieties, performing parentage analyses, studying population structure (e.g. Merdinoglu et al. 2005; Zyprian and Töpfer 2005; Cipriani et al. 2008). Chloroplast SSR markers are also available for grapevine genotyping for chlorotype and maternal lines determination (e.g. Arroyo-García et al. 2006; Riaz et al. 2019). Nine nSSRs were proposed as a standard set for grapevine genotyping and identification (This et al. 2004; Maul et al. 2012). This choice has proved strategic, given the great conservation of microsatellite loci within the Vitis genus (Di Gaspero et al. 2000); so, the comparison of grapevine germplasm was speeded up simply searching for matching molecular profiles, i.e., comparing strings of few numbers. This information allowed for the evaluation of the real extent of grapevine genetic resources. Nowadays, an increasing number of SSR profiles are available in the literature and in huge public molecular databases, such as the Vitis International Variety Catalogue (VIVC, www.vivc.de), presently hosting more than 5,700 genetic profiles. SSRs are highly polymorphic, co-dominant, can be multiplexed, but require accuracy in allele calling. Moreover, two of the nine used as the standard set for identification showed to be less suitable for non-vinifera germplasm (Migliaro et al. 2019).
Careful ampelographic description of vines combined with identification based on SSR markers allowed a more accurate definition of what a grapevine cultivar is, and according to Boursiquot and This (1999) we consider as belonging to the same variety (cépage) all grapevine plants derived from the same single seedling after vegetative propagation. Therefore, SSR markers assisted in distinguishing cultivar phenotypic diversity, as based on somatic variation or on genetic recombination through sexual reproduction. Indeed, SSRs are not useful to distinguish somatic variants (e.g. Vargas et al. 2007; Giannetto et al. 2008; Crespan et al. 2016), that is to say clones and/or mutant cultivars within a variety, while they distinguish very well the varieties derived from sexual reproduction, both by crossing or by self-fertilization. For instance, ‘Garnacha rosada’ and ‘Garnacha blanca’ (two cultivars resulting in mutations from the cépage ‘Garnacha tinta’) have the same SSR profile, but ‘Caladoc’ (resulting from a cross between ‘Garnacha tinta’ and ‘Cot’) has a clearly different one.
Resequencing provided hundreds of thousands of SNP markers in all the Vitis spp. genomes. After the first sets of 48 and 332 SNPs obtained from V. v. subsp. sativa and subsp. sylvestris (Lijavetzky et al. 2007; Cabezas et al. 2011), larger chip arrays were developed and are increasingly being used for grapevine genotyping. The currently available SNP-chip arrays encompass: (i) the Vitis9kSNP (Myles et al. 2010), having 9,000 SNPs detected in 11 V. vinifera varieties and six wild Vitis species, and (ii) the Vitis18kSNP, with around 18,000 SNPs obtained from 47 V. vinifera varieties, 12 wild Vitis species and 5 M. rotundifolia varieties (Le Paslier et al. 2013); 37 k SNPs were also discovered by Marrano et al. (2017) by analyzing 51 V. v. subsp. sativa cultivars and 44 V. v. subsp. sylvestris vines. SNPs low polymorphism is counterbalanced by their number and qualitative information, not requiring reference varieties for allele calling; they are also high throughput, co-dominant and cost effective, even if the chip arrays are not manageable in molecular labs with ordinary equipment, and costs are unsustainable for one or few samples genotyping.
4.7.2 Relevance of Germplasm Characterization and Conservation: Extent of Genetic Diversity Within Vitis
Information on grapevine genetic diversity is scattered in hundreds of scientific papers, the most comprehensive ones being based on large grapevine collections analyses, encompassing hundreds or even thousands of genotypes. Evaluations performed at the beginning of the 1990ies estimated about 10,000 sativa varieties around the world, but This et al. (2006) halved this number to 5,000 based on extensive DNA profiling results. Active searching of local and relict varieties in old (sometimes prephylloxera) vineyards in many countries is increasing this number. The massive use of genotyping with molecular markers has gained a lot of new information on homonym and synonym ambiguities, both previously hypothesized by ampelographers or unknown. This knowledge is important to join and compare the information available for single varieties, even if disseminated under different names. A good example is the case of ‘Malvasia delle Lipari’ and its synonyms: this ancient cultivar represents one of the oldest known Malvasias, scattered all over the Mediterranean area, thousands of kilometers away, along sea routes, from Dubrovnik to Eolie, Ischia and Sardinia islands, to Spain, Balearic Islands, and even to the Canaries and Madeira (Crespan et al. 2006).
Currently, cultivated grapevine genetic diversity is mostly confined to germplasm collections (This et al. 2006). Assessing the genetic diversity and structure of grapevine germplasm supports the management of collections and the exploitation of genetic resources. A strategic approach facilitating the study of complex traits on a large number of genotypes is the establishment of a genetic core collection, i.e., a subsample containing the minimum number of individuals representing the whole genetic diversity of the original collection (Brown 1995). Some genetic core collections have been established, mainly for V. v. subsp. sativa (Cipriani et al. 2008; Le Cunff et al. 2008; Emanuelli et al. 2013), and for non-vinifera (Migliaro et al. 2019).
Despite the widespread family structure in the sativa subspecies (see below), an excess of heterozygosity was found, with observed values higher than expected (Cipriani et al. 2010; Bacilieri et al. 2013; De Lorenzis et al. 2015; Marrano et al. 2017; Laucou et al. 2018). This finding is explainable with a heavy genetic load (presence of deleterious alleles) already recognized by Olmo (1976) and confirmed by Zhou et al. (2017), as well as with events of migration or balancing selection (Marrano et al. 2017).
Sylvestris subspecies currently shows the lowest genetic diversity, probably due to heavy erosion linked to human activities. Moreover, a clear genetic distance was found between the Western European and the Caucasian sylvestris (Riaz et al. 2018b). Interestingly, genetic diversity studies suggested that gene flow between sylvestris and sativa was not unidirectional and still continues, since several cases of introgression of sylvestris into cultivated germplasm have been evidenced, as well as of recent sylvestris vine selection for cultivation (e.g. Di Vecchi-Staraz et al. 2009; De Lorenzis et al. 2015; Cunha et al. 2020; D’Onofrio 2020; Maraš et al. 2020).
Rootstocks (derived from simple American interspecific Vitis crosses) were found more diverse than hybrids (coming from complex crosses between American Vitis spp. and Eurasian V. vinifera). Both types are in turn more diverse than V. v. subsp. sativa, which is more diverse than V. v. subsp. sylvestris (e.g. Laucou et al. 2011; Emanuelli et al. 2013; Nicolas et al. 2016). A clear distinction was found between V. vinifera and non-vinifera germplasm (other Vitis species, rootstocks and fruiting hybrids) as well as a larger genetic diversity in allelic richness and heterozygosity in the wild Vitis than in the cultivated one (Crespan et al. 2009; Myles et al. 2010; Laucou et al. 2011; Emanuelli et al. 2013; Bianchi et al. 2020). The larger genetic diversity in non-vinifera germplasm (dozens of American and Asian species) than in vinifera (one Eurasian species) is shown also by chlorotype numbers, being eight those found by Arroyo-García et al. (2006) analyzing 513 V. v. subsp. sativa and 688 V. v. subsp. sylvestris samples, and 63 those detected by Riaz et al. (2019) analyzing 140 non-vinifera samples.
4.7.3 Genetic Structure Analysis: Relationships with Geographical Distribution and Use
Several studies with different molecular markers using large amounts of genotypes have shown that the diversity contained in V. v. subsp. sativa (that includes most of the cultivated varieties) is structured. And this genetic structure is mainly related, as the pioneer Negrul (1946) suggested, to berry morphology and use (wine/table) and geographical origin (Arroyo-García et al. 2006; Myles et al. 2011; Bacilieri et al. 2013; Emanuelli et al. 2013; Laucou et al. 2018). Excluding grapes obtained from breeding, a first division into three/four genetic groups was generally found: (i) East table grapes (from Eastern Mediterranean, Caucasus, Middle and Far East countries), showing the highest genetic diversity, according to the hypothesis of Caucasus and Fertile Crescent as the cradle of grapevine domestication; (ii) West wine grapes (from West and Central Europe), the second most diverse, resulting from introgression from local wilds; (iii) East wine grapes (from the Balkans and East Europe), with further down diversity; (iv) wine and table grapes from Iberian peninsula and Maghreb (Aradhya et al. 2003; Bacilieri et al. 2013; Emanuelli et al. 2013; Laucou et al. 2018). A second division level was found into 8–16 subgroups corresponding mainly to regional and/or grape use stratification (Aradhya et al. 2003; Bacilieri et al. 2013; Laucou et al. 2018).
Genetic structure studies show V. v. subsp. sylvestris as a close genetic group to V. v. sativa, as expected since the accepted hypothesis for the origin of the cultivated grapes is the domestication of V. v. sylvestris (Grassi and Arroyo-García 2020). It is common to find genetically intermediate genotypes both in individuals sampled as cultivated and as wild vines, and even several authors found cultivars genetically closest to V. v. sylvestris than to V. v. sativa (De Andrés et al. 2012; Emanuelli et al. 2013; Ekhvaia et al. 2014; De Lorenzis et al. 2015; Riaz et al. 2018b; Cunha et al. 2020; D’Onofrio 2020; Maraš et al. 2020; Zdunić et al. 2020). Some of these cultivars, like ‘Savagnin’ (syn. ‘Traminer’) or ‘Cabernet Franc’, are representatives of viticulture from ancient times; in other cases introgressions are evidenced of sylvestris on sativa at local scale, like Amaral in Portugal (Cunha et al. 2020), ‘Petit Verdot’ in France or ‘Greco Bianco’ in Italy (D’Onofrio 2020), showing the existence of genetic flow between the two subspecies in the past. This genetic flow is still active in some places, like Montenegro, where domestication-like processes are ongoing (Maraš et al. 2020). Genetic flow from subspecies sativa to sylvestris is also present, as demonstrated by the high number of plants growing wild, supposed to be sylvestris because of their habitat, flower sex and leaf morphology, that are genetically intermediate between sylvestris and sativa. This introgression is getting worse the situation of the sylvestris subspecies, already in risk of extinction in many regions, and could prevent its potential use for the grapevine improvement in some traits of interest, in particular it has been reported a higher tolerance of sylvestris vines to some diseases, compared to sativa (Guan et al. 2016) (see Sect. 4.5.2.3).
In wider studies including several species of genus Vitis, V. v. subsp. sativa and sylvestris cluster together and separate from other species, as expected according to their inclusion within the same species (Aradhya et al. 2013; Emanuelli et al. 2013), but there are some exceptions. Using data from four chloroplast spacers, Zecca et al. (2012) found V. v. subsp. sylvestris to be sister to the Asian Vitis species. A similar result was found using SNP genotyping data from the Vitis9kSNP array, although this was attributed to the ascertainment bias inherent in the genotype calls; V. v. subsp. sativa and sylvestris clustered together when using quantitative genotypes from the hybridization intensities to estimate genetic distances among species (Miller et al. 2013). In a whole-genome resequencing survey, the first stratification level separated V. v. subsp. sativa from V. v. subsp. sylvestris and all studied North American and Asian Vitis species (Liang et al. 2019). More studies are needed to understand the meaning of such an unexpected phylogenetics result. Recent studies using molecular markers have also shed light on the origin and relationships within the Vitis genus, particularly subgenus Vitis (formerly Euvitis), which was easily distinguished from subgenus Muscadinia (e.g. M. rotundifolia). Although previous studies supported the existence of separated origins for the North American and the Eurasian species (Miller et al. 2013), the monophyletic hypothesis for the Vitis genus is more spread, with some data indicating that subgenus Vitis originated in Asia (Tröndle et al. 2010; Peros et al. 2011), while other surveys indicate that subgenus Vitis was originated in the New World from where it migrated to the Old World, Eurasia (Zecca et al. 2012; Wan et al. 2013; Liu et al. 2016a; Ma et al. 2018). Most studies, including recent ones based on genomics analyses, distinguish between three main groups consistent with geography: North America, East Asia and Europe, with the European V. vinifera closest to the Asian species (Peros et al. 2011; Zecca et al. 2012; Liu et al. 2016a; Klein et al. 2018; Ma et al. 2018; Liang et al. 2019). Vitis species from North America includes two or three sub-clades, which can be roughly related to their geographical dispersion (Wan et al. 2013; Miller et al. 2013; Klein et al. 2018; Ma et al. 2018; Riaz et al. 2020b), while the use of ca. 4000 SNPs allowed to confirm the existence of the hybrid species previously defined: V. × champinii, V. × doaniana, V. × novae-angliae, and V. × slavinii (Zecca et al. 2020). Very recent studies on North American Vitis species found that this molecular diversity between geographical origins also reflects variation for resistance: Riaz et al. (2020b) found an east–west genetic border that divides North American Vitis species and that mirrors their resistance to PD, strongest in the southwestern species; Péros et al. (2021) found large genetic differences between V. aestivalis, V. cinerea and V. riparia, and, interestingly, variation for DM resistance, both within and between species.
4.7.4 Pedigree Studies: Relationships Between Vinifera Varieties
The basal level for the genetic stratification of grapevine varieties is provided by the kinship relationships. The widest pedigree studies within V. v. subsp. sativa have shown that most of the varieties are related to at least one other variety by a first-degree relationship: from 58.5% in a study with 783 varieties (unique genotypes) (Laucou et al. 2018) to 74.8% in a study with 583 varieties (Myles et al. 2011), up to 88% in a study of 2344 varieties from INRAE ‘Domaine de Vassal’ Grape Germplasm Repository (France) (Lacombe et al. 2013). These genetic materials include both traditional and “modern” (obtained by breeding) cultivars, the latest mostly being table grapes. The existence of those abundant first-degree relationships for traditional varieties confirms the long-term cultivation of many varieties, such as recently proven for ‘Savagnin/Traminer’, found in an archaeological sample dated in 1050–1200 CE (Ramos-Madrigal et al. 2019). On the other hand, the true knowledge of the pedigrees of many bred cultivars and their quality traits is useful for future breeding programs in the selection of the most appropriate cultivars according to their performance as genitors.
When considering only traditional varieties, pedigree studies at a regional or local scale may contribute to clarify their genetic origins and relationships. National and regional pedigree studies have been done in many countries and have allowed to discover the origins of iconic varieties like ‘Cabernet-Sauvignon’ (Bowers and Meredith 1997), ‘Chardonnay’ (Bowers et al. 1999), ‘Merlot’ (Boursiquot et al. 2009), or ‘Tempranillo’ (Ibáñez et al. 2012). These are only representative examples present in very intricate pedigree networks with tens or hundreds of varieties like that recently published for Italian varieties (D’Onofrio et al. 2021). The complex regional pedigree structures become even more complex when the pedigree analyses are done in a global framework, including foreign varieties that, although have their geographic origin attributed to another place, could be considered “traditional” in other locations. That is the case of some ancient varieties which travelled through the Mediterranean basin like ‘Savagnin/Traminer’, ‘Malvasia Aromática’ (delle Lipari, di Sardegna, Dubrovacka), or ‘Heptakilo’ (a parent of Muscat of Alexandria). These varieties have contributed to the local diversification of grape varieties in certain regions, such as ‘Savagnin/Traminer’ in Portugal (Cunha et al. 2020), the Turkish Parmak Cerven in Montenegro (Maraš et al. 2020) or the Austrian Vulpea in northeastern Italy (Crespan et al. 2020).
A very common characteristic found at local, regional, and/or national scale in the European viticulture is the existence of varietal families, in many cases created around founder cultivars, varieties whose offspring were recurrently selected and multiplied to become new varieties. The VIVC (www.vivc.de, accessed March 17, 2021) contained 182 varieties (Prime names), each one involved in more than 20 known pedigrees (more than 12,000 summing up all of them). These data include mostly bred varieties, and also some varieties from non-vinifera Vitis species and Vitis interspecific hybrids, but also a number of traditional varieties that acted as founders in different regions, as ‘Gouais blanc’, ‘Cabernet franc’, and ‘Pinot’ in France (Bowers et al. 1999; Lacombe et al. 2013); ‘Heben’, ‘Traminer’, ‘Alfrocheiro’, ‘Marufo’, and ‘Cayetana blanca’ in the Iberian Peninsula (Lacombe et al. 2013; Cunha et al. 2015, 2020; Zinelabidine et al. 2015); ‘Sangiovese’, ‘Visparola’, ‘Garganega’, ‘Bombino bianco’, and ‘Refosco Nostrano’ in Italy (Crespan et al. 2020; D’Onofrio et al. 2021); or ‘Kratošija’ in the Western Balkans (Maraš et al. 2020). Even in the New World, two traditional Mediterranean varieties such as ‘Muscat of Alexandria’ and ‘Listan Prieto’ gave place to the “Criollas” varieties group, representative of the Latin American viticulture (Aliquó et al. 2017). One pending question is whether or not a regional wine typicity may be related to the existence of these varietal families.
4.7.5 Hybrids Between Vitis Vinifera and Other Vitis Species
Phylloxera was introduced to Europe in 1863 where it had dramatic consequences and destroyed many vineyards. In France alone, one third of the total vineyard area was destroyed, and wine production plummeted from 85 million hl in 1875 to 23 million hl in 1889, a 73% decrease (Meloni and Swinnen 2013). That crisis promoted, during the first half of the twentieth century, an active development of wine grape cultivars resistant to pests and diseases by hybridizing V. vinifera with many American wild species (called “French hybrids”). Even earlier, hybrids were generated in North America by crossing local Vitis species with V. vinifera (American hybrids). The resultant hybrids were massively planted but, because they produced wine of low quality, they were gradually discarded and breeding was discouraged in many places (Bouquet 2011). In fact, the low quality of the wines produced by this first generation of hybrids prompted the development of specific laws that banned most (not all) hybrids, first in France and later in the whole EU. Currently, hybrids are outlawed from the highest quality level category throughout the EU (Meloni and Swinnen 2013), whereas outside of the EU there are few restrictions, if any, on what cultivars may be planted (see Sect. 4.14). At a global level, the cultivated surface of these hybrids is not very relevant, with the exception of the table grape Kyoho, the most cultivated variety in the world, with about 365,000 ha in China (OIV 2017). For wine production, the most planted hybrid variety in 2016 was Isabella (cultivar forbidden in the EU) with about 18,000 ha (mostly in Brazil), followed by Concord, 10,000 ha mostly in USA (Anderson and Nelgen 2020).
The situation started to change in the twenty-first century in many places, including the EU, for two kinds of reasons. On the one hand, environmental, health and cost concerns regarding the use of chemicals (mostly fungicides) in the vineyard; on the other hand, the latest generation of new bred varieties, bearing more pest and disease tolerance genes and able to produce wine of quality, as exemplified by Regent. This variety was released in the 1990 decade and originated from a seed produced in Germany 30 years earlier from a cross between the V. vinifera cv. ‘Diana’ and the French hybrid ‘Chambourcin’. Thus, although part of ‘Regent’ is non-vinifera (conferring resistance to PM and DM), most of its genome is vinifera that has been estimated at about 68% (Migicovsky et al. 2016). In 1996, Regent was approved in Germany for quality wine production. Later, the EU started to relax regulations prohibiting hybrids and currently there is a Proposal for a Regulation of the European Parliament and of the Council, amending previous Regulations, “to enable producers to use vine varieties that are better adapted to changing climatic conditions and with higher resistance to diseases, provision should be made permitting products using designations of origin not only from vine varieties belonging to V. vinifera but also from vine varieties stemming from a cross between V. vinifera and other species of the genus Vitis” (COM (2018) 394 final/2).
Many different hybrids have been released to the market in the past ten years, accompanied by public or private promotion, like the international working group for the promotion of fungus-resistant grape varieties called “PIWI International” (see Sect. 4.1). There are already fungus-resistant grape varieties available that originated from and were registered in Germany, Italy, France, Austria, Hungary or Czech Republic, and more breeding programs with that goal are progressing. Beyond the EU, hybrid scion cultivars have been released in North and South America as well as in Asia. All the signals point to these hybrid varieties as being an important part of the future of viticulture and thus it would be important to think and debate on different aspects related to this issue. For instance, the appropriateness of naming the new varieties using compound names containing the name of the vinifera “noble” parent with some additional words, or the convenience of the use of founder cultivars or local varieties in the breeding programs to help keep regional wine typicity in certain cases.
4.8 Brief Account of Molecular Mapping of Resistance Genes and QTLs
4.8.1 Exploring the Genetic Architecture of Grapevine Disease and Pest Resistance Traits via Molecular Mapping
Understanding the complex mechanisms of plant-pathogen interaction is fundamental to exploit resistant traits and to achieve a durable resistance, which is particularly important for a perennial crop such as grapevine (Merdinoglu et al. 2018). The identification of grapevine genomic loci associated with complex phenotypic traits has been enabled by the development of QTL mapping approaches, which provided relevant information such as the number, location, effect and identity of the loci involved in grapevine response to relevant disease agents. The use of high-throughput genotyping tools, together with large segregating progenies, increases the resolution of QTL confidence intervals, which facilitate the identification of candidate resistance genes. As recently reviewed by Vezzulli et al. (2019b), the identification of QTLs for disease resistance has been a highly relevant topic in grapevine research in the last two decades, with DM and PM being the two diseases most commonly analyzed by QTL mapping. This approach is mostly performed using segregating biparental populations, whose parental genotypes differ with regard to the trait of interest (Würschum 2012). The cultivated European V. vinifera varieties are highly appreciated worldwide due to their high fruit quality, but they are susceptible to the majority of diseases caused by pathogens and pests (Armijo et al. 2016; Wilcox et al. 2015). On the contrary, resistance traits can be often found in other North American and Asian grapevine species, which are resistant to fungi and insects, and tolerant to viruses and some bacteria adapted through different mechanisms by centuries of coevolution (Armijo et al. 2016; Tello and Forneck 2019). Therefore, biparental populations to map disease resistance traits have been commonly generated through the cross between a susceptible V. vinifera parental genotype and a resistant (or tolerant) non-vinifera parental genotype, or with breeding material that has already a certain level of resistance. In fact, crossing susceptible V. vinifera accessions with resistant non-vinifera accessions to introduce resistance traits in V. vinifera has been performed since the arrival of PM and DM in Europe (Buonassisi et al. 2017). In contrast, most rootstocks created to overcome the devastating effects grape phylloxera over 100 years ago are still used (Migliaro et al. 2019; Riaz et al. 2019).
To date, loci associated to resistance to Agrobacterium/Allorizhobium species (Rcg1), C. gloeosporioides (Cgr1), Coniothyrium diplodiella, D. ampelina (Rda1-2), E. necator (Ren1-19, Run1, 2.1, 2.2 and Sen1), P. ampelicida (Rgb1-2), X. fastidiosa (Pdr1), P. viticola (Rpv1-31), D. vitifoliae (Rdv1-8), M. javanica (MjR1) and X. index (XiR1-4) have been reported by QTL mapping, which were found to be spread over almost all the 19 grapevine chromosomes. GWA approaches have been adopted to explore the genetic architecture of grapevine response against the following disease agents: Coniella diplodiella, Colleotrichum acutatum and C. gloeosporiodes, and P. viticola. The main QTLs reported for grapevine resistance to pests and pathogens before 2019 have been recently reviewed (see Merdinoglu et al. 2018, Vezzulli et al. 2019b and Dry et al. 2019) and new findings, especially concerning the V. vinifera species, are reported in the following sections.
4.8.2 Genetic Architecture of Disease and Pest Resistance Traits
Thanks to natural selection, plants are resistant to most pathogens and pests (Hammond-Kosack and Jones 2015). Indeed, plant co-evolution with pests and pathogens started early before crop plant domestication and led to the development of an immune system. This was first described in the twentieth century by Flor (1942), who proposed a gene-for-gene model that was further disentangled in the zig-zag model by Jones and Dangl (2006). According to the gene-for-gene model, for each gene conditioning the plant reaction (resistance gene or R-gene), there is a specific gene conditioning pathogenicity in the parasite (avirulence gene or Avr-gene). In particular, plant resistance only occurs when a plant possesses a dominant R-gene and the pathogen expresses the complementary Avr-gene (Hammond-Kosack and Jones 2015). The interaction between the products of R and Avr genes leads to pathogen recognition and triggers the plant resistance response. In this model, resistance is inherited as a monogenic trait following the laws of classical Mendelian genetics (Keller et al. 2000). The plant immune system summarized in the zig-zag model is based on two levels of resistance, triggered by the recognition by the host of different pathogen features. In the first level, named pattern-triggered immunity (PTI), the host cell recognizes pathogen-, microbe- or damage-associated molecular patterns (PAMPs, MAMPs or DAMPs) through pattern recognition receptors (PRRs) that are located on the cell surface. PTI mounts a basal immunity response that limits pathogen colonization through a complex signaling network (Bigeard et al. 2015; Hou et al. 2019; Zhang and Zhou 2010; Zipfel 2008). The plant response includes the synthesis of reactive oxygen species (ROS) with a toxic action against the invader, and the apposition of physical barriers, such as callose appositions (Boller and Felix 2009). The second immunity level (effector-triggered immunity, ETI) is activated when the pathogen produces effector molecules (encoded by the Avr genes described above) that are recognized by host-specific receptors (encoded by R-genes). R-proteins are in most cases intracellular receptor proteins of the NB-LRR type. ETI induces a more rapid and intense action against the pathogen than PTI, and often leads to the HR, a rapid localized cell death that occurs at the point of pathogen penetration (Balint-Kurti 2019). PTI and ETI mechanisms can be associated with quantitative and qualitative (gene-for-gene) resistance, respectively. Qualitative disease resistance is typically controlled by one (in monogenic resistance) or few (in oligogenic resistance) major resistance genes and leads to the absence of disease (St.Clair 2010). This type of resistance causes a strong directional selection on the pathogen population, by favoring the individuals that possess mutations in the Avr-gene that prevent pathogen recognition and the resistance response onset, leading to resistance breakdown (McDonald and Linde 2002). In contrast, polygenic resistance (quantitative disease resistance) is determined by many genes and leads to reduction in the disease, not in its complete absence (St.Clair 2010). Quantitative disease resistance is more often associated with a durable resistance (Mundt 2014; St.Clair 2010), even if cases of durable resistance associated with monogenic traits have been reported, such as in the grapevine-E. necator pathosystem (Pessina et al. 2016). Among the recently reviewed PAMP recognized by grapevine (Héloir et al. 2019), the fungal chitin, chitosan, and β-1,3 glucans (laminarin) (Brulé et al. 2019; Héloir et al. 2019), and the bacterial flagellin and rhamnolipids (Varnier et al. 2009; Trdá et al. 2014) are reported.
Susceptibility genes (S-genes) are necessary for the onset of pathogenesis by facilitating infection and supporting compatibility with the pathogen (Van Schie and Takken 2014), and their knockdown can be a strategy for achieving a durable disease resistance (Pavan et al. 2010). The most known susceptibility gene, whose knockdown associates with resistance to PM in grapevine, is the mlo gene (Pessina et al. 2016). Recently, two genes that are putatively involved in susceptibility to the DM agent in V. vinifera have been reported (Pirrello et al. 2021; Toffolatti et al. 2020), highlighting the increasing interest of research in this aspect of resistance mechanisms.
As summarized here, plant resistance to pathogens and pests is determined by a complex network of genes involved in recognition, signal transduction and defense mechanisms. Gene expression analyses and proteomic studies provide a precise picture of the genes that are involved in that response (Buonassisi et al. 2017), but the most effective approach to dissect the genetic basis of polygenic forms of resistance is still via QTL mapping (Young 1996).
4.8.3 QTL Mapping for Disease and Pest Resistances: An Update
During the last years, several QTLs related to grapevine disease resistance against both pathogens and pests have been identified in biparental populations. As stated above, three recent reviews provide an extensive overview of the resistance QTLs detected for multiple disease agents until 2019. As an update, details on those identified from 2019 to date are indicated in Table 4.2, whilst for a constantly updated list, the reader is referred to the VIVC website (https://www.vivc.de/loci).
Colletotrichum gloeosporioides is the causal agent of grape ripe rot disease. The solely QTL reported for resistance to this pathogen (Cgr1) has been recently identified in a F1 biparental population of V. vinifera cv. ‘Cabernet-Sauvignon’ (susceptible) × V. amurensis cv. ‘Shuang Hong’ (resistant), using 934 SNPs for genetic map construction. This QTL is located on chromosome 14 and explains ca. 20% of phenotypic variance. The detailed analysis of the Cgr1 region identified 11 genes with NBS and/or LRR domains, arising as interesting candidate genes responsible for grapevine resistance against this disease agent (Fu et al. 2019). Besides, Su et al. (2021) recently explored the genetic basis of grape white rot disease caused by C. diplodiella in a progeny of 177 individuals obtained from the cross of a resistant (V. vinifera × V. labrusca cv. ‘Zhuosexiang’) and a susceptible (V. vinifera cv. ‘Victoria’) cultivar, using 6,249 SNPs obtained by restriction site-associated DNA sequencing (RAD-seq) for genetic map construction. QTL mapping detected one stable QTL of minor effect (ca. 13% of phenotypic variance) on chromosome 14, which carries a series of R-genes containing NB and LRR domains (Table 4.2).
As indicated before, the genetic basis of DM resistance is a relevant topic in grapevine research (Buonassisi et al. 2017; Vezzulli et al. 2019b). Only in the last two years, seven novel Rpv loci (Rpv22–Rpv27) have been identified by QTL mapping (Lin et al. 2019; Sapkota et al. 2019a; Fu et al. 2020; Bhattarai et al. 2021), which add complexity to the already known complex architecture of the grapevine resistance mechanisms against P. viticola (Table 4.2). Rpv22, Rpv23, Rpv24, Rpv25 and Rpv26 loci are suggested to derive from V. amurensis. The first three loci were detected using the same population described for Cgr1 (Fu et al. 2019, 2020), whereas the last two loci were identified in a biparental population derived from the susceptible V. vinifera cv. ‘Red Globe’ and the resistant V. amurensis cv. ‘Shuangyou’ (Lin et al. 2019). Rpv22, Rpv23, Rpv24 were detected via genotyping-by-sequencing (GBS)-based QTL analysis, and they were mapped on chromosomes 2, 15 and 18, respectively. The range of phenotypic variance explained by these three QTLs varied from 26 to 30%, and results indicated a total of seven candidate resistance genes associated with these loci. Besides, Rpv25 and Rpv26 have been mapped on different genomic regions of chromosome 15, using a high-density specific length amplified fragment (SLAF)-marker genetic map. Rpv25 explained less than 20% of total phenotypic variance, whilst Rpv26 can be included in the list of major QTLs, due to the high percentage of variability explained, ca. 64%. Candidate genes responsible for these loci include three cysteine-rich receptor protein kinases (in the Rpv25 region) and a gene encoding a LRR-RLK (receptor-like protein kinases) family protein (in the Rpv26 region). On chromosome 18, Sapkota et al. (2019a) mapped a new resistance locus for P. viticola (Rpv27) in a population originated from a cross between the resistant V. aestivalis × V. vinifera-derived cv. ‘Norton’ and the susceptible V. vinifera cv. ‘Cabernet-Sauvignon’, based on a high-resolution linkage map obtained with SNP and SSR markers. Rpv27 can be considered a major QTL, because it can explain around 34% of the phenotypic variance (Table 4.2). Lastly, Rpv28 was mapped on chromosome 10 of V. rupestris, and it explained up to 67% of the phenotypic variance (Bhattarai et al. 2021).
Regarding the genetic determinants of resistance to root-feeding forms of grape phylloxera (D. vitifoliae), three new loci (Rdv6, Rdv7 and Rdv8) have been recently identified in a progeny between the resistant hybrid VRH8771 (V. vinifera × M. rotundifolia) and the susceptible V. vinifera cv. ‘Cabernet-Sauvignon’ (Rubio et al. 2020). Rdv6, identified on chromosome 7, is a major QTL because of the high trait variance it explains. The other two QTLs, mapped on chromosome 3 and 10, explain less phenotypic variance. These results indicate that various QTLs influence grapevine resistance to D. vitifoliae attack. The same population has been used to characterize the genetic determinants of grapevine resistance to the dagger nematode X. index (Rubio et al. 2020). Three new loci were mapped on chromosome 9 (XiR2), 10 (XiR3) and 18 (XiR4), explaining around 23, 21 and 13% of the total phenotypic variance, respectively (Table 4.2).
4.8.4 Association Mapping for QTL Detection: Benefits and Drawbacks
Linkage disequilibrium (LD)-based association mapping (AM) is a complementary approach to the conventional QTL mapping performed in biparental mapping populations to understand the global genetic architecture of complex traits. In contrast to QTL mapping, AM searches for functional variation in a broader context, using an association panel of diverse genotypes derived from germplasm collections and/or breeding programs, which are selected by carrying most of the genetic variability available for the trait of interest (Zhu et al. 2008). The genetic diversity analysed in the association panel is the result of numerous historical and evolutionary recombination events happening across generations, so AM provides higher resolution during QTL mapping than conventional approaches under an adequate marker density (Myles et al. 2009). As a result, whilst findings from biparental crosses tend to be specific to the same or closely related populations, results from AM are more applicable to a much wider level (Zhu et al. 2008). As in other species with long generation cycles, AM is of special interest in grapevine genetics since it reduces research time by not requiring the generation of new mapping populations (Myles et al. 2009).
Despite its multiple advantages, AM also presents a series of drawbacks, including the risk of spurious marker-trait associations due to population genetic structure and family relatedness effects. Association panels often are a compendium of non-independent genotypes with varying levels of pedigree relationships, common geographical origin and local adaptation and breeding history (Zhu et al. 2008). As a result, any phenotypic trait that correlates with the underlying population structure or pedigree relatedness at neutral loci will show an inflated number of spurious associations (Balding 2006). This problem is widely known, and many statistical methods have been developed to reduce the confounding effect of these two factors on AM (Tibbs Cortes et al. 2021). For instance, the popular unified mixed linear model (MLM) proposed by Yu et al. (2006) simultaneously controls for both population structure and kinship effects, providing a powerful solution for trait dissection. Nonetheless, correcting for population structure (and kinship) might increase the number of false negatives (Tibbs Cortes et al. 2021). The presence of high LD between genotyped markers is another factor that might reduce the potential of AM to detect true biological signals (Gao et al. 2010). Therefore, determining an adequate statistical significance threshold is fundamental to identify markers truly associated with the target trait. This is commonly done by Bonferroni correction and false discovery rate (FDR) approaches, which assume independence between association tests. Nevertheless, it is not habitual in AM studies (Tibbs Cortes et al. 2021), resulting in the use of overly stringent thresholds that increase the number of false negatives. Among others, alternative approaches based on the dependency among markers (Duggal et al. 2008; Gao et al. 2010) or marker-based heritability values (Kaler and Purcell 2019) have been proposed to calculate more appropriate significance thresholds for AM.
4.8.5 Extent of Linkage Disequilibrium in Grapevine
The degree of non-random association between alleles at two genetic loci within a population is known as LD (Zhu et al. 2008). AM performance relies on the LD between the genotyped markers and the functional polymorphism in the causative gene, and on the rate of LD decay over a specific genetic distance. In general, markers near the causative locus used to be in high LD with the functional polymorphism, and thus associate with the phenotype of interest. AM detects these associations and marks up the genomic regions harboring these significant markers and the potentially implicated genes (Myles et al. 2009). Therefore, LD is an important factor to determine the number and density of genetic markers needed to reach an adequate statistical power at the whole-genome level. In particular, marker density needs to overcome the underlying LD structure to ensure that all responsible polymorphisms are in linkage with (at least) one genotyped marker. Studies describing patterns of LD in grapevine indicate a fast decay in the sativa subspecies, reaching r2 values below 0.2 within short physical distances. To cite some examples, Nicolas et al. (2016) observed a LD decay below r2 = 0.2 at 43 kbp in a diversity panel of 279 grape cultivars, whilst Marrano et al. (2017) reported a LD extent of 20 kbp to reach the same LD decay through the analysis of 14 k RAD-seq-derived SNPs screened in 51 grapevine cultivars. These estimates are comparable to those indicated by Laucou et al. (2018) through the analysis of 10 k SNPs in 783 genotypes (LD decay below r2 = 0.2 at 29–58 kbp). These short LD values suggest the need of genotyping a very high number of well-scattered markers to have a proper genome-wide statistical power. In fact, Nicolas et al. (2016) suggested the need of genotyping (at least) one marker per kbp, which undoubtedly implies the use of high-throughput genotyping methods like reduced representation sequencing (using technologies like GBS, RAD-seq and double-digested (dd) RAD-seq), or whole genome resequencing to reach such marker densities (Pavan et al. 2020). More recently, the whole genome resequencing of a panel of table, wine and multi-purpose cultivars has suggested that the rate of LD decay in the cultivated grapevine might be faster than previously reported (Kui et al. 2020; Liang et al. 2019), entailing the need of using an even higher number of markers for an adequate genome-wide coverage.
4.8.6 Association Mapping Software and Statistical Models
As stated before, the need to control for population structure and kinship effects in AM studies led to the development of multiple statistical models, and this continues to be an important research topic (see Tibbs Cortes et al. (2021) for a recent review). Nowadays, there is a trend through the development of multi-locus models, which improve mapping resolution compared to single-locus methods by the simultaneous incorporation of multiple markers in the model as covariates. This approach was firstly implemented in the multi-locus mixed model (MLMM) developed by Segura et al. (2012), which paved the way to other derived models, like FarmCPU (Liu et al. 2016b), mrMLM (Wang et al. 2016a) or ISIS EM-BLASSO (Tamba et al. 2017), already tested or with great potential for AM studies in grapevine. More recently, different multi-trait multi-locus models capable of dealing with complex underlying associations between markers and traits have been developed too, such as the penalized MTMM (Liu et al. 2016c) or the mtmlSEM (Igolkina et al. 2020) models. Many mixed linear models and multi-locus models have been progressively incorporated into common software packages like TASSEL (Bradbury et al. 2007), PLINK (Purcell et al. 2007), GAPIT (Lipka et al. 2012) and GEMMA (Zhou and Stephens 2014) to ease data processing and results interpretation. As a result, these packages now implement multiple statistical models to allow a direct comparison between AM solutions. Nevertheless, the development of new methods and tools for AM continues, and, attending to the many works posted as preprints in open-access servers, many more will be released soon, providing a new framework for AM studies in the next few years.
4.8.7 Candidate-Gene and Genome-Wide Association Studies of Grapevine Resistance
For AM studies, two widely used approaches are used: genome-wide and candidate-gene association studies (GWAS and CGAS, respectively). GWAS are mostly used as exploratory analyses to examine the genetic architecture of the trait of interest. On the contrary, CGAS assumes some previous understanding of the genetics of the trait, and commonly the candidate gene derives from a QTL identified through a previous GWAS (Zhu et al. 2008). In other cases, candidate gene selection is based on information obtained from genetic, biochemical or physiological studies in related or model crop species. CGAS are ultimately used to move from QTLs to quantitative trait nucleotides (QTNs), to ideally identify the functional genetic variant responsible for phenotypic variation (Myles et al. 2009).
Different CGAS are available exploring relevant traits for grape breeding like muscat flavor, berry colour, or cluster characteristics (see Vezzulli et al. 2019b and references therein). Nevertheless, this approach has not been applied yet to the analysis of disease resistance in grapevines. On the contrary, several works dealing with this issue via GWAS can already be found in the literature. Zhang et al. (2020) examined the genetic architecture of grapevine white rot disease resistance caused by C. diplodiella through the analysis of 386 genotypes and 88.877 SNPs detected by RAD-seq. MLM results indicated six SNPs located on chromosomes 1, 2, 4, 13, 16 and 17 significantly associated with disease symptoms. The analysis of the neighbouring regions derived in the detection of eight candidate genes, five on them with putative functions related to plant resistance mechanisms. Through a GWAS using 350 cultivars and 77,126 SNPs detected by GBS, Jang et al. (2020) explored the ripe rot disease resistance mechanisms caused by the fungal pathogens C. acutatum and C. gloeosporiodes. Results identified 26 and 44 SNPs significantly associated with C. acutatum and C. gloeosporiodes disease symptoms respectively, which subsequently led to the identification of two genes that code for two coiled coil (CC)–NBS–LRR proteins capable to recognize specific pathogen-derived products to start a complex resistance response. Lastly, after testing both single- and multi-locus GWA models, Sargolzaei et al. (2020) identified three new genomic loci associated with grapevine resistance to P. viticola (Rpv29, Rpv30, and Rpv31, in chromosomes 14, 3, and 16, respectively) coupling information from a breeding population obtained by self-pollination of ‘Mgaloblishvili’ (a V. vinifera cultivar resistant to DM (Toffolatti et al. 2018)) and a series of Georgian cultivars, genotyped with the Vitis18K SNP-chip array (Laucou et al. 2018). These three new loci co-localize in genomic regions enriched of genes associated with plant defense mechanisms against biotic stress, including receptors of pathogen effectors, signaling mechanisms mediated by protein ubiquitination, and a cluster of Lr10-like (NB-LRR) effector receptors.
4.8.8 Potential Application of QTL Results for Assisted Germplasm Enhancement
Decades of research support the usefulness of genetic mapping to detect QTLs and underlying genes involved in grapevine biotic resistance mechanisms, results that have been eventually used as a starting point to guide MAB and germplasm enhancement programs and pyramid breeding. In general, to avoid the resistance breakdown risk that is associated with monogenic and oligogenic resistance, breeding programs aim at pyramiding different QTLs in single prebreeding varieties, used as donors of traits in breeding programs (Zini et al. 2019). Indeed, durability of resistance to the nematode X. index (vector of GFLV) was confirmed in grapevine rootstocks possessing three QTLs (XiR2, XiR3 and XiR4) (Nguyen et al. 2020). In contrast, P. viticola strains that are able to overcome resistance due to a single QTL (Rpv3 and its allelic variants Rpv3.1 and Rpv3.2) have been isolated in Czechia, Italy and Germany (Eisenmann et al. 2019; Peressotti et al. 2010; Toffolatti et al. 2012). The recent discovery that resistance traits can also be found in V. vinifera cultivars (as observed with DM (Toffolatti et al. 2016)) opens the way to new breeding perspectives within this species.
Now, AM potentially allows discovering the QTNs responsible for phenotypic variation, which enables the development of highly efficient functional markers to track relevant traits in grapevine breeding (Emanuelli et al. 2014). Nevertheless, and even under strong statistical evidence and/or theoretical support, the risk of false positive associations is still present, which calls for an independent validation process of the candidate gene/s. This stage could be approached through a cross validation in different populations, as the potential usefulness of an association would be higher if it is found in independent genetic backgrounds. Alternatively, laboratory experiments (like candidate gene knock-out or overexpression studies) could be determinant to bind a candidate gene or candidate mutation to a trait of interest, as proved for the grapevine mlo genes and PM susceptibility (Pessina et al. 2016). In this regard, emerging genome editing technologies such as the CRISPR/Cas9 system are suggested to speed up this process, as they are expected to provide determinant functional information to validate the role of a candidate gene (or mutation) on a specific trait (see Sect. 4.13.2).
4.9 Hints About Map-Based Cloning of Resistance Genes
4.9.1 Genomic DNA Libraries and Physical Mapping
Unlike the plethora of genetic loci identified in grapevine, single genes controlling important agronomic traits–such as biotic stress resistance–are barely unknown. A physical map is essential to positionally clone such genes and instrumental in a genome sequencing project. Bacterial artificial chromosome (BAC) libraries are the large DNA insert libraries of choice and an indispensable tool for map-based cloning, physical mapping, molecular cytogenetics, comparative genomics and genome sequencing. In contrast to their name, BACs are not artificial chromosomes per se, but rather are artificial bacterial F factor derived constructs (Ren et al. 2005).
Based on an automated protocol, an initial physical map was constructed based on 29,727 BAC clones derived from the cultivar 'Cabernet Sauvignon'. Despite some limitations that interfere with the correct assembly of heterozygous clones into contigs, this physical map is a useful and reliable intermediary step between a genetic map and the genome sequence. This tool was successfully exploited for a quick mapping of complex families of genes, and it strengthened previous clues of co-localisation of major NBS-LRR clusters and R-loci in grapevine (Moroldo et al. 2008). Then, the first whole genome physical map of grapevine was built using high information content fingerprinting of 49,104 BAC clones from the cultivar Pinot Noir. Prior computer simulations, the experimental assembly results were in full agreement with the theoretical expectations, given the heterozygosity levels reported for grape (Scalabrin et al. 2010). The physical map was anchored to a dense linkage map based also on BAC-end sequence (BES) markers (Troggio et al. 2007), paving the way to a new era in both grapevine genetics and GAB.
4.9.2 Positional Cloning of R-Genes
The ultimate goal of mapping agronomical traits is the identification and isolation of the underlying genes. This approach aiming to associate a phenotype with a genotype is known as “forward genetics”. In forward genetics, map-based cloning (or positional cloning) is a commonly used strategy to isolate genes governing major traits. Initially, molecular markers physically linked to the trait of interest have to be identified through whole genome or local genetic mapping. Traditionally, a local fine map is constructed with the aim to assign the trait to the smallest possible genetic region, thus reducing subsequent sequencing efforts. This means that the region of interest will be targeted and densely covered by molecular markers in order to detect markers flanking the gene of interest as closely as possible. When a marker is closely related to a gene, low recombination frequency is expected and large segregating populations are required to detect the rare recombinant (Welter et al. 2011).
This strategy was successfully used to physically map the resistance locus Run1. At first, a local map around the Run1 locus was constructed employing the BSA strategy (Pauquet et al. 2001). Secondly, using the same strategy, resistance gene analog (RGA)-based markers tightly linked to the Run1 locus were detected (Donald et al. 2002). For fine mapping, three independent populations segregating for the resistance locus Run1, in total 996 recombinant individuals, were employed (Barker et al. 2005). Fine mapping allowed the identification of two flanking microsatellite markers showing a very low recombination frequency (tight linkage) with the Run1 locus, thus defining a short genetic interval for the locus. These two flanking markers were used together with three markers that co-segregated with Run1, to screen a BAC-library constructed from the genomic DNA of a plant carrying Run1. In this way the physical mapping of the region spanning the resistance locus was performed. Marker-carrying BAC clones were end-sequenced and assembled into extended contigs after identification of overlapping BACs. This allowed an overall coverage of the region spanning the locus Run1. After the sequencing of the contig spanning the genomic region associated with Run1, positional candidate genes were selected for functional analysis. The identified gene responsible for PM resistance has shown to encode a TIR–NB-LRR domain protein which represents the most important class of R proteins in plants (Feechan et al. 2013). Further examples of map-based cloned genes responsible for biotic stress resistance have concerned the PdR (Riaz et al. 2008a), Ren1 (Hoffmann et al. 2008), and XiR (Hwang et al. 2010) loci.
The availability of the grapevine genome sequence opens up new options to accelerate the map-based cloning of genes. Molecular markers surrounding the trait of interest may be anchored to the genome sequence. This genomic region may then be searched for positional candidate genes, based on their predicted functional role. Molecular markers tagging such candidate genes can be developed and tested for their association to the trait. Additionally, markers (e.g. SNPs) with a progressive physical distance from the trait of interest may be developed in both directions and used to estimate their recombination frequencies to the trait. However, it is important to be aware of the fact that specific genes such as resistance genes may not be present in the currently elaborated model genome sequences, as these have been derived from susceptible grapevines. It remains to be seen from further grapevine genome sequencing to what extent the reference genome sequence of ‘PN40024’ are colinear to other grapevine cultivars, breeding material and Vitis wild species accessions on large- andfine scale levels (Anderson et al. 2011). Actually, the milestone of conventional BAC-based physical mapping seems not to be overcome with the advent of genome sequencing and the boost in next-generation sequencing technologies. Lately, complementary strategies for assembling the Rpv3.1 haplotype were adopted combining whole genome shotgun (WGS) sequencing of a Vitis accession, homozygous for Rpv3.1, and restriction-based fingerprinting and sequencing of BAC clone inserts across the Rpv3.1 region. This approach allowed for mapping the causal factor for DM resistance to an interval containing a TIR-NB-LRR (TNL) gene pair (Foria et al. 2020).
From the applied point of view, positional cloning of the resistance genes provides sequence information that can be used to design perfect genetic markers, which will maximize the efficiency of MAS approaches. In addition, map-based cloning offers the possibility to introduce these genes into existing elite wine grape cultivars, which might not be hybridized, by transgenesis without affecting wine quality.
4.10 Marker-Assisted Breeding for Resistance
4.10.1 Development and Evaluation of Robust Molecular Markers
QTL experiments provide relevant information about the genetics of the trait of interest, circumscribed by the experimental design used. Loci obtained from QTL mapping studies differ in percentage of phenotypic variance explained by the locus (r2), allele effect, and confidence intervals (see Table 4.2). Although loci with high r2 are preferred, a breeding strategy could use loci explaining different percentages of the phenotypic variance. Before deployment in breeding programs with more diverse genetic background, the effect of the desired alleles requires confirmation, even in the case of a major QTL (Bernardo 2014).
In grapevines, a well-documented example of locus validation is the p3-VvAGL11 marker linked to the seedlessness (sdl) locus, which has been successfully adopted by several table grape breeding programs (Bergamini et al. 2013; Ocarez et al. 2020). Biotic resistance is a more complex trait that requires further study. In a recent study, SSR alleles linked to 11 R-loci were investigated among 102 grapevine accessions displaying different levels of PM and DM disease severity over 3 seasons, with only a few of them being further suitable for marker-assisted parent selection (MAPS) or marker-assisted seedling selection (MASS) (Zini et al. 2019). Selection accuracy is related to the genetic distance between the causal polymorphism and the molecular marker selected. Two markers flanking the loci can be used to increase accuracy, as well as haplotypes instead of a single marker, as recently reported for the Rpv3 locus (Wairich et al. 2021). Highly saturated genetic maps and controlled phenotyping assays allow precise loci dissection. Fine mapping of the Ren3 locus narrowed the location to a 7 Mb segment that included both Ren3 and Ren9 (Zendler et al. 2017), and subsequently into separate intervals of 3.1 and 0.6 Mb, respectively (Zendler et al. 2021b). In this study, new molecular markers tightly linked to these two regions were described, allowing breeders to track the presence of these two sources of resistance independently.
As described in Sect. 4.8, new genotyping technologies based on NGS, such as GBS and RAD-seq, have improved genetic mapping. But the strategy is not well suited for detailed characterization in grapevines, due to the high percentage of missing data, sparse coverage and heterozygote undercalling (Hyma et al. 2015; Barba et al. 2014). For grapevine breeding, loci identified with this strategy have been further studied to: (i) assess the linkage to existing markers located nearby, such as SSRs (Barba et al. 2018), (ii) develop fluorescence hybridization-based genotyping assays, such as kompetitive allele-specific PCR (KASP™) Genotyping (Wairich et al. 2021) or high resolution melting curves (Jang et al. 2020), and (iii) development of genotyping assays based on NGS, such as AmpSeq (Yang et al. 2016) or rhAmpSeq (Zou et al. 2020). Given the high-throughput nature of NGS, these assays allow the screening of up to 2,000 sites in one experiment, but require specific pipelines for data analysis (Fresnedo-Ramírez et al. 2019). Next-generation sequencing technologies have also enabled the study of heterozygous genomes. In a recent study, a partially-phased reference genome of the resistant interspecific hybrid ‘Börner’ has been developed and utilized for marker discovery (Holtgräwe et al. 2020). Their study found 10,820 putative SSR marker positions, with more than 4,000 tri-allelic SSR candidates. As a proof of concept, 19 single nucleotide variant (SNV) markers were developed around the Rpv14 locus and used to narrow it down to 330 kbp using bulked segregant analysis (BSA) of the V3125 x ‘Börner’ progeny.
4.10.2 Marker-Assisted Selection as a Tool for Marker-Assisted Breeding
The use of molecular markers tightly linked to R-loci obtained from QTL studies helps optimize resources, select progeny with several R-loci for durable resistance, and expedite breeding of new resistant grapevine cultivars through MASS (Dry et al. 2019; Vezzulli et al. 2019b). As useful new loci are identified and publication becomes imminent, they are named and listed here for use by the Vitis research community: https://www.vivc.de/index.php?r=loci%2Findex (Table of Loci for Traits of Grapevine; Maul et al. 2021).
In recent years, several programs have tested markers linked to Ren/Run and Rpv loci to determine the presence of alleles associated with resistance, using germplasm of diverse origins, such as Russia (Ilnitskaya et al. 2020), Kazakhstan (Pozharskiy et al. 2020) and Italy (Vezzulli et al. 2019c). Beyond PM and DM, breeders have expanded the use of MAS to other pathogens and pests, such as the Mjr1 locus for M. javanica resistance and D. vitifoliae resistance (Table 4.3). In most cases, the R-loci originated in Vitis spp. other than V. vinifera, thus the introgression into the cultivated genetic background is required.
4.10.3 Marker-Assisted Gene Introgression
Marker-assisted seedling backcross and MAPS have been used very effectively for rapid gene introgression. Modified backcross (MBC) breeding is used for Vitis improvement, whereby the recurrent parent is a different unrelated genotype in each generation of crossing. When introgression of a single locus from a wild species is the goal, the trait of interest would normally be assayed prior to crossing to produce the next backcross generation. Where the trait may be expensive to assay or dependent on fruit phenotyping, additional time and funding are required. Yet if tightly linked markers are available, then selections may be made in the weeks following germination, once DNA samples are extracted and analyzed. An excellent example comes from the introgression of PdR1b (encoding PD resistance) on chromosome 14 from V. arizonica into V. vinifera wine grapes (Riaz et al. 2008b). Generation time was reduced to 2 years, or 4 generations in 10 years (Walker and Tenscher 2019) through optimized growing practices. With each generation, presence of the resistance locus was assayed using SSR markers (VVIP26 and CH14-77) soon after seed germination (Riaz et al. 2008b, 2009; Walker et al. 2021a, b). Greenhouse assays were also used to confirm resistance.
As a result of this work, the top selections ranging from 88% (MBC2) to 97% (MBC4) V. vinifera background were field-tested at multiple locations in California and Texas. Evaluations of viticultural traits and enological qualities led to the release of five new cultivars in 2019 (Walker et al. 2021a, b caes.ucdavis.edu). The project took ~20 years, including the time it took for four cycles of backcrossing (Walker and Tenscher 2019).
V. rotundifolia (selection G52) has strong resistance to PM of grapevine leaves, petioles, canes and fruit. The difficult cross between this muscadine grape (2n = 40) and V. vinifera (2n = 38) was made in North Carolina prior to 1919 (Detjen 1919) and a resulting hybrid (NC 6–15) remained under-utilized for many years. Backcrossing efforts to introgress PM resistance into V. vinifera began in the 1980s (Bouquet 1986), whereupon the single locus for PM resistance, Run1, was first identified. Initially, leaf disc assays and field phenotyping were required to test for the presence of the muscadine resistance source. Later, AFLP markers were established (Pauquet et al. 2001) and proposed for MAS. The causal gene was later identified, and improved, tightly-linked markers were reported (Feechan et al. 2013).
As a result of the efforts of Bouquet, along with the teams developing molecular markers for the Run1 locus, it is now used widely by breeders in Hungary (Hajdu 2015), New York (Cadle-Davidson et al. 2011), and elsewhere. In France, the INRA-ResDur1 breeding program (Merdinoglu et al. 2018) developed and released four cultivars incorporating the Run1 locus along with another locus for PM resistance, and two loci conferring DM resistance. Cornell University test selections with the Run1 locus are currently in field trials in consideration of future release (Reisch, pers. comm.).
4.10.4 Gene Stacking
To increase the durability of resistance, gene stacking (“pyramiding”) has been proposed (Mundt 2018; Dry et al. 2019). In grapevines, this approach is enabled by the identification of multiple R-loci for both PM and DM resistance (Dry et al. 2010). However, the race-specificities of the many available loci are only known in a small number of cases, and there are likely more races in situ than have been isolated for laboratory assays. Nevertheless, Stam and McDonald have estimated that four R-genes stacked together would be virtually impossible for cereal PM to overcome (Stam and McDonald 2018). With long-lived perennial crops like grapes, it is prudent to consider measures to make sure that resistance features of new cultivars are not easily overcome.
For this reason, some have also proposed combining management strategies (including limited fungicide applications) in combination with deployment of resistant cultivars as a pathway to better assure the durability of resistance over time (Feechan et al. 2015). If left completely uncontrolled, conditions for the pathogen to mutate are optimized.
An excellent example of a gene stacking strategy is outlined by Dry et al. (2019) in which the INRA-ResDur program, working with the Julius Kühn-Institut (Germany), Staatliches Weinbauinstitut, (Germany) and Agroscope (Switzlerand) obtained lines with three loci for PM resistance plus three loci for DM resistance. Also in Italy Fondazione Edmund Mach (FEM) obtained stacked (“pyramided”) genotypes carrying two or three loci for DM and/or PM resistance, up to seven combined R-loci in total (Vezzulli et al. 2019c); moreover, Vivai Cooperativi Rauscedo (VCR), along with University of Udine, developed resistant genotypes derived from “elite” cultivars carrying two Rpv loci coupled with two Run/Ren loci (Foria et al. 2019).
4.10.5 Up-and-Coming Exploitation of Susceptibility Genes
Susceptibility genes in plants facilitate infection and compatible interaction with the pathogen (see Sect. 4.13). Development of cultivars homozygous for non-functional alleles has been a successful breeding strategy for durable PM resistance in barley using the MLO gene (Jørgensen 1992), and DM resistance using the DMR and DLO genes, first identified in Arabidopsis thaliana (Van Damme et al. 2008). In grapevines, gene homologues have been studied and described in (Dry et al. 2019). Screening of non-functional mutants among the natural rich diversity of Vitis, or its creation using new breeding techniques such as EcoTILLING or gene editing could be valuable for the development of biotic stress resistant grapevines (Dry et al. 2019; Merdinoglu et al. 2018; Pirrello et al. 2021).
Given high heterozygosity and inbreeding depression, the adoption of susceptibility gene strategies in traditional grapevine breeding will require the use of a modified backcross scheme. To this end, use of donors with diverse genetic backgrounds and co-dominant markers linked to the susceptibility loci (to allow selection on the absence of the phenotype among heterozygous seedlings) will expedite the process.
4.10.6 Limitations and Prospects of Molecular-Assisted Breeding Applications
To be most useful, genetic markers should be (i) tightly linked, (ii) flanking the locus of interest, (iii) inexpensive, (iv) rapid to assay, and (v) readily transferable between populations. Microsatellite loci have been most widely used (Vezzulli et al. 2019b), but SNP-based assays are compatible with high throughput analyses, automated pipelines, and may increasingly become the system of choice. There are dual roles for MAS: both for selection of desirable seedlings within weeks of germination (MASS), and for selection of parents with desirable traits (MAPS). Both roles present breeders with the potential to reduce phenotyping errors, and to accelerate the identification of desired genotypes. Yet the overall progress in grapevine breeding is not always accelerated by MAB since multi-year field trials are still required to assess the totality of viticultural (and often enological) performance. The best example of accelerated breeding is given in the above discussion of Pdr1b locus introgression from a wild species into cultivated wine grapes at the rate of four generations per 10 years. Marker-assisted breeding is also advantageous for gene stacking approaches where trait phenotyping is unable to determine the marker complement in each seedling (e.g. presence of multiple loci for PM resistance). Yet knowledge of which loci are complementary to each other, and which loci will lead to durable stress resistance, is quite lacking.
Marker-assisted breeding is most useful for selection of single loci of moderate to strong effects. It is not useful where traits are under quantitative genetic control, e.g. berry weight, cluster size, etc. If minor QTL are selected for MAB, stability should be evaluated across environments prior to use (Vezzulli et al. 2019b). Where the goal is to manipulate polygenic traits, genomic selection (GS) may be the preferred approach but requires further investigation in Vitis. High-density marker sets are available through GBS, RAD-seq, AmpSeq, and rhAmpSeq methods, but the calibration and validation of GS models remains in order to investigate the potential for DNA analyses to inform the breeding for complex, polygenic traits.
4.11 Towards Genomics-Assisted Breeding for Resistance Traits
4.11.1 Excursus in Genome Sequencing
The number of grapevine and pathogen/pest genome sequences as well as tools for functional genomics has increased dramatically in the 15 years since the first grapevine sequence assemblies were released. In 2007, two genome assemblies, a descendant of ‘Pinot Noir’ and a heterozygous ‘Pinot Noir’ clone, were the fourth assemblies for flowering plant and first woody fruit crop genome assemblies to be released (Jaillon et al. 2007; Velasco et al. 2007). Grapevine is very heterozygous and the highly homozygous ‘PN40024’ (~7% heterozygosity) made it easier to develop an assembly with haploid consensus sequences (Jaillon et al. 2007). Further sequencing, re-assembly, gene annotation, and an updated genome browser made this a very useful resource in identifying disease related genes and supporting large-scale transcriptomic and proteomic analyses (Canaguier et al. 2017). Subsequently, several short-read assemblies with greater genome coverage were produced for cultivars and species; however, due to the heterozygosity of the grapevine these resulted in fragmented sequences, large number of contigs, large assembly size and inaccuracies in gene models and copy number (e.g. Da Silva et al. 2013; Di Genova et al. 2014; Patel et al. 2020). Regardless, the benefit of these and other newly generated genome sequences was shown in the alignment of multiple species and cultivars that enabled the identification of core sequences and development of a set of universal multiallelic genetic markers (rhAmpSeq) with greater transferability across interspecific grapevine populations (Zou et al. 2020).
The divergence of sequences between cultivars and species, which has been acquired during speciation, hybridization and selection, cannot be encapsulated in a few consensus genome assemblies; thus, de novo assemblies conserving the characteristics of both haplotypes for many additional genotypes (including interspecific genotypes) is needed. Several short read and phased long read assemblies are publicly available for 12 V. v. subsp. sativa cultivars (‘Pinot Noir’, ‘Black Corinth’ (seeded and seedless), ‘Cabernet-Sauvignon’, ‘Carménère’, ‘Chardonnay’, ‘Merlot’, ‘Reisling’, ‘Semillion’, ‘Sultanina’, ‘Tannat’, and ‘Zinfandel’), some accessions of V. v. subsp. sylvestris, and six species (V. arizonica, V. cinerea, V. rupestris, V. riparia, V. amurensis and M. rotundifolia), providing the opportunity to mine disease resistance genes. The largest single access point for grape and grape pathogen genomes that are browser enabled can be found at the http://grapegenomics.com website (D. Cantu Laboratory, UC-Davis, USA). Recent assemblies using long read sequencing technology have provided the opportunity to assemble the entirety of a genome with both haplotypes, thereby identifying structural variation within coding sequences (Minio et al. 2017). Development of diploid genome assemblies with phased haplotypes for many cultivars and species provides the opportunity to identify unique genes that are important in phenotypic variation in response to biotic stressors in grapevine. It is important to sequence multiple genotypes of each of the 60–70 dioescious wild species, develop diploid genome assemblies and mine structural variation and gene content as the wild species are the primary source of pathogen and pest resistance genes.
A survey of common grapevine pathogens indicates a growing number of draft genomes of the major fungal and bacterial pathogens as well as numerous viral and antagonistic pathogens of grapevine pathogens. There are no database resources for these pathogen genomes making it difficult to conduct comparative analysis within a species or between strains. Development of such a resource would provide a greater opportunity to improve detection and understanding of host–pathogen dynamic interactions (Näpflin et al. 2019).
4.11.2 Gene Prediction and Annotation
Protein coding gene prediction and annotation are ongoing efforts with new annotation technologies, greater understanding of the structural components within a genome, and new genomes being assembled. Gene prediction using automatic gene annotation has identified 27,000 to 33,568 genes for the ‘PN40024’ 12X.v2 consensus reference genome VCOSTv3 annotation (Canaguier et al. 2017). Transcriptional evidence—expression sequence tags (ESTs)s and cDNA—provides further validation of gene models, emphasizing the importance of paired end and full-length cDNA transcriptomes for identifying novel genes (Minio et al. 2019a). Gene content differences between cultivars and species can be assessed by alignment with well annotated genomes such as ‘PN40024’ and ‘Cabernet-Sauvignon’ (Canaguier et al. 2017; Minio et al. 2017). However, it is important to keep in mind that the inbred ‘PN40024’ likely represents of fraction of the greater diversity of Vitis species and cultivar gene information. Recent comparative analysis of ‘Carménère’ indicated numerous structural variants relative to ‘PN40024’ and ‘Cabernet-Sauvignon' genomes likely contributed to its differences in gene content (absence 494 and 253; novel 1561 and 449) in ‘Carménère’, respectively to ‘PN40024’ and ‘Cabernet Sauvignon’ (Minio et al. 2019b). Increased availability of annotated phased diploid assemblies, that are more complete than ‘PN40024’, enables identification of homozygous and heterozygous gene regions and gene regions unique to one haplotype allowing the exploration of allelic differences that affect phenotypic differences between genotypes (Smit et al. 2020).
4.11.3 Updates on Transcriptomics, Proteomics and Metabolomics Databases
High-throughput ‘omic technologies are rapidly generating huge datasets (transcriptome, metabolome, proteome, and other components of the grapevine phenome); however, access and utilization of these datasets is limited by the need for searchable databases. Public sequence repositories for RNA and protein sequences provide an important data repository for grapevine information but have limitations in comparative analysis. VitisNet provided a database of gene pathways based on the ‘PN40024’ genome V2 annotation aiding in the functional understanding of gene expression and proteomic studies (Grimplet et al. 2009, 2012). A gene co-expression database utilizing grapevine microarray expression data (~29,000 genes) allowed identification of potential regulatory mechanisms and infer gene function (Wong et al. 2013). Moretto et al. (2016) established the VESPUCCI grapevine gene expression database (http://vespucci.colombos.fmach.it/; 1608 microarray and 135 RNA-seq samples) allowing the user to explore, analyze and visualize the expression data. Grape RNA is a database of RNAseq and microRNA with 25 experimental conditions, assembled transcriptomes, GO, KEGG and NCBI annotations and tools to search, conditions was recently established expanding the RNA-seq data that can be queried using ‘PN40024’ V2 annotation (Wang et al. 2020a). These historical databases provide context for much of the early transcriptomic data; however, strategies for maintaining current and historical data in a useable format is a challenge. Databases quickly become obsolete with the rapid development of genomes and new data acquisition technologies. The heterogenous nature of the phenomic datasets further challenges the grapevine research community to organize, fund, and maintain both historical and new databases. A framework for a distributed grapevine information system has been envisioned, and remains to be implemented with metadata standards for all data types to promote dataset access and organization in future databases (Adam-Blondon et al. 2016).
4.11.4 Methylomics
Inspired by seminal studies in Arabidopsis and cereals and enabled by the revolution of NGS in the early 2000s, research in fruit plant science has started investigating chromatin properties and related epigenetic phenomena more deeply, although the discipline is still in its infancy as far as grapevine is concerned. Chromatin modifications, including DNA methylation, histone modifications and nucleosome spatial rearrangements, are deployed by eukaryotic cells to modulate gene expression and maintain genome stability through the control of chromatin tridimensional structure and accessibility. The mode of replication and conservation of such modifications during cell division is well-suited for the potential transmission of gene regulatory information through cell lineages during the development of a plant body plan or through vegetative and sexual reproduction, posing the precondition for long lasting memory or even transgenerational inheritance of regulatory setups for gene expression (reviewed in Lämke and Bäurle 2017). The role of chromatin modifications in mediating fundamental genomic functions, recording environmental signals and stabilizing molecular responses and phenotypes over time are all different nuances of a unique body of mechanisms that convey genetic information and elicit genetic differentiation without involving changes in the DNA primary sequence, hence the term epigenetics (literally “beyond genetics”) used to refer to these phenomena.
The early steps of epigenetic research in Vitis put a major emphasis on a number of epigenetic features as well as on specific scientific questions raised by the biology of the species and its exploitation, namely (i) the identification of molecular factors known to be responsible for epigenetic modifications and the genetic basis thereof; (ii) the profiling of chromatin modifications at the genome scale (the so called “epigenome”); (iii) the molecular changes triggered by in vitro regeneration as a source for somaclonal variation; (iv) the capability of epigenetic modifications to generate diversity despite an invariable genetic background; and (v) the effects of environmental cues on the epigenome as a molecular foundation for the concept of terroir. In recent studies tackling the above aspects, DNA methylation data have been collected from grapevine samples using NGS approaches, mainly based on the golden standard method of bisulfite sequencing (BS-seq), at the whole genome level or using reduced representation techniques such as reduced representation bisulfite sequencing (RRBS) (Niederhuth et al. 2016; Celii 2016; Xie et al. 2017; Dal Santo et al. 2018; Williams et al. 2020; Varela et al. 2021 and others mentioned hereafter). Some studies also exploited the technique of methylation sensitive amplification polymorphisms (MSAP) (Xie et al. 2017, Valeraet al. 2021) based on methylation sensitive restriction enzymes, which is less informative but suitable for a cheaper pre-screening. Compared with DNA methylation, histone modifications and chromatin accessibility have been poorly investigated in grapevine thus far. A notable exception is represented by the fruitENCODE project, which analyzed the regulatory circuits of ripening in different climacteric and non-climacteric fleshy fruit species, including grape, by means of ChIP-seq and DNaseI-seq methods (Lü et al. 2018). To date, this study is the most comprehensive source of information on histone modifications in grapevine with a major focus on the H3K27me3 and H3K43me3 modifications that are associated with gene silencing and active transcription, respectively. Histone modifiers (HMs) involved in the deposition and removal of histone modifications are several and much diversified in plants; a bioinformatic search in the grapevine genome (Wang et al. 2020b) identified 117 HM genes sorted into 11 subfamilies and differentially expressed according to anatomy, hormone treatment and exposure to fungal infection. Tandem and segmental duplications seem to account for 21% of all the HM genes identified, suggesting a possible diversification of functions following species-specific genome rearrangements.
The epigenetic regulation of gene expression also involves non-coding RNAs (ncRNA), which comprises ~ 20–24 nt small interfering small RNAs (siRNA) capable of guiding the RNA-induced Silencing Complex (RISC) toward the genomic sites to be epigenetically silenced. The biogenesis and function of siRNAs requires RNA-dependent RNA polymerases (RDR), riboendonucleases of the DICER-like (DCL) family and ARGONAUTE (AGO) proteins that are catalytic components of the RISC complex. According to a bioinformatic survey, a total of five VvRDR, four VvDCL and thirteen VvAGO genes are present in the grapevine genome (Zhao et al. 2015), including the peculiar VvAGO10a gene, only expressed in the stem and possibly involved in the systemic movement of siRNAs (Melnyk et al. 2011). Next-generation sequencing was used in some studies (Chávez Montes et al. 2014; Zhu et al. 2018) to identify large numbers of endogenous siRNAs from multiple grapevine tissues/organs and provide confirmatory results about the features and genomic distribution of this small RNA category in addition to promising information on their differential expression across developmental stages.
Grapevine is one of a few dicot species investigated thus far that stand out for the peculiar DNA methylation pattern observed at the genomic level (Niederhuth et al. 2016). Indeed, DNA methylation in plants occurs by the covalent addition of a methyl group to the 5th carbon position of cytosines in different sequence contexts, namely CG, CHG and CHH (where H represents any base other than G). The three contexts are associated with different DNA methylation pathways and the variation in the amount of average methylation at each context may represent predominance for a specific pathway or the lack of it. The cultivated grapevine is characterized by a lower-than-average level of CHH methylation, which is usually associated with siRNA-mediated RNA-dependent DNA methylation, and it shares this feature with other clonally propagated species (Niederhuth et al. 2016). It is not clear to which extent the mode of reproduction is causally related to CHH methylation, but it has been verified that higher levels of CHH methylation cannot be promptly rescued by a single round of sexual reproduction. As a result, in grapevine, the poor levels of CHH methylation alone can hardly distinguish transcriptionally active regions of the genome from repetitive DNA that is targeted for gene silencing by DNA methylation. However, the meta-analysis of several transposable elements of different families still shows a significant enrichment of CHH methylation, although the effect is much more apparent for the CG and CHG context, suggesting that the underlying function of this pathway may be preserved irrespective of the overall depletion (Celii 2016).
Epigenetic marks can add a further level of diversity among individual plants, even if epigenetic changes may be more stochastic and unstable in contrast with strict-sense genetic polymorphisms altering the DNA sequence. The reversibility of DNA methylation changes, for example, is a distinct feature of in vitro propagation, in which epigenetics has long been considered a useful source of phenotypic variability (Schellenbaum 2008; Baránek et al. 2010, 2015). However, it should be noticed that part of the existing epigenetic variation could be ultimately ascribed to genetic variation, such as genomic structural variation caused by transposable element insertions that are target of epigenetic silencing. For instance, spreading of DNA methylation from repetitive DNA into the surrounding regions (up to 2Kbp) has been observed in the grapevine genome by comparing haplotype-specific DNA methylation profiles at sites of hemizygous TE insertions in the ‘Pinot noir’ cultivar (Celii 2016). In these sites, the haplotype presenting the TE insertion displayed higher DNA methylation levels in the flanking regions than in the homologous regions of the parental haplotype. These and similar processes capable of creating obligatory epialleles, that is alleles that differ in their epigenetic state because of associated genetic differentiation, may account for some, probably most, of the epigenetic variation observed between grapevine cultivars. Examples of DNA methylation diversity have been observed between ‘Cabernet-Sauvignon’ and ‘Sangiovese’ plants grown in Italy (Dal Santo et al. 2018) and among clones of the ‘Malbec' cultivar grown in Mendoza, Argentina (Varela et al. 2021). However, the paucity of such studies and the different methodological approaches adopted prevent a systematic evaluation of the phenomenon and its biological consequences. Interestingly, DNA methylation is also a predictor of variation in gene expression between grapevine genotypes, but from a different angle that challenges the traditional view of a negative association between DNA methylation and expression. Indeed, when gene body CG methylation is considered, rather than methylation in promoters or in detrimental repetitive DNA elements, the function of such epigenetic mark does not seem to be linked to silencing and is rather enriched in the transcribed region (gene body) of housekeeping genes that are stably and constitutive expressed in most cultivars (Magris et al. 2019). This observation, which represents an example of a more general phenomenon detected in most plants (Bewick and Schmitz 2017, Zilberman 2017), suggests that the lack of DNA methylation might be the signal to look for when searching for epigenetic marks of potential variability in the molecular phenotype.
Whether epigenetic mechanisms can contribute to translate soil physicochemical properties, climate parameters and system of vine management into a chromatin signature that is representative of a specific terroir is a fascinating question that still compels a definitive answer, although promising observations are already available. To date, three major studies addressed this question, namely a comparison of DNA methylation profiling between clones of the ‘Shiraz' cultivar grown in six wine sub-regions of the Barossa, South Australia (Xie et al. 2017), a multi-omic comparison between clones of ‘Cabernet-Sauvignon’ and ‘Sangiovese’ both grown in three different environments of North Italy (Dal Santo et al. 2018) and an analysis of clones of the ‘Malbec’ cultivar grown in two contrasting vineyards in the area of Mendoza, Argentina (Varela et al. 2021). Except for the second study, which convincingly demonstrated genotype x environment interactions only at the transcriptional level, the other two showed that DNA methylation data could be used to segregate individual plants based on environmental covariates and in two very distinct geographical scenarios.
4.11.5 Integration of Different ‘Omic and Phenomic Data
Integration of biotic susceptibility or resistance responses (transcriptomic, proteomic or metabolomic) with population genetic studies provides potential to explore the genetic architecture of the phenotypes and identify candidate genes or infer gene roles in response phenotypes. Metabolic biomarkers have been identified by characterizing metabolic profiles associated with disease resistance (Maia et al. 2020; Viret et al. 2018). Histochemical, transcriptomic, and metabolomic analyses of DM response in susceptible and resistant cultivars showed strong correlation between stilbenoid biosysnthesis related genes, stilbene accumulation, and pathogen growth inhibition (Eisenmann et al. 2019). Targeted metabolomic analysis showed a significant induction of stilbenoids in a population segregating for DM resistance (Vezzulli et al. 2019a). Targeted analysis of stilbenoids identified metabolite (m)QTL hotspots with disease-resistance motifs on chromosome 18 and theses mQTLs overlapped a reported PM resistance locus (Riaz et al. 2011; Teh et al. 2019) and the DM resistance region described by Vezzulli et al. (2019b). These complementary studies indicate the power of integrating transcriptomic and metabolomic datasets, collected in conjunction with disease phenotyping in QTL mapping to gain a better understanding of resistance mechanisms. The growing number of genomes and transcriptomic, proteomic, and metabolomic analyses in response to pathogens and pests will increase the opportunity to identify new candidate genes and biomarkers that underly the pathogen and pest response enabling greater understanding of their genetic architecture and development of new cultivar selection and disease/pest control strategies.
New initiatives aiming to coordinate Vitis ‘omic data integrations are being funded (see COST CA17111 INTEGRAPE: Data integration to maximize the power of ‘omics for grapevine improvement, http://www.cost.eu/COST_Actions/ca/CA17111 as an example).
4.12 Brief on Genetic Engineering for Resistance Genes
4.12.1 Target Traits and Alien Genes
Over time, plants have evolved many responses to the different biotic factors (fungi, bacteria, phytoplasmas, viruses, nematodes, insects, and mites) ranging from the initial recognition of the pathogen/pest to the activation of specific containment mechanisms. The precise knowledge of the molecular pathways underlying defense and/or tolerance responses are essential to identify specific genes involved in these mechanisms for their exploitation through established techniques of genetic modification (ETGMs), i.e. recombinant DNA techniques enabling the insertion of genetic information into an organism regardless of sexual compatibility. The difficulties linked to the genetic transformation process in grapevine, including grapevine regeneration problems, genotype effect and long times to obtain stable transgenesis often led to testing candidate genes in herbaceous model plants or in transient expression approaches before the stable transformation in grapevine (Table 4.4). This section reflects a succession of approaches that has often been observed over the years, namely the identification of gene in different grapevine species and characterization in model herbaceous plants, stable transgenic expression in V. vinifera and finally the last step, the future perspective of cisgenesis, i.e. the use of the most promising genes with their native regulatory sequences.
In the last 25 years the genetic transformation in grapevine has been reported in several works. In this section we will report some of the most significant experiences of the last 5 years related to resistance/tolerance to pathogens, while for a more exhaustive overview of transgenesis in grapevine, different reviews are available (Laimer et al. 2009; Gray et al. 2014; Abu Qamar et al. 2017; Capriotti et al. 2020). In recent years, the search for new traits or alien genes for tolerance and/or resistance to grapevine pathogens have essentially, if not exclusively, focused on the fungus/oomycete responsible of DM and PM for the objective of reducing antifungals treatments in vineyards and for a viticulture with low environmental impact. The search for new sources of resistance/tolerance has been intensified by exploiting wild Chinese grape varieties, a germplasm very interesting for breeding new cultivars and studying special resistance mechanisms. In particular, V. pseudoreticulata ‘Baihe-35–1’, V. quinquangularis and V. amurensis were the species whose resistance mechanisms were most exploited (Table 4.4). V. pseudoreticulata ‘Baihe-35–1’ is a valuable germplasm resource for PM resistance (Yu et al. 2013), as well as for DM, anthracnose (Elsinoe ampelina) and Meloidogyne incognita tolerance; V. quinquangularis with a higher resveratrol content compared to other wild Chinese species showed resistance to PM, anthracnose and M. incognita, while V. amurensis is tolerant to cold, DM and anthracnose (Xu and Wang 2014). Through transcriptomic approaches potential traits involved in the tolerance mechanisms were identified in these species, genes such as pathogenesis-related proteins (PRs), transcriptional factors (WRKYs, DREBs, ERFs), heat shock proteins and stilbene synthases. For example, thaumatin-like protein (TLP) belonging to PR protein family 5 was characterized in Chinese grapevine comparing the expression profiles of this gene in disease resistant and susceptible grape species infected with anthracnose, E. necator or B. cinerea. The expression level of VqTLP29 from V. quinquangularis increased following the pathogen inoculations, and the over-expression in Arabidopsis thaliana enhances resistance to PM and the bacterium Pseudomonas syringae pv. tomato DC3000 suggesting that VqTLP29 may be involved in the SA and JA/ET pathways (Yan et al. 2017). VqJAZ7 gene from V. quinquangularis ‘Shang-24’ encodes for a jasmonate ZIM-domain (JAZ) protein, a protein family acting as a negative regulator of JA signalling. In preliminary characterization in Arabidopsis, transgenic lines overexpressing VqJAZ7 enhanced resistance to biotrophic fungus Golovinomyces cichoracearum (responsible of PM in Arabidopsis), but was ineffective against the necrotrophic fungus B. cinerea, and P. syringae pv. tomato DC3000 (Hanif et al. 2018). The results suggested that VqJAZ7 altered the SA-dependent and JA-dependent responses in plants, and it can be useful in grapevine, albeit only against some classes of pathogens.
Some ET response factor (ERF) transcription factors play important roles in the regulation of immune responses in plants. VaERF20 isolated from V. amurensis cv. ‘Shuangyou’ (Wang et al. 2018a) and VqERF112, VqERF114 and VqERF072 from V. quinquangularis (Wang et al. 2020c) were induced by fungal infection. Over-expressed in Arabidopsis they increased the resistance to B. cinerea and P. syringae pv. tomato DC3000. The overexpression of these genes induced the activation of both SA and JA/ET defense genes, callose accumulation and pattern-triggered immunity (PTI) genes, suggesting that the ERF genes isolated from grapevine resistant genotypes are involved in different signal transduction pathways favouring the plant immune responses (Wang et al. 2018a).
Among pattern recognition receptors (PRRs), in V. amurensis cv. ‘Shuanghong’ infected with P. viticola, the VaHAESA gene belonging to the LRR-RLK (leucine-rich repeat receptor-like protein kinase) family was identified through transcriptome analysis. In transient expression in V. vinifera, the gene induced the increase of H2O2, NO, and callose levels and the stable expression in Arabidospis confirmed the activation of PAMP-triggered immunity improving the resistance against DM (Liu et al. 2018). From the same grapevine species, a TIR-NBS-LRR gene (VaRGA1) overexpressed in Arabidopsis induced resistance to the biotrofic Hyaloperonospora arabidopsidis (responsible of DM in Arabidopsis) and P. syringae pv. tomato DC3000, while increases the disease induced by the necrotrophic B. cinerea. The resistance mechanisms induced by VaRGA1 involved the activation of SA signaling, while inhibiting the JA pathway (Tian et al. 2020). Another resistance gene from the family TIR-NB-ARC-LRR (VpTNL1) from V. pseudoreticulata, identified from a transcriptomic analysis of leaves inoculated with PM was characterized in herbaceous host. In Arabidopsis, it enhanced the tolerance to G. cichoracearum and P. syringae pv. tomato DC3000, and in Nicotiana tabacum was found to confer resistance to tobacco Erysiphe cichoacearum DC. This gene represents an interesting candidate for PM resistance in grapevine (Wen et al. 2017).
Among transcription factors, bZIP family plays a crucial role in response to abiotic stress and plant development, however in recent years evidence demonstrates the bZIP can be involved also in plant immune response. VvbZIP60 isolated from V. vinifera cv. ‘Jing Xiu’ was accumulated in leaves in response to pathogens, or to exogenous application of SA and JA. Interestingly, overexpression of VvbZIP60 increased in Arabidopsis the resistance to PM by accumulating PR1 and inducing several genes involved in the SA-signaling pathway (Yu et al. 2019a). WRKY transcription factors are involved with SA in the response against fungi. VvWRKY53 plays a role in the resistance response during the early stage of infection of PM, and other WRKYs isolated from V. pseudoreticulata ‘Baihe-35–1’ and V. labrusca x V. vinifera cv. ‘Kyoho’ improved the resistance to biotic stress (Zhu et al. 2012; Guo et al. 2014, 2018). VdWRKY53 isolated from V. davidii (a wild Chinese grapevine species showing high level of resistance to white rot caused by the fungus Coniella diplodiella (Speg.) Petr. and Syd.) and inserted in Arabidopsis showed greater resistance to C. diplodiella, P. syringae pv. tomato DC3000 and G. cichoracearum (Zhang et al. 2019), confirming that WRKY transcription factors are strong activators of defense-related genes.
Melatonin (N-acetyl-5-methoxytryptamine) in plant plays a key role in different developmental processes including responses to biotic and abiotic stresses (Nawaz et al. 2016), and serotonin N-acetyltransferase (SNAT) is one of the key enzymes for melatonin synthesis from L-tryptophan. The expression of VvSNAT2 isolated form V. vinifera cv. ‘Cabernet-Sauvignon’ was induced by pathogen inoculation, and transgenic Arabidopsis overexpressing VvSNAT2 showed high levels of melatonin and chlorophyll and improved the resistance to PM through the activation of SA signaling (Yu et al. 2019b).
C-repeat-binding factor dehydration-responsive element-binding factor 1C (CBF2/DREB1C) is a transcription factor family well known to play an important role in freezing tolerance and cold acclimation of plants. Recently, its involvement in the early response to DM in grapevine was demonstrated. The gene isolated from M. rotundifolia (MrCBF2) introduced in Arabidopsis showed an increased resistance to DM linked to the accumulation of SA and PR transcripts. However, MrCBF2-overexpressing plants exhibited an altered phenotype such as growth retardation, dwarfism, late flowering, and prone rosette leaves (Wu et al. 2017).
These are representative examples of functional characterization of candidate genes for tolerance to pathogens in recent years, some of them may be interesting genes to be tested on transgenic grapevines (essentially in V. vinifera) as well as being candidates for future cisgenic approaches.
4.12.2 Genetic Transformation for Biotic Stress Resistance
Alongside functional characterization in herbaceous hosts, new resistance or tolerance genes have been characterized directly in V. vinifera. Although fungi are the pathogens on which transgenic research has focused the most efforts in recent years, the fight against other grapevine pathogens has continued (Table 4.4). Grapevine breeding has remained ineffective against viruses due to the absence of confirmed sources of viral resistance or tolerance in the Vitis germplasm (Oliver and Fuchs 2011). For this reason, genetic engineering has played an important role in the attempts to incorporate resistance to grapevine viruses using pathogen-derived resistance and RNA-mediated resistance approaches (Gambino and Gribaudo 2012). Historically the first attempts at engineering in grapevine have been made against grapevine viruses, in particular against GFLV, a nematode-transmitted icosahedral virus and causal agent of fanleaf degeneration. However, only partial successes have been reported (Gambino and Gribaudo 2012) and with the exception of to GFLV-resistant transgenic rootstocks (Vigne et al. 2004), virus-resistant engineered grapevines have rarely been obtained. In recent years, the works in this field have been significantly reduced (Dal Bosco et al. 2018), due to the poor success achieved in the past and to the bad reputation of classic genetically modified organisms (GMOs) among consumers. However, the safety of the classical transgenic approach against viruses has been confirmed recently, indeed no statistically significant differences in the genetic diversity of virus strains and microbiome were associated with GFLV-resistant transgenic rootstocks cultivated in commercial vineyard soil for several years (Hily et al. 2018). Recently Hemmer et al. (2018) proposed an interesting new antiviral approach adopting single-domain antigen-binding fragments of camelid-derived heavy-chain only antibodies, also known as nanobodies (Nbs). Nanobodies specific to GFLV expressed in Nicotiana benthamiana and in grapevine 41B rootstock, conferred effective resistance against a wide range of GFLV isolates neutralizing the virus at an early step of the life cycle, prior to cell-to-cell movement.
In addition to viruses, transgenic grapevines expressing a pear polygalacturonase inhibitory protein (PGIP) (Agüero et al. 2005) or a chimeric antimicrobial protein (CAP) (Dandekar et al. 2012) were previously obtained, and they resulted tolerant to PD, an insect-transmitted bacterial disease caused by X. fastidiosa. Dandekar et al. (2019) tested in field conditions these transgenic lines of V. vinifera cv. ‘Thompson Seedless’, and they demonstrated that PGIP and CAP secreted into the xylem can migrate into scion through the graft union. Interestingly, the tolerance to PD was observed also in the untransformed scion thanks to the transfer from the transgenic rootstock of PGIP or CAP. The trans-graft protection is then effective under field conditions, and this approach could be interesting for further application since the scion would not be transformed. Another approach against PD involved the gene rpfF isolated from both X. fastidiosa and Xanthomonas campestris pv. Campestris, encoding a synthase for diffusible signal factor (DSF). The genes expressed ectopically in ‘Freedom’ rootstock induced a reduced mobility of X. fastidiosa and in field conditions these plants showed a reduction of PD incidence two- to four-fold lower than that of untransformed plants (Lindow et al. 2014). However, Graphocephala atropunctata, one of the leafhopper vector of X. fastidiosa, showed greater colonization efficiency on DSF transgenic grapevines even though DSF plants maintained low levels of X. fastidiosa populations. These contrasting results between insect vector and bacterium suggested that under some conditions DSF transgenic plants could facilitate the X. fastidiosa spread and thus hinder the disease management (Zeilinger et al. 2018). This also highlights the importance of field tests for new genotypes obtained through transgenic and other new breeding techniques, because only in this way it will be possible to correctly evaluate all aspects related to the viticultural ecosystem.
An interesting approach has been adopted against the European grapevine moth L. botrana, which is becoming a key pest for grapevine. The insect is attracted to the odour profile of plants characterized by a specific ratio of volatile terpenoids. An (E)-β-farnesene synthase (Aaβ-FS) inserted in V. vinifera cv. ‘Brachetto’modified the emission of (E)-β-caryophyllene and (E)-β-farnesene, transgenic plants showed an alteration of the natural ratio of these compounds and these plants resulted less attractive to insect. The results suggested that the volatile ratio modification might represent a new and effective pest control strategy (Salvagnin et al. 2018). Different genes involved in the response to stress and secondary metabolism have been used against fungi. For example, VpPR4 was strongly induced in V. pseudoreticulata ‘Baihe 35–1’ after 24 h from E. necator inoculation at significantly higher levels in respect to its homologous VvPR4 in V. vinifera. The overexpression of VpPR4 in V. vinifera cv. ‘Red Globe’ under constitutive 35S promoter induced a significant reduction of E. necator hyphal growth, although a complete inhibition was not reached, since the PM resistance is likely regulated by a multi-gene complex (Dai et al. 2016). Other PR proteins isolated from V. pseudoreticulata have been shown to be effective in improving the tolerance to DM. VpPR10.1 inserted in V. vinifera cv. ‘Thompson Seedless’ did not induce macroscopic phenotypic alterations in transgenic lines, but the overexpression of this gene significantly improved DM resistance by reducing the spread and intensity of P. viticola sporulation (He et al. 2013; Su et al. 2018). VpPR10.1, interacting with a voltage-dependent anion channel 3 (DAC3), was more active in defense responses than its homologous in V. vinifera. The complex VpPR10.1/VpVDAC3 regulates the defence response to P. viticola through a cell death-related immunity mechanism (Ma et al. 2018). Another PR isolated from V. amurensis cv. ‘Zhuoshan-1’ (VaTLP) and transferred in V. vinifera cv. ‘Thompson Seedless’ under constitutive promoter improved resistance against DM by inhibiting both hyphal growth and asexual reproduction of P. viticola (He et al. 2017).
In Arabidopsis, resistance to PM 8 (RPW8) are atypical R genes without NB or LRR domains but able to induce SA-dependent responses and resistance to PM (Wang et al. 2009). RPW8 genes isolated from V. pseudoreticulatawere strongly overexpressed after P. viticola infection, while their homologs in V. vinifera were not activated by the pathogen. In particular, VpRPW8-d overexpressed in N. benthamiana enhanced resistance to Phytophthora capsici (Lai et al. 2018), and the Arabidopsis AtRPW8.2 improved resistance to PM in V. vinifera cv. ‘Thompson Seedless’. The resistance was associated with H2O2 accumulation, activation of SA signalling and altered expression of other phytohormone-associated genes (Hu et al. 2018a).
Stilbene synthases (STSs) codifying for stilbenoids in response to biotic and abiotic stresses in grapevine can increase the resistance or tolerance to different pathogens. Transgenic V. vinifera cv. ‘Thompson Seedless’ plants overexpressing VqSTS6 from V. quinquangularis showed higher stilbenoid content and enhanced resistance to PM (Cheng et al. 2016). Interestingly, if these transgenic lines were used as rootstock, high stilbenoids accumulation was observed in the untransformed scion associated with high callose deposition and increased tolerance to PM (Liu et al. 2019). The results suggested that, in addition to the leaf production, stilbenoids are also root–shoot transported, and this may be useful for inducing resistance/tolerance to wild type scions exploiting the trans-graft protection. Other STS genes VpSTS29/STS2 isolated from V. pseudoreticulata and inserted always in V. vinifera cv. ‘Thompson Seedless’ improved tolerance to E. necator. The overexpression of these genes in V. vinifera reprogrammed the transcriptome and oriented the global gene expression towards the defence, indeed the SA signalling pathway and several endogenous STS genes were transcriptionally activated in leaf-infected tissues inducing programmed cell death (Xu et al. 2019).
Calcium ions are ubiquitous second messengers in plants, and calcium-dependent protein kinases (CDPKs) are important signal transmitters of Ca2+ changes in response to environmental stimuli such as biotic ones (Hu et al. 2018b). VpCDPK9 and VpCDPK13 isolated from V. pseudoreticulata induced in overexpressing V. vinifera cv. ‘Thompson Seedless’ an increased resistance to PM involving SA and ET regulation and accumulation of H2O2 and callose in the cells surrounding the site of infection (Hu et al. 2021).
An F-box protein (VpEIFP1) induced by E. necator was isolated from V. pseudoreticulata and inserted in Arabidopisis and V. vinifera cv. ‘Red Globe’. In the transgenic plants, the accumulation of H2O2 and the increase of NPR1 and PR1 expression levels induced the suppression of E. necator germination and growth. Likely, the mechanism of action induced by VpEIFP1involved the degradation of thioredoxin z (VpTrxz) via the ubiquitin/26S proteasome system (Wang et al. 2017a).
The ubiquitination plays important roles in disease resistance in plants, and some attempts have been made to exploit this system to improve tolerance to pathogens in grapevine. V. riparia E3 ubiquitin ligase gene (VriATL156), a gene belonging to the ATL subfamily, associated with defense in Arabidopsis, was activated during P. viticola infection as one of the principal signal transduction components of the plant response to the pathogen. Transgenic V. vinifera cv. ‘Shiraz’ overexpressing VriATL156 showed a transcriptional reprogramming of the plant responses in the earliest stages of pathogen infection and provided an effective control of DM (Vandelle et al. 2021). Another example is the VpRH2, a RING-H2-type ubiquitin ligase gene from V. pseudoreticulata inserted in V. vinifera cv. ‘Thompson Seedless’ that induced an increase of PM tolerance. During the infection process, the interaction of VpRH2 with VpGRP2A, a glycine-rich RNA-binding protein, played an important role in PM-resistant gene cascades (Wang et al. 2017b).
4.12.3 Future Challenges
The concept of cisgenic plants was introduced in 2006 (Schouten et al. 2006) and refers to genetically modified plants with genes in sense orientation containing their native regulatory sequences (i.e. introns, promoter and terminator) isolated from the same species or species compatible for sexual hybridization. In addition, foreign sequences such as selection genes and vector-backbone sequences must be absent. In parallel with cisgenesis, the concept of intragenesis was also developed, which differ from the first because it allows to use new combination of functional genetic elements (gene, promoter, terminator always from species capable of sexual hybridization) obtained by in vitro cloning (Holme et al. 2013). The cisgenic approach has been taken up in several reviews (Espinoza et al. 2013; Dalla Costa et al. 2017; Limera et al. 2017; Eckerstorfer et al. 2019), although concrete examples in grapevine are almost absent. The first obstacle is to obtain marker-free plants without vector-backbone sequences. While for annual plants these sequences can be eliminated by self-fertilization and segregation in the offspring, in a woody species such as grapevine, approaches involving the use of recombinases have been adopted. The most frequently used recombinase/recognition sites are the Cre/loxP and the yeast Flp/FRT. The bacteriophage P1 Cre recombinase, which specifically recognizes loxP sites, was placed under the control of an estrogen receptor-based fusion transactivator (XVE system) and activated by 17-β-estradiol (Zuo et al. 2001). The excision system was adopted in V. vinifera cv. ‘Brachetto’ with positive results and effective excision of neomycin phosphotransferase (nptII) gene (Dalla Costa et al. 2010). In the same genotype, Flp/FRT system driven by the soybean Gmhsp17.5-E promoter effectively mediated site-specific excision of nptII gene by heat-shock treatment (Dalla Costa et al. 2016). Another obstacle to the development of the cisgenic approach in grapevine is often the lack of knowledge of the regulatory sequences of resistance/tolerance genes. However, in recent years, in addition to the classic functional analysis of the coding sequences, several studies have also been conducted on grapevine promoters (Wen et al. 2017; Tian et al. 2020; Zhang et al. 2019; Wang et al. 2020c; Vandelle et al. 2021) improving the basic knowledge in this research field. Although there are still no published articles on cisgenic grapevines for resistance to pathogens, several research groups around the world are working intensively towards this goal and cisgenic plants will be available soon in the near future. Likely, the first plants will be resistant to major fungal and oomycete pathogens (PM and DM) using the classical resistance genes, for example, MrRUN1 and MrRPV1 isolated from M. rotundifolia (Feechan et al. 2013).
The future of genetic engineering in grapevine for resistance to pathogens seems to have taken a main road in recent years, that is the study of the resistance mechanisms of grapevine species naturally tolerant to PM and DM in order to insert them in the European grapevine (V. vinifera) which is very sensitive to these diseases. In addition to classic TIR-NB-LRR resistance genes used in the traditional breeding of grapevine (e.g. Run/Ren loci), likely new tolerance genes that act against a broad spectrum of pathogens, and therefore do not confer complete resistance, but different levels of tolerance will be used. This alternative approach to classical resistance can be interesting as it would: (i) in any case reduce the impact of treatments in viticulture, (ii) ensure broad-spectrum protection against multiple pathogens (not only fungi) and (iii) make it more difficult for pathogens to overcome resistance. The road to using these engineered genotypes in the vineyard is still long as it will be necessary to: (i) improve the transformation processes, as few genotypes were currently utilised (e.g. V. vinifera cv. ‘Thompson Seedless’ seem to be the elite genotype to test the effects of transgenes in V. vinifera, Table 4.4); (ii) implement cisgenesis, an approach so far mostly ventilated rather than used in practice, the only approach with genome editing that could be more acceptable to consumers (see Table 4.5); (iii) analyze the new engineered genotypes directly in the vineyard to fully understand the levels of resistance/tolerance, the undesirable effects against the plant microbiome and the effects on the yield and the quality of the productions, aspects almost completely unknown for all transgenic grapevines produced. However, the research progress in recent years suggests a rapid overcoming of many problems related to grapevine transformation and with the support of Government Institutions, the practical use of these new-engineered genotypes might no longer be a utopia.
4.13 Recent Concepts and Strategies Developed
4.13.1 Advent of New Breeding Technologies
The term “new breeding technologies” (NBTs) refers to a group of new-generation biotechnological techniques, which resemble traditional breeding techniques but require shorter times, especially at the initial steps, and do not alter the genetic heritage of the cultivar of interest. The best-known NBTs are the “cisgenesis” and the “genome editing through site-directed nucleases”. In a cisgenic plant (see Sect. 4.12.3), only species-specific genes that could also have been transferred by traditional breeding are retained (Schouten et al. 2006). In an edited plant, a programmable nuclease will induce a double stranded break which is repaired by the cell natural repair mechanism, either non-homologous end joining (NHEJ), which may introduce nucleotide variation, or homologous direct repair (HDR) when a donor DNA with homologous arms is present (Xiong et al. 2015).
The introgression of desired traits through conventional breeding into commercial varieties of woody fruit crops with a long juvenile phase usually requires a lapse of time that can last several decades and the application of ETGM to overcome these constraints has raised a great deal of ethical criticisms. NBTs may represent a valid alternative to these approaches to produce a genetically improved plant without the introgression of foreign DNA.
Compared to other woody perennial species, the grapevine features several characteristics that make it a suitable system for the application of NBTs. This holds true in particular for genotypes used by the wine industry, represented by elite cultivars selected over the years in the various traditional viticultural areas and vegetatively propagated for decades or centuries. Cultural aspects, varietal traditions and consumer demands hinder the application of genetic engineering to these genotypes, as well as of classical breeding techniques because crossing would alter their peculiar agronomical and qualitative features. Therefore, the genetic improvement of wine grapes may gain a great benefit from NBTs, which require shorter times than traditional breeding techniques and do not alter the genetic heritage of the cultivar (Table 4.5).
Grapevine was one of the first plant species included in genome sequencing programs and a well annotated, constantly improved reference genome has been publicly available since 2007 (Jaillon et al. 2007). This important achievement has allowed a large number of transcriptomic and gene functional analyses, which represent a valuable base of knowledge facilitating the selection of candidate genes and the application of NBTs. Indeed, many efforts have already been made to identify interesting genes to be engineered with both ETGM and NBTs (Capriotti et al. 2020). Moreover, protocols for culturing and transforming cell suspensions, hairy roots or embryogenic calli and for regenerating whole plants starting from various explants, including protoplast, have been set up and improved to accomplish the specific requirements of some cultivars (Bertini et al. 2019; Dalla Costa et al. 2019; Scintilla et al. 2021).
However, there are still several technical challenges, which may hamper the application of cisgenesis and genome editing to the grapevine. The setting-up of efficient tissue culture procedures requires further development and optimization of several technical aspects, especially for many economically important élite cultivars proved recalcitrant to gene transfer and/or regeneration (Gribaudo et al. 2017). Another limiting factor is the chimerical integration of exogenous DNA and somaclonal variation as an outcome of tissue culture. In grapevine, somaclonal variation is frequently observed among plants regenerated through somatic embryogenesis, resulting in a wide range of traits regarding chlorophyll deficiencies, morphogenetic development, leaf shape and flower type (Martinelli and Gribaudo 2001). An important aspect to take in consideration for the choice of the target sequence is the high heterozygosity of the grapevine genome and the high degree of intraspecific genetic variation among cultivars and accessions. Finally, despite the many efforts made to explore the grapevine genome, the list of interesting candidate genes to be edited is still small and further studies are needed to enlarge the number of candidates, sufficiently characterized to be an object of NBT application.
Given the above mentioned limitations, no NBT-derived grapevines have been obtained to date. However, considering the rapid biotechnological advancements achieved in the last few years, the successful production of cisgenic and DNA-free genome edited vines is around the corner. A primary goal for cisgenesis could be the transfer of genes which confer resistance to the major fungal and oomycete pathogens in cultivated grapevine (PM and DM) (Capriotti et al. 2020; Dalla Costa et al. 2019), whereas for the genome editing approach could be the silencing of susceptibility genes, whose knock-down confer resistance to pathogens (Pirrello et al. 2021). An up-to-date overview of the attempts to apply genome editing is presented in the next sections.
4.13.2 CRISPR/Cas System for Gene Editing
Genome editing is carried out using sequence-specific nucleases, including zinc finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs) and, more recently, the CRISPR/Cas9 system (Yin et al. 2017). The double-stranded breaks induced by the sequence-specific nucleases at targeted genome sites are generally repaired by NHEJ or HDR, which lead to gene knockout or gene replacement, respectively. In particular, CRISPR/Cas9 technology is a cutting-edge approach comprising a Cas9 effector protein and a single guide RNA (sgRNA). The Cas9 nuclease is guided to the target DNA by the sgRNA which contains a sequence that matches the sequence to be cleaved. Essential for cleavage is a three-nucleotide sequence motif (NGG) immediately downstream on the 3’ end of the target region, known as the protospacer-adjacent motif (PAM). RNA-guided Cas9 creates site-specific double-stranded DNA breaks, which are then repaired by either non-homologous end joining or homologous recombination (Jinek et al. 2012).
Prime editing is the most recent CRISPR genome-engineering tool, described as a “search-and-replace” genome editing technology, that represents a novel approach to expand the scope of donor-free precise DNA editing to not only all transition and transversion mutations, but small insertion and deletion mutations as well (Kantor et al. 2020). Prime editing involves a longer-than-usual gRNA, known as pegRNA, and a fusion protein consisting of Cas9 H840A nickase fused to an engineered reverse transcriptase (RT) enzyme. The Cas9 element of the prime editor digests the genomic DNA and the RT element polymerises DNA onto the nicked strand based on the pegRNA sequence (Matsoukas 2020).
Genome editing in tree crops by CRISPR/Cas9 is an emerging field and in grapevine very few examples of successful application to dissect gene functions and enhance plant traits have been reported. The first proof-of-concept was provided by Ren et al. (2016) who modified the metabolism of tartaric acid. The mutation of the L-idonate dehydrogenase (IdnDH) enzyme was obtained by stable integration of the genetic components of CRISPR/Cas9 system through A. tumefaciens gene transfer of ‘Chardonnay’ suspension cells. In 2017, ‘Neo Muscat’ somatic embryos were transformed with a CRISPR/Cas9 editing construct targeting the phytoene desaturase gene and plants with albino leaves were produced (Nakajima et al. 2017). In 2020 embryogenic grape cells derived from 41B rootstock were used to knock out CCD8 genes, involved in the control of shoot architecture in grapevine (Ren et al. 2020). Wang et al. (2018b) generated transgenic ‘Thompson Seedless’ lines with biallelic mutation of the WRKY52 transcription factor, with improved resistance to noble rot caused by B. cinerea. The authors therefore provided the first strong evidence in using CRISPR/Cas9 to enhance disease resistance in grapevines. Subsequently, Sunitha and Rock (2020) produced resistant transgenic plants with CRISPR/Cas9 targeting TAS4b, a molecular determinant of GRB virus and PD host susceptibility, and the anthocyanin regulator MYBA7 in the 101–14 rootstock. Wan et al. (2020) showed that CRISPR/Cas9 MLO3-edited grape lines had enhanced resistance to grapevine PM, and Li et al. (2020) reported that PR4b loss-of-function lines had decreased resistance to P. viticola.
Still, the wide application of genome editing to grapevine faces many challenges, above all, the previously mentioned bottlenecks of the length and the variety-specific efficiency of the regeneration process. In addition, a general constraint is represented by the need to improve several aspects for a precise and efficient editing. Ren et al. (2021) demonstrated that the use of the identified U3/U6 and UBQ2 promoters could significantly increase the editing efficiency in grapevine by improving the expression of sgRNA and Cas9, respectively. Moreover, this study represented the first example of multiplex genome editing in grapevine through the simultaneous editing of the sugar-related tonoplastic monosaccharide transporter (TMT) family genes TMT1 and TMT2. The occurrence of the off-targets, which are highly undesired, was recently investigated by Wang et al. (2021) in grapevine, showing that their frequency is likely insignificant compared with variations caused by tissue culturing and/or Agrobacterium infection. However, the use of Cas variants with higher specificity, accuracy and ability to recognize different PAM sequences, may further reduce the occurrence of these unfavorable events. The requirement of a specific PAM site (NGG) adjacent to the target site limits the number of potential targets. Up to now, five types of CRISPR/Cas9 target sites have been identified and characterized and a user-friendly database for editing grape genomes has been developed (Wang et al. 2016b).
Additional general drawbacks of CRISPR/Cas9 technology are represented by the pleiotropic effects associated with the knockout of target genes, the challenging of gene knock-in approach compared to knock-out and the need of a biallelic editing in the case of recessive mutation. These aspects represent a severe limitation in grapevine, species in which functional analysis has been performed on a relatively small number of genes. An alternative strategy to limit pleiotropic effects associated with an abolished gene function is represented by the quantitative regulation of gene expression achieved with genome editing on cis-regulatory elements (Rodríguez-Leal et al. 2017).
For selecting the best target sequences the grapevine reference genome can be used. However, after this step, a subsequent sequencing of those regions in the genotypes of interest is needed to avoid bumping into SNPs which can prevent an optimal recognition by the endonuclease and consequently the DNA cleavage.
4.13.3 Towards the Generation of Transgene-Free Resistant Grapevines
The adoption of NBTs might represent a revolutionary cutting-edge in worldwide grapevine breeding and cultivation, but this requires government support in setting up an updated regulatory framework. Some non-European countries (e.g. USA and Australia) have established that if no foreign DNA is present in a genome edited variety it will not be subject to additional regulation and risk assessment, whereas European Union strongly reaffirmed the precautionary principle and genome edited organisms are currently included in the strict legal framework of genetically modified organisms. Yet, a recent document by the EU announced the revision of the legal status of NBT products (https://ec.europa.eu/food/sites/food/files/plant/docs/gmo_mod-bio_ngt_exec-sum_en.pdf) which may facilitate their diffusion in the near future. In any case, the absence of foreign DNA is mandatory for complying with legal clues on organisms subjected to NBTs. The production of DNA-free edited plants is mainly based on two strategies (Zhou et al. 2020). The first one involves the stable integration in the genome of the CRISPR/Cas9 genetic components through A. tumefaciens T-DNA gene transfer and their subsequent removal. While for annual plants these components can be eliminated by self-fertilization and segregation in the first generation of offspring, in case of vegetatively propagated woody fruit crops this could be achieved by the use of site-specific recombination systems which leads to the excision of the foreign DNA.
In grapevine, some preliminary studies successfully tested the mechanisms for the removal of foreign DNA. Dalla Costa et al. (2016) set up the experimental conditions for the application of a heat-shock treatment to activate the site-specific recombinase Flippase (Flp) which recognizes the target sequences. Heat-shock induction for the removal of the kanamycin resistance gene nptII was carried out on genetically modified ‘Brachetto’ in vitro lines. The same research group recently proposed an alternative approach based on heat-shock treatment (Dalla Costa et al. 2020). The procedure exploits the cleavage activity of Cas9 not only to edit the endogenous target site, but also to remove foreign DNA by placing two additional synthetic target sites next to the left and right borders. In this study the CRISPR/Cas9 system was used to knock-out the MLO7 gene involved in the susceptibility of PM and a very detailed analysis the bacterial T-DNA molecular features at the insertion site, crucial for the excision of T-DNA cassette, was performed.
The second strategy to obtain DNA-free edited plants is based on the transient expression of the CRISPR/Cas9 components from non-integrating foreign DNA, not yet applied in grapevine, or on the direct delivery of the in vitro assembled ribonucleoprotein (RNP) composed by purified Cas9 protein and gRNAs. This approach was adopted by Malnoy et al. (2016) demonstrating that the direct delivery of CRISPR/Cas9 RNPs to ‘Chardonnay’ protoplasts enables targeted gene editing, despite whole plants from edited protoplasts could not be regenerated. Subsequently, Osakabe et al. (2018) provided a stepwise protocol for the design and transfer of CRISPR/Cas9 components to apple and grapevine protoplasts, and Ren et al. (2019) helped to optimize CRISPR/Cas9 performance in grape by showing that sgRNAs with high GC content improved editing efficiency in grapevine and that editing efficiency also depends on selecting the appropriate cultivar. However, several additional important technological steps still need further development. For instance, a selectable marker-free method is required for the recovery of edited plants at high frequencies. Furthermore, the delivery of RNP complexes via protoplasts transfection is limited only to some plant species and in grapevine the regeneration process from single cell is a major bottleneck. The isolation of protoplasts from embryogenic grapevine tissue and the regeneration of these protoplasts into plants were successfully reported in few early works for two V. vinifera cultivars ‘Seyval blanc’ (Reustle et al. 1994) and ‘Koshusanjaku’ (Zhu et al. 1997). Recently, a stepwise protocol for the regeneration of whole plants from embryogenic callus-derived protoplasts of two Italian varieties, ‘Garganega’ and ‘Sangiovese’ was developed (Bertini et al. 2019). These works paved the way to further specific development of regeneration protocols, broadening the spectrum of varieties prone to be edited by CRISPR/Cas9 technology.
4.13.4 Future Challenges
Conventional breeding has been fundamental for grapevine domestication and improvement, resulting in a huge amount of different varieties adapted to the cultivation in a wide range of pedoclimatic conditions. However, due to the low accessibility of desirable variations or gene combinations, cross and mutation breeding of new cultivars requires the production and analysis of large numbers of offspring, and takes a long time. The gene editing technology developed in recent years allows precise engineering of desirable variants with unprecedentedly high efficiency and resolution, greatly expanding the range of variations available and reducing our reliance on naturally existing mutations. Efficient transformation and regeneration procedures have been established for a number of grape varieties and the strength of the artificially generated variations driven by CRISPR/Cas9 technology has been demonstrated. However, despite several biotechnological approaches covering all the necessary steps, they need to be combined in complete procedures for the generation of cisgenic or edited plants. Finally, the adoption of high-throughput techniques, still applied to a limited extent to the grapevine, could help characterize the phenotype of plants obtained by NBTs in the vineyard and in a controlled environment, increasing the objectivity, automation and precision of data collected. A new biotechnology-driven revolution in viticulture could be just around the corner.
4.14 International Hybrid Regulations: Status and Background
4.14.1 Hybrids in Viticulture
The use of hybrids in viticulture is under continuous discussion, even if this sector has a big challenge to maintain sustainable production under climate change scenarios and foster its adaptation. The use of new cultivars could be a medium-long term adaptation technique and provide new solutions to reducing pesticides, controlling vine decay, water management and stress, etc. Hybrids were used in the past for trying to solve relevant problems, like the phylloxera plague in Europe during the second half of the nineteenth century. It should be remarked that one of the possible solutions discussed was hybridization because American species are often naturally phylloxera-resistant or tolerant (V. aestivalis, V. rupestris, V. riparia and V. labrusca). Even if this problem was generally solved by grafting American rootstock with European varieties, some areas had a great dispersion of these American direct producer vine varieties and hybrids were so successful (see Sect. 4.6.1). For instance, in 1958, 30% of French vineyards (400,000 ha) were covered by fungus-resistant grape varieties or in Pontevedra (Galicia, Spain) in 1983, covering 73% of total vineyard area (Martínez and Pérez 1995). Nevertheless, due to organoleptic (insufficient quality, foxy aroma and strawberry flavours in wine), compositional (high methanol or malvidin content) or ethical (genetic modifications) questions, not all countries accept this material for making commercial wines.
4.14.2 Hybrid Wine Profiles
American and Asian species are repeatedly backcrossed to V. vinifera cultivars with the main objective of transferring their resistances (mainly powdery and downy mildew) into the vinifera genetic background, restoring or improving their traits and of course, maintaining their oenological aptitude (de la Fuente 2018). A typical characteristic of the red wines produced from these cultivars is the malvidin content in their wines. Malvidin is an anthocyanin (principal red colouring matter), present in the form of malvidin-3,5-diglycoside, and being a common natural compound in the red grapevine skins, together with the other coloring pigments. Due to this discussion, the OIV defined in this “compendium of international methods of wine and must analysis”, a clear methodology for analysing malvidin glucoside (OIV-MA-AS315-03; 377/2009) content and fixed the maximum acceptable malvidin limit (15 mg/L) for a wine. Besides this, the true situation is that new cultivars or some hybrids mainly based on V. vinifera, but with one or more backcrossed genes from wild American species from breeding programs (e. g. ‘Regent’), even if they are classified as V. vinifera, they exceed the OIV threshold for malvidin diclucosides. Some cultivars are rejected for the production of quality wine at the European level (Art. 81 b CE 1308/2013). This discussion is never ending at the international level, but it should keep on being discussed because scientific limits are not well applied to this topic at present, and OIV standards usually help to harmonize these questions.
4.14.3 Evidence of Hybrid Utilization Worldwide
Despite the above mentioned problems concerning hybrid wines, the use of hybrids is common in non-European countries. For instance, sparkling wines in Brazil are frequently based on V. labrusca and hybrids (Caliari et al. 2014), being in some regions more than 90% of the total vineyard area for wine production. It should be noted that Brazil has a total surface vineyard of 79.094 ha, in which close to half are planted with hybrids (data source OIV; 2015). China, which started in the 1950s a great breeding program facing the cold resistance for wine and fresh grapevine varieties, has arisen more than 70 new cultivars, today which means thousands of ha (Li 2014). Furthermore, the Korean hybrid grape cv. ‘Cheongsoo’ was selected in 2005 and spread for its good winemaking performance (Chang et al. 2014; Kim et al. 2015).
In the United States, several breeding programs have released well-adapted wine and table grape cultivars with improved disease and virus resistance (Reisch and Reynolds 2015), and these have been widely adopted, especially in regions where it is more difficult to grow V. vinifera cultivars. Additionally, the University of California, Davis, has released five new winegrape cultivars derived from a PD resistant genotype of V. arizonica. There are no regulations restricting the planting of any of these new cultivars, other than the need to register the name with the U.S. Tax and Trade Bureau if the name is intended for use on a wine label (https://www.ttb.gov/wine/grape-variety-designations-on-american-wine-labels).
Even in Europe, there are some interspecific hybrids produced by crossing V. vinifera and other species, which can provide certain traits of Vitis spp. The complex hybrid ‘Aletta’ was qualified in 2009 by the Hungarian register, and in 2012 the surface area was 423 ha and now has risen to 1,300 ha (Hajdu 2015). France created the national observatory for the development of resistant vine varieties in 2016 (OsCar; https://observatoire-cepages-resistants.fr/en/), a participatory network under a minimum of 6 years program for studying and validating the potential of these new cultivars in different locations and their adaptation before establishing them as new commercial varieties, under a temporary or final registration. This observatory is a great example of public and private collaboration between research centres (INRAE-IFV) and producers associations, linked to the French denomination of origin institution (INAO). The collaboration resulted in four INRA ResDurI cultivars (‘Voltis’, ‘Artaban’, ‘Vidoc’, and ‘Floreal’), which have a significant impact to the French wine sector. Germany also has more than 50 years of breeding programs with some cases of success (e.g. PIWI family). Cooperative (VCR) breeding efforts in Italy resulted in several new resistant cultivars currently being plant in Europe and North America, including ‘Fleurtai’, ‘Soreli’, ‘Merlot Kanthus’, and ‘Cabernet Volos’. Including the four new varieties ‘Termantis’, ‘Nermantis’, ‘Valnosia’, and ‘Charvir’ recently released by FEM, a total of 31 mildew resistant/tolerant cultivars are now registered at the Italian grapevine variety catalogue (http://catalogoviti.politicheagricole.it/catalogo.php).
4.14.4 Regulation Framework
This complexity of existing and used Vitis species (see Sect. 4.5.1) offers a great number of different inter- and intra specific crossings and adds difficulties to establish definitions and regulations on this topic. Under this composite framework, the OIV is working (since 2016) on defining relevant concepts concerning the classification of grapevine material used. The genetic resources and vine selection (GENET) group of experts from Commission has drafted the first version of this document (VITI-GENET 19–610), including a provisional definition of cultivated variety, grapevine variety, clone, direct producer hybrid, etc. This will be a relevant work from the experts, and it could have a great impact on the world wine sector. Some discussions remain at the group and by now this project resolution is kept at step 3. Together with this resolution, Commission I is also working on another draft concerning management of plant material exchanges: VITI 14–565. OIV Guidelines for production and exchange of viticultural plant material: phytosanitary and genetic aspects. OIV has recently approved other resolutions, which have a real impact on the selection process: OIV process for the clonal selection of vines (OIV-VITI 564A-2017) and OIV process for the recovery and conservation of the intravarietal diversity and the polyclonal selection of the vine in grape varieties with wide genetic variability OIV-VITI 564B-2019). The first one is the updated version of the previous resolution (OIV-VITI 1–1991) including one definition for the selected clone: “A clone is the vegetative progeny of a single vine plant. For selection purposes this single plant is chosen for its varietal identity, its phenotypic traits and its sanitary state”. Other definitions should be addressed in future resolutions as mentioned before. The second one defines the polyclonal selection protocol as a process for the recovery and conservation of intravarietal diversity and polyclonal selection of grapevines on vines with wide genetic variability. In annex I, this resolution has a glossary of concepts that could be used at an international level, but this glossary should only be applied within the scope of this resolution.
At the European level, the use of hybrids for wine production under a protected designation of origin (PDO) label is currently banned, however, the interest in these varieties and wines produced, either under the protected geographical indication (PGI), without a quality seal or in third countries is increasing (Aranda and Armengol 2019).
Since 2013, the regulation on PDOs and PGIs has been established by the Parliament and European Council Regulation (EU) 1308/2013, establishing the common market organization (CMO) for agricultural products, and specifically for wine products (in Articles 92 to 111). Wines labelled under a PDO must be produced by a V. vinifera variety (Article 93) and wines qualified under a PGI, must be produced by V. vinifera or hybridizations between V. vinifera and other species of Vitis genus. Even more, some specific hybrids are detailed in this legislation as forbidden varieties for wine production: ‘Noah’, ‘Othello’, ‘Isabella’, ‘Jacquez’, ‘Clinton’, and ‘Herbemont’ (Article 81), without any label consideration. In some Countries, new disease resistant cultivars are registered as V. vinifera.
4.14.5 New Regulations and Perspectives
During recent years, a new common agricultural policy (CAP) and therefore a new CMO, are being drafted and discussed by European countries, and they are expected in 2023 (at the latest). Discussion is on the table, and some grapevine varieties for wine production could be allowed, with the main objective to “increase resistance to diseases and improve the grapevine adaptation to climate change”. In that case, some hybridizations or backcrossed varieties from V. labrusca or others could be planted and produced. In 2018, within the proposal on the CMO regulation, some relevant modifications were proposed to the Commission, including the possibility to accept the six forbidden hybrids. On the positive side, these varieties could be beneficial for both producers and consumers, by significantly limiting the amount of pesticides used and therefore, having a positive impact on the environment and profit margins of the farmers. On the other hand, it was underlined that opening of these wine grape varieties belonging to V. labrusca and of the six forbidden varieties would decrease the quality of the wine products and therefore, could affect the reputation of European wines.
Discussions showed a clear difference of opinion between the main wine-producing countries (strong and motivated opposition) and the rest of the Member States (more flexible or keen to accept it). This point is still being discussed today. Disease resistant vines and future vineyards underline the role of further research and innovation in the sector and the need to (a) find new genetically resistant varieties to reduce the negative aspects linked to the use of pesticides or (b) explore current varieties for wine production with enough genetic variability, necessary to face adaptation needs. With current technological advances and specialized knowledge, there is a large potential for R&D in viticulture, boosting some sustainable varieties which would adapt to agronomic conditions and which could be properly selected with wine tasting quality criteria. It should be remarked that the OIV definition of wine is as follows: “Wine is the beverage resulting exclusively from the partial or complete alcoholic fermentation of fresh grapes, whether crushed or not, or of grape must” and the official OIV grapevine varieties and synonyms database does not differentiate between V. vinifera and other cultivars (de la Fuente 2018).
4.15 Future Perspectives
Growers can now utilize grape varieties (from inter- and intraspecific hybridization) and rootstocks (normally hybrids of American Vitis spp.) with tolerance to biotic stresses. This resilience can be further improved by breeding programmes selecting new clones and producing new varieties and rootstocks. The history of grapevine breeding for disease resistance (mostly for wine grapes) began with the French-American hybrids and demonstrated the technical, societal and regulatory hurdles that need to be overcome (Bavaresco 2019). Climate change will likely push forward the demand for wine and table grape varieties with better adaptation to warmer and likely drier climates, and for rootstocks with better adaptation to drought and salinity. For wine and table grapes this process may depend on gene editing techniques capable of altering gene expression to improve fruit and wine quality under new climates. At any rate, it seems likely that climate change will spur on the discussion of whether and where we will need better adapted fruit and rootstock varieties, and what breeding and improvement strategies we will use to achieve these goals.
Considering fruiting varieties, about the technical aspects, a wider exploitation of intravarietal variation is needed, particularly of the ancient V. vinifera varieties; we cannot select resistant or tolerant clones, but less susceptible individuals are known and could be exploited in clonal selection programs. Moreover, a larger utilization of all the available V. vinifera germplasm should be encouraged, because unexpected findings do occur, such as the discovery of strong PM resistance genes in Central Asian vinifera. Traditional breeding methods have been improved and time-consuming procedures utilized for classical hybridization have been improved and accelerated by the application of MAB approaches, and may improve further with the application of transgenesis, cisgenesis, and genome editing. Marker-assisted breeding is a very useful if precise phenotyping analysis is performed. Phenomics is a promising approach with the capacity to greatly improve breeding progress. It will be crucial for exploiting and linking in a correct way the genomic information to the plant’s physiology and behaviour. On the other hand, only a thorough genetic characterization will allow for an understanding of which genes are involved in the resistance response of the plant. Marker-assisted selection currently supports both the choice of the parents to be hybridized and the selection of the proper individuals in the segregating populations (progeny). Regarding the selection of the new progeny, MAS reduces the number of seedlings to be grown, but field trials are still necessary to verify the behaviour of the new plants under different growing conditions. Since MAS allows a reduction in the number of seedlings to be grown, a larger number of seeds can be managed and this is positive because the higher the number of seeds, the greater the possibility of selecting the optimum individuals. Moreover, within the whole “economy” of a specific breeding program, MAS will allow many more pathogen and pest resistance evaluations to be conducted, and will allow greater progress beyond resistance to PM, DM and PD. It is crystal clear that for a thorough genetic improvement a comprehensive program is required, linking a variety of ad hoc programs. Multiple parallel programs shall consider both quality- and resistance-related traits, focusing on a final shared goal. It will be challenging to obtain a new individual completely and durably resistant to all the pathogens and pests but, even though a few treatments are required, the overall impact of resistant varieties will be a healthier environment and populace, and a more sustainable wine industry. Another aspect to be emphasized is the need to develop local breeding programs in order to obtain new individuals well adapted to that specific environment. In fact, other pedoclimatic conditions (with different pathogen/pest races and pressures) might modify the vine response in terms of both grape quality (ratio sugars/acids, aromas, etc.) and of disease resistance. Thus far, the reaction of wine growers and makers to new disease resistant wine varieties has not been highly appreciative because of the relatively poor quality of the earliest hybrids when compared to traditional pure vinifera varieties. Marker-assisted breeding has the potential of maintaining high resistance by following marker expression over multiple generations, allowing the breeder to focus on fruit and wine quality in the later generations when the percentage of vinifera in the advanced selections is high. Future breeding programs for biotic stress resistance, in which breeders can focus on fruit and wine quality while selecting in the early generations, should carefully select against poor quality fruit characteristics and for high wine quality. The NBTs, especially genome editing, are very promising methods to help with the development of improved fruit and wine quality through the alteration of gene expression and biochemical pathways. We are at the dawn of a new (scientific) revolution, enabling scientists to alter plants for suitable and sustainable cultivation. The potential “plus” given by genome editing consists of the possibility to rearrange only the disease/pest resistant genes without manipulating the other traits of the plant, especially those related to the quality parameters for wine and table grapes. Gene editing may have a very large impact on the expression of biochemical by-products we consider responsible for high wine quality. However, a thorough understanding of the genome and phenome and very effective phenotyping and genotyping techniques will be needed.
When societal issues are considered, few problems are likely to arise with the selection of new clones, while the acceptance of new disease resistant or gene-edited varieties may be problematic for wine grapes, regardless of the technologies with which they were obtained. Table and raisin grapes, being a regular fruit, may encounter fewer issues, if obtained by the CBTs. In addition, new wine variety names might create mistrust among consumers, but only if the wine is sold with the variety name; by contrast, for table grapes a new name might be very attractive. Much more problematic will be dealing with the NBTs, for both wine and table grapes. Although genome/gene editing techniques do not transfer foreign genetic material and are based on the use of molecular scissors (biological mutation), they do recall a genetic manipulation and therefore a hostile reproach by the public is likely to occur, even in the countries where NBTs will be allowed. That is why education as well as open dialogue between scientists and the public on a rational basis is relevant. All of the participants in the wine chain, including retailers and the consumers, will need to be convinced of the science behind gene editing for this technology to become a real innovation. Moreover, there is the need to move the policy discussion from the national/international level to local communities, which will be the first to feel the context-dependent impacts of any release. In other words, we need collective oversight (Kofler et al. 2018).
Concerning the regulatory aspects, new disease resistant wine grape varieties will have no restriction in countries outside the EU, while in EU they will be allowed to produce table and PGI wines and most likely also PDO wines. In the next review of the wine CMO, clearer provisions will be established on this topic. In France, for instance, INAO created a new category of wine grape varieties called “grapes for climate and environmental adaptation” (including the new disease resistant varieties) to be used in the production of PDO wines and approval is expected by the EU in the next CMO. In the case of new resistant varieties for table/raisin grapes, no international cultivation restrictions are present, and in the countries with a national grapevine register (NGR), their registration is only mandatory to allow propagation by nurseries. Derived from both classical breeding and MAB, a new grape variety can undergo the patent process. In the EU, the community plant variety office (CPVO), a self-financed EU agency, is responsible for the management of the community plant variety rights system, covering the 27 Member States. Located in Angers, France, the CPVO was created by the Council Regulation 2100/94Footnote 1 and has been operational since April 1995. The grape varieties (including the resistant ones) protected by plant variety rights are registered and can be found in the CPVO database (https://cpvoextranet.cpvo.europa.eu/Denominations). On paper, it is impossible to distinguish the resistant varieties from others, because they are registered as V. vinifera. A new vine coming from the NBTs will be most likely considered as a clone, bearing precise distinguishable characteristics.
Considering rootstocks, the perspectives are to improve the resistance/tolerance toward current/emerging pathogens and pests by both CBTs, ETGMs and NBTs. No societal problems are expected to occur with NBTs since the rootstocks are not the fruit bearing part of the plant. Concerning the regulatory aspects, the registration of new rootstocks is under the same requirements as for the fruiting varieties (Directive 2004/29/CE of 4 March 2004). The nurseries (in the EU) have some rules to follow, i.e. to propagate genotypes registered in the NGR while the grape growers can use the rootstock they wish, without any restriction.
Notes
- 1.
COUNCIL REGULATION (EC) No 2100/94 of 27 July 1994 on Community plant variety rights. (OJ L 227, 1.9.1994, p.1).
Abbreviations
- Aaβ-FS:
-
(E)-β-farnesene synthase
- AGO:
-
ARGONAUTE protein
- AM:
-
Association mapping
- AmpSeq:
-
Amplicon sequencing
- ATL:
-
Arabidopsis tóxicos en levadura
- ATP:
-
Adenosine triphosphate
- AUDPC:
-
Area under the disease progress curve
- AWPM:
-
Area wide pest management
- BAC:
-
Bacterial artificial chromosome
- BBR:
-
Botrytis bunch rot
- BCA:
-
Biological control agents
- BES:
-
BAC end sequence
- BLAST:
-
Basic local alignment search tool
- BN:
-
Bois noir
- BR:
-
Black rot
- BSA:
-
Bulked segregant analysis
- BS-seq:
-
Bisulfite sequencing
- bZIP:
-
basic Leucine zipper
- CAP:
-
Common agricultural policy
- CAP:
-
Chimeric antimicrobial protein
- CBF2/DREB1C:
-
C-repeat-binding factor dehydration-responsive element-binding factor 1C
- CBTs:
-
Conventional breeding technologies
- CCD:
-
Carotenoid cleavage dioxygenase
- CDPKs:
-
Calcium-dependent protein kinases
- CGAS:
-
Candidate gene association study
- ChIP-seq:
-
Chromatin immunoprecipitation followed by sequencing
- chr:
-
chromosome
- CIVIT:
-
Consorzio Innovazione Vite
- CMO:
-
Common Market Organization
- CNV:
-
Copy numbers variation
- CPVO:
-
Community Plant Variety Office
- CRISPR/Cas9:
-
Clustered regularly interspaced short palindromic repeats-associated protein 9
- cv.:
-
cultivar
- DAC3:
-
voltage-dependent anion channel 3
- DAMP:
-
Damage-associated molecular pattern
- DCL:
-
DICER-like
- ddRAD-seq:
-
double-digested RAD-seq
- DDS:
-
Decision support system
- DLO :
-
DMR-like oxygenase gene
- DM:
-
Downy mildew
- DMI:
-
Demethylase inhibitor
- DMR :
-
Downy mildew-resistant gene
- DREBs:
-
Dehydration-responsive element-binding proteins
- DSF:
-
Diffusible signal factor
- DSS:
-
Decision support system
- EcoTILLING:
-
Ecotype-Target induced local lesions in genomes
- ELISA:
-
Enzyme-linked immunosorbent assays
- EPPO:
-
European and Mediterranean Plant Protection Organization
- eQTL:
-
expression QTL
- ERFs:
-
Ethylene-responsive factors
- EST:
-
Expression sequence tag
- ET:
-
Ethylene
- ETGMs:
-
Established techniques of genetic modification
- ETI:
-
Effector-triggered immunity
- EU:
-
European Union
- FAIR:
-
Findability, accessibility, interoperability and reusability
- FAO:
-
Food and Agriculture Organization
- FAOSTAT:
-
Food and Agriculture Organization Corporate Statistical Database
- FD:
-
Flavescence dorée
- FDR:
-
False discovery rate
- FEM:
-
Fondazione Edmund Mach
- Flp:
-
Flippase
- Flp/FRT:
-
Flippase recombinase/Flippase recognition target sites
- GAB:
-
Genomics-assisted breeding
- GBS:
-
Genotyping-by-sequencing
- GC:
-
Guanine-cytosine
- GEM:
-
Grape erineum mite
- GENET:
-
Genetic resources and vine selection
- GFLV:
-
Grapevine Fanleaf Virus
- GINV:
-
Grapevine berry Inner Necrosis Virus
- GLRaV:
-
Grapevine Leafroll associated Virus
- GMO:
-
Genetically modified organisms
- GO:
-
Gene Ontology
- GPGV:
-
Grapevine Pinot gris Virus
- GRBV:
-
Grapevine Red Blotch virus
- GRSPaV:
-
Grapevine Rupestris Stem Pitting associated Virus
- GS:
-
Genomic selection
- GTD:
-
Grapevine trunk diseases
- GVA:
-
Grapevine Virus A
- GVB:
-
Grapevine Virus B
- GWA:
-
Genome wide association
- GWAS:
-
Genome wide association studies
- GxE:
-
Genotype x environment interaction
- GY:
-
Grapevine yellow
- H2O2:
-
Hydrogen peroxide
- H3K27me3:
-
Histone 3 lysine 27 trimethylation
- H3K43me3:
-
Histone 3 lysine 43 trimethylation
- HDR:
-
Homologous direct repair
- HM:
-
Histone modifier
- HR:
-
Hypersensitive response
- IdnDH:
-
L-idonate dehydrogenase enzyme
- IFV:
-
Institut Français de la Vigne et du Vin
- IGGP:
-
International Grapevine Genome Program
- INAO:
-
Institut national de l'origine et de la qualité
- INRA:
-
Institut National de la Recherche Agronomique
- INRAE:
-
France's National Research Institute for Agriculture, Food and Environment
- IPCC:
-
Intergovernmental panel on climate change
- IPM:
-
Integrated pest management
- ITS:
-
Internal transcribed spacer regions
- JA:
-
Jasmonic acid
- JAZ:
-
JASMONATE-ZIM DOMAIN protein
- KASP:
-
Kompetitive allele-specific PCR
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- LAMP:
-
Loop-mediated isothermal amplification
- LD:
-
Linkage disequilibrium
- LRR:
-
Leucine-rich repeat
- MAB:
-
Marker assisted breeding
- MABC:
-
Marker-assisted backcrossing
- MAMP:
-
Microbe-associated molecular pattern
- MAPS:
-
Marker-assisted parent selection
- MAS:
-
Marker-assisted selection
- MASS:
-
Marker-assisted seedling selection
- Mb:
-
Megabases
- MBC:
-
Modified backcross
- MD:
-
Mating disruption
- MIAPPE:
-
Minimum information about a plant phenotyping experiment
- miRNA:
-
microRNA
- MLM:
-
Mixed linear model
- MLMM:
-
Multi-locus mixed model
- MLO :
-
Mildew locus O gene
- mQTL:
-
metabolite QTL
- MSAP:
-
Methylation sensitive amplification polymorphism
- mtmlSEM:
-
multi-trait multi-locus Structural equation modeling
- MTMM:
-
Multi-trait mixed model
- NBS:
-
Nucleotide-binding site
- Nbs:
-
Nanobodies
- NBTs:
-
New breeding techniques
- NCBI:
-
National Center for Biotechnology Information
- ncRNA:
-
non-coding RNA
- NGR:
-
National Grapevine Register
- NGS:
-
Next-generation sequencing
- NHEJ:
-
Non-homologous end joining
- NO:
-
Nitric oxide
- NPTII:
-
Neomycin phosphotransferase
- nSSR:
-
nuclear SSR
- OIV:
-
Organisation Internationale de la Vigne et du Vin
- ORF:
-
Open reading frame
- PAM:
-
Protospacer-adjacent motif
- PAMP:
-
Pathogen-associated molecular pattern
- PCLS:
-
Phomopsis cane and leaf spot
- PCR:
-
Polymerase chain reaction
- PD:
-
Pierce's disease
- PDO:
-
Protected designation of origin
- pegRNA:
-
Prime editing guide RNA
- PGI:
-
Protected geographical indication
- PGIP:
-
Polygalacturonase inhibitory protein
- PIWI:
-
pilzwiderstandsfähig
- PM:
-
Powdery mildew
- PNN:
-
Plant-parasitic nematode
- PR4b:
-
Pathogenesis Related 4b protein
- PR:
-
Pathogenesis-related protein
- PRR:
-
Pattern recognition receptor
- PTI:
-
PAMP-triggered immunity
- pv.:
-
pathovar
- QoI:
-
Quinone outside inhibitor
- qPCR:
-
quantitative PCR
- QTL:
-
Quantitative trait locus
- QTLs:
-
Quantitative trait loci
- QTN:
-
Quantitative trait nucleotide
- RAD-seq:
-
Restriction site-associated DNA sequencing
- RAPD:
-
Random amplified polymorphic DNA
- RDR:
-
RNA-dependent RNA polymerase
- RGA:
-
Resistance gene analog
- R-gene :
-
Resistance gene
- rhAmpSeq:
-
RNase H2 enzyme-dependent amplicon sequencing
- RISC:
-
RNA-induced silencing complex
- RLK:
-
Receptor-like protein kinase
- RNA-seq:
-
RNA sequencing
- RNP:
-
Ribonucleoprotein
- rpfF:
-
regulation of pathogenicity factor F
- RPW8 :
-
Resistance to powdery mildew 8 gene
- RRBS:
-
Reduced representation bisulfite sequencing
- RT:
-
Reverse transcriptase enzyme
- SA:
-
Salicylic acid
- SEM:
-
Structural equation modeling
- sgRNA:
-
single guide RNA
- siRNA:
-
small interfering RNA
- SLAF:
-
Specific length amplified fragment
- SNAT:
-
Serotonin N-acetyltransferase
- SNP:
-
Single nucleotide polymorphism
- SNV:
-
Single nucleotide variant
- SSR:
-
Simple sequence repeat
- STS:
-
Stilbene synthase
- syn.:
-
synonymous
- TALEN:
-
Transcription activator like effector nuclease
- TAS4b :
-
Trans-acting small interfering RNA gene 4
- T-DNA:
-
Transfer DNA
- TIR:
-
Toll/interleukin-1 receptor
- TIR-NB-ARC-LRR:
-
Toll-interleukin-1-receptor - nucleotide-binding - adaptor shared by APAF-1 (apoptotic protease-activating factor-1), R proteins, and CED-4 (Caenorhabditis elegans death-4 protein) - leucine-rich repeat
- TLP:
-
Thaumatin-like protein
- TMT:
-
Tonoplastic monosaccharide transporter
- UBQ:
-
Ubiquitin
- UPOV:
-
International Union for the Protection of New Varieties of Plants
- USDA:
-
United States Department of Agriculture
- VCR:
-
Vivai Cooperativi Rauscedo
- VESPUCCI:
-
Vitis Expression Studies Platform Using COLOMBOS Compendia Instances
- VIVC:
-
Vitis International Variety Catalog
- VpGRP2A :
-
Vitis pseudoreticulata glycine-rich protein 2A gene
- VpRH2 :
-
Vitis pseudoreticulata RING-H2-type ubiquitin ligase gene
- VpTNL1 :
-
Vitis pseudoreticulata TIR-NB-LRR gene 1
- VOC:
-
Volatile organic compound
- WGS:
-
Whole genome shotgun
- WRKY:
-
WRKY family protein
- XVE:
-
LexA-VP16-ER
- ZFN:
-
Zinc finger nuclease
References
The authors apologize to the scientists that are not cited because of space limitation.
Aballay E, Mårtensson A, Persson P (2011) Screening of rhizosphere bacteria from grapevine for their suppressive effect on Xiphinema index Thorne & Allen on In vitro grape plants. Plant Soil 347:313–325. https://doi.org/10.1007/s11104-011-0851-6
Aballay E, Ordenes P, Mårtensson A, Persson P (2013) Effects of rhizobacteria on parasitism by Meloidogyne ethiopica on grapevines. Eur J Plant Pathol 135:137–145. https://doi.org/10.1007/s10658-012-0073-7
Aballay E, Prodan S, Zamorano A, Castaneda-Alvarez C (2017) Nematicidal effect of rhizobacteria on plant-parasitic nematodes associated with vineyards. World J Microbiol Biotechnol 33:131. https://doi.org/10.1007/s11274-017-2303-9
AbuQamar S, Moustafa K, Tran LSP (2017) Mechanisms and strategies of plant defense against Botrytis cinerea. Crit Rev Biotechnol 37:262–274. https://doi.org/10.1080/07388551.2016.1271767
Adam-Blondon AF, Alaux M, Pommier C, Cantu D, Cheng ZM et al (2016) Towards an open grapevine information system. Hortic Res 3:22. https://doi.org/10.1038/hortres.2016.56
Agarwal A, Cunningham JP, Valenzuela I, Blacket MJ (2020) A diagnostic LAMP assay for the destructive grapevine insect pest, phylloxera (Daktulosphaira vitifoliae). Sci Rep 10:1–10. https://doi.org/10.1038/s41598-020-77928-9
Agrios GN (2004) Plant pathology, 5th edn. Elsevier Academic Press, Burlington, MA
Agüero CB, Uratsu SL, Greve C, Powell ALT, Labavitch JM et al (2005) Evaluation of tolerance to Pierce’s disease and Botrytis in transgenic plants of Vitis vinifera L. expressing the pear PGIP gene. Mol Plant Pathol 6:43–51. https://doi.org/10.1111/J.1364-3703.2004.00262.X
Algarra Alarcon A, Lazazzara V, Cappellin L, Bianchedi PL, Schuhmacher R et al (2015) Emission of volatile sesquiterpenes and monoterpenes in grapevine genotypes following Plasmopara viticola inoculation in vitro. J Mass Spectrom 50:1013–1022. https://doi.org/10.1002/jms.3615
Ali K, Maltese F, Choi YH, Verpoorte R (2010) Metabolic constituents of grapevine and grape-derived products. Phytochem Rev 9:357–378. https://doi.org/10.1007/s11101-009-9158-0
Aliquó G, Torres R, Lacombe T, Boursiquot J-M, Laucou V et al (2017) Identity and parentage of some South American grapevine cultivars present in Argentina. Aust J Grape Wine Res 23:452–460. https://doi.org/10.1111/ajgw.12282
Alleweldt G, Possingham JV (1988) Progress in grapevine breeding. Theor Appl Genet 75:669–673. https://doi.org/10.1007/BF00265585
Alma A, Lessio F, Gonella E, Picciau L, Mandrioli M, Tota F (2018) New insights in phytoplasma-vector interaction: acquisition and inoculation of flavescence dorée phytoplasma by Scaphoideus titanus adults in a short window of time. Ann Appl Biology 173(1):55–62
Almada R, Cabrera N, Casaretto JA, Peña-Cortés H, Ruiz-Lara S et al (2011) Epigenetic repressor-like genes are differentially regulated during grapevine (Vitis vinifera L.) development. Plant Cell Rep 30:1959–1968. https://doi.org/10.1007/s00299-011-1104-0
Amala U, Chinniah C, Sawant IS, Yadav DS, Phad DM (2016) Comparative biology and fertility parameters of two spotted spider mite, Tetranychus urticae Koch. on different grapevine varieties. VITIS J Grapevine Res 55:31–36. https://doi.org/10.5073/vitis.2016.55.31-36
Anco DJ, Madden LV, Ellis MA (2012) Temporal patterns of sporulation potential of Phomopsis viticola on infected grape shoots, canes, and rachises. Plant Dis 96:1297–1302. https://doi.org/10.1094/PDIS-09-11-0806-RE
Anco DJ, Madden LV, Ellis MA (2013) Effects of temperature and wetness duration on the sporulation rate of Phomopsis viticola on infected grape canes. Plant Dis 97:579–589. https://doi.org/10.1094/PDIS-07-12-0666-RE
Anderson C, Choisne N, Adam-Blondon A-F, Dry IB (2011) Positional cloning of disease resistance genes in Grapevine. In: Adam-Blondon A-F, Martinez-Zapater J-M, Kole C (eds) Genetics, genomics, and breeding of grapes. Sciences Publishers, New York, NY, pp 186–210
Anderson K, Nelgen S (2020) Which winegrape varieties are grown where? A Global Empirical Picture, Revised. University of Adelaide Press, Adelaide, SA
Anfora G, Tasin M, de Cristofaro A, Ioriatti C, Lucchi A (2009) Synthetic grape volatiles attract mated Lobesia botrana females in laboratory and field bioassays. J Chem Ecol 35:1054–1062. https://doi.org/10.1007/s10886-009-9686-5
Antolín MC, Santesteban H, Ayari M, Aguirreolea J, Sánchez-Díaz M (2010) Grapevine fruiting cuttings: an experimental system to study grapevine physiology under water deficit conditions. In: Delrot S, Medrano H, Or E, Bavaresco L, Grando S (eds) Methodologies and results in grapevine research. Springer, Dordrecht, Netherlands, pp 151–163. https://doi.org/10.1007/978-90-481-9283-0_11
Antonić B, Jančíková S, Dordević D, Tremlová B (2020) Grape pomace valorization: a systematic review and meta-analysis. Foods 9:1627. https://doi.org/10.3390/foods9111627
Antonielli L, Compant S, Strauss J, Sessitsch A, Berger H (2014) Draft genome sequence of Phaeomoniella chlamydospora strain RR-HG1, a grapevine trunk disease (esca)-related member of the ascomycota. Genome Announc 2:10–12. https://doi.org/10.1128/genomeA.00098-14
Anzalone AV, Randolph PB, Davis JR, Sousa AA, Koblan LW et al (2019) Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576:149–157. https://doi.org/10.1038/s41586-019-1711-4
Aradhya MK, Dangl GS, Prins BH, Boursiquot J-M, Walker MA et al (2003) Genetic structure and differentiation in cultivated grape, Vitis vinifera L. Genet Res 81:179–192. https://doi.org/10.1017/S0016672303006177
Aradhya MK, Wang Y, Walker MA, Prins BH, Koehmstedt AM et al (2013) Genetic diversity, structure, and patterns of differentiation in the genus Vitis. Plant Syst Evol 299:317–330. https://doi.org/10.1007/s00606-012-0723-4
Arancibia C, Riaz S, Agüero C, Ramirez-Corona B, Alonso R et al (2018) Grape phylloxera (Daktulosphaira vitifoliae Fitch) in Argentina: ecological associations to diversity, population structure and reproductive mode. Aust J Grape Wine Res 24:284–291. https://doi.org/10.1111/ajgw.12337
Aranda X, Armengol J (2019) La reglamentación sobre el registro y uso de variedades de vid para vinificación. Enoviticolura Los Recur Genéticos Vitic Ante El Cambio Glob 59:92–100
Arias GA, Nieto JC (1980) Observaciones sobre la biología de la araña amarilla (Tetranychus urticae Koch) en los viñedos de “Tierra de Barros” (Badajoz) durante 1978 y 1979. Ministerio de Agricultura, Dirección general de la producción agraria. Serv Def Plagas Insp Fitopatol 4:1–39
Arias GA, Nieto JC (1981) Observaciones sobre la biología de la “araña amarilla” (Tetranychus urticae Koch) y correlación entre sintomas y pérdidas en una viña de “Tierra de Barros” (Badajoz) durante 1980. Serv Def Plagas Insp Fitopatol Comun 9:1–41
Armijo G, Schlechter R, Agurto M, Muñoz D, Nuñez C et al (2016) Grapevine pathogenic microorganisms: understanding infection strategies and host response scenarios. Front Plant Sci 7. https://doi.org/10.3389/fpls.2016.00382
Arroyo-García R, Ruiz-García L, Bolling L, Ocete R, López MA et al (2006) Multiple origins of cultivated grapevine (Vitis vinifera L. ssp. sativa) based on chloroplast DNA polymorphisms. Mol Ecol 15:3707–3714. https://doi.org/10.1111/j.1365-294X.2006.03049.x
Aruani C, Ruiz VS, Eibach R (2015) Les variétés de vigne. Origine, évolution et identification. Rev Des Oenologues Des Tech Vitivinic Oenologiques Mag Trimest D’information Prof 42:21–22
Asghari S, Harighi B, Ashengroph M, Clement C, Aziz A et al (2020) Induction of systemic resistance to Agrobacterium tumefaciens by endophytic bacteria in grapevine. Plant Pathol 69:827–837. https://doi.org/10.1111/ppa.13175
Asplen MK, Anfora G, Biondi A, Choi D-SS, Chu D et al (2015) Invasion biology of spotted wing Drosophila (Drosophila suzukii): a global perspective and future priorities. J Pest Sci 2004(88):469–494. https://doi.org/10.1007/s10340-015-0681-z
Ayres MR, Wicks TJ, Scott ES, Sosnowski MR (2017) Developing pruning wound protection strategies for managing Eutypa dieback. Aust J Grape Wine Res 23:103–111. https://doi.org/10.1111/ajgw.12254
Baccari C, Antonova E, Lindow S (2019) Biological control of Pierce’s disease of grape by an endophytic bacterium. Phytopathology 109:248–256. https://doi.org/10.1094/PHYTO-07-18-0245-FI
Bacilieri R, Lacombe T, Le Cunff L, Di Vecchi-Staraz M, Laucou V et al (2013) Genetic structure in cultivated grapevines is linked to geography and human selection. BMC Plant Biol 13:25. https://doi.org/10.1186/1471-2229-13-25
Badieinia F, Khajehali J, Nauen R, Dermauw W, Van Leeuwen T (2020) Metabolic mechanisms of resistance to spirodiclofen and spiromesifen in Iranian populations of Panonychus ulmi. Crop Prot 134:105166. https://doi.org/10.1016/j.cropro.2020.105166
Baginsky C, Contreras A, Covarrubias JI, Seguel O, Aballay E (2013) Control of plant-parasitic nematodes using cover crops in table grape cultivation in Chile. Int J Agric Nat Resour 40:547–557
Bagnoli B, Gargani E (2011) Survey on Scaphoideus titanus egg distribution on grapevine. Integr Prot Prod Vitic IOBC/Wprs Bull 67:233–237
Bajda S, Dermauw W, Greenhalgh R, Nauen R, Tirry L et al (2015) Transcriptome profiling of a spirodiclofen susceptible and resistant strain of the European red mite Panonychus ulmi using strand-specific RNA-seq. BMC Genomics 16:974. https://doi.org/10.1186/s12864-015-2157-1
Balding DJ (2006) A tutorial on statistical methods for population association studies. Nat Rev Genet 7:781–791. https://doi.org/10.1038/nrg1916
Balint‐Kurti P (2019) The plant hypersensitive response: concepts, control and consequences. Mol Plant Pathol 20:mpp.12821. https://doi.org/10.1111/mpp.12821
Bao LV, Scatoni IB, Gaggero C, Gutiérrez L, Monza J et al (2015) Genetic diversity of grape phylloxera leaf-galling populations on Vitis species in Uruguay. Am J Enol Vitic 66:46–53. https://doi.org/10.5344/ajev.2014.14026
Baránek M, Křižan B, Ondrušíková E, Pidra M (2010) DNA-methylation changes in grapevine somaclones following in vitro culture and thermotherapy. Plant Cell Tissue Organ Cult 101:11–22. https://doi.org/10.1007/s11240-009-9656-1
Baránek M, Čechová J, Raddová J, Holleinová V, Ondrušíková E et al (2015) Dynamics and reversibility of the DNA methylation landscape of grapevine plants (Vitis vinifera) stressed by in vitro cultivation and thermotherapy. PLoS ONE 10:1–16. https://doi.org/10.1371/journal.pone.0126638
Barba P, Cadle-Davidson L, Harriman J, Glaubitz J, Brooks S et al (2014) Grapevine powdery mildew resistance and susceptibility loci identified on a high-resolution SNP map. Theor Appl Genet 127:73–84. https://doi.org/10.1007/s00122-013-2202-x
Barba P, Lillis J, Luce RS, Travadon R, Osier M et al (2018) Two dominant loci determine resistance to Phomopsis cane lesions in F1 families of hybrid grapevines. Theor Appl Genet 131:1173–1189. https://doi.org/10.1007/s00122-018-3070-1
Barba P, Loughner R, Wentworth K, Nyrop JP, Loeb GM et al (2019) A QTL associated with leaf trichome traits has a major influence on the abundance of the predatory mite Typhlodromus pyri in a hybrid grapevine population. Hortic Res 6. https://doi.org/10.1038/s41438-019-0169-8
Barker CL, Donald T, Pauquet J, Ratnaparkhe MB, Bouquet A et al (2005) Genetic and physical mapping of the grapevine powdery mildew resistance gene, Run1, using a bacterial artificial chromosome library. Theor Appl Genet 111:370–377. https://doi.org/10.1007/s00122-005-2030-8
Barnett DE (1976) A revision of the Nearctic species of the genus Scaphoideus (Homoptera: Cicadellidae). Trans Am Entomol Soc 102:485–593
Barrett HC (1953) Survey of black rot resistance of the foliage of wild grape species. In: Proceedings of the American society for horticultural science, College Park, MD
Baser N, Broutou O, Verrastro V, Porcelli F, Ioriatti C et al (2018) Susceptibility of table grape varieties grown in South-Eastern Italy to Drosophila suzukii. J Appl Entomol 142:465–472. https://doi.org/10.1111/jen.12490
Batovska DI, Todorova IT, Parushev SP, Nedelcheva DV, Bankova VS et al (2009) Biomarkers for the prediction of the resistance and susceptibility of grapevine leaves to downy mildew. J Plant Physiol 166:781–785. https://doi.org/10.1016/j.jplph.2008.08.008
Baumgartner K, Fujiyoshi PT, Travadon R, Castlebury LA, Wilcox WF et al (2013) Characterization of species of Diaporthe from wood cankers of grape in Eastern North American vineyards. Plant Dis 97:912–920. https://doi.org/10.1094/PDIS-04-12-0357-RE
Bavaresco L (2019) Impact of grapevine breeding for disease resistance on the global wine industry. Acta Hortic 1248:7–14. https://doi.org/10.17660/ActaHortic.2019.1248.2
Bazzi C, Alexandrova M, Stefani E, Anaclerio F, Burr TJ (1999) Biological control of Agrobacterium vitis using non-tumorigenic Agrobacteria. VITIS J Grapevine Res 38:31–35
Belli G, Bianco PA, Conti M (2010) Grapevine yellows in Italy: past, present and future. J Plant Pathol 92:303–326. https://doi.org/10.4454/jpp.v92i2.172
Bellin D, Peressotti E, Merdinoglu D, Wiedemann-Merdinoglu S, Adam-Blondon A-F et al (2009) Resistance to Plasmopara viticola in grapevine 'Bianca' is controlled by a major dominant gene causing localised necrosis at the infection site. Theor Appl Genet 120:163–176. https://doi.org/10.1007/s00122-009-1167-2
Bello A, Arias M, López-Pérez JA, García-Álvarez A, Fresno J et al (2004) Biofumigation, fallow and nematode management in vineyard replant. Nematropica 34:53–64
Benelli L, Thomson I (2019) Sex pheromone aerosol devices for mating disruption: challenges for a brighter future. Insects 10:308. https://doi.org/10.3390/insects10100308
Benheim D, Rochfort S, Ezernieks V, Korosi GA, Powell KS et al (2011) Early detection of grape phylloxera (Daktulosphaira vitifoliae Fitch) infestation through identification of chemical biomarkers. Acta Hortic 904:17–24. https://doi.org/10.17660/ActaHortic.2011.904.2
Berbegal M, Ramón-Albalat A, León M, Armengol J (2020) Evaluation of long-term protection from nursery to vineyard provided by Trichoderma atroviride SC1 against fungal grapevine trunk pathogens. Pest Manag Sci 76:967–977. https://doi.org/10.1002/ps.5605
Bergamini C, Cardone MF, Anaclerio A, Perniola R, Pichierri A et al (2013) Validation assay of p3-VvAGL11 marker in a wide range of genetic background for early selection of stenospermocarpy in Vitis vinifera L. Mol Biotechnol 54:1021–1030. https://doi.org/10.1007/s12033-013-9654-8
Bernard MB, Horne PA, Hoffmann AA (2005) Eriophyoid mite damage in Vitis vinifera (grapevine) in Australia: Calepitrimerus vitis and Colomerus vitis (Acari: Eriophyidae) as the common cause of the widespread ‘Restricted Spring Growth’ syndrome. Exp Appl Acarol 35:83–109. https://doi.org/10.1007/s10493-004-1986-4
Bernardo R (2014) Essentials of plant breeding. Stemma Press, Woodbury, Minnesota
Bertazzon N, Borgo M, Angelini E (2010) The complete genome sequence of the BD variant of Grapevine Leafroll-associated Virus 2. Arch Virol 155:1717–1719. https://doi.org/10.1007/s00705-010-0769-y
Bertazzon N, Bagnaresi P, Forte V, Mazzucotelli E, Filippin L et al (2019) Grapevine comparative early transcriptomic profiling suggests that Flavescence dorée phytoplasma represses plant responses induced by vector feeding in susceptible varieties. BMC Genomics 20:1–27. https://doi.org/10.1186/s12864-019-5908-6
Bertin S, Cavalieri V, Gribaudo I, Sacco D, Marzachì C et al (2016) Transmission of Grapevine Virus A and Grapevine Leafroll-associated Virus 1 and 3 by Heliococcus bohemicus (Hemiptera: Pseudococcidae) Nymphs from Plants with Mixed Infections. J Econ Entomol 109:1504–1511. https://doi.org/10.1093/jee/tow120
Bertini E, Tornielli GB, Pezzotti M, Zenoni S (2019) Regeneration of plants from embryogenic callus-derived protoplasts of Garganega and Sangiovese grapevine (Vitis vinifera L.) cultivars. Plant Cell Tissue Organ Cult 138:239–246. https://doi.org/10.1007/s11240-019-01619-1
Bettiga LJ, Gubler WD (2015) “Bunch rots”. In: Bettiga LJ (ed) Grape pest management. University of California, Agriculture and Natural Resources, Davis, CA pp 3343:93–103
Bewick AJ, Schmitz RJ (2017) Gene body DNA methylation in plants. Curr Opin Plant Biol 36:103–110. https://doi.org/10.1016/j.pbi.2016.12.007
Bhattarai G, Fennell A, Londo JP, Coleman C, Kovacs LG (2021) A novel grape downy mildew resistance locus from Vitis rupestris. Am J Enol Vitic 72:12–20. https://doi.org/10.5344/ajev.2020.20030
Bianchi D, Brancadoro L, De Lorenzis G (2020) Genetic diversity and population structure in a Vitis spp. Core collection investigated by SNP Markers. Diversity 12:103. https://doi.org/10.3390/d12030103
Bierman A, LaPlumm T, Cadle-Davidson L, Gadoury D, Martinez D et al (2019) A high-throughput phenotyping system using machine vision to quantify severity of grapevine powdery mildew. Plant Phenomics 2019:1–13. https://doi.org/10.34133/2019/9209727
Bigeard J, Colcombet J, Hirt H (2015) Signaling mechanisms in pattern-triggered immunity (PTI). Mol Plant 8:521–539. https://doi.org/10.1016/j.molp.2014.12.022
Billet K, Malinowska MA, Munsch T, Unlubayir M, Adler S et al (2020) Semi-targeted metabolomics to validate biomarkers of grape downy mildew infection under field conditions. Plants 9:1008. https://doi.org/10.3390/plants9081008
Billones-Baaijens R, Jones EE, Ridgway HJ, Jaspers MV (2014) Susceptiblity of common rootstock and scion varieties of grapevines to Botryosphaeriaceae species. Australas Plant Pathol 43:25–31. https://doi.org/10.1007/s13313-013-0228-9
Bini F, Kuczmog A, Putnoky P, Otten L, Bazzi C et al (2008) Novel pathogen-specific primers for the detection of Agrobacterium vitis and Agrobacterium tumefaciens. VITIS J Grapevine Res 47:181–189. https://doi.org/10.5073/vitis.2008.47.181-189
Bird G, Diamond C, Warner F, Davenport J (1994) Distribution and regulation of meloidogyne nataliei. J Nematol 26:727–730
Bitsadze N, Aznarashvili M, Vercesi A, Chipashvili R, Failla O et al (2015) Screening of Georgian grapevine germplasm for susceptibility to downy mildew (Plasmopara viticola). VITIS J Grapevine Res 54:193–196. https://doi.org/10.5073/vitis.2015.54.special-issue
Blanc S, Wiedemann-Merdinoglu S, Dumas V, Mestre P, Merdinoglu D (2012) A reference genetic map of Muscadinia rotundifolia and identification of Ren5, a new major locus for resistance to grapevine powdery mildew. Theor Appl Genet 125:1663–1675. https://doi.org/10.1007/s00122-012-1942-3
Blanc-Mathieu R, Perfus-Barbeoch L, Aury J-M, Da Rocha M, Gouzy J et al (2017) Hybridization and polyploidy enable genomic plasticity without sex in the most devastating plant-parasitic nematodes. PLOS Genet 13:e1006777. https://doi.org/10.1371/journal.pgen.1006777
Blanco-Ulate B, Allen G, Powell ALT, Cantu D (2013a) Draft genome sequence of Botrytis cinerea BcDW1, inoculum for noble rot of grape berries. Genome Announc 1. https://doi.org/10.1128/genomeA.00252-13
Blanco-Ulate B, Vincenti E, Powell ALT, Cantu D (2013b) Tomato transcriptome and mutant analyses suggest a role for plant stress hormones in the interaction between fruit and Botrytis cinerea. Front Plant Sci 4:1–16. https://doi.org/10.3389/fpls.2013.00142
Blanco-Ulate B, Rolshausen P, Cantu D (2013c) Draft genome sequence of the ascomycete Phaeoacremonium aleophilum strain UCR-PA7, a causal agent of the esca disease complex in Grapevines. Genome Announc 1:e00390–e413. https://doi.org/10.1128/genomeA.00390-13
Blank L, Wolf T, Eimert K, Schröder M-B (2009) Differential gene expression during hypersensitive response in phylloxera -resistant rootstock ‘Börner’ using custom oligonucleotide arrays. J Plant Interact 4:261–269. https://doi.org/10.1080/17429140903254697
Blasi P, Blanc S, Wiedemann-Merdinoglu S, Prado E, Rühl EH et al (2011) Construction of a reference linkage map of Vitis amurensis and genetic mapping of Rpv8, a locus conferring resistance to grapevine downy mildew. Theor Appl Genet 123:43–53. https://doi.org/10.1007/s00122-011-1565-0
Body MJA, Appel HM, Edger PP, Schultz JC (2019) A gall-forming insect manipulates hostplant phytohormone synthesis, concentrations, and signaling. bioRxiv 658823. https://doi.org/10.1101/658823
Bois B, Zito S, Calonnec A, Ollat, N. (2017). Climate vs grapevine pests and diseases worldwide: the first results of a global survey. J Int des Sci la Vigne du Vin 51:133–139. https://doi.org/10.20870/oeno-one.2016.0.0.1780
Boller E, Janser E, Zahner S, Potter C (1985) Kann Herbizideinsatz im Weinbau Spinnmilbenprobleme verursachen? Schweiz Zeit Obs Weinb 121:527–531
Boller T, Felix G (2009) A renaissance of elicitors: perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Annu Rev Plant Biol 60:379–406. https://doi.org/10.1146/annurev.arplant.57.032905.105346
Borgo M, Pegoraro G, Sartori E (2016) Susceptibility of grape varieties to esca disease. BIO Web Conf 7:01041. https://doi.org/10.1051/bioconf/20160701041
Borie B, Jacquiot L, Jamaux-Despréaux I, Larignon P, Péros JP (2002) Genetic diversity in populations of the fungi Phaeomoniella chlamydospora and Phaeoacremonium aleophilum on grapevine in France. Plant Pathol 51:85–96. https://doi.org/10.1046/j.0032-0862.2001.658.x
Boso S, Santiago JL, Martínez MC (2005) A method to evaluate downy mildew resistance in Grapevine. Agron Sustain Dev 25:163–165. https://doi.org/10.1051/agro:2004062
Boso S, Alonso-Villaverde V, Gago P, Santiago JL, Martínez MC (2011) Susceptibility of 44 grapevine (Vitis vinifera L.) varieties to downy mildew in the field. Aust J Grape Wine Res 17:394–400. https://doi.org/10.1111/j.1755-0238.2011.00157.x
Boso S, Alonso-Villaverde V, Gago P, Santiago JL, Martínez MC (2014) Susceptibility to downy mildew (Plasmopara viticola) of different Vitis varieties. Crop Prot 63:26–35. https://doi.org/10.1016/j.cropro.2014.04.018
Bouquet A (1986) Introduction dans l’espèce Vitis vinifera L. d’un caractère de résistance à l’oidium (Uncinula necator Schw. Burr.) issu de l’espèce Muscadinia rotundifolia (Michx.) Small. Vignevini 12:141–146
Bouquet A (2011) Grapevines and viticulture. In: Adam-Blondon AM, Martinez-Zapater J, Kole C (eds) Genetics, genomics, and breeding of grapes. Science Publishers, pp 1–29. https://doi.org/10.1201/b10948-2
Boursiquot JM, Lacombe T, Laucou V, Julliard S, Perrin FX et al (2009) Parentage of Merlot and related winegrape cultivars of southwestern France: discovery of the missing link. Aust J Grape Wine Res 15:144–155. https://doi.org/10.1111/j.1755-0238.2008.00041.x
Bova S, Rosenthal Y, Liu Z, Godad SP, Yan M (2021) Seasonal origin of the thermal maxima at the Holocene and the last interglacial. Nature 589:548–553. https://doi.org/10.1038/s41586-020-03155-x
Bove F, Bavaresco L, Caffi T, Rossi V (2019) Assessment of resistance components for improved phenotyping of grapevine varieties resistant to downy mildew. Front Plant Sci 10. https://doi.org/10.3389/fpls.2019.01559
Bovey RM, Baggiolini A, Bolay E, Bovay R, Corbaz G et al (1979) La défense des plantes cultivées. Ed. Payot, Lausanne, Suisse
Bowers JE, Dangl GS, Vignani R, Meredith CP (1996) Isolation and characterization of new polymorphic simple sequence repeat loci in grape (Vitis vinifera L.). Genome 39:628–633. https://doi.org/10.1139/g96-080
Bowers JE, Meredith CP (1997) The parentage of a classic wine grape, Cabernet Sauvignon. Nat Genet 16:84–87. https://doi.org/10.1038/ng0597-84
Bowers JE, Boursiquot JM, This P, Chu K, Johansson H et al (1999) Historical genetics: the parentage of chardonnay, gamay, and other wine grapes of Northeastern France. Science (80-) 285:1562–1565. https://doi.org/10.1126/science.285.5433.1562
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y et al (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635. https://doi.org/10.1093/bioinformatics/btm308
Brilli M, Asquini E, Moser M, Bianchedi PL, Perazzolli M et al (2018) A multi-omics study of the grapevine-downy mildew (Plasmopara viticola) pathosystem unveils a complex protein coding- and noncoding-based arms race during infection. Sci Rep 8. https://doi.org/10.1038/s41598-018-19158-8
Brown AHD (1995) The core collection at the crossroads. In: Hodgkin, T, Brown AHD, van Hintum TJL, Morales EAV (eds) Core collections of plant genetic resources. John Wiley and Sons, Chichester, UK, pp 3–19
Brown DJF, Dalmasso A, Trudgill DL (1993) Nematode pests of soft fruits and vines. In: Evans K, Trudgill DL, Webster JM (eds) Plant parasitic nematodes in temperate agriculture. CAB International, Wallingford UK, pp 427–462
Brown MV., Moore JN, Fenn P, McNew RW (1999) Comparison of leaf disk, greenhouse, and field screening procedures for evaluation of grape seedlings for downy mildew resistance. HortScience 34:331–333. https://doi.org/10.21273/hortsci.34.2.331
Brown AA, Lawrence DP, Baumgartner K (2020) Role of basidiomycete fungi in the grapevine trunk disease esca. Plant Pathol 69:205–220. https://doi.org/10.1111/ppa.13116
Brown AA, Travadon R, Lawrence DP, Torres G, Zhuang G et al (2021) Pruning-wound protectants for trunk-disease management in California table grapes. Crop Prot 141:105490. https://doi.org/10.1016/j.cropro.2020.105490
Bruez E, Lecomte P, Grosman J, Doublet B, Bertsch C et al (2013) Overview of grapevine trunk diseases in France in the 2000s. Phytopathol Mediterr 52:262–275. https://doi.org/10.14601/Phytopathol_Mediterr-11578
Brulé D, Villano C, Davies LJ, Trdá L, Claverie J et al (2019) The grapevine (Vitis vinifera) LysM receptor kinases VvLYK1-1 and VvLYK1-2 mediate chitooligosaccharide-triggered immunity. Plant Biotechnol J 17:812–825. https://doi.org/10.1111/pbi.13017
Buonassisi D, Colombo M, Migliaro D, Dolzani C, Peressotti E et al (2017) Breeding for grapevine downy mildew resistance: a review of “omics” approaches. Euphytica 213:103. https://doi.org/10.1007/s10681-017-1882-8
Buonassisi D, Cappellin L, Dolzani C, Velasco R, Peressotti E et al (2018) Development of a novel phenotyping method to assess downy mildew symptoms on grapevine inflorescences. Sci Hortic 236:79–89. https://doi.org/10.1016/j.scienta.2018.03.023
Burrack HJ, Fernandez GE, Spivey T, Kraus DA (2013) Variation in selection and utilization of host crops in the field and laboratory by Drosophila suzukii Matsumara (Diptera: Drosophilidae), an invasive frugivore. Pest Manag Sci 69:1173–1180. https://doi.org/10.1002/ps.3489
Cabezas JA, Ibáñez J, Lijavetzky D, Vélez MD, Bravo G et al (2011) A 48 SNP set for grapevine cultivar identification. BMC Plant Biol 11:153. https://doi.org/10.1186/1471-2229-11-153
Cadle-Davidson L (2008) Variation within and between Vitis spp. for foliar resistance to the downy mildew pathogen Plasmopara viticola. Plant Dis 92:1577–1584. https://doi.org/10.1094/PDIS-92-11-1577
Cadle-Davidson L, Mahani S, Gadoury DM, Kozma P, Reisch BI (2011) Natural infection of Run1 -positive vines by naive genotypes of Erysiphe necator. VITIS J Grapevine Res 50:173–175
Cadle-Davidson L, Londo J, Martinez D, Sapkota S, Gutierrez B (2019) From phenotyping to phenomics: present and future approaches in grape trait analysis to inform grape gene function. In: Cantu D, Walker MA (eds) The grape genome. Springer, Cham, Switzerland, pp 199–222. https://doi.org/10.1007/978-3-030-18601-2_10
Caffarra A, Rinaldi M, Eccel E, Rossi V, Pertot I (2012) Modelling the impact of climate change on the interaction between grapevine and its pests and pathogens: European grapevine moth and powdery mildew. Agric Ecosyst Environ 148:89–101. https://doi.org/10.1016/j.agee.2011.11.017
Caffi T, Rossi V, Legler SE, Bugiani R (2011) A mechanistic model simulating ascosporic infections by Erysiphe necator, the powdery mildew fungus of grapevine. Plant Pathol 60:522–531. https://doi.org/10.1111/j.1365-3059.2010.02395.x
Caffi T, Legler SE, Rossi V, Bugiani R (2012) Evaluation of a warning system for early-season control of grapevine powdery mildew. Plant Dis 96:104–110. https://doi.org/10.1094/PDIS-06-11-0484
Caffi T, Legler SE, Bugiani R, Rossi V (2013) Combining sanitation and disease modelling for control of grapevine powdery mildew. Eur J Plant Pathol 135:817–829. https://doi.org/10.1007/s10658-012-0124-0
Caffi T, Legler S, González-Domınguez E, Rossi V (2017) Sustainable management of vineyards: the experience of a large-scale application of a web-based decision support system. In: Proceedings of 8th international workshop grapevine downy and powdery mildew. Oregon State University Campus, Corvallis, OR
Caliari V, Burin VM, Rosier JP, Bordignon Luiz MT (2014) Aromatic profile of Brazilian sparkling wines produced with classical and innovative grape varieties. Food Res Int 62:965–973. https://doi.org/10.1016/j.foodres.2014.05.013
Calonnec A, Cartolaro P, Naulin JM, Bailey D, Langlais M (2008) A host-pathogen simulation model: Powdery mildew of grapevine. Plant Pathol 57:493–508. https://doi.org/10.1111/j.1365-3059.2007.01783.x
Camporese P, Duso C (1996) Different colonization patterns of phytophagous and predatory mites (Acari: Tetranychidae, Phytoseiidae) on three grape varieties: a case study. Exp Appl Acarol 20:1–22
Canaguier A, Grimplet J, Di Gaspero G, Scalabrin S, Duchêne E et al (2017) A new version of the grapevine reference genome assembly (12X.v2) and of its annotation (VCost.v3). Genomics Data 14:56–62. https://doi.org/10.1016/j.gdata.2017.09.002
Capriotti L, Baraldi E, Mezzetti B, Limera C, Sabbadini S (2020) Biotechnological approaches: gene overexpression, gene silencing, and genome editing to control fungal and oomycete diseases in Grapevine. Int J Mol Sci 21:5701. https://doi.org/10.3390/ijms21165701
Carisse O, Bacon R, Lefebvre A (2009) Grape powdery mildew (Erysiphe necator) risk assessment based on airborne conidium concentration. Crop Prot 28:1036–1044. https://doi.org/10.1016/j.cropro.2009.06.002
Carisse O, Lefebvre A (2011a) A model to estimate the amount of primary inoculum of Elsinoë ampelina. Plant Dis 95:1167–1171. https://doi.org/10.1094/PDIS-11-10-0798
Carisse O, Lefebvre A (2011b) Evaluation of Northern grape hybrid cultivars for their susceptibility to anthracnose caused by Elsinoë ampelina. Plant Heal Prog 12:9. https://doi.org/10.1094/php-2011-0805-01-rs
Carisse O, Morissette-Thomas V (2013) Epidemiology of grape anthracnose: factors associated with defoliation of grape leaves infected by Elsinoë ampelina. Plant Dis 97:222–230. https://doi.org/10.1094/PDIS-04-12-0393-RE
Carle P, Malembic-Maher S, Arricau-Bouvery N, Desqué D, Eveillard S et al (2011) “Flavescence dorée” phytoplasma genome: A metabolism oriented towards glycolysis and protein degradation. Bull Insectology 64:13–14
Carneiro RMDG, Randig O, Almeida MRA, Gomes ACMM (2004) Additional information on Meloidogyne ethiopica Whitehead, 1968 (Tylenchida: Meloidogynidae), a root-knot nematode parasitising kiwi fruit and grapevine from Brazil and Chile. Nematology 6:109–123. https://doi.org/10.1163/156854104323072982
Castillo P, Rapoport HF, Rius JEP, Díaz RMJ (2008) Suitability of weed species prevailing in Spanish vineyards as hosts for root-knot nematodes. Eur J Plant Pathol 120:43–51. https://doi.org/10.1007/s10658-007-9195-8
Castillo P, Gutiérrez-Gutiérrez C, Palomares-Rius JE, Cantalapiedra Navarrete C, Landa BB (2009) First report of root-knot nematode Meloidogyne hispanica infecting grapevines in Southern Spain. Plant Dis 93:1353–1353. https://doi.org/10.1094/PDIS-93-12-1353B
Cavaco AR, Maia M, Laureano G, Duarte B, Caçador I et al (2018) P-285-Identifying grapevine lipid biomarkers of resistance/susceptibility to Plasmopara viticola; towards a sustainable viticulture. Free Radic Biol Med 120:S131. https://doi.org/10.1016/j.freeradbiomed.2018.04.432
Celii M (2016) Analysis of epigenomic variability in grapevine and its relation with structural variation. Università degli studi di Udine
Chacón-Vozmediano JL, Gramaje D, León M, Armengol J, Moral J et al (2021) Cultivar susceptibility to natural infections caused by fungal grapevine trunk pathogens in La Mancha Designation of Origin (Spain). Plants 10:1171. https://doi.org/10.3390/plants10061171
Chang E-H, Jung S-M, Park S-J, Noh JH, Hur Y-Y et al (2014) Wine quality of grapevine “Cheongsoo” and the related metabolites on proton nuclear magnetic resonance (NMR) spectroscoscopy at the different harvest times. Plant Omics 7:80–86
Chávez Montes RA, De Fátima Rosas-Cárdenas F, De Paoli E, Accerbi M, Rymarquis LA et al (2014) Sample sequencing of vascular plants demonstrates widespread conservation and divergence of microRNAs. Nat Commun 5:3722. https://doi.org/10.1038/ncomms4722
Cheng S, Xie X, Xu Y, Zhang C, Wang X et al (2016) Genetic transformation of a fruit-specific, highly expressed stilbene synthase gene from Chinese wild Vitis quinquangularis. Planta 243:1041–1053. https://doi.org/10.1007/s00425-015-2459-1
Chitarrini G, Soini E, Riccadonna S, Franceschi P, Zulini L et al (2017) Identification of biomarkers for defense response to Plasmopara viticola in a resistant grape variety. Front Plant Sci 8. https://doi.org/10.3389/fpls.2017.01524
Chitarrini G, Riccadonna S, Zulini L, Vecchione A, Stefanini M et al (2020) Two-omics data revealed commonalities and differences between Rpv12- and Rpv3-mediated resistance in grapevine. Sci Rep 10:1–15. https://doi.org/10.1038/s41598-020-69051-6
Chiu JC, Jiang X, Zhao L, Hamm CA, Cridland JM et al (2013) Genome of Drosophila suzukii, the spotted wing drosophila. G3 Genes|Genomes|Genetics 3:2257–2271. https://doi.org/10.1534/g3.113.008185
Chuche J, Thiéry D (2014) Biology and ecology of the Flavescence dorée vector Scaphoideus titanus: A review. Agron Sustain Dev 34:381–403. https://doi.org/10.1007/s13593-014-0208-7
Ciliberti N, Fermaud M, Roudet J, Rossi V (2015) Environmental conditions affect Botrytis cinerea infection of mature grape berries more than the strain or transposon genotype. Phytopathology 105:1090–1096. https://doi.org/10.1094/PHYTO-10-14-0264-R
Cini A, Ioriatti C, Anfora G (2012) A review of the invasion of Drosophila suzukii in Europe and a draft research agenda for integrated pest management. Bull Insectology 65:149–160
Cini A, Anfora G, Escudero-Colomar LA, Grassi A, Santosuosso U et al (2014) Tracking the invasion of the alien fruit pest Drosophila suzukii in Europe. J Pest Sci 87:559–566. https://doi.org/10.1007/s10340-014-0617-z
Cipriani G, Marrazzo MT, Di Gaspero G, Pfeiffer A, Morgante M et al (2008) A set of microsatellite markers with long core repeat optimized for grape (Vitis spp.) genotyping. BMC Plant Biol 8:127. https://doi.org/10.1186/1471-2229-8-127
Cipriani G, Spadotto A, Jurman I, Di Gaspero G, Crespan M et al (2010) The SSR-based molecular profile of 1005 grapevine (Vitis vinifera L.) accessions uncovers new synonymy and parentages, and reveals a large admixture amongst varieties of different geographic origin. Theor Appl Genet 121:1569–1585. https://doi.org/10.1007/s00122-010-1411-9
Clark MD, Teh SL, Burkness E, Moreira L, Watson G et al (2018) Quantitative Trait Loci identified for foliar phylloxera resistance in a hybrid grape population. Aust J Grape Wine Res 24:292–300. https://doi.org/10.1111/ajgw.12341
Clarke CW, Norng S, Yuanpeng D, Carmody BM, Powell KS (2018) Efficacy of steam and hot water disinfestation treatments against genetically diverse strains of grape phylloxera Daktulosphaira vitifoliae Fitch (Hemiptera: Phylloxeridae) on viticulture equipment and machinery. Aust J Grape Wine Res 24:275–281. https://doi.org/10.1111/ajgw.12329
Clarke CW, Norng S, Carmody BM, Yuanpeng D, Powell KS (2019) Hot water immersion as a disinfestation treatment for grapevine root cuttings against genetically diverse grape phylloxera Daktulosphaira vitifoliae Fitch. Aust J Grape Wine Res 25:396–403. https://doi.org/10.1111/ajgw.12407
Claverie M, Notaro M, Fontaine F, Wéry J (2020) Current knowledge on Grapevine Trunk Diseases with complex etiology: a systemic approach. Phytopathol Mediterr 59:29–53. https://doi.org/10.14601/Phyto-11150
Clayton CN, Ridings W (1970) Grape rust, Physopella ampelopsidis, on Vitis rotundifolia in North Carolina. Phytopathology 60:1022–1023
Cloete M, Fischer M, Mostert L, Halleen F (2015) Hymenochaetales associated with esca-related wood rots on grapevine with a special emphasis on the status of esca in South African vineyards. Phytopathol Mediterr 54:299–312. https://doi.org/10.14601/Phytopathol_Mediterr-16364
Cobos R, Mateos RM, Álvarez-Pérez JM, Olego MA, Sevillano S et al (2015) Effectiveness of natural antifungal compounds in controlling infection by grapevine trunk disease pathogens through pruning wounds. Appl Environ Microbiol 81:6474–6483. https://doi.org/10.1128/AEM.01818-15
Cocco A, Pacheco da Silva VC, Benelli G, Botton M, Lucchi A et al (2021) Sustainable management of the vine mealybug in organic vineyards. J Pest Sci 2004(94):153–185. https://doi.org/10.1007/s10340-020-01305-8
Cola G, Mariani L, Salinari F, Civardi S, Bernizzoni F et al (2014) Description and testing of a weather-based model for predicting phenology, canopy development and source-sink balance in Vitis vinifera L. cv. Barbera Agric Meteorol 184:117–136. https://doi.org/10.1016/j.agrformet.2013.09.008
Coleman C, Copetti D, Cipriani G, Hoffmann S, Kozma P et al (2009) The powdery mildew resistance gene Ren1 co-segregates with an NBS-LRR gene cluster in two Central Asian grapevines. BMC Genet 10:1–20. https://doi.org/10.1186/1471-2156-10-89
Comont G, Corio-Costet MF, Larignon P, Delmotte F (2010) AFLP markers reveal two genetic groups in the French population of the grapevine fungal pathogen Phaeomoniella chlamydospora. Eur J Plant Pathol 127:451–464. https://doi.org/10.1007/s10658-010-9611-3
Cooper ML, Hobbs MB, Strode B, Varela LG (2020) Grape erineum mite: postharvest sulfur use reduces subsequent leaf blistering. Calif Agric 74:94–100. https://doi.org/10.3733/ca.2020a0012
Correa J, Mamani M, Muñoz-Espinoza C, Laborie D, Muñoz C et al (2014) Heritability and identification of QTLs and underlying candidate genes associated with the architecture of the grapevine cluster (Vitis vinifera L.). Theor Appl Genet 127:1143–1162. https://doi.org/10.1007/s00122-014-2286-y
Corrie AM, Crozier RH, Van Heeswijck R, Hoffmann AA (2002) Clonal reproduction and population genetic structure of grape phylloxera, Daktulosphaira vitifoliae, in Australia. Heredity (edinb) 88:203–211. https://doi.org/10.1038/sj.hdy.6800028
Cottral E, Ridgway HJ, Pascoe I, Edwards J, Taylor P (2001) UP-PCR analysis of Australian isolates of Phaeomoniella chlamydospora and Phaeoacremonium aleophilum. Phytopathol Mediterr 40. https://doi.org/10.1400/14670
COUNCIL REGULATION (EC) No 2100/94 of 27 July 1994 on Community plant variety rights. (OJ L 227, 1.9.1994, p 1)
Crava CM, Zanini D, Amati S, Sollai G, Crnjar R et al (2020) Structural and transcriptional evidence of mechanotransduction in the Drosophila suzukii ovipositor. J Insect Physiol 125:104088. https://doi.org/10.1016/j.jinsphys.2020.104088
Crespan M, Cabello F, Giannietto S, Ibanez J, Kontic JK et al (2006) Malvasia delle Lipari, Malvasia di Sardegna, Greco di Gerace, Malvasia de Sitges and Malvasia dubrovačka–synonyms of an old and famous grape cultivar. VITIS J Grapevine Res 45:69–73
Crespan M, Meneghetti S, Cancellier S (2009) Identification and genetic relationship of the principal rootstocks cultivated in Italy. Am J Enol Vitic 60:349–356
Crespan M, Carraro R, Giust M, Migliaro D (2016) Origin of Termarina cultivar, another grapevine (Vitis vinifera L.) parthenocarpic somatic variant. Aust J Grape Wine Res 22:489–493. https://doi.org/10.1111/ajgw.12236
Crespan M, Migliaro D, Larger S, Pindo M, Petrussi C et al (2020) Unraveling the genetic origin of ‘Glera’, ‘Ribolla Gialla’ and other autochthonous grapevine varieties from Friuli Venezia Giulia (Northeastern Italy). Sci Rep. 10. https://doi.org/10.1038/s41598-020-64061-w
Cunha J, Zinelabidine LH, Teixeira-Santos M, Brazao J, Fevereiro P et al (2015) Grapevine cultivar “Alfrocheiro” or “Brunal” plays a primary role in the relationship among Iberian grapevines. VITIS J Grapevine Res 54:59–65
Cunha J, Ibáñez J, Teixeira-Santos M, Brazão J, Fevereiro P et al (2020) Genetic relationships among portuguese cultivated and Wild Vitis vinifera L. Germplasm Front Plant Sci 11:27. https://doi.org/10.3389/fpls.2020.00127
D’Onofrio C (2020) Introgression among cultivated and wild grapevine in Tuscany. Front Plant Sci 11. https://doi.org/10.3389/fpls.2020.00202
D’Onofrio C, Tumino G, Gardiman M, Crespan M, Bignami C et al (2021) Parentage atlas of Italian grapevine varieties as inferred from SNP genotyping. Front Plant Sci 11:605934. https://doi.org/10.3389/fpls.2020.605934
Da Silva C, Zamperin G, Ferrarini A, Minio A, Dal Molin A et al (2013) The high polyphenol content of Grapevine cultivar tannat berries is conferred primarily by Genes that are not shared with the reference genome. Plant Cell 25:4777–4788. https://doi.org/10.1105/tpc.113.118810
De la Fuente Lloreda M (2018) Use of hybrids in viticulture. A challenge for the OIV. OENO One 52:231–234. https://doi.org/10.20870/oeno-one.2018.52.3.2312
Daane KM, Almeida RPP, Bell VA, Walker JTS, Botton M et al (2012) Biology and management of mealybugs in Vineyards. In: Arthropod management in vineyards: Springer, Dordrecht, Netherlands, pp 271–307. https://doi.org/10.1007/978-94-007-4032-7_12
Daane KM, Vincent C, Isaacs R, Ioriatti C (2018) Entomological opportunities and challenges for sustainable viticulture in a global market. Annu Rev Entomol 63:193–214. https://doi.org/10.1146/annurev-ento-010715-023547
Daane KM, Yokota GY, Walton VM, Hogg BN, Cooper ML et al (2020) Development of a mating disruption program for a mealybug, Planococcus ficus, in Vineyards. Insects 11:635. https://doi.org/10.3390/insects11090635
Dai L, Wang D, Xie X, Zhang C, Wang X et al (2016) The novel gene VpPR4-1 from Vitis pseudoreticulata increases powdery mildew resistance in Transgenic Vitis vinifera L. Front Plant Sci 7:695. https://doi.org/10.3389/fpls.2016.00695
Dal Bosco D, Sinski I, Ritschel PS, Camargo UA, Fajardo TVM et al (2018) Expression of disease resistance in genetically modified grapevines correlates with the contents of viral sequences in the T-DNA and global genome methylation. Transgenic Res 27:379–396. https://doi.org/10.1007/s11248-018-0082-1
Dal Santo S, Zenoni S, Sandri M, De Lorenzis G, Magris G et al (2018) Grapevine field experiments reveal the contribution of genotype, the influence of environment and the effect of their interaction (G×E) on the berry transcriptome. Plant J 93:1143–1159. https://doi.org/10.1111/tpj.13834
Dalbó MA, Weeden NF, Reisch BI (2000) QTL analysis of disease resistance in interspecific hybrid grapes. Acta Hortic 528:217–222. https://doi.org/10.17660/ActaHortic.2000.528.29
Dalbó MA, Ye GN, Weeden NF, Wilcox WF, Reisch BI (2001) Marker-assisted selection for powdery mildew resistance in grapes. J Am Soc Hortic Sci 126:83–89. https://doi.org/10.21273/JASHS.126.1.83
Dalla Costa L, Mandolini M, Poletti V, Martinelli L (2010) Comparing 17-β-estradiol supply strategies for applying the XVE-Cre/loxP system in grape gene transfer (Vitis vinifera L.). VITIS J Grapevine Res 49:201–208
Dalla Costa L, Piazza S, Campa M, Flachowsky H, Hanke M-V et al (2016) Efficient heat-shock removal of the selectable marker gene in genetically modified grapevine. Plant Cell Tissue Organ Cult 124:471–481. https://doi.org/10.1007/s11240-015-0907-z
Dalla Costa L, Malnoy M, Gribaudo I (2017) Breeding next generation tree fruits: technical and legal challenges. Hortic Res 4:1–11. https://doi.org/10.1038/hortres.2017.67
Dalla Costa L, Malnoy M, Lecourieux D, Deluc L, Ouaked-Lecourieux F et al (2019) The state-of-the-art of grapevine biotechnology and new breeding technologies (NBTS). OENO One 53:205–228. https://doi.org/10.20870/oeno-one.2019.53.2.2405
Dalla Costa L, Piazza S, Pompili V, Salvagnin U, Cestaro A et al (2020) Strategies to produce T-DNA free CRISPRed fruit trees via Agrobacterium tumefaciens stable gene transfer. Sci Rep 10:1–14. https://doi.org/10.1038/s41598-020-77110-1
Dandekar AM, Gouran H, Ibáñez AM, Uratsu SL, Agüero CB et al (2012) An engineered innate immune defense protects grapevines from Pierce's disease. Proc Natl Acad Sci USA 109:3721–3725. https://doi.org/10.1073/pnas.1116027109
Dandekar AM, Jacobson A, Ibáñez AM, Gouran H, Dolan DL et al (2019) Trans-graft protection against Pierce’s disease mediated by Transgenic Grapevine Rootstocks. Front Plant Sci 10:1–10. https://doi.org/10.3389/fpls.2019.00084
Das M, Bhowmick TS, Ahern SJ, Young R, Gonzalez CF (2015) Control of Pierce’s disease by phage. PLoS ONE 10:1–15. https://doi.org/10.1371/journal.pone.0128902
Daykin ME (1984) Histopathology of ripe rot caused by Colletotrichum gloeosporioides on muscadine grape. Phytopathology 74:1339. https://doi.org/10.1094/Phyto-74-1339
De Andrés MT, Benito A, Pérez-Rivera G, Ocete R, Lopez MA et al (2012) Genetic diversity of wild grapevine populations in Spain and their genetic relationships with cultivated grapevines. Mol Ecol 21:800–816. https://doi.org/10.1111/j.1365-294X.2011.05395.x
De Cleene M, De Ley J (1976) The host range of crown gall. Bot Rev 42:389–466. https://doi.org/10.1007/BF02860827
de Lillo E, Pozzebon A, Valenzano D, Duso C (2018) An intimate relationship between eriophyoid mites and their host plants–a review. Front Plant Sci 9:1–14. https://doi.org/10.3389/fpls.2018.01786
De Lorenzis G, Chipashvili R, Failla O, Maghradze D (2015) Study of genetic variability in Vitis vinifera L. germplasm by high-throughput Vitis18kSNP array: the case of Georgian genetic resources. BMC Plant Biol 15:154. https://doi.org/10.1186/s12870-015-0510-9
Delmas CEL, Fabre F, Jolivet J, Mazet ID, Richart Cervera S et al (2016) Adaptation of a plant pathogen to partial host resistance: Selection for greater aggressiveness in grapevine downy mildew. Evol Appl 9:709–725. https://doi.org/10.1111/eva.12368
Delmas CEL, Dussert Y, Delière L, Couture C, Mazet ID et al (2017) Soft selective sweeps in fungicide resistance evolution: recurrent mutations without fitness costs in grapevine downy mildew. Mol Ecol 26:1936–1951. https://doi.org/10.1111/mec.14006
Delrot S, Grimplet J, Carbonell-bejerano P, Schwandner A, Bert P et al (2020) Genetic and genomic approaches for adaptation of grapevine to climate change. In: Kole C (ed) Genomic designing of climate-smart fruit crops. Springer, Cham, Switzerland, pp 157–270
Demangeat G, Voisin R, Minot J-C, Bosselut N, Fuchs M et al (2005) Survival of Xiphinema index in vineyard soil and retention of Grapevine Fanleaf Virus over extended time in the absence of host plants. Phytopathology 95:1151–1156. https://doi.org/10.1094/PHYTO-95-1151
Deprá M, Poppe JL, Schmitz HJ, De Toni DC, Valente VLS (2014) The first records of the invasive pest Drosophila suzukii in the South American continent. J Pest Sci 87:379–383. https://doi.org/10.1007/s10340-014-0591-5
Detjen LR (1919) Some F1 hybrids of Vitis rotundifolia with related species and genera. North Carolina Agri Exp Stat Technical Bull 18:1–50
Deytieux-Belleau C, Geny L, Roudet J, Mayet V, Donèche B et al (2009) Grape berry skin features related to ontogenic resistance to Botrytis cinerea. Eur J Plant Pathol 125:551–563. https://doi.org/10.1007/s10658-009-9503-6
Di Gaspero G, Peterlunger E, Testolin R, Edwards KJ, Cipriani G (2000) Conservation of microsatellite loci within the genus Vitis. Theor Appl Genet 101:301–308. https://doi.org/10.1007/s001220051483
Di Gaspero G, Cattonaro F (2010) Application of genomics to grapevine improvement. Aust J Grape Wine Res 16:122–130. https://doi.org/10.1111/j.1755-0238.2009.00072.x
Di Gaspero G, Copetti D, Coleman C, Castellarin SD, Eibach R et al (2012) Selective sweep at the Rpv3 locus during grapevine breeding for downy mildew resistance. Theor Appl Genet 124:277–286. https://doi.org/10.1007/s00122-011-1703-8
Di Genova A, Almeida A, Muñoz-Espinoza C, Vizoso P, Travisany D et al (2014) Whole genome comparison between table and wine grapes reveals a comprehensive catalog of structural variants. BMC Plant Biol 14:7. https://doi.org/10.1186/1471-2229-14-7
Di Marco S, Osti F, Cesari A (2004) Experiments on the control of esca by Trichoderma. Phytopathol Mediterr 43:108–115. https://doi.org/10.14601/Phytopathol_Mediterr-1730
Di Marco S, Osti F, Calzarano F, Roberti R, Veronesi A et al (2011) Effects of grapevine applications of fosetyl-aluminium formulations for downy mildew control on “esca” and associated fungi. Phytopathol Mediterr 50:285–299. https://doi.org/10.14601/Phytopathol_Mediterr-9802
Di Vecchi-Staraz M, Laucou V, Bruno G, Lacombe T, Gerber S et al (2009) Low level of pollen-mediated gene flow from cultivated to wild grapevine: consequences for the evolution of the endangered subspecies Vitis vinifera L. subsp. sylvestris. J Hered 100:66–75. https://doi.org/10.1093/jhered/esn084
Díaz GA, Latorre BA (2013) Efficacy of paste and liquid fungicide formulations to protect pruning wounds against pathogens associated with grapevine trunk diseases in Chile. Crop Prot 46:106–112. https://doi.org/10.1016/j.cropro.2013.01.001
Díaz-Riquelme J, Zhurov V, Rioja C, Pérez-Moreno I, Torres-Pérez R et al (2016) Comparative genome-wide transcriptome analysis of Vitis vinifera responses to adapted and non-adapted strains of two-spotted spider mite, Tetranyhus urticae. BMC Genomics 17:1–15. https://doi.org/10.1186/s12864-016-2401-3
Divilov K, Barba P, Cadle-Davidson L, Reisch BI (2018) Single and multiple phenotype QTL analyses of downy mildew resistance in interspecific grapevines. Theor Appl Genet 131:1133–1143. https://doi.org/10.1007/s00122-018-3065-y
Donald TM, Pellerone F, Adam-Blondon A-F, Bouquet A, Thomas MR et al (2002) Identification of resistance gene analogs linked to a powdery mildew resistance locus in grapevine. TAG Theor Appl Genet 104:610–618. https://doi.org/10.1007/s00122-001-0768-1
Downie DA (1999) Performance of native grape phylloxera on host plants within and among terrestrial islands in Arizona, USA. Oecologia 121:527. https://doi.org/10.1007/s004420050959
Downie DA (2010) Baubles, bangles, and biotypes: a critical review of the use and abuse of the biotype concept. J Insect Sci 10:1–18. https://doi.org/10.1673/031.010.14136
Dry IB, Feechan A, Anderson C, Jermakow AM, Bouquet A et al (2010) Molecular strategies to enhance the genetic resistance of grapevines to powdery mildew. In: Gerling C (ed) Environmental sustainable viticulture practices and practicality. Apple Academic Press Inc, Oakville, Canada, pp 94–105. https://doi.org/10.1111/j.1755-0238.2009.00076.x
Dry I, Riaz S, Fuchs M, Sosnowski M, Thomas M (2019) Scion breeding for resistance to biotic stresses. In: Cantu D, Walker MA (eds) The grape genome. Springer, Cham, Switzerland, pp 319–347. https://doi.org/10.1007/978-3-030-18601-2_15
Du YP, Wang ZS, Zhai H (2011) Grape root cell features related to phylloxera resistance and changes of anatomy and endogenous hormones during nodosity and tuberosity formation. Aust J Grape Wine Res 17:291–297. https://doi.org/10.1111/j.1755-0238.2011.00131.x
Du Y, Dai W, Dietrich CH (2017) Mitochondrial genomic variation and phylogenetic relationships of three groups in the genus Scaphoideus (Hemiptera: Cicadellidae: Deltocephalinae). Sci Rep 7:16908. https://doi.org/10.1038/s41598-017-17145-z
Duggal P, Gillanders EM, Holmes TN, Bailey-Wilson JE (2008) Establishing an adjusted p-value threshold to control the family-wide type 1 error in genome wide association studies. BMC Genomics 9:516. https://doi.org/10.1186/1471-2164-9-516
Duso C, Pasqualetto C (1993) Factors affecting the potential of phytoseiid mites (Acari: Phytoseiidae) as biocontrol agents in North-Italian vineyards. Exp Appl Acarol 17:241–258. https://doi.org/10.1007/BF02337274
Duso C, de Lillo E (1996) Damage and control of eriophyoid mites, grape. In: Lindquist EE, Bruin J, Sabelis MW (eds) Eriophyid mites. Their biology, natural enemies and control. Elsevier Science B.V., Amsterdam, Netherlands, pp 571–582
Duso C, Vettorazzo E (1999) Mite population dynamics on different grape varieties with or without phytoseiids released (Acari: Phytoseiidae). Exp Appl Acarol 23:741–763. https://doi.org/10.1023/A:1006297225577
Duso C, Castagnoli M, Simoni S, Angeli G (2010) The impact of eriophyoids on crops: recent issues on Aculus schlechtendali, Calepitrimerus vitis and Aculops lycopersici. Exp Appl Acarol 51:151–168. https://doi.org/10.1007/s10493-009-9300-0
Duso C, Pozzebon A, Kreiter S, Tixier M-SS, Candolfi M (2012) Management of phytophagous mites in European vineyards. In: Bostanian N, Vincent C, Isaacs R (eds) Arthropod management in vineyards: pests, approaches, and future directions. Springer, Dordrecht, Netherlands, pp 191–217. https://doi.org/10.1007/978-94-007-4032-7_9
Duso C, Van Leeuwen T, Pozzebon A (2020) Improving the compatibility of pesticides and predatory mites: recent findings on physiological and ecological selectivity. Curr Opin Insect Sci 39:63–68. https://doi.org/10.1016/j.cois.2020.03.005
Dussert Y, Mazet ID, Couture C, Gouzy J, Piron MC et al (2019) A high-quality grapevine downy mildew genome assembly reveals rapidly evolving and lineage-specific putative host adaptation genes. Genome Biol Evol 11:954–969. https://doi.org/10.1093/gbe/evz048
Eckerstorfer MF, Heissenberger A, Reichenbecher W, Steinbrecher RA, Waßmann F (2019) An EU perspective on biosafety considerations for plants developed by genome editing and other new genetic modification techniques (nGMs). Front Bioeng Biotechnol 7. https://doi.org/10.3389/fbioe.2019.00031
Eibach R, Zyprian E, Welter L, Töpfer R (2007) The use of molecular markers for pyramiding resistance genes in grapevine breeding. VITIS J Grapevine Res 46:120–124
Eibach R, Töpfer R, Hausmann L (2010) Use of genetic diversity for grapevine resistance breeding. Mitteilungen Klosterneubg 60:332–337
Eichmeier A, Pečenka J, Peňázová E, Baránek M, Català-García S et al (2018) High-throughput amplicon sequencing-based analysis of active fungal communities inhabiting grapevine after hot-water treatments reveals unexpectedly high fungal diversity. Fungal Ecol 36:26–38. https://doi.org/10.1016/j.funeco.2018.07.011
Eisenmann B, Czemmel S, Ziegler T, Buchholz G, Kortekamp A et al (2019) Rpv3–1 mediated resistance to grapevine downy mildew is associated with specific host transcriptional responses and the accumulation of stilbenes. BMC Plant Biol 19:343. https://doi.org/10.1186/s12870-019-1935-3
Eitle MW, Carolan JC, Griesser M, Forneck A (2019a) The salivary gland proteome of root-galling grape phylloxera (Daktulosphaira vitifoliae Fitch) feeding on Vitis spp. PLoS ONE 14:1–18. https://doi.org/10.1371/journal.pone.0225881
Eitle MW, Griesser M, Vankova R, Dobrev P, Aberer S et al (2019b) Grape phylloxera (D. vitifoliae) manipulates SA/JA concentrations and signalling pathways in root galls of Vitis spp. Plant Physiol Biochem 144:85–91. https://doi.org/10.1016/j.plaphy.2019.09.024
Eitle MW, Loacker J, Meng-Reiterer J, Schuhmacher R, Griesser M et al (2019c) Polyphenolic profiling of roots (Vitis spp.) under grape phylloxera (D. vitifoliae Fitch) attack. Plant Physiol Biochem 135:174–181. https://doi.org/10.1016/j.plaphy.2018.12.004
Ekhvaia J, Gurushidze M, Blattner FR, Akhalkatsi M (2014) Genetic diversity of Vitis vinifera in Georgia: relationships between local cultivars and wild grapevine, V. vinifera L. subsp. sylvestris. Genet Resour Crop Evol 61:1507–1521. https://doi.org/10.1007/s10722-014-0125-2
Elad Y, Vivier M, Fillinger S (2016) Botrytis, the good, the bad and the ugly. In: Fillinger S, Elad Y (eds) Botrytis – the fungus, the pathogen and its management in agricultural systems. Springer, Cham, Switzerland, pp 1–15. https://doi.org/10.1007/978-3-319-23371-0_1
Elbeaino T, Kiyi H, Boutarfa R, Minafra A, Martelli GP et al (2014) Phylogenetic and recombination analysis of the homing protein domain of Grapevine Fanleaf virus (GFLV) isolates associated with ‘yellow mosaic’ and ‘infectious malformation’ syndromes in grapevine. Arch Virol 159:2757–2764. https://doi.org/10.1007/s00705-014-2138-8
Elfert M, Ulrich D, Fischer M, Hoffmann C, Strumpf T (2013) Auf der suche nach Biomarkern im Weinblattmetabolom. J Fur Kult 65:19–23. https://doi.org/10.5073/JFK.2013.01.03
Emanuelli F, Lorenzi S, Grzeskowiak L, Catalano V, Stefanini M et al (2013) Genetic diversity and population structure assessed by SSR and SNP markers in a large germplasm collection of grape. BMC Plant Biol 13:39. https://doi.org/10.1186/1471-2229-13-39
Emanuelli F, Sordo M, Lorenzi S, Battilana J, Grando MS (2014) Development of user-friendly functional molecular markers for VvDXS gene conferring muscat flavor in grapevine. Mol Breed 33:235–241. https://doi.org/10.1007/s11032-013-9929-6
EPPO (2020a) Drosophila suzukii Datasheet. EPPO Global Database. https://gd.eppo.int/taxon/DROSSU/datasheet
EPPO (2020b) https://www.eppo.int/RESOURCES/eppo_standards
EPPO (2021) Scaphoideus titanus World distribution. EPPO Global Database.
Eriksson A, Anfora G, Lucchi A, Lanzo F, Virant-Doberlet M et al (2012) Exploitation of insect vibrational signals reveals a new method of pest management. PLoS ONE 7:32954. https://doi.org/10.1371/journal.pone.0032954
Erincik O, Madden LV, Ferree DC, Ellis MA (2003) Temperature and wetness-duration requirements for grape leaf and cane infection by Phomopsis viticola. Plant Dis 87:832–840. https://doi.org/10.1094/PDIS.2003.87.7.832
Esmenjaud D, Bouquet A (2009) Selection and application of resistant germplasm for grapevine nematodes management. In: Ciancio A, Mukerji KG (eds) Integrated management of fruit crops nematodes. Springer, Dordrecht, Netherlands, pp 195–214. https://doi.org/10.1007/978-1-4020-9858-1_8
Espinoza C, Schlechter R, Herrera D, Torres E, Serrano A et al (2013) Cisgenesis and intragenesis: new tools for improving crops. Biol Res 46:323–331. https://doi.org/10.4067/S0716-97602013000400003
EU Explanatory Note (2017) New techniques in Agricultural Biotechnology
EUROSTAT (2019) Agriculture, forestry and fishery statistics. doi:10.2785/743056
Eveillard S, Jollard C, Labroussaa F, Khalil D, Perrin M et al (2016) Contrasting susceptibilities to Flavescence dorée in Vitis vinifera, rootstocks and wild Vitis species. Front Plant Sci 7:1–12. https://doi.org/10.3389/fpls.2016.01762
Luo F, Zang F (1990) Grape breeding in China. VITIS J Grapevine Res 29:212. https://doi.org/10.5073/vitis.1990.29.special-issue.212-218
FAOSTAT (2014) Sustainability Dimensions. FAO Statistical Yearbook 2014:105–130
FAO-OIV (2016) Table and Dried Grapes. Non-alcoholic products of the vitivinicultural sector intended for human consumption.
Fedele G, González-Domínguez E, Delière L, Díez-Navajas AM, Rossi V (2020) Consideration of latent infections improves the prediction of Botrytis bunch rot severity in vineyards. Plant Dis 104:1291–1297. https://doi.org/10.1094/PDIS-11-19-2309-RE
Feechan A, Anderson C, Torregrosa L, Jermakow A, Mestre P et al (2013) Genetic dissection of a TIR-NB-LRR locus from the wild North American grapevine species Muscadinia rotundifolia identifies paralogous genes conferring resistance to major fungal and oomycete pathogens in cultivated grapevine. Plant J 76:661–674. https://doi.org/10.1111/tpj.12327
Feechan A, Kocsis M, Riaz S, Zhang W, Gadoury DM et al (2015) Strategies for Run1 deployment using Run2 and Ren2 to manage grapevine powdery mildew informed by studies of race specificity. Phytopathology 105:1104–1113. https://doi.org/10.1094/PHYTO-09-14-0244-R
Feliciano AJ, Eskalen A, Gubler WD (2004) Differential susceptibility of three grapevine cultivars to Phaeoacremonium aleophilum and Phaeomoniella chlamydospora in California. Phytopathol Mediterr 43:66–69. https://doi.org/10.14601/Phytopathol_Mediterr-1727
Ferrin DM (1977) Ascospore dispersal and infection of grapes by Guignardia bidwellii, the causal agent of grape black rot disease. Phytopathology 77:1501. https://doi.org/10.1094/Phyto-67-1501
Ferris H, Zheng L, Walker MA (2012) Resistance of grape rootstocks to plant-parasitic nematodes. J Nematol 44:377–386
Ferris H, Zheng L, Walker MA (2013) Soil temperature effects on the interaction of grape rootstocks and plant-parasitic nematodes. J Nematol 45:49–57
Firrao G, Andersen M, Bertaccini A, Boudon E, Bové JM et al (2004) “Candidatus Phytoplasma”, a taxon for the wall-less, non-helical prokaryotes that colonize plant phloem and insects. Int J Syst Evol Microbiol 54:1243–1255. https://doi.org/10.1099/ijs.0.02854-0
Fischer BM, Salakhutdinov I, Akkurt M, Eibach R, Edwards KJ et al (2004) Quantitative Trait Locus analysis of fungal disease resistance factors on a molecular map of grapevine. Theor Appl Genet 108:501–515. https://doi.org/10.1007/s00122-003-1445-3
Fischer M, Ashnaei SP (2019) Grapevine, esca complex, and environment: the disease triangle. Phytopathol Mediterr 58:17–37. https://doi.org/10.13128/Phytopathol_Mediterr-25086
Flor HH (1942) Inheritance of pathogenicity in Melampsora lini. Phytopathology 32:653–669
Fong G, Walker MA, Granett J (1995) RAPD assessment of California phylloxera diversity. Mol Ecol 4:459–464. https://doi.org/10.1111/j.1365-294X.1995.tb00239.x
Fontaine MC, Austerlitz F, Giraud T, Labbé F, Papura D et al (2013) Genetic signature of a range expansion and leap-frog event after the recent invasion of Europe by the grapevine downy mildew pathogen Plasmopara viticola. Mol Ecol 22:2771–2786. https://doi.org/10.1111/mec.12293
Fontaine MC, Labbé F, Dussert Y, Delière L, Richart-Cervera S et al (2021) Europe as a bridgehead in the worldwide invasion history of grapevine downy mildew, Plasmopara viticola. Curr Biol 1–12. https://doi.org/10.1016/j.cub.2021.03.009
Foria S, Monte C, Testolin R, Di Gaspero G, Cipriani G (2019) Pyramidizing resistance genes in grape: a breeding program for the selection of elite cultivars. Acta Hortic 1248:549–554. https://doi.org/10.17660/ActaHortic.2019.1248.73
Foria S, Copetti D, Eisenmann B, Magris G, Vidotto M et al (2020) Gene duplication and transposition of mobile elements drive evolution of the Rpv3 resistance locus in grapevine. Plant J 101:529–542. https://doi.org/10.1111/tpj.14551
Forneck A, Huber L (2009) (A)sexual reproduction-a review of life cycles of grape phylloxera, Daktulosphaira vitifoliae. Entomol Exp Appl 131:1–10. https://doi.org/10.1111/j.1570-7458.2008.00811.x
Forneck A, Anhalt UCM, Mammerler R, Griesser M (2015) No evidence of superclones in leaf-feeding forms of Austrian grape phylloxera (Daktulosphaira vitifoliae). Eur J Plant Pathol 142:441–448. https://doi.org/10.1007/s10658-015-0624-9
Forneck A, Powell KS, Walker MA (2016) Scientific opinion: improving the definition of grape phylloxera biotypes and standardizing biotype screening protocols. Am J Enol Vitic 67:371–376. https://doi.org/10.5344/ajev.2016.15106
Forneck A, Dockner V, Mammerler R, Powell KS, Kocsis L et al (2017) PHYLLI–an international database for grape phylloxera (Daktulosphaira vitifoliae Fitch). Integr Prot Prod Vitic 128:45–51
Fraga H, Pinto JG, Santos JA (2019) Climate change projections for chilling and heat forcing conditions in European vineyards and olive orchards: a multi-model assessment. Clim Change 152:179–193. https://doi.org/10.1007/s10584-018-2337-5
Frenkel O, Brewer MT, Milgroom MG (2010) Variation in pathogenicity and aggressiveness of Erysiphe necator from different Vitis spp. and geographic origins in the Eastern United States. Phytopathology 100:1185–1193. https://doi.org/10.1094/PHYTO-01-10-0023
Fresnedo-Ramírez J, Yang S, Sun Q, Karn A, Reisch BI et al (2019) Computational analysis of ampseq data for targeted, high-throughput genotyping of amplicons. Front Plant Sci 10:599. https://doi.org/10.3389/fpls.2019.00599
Fritschi FB, Lin H, Walker MA (2007) Xylella fastidiosa population dynamics in grapevine genotypes differing in susceptibility to Pierce’s disease. Am J Enol Vitic 58:326–332
Fu P, Tian Q, Lai G, Li R, Song S et al (2019) Cgr1, a ripe rot resistance QTL in Vitis amurensis ‘Shuang Hong’ grapevine. Hortic Res 6:67. https://doi.org/10.1038/s41438-019-0148-0
Fu P, Wu W, Lai G, Li R, Peng Y et al (2020) Identifying Plasmopara viticola resistance loci in grapevine (Vitis amurensis) via genotyping-by-sequencing-based QTL mapping. Plant Physiol Biochem 154:75–84. https://doi.org/10.1016/j.plaphy.2020.05.016
Fussler L, Kobes N, Bertrand F, Maumy M, Grosman J et al (2008) A characterization of grapevine trunk diseases in France from data generated by the National Grapevine Wood Diseases Survey. Phytopathology 98:571–579. https://doi.org/10.1094/PHYTO-98-5-0571
Gabler FM, Smilanick JL, Mansour M, Ramming DW, Mackey BE (2003) Correlations of morphological, anatomical, and chemical features of grape berries with resistance to Botrytis cinerea. Phytopathology 93:1263–1273. https://doi.org/10.1094/PHYTO.2003.93.10.1263
Gadoury DM, Pearson CP (1991) Heterothallism and pathogenic specialization in Uncinula necator. Phytopathology 81:1287. https://doi.org/10.1094/Phyto-81-1287
Gadoury DM, Cadle-Davidson L, Wilcox WF, Dry IB, Seem RC et al (2012) Grapevine powdery mildew (Erysiphe necator): A fascinating system for the study of the biology, ecology and epidemiology of an obligate biotroph. Mol Plant Pathol 13:1–16. https://doi.org/10.1111/j.1364-3703.2011.00728.x
Gadoury DM, Wilcox WF, Rumbolz J, Gubler WD (2019) Powdery mildew. In: Wilcox WF, Gubler WD, Uyemoto JK (eds) Compendium of grape diseases, disorders, and pests, 2nd edn. APS Press, St. Paul, MN, pp 75–83
Galambos A, Zok A, Kuczmog A, Oláh R, Putnoky P et al (2013) Silencing Agrobacterium oncogenes in transgenic grapevine results in strain-specific crown gall resistance. Plant Cell Rep 32:1751–1757. https://doi.org/10.1007/s00299-013-1488-0
Gale, G Sandler, M Pinder R (2002) Saving the vine from Phylloxera: a never-ending battle. Wine a Sci Explor 70–91
Galet P (1988) Les vignes américaines. In: Cépages et vignobles de France. Charles Dehan, Montpellier, France
Gambino G, Gribaudo I (2012) Genetic transformation of fruit trees: current status and remaining challenges. Transgenic Res 21:1163–1181. https://doi.org/10.1007/s11248-012-9602-6
Gao X, Becker LC, Becker DM, Starmer JD, Province MA (2010) Avoiding the high Bonferroni penalty in genome-wide association studies. Genet Epidemiol 34:100–105. https://doi.org/10.1002/gepi.20430
Gessler C, Pertot I, Perazzolli M (2011) Plasmopara viticola: A review of knowledge on downy mildew of grapevine and effective disease management. Phytopathol Mediterr 50:3–44. https://doi.org/10.14601/Phytopathol_Mediterr-9360
Giannetto S, Velasco R, Troggio M, Malacarne G, Storchi P et al (2008) A PCR-based diagnostic tool for distinguishing grape skin color mutants. Plant Sci 175:402–409. https://doi.org/10.1016/j.plantsci.2008.05.010
Gilligan TM, Epstein ME, Passoa SC, Powell JA, Sage OC et al (2011) Discovery of Lobesia botrana ([Denis & Schiffermller]) in California: An invasive species new to North America (Lepidoptera: Tortricidae). Proc Entomol Soc Washingt 113:14–30. https://doi.org/10.4289/0013-8797.113.1.14
Gindro K, Alonso-Villaverde V, Viret O, Spring J-L, Marti G et al (2012) Stilbenes: biomarkers of grapevine resistance to disease of high relevance for agronomy, oenology and human health. In: Mérillon JM, Ramawat KG (eds) Plant defence: biological control. Springer, Dordrecht, Netherlands, pp 25–54. https://doi.org/10.1007/978-94-007-1933-0_2
Girolami V (1981) Danni, soglie di intervento, controllo degli acari della vite. In: Proceedings, III incontro sulla difesa integrata della vite, 3–4 December 1981, Latina, Italy, pp 111–143
Gobbin D, Pertot I, Gessler C (2003) Genetic structure of a Plasmopara viticola population in an isolated Italian mountain vineyard. J Phytopathol 151:636–646. https://doi.org/10.1046/j.0931-1785.2003.00779.x
Gobbin D, Rumbou A, Linde CC, Gessler C (2006) Population genetic structure of Plasmopara viticola after 125 years of colonization in European vineyards. Mol Plant Pathol 7:519–531. https://doi.org/10.1111/j.1364-3703.2006.00357.x
Godefroid M, Cruaud A, Streito J-C, Rasplus J-Y, Rossi J-P (2019) Xylella fastidiosa: climate suitability of European continent. Sci Rep 9:8844. https://doi.org/10.1038/s41598-019-45365-y
Gomes BR, Bogo A, Copatti A, Guginski-Piva CA, de Morais AC et al (2019) Assessment of grapevine germoplasm collection for resistance to grape leaf rust (Phakopsora euvitis) using a leaf disc assay. Euphytica 215:1–11. https://doi.org/10.1007/s10681-019-2514-2
Gonzalez M (2010) Lobesia botrana: Polilla de la uva. Enología 1–5
González-Domínguez E, Caffi T, Ciliberti N, Rossi V (2015) A mechanistic model of Botrytis cinerea on grapevines that includes weather, vine growth stage, and the main infection pathways. PLoS ONE 10(10):e0140444. https://doi.org/10.1371/journal.pone.0140444
Gonzalez-Dominguez E, Caffi T, Languasco L, Latinovic N, Latinovic J et al (2021) Dynamics of Diaporthe ampelina conidia released from grape canes that overwintered in the vineyard. Plant Dis 1–38. https://doi.org/10.1094/pdis-12-20-2639-re
Gramaje D, García-Jiménez J, Armengol J (2010) Field evaluation of grapevine rootstocks inoculated with fungi associated with petri disease and esca. Am J Enol Vitic 61:512–520. https://doi.org/10.5344/ajev.2010.10021
Gramaje D, Mostert L, Armengol J (2011) Characterization of Cadophora luteo-olivacea and C. melinii isolates obtained from grapevines and environmental samples from grapevine nurseries in Spain. Phytopathol Mediterr 50:112–126. https://doi.org/10.14601/Phytopathol_Mediterr-8723
Gramaje D, Armengol J (2011) Fungal trunk pathogens in the grapevine propagation process: potential inoculum sources, detection, identification, and management strategies. Plant Dis 95:1040–1055. https://doi.org/10.1094/PDIS-01-11-0025
Gramaje D, Armengol J, Ridgway HJ (2013) Genetic and virulence diversity, and mating type distribution of Togninia minima causing grapevine trunk diseases in Spain. Eur J Plant Pathol 135:727–743. https://doi.org/10.1007/s10658-012-0110-6
Gramaje D, León M, Santana M, Crous PW, Armengol J (2014) Multilocus ISSR markers reveal two major genetic groups in Spanish and South African populations of the grapevine fungal pathogen Cadophora luteo-olivacea. PLoS ONE 9(10):e0110417. https://doi.org/10.1371/journal.pone.0110417
Gramaje D, Di Marco S (2015) Identifying practices likely to have impacts on grapevine trunk disease infections: a European nursery survey. Phytopathol Mediterr 54:313–324. https://doi.org/10.14601/Phytopathol_Mediterr-16317
Gramaje D, Mostert L, Groenewald JZ, Crous PW (2015) Phaeoacremonium: from esca disease to phaeohyphomycosis. Fungal Biol 119:759–783. https://doi.org/10.1016/j.funbio.2015.06.004
Gramaje D, Urbez-Torres JR, Sosnowski MR (2018) Managing grapevine trunk diseases with respect to etiology and epidemiology: current strategies and future prospects. Plant Dis 102:12–39. https://doi.org/10.1094/PDIS-04-17-0512-FE
Granett J, Timper P, Lider LA (1985) Grape Phylloxera (Daktulosphaira vitifoliae) (Homoptera: Phylloxeridae) Biotypes in California. J Econ Entomol 78:1463–1467. https://doi.org/10.1093/jee/78.6.1463
Granett J, Walker MA, Kocsis L, Omer AD (2001) Biology and management of grape phylloxera. Annu Rev Entomol 46:387–412. https://doi.org/10.1146/annurev.ento.46.1.387
Grassi F, Arroyo-García R (2020) Editorial: Origins and domestication of the grape. Front Plant Sci 11:1176. https://doi.org/10.3389/fpls.2020.01176
Gray DJ, Li ZT, Dhekney SA (2014) Precision breeding of grapevine (Vitis vinifera L.) for improved traits. Plant Sci 228:3–10. https://doi.org/10.1016/j.plantsci.2014.03.023
Gribaudo I, Gambino G, Boccacci P, Perrone I, Cuozzo D (2017) A multi-year study on the regenerative potential of several Vitis genotypes. Acta Hortic 1155:45–50. https://doi.org/10.17660/ActaHortic.2017.1155.5
Griesser M, Lawo NC, Crespo-Martinez S, Schoedl-Hummel K, Wieczorek K et al (2015) Phylloxera (Daktulosphaira vitifoliae Fitch) alters the carbohydrate metabolism in root galls to allowing the compatible interaction with grapevine (Vitis ssp.) roots. Plant Sci 234:38–49. https://doi.org/10.1016/j.plantsci.2015.02.002
Grimplet J, Cramer GR, Dickerson JA, Mathiason K, van Hemert J et al (2009) VitisNet: “Omics” integration through grapevine molecular networks. PLoS ONE 4:e8365. https://doi.org/10.1371/journal.pone.0008365
Grimplet J, Van Hemert J, Carbonell-Bejerano P, Díaz-Riquelme J, Dickerson J et al (2012) Comparative analysis of grapevine whole-genome gene predictions, functional annotation, categorization and integration of the predicted gene sequences. BMC Res Notes 5:1–10. https://doi.org/10.1186/1756-0500-5-213
Guan X, Essakhi S, Laloue H, Nick P, Bertsch C et al (2016) Mining new resources for grape resistance against Botryosphaeriaceae: a focus on Vitis vinifera subsp. sylvestris. Plant Pathol 65:273–284. https://doi.org/10.1111/ppa.12405
Guarnaccia V, Groenewald JZ, Woodhall J, Armengol J, Cinelli T et al (2018) Diaporthe diversity and pathogenicity revealed from a broad survey of grapevine diseases in Europe. Persoonia Mol Phylogeny Evol Fungi 40:135–153. https://doi.org/10.3767/persoonia.2018.40.06
Gubler WD, Rademacher MR, Vasquez SJ, Thomas CS (1999) Control of powdery mildew using the UC Davis powdery mildew risk index. Apsnet Featur. https://doi.org/10.1094/APSnetFeature-1999-0199
Gubler WD, Leavitt GM, Bettiga LJ (2015) Downy mildew. In: Bettiga LJ (ed) Grape pest management. University of California, Agriculture and Natural Resources, Davis, CA, pp 117–119
Guo C, Guo R, Xu X, Gao M, Li X et al (2014) Evolution and expression analysis of the grape (Vitis vinifera L.) WRKY gene family. J Exp Bot 65:1513–1528. https://doi.org/10.1093/jxb/eru007
Guo R, Qiao H, Zhao J, Wang X, Tu M et al (2018) The grape VlWRKY3 gene promotes abiotic and biotic stress tolerance in transgenic Arabidopsis thaliana. Front Plant Sci 9:1–16. https://doi.org/10.3389/fpls.2018.00545
Gur L, Reuveni M, Cohen Y, Cadle-Davidson L, Kisselstein B et al (2021) Population structure of Erysiphe necator on domesticated and wild vines in the Middle East raises questions on the origin of the grapevine powdery mildew pathogen. Environ Microbiol. https://doi.org/10.1111/1462-2920.15401
Hajdu E (2015) Grapevine breeding in Hungary. In: Reynolds A (ed) Grapevine breeding programs for the wine industry. Elsevier, Woodhead Publishing, Sawston, UK, pp 103–134. https://doi.org/10.1016/B978-1-78242-075-0.00006-5
Halleen F, Fourie PH (2016) An integrated strategy for the proactive management of grapevine trunk disease pathogen infections in grapevine nurseries. South African J Enol Vitic 37:104–114. https://doi.org/10.21548/37-2-825
Hammond-Kosack KE, Jones JDG (2015) Responses to plant pathogens. In: Buchanan BB, Gruissem W, Jones LR (eds) Biochemistry & molecular biology of plants. John Wiley & Sons, Ltd, Hoboken, NJ
Hanif M, Rahman M, Gao M, Yang J, Ahmad B et al (2018) Heterologous expression of the grapevine JAZ7 gene in Arabidopsis confers enhanced resistance to powdery mildew but not to Botrytis cinerea. Int J Mol Sci 19(12):3889. https://doi.org/10.3390/ijms19123889
Hao Z, Fayolle L, van Tuinen D, Chatagnier O, Li X et al (2012) Local and systemic mycorrhiza-induced protection against the ectoparasitic nematode Xiphinema index involves priming of defence gene responses in grapevine. J Exp Bot 63:3657–3672. https://doi.org/10.1093/jxb/ers046
Harlan JR, Wet JMJ (1971) Toward a rational classification of cultivate plants. Taxon 20:509–517. https://doi.org/10.2307/1218252
Hart KH, Magarey RD, Emmett RW, Magarey PA (1993) Susceptibility of grapevine selections to black spot (anthracnose), Elsinöe ampelina. Aust Grapegrow Winemak 352:85–87
Hausmann L, Eibach R, Zyprian E, Töpfer R (2011) Genetic analysis of phylloxera root resistance in cultivar “Börner.” Acta Hortic 904:47–52. https://doi.org/10.17660/ActaHortic.2011.904.6
Hausmann L, Eibach R, Zyprian E, Töpfer R (2014) Sequencing of the phylloxera resistance locus Rdv1 of cultivar “Börner.” Acta Hortic 1046:73–78. https://doi.org/10.17660/ActaHortic.2014.1046.7
Hausmann L, Rex F, Töpfer R (2017) Evaluation and genetic analysis of grapevine black rot resistances. Acta Hortic 1188:285–290. https://doi.org/10.17660/ActaHortic.2017.1188.37
Hausmann L, Maul E, Ganesch A, Töpfer R (2019) Overview of genetic loci for traits in grapevine and their integration into the VIVC database. Acta Hortic 1248:221–226. https://doi.org/10.17660/ActaHortic.2019.1248.32
Haye T, Girod P, Cuthbertson AGS, Wang XG, Daane KM et al (2016) Current SWD IPM tactics and their practical implementation in fruit crops across different regions around the world. J Pest Sci (2004)89:643–651. https://doi.org/10.1007/s10340-016-0737-8
He M, Xu Y, Cao J, Zhu Z, Jiao Y et al (2013) Subcellular localization and functional analyses of a PR10 protein gene from Vitis pseudoreticulata in response to Plasmopara viticola infection. Protoplasma 250:129–140. https://doi.org/10.1007/s00709-012-0384-8
He R, Wu J, Zhang Y, Agüero CB, Li X et al (2017) Overexpression of a thaumatin-like protein gene from Vitis amurensis improves downy mildew resistance in Vitis vinifera grapevine. Protoplasma 254:1579–1589. https://doi.org/10.1007/s00709-016-1047-y
Héloir M-C, Adrian M, Brulé D, Claverie J, Cordelier S et al (2019) Recognition of elicitors in grapevine: from MAMP and DAMP perception to induced resistance. Front Plant Sci 10:e1117. https://doi.org/10.3389/fpls.2019.01117
Hemmer C, Djennane S, Ackerer L, Hleibieh K, Marmonier A et al (2018) Nanobody-mediated resistance to Grapevine Fanleaf Virus in plants. Plant Biotechnol J 16:660–671. https://doi.org/10.1111/pbi.12819
Hennessy CR, Daly AM, Hearnden MN (2007) Assessment of grapevine cultivars for resistance to Phakopsora euvitis. Australas Plant Pathol 36:313–317. https://doi.org/10.1071/AP07028
Herzog K, Wind R, Töpfer R (2015) Impedance of the grape berry cuticle as a novel phenotypic trait to estimate resistance to Botrytis cinerea. Sensors (Switzerland) 15:12498–12512. https://doi.org/10.3390/s150612498
Hewitt WB, Pearson RC (1988) Phomopsis cane and leaf spot. In: Pearson RC, Goheen AC (eds) Compendium of grape diseases. APS Press, St. Paul, MN, pp 16–18
Highet A, Wicks TJ (1998) The incidence of Eutypa dieback in South Australian vineyards. Aust Grapegrow Winemak Annu Tech Issue 441a:135–136
Hill GN, Evans KJ, Beresford RM, Dambergs RG (2014) Comparison of methods for the quantification of Botrytis bunch rot in white wine grapes. Aust J Grape Wine Res 20:432–441. https://doi.org/10.1111/ajgw.12101
Hily JM, Demanèche S, Poulicard N, Tannières M, Djennane S et al (2018) Metagenomic-based impact study of transgenic grapevine rootstock on its associated virome and soil bacteriome. Plant Biotechnol J 16:208–220. https://doi.org/10.1111/pbi.12761
Hoffman LE, Wilcox WF, Gadoury DM, Seem RC, Riegel DG (2004) Integrated control of grape black rot: influence of host phenology, inoculum availability, sanitation, and spray timing. Phytopathology 94:641–650. https://doi.org/10.1094/PHYTO.2004.94.6.641
Hoffmann S, Di Gaspero G, Kovács L, Howard S, Kiss E et al (2008) Resistance to Erysiphe necator in the grapevine “Kishmish vatkana” is controlled by a single locus through restriction of hyphal growth. Theor Appl Genet 116:427–438. https://doi.org/10.1007/s00122-007-0680-4
Holme IB, Wendt T, Holm PB (2013) Intragenesis and cisgenesis as alternatives to transgenic crop development. Plant Biotechnol J 11:395–407. https://doi.org/10.1111/pbi.12055
Holtgräwe D, Rosleff Soerensen T, Hausmann L, Pucker B, Viehöver P et al (2020) A partially phase-separated genome sequence assembly of the Vitis rootstock ‘Börner’ (Vitis riparia × Vitis cinerea) and its exploitation for marker development and targeted mapping. Front Plant Sci 11:e00156. https://doi.org/10.3389/fpls.2020.00156
Hopkins DL (2005) Biological control of Pierce’s disease in the vineyard with strains of Xylella fastidiosa benign to grapevine. Plant Dis 89:1348–1352. https://doi.org/10.1094/PD-89-1348
Hopkins DL, Harris JW (2000) A Greenhouse Method for screening grapevine seedlings for resistance to anthracnose. HortScience 35:89–91. https://doi.org/10.21273/HORTSCI.35.1.89
Hou S, Liu Z, Shen H, Wu D (2019) Damage-associated molecular pattern-triggered Immunity in Plants. Front Plant Sci 10. https://doi.org/10.3389/fpls.2019.00646
Houle D, Govindaraju DR, Omholt S (2010) Phenomics: the next challenge. Nat Rev Genet 11:855–866
Hu Y, Li Y, Hou F, Wan D, Cheng Y et al (2018) Ectopic expression of Arabidopsis broad-spectrum resistance gene RPW8.2 improves the resistance to powdery mildew in grapevine (Vitis vinifera). Plant Sci 267:20–31. https://doi.org/10.1016/j.plantsci.2017.11.005
Hu Q, Zhu L, Zhang X, Guan Q, Xiao S et al (2018) GhCPK33 negatively regulates defense against Verticillium dahliae by phosphorylating GhOPR3. Plant Physiol 178:876–889. https://doi.org/10.1104/pp.18.00737
Hu Y, Gao YR, Yang LS, Wang W, Wang YJ et al (2019) The cytological basis of powdery mildew resistance in wild Chinese Vitis species. Plant Physiol Biochem 144:244–253. https://doi.org/10.1016/j.plaphy.2019.09.049
Hu Y, Cheng Y, Yu X, Liu J, Yang L et al (2021) Overexpression of two CDPKs from wild Chinese grapevine enhances powdery mildew resistance in Vitis vinifera and Arabidopsis. New Phytol 230:2029–2046. https://doi.org/10.1111/nph.17285
Hwang C-F, Xu K, Hu R, Zhou R, Riaz S et al (2010) Cloning and characterization of XiR1, a locus responsible for dagger nematode resistance in grape. Theor Appl Genet 121:789–799. https://doi.org/10.1007/s00122-010-1349-y
Hyma KE, Barba P, Wang M, Londo JP, Acharya CB et al (2015) Heterozygous mapping strategy (HetMappS) for high resolution genotyping-by-sequencing markers: a case study in grapevine. PLoS ONE 10:e0134880. https://doi.org/10.1371/journal.pone.0134880
Ibáñez J, Muñoz-Organero G, Zinelabidine LH, de Andrés MT, Cabello F et al (2012) Genetic origin of the grapevine cultivar Tempranillo. Am J Enol Vitic 63:549–553. https://doi.org/10.5344/ajev.2012.12012
Igolkina AA, Meshcheryakov G, Gretsova MV, Nuzhdin SV, Samsonova MG (2020) Multi-trait multi-locus SEM model discriminates SNPs of different effects. BMC Genomics 21:490. https://doi.org/10.1186/s12864-020-06833-2
Ilnitskaya E, Makarkina M, Tokmakov S, Kotlyar V (2020) DNA-marker identification of Rpv3 and Rpv12 resistance loci in genotypes of table and seedless grape varieties. In: BIO web of conferences. EDP sciences, vol 25, pp 03004. https://doi.org/10.1051/bioconf/20202503004
Ioriatti C, Anfora G, Tasin M, De Cristofaro A, Witzgall P et al (2011) Chemical ecology and management of Lobesia botrana (Lepidoptera: Tortricidae). J Econ Entomol 104:1125–1137. https://doi.org/10.1603/EC10443
Ioriatti C, Walton V, Dalton D, Anfora G, Grassi A et al (2015) Drosophila suzukii (Diptera: Drosophilidae) and its potential impact to wine grapes during harvest in two cool climate wine grape production regions. J Econ Entomol 108:1148–1155. https://doi.org/10.1093/jee/tov042
Ioriatti C, Lucchi A (2016) Semiochemical strategies for tortricid moth control in apple orchards and vineyards in Italy. J Chem Ecol 42:571–583. https://doi.org/10.1007/s10886-016-0722-y
Ioriatti C, Guzzon R, Anfora G, Ghidoni F, Mazzoni V et al (2018) Drosophila suzukii (Diptera: Drosophilidae) contributes to the development of sour rot in grape. J Econ Entomol 111:283–292. https://doi.org/10.1093/jee/tox292
IPCC (2013) Climate change 2013 the physical science basis: working group I contribution to the fifth assessment report of the intergovernmental panel on climate change. Cambridge University Press UK
Jang HA, Oo MM, Kim D-GG, Yoon H-YY, Kim M-RR et al (2020) CC-NBS-LRR, a set of VvCRP markers, can distinguish cultivars with ripe rot resistance to Colletotrichum pathogens in grapevine. Hortic Environ Biotechnol 61:915–927. https://doi.org/10.1007/s13580-020-00290-2
Jang MH, Moon YS, Noh JH, Kim SH, Hong SK et al (2011) In vitro evaluation system for varietal resistance against ripe rot caused by Colletotrichum acutatum in grapevines. Hortic Environ Biotechnol 52:52–57. https://doi.org/10.1007/s13580-011-0190-9
Jarausch W, Angelini E, Eveillard, Sandrine Malembic-Maher S (2013) Management of fruit tree and grapevine phytoplasma diseases through genetic resistance. In: COST Action FA0807 integrated management of phytoplasma epidemics in different crop systems. New perspectives in phytoplasma disease management: Book of abstracts of presentations Work Groups 2–3. p 56
Javadi Khederi S, Khanjani M, Fayaz BA (2014a) Resistance of three grapevine cultivars to Grape Erineum Mite, Colomerus vitis (Acari: Eriophyidae), in field conditions. Persian J Acarol 3(1). https://doi.org/10.22073/pja.v3i1.10132
Javadi Khederi S, de Lillo E, Khanjani M, Gholami M (2014b) Resistance of grapevine to the erineum strain of Colomerus vitis (Acari: Eriophyidae) in Western Iran and its correlation with plant features. Exp Appl Acarol 63:15–35. https://doi.org/10.1007/s10493-014-9778-y
Javadi Khederi S, Khanjani M, Gholami M, de Lillo E (2018a) Impact of the erineum strain of Colomerus vitis (Acari: Eriophyidae) on the development of plants of grapevine cultivars of Iran. Exp Appl Acarol 74:347–363. https://doi.org/10.1007/s10493-018-0245-z
Javadi Khederi S, Khanjani M, Gholami M, Panzarino O, de Lillo E (2018b) Influence of the erineum strain of Colomerus vitis (Acari: Eriophyidae) on grape (Vitis vinifera) defense mechanisms. Exp Appl Acarol 75:1–24. https://doi.org/10.1007/s10493-018-0252-0
Javadi Khederi S, Khanjani M, Gholami M, Bruno GL (2018c) Study of defense-related gene expression in grapevine infested by Colomerus vitis (Acari: Eriophyidae). Exp Appl Acarol 75:25–40. https://doi.org/10.1007/s10493-018-0255-x
Javadi Khederi S, Khanjani M, Gholami M, De Lillo E (2018d) Sources of resistance to the erineum strain of Colomerus vitis (Acari: Eriophyidae) in grapevine cultivars. Syst Appl Acarol 23:405. https://doi.org/10.11158/saa.23.3.1
Jeger M, Bragard C, Caffier D, Candresse T, Chatzivassiliou E et al (2016) Risk to plant health of Flavescence dorée for the EU territory. EFSA J 14:12. https://doi.org/10.2903/j.efsa.2016.4603
Jelly NS, Schellenbaum P, Walter B, Maillot P (2012) Transient expression of artificial microRNAs targeting Grapevine Fanleaf Virus and evidence for RNA silencing in grapevine somatic embryos. Transgenic Res 21:1319–1327. https://doi.org/10.1007/s11248-012-9611-5
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA et al (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science (80-) 337:816–821. https://doi.org/10.1126/science.1225829
Johann L, do Nascimento JM, da Silva GL, Silva Carvalho G, Juarez Ferla N (2019) Life history and life table parameters of Panonychus ulmi (Acari: Tetranychidae) on two European grape cultivars. Phytoparasitica 47:79–86. https://doi.org/10.1007/s12600-018-00709-8
Jollard C, Malembic-Maher S, Labroussaa F, Khalil D, Perrin M et al (2019) Contrasting susceptibilities to Flavescence dorée in Vitis vinifera cultivars and progenies suggest segregation of genetic traits involved in disease response. Acta Hortic 1248:601–606. https://doi.org/10.17660/ActaHortic.2019.1248.81
Jones JDG, Dangl JL (2006) The plant immune system. Nature 444:323–329. https://doi.org/10.1038/nature05286
Jones L, Riaz S, Morales-Cruz A, Amrine KC, McGuire B et al (2014) Adaptive genomic structural variation in the grape powdery mildew pathogen, Erysiphe necator. BMC Genomics 15:1081. https://doi.org/10.1186/1471-2164-15-1081
Jørgensen IH (1992) Discovery, characterization and exploitation of MLO powdery mildew resistance in barley. Euphytica 63:141–152. https://doi.org/10.1007/BF00023919
Jung J-H, Joe M-H, Kim D-H, Park H, Choi J et al (2018) Complete genome sequence of Planococcus sp. PAMC21323 isolated from Antarctica and its metabolic potential to detoxify pollutants. Stand Genomic Sci 13:31. https://doi.org/10.1186/s40793-018-0334-y
Jürges G, Kassemeyer HH, Dürrenberger M, Düggelin M, Nick P (2009) The mode of interaction between Vitis and Plasmopara viticola Berk. & Curt. Ex de Bary depends on the host species. Plant Biol 11:886–898. https://doi.org/10.1111/j.1438-8677.2008.00182.x
Kaler AS, Purcell LC (2019) Estimation of a significance threshold for genome-wide association studies. BMC Genomics 20:618. https://doi.org/10.1186/s12864-019-5992-7
Kaliterna J, Miličević T, Cvjetković B (2012) Grapevine trunk diseases associated with fungi from the Diaporthaceae family in Croatian vineyards. Arh Hig Rada Toksikol 63:471–479. https://doi.org/10.2478/10004-1254-63-2012-2226
Kantor A, McClements M, MacLaren R (2020) CRISPR-Cas9 DNA base-editing and prime-editing. Int J Mol Sci 21:6240. https://doi.org/10.3390/ijms21176240
Kaplan J, Travadon R, Cooper M, Hillis V, Lubell M et al (2016) Identifying economic hurdles to early adoption of preventative practices: the case of trunk diseases in California winegrape vineyards. Wine Econ Policy 5:127–141. https://doi.org/10.1016/j.wep.2016.11.001
Karn A, Zou C, Brooks S, Fresnedo-Ramírez J, Gabler F, Sun Q, Ramming D, Naegele R, Ledbetter C, Cadle-Davidson L (2021) Discovery of the Ren11 locus from Vitis aestivalis for stable resistance to grapevine powdery mildew in a family segregating for several unstable and tissue-specific quantitative resistance loci. Front Plant Sci 12:733899. https://doi.org/10.3389/fpls.2021.733899
Kassemeyer H-H, Gadoury DM, Hill G, Wilcox WF (2019) Downy mildew. In: Wilcox WF, Gubler WD, Uyemoto JK (eds) Compendium of grape diseases, disorders, and pests. APS Press, St. Paul, MN, pp 46–52
Keller B, Feuillet C, Messmer M (2000) Genetics of disease resistance. In: Slusarenko AJ, Fraser RSS, van Loon LC (eds) Mechanisms of resistance to plant diseases. Springer, Dordrecht, Netherlands, pp 101–160. https://doi.org/10.1007/978-94-011-3937-3_5
Kellow AV (2004) Interaction between Vitis vinifera and grape phylloxera: changes in root tissue during nodosity formation. Ann Bot 93:581–590. https://doi.org/10.1093/aob/mch082
Kim ES, Chang EH, Hur YY, Kim TW, Jung SM (2015) Anthocyanin contents and composition of VlmybA1–2 and VlmybA2 genes in Vitis labrusca hybrid grape cultivars and cross seedlings. Plant Omics 8:472–478. https://doi.org/10.3316/INFORMIT.516994550945748
Kim GH, Yun HK, Choi CS, Park JH, Jung YJ et al (2008) Identification of AFLP and RAPD markers linked to anthracnose resistance in grapes and their conversion to SCAR markers. Plant Breed 127:418–423. https://doi.org/10.1111/j.1439-0523.2008.01488.x
Klein LL, Miller AJ, Ciotir C, Hyma K, Uribe-Convers S et al (2018) High-throughput sequencing data clarify evolutionary relationships among North American Vitis species and improve identification in USDA Vitis germplasm collections. Am J Bot 105:215–226. https://doi.org/10.1002/ajb2.1033
Kofler N, Collins JP, Kuzma J, Marris E, Esvelt K et al (2018) Editing nature: local roots of global governance. Science 362:527–529. https://doi.org/10.1126/science.aat4612
Koleda I (1975) Ergebnisse von Kreuzungen zwischen Vitis amurensis und Vitis vinifera in der Züchtung frostwiderstandsfähiger Reben. VITIS J Grapevine Res 14:1–5
Kono A, Sato A, Nakano M, Yamada M, Mitani N et al (2012) Evaluating grapevine cultivars for resistance to anthracnose based on lesion number and length. Am J Enol Vitic 63:262–268. https://doi.org/10.5344/ajev.2012.11109
Kono A, Sato A, Ban Y, Mitani N (2013) Resistance of Vitis germplasm to Elsinoë ampelina (de Bary) Shear evaluated by lesion number and diameter. HortScience 48:1433–1439. https://doi.org/10.21273/hortsci.48.12.1433
Kono A, Ban Y, Mitani N, Fujii H, Sato S et al (2018) Development of SSR markers linked to QTL reducing leaf hair density and grapevine downy mildew resistance in Vitis vinifera. Mol Breed 38:138. https://doi.org/10.1007/s11032-018-0889-8
Koopman T, Linde CC, Fourie PH, Mcleod A (2007) Population genetic structure of Plasmopara viticola in the Western Cape province of South Africa. Mol Plant Pathol 8:723–736. https://doi.org/10.1111/j.1364-3703.2007.00429.x
Korbuly J (2000) Results of breeding for resistance to winter frosts and different pathogens using Vitis amurensis. Acta Hortic 528:551–557. https://doi.org/10.17660/ActaHortic.2000.528.80
Kortekamp A, Wind R, Zyprian E (1998) Investigation of the interaction of Plasmopara viticola with susceptible and resistant grapevine cultivars. J Plant Dis Prot 105:475–488. https://www.jstor.org/stable/43386544
Krivanek AF, Riaz S, Walker MA (2006) Identification and molecular mapping of PdR1 a primary resistance gene to Pierce’s disease in Vitis. Theor Appl Genet 112:1125–1131. https://doi.org/10.1007/s00122-006-0214-5
Kruger DHM, Fourie JC, Malan AP (2015) The effect of cover crops and their management on plant-parasitic nematodes in vineyards. South African J Enol Vitic 36(2):195–209
Kuczmog A, Galambos A, Horváth S, Mátai A, Kozma P et al (2012) Mapping of crown gall resistance locus Rcg1 in grapevine. Theor Appl Genet 125:1565–1574. https://doi.org/10.1007/s00122-012-1935-2
Kui L, Tang M, Duan S, Wang S, Dong X (2020) Identification of selective sweeps in the domesticated table and wine grape (Vitis vinifera L.). Front Plant Sci 11:e00572. https://doi.org/10.3389/fpls.2020.00572
Kulakiotu EK, Thanassoulopoulos CC, Sfakiotakis EM (2004) Postharvest biological control of Botrytis cinerea on kiwifruit by volatiles of “Isabella” grapes. Phytopathology 94:1280–1285. https://doi.org/10.1094/PHYTO.2004.94.12.1280
Kumar J, Pratap A, Kumar S (2015) Plant phenomics: an overview. In: Kumar J, Pratap A, Kumar S (eds) Phenomics in crop plants: trends, options and limitations. Springer, New Delhi, India. https://doi.org/10.1007/978-81-322-2226-2_1
Kunde RM, Lider LA, Schmitt RV (1968) A test of Vitis resistance to Xiphinema index. Am J Enol Vitic 19:30–36
Kuzmanović N, Biondi E, Overmann J, Puławska J, Verbarg S et al (2020) Revisiting the taxonomy of Allorhizobium vitis (i.e. Agrobacterium vitis) using genomics - emended description of all vitis sensu stricto and description of Allorhizobium ampelinum sp. nov. bioRxiv. https://doi.org/10.1101/2020.12.19.423612
Kyrkou I, Pusa T, Ellegaard-Jensen L, Sagot MF, Hansen LH (2018) Pierce’s disease of grapevines: a review of control strategies and an outline of an epidemiological model. Front Microbiol 9:1–23. https://doi.org/10.3389/fmicb.2018.02141
Lacombe T, Boursiquot J-M, Laucou V, Di Vecchi-Staraz M, Péros J-P et al (2013) Large-scale parentage analysis in an extended set of grapevine cultivars (Vitis vinifera L.). Theor Appl Genet 126:401–414. https://doi.org/10.1007/s00122-012-1988-2
Lai G, Fu P, Liu Y, Xiang J, Lu J (2018) Molecular characterization and overexpression of VpRPW8s from Vitis pseudoreticulata enhances resistance to Phytophthora capsici in Nicotiana benthamiana. Int J Mol Sci 19(3):839. https://doi.org/10.3390/ijms19030839
Laimer M, Lemaire O, Herrbach E, Goldschmidt V, Minafra A et al (2009) Resistance to viruses, phytoplasmas and their vectors in the grapevine in Europe: a review. J Plant Pathol 91:7–23. https://doi.org/10.4454/jpp.v91i1.620
Lämke J, Bäurle I (2017) Epigenetic and chromatin-based mechanisms in environmental stress adaptation and stress memory in plants. Genome Biol 18:124. https://doi.org/10.1186/s13059-017-1263-6
Landi L, Murolo S, Romanazzi G (2012) Colonization of Vitis spp. wood by sGFP-transformed Phaeomoniella chlamydospora, a tracheomycotic fungus involved in esca disease. Phytopathology 102:290–297. https://doi.org/10.1094/PHYTO-06-11-0165
Laucou V, Lacombe T, Dechesne F, Siret R, Bruno J-PP et al (2011) High throughput analysis of grape genetic diversity as a tool for germplasm collection management. Theor Appl Genet 122:1233–1245. https://doi.org/10.1007/s00122-010-1527-y
Laucou V, Launay A, Bacilieri R, Lacombe T, Adam-Blondon A-F et al (2018) Extended diversity analysis of cultivated grapevine Vitis vinifera with 10K genome-wide SNPs. PLoS ONE 13:27. https://doi.org/10.1371/journal.pone.0192540
Lawo NC, Weingart GJF, Schuhmacher R, Forneck A (2011) The volatile metabolome of grapevine roots: first insights into the metabolic response upon phylloxera attack. Plant Physiol Biochem 49:1059–1063. https://doi.org/10.1016/j.plaphy.2011.06.008
Lawrence DP, Travadon R, Baumgartner K (2015) Diversity of Diaporthe species associated with wood cankers of fruit and nut crops in Northern California. Mycologia 107:926–940. https://doi.org/10.3852/14-353
Le Cunff L, Fournier-Level A, Laucou V, Vezzulli S, Lacombe T et al (2008) Construction of nested genetic core collections to optimize the exploitation of natural diversity in Vitis vinifera L. subsp. sativa. BMC Plant Biol 8:31. https://doi.org/10.1186/1471-2229-8-31
Le Paslier M-C, Choisne N, Bacilieri R, Bounon R, Boursiquot J-M et al (2013) The GrapeReSeq 18k Vitis genotyping chip. In: 9th international symposium grapevine physiology and biotechnology. International Society for Horticultural Science, La Serena Chile, p 123
Lecomte P, Darrieutort G, Liminana JM, Comont G, Muruamendiaraz A et al (2012) New insights into esca of grapevine: the development of foliar symptoms and their association with xylem discoloration. Plant Dis 96:924–934. https://doi.org/10.1094/PDIS-09-11-0776-RE
Lee JC, Wang X, Daane KM, Hoelmer KA, Isaacs R et al (2019) Biological control of spotted-wing Drosophila (Diptera: Drosophilidae)-current and pending tactics. J Integr Pest Manag 10(1):13. https://doi.org/10.1093/jipm/pmz012
Lesuthu P, Mostert L, Spies CFJ, Moyo P, Regnier T et al (2019) Diaporthe nebulae sp. nov. and first report of D. cynaroidis, D. novem, and D. serafiniae on grapevines in South Africa. Plant Dis 103:808–817. https://doi.org/10.1094/PDIS-03-18-0433-RE
Li H (1993) Studies on the resistance of grapevine to powdery mildew. Plant Pathol 42:792–796. https://doi.org/10.1111/j.1365-3059.1993.tb01566.x
Li H-Y, Yang G-D, Shu H-R, Yang Y-T, Ye B-X et al (2006) Colonization by the arbuscular mycorrhizal fungus Glomus versiforme induces a defense response against the root-knot nematode Meloidogyne incognita in the grapevine (Vitis amurensis Rupr.), which includes transcriptional activation of the class III Chitin. Plant Cell Physiol 47:154–163. https://doi.org/10.1093/pcp/pci231
Li D, Wan Y, Wang Y, He P (2008) Relatedness of resistance to anthracnose and to white rot in Chinese wild grapes. VITIS J Grapevine Res 47:213–215. https://doi.org/10.5073/vitis.2008.47.213-215
Li D (2014) The history of Chinese winegrowing and winemaking-part 2. Decant China
Li X, Yin L, Ma L, Zhang Y, An Y et al (2016) Pathogenicity variation and population genetic structure of Plasmopara viticola in China. J Phytopathol 164:863–873. https://doi.org/10.1111/jph.12505
Li Z, Fan Y, Chang P, Gao L, Wang X (2020) Genome sequence resource for Elsinoë ampelina, the causal organism of grapevine anthracnose. Mol Plant-Microbe Interact 33:576–579. https://doi.org/10.1094/MPMI-12-19-0337-A
Liang ZC, Duan SC, Sheng J, Zhu SS, Ni XM et al (2019) Whole-genome resequencing of 472 Vitis accessions for grapevine diversity and demographic history analyses. Nat Commun 10:1190. https://doi.org/10.1038/s41467-019-09135-8
Lijavetzky D, Cabezas J, Ibáñez A, Rodríguez V, Martínez-Zapater JM (2007) High throughput SNP discovery and genotyping in grapevine (Vitis vinifera L.) by combining a re-sequencing approach and SNPlex technology. BMC Genomics 8:424. https://doi.org/10.1186/1471-2164-8-424
Likert R (1932) A technique for the measurement of attitudes. Arch Psychol 140:44–53
Limera C, Sabbadini S, Sweet JB, Mezzetti B (2017) New biotechnological tools for the genetic improvement of major woody fruit species. Front Plant Sci 8:1–16. https://doi.org/10.3389/fpls.2017.01418
Lin H, Islam MS, Morano L, Groves R, Bextine B et al (2013) Genetic variation of Xylella fastidiosa associated with grapevines in two major viticultural regions in the United States: California and Texas. J Plant Pathol 95:329–337. https://doi.org/10.4454/JPP.V95I2.028
Lin H (2017) Genetic analysis of Pierce’s disease resistant progeny of N18-6 x Flame seedless grapevine breeding population. In: Research progress reports: Pierce’s disease and other designated pests and diseases of winegrapes. California Department of Food and Agriculture, Sacramento, CA, p 148
Lin H, Leng H, Guo Y, Kondo S, Zhao Y et al (2019) QTLs and candidate genes for downy mildew resistance conferred by interspecific grape (V. vinifera L. × V. amurensis Rupr.) crossing. Sci Hortic 244:200–207. https://doi.org/10.1016/j.scienta.2018.09.045
Lindow S, Newman K, Chatterjee S, Baccari C, Lavarone AT et al (2014) Production of Xylella fastidiosa diffusible signal factor in transgenic grape causes pathogen confusion and reduction in severity of Pierce’s disease. Mol Plant-Microbe Interact 27:244–254. https://doi.org/10.1094/MPMI-07-13-0197-FI
Lipka AE, Tian F, Wang Q, Peiffer J, Li M et al (2012) GAPIT: genome association and prediction integrated tool. Bioinformatics 28:2397–2399. https://doi.org/10.1093/bioinformatics/bts444
Liu X-Q, Ickert-Bond SM, Nie Z-L, Zhou Z, Chen L-Q et al (2016a) Phylogeny of the Ampelocissus-Vitis clade in Vitaceae supports the New World origin of the grape genus. Mol Phylogenet Evol 95:217–228. https://doi.org/10.1016/j.ympev.2015.10.013
Liu X, Huang M, Fan B, Buckler ES, Zhang Z (2016b) Iterative usage of fixed and random effect models for powerful and efficient genome-wide association studies. PLOS Genet 12:e1005767. https://doi.org/10.1371/journal.pgen.1005767
Liu J, Yang C, Shi X, Li C, Huang J et al (2016c) Analyzing association mapping in pedigree-based GWAS using a penalized multitrait mixed model. Genet Epidemiol 40:382–393. https://doi.org/10.1002/gepi.21975
Liu S, Zhang C, Chao N, Lu J, Zhang Y (2018) Cloning, characterization, and functional investigation of VaHAESA from Vitis amurensis inoculated with Plasmopara viticola. Int J Mol Sci 19(4):1204. https://doi.org/10.3390/ijms19041204
Liu M, Ma F, Wu F, Jiang C, Wang Y (2019) Expression of stilbene synthase VqSTS6 from wild Chinese Vitis quinquangularis in grapevine enhances resveratrol production and powdery mildew resistance. Planta 250:1997–2007. https://doi.org/10.1007/s00425-019-03276-2
Loose S, Pabst E (2019) ProWein business report: “Climate Change”
Loose S (2020) ProWein business report: Covid-19 is the biggest challenge for the global wine industry
Loschiavo A, Sosnowski M, Wicks T (2007) Incidence of eutypa dieback in the Adelaide Hills. Aust New Zeal Grapegrow Winemak 08:26–29
Louime C, Lu J, Onokpise O, Vasanthaiah HKN, Kambiranda D et al (2011) Resistance to Elsinoë ampelina and expression of related resistant genes in Vitis rotundifolia Michx. grapes. Int J Mol Sci 12:3473–3488. https://doi.org/10.3390/ijms12063473
Lovato A, Zenoni S, Tornielli GB, Colombo T, Vandelle E et al (2019) Plant and fungus transcriptomic data from grapevine berries undergoing artificially-induced noble rot caused by Botrytis cinerea. Data Br 25:104150. https://doi.org/10.1016/j.dib.2019.104150
Lu J, Liu C (2015) Grapevine breeding in China. In: Reynolds A (ed) Grapevine breeding programs for the wine industry. Elsevier, Woodhead Publishing, Sawston, UK, pp 273–310. https://doi.org/10.1016/B978-1-78242-075-0.00012-0
Lu W, Newlands NK, Carisse O, Atkinson DE, Cannon AJ (2020) Disease risk forecasting with Bayesian learning networks: application to grape powdery mildew (Erysiphe necator) in vineyards. Agronomy 10:1–29. https://doi.org/10.3390/agronomy10050622
Lü P, Yu S, Zhu N, Chen YR, Zhou B et al (2018) Genome encode analyses reveal the basis of convergent evolution of fleshy fruit ripening. Nat Plants 4:784–791. https://doi.org/10.1038/s41477-018-0249-z
Lund K, Riaz S, Walker MA (2017) Population structure, diversity and reproductive mode of the grape phylloxera (Daktulosphaira vitifoliae) across its native range. PLoS ONE 12:1–21. https://doi.org/10.1371/journal.pone.0170678
Luttrell ES (1946) Black rot of muscadine grapes. Phytopathology 36:905–926
Luttrell ES (1948) Physiologic specialization in Guignardia bidwellii, cause of black rot of Vitis and Parthenocissus species. Phytopathology 38:716–723
Ma H, Xiang G, Li Z, Wang Y, Dou M et al (2018) Grapevine VpPR10.1 functions in resistance to Plasmopara viticola through triggering a cell death-like defence response by interacting with VpVDAC3. Plant Biotechnol J 16(8):1488–1501. https://doi.org/10.1111/pbi.12891
Maddalena G, Delmotte F, Bianco PA, De Lorenzis G, Toffolatti SL (2020) Genetic structure of Italian population of the grapevine downy mildew agent, Plasmopara viticola. Ann Appl Biol 176:257–267. https://doi.org/10.1111/aab.12567
Magris G, Di Gaspero G, Marroni F, Zenoni S, Tornielli GB et al (2019) Genetic, epigenetic and genomic effects on variation of gene expression among grape varieties. Plant J 99:895–909. https://doi.org/10.1111/tpj.14370
Mahanil S, Ramming D, Cadle-Davidson M, Owens C, Garris A et al (2012) Development of marker sets useful in the early selection of Ren4 powdery mildew resistance and seedlessness for table and raisin grape breeding. Theor Appl Genet 124:23–33. https://doi.org/10.1007/s00122-011-1684-7
Maia M, Ferreira AEN, Nascimento R, Monteiro F, Traquete F et al (2020) Integrating metabolomics and targeted gene expression to uncover potential biomarkers of fungal/oomycetes-associated disease susceptibility in grapevine. Sci Rep 10:1–15. https://doi.org/10.1038/s41598-020-72781-2
Malagnini V, de Lillo E, Saldarelli P, Beber R, Duso C et al (2016) Transmission of Grapevine Pinot gris Virus by Colomerus vitis (Acari: Eriophyidae) to grapevine. Arch Virol 161:2595–2599. https://doi.org/10.1007/s00705-016-2935-3
Malnoy M, Viola R, Jung MH, Koo OJ, Kim S et al (2016) DNA-free genetically edited grapevine and apple protoplast using CRISPR/Cas9 ribonucleoproteins. Front Plant Sci 7:1–9. https://doi.org/10.3389/fpls.2016.01904
Manawasinghe IS, Dissanayake AJ, Li X, Liu M, Wanasinghe DN et al (2019) High genetic diversity and species complexity of diaporthe associated with grapevine dieback in China. Front Microbiol 10:1–28. https://doi.org/10.3389/fmicb.2019.01936
Mansour R, Belzunces LP, Suma P, Zappalà L, Mazzeo G et al (2018) Vine and citrus mealybug pest control based on synthetic chemicals. A review. Agron Sustain Dev 38:1–20. https://doi.org/10.1007/s13593-018-0513-7
Maraš V, Tello J, Gazivoda A, Mugoša M, Perišić M et al (2020) Population genetic analysis in old Montenegrin vineyards reveals ancient ways currently active to generate diversity in Vitis vinifera. Sci Rep 10:15000. https://doi.org/10.1038/s41598-020-71918-7
Marchi G (2001) Susceptibility to esca of various grapevine (Vitis vinifera) cultivars grafted on different rootstocks in a vineyard in the province of Siena (Italy). Phytopathol Mediterr 40:27–36. https://doi.org/10.14601/Phytopathol_Mediterr-1589
Marguerit E, Boury C, Manicki A, Donnart M, Butterlin G et al (2009) Genetic dissection of sex determinism, inflorescence morphology and downy mildew resistance in grapevine. Theor Appl Genet 118:1261–1278. https://doi.org/10.1007/s00122-009-0979-4
Marrano A, Birolo G, Prazzoli ML, Lorenzi S, Valle G et al (2017) SNP-discovery by RAD-sequencing in a germplasm collection of wild and cultivated grapevines (V. vinifera L.). PLoS ONE 12:19. https://doi.org/10.1371/journal.pone.0170655
Martelli GP, Taylor CE (1990) Distribution of viruses and their nematode vectors. In: Harris KF (ed) Advances in disease vector research, vol 6. Springer, New York, NY, pp 151–189. https://doi.org/10.1007/978-1-4612-3292-6_6
Martelli GP (1997) Infectious diseases and certification of grapevines. In: Martelli GP, Digiaro M (eds) Mediterranean network on grapevine Closteroviruses 1992–1997 and the viroses and virus-like diseases of the grapevine a bibliographic report, 1985–1997. Options Méditerranéennes Série B. Etudes et Recherches n. 29. CHIEAM, Bari, Italy, pp 47–64
Martelli GP, Abou Ghanem-Sabanadzovic N, Agranovsky AA, Al Rwahnih M, Dolja VV et al (2012) Taxonomic revision of the family closteroviridae with special reference to the grapevine leafroll-associated members of the genus Ampelovirus and the putative species unassigned to the family. J Plant Pathol 94:7–19. https://doi.org/10.4454/jpp.fa.2012.022
Martelli GP (2014) Directory of virus and virus-like diseases of the grapevine and their agents. J Plant Pathol 96:1–4
Martelli GP (2017) An overview on grapevine viruses, viroids, and the diseases they cause. In: Meng B, Martelli GP, Golino DA, Fuchs M (eds) Grapevine viruses: molecular biology, diagnostics and management. Springer, Cham, Switzerland, pp 31–46. https://doi.org/10.1007/978-3-319-57706-7_2
Martín L, Sáenz de Miera LE, Martín MT (2014) AFLP and RAPD characterization of Phaeoacremonium aleophilum associated with Vitis vinifera decline in Spain. J Phytopathol 162:245–257. https://doi.org/10.1111/jph.12180
Martinelli L, Gribaudo I (2001) Somatic embryogenesis in Grapevine. In: Molecular biology & biotechnology of the grapevine. Springer, Dordrecht, Netherlands, pp 327–351. https://doi.org/10.1007/978-94-017-2308-4_13
Martinez MC, Perez JE (1995) “Catalan Blanco”, “Catalan Roxo”, “Folla Redonda”: Vitis vinifera L. o hibridos productores directo? Investig Agrar: Prod Prot Veg 10:5–14
Martínez-Diz M del P, Díaz-Losada E, Barajas E, Ruano-Rosa D, Andrés-Sodupe M et al (2019) Screening of Spanish Vitis vinifera germplasm for resistance to Phaeomoniella chlamydospora. Sci Hortic 246:104–109. https://doi.org/10.1016/j.scienta.2018.10.049
Martínez-Diz MP, Díaz-Losada E, Díaz-Fernández Á, Bouzas-Cid Y, Gramaje D (2021a) Protection of grapevine pruning wounds against Phaeomoniella chlamydospora and Diplodia seriata by commercial biological and chemical methods. Crop Prot 143:125. https://doi.org/10.1016/j.cropro.2020.105465
Martínez-Diz MP, Díaz-Losada E, Andrés-Sodupe M, Bujanda R, Maldonado-González MM et al (2021b) Field evaluation of biocontrol agents against black-foot and Petri diseases of grapevine. Pest Manag Sci 77:697–708. https://doi.org/10.1002/ps.6064
Matsoukas IG (2020) Prime editing: genome editing for rare genetic diseases without double-strand breaks or donor DNA. Front Genet 11:1–6. https://doi.org/10.3389/fgene.2020.00528
Maul E, Sudharma KN, Kecke S, Marx G, Müller C et al (2012) The European *Vitis* database (www.eu-vitis-de)-a technical innovation through an online uploading and interactive modification system. VITIS J Grapevine Res 51:79–85. https://doi.org/10.5073/vitis.2012.51.79-85
Maul E et al (2021) Vitis International Variety Catalogue. www.vivc.de
Mazzoni V, Ioriatti C, Trona F, Lucchi A, De Cristofaro A et al (2009a) Study on the role of olfaction in host plant detection of Scaphoideus titanus (Hemiptera: Cicadellidae) Nymphs. J Econ Entomol 102:974–980. https://doi.org/10.1603/029.102.0316
Mazzoni V, Prešern J, Lucchi A, Virant-Doberlet M (2009b) Reproductive strategy of the nearctic leafhopper Scaphoideus titanus Ball (Hemiptera: Cicadellidae). Bull Entomol Res 99:401–413. https://doi.org/10.1017/S0007485308006408
Mazzoni V, Nieri R, Eriksson A, Virant-Doberlet M, Polajnar J et al (2019) Mating disruption by vibrational signals: state of the field and perspectives. In: Hill PSM, Lakes-Harlan R, Mazzoni V, Narins PM, Virant-Doberlet M et al (eds) Biotremology: studying vibrational behavior, vol 6. Springer, Cham, Switzerland, pp 331–354. https://doi.org/10.1007/978-3-030-22293-2_17
McDonald BA, Linde C (2002) Pathogen population genetics, evolutionary potential, and durable resistance. Annu Rev Phytopathol 40:349–379. https://doi.org/10.1146/annurev.phyto.40.120501.101443
Melnyk CW, Molnar A, Baulcombe DC (2011) Intercellular and systemic movement of RNA silencing signals. EMBO J 30:3553–3563. https://doi.org/10.1038/emboj.2011.274
Meloni G, Swinnen J (2013) The political economy of European wine regulations. J Wine Econ 8:244–284. https://doi.org/10.1017/jwe.2013.33
Meng B, Martelli GP, Golino DA, Fuchs M (2017) Grapevine viruses: molecular biology, diagnostics and management. Springer, Cham, Switzerland https://doi.org/10.1007/978-3-319-57706-7
Mercier A, Carpentier F, Duplaix C, Auger A, Pradier JM et al (2019) The polyphagous plant pathogenic fungus Botrytis cinerea encompasses host-specialized and generalist populations. Environ Microbiol 21:4808–4821. https://doi.org/10.1111/1462-2920.14829
Merdinoglu D, Wiedeman-Merdinoglu S, Coste P, Dumas V, Haetty S et al (2003) Genetic analysis of downy mildew resistance derived from Muscadinia rotundifolia. Acta Hortic 603_57:451–456. https://doi.org/10.17660/ActaHortic.2003.603.57
Merdinoglu D, Butterlin G, Bevilacqua L, Chiquet V, Adam-Blondon A-F et al (2005) Development and characterization of a large set of microsatellite markers in grapevine (Vitis vinifera L.) suitable for multiplex PCR. Mol Breed 15:349–366. https://doi.org/10.1007/s11032-004-7651-0
Merdinoglu D, Schneider C, Prado E, Wiedemann-Merdinoglu S, Mestre P (2018) Breeding for durable resistance to downy and powdery mildew in grapevine. OENO One 52:203–209. https://doi.org/10.20870/oeno-one.2018.52.3.2116
Migicovsky Z, Sawler J, Money D, Eibach R, Miller AJ et al (2016) Genomic ancestry estimation quantifies use of wild species in grape breeding. BMC Genomics 17:478. https://doi.org/10.1186/s12864-016-2834-8
Migliaro D, De Lorenzis G, Di Lorenzo GS, De Nardi B, Gardiman M et al (2019) Grapevine non-vinifera genetic diversity assessed by simple sequence repeat markers as a starting point for new rootstock breeding programs. Am J Enol Vitic 70:390–397. https://doi.org/10.5344/ajev.2019.18054
Miller AJ, Matasci N, Schwaninger H, Aradhya MK, Prins B et al (2013) Vitis phylogenomics: hybridization intensities from a SNP array outperform genotype calls. PLoS ONE 8:e78680. https://doi.org/10.1371/journal.pone.0078680
Minio A, Lin J, Gaut BS, Cantu D (2017) How single molecule real-time sequencing and haplotype phasing have enabled reference-grade diploid genome assembly of wine grapes. Front Plant Sci 8. https://doi.org/10.3389/fpls.2017.00826
Minio A, Massonnet M, Figueroa-Balderas R, Vondras AM, Blanco-Ulate B et al (2019a) Iso-Seq allows genome-independent transcriptome profiling of grape berry development. G3 Genes|Genomes|Genetics 9:755–767. https://doi.org/10.1534/g3.118.201008
Minio A, Massonnet M, Figueroa-Balderas R, Castro A, Cantu D (2019b) Diploid genome assembly of the wine grape Carménère. G3 Genes|Genomes|Genetics 9:1331–1337. https://doi.org/10.1534/g3.119.400030
Mirica I (1988) Anthracnose. In: Pearson RC, Goheen AC (eds) Compendium of grape diseases. APS Press, St. Paul, MN, pp 18–19. https://doi.org/10.1094/9780890544815
Modesto LR, Steiner DRM, Menon JK, Nodari RO, Welter LJ et al (2020) Standard area diagram set for anthracnose severity on grapevine bunches and shoots. Australas Plant Pathol 49:561–569. https://doi.org/10.1007/s13313-020-00728-2
Molina A, Miedes E, Bacete L et al (2021) Arabidopsis cell wall composition determines disease resistance specificity and fitness. Proc Natl Acad Sci 118:e2010243118. https://doi.org/10.1073/pnas.2010243118
Molitor D, Beyer M (2014) Epidemiology, identification and disease management of grape black rot and potentially useful metabolites of black rot pathogens for industrial applications-a review. Ann Appl Biol 165:305–317. https://doi.org/10.1111/aab.12155
Moralejo E, Borràs D, Gomila M, Montesinos M, Adrover F et al (2019) Insights into the epidemiology of Pierce’s disease in vineyards of Mallorca, Spain. Plant Pathol 68:1458–1471. https://doi.org/10.1111/ppa.13076
Morales-Cruz A, Amrine KC, Blanco-Ulate B, Lawrence DP, Travadon R et al (2015) Distinctive expansion of gene families associated with plant cell wall degradation, secondary metabolism, and nutrient uptake in the genomes of grapevine trunk pathogens. BMC Genomics 16:469. https://doi.org/10.1186/s12864-015-1624-z
Morán F, Olmos A, Lotos L, Predajňa L, Katis N et al (2018) A novel specific duplex real-time RT-PCR method for absolute quantitation of Grapevine Pinot gris Virus in plant material and single mites. PLoS ONE 13:1–14. https://doi.org/10.1371/journal.pone.0197237
Moreira FM, Madini A, Marino R, Zulini L, Stefanini M et al (2011) Genetic linkage maps of two interspecific grape crosses (Vitis spp.) used to localize Quantitative Trait Loci for downy mildew resistance. Tree Genet Genomes 7:153–167. https://doi.org/10.1007/s11295-010-0322-x
Moretto M, Sonego P, Pilati S, Malacarne G, Costantini L et al (2016) VESPUCCI: exploring patterns of gene expression in grapevine. Front Plant Sci 7:633. https://doi.org/10.3389/fpls.2016.00633
Moroldo M, Paillard S, Marconi R, Fabrice L, Canaguier A et al (2008) A physical map of the heterozygous grapevine “Cabernet Sauvignon” allows mapping candidate genes for disease resistance. BMC Plant Biol 8:66. https://doi.org/10.1186/1471-2229-8-66
Mortensen JA (1981) Sources and inheritance of resistance to anthracnose in Vitis. J Hered 72:423–426. https://doi.org/10.1093/oxfordjournals.jhered.a109545
Mosa KA, Ismail A, Helmy M (2017) Omics and system biology approaches in plant stress research. In: Plant stress tolerance: an integrated omics approach. Springer, Cham, Switzerland, pp 21–34. https://doi.org/10.1007/978-3-319-59379-1_2
Mostert L, Crous PW, Kang J-C, Phillips AJL (2001) Species of Phomopsis and a Libertella sp. occurring on grapevines with specific reference to South Africa: morphological, cultural, molecular and pathological characterization. Mycologia 93:146–167. https://doi.org/10.1080/00275514.2001.12061286
Mugnai L, Graniti A, Surico G (1999) Esca (Black measles) and brown wood-streaking: Two old and elusive diseases of grapevines. Plant Dis 83:404–418. https://doi.org/10.1094/PDIS.1999.83.5.404
Mullins MG, Rajasekaran K (1981) Fruiting cuttings: revised method for producing test plants of grapevine cultivars. Am J Enol Vitic 23:35–40
Mundt CC (2014) Durable resistance: a key to sustainable management of pathogens and pests. Infect Genet Evol 27:446–455. https://doi.org/10.1016/j.meegid.2014.01.011
Mundt CC (2018) Pyramiding for resistance durability: theory and practice. Phytopathology 108:792–802. https://doi.org/10.1094/PHYTO-12-17-0426-RVW
Murolo S, Romanazzi G (2014) Effects of grapevine cultivar, rootstock and clone on esca disease. Australas Plant Pathol 43:215–221. https://doi.org/10.1007/s13313-014-0276-9
Myles S, Peiffer J, Brown PJ, Ersoz ES, Zhang Z et al (2009) Association mapping: critical considerations shift from genotyping to experimental design. Plant Cell 21:2194–2202. https://doi.org/10.1105/tpc.109.068437
Myles S, Chia J-M, Hurwitz B, Simon C, Zhong GY et al (2010) Rapid genomic characterization of the genus Vitis. PLoS ONE 5:e8219. https://doi.org/10.1371/journal.pone.0008219
Myles S, Boyko AR, Owens CL, Brown PJ, Grassi F et al (2011) Genetic structure and domestication history of the grape. Proc Natl Acad Sci USA 108:3457–3458. https://doi.org/10.1073/pnas.1009363108
Nabity PD, Haus MJ, Berenbaum MR, DeLucia EH (2013) Leaf-galling phylloxera on grapes reprograms host metabolism and morphology. Proc Natl Acad Sci USA 110:16663–16668. https://doi.org/10.1073/pnas.1220219110
Naegele RP (2018) Evaluation of host resistance to Botrytis bunch rot in Vitis spp. and its correlation with Botrytis leaf spot. HortScience 53:204–207. https://doi.org/10.21273/HORTSCI12655-17
Nakajima I, Ban Y, Azuma A, Onoue N, Moriguchi T et al (2017) CRISPR/Cas9-mediated targeted mutagenesis in grape. PLoS ONE 12:1–16. https://doi.org/10.1371/journal.pone.0177966
Namba S (2019) Molecular and biological properties of phytoplasmas. Proc Jpn Acad Ser B 95:401–418. https://doi.org/10.2183/pjab.95.028
Näpflin K, O’Connor EA, Becks L, Bensch S, Ellis VA et al (2019) Genomics of host-pathogen interactions: challenges and opportunities across ecological and spatiotemporal scales. PeerJ 7:e8013. https://doi.org/10.7717/peerj.8013
Narduzzi-Wicht B, Jermini M, Gessler C, Broggini GAL (2014) Microsatellite markers for population studies of the ascomycete Phyllosticta ampelicida, the pathogen causing grape black rot. Phytopathol Mediterr 53:470–479. https://doi.org/10.14601/phytopathol_mediterr-14481
Nascimento-Gavioli MCA, Rockenbach MF, Welter LJ, Guerra MP (2020) Histopathological study of resistant (Vitis labrusca L.) and susceptible (Vitis vinifera L.) cultivars of grapevine to the infection by downy mildew. J Hortic Sci Biotechnol 95:521–531. https://doi.org/10.1080/14620316.2019.1685411
Nawaz MA, Huang Y, Bie Z, Ahmed W, Reiter RJ et al (2016) Melatonin: current status and future perspectives in plant science. Front Plant Sci 6:1–13. https://doi.org/10.3389/fpls.2015.01230
Negrul AM (1946) Origin of cultivated grapevine and its classification. In: Frolov-Bagreev AM (ed) Ampelography of the Soviet Union, Moscow, Pishchepromizdat, pp 159–216
Negrel L, Halter D, Wiedemann-Merdinoglu S, Rustenholz C, Merdinoglu D et al (2018) Identification of lipid markers of Plasmopara viticola infection in grapevine using a non-targeted metabolomic approach. Front Plant Sci 9:360. https://doi.org/10.3389/fpls.2018.00360
Nguyen VC, Villate L, Gutierrez-Gutierrez C, Castillo P, Van Ghelder C et al (2019) Phylogeography of the soil-borne vector nematode Xiphinema index highly suggests Eastern origin and dissemination with domesticated grapevine. Sci Rep 9:1–11. https://doi.org/10.1038/s41598-019-43812-4
Nguyen VC, Tandonnet J-P, Khallouk S, Van Ghelder C, Portier U et al (2020) Grapevine resistance to the nematode Xiphinema index is durable in muscadine-derived plants obtained from hardwood cuttings but not from in vitro. Phytopathology 110:1565–1571. https://doi.org/10.1094/PHYTO-01-20-0008-R
Nicol JM, Stirling GR, Rose BJ, May P, Heeswijck R (1999) Impact of nematodes on grapevine growth and productivity: current knowledge and future directions, with special reference to Australian viticulture. Aust J Grape Wine Res 5:109–127. https://doi.org/10.1111/j.1755-0238.1999.tb00295.x
Nicolas SD, Péros J-P, Lacombe T, Launay A, Le Paslier M-C et al (2016) Genetic diversity, linkage disequilibrium and power of a large grapevine (Vitis vinifera L) diversity panel newly designed for association studies. BMC Plant Biol 16:74. https://doi.org/10.1186/s12870-016-0754-z
Niederhuth CE, Bewick AJ, Ji L, Alabady MS, Do KK et al (2016) Widespread natural variation of DNA methylation within angiosperms. Genome Biol 17:194. https://doi.org/10.1186/s13059-016-1059-0
Nieri R, Anfora G, Mazzoni V, Rossi Stacconi MV (2021) Semiochemicals, semiophysicals and their integration for the development of innovative multi-modal systems for agricultural pests’ monitoring and control. Entomol Gen. https://doi.org/10.1127/entomologia/2021/1236
Niks RE, Rubiales D (2002) Potentially durable resistance mechanisms in plants to specialised fungal pathogens. Euphytica 124:201–216. https://doi.org/10.1023/A:1015634617334
Nita M, Ellis MA, Wilson LL, Madden LV (2006) Evaluation of a disease warning system for Phomopsis cane and leaf spot of grape: a field study. Plant Dis 90:1239–1246. https://doi.org/10.1094/PD-90-1239
Ocarez N, Jiménez N, Núñez R, Perniola R, Marsico AD et al (2020) Unraveling the deep genetic architecture for seedlessness in grapevine and the development and validation of a new set of markers for VviAGL11- based gene-assisted selection. Genes (basel) 11:151. https://doi.org/10.3390/genes11020151
Ochssner I, Hausmann L, Töpfer R (2016) Rpv14, a new genetic source for Plasmopara viticola resistance conferred by Vitis cinerea. VITIS J Grapevine Res 55:79–81. https://doi.org/10.5073/vitis.2016.55.79-81
OIV (2009) 2nd edition of the OIV Descriptor list for grape varieties and Vitis species
OIV (2017) Focus OIV 2017 Distribution of the world’s grapevine varieties. OIV-International organization of vine and wine, Paris, France
OIV (2020) 2020 Wine prodution-OIV First Estimates. Int Organ Vine Wine, pp 1–8
OIV-MA-AS315-03; 377/2009 (2009) Malvidin diglucoside. In: Compendium of international methods of analysis. https://www.oiv.int/public/medias/2530/oiv-ma-as315-03.pdf
OIV-VITI 1-1991 (1991) Clonal Selection. In: Resolution OIV-VITI 1-1991
OIV-VITI 564A-2017 (2017) OIV Process for the clonal selection of vines
OIV-VITI 564B-2019 OIV process for the recovery and conservation of the intravarietal diversity and the polyclonal selection of the vine in grape varieties with wide genetic variability
Oliver JE, Fuchs M (2011) Tolerance and resistance to viruses and their vectors in Vitis sp.: a virologist’s perspective of the literature. Am J Enol Vitic 62:438–451. https://doi.org/10.5344/ajev.2011.11036
Ollat N, Claverie M, Esmenjaud D, Demangeat G, Jacquet O (2011) Dossier Vigne: Un porte-greffe pour lutter contre le court-noué. Phytoma-La Défense Des Végétaux 649:29–33
Ollat N, Peccoux A, Papura D, Esmenjaud D, Marguerit E et al (2016) Rootstocks as a component of adaptation to environment. In: Gerós H, Chaves MM, Gil HM, Delrot S (eds) Grapevine in a changing environment. John Wiley & Sons, Ltd., Hoboken, NJ, pp 68–108. https://doi.org/10.1002/9781118735985.ch4
Olmo HP (1976) Grapes. In: Simmonds NW (ed) Evolution of crop plants. Longman, London, UK, pp 294–298
Ometto L, Cestaro A, Ramasamy S, Grassi A, Revadi S et al (2013) Linking genomics and ecology to investigate the complex evolution of an invasive Drosophila pest. Genome Biol Evol 5:745–757. https://doi.org/10.1093/gbe/evt034
Onesti G, González-Domínguez E, Rossi V (2016) Accurate prediction of black rot epidemics in vineyards using a weather-driven disease model. Pest Manag Sci 72:2321–2329. https://doi.org/10.1002/ps.4277
Ormeño-Orrillo E, Servín-Garcidueñas LE, Rogel MA, González V, Peralta H et al (2015) Taxonomy of rhizobia and Agrobacteria from the Rhizobiaceae family in light of genomics. Syst Appl Microbiol 38:287–291. https://doi.org/10.1016/j.syapm.2014.12.002
Osakabe Y, Liang Z, Ren C, Nishitani C, Osakabe K et al (2018) CRISPR–Cas9-mediated genome editing in apple and grapevine. Nat Protoc 13:2844–2863. https://doi.org/10.1038/s41596-018-0067-9
Ouantar M, Anfora G, Bouharoud R, Chebli B (2020) First report of Drosophila suzukii (Diptera: Drosophiladae) in North Africa. Moroccan J Agric Sci 1:277–279
Pacifico D, Gaiotti F, Giusti M, Tomasi D (2013) Performance of interspecific grapevine varieties in North-East Italy. Agric Sci 04:91–101. https://doi.org/10.4236/as.2013.42015
Palmieri MC, Perazzolli M, Matafora V, Moretto M, Bachi A et al (2012) Proteomic analysis of grapevine resistance induced by Trichoderma harzianum T39 reveals specific defence pathways activated against downy mildew. J Exp Bot 63:6237–6251. https://doi.org/10.1093/jxb/ers279
Pap D, Riaz S, Dry IB, Jermakow A, Tenscher AC et al (2016) Identification of two novel powdery mildew resistance loci, Ren6 and Ren7, from the wild Chinese grape species Vitis piasezkii. BMC Plant Biol 16:1–19. https://doi.org/10.1186/s12870-016-0855-8
Papura D, Burban C, van Helden M, Giresse X, Nusillard B et al (2012) Microsatellite and mitochondrial data provide evidence for a single major introduction for the neartic leafhopper Scaphoideus titanus in Europe. PLoS ONE 7:e36882. https://doi.org/10.1371/journal.pone.0036882
Paris M, Boyer R, Jaenichen R, Wolf J, Karageorgi M et al (2020) Near-chromosome level genome assembly of the fruit pest Drosophila suzukii using long-read sequencing. Sci Rep 10:11227. https://doi.org/10.1038/s41598-020-67373-z
Patel S, Robben M, Fennell A, Londo JP, Alahakoon D et al (2020) Draft genome of the Native American cold hardy grapevine Vitis riparia Michx. ‘Manitoba 37.’ Hortic Res 7:92. https://doi.org/10.1038/s41438-020-0316-2
Patil SG, Honrao BK, Karkamar S (1998) Reaction of grape germplasm against rust diseases. J Maharashtra Agric Univ 23:138–140
Pauquet J, Bouquet A, This P, Adam-Blondon AF (2001) Establishment of a local map of AFLP markers around the powdery mildew resistance gene Run1 in grapevine and assessment of their usefulness for marker assisted selection. Theor Appl Genet 103:1201–1210. https://doi.org/10.1007/s001220100664
Pavan S, Jacobsen E, Visser RGF, Bai Y (2010) Loss of susceptibility as a novel breeding strategy for durable and broad-spectrum resistance. Mol Breed 25:1–12. https://doi.org/10.1007/s11032-009-9323-6
Pavan S, Delvento C, Ricciardi L, Lotti C, Ciani E et al (2020) Recommendations for choosing the genotyping method and best practices for quality control in crop genome-wide association studies. Front Genet 11:447. https://doi.org/10.3389/fgene.2020.00447
Pavloušek P (2012) Evaluation of foliar resistance of grapevine genetic resources to downy mildew (Plasmopara viticola). Acta Univ Agric Silvic Mendelianae Brun 60:191–198. https://doi.org/10.11118/actaun201260080191
Peleman JD, Sørensen AP, van der Voort JR (2005) Breeding by design: exploiting genetic maps and molecular markers through marker-assisted selection. In: Meksem K, Kahl G (eds) The handbook of plant genome mapping. Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, FRG, pp 109–129. https://doi.org/10.1002/3527603514.ch5
Pennington T, Kraus C, Alakina E, Entling M, Hoffmann C (2017) Minimal pruning and reduced plant protection promote predatory mites in Grapevine. Insects 8:86. https://doi.org/10.3390/insects8030086
Perazzolli M, Palmieri MC, Matafora V, Bachi A, Pertot I (2016) Phosphoproteomic analysis of induced resistance reveals activation of signal transduction processes by beneficial and pathogenic interaction in grapevine. J Plant Physiol 195:59–72. https://doi.org/10.1016/j.jplph.2016.03.007
Peressotti E, Wiedemann-Merdinoglu S, Delmotte F, Bellin D, Di Gaspero G et al (2010) Breakdown of resistance to grapevine downy mildew upon limited deployment of a resistant variety. BMC Plant Biol 10:147. https://doi.org/10.1186/1471-2229-10-147
Peros JP, Berger G, Portemont A, Boursiquot JM, Lacombe T (2011) Genetic variation and biogeography of the disjunct Vitis subg. Vitis (Vitaceae). J Biogeogr 38:471–486. https://doi.org/10.1111/j.1365-2699.2010.02410.x
Péros JP, Cousins P, Launay A, Cubry P, Walker A et al (2021) Genetic diversity and population structure in Vitis species illustrate phylogeographic patterns in Eastern North America. Mol Ecol 30:2333–2348. https://doi.org/10.1111/mec.15881
Pertot I, Caffi T, Rossi V, Mugnai L, Hoffmann C et al (2017) A critical review of plant protection tools for reducing pesticide use on grapevine and new perspectives for the implementation of IPM in viticulture. Crop Prot 97:70–84. https://doi.org/10.1016/j.cropro.2016.11.025
Pertot I, Prodorutti D, Colombini A, Pasini L (2016) Trichoderma atroviride SC1 prevents Phaeomoniella chlamydospora and Phaeoacremonium aleophilum infection of grapevine plants during the grafting process in nurseries. Biocontrol 61:257–267. https://doi.org/10.1007/s10526-016-9723-6
Pessina S, Lenzi L, Perazzolli M, Campa M, Dalla Costa L et al (2016) Knockdown of MLO genes reduces susceptibility to powdery mildew in grapevine. Hortic Res 3:16016. https://doi.org/10.1038/hortres.2016.16
Piper MC, van Helden M, Court LN, Tay WT (2016) Complete mitochondrial genome of the European grapevine moth (EGVM) Lobesia botrana (Lepidoptera: Tortricidae). Mitochondrial DNA Part A 27:3759–3760. https://doi.org/10.3109/19401736.2015.1079893
Pirrello C, Mizzotti C, Tomazetti TC, Colombo M, Bettinelli P et al (2019) Emergent ascomycetes in viticulture: an interdisciplinary overview. Front Plant Sci 10:1–30. https://doi.org/10.3389/fpls.2019.01394
Pirrello C, Zeilmaker T, Bianco L, Giacomelli L, Moser C et al (2021) Mining grapevine downy mildew susceptibility genes: a resource for genomics-based breeding and tailored gene editing. Biomolecules 11:181. https://doi.org/10.3390/biom11020181
Poland JA, Nelson RJ (2011) In the eye of the beholder: the effect of rater variability and different rating scales on QTL mapping. Phytopathology 101:290–298. https://doi.org/10.1094/PHYTO-03-10-0087
Polesani M, Bortesi L, Ferrarini A, Zamboni A, Fasoli M et al (2010) General and species-specific transcriptional responses to downy mildew infection in a susceptible (Vitis vinifera) and a resistant (V. riparia) grapevine species. BMC Genomics 11:117. https://doi.org/10.1186/1471-2164-11-117
Poolsawat O, Tharapreuksapong A, Wongkaew S, Chaowiset W, Tantasawat P (2012) Laboratory and field evaluations of resistance to Sphaceloma ampelinum causing anthracnose in grapevine. Australas Plant Pathol 41:263–269. https://doi.org/10.1007/s13313-012-0127-5
Poolsawat O, Tharapreuksapong A, Wongkaew S, Reisch B, Tantasawat P (2010) Genetic diversity and pathogenicity analysis of Sphaceloma ampelinum causing grape anthracnose in Thailand. J Phytopathol 158:837–840. https://doi.org/10.1111/j.1439-0434.2010.01696.x
Possamai T, Wiedemann-Merdinoglu S, De Mori G, Cipriani G, Velasco R (2021) Construction of a high-density genetic map and detection of a major QTL of resistance to powdery mildew (Erysiphe necator Sch.) in Caucasian grapes (Vitis vinifera L.). BMC Plant Biol 21:528. https://doi.org/10.1186/s12870-021-03174-4
Pouzoulet J, Scudiero E, Schiavon M, Rolshausen PE (2017) Xylem vessel diameter affects the compartmentalization of the vascular pathogen Phaeomoniella chlamydospora in grapevine. Front Plant Sci 8:1–13. https://doi.org/10.3389/fpls.2017.01442
Pouzoulet J, Rolshausen PE, Charbois R, Chen J, Guillaumie S et al (2020) Behind the curtain of the compartmentalization process: exploring how xylem vessel diameter impacts vascular pathogen resistance. Plant Cell Environ 43:2782–2796. https://doi.org/10.1111/pce.13848
Poveda J, Abril-Urias P, Escobar C (2020) Biological control of plant-parasitic nematodes by filamentous fungi inducers of resistance: Trichoderma. Mycorrhizal and Endophytic Fungi. Front Microbiol 11:992. https://doi.org/10.3389/fmicb.2020.00992
Powell KS (2012) A holistic approach to future management of grapevine phylloxera. In: Bostanian N, Vincent C, Isaacs R (eds) Arthropod management in Vineyards: Springer, Dordrecht, Netherlands, pp 219–251. https://doi.org/10.1007/978-94-007-4032-7_10
Powell KS, Cooper PD, Forneck A (2013) The biology, physiology and host-plant interactions of grape phylloxera Daktulosphaira vitifoliae. In: Advances in insect physiology. Academic Press Inc., pp 45:159–218. https://doi.org/10.1016/B978-0-12-417165-7.00004-0
Pozharskiy AS, Aubakirova KP, Gritsenko DA, Tlevlesov NI, Karimov NZ et al (2020) Genotyping and morphometric analysis of Kazakhstani grapevine cultivars versus Asian and European cultivars. Genet Mol Res 19(1):gmr18482. https://doi.org/10.4238/gmr18482
Pozzebon A, Tirello P, Moret R, Pederiva M, Duso C (2015) A fundamental step in IPM on grapevine: evaluating the side effects of pesticides on predatory mites. Insects 6:847–857. https://doi.org/10.3390/insects6040847
Prajongjai T, Poolsawat O, Pombungkerd P, Wongkaew S, Tantasawat PA (2014) Evaluation of grapevines for resistance to Downy Mildew (Plasmopara viticola) under laboratory and field conditions. South African J Enol Vitic 35:43–50. https://doi.org/10.21548/35-1-983
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575. https://doi.org/10.1086/519795
Rahman MU, Ma Q, Ahmad B, Hanif M, Zhang Y (2020) Histochemical and microscopic studies predict that grapevine genotype “Ju mei gui” is highly resistant against Botrytis cinerea. Pathogens 9(4):253. https://doi.org/10.3390/pathogens9040253
Rameshgar F, Khajehali J, Nauen R, Bajda S, Jonckheere W et al (2019a) Point mutations in the voltage-gated sodium channel gene associated with pyrethroid resistance in Iranian populations of the European red mite Panonychus ulmi. Pestic Biochem Physiol 157:80–87. https://doi.org/10.1016/j.pestbp.2019.03.008
Rameshgar F, Khajehali J, Nauen R, Dermauw W, Van Leeuwen T (2019b) Characterization of abamectin resistance in Iranian populations of European red mite, Panonychus ulmi Koch (Acari: Tetranychidae). Crop Prot 125:104903. https://doi.org/10.1016/j.cropro.2019.104903
Ramming DW, Gabler F, Smilanick J, Cadle-Davidson M, Barba P et al (2011) A single dominant locus, Ren4, confers rapid non-race-specific resistance to grapevine powdery mildew. Phytopathology 101:502–508. https://doi.org/10.1094/PHYTO-09-10-0237
Ramming DW, Gabler F, Smilanick JL, Margosan D, Cadle-Davidson MM et al (2012) Identification of race-specific resistance in North American Vitis spp. limiting Erysiphe necator hyphal growth. Phytopathology 102:83–93. https://doi.org/10.1094/PHYTO-03-11-0062
Ramos-Madrigal J, Runge AKW, Bouby L, Lacombe T, Samaniego Castruita JA et al (2019) Palaeogenomic insights into the origins of French grapevine diversity. Nat Plants 5:595–603. https://doi.org/10.1038/s41477-019-0437-5
Ramsdell DC, Milholland RD (1988) Black rot. In: Pearson RC, Goheen AC (eds) Compendium of grape diseases. APS Press, St. Paul, MN, pp 15–16
Raski DJ, Hewitt WB, Goheen AC, Taylor CE, Taylor RH (1965) Survival of Xiphinema index and reservoirs of fanleaf virus in fallowed vineyard soil. Nematologica 11:349–352. https://doi.org/10.1163/187529265X00267
Reineke A, Assaf HA, Kulanek D, Mori N, Pozzebon A et al (2015) A novel set of microsatellite markers for the European grapevine moth Lobesia botrana isolated using next-generation sequencing and their utility for genetic characterization of populations from Europe and the Middle East. Bull Entomol Res 105:408–416. https://doi.org/10.1017/S0007485315000267
Ren C, Xu Z, Sun S, Lee M-K, Wu C, et al. (2005) Genomic DNA libraries and physical mapping. In: Meksem K, Kahl G (eds) The handbook of plant genome mapping. Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, FRG, pp 173–213. https://doi.org/10.1002/3527603514.ch8
Ren C, Liu X, Zhang Z, Wang Y, Duan W et al (2016) CRISPR/Cas9-mediated efficient targeted mutagenesis in Chardonnay (Vitis vinifera L.). Sci Rep 6:1–9. https://doi.org/10.1038/srep32289
Ren C, Guo Y, Kong J, Lecourieux F, Dai Z et al (2020) Knockout of VvCCD8 gene in grapevine affects shoot branching. BMC Plant Biol 20:1–8. https://doi.org/10.1186/s12870-020-2263-3
Ren C, Liu Y, Guo Y, Duan W, Fan P et al (2021) Optimizing the CRISPR/Cas9 system for genome editing in grape by using grape promoters. Hortic Res 8:52. https://doi.org/10.1038/s41438-021-00489-z
Ren F, Ren C, Zhang Z, Duan W, Lecourieux D et al (2019) Efficiency optimization of CRISPR/Cas9-mediated targeted mutagenesis in grape. Front Plant Sci 10:1–8. https://doi.org/10.3389/fpls.2019.00612
Reustle G, Harst M, Alleweldt G (1994) Regeneration of grapevine (Vitis sp.) protoplasts. VITIS J Grapevine Res 33:173–174. https://doi.org/10.5073/vitis.1994.33.173-174
Rex F, Fechter I, Hausmann L, Töpfer R (2014) QTL mapping of black rot (Guignardia bidwellii) resistance in the grapevine rootstock “Börner” (V. riparia Gm183 × V. cinerea Arnold). Theor Appl Genet 127:1667–1677. https://doi.org/10.1007/s00122-014-2329-4
Reynolds AG, Reisch BI (2015) Grapevine breeding in the Eastern United States. In: Reynolds A (ed) Grapevine breeding programs for the wine industry. Elsevier, Woodhead Publishing, Sawston, UK, pp 345–358. https://doi.org/10.1016/B978-1-78242-075-0.00014-4
Riaz S, Krivanek AF, Xu K, Walker MA (2006) Refined mapping of the Pierce’s disease resistance locus, PdR1, and Sex on an extended genetic map of Vitis rupestris x V. arizonica. Theor Appl Genet 113:1317–1329. https://doi.org/10.1007/s00122-006-0385-0
Riaz S, Tenscher AC, Rubin J, Graziani R, Pao SS et al (2008) Fine-scale genetic mapping of two Pierce’s disease resistance loci and a major segregation distortion region on chromosome 14 of grape. Theor Appl Genet 117:671–681. https://doi.org/10.1007/s00122-008-0802-7
Riaz S, Tenscher AC, Graziani R, Andrew Walker M (2008) Using marker-assisted selection to breed for Pierce's disease resistance in grape. Am J Enol Vitic 59:341
Riaz S, Tenscher AC, Graziani R, Krivanek AF, Ramming DW et al (2009) Using marker-assisted selection to breed Pierce’s disease-resistant grapes. Am J Enol Vitic 60:199–207
Riaz S, Tenscher AC, Ramming DW, Walker MA (2011) Using a limited mapping strategy to identify major QTLs for resistance to grapevine powdery mildew (Erysiphe necator) and their use in marker-assisted breeding. Theor Appl Genet 122:1059–1073. https://doi.org/10.1007/s00122-010-1511-6
Riaz S, Boursiquot J-M, Dangl GS, Lacombe T, Laucou V et al (2013) Identification of mildew resistance in wild and cultivated Central Asian grape germplasm. BMC Plant Biol 13:149. https://doi.org/10.1186/1471-2229-13-149
Riaz S, Lund KT, Granett J, Walker MA (2017) Population diversity of grape phylloxera in California and evidence for sexual reproduction. Am J Enol Vitic 68:218–227. https://doi.org/10.5344/ajev.2016.15114
Riaz S, Huerta-Acosta K, Tenscher AC, Walker MA (2018a) Genetic characterization of Vitis germplasm collected from the Southwestern US and Mexico to expedite Pierce’s disease-resistance breeding. Theor Appl Genet 131:1589–1602. https://doi.org/10.1007/s00122-018-3100-z
Riaz S, De Lorenzis G, Velasco D, Koehmstedt A, Maghradze D et al (2018b) Genetic diversity analysis of cultivated and wild grapevine (Vitis vinifera L.) accessions around the Mediterranean basin and Central Asia. BMC Plant Biol 18:137. https://doi.org/10.1186/s12870-018-1351-0
Riaz S, Pap D, Uretsky J, Laucou V, Boursiquot J-MM et al (2019) Genetic diversity and parentage analysis of grape rootstocks. Theor Appl Genet 132:1847–1860. https://doi.org/10.1007/s00122-019-03320-5
Riaz S, Menéndez CM, Tenscher A, Pap D, Walker MA (2020a) Genetic mapping and survey of powdery mildew resistance in the wild Central Asian ancestor of cultivated grapevines in Central Asia. Hortic Res 7. https://doi.org/10.1038/s41438-020-0335-z
Riaz S, Tenscher AC, Heinitz CC, Huerta-Acosta KG, Walker MA (2020b) Genetic analysis reveals an East-West divide within North American Vitis species that mirrors their resistance to Pierce’s disease. PLoS ONE 15:e0243445. https://doi.org/10.1371/journal.pone.0243445
Richter R, Gabriel D, Rist F, Töpfer R, Zyprian E (2019) Identification of co-located QTLs and genomic regions affecting grapevine cluster architecture. Theor Appl Genet 132:1159–1177. https://doi.org/10.1007/s00122-018-3269-1
Ridgman WJ (1991) The status of Eutypa lata as a Pathogen. CABI International, Wallingford, UK
Rilling G (1989) Differential response of grapevine cultivars to European red mite (Panonychus ulmi Koch) - elaboration of a screening method. VITIS J Grapevine Res 28(2):97–97. https://doi.org/10.5073/VITIS.1989.28.97-110
Rinaldi PA, Paffetti D, Comparini C, Broggini GAL, Gessler C et al (2017) Genetic variability of Phyllosticta ampelicida, the agent of black rot disease of grapevine. Phytopathology 107:1406–1416. https://doi.org/10.1094/PHYTO-11-16-0404-R
Rispe C, Legeai F, Nabity PD, Fernández R, Arora AK et al (2020) The genome sequence of the grape phylloxera provides insights into the evolution, adaptation, and invasion routes of an iconic pest. BMC Biol 18:1–25. https://doi.org/10.1186/s12915-020-00820-5
Rispe C, Legeai F, Papura D, Bretaudeau A, Hudaverdian S et al (2016) De novo transcriptome assembly of the grapevine phylloxera allows identification of genes differentially expressed between leaf- and root-feeding forms. BMC Genomics 17:1–15. https://doi.org/10.1186/s12864-016-2530-8
Rodríguez-Leal D, Lemmon ZH, Man J, Bartlett ME, Lippman ZB (2017) Engineering quantitative trait variation for crop improvement by genome editing. Cell 171:470–480.e8. https://doi.org/10.1016/j.cell.2017.08.030
Rojas V, Jiménez H, Palma-Millanao R, González-González A, Machuca J et al (2018) Analysis of the grapevine moth Lobesia botrana antennal transcriptome and expression of odorant-binding and chemosensory proteins. Comp Biochem Physiol Part D Genomics Proteomics 27:1–12. https://doi.org/10.1016/j.cbd.2018.04.003
Rolshausen PE, Úrbez-Torres JR, Rooney-Latham S, Eskalen A, Smith RJ et al (2010) Evaluation of pruning wound susceptibility and protection against fungi associated with grapevine trunk diseases. Am J Enol Vitic 61:113–119
Rooney-Latham S, Eskalen A, Gubler WD (2005) Occurrence of Togninia minima perithecia in esca-affected vineyards in California. Plant Dis 89:867–871. https://doi.org/10.1094/PD-89-0867
Rossi V, Caffi T, Legler SE, Carotenuto E, Bigot G (2014) Large-scale application of a web-based Decision Support System for sustainable viticulture. IOBC/WPRS Bull
Rossi Stacconi MV, Grassi A, Ioriatti C, Anfora G (2019) Augmentative releases of Trichopria drosophilae for the suppression of early season Drosophila suzukii populations. Biocontrol 64:9–19. https://doi.org/10.1007/s10526-018-09914-0
Rossman AY, Crous PW, Hyde KD, Hawksworth DL, Aptroot A et al (2015) Recommended names for pleomorphic genera in Dothideomycetes. IMA Fungus 6:507–523. https://doi.org/10.5598/imafungus.2015.06.02.14
Rossmann S, Richter R, Sun H, Schneeberger K, Töpfer R et al (2020) Mutations in the miR396 binding site of the growth-regulating factor gene VvGRF4 modulate inflorescence architecture in grapevine. Plant J 101:1234–1248. https://doi.org/10.1111/tpj.14588
Roush TL, Granett J, Walker MA (2007) Inheritance of gall formation relative to phylloxera resistance levels in hybrid grapevines. Am J Enol Vitic 58:234–241
Rouxel M, Mestre P, Comont G, Lehman BL, Schilder A et al (2013) Phylogenetic and experimental evidence for host-specialized cryptic species in a biotrophic oomycete. New Phytol 197:251–263. https://doi.org/10.1111/nph.12016
Rouxel M, Mestre P, Baudoin A, Carisse O, Delière L et al (2014) Geographic distribution of cryptic species of Plasmopara viticola causing downy mildew on wild and cultivated grape in Eastern North America. Phytopathology 104:692–701. https://doi.org/10.1094/PHYTO-08-13-0225-R
Rubio B, Lalanne-Tisné G, Voisin R, Tandonnet J-P, Portier U et al (2020) Characterization of genetic determinants of the resistance to phylloxera, Daktulosphaira vitifoliae, and the dagger nematode Xiphinema index from muscadine background. BMC Plant Biol 20:1–15. https://doi.org/10.1186/s12870-020-2310-0
Sáenz-Romo MG, Martínez-García H, Veas-Bernal A, Carvajal-Montoya LD, Martínez-Villar E et al (2019) Effect of ground-cover management on predatory mites (Acari: Phytoseiidae) in a Mediterranean vineyard. VITIS J Grapevine Res 58:25–32. https://doi.org/10.5073/vitis.2019.58.special-issue.25-32
Salvagnin U, Carlin S, Angeli S, Vrhovsek U, Anfora G et al (2016) Homologous and heterologous expression of grapevine E-(β)-caryophyllene synthase (VvGwECar2). Phytochemistry 131:76–83. https://doi.org/10.1016/j.phytochem.2016.08.002
Salvagnin U, Malnoy M, Thöming G, Tasin M, Carlin S et al (2018) Adjusting the scent ratio: using genetically modified Vitis vinifera plants to manipulate European grapevine moth behaviour. Plant Biotechnol J 16:264–271. https://doi.org/10.1111/pbi.12767
Sambucci O, Alston JM, Fuller KB, Lusk J (2019) The pecuniary and nonpecuniary costs of powdery mildew and the potential value of resistant grape varieties in California. Am J Enol Vitic 70:177–187. https://doi.org/10.5344/ajev.2018.18032
Santos JM, Correia VG, Phillips AJL (2010) Primers for mating-type diagnosis in Diaporthe and Phomopsis: their use in teleomorph induction in vitro and biological species definition. Fungal Biol 114:255–270. https://doi.org/10.1016/j.funbio.2010.01.007
dos Santos RF, Spósito MB, Ayres MR, Sosnowski MR (2018) Phylogeny, morphology and pathogenicity of Elsinoë ampelina, the causal agent of grapevine anthracnose in Brazil and Australia. J Phytopathol 166:187–198. https://doi.org/10.1111/jph.12675
dos Santos RF, Ciampi-Guillardi M, Amorim L, Massola NS, Spósito MB (2018) Aetiology of anthracnose on grapevine shoots in Brazil. Plant Pathol 67:692–706. https://doi.org/10.1111/ppa.12756
Sapkota S, Chen LL, Yang S, Hyma KE, Cadle-Davidson L et al (2019a) Construction of a high-density linkage map and QTL detection of downy mildew resistance in Vitis aestivalis-derived ‘Norton.’ Theor Appl Genet 132:137–147. https://doi.org/10.1007/s00122-018-3203-6
Sapkota SD, Chen LL, Yang S, Hyma KE, Cadle-Davidson LE et al (2019b) Quantitative Trait Locus mapping of downy mildew and Botrytis bunch rot resistance in a Vitis aestivalis-derived ‘Norton’-based population. Acta Hortic 1248:305–311. https://doi.org/10.17660/ActaHortic.2019.1248.44
Sargolzaei M, Maddalena G, Bitsadze N, Maghradze D, Bianco PA et al (2020) Rpv29, Rpv30 and Rpv31: three novel genomic loci associated with resistance to Plasmopara viticola in Vitis vinifera. Front Plant Sci 11:1–16. https://doi.org/10.3389/fpls.2020.562432
Sargolzaei M, Rustioni L, Cola G, Ricciardi V, Bianco PA et al (2021) Georgian grapevine cultivars: ancient biodiversity for future viticulture. Front Plant Sci 12:630122. https://doi.org/10.3389/fpls.2021.630122
Saucet SB, Van Ghelder C, Abad P, Duval H, Esmenjaud D (2016) Resistance to root-knot nematodes Meloidogyne spp. in woody plants. New Phytol 211:41–56. https://doi.org/10.1111/nph.13933
Savi T, García González A, Herrera JC, Forneck A (2019) Gas exchange, biomass and non-structural carbohydrates dynamics in vines under combined drought and biotic stress. BMC Plant Biol 19:1–11. https://doi.org/10.1186/s12870-019-2017-2
Savoi S, Eitle MW, Berger H, Curto M, Meimberg H et al (2020) Comparative transcriptome analysis of two root- feeding grape phylloxera (D. vitifoliae) lineages feeding on a rootstock and V. vinifera. Insects 11:1–21. https://doi.org/10.3390/insects11100691
Scalabrin S, Troggio M, Moroldo M, Pindo M, Felice N et al (2010) Physical mapping in highly heterozygous genomes: a physical contig map of the Pinot Noir grapevine cultivar. BMC Genomics 11:204. https://doi.org/10.1186/1471-2164-11-204
Schellenbaum P, Mohler V, Wenzel G, Walter B (2008) Variation in DNA methylation patterns of grapevine somaclones (Vitis vinifera L.). BMC Plant Biol 8:78. https://doi.org/10.1186/1471-2229-8-78
Scheper RWA (2001) Studies on the biology and genetic variation of Phomopsis on grapevine. PhD Thesis University of Adelaide, SA
Schmid J, Manty F, Cousins P (2009) Collecting Vitis berlandieri from native habitat sites. Acta Hortic 827:151–154. https://doi.org/10.17660/ActaHortic.2009.827.22
Schmidt RA (2014) Leaf structures affect predatory mites (Acari: Phytoseiidae) and biological control: a review. Exp Appl Acarol 62:1–17. https://doi.org/10.1007/s10493-013-9730-6
Schmitt-Keichinger C, Hemmer C, Berthold F, Ritzenthaler C (2017) Molecular, cellular, and structural biology of Grapevine Fanleaf Virus. In: Meng B, Martelli GP, Golino DA, Fuchs M (eds) Grapevine viruses: molecular biology, diagnostics and management. Springer, Cham, Switzerland, pp 83–107. https://doi.org/10.1007/978-3-319-57706-7_4
Schouten HJ, Krens FA, Jacobsen E (2006) Cisgenic plants are similar to traditionally bred plants: international regulations for genetically modified organisms should be altered to exempt cisgenesis. EMBO Rep 7:750–753. https://doi.org/10.1038/sj.embor.7400769
Schreiner U (1984) Untersuchungen zur Anfälligkeit verschiedener Rebsorten, Pfropfkombinationen und Unterlagen gegenüber Tetranychus urticae und Panonychus ulmi. PhD Thesis Universität Kaiserslautern, Germany
Schröder S, Telle S, Nick P, Thines M (2011) Cryptic diversity of Plasmopara viticola (Oomycota, Peronosporaceae) in North America. Org Divers Evol 11:3–7. https://doi.org/10.1007/s13127-010-0035-x
Schruft G, Mittenmüller K, Stärk O (1979) Die Überwinterung der Gemeinen Spinnmilbe Tetranychus urticae Koch an der Rebe und ihr Auftreten im Frühjahr. Wein-Wissen 34:55–60
Schultz JC, Edger PP, Body MJA, Appel HM (2019) A galling insect activates plant reproductive programs during gall development. Sci Rep 9:1833. https://doi.org/10.1038/s41598-018-38475-6
Schwander F, Eibach R, Fechter I, Hausmann L, Zyprian E et al (2012) Rpv10: a new locus from the Asian Vitis gene pool for pyramiding downy mildew resistance loci in grapevine. Theor Appl Genet 124:163–176. https://doi.org/10.1007/s00122-011-1695-4
Scintilla S, Salvagnin U, Giacomelli L, Zeilmaker T, Malnoy MA et al (2021) Regeneration of plants from DNA-free edited grapevine protoplasts. bioRxiv 2021.07.16.452503
Sefc KM, Regner F, Turetschek E, Glössl J, Steinkellner H (1999) Identification of microsatellite sequences in Vitis riparia and their applicability for genotyping of different Vitis species. Genome 42:367–373
Segura V, Vilhjálmsson BJ, Platt A, Korte A, Seren Ü et al (2012) An efficient multi-locus mixed-model approach for genome-wide association studies in structured populations. Nat Genet 44:825–830. https://doi.org/10.1038/ng.2314
Sergeeva V, Nair NG, Barchia I, Priest M, Spooner-Hart R (2003) Germination of β conidia of Phomopsis viticola. Australas Plant Pathol 32:105–107. https://doi.org/10.1071/AP02072
Šeruga Musić M, Samarzija I, Hogenhout SA, Haryono M, Cho ST et al (2019) The genome of ‘Candidatus Phytoplasma solani’ strain SA-1 is highly dynamic and prone to adopting foreign sequences. Syst Appl Microbiol 42:117–127. https://doi.org/10.1016/j.syapm.2018.10.008
Shapira I, Keasar T, Harari AR, Gavish-Regev E, Kishinevsky M et al (2018) Does mating disruption of Planococcus ficus and Lobesia botrana affect the diversity, abundance and composition of natural enemies in Israeli vineyards? Pest Manag Sci 74:1837–1844. https://doi.org/10.1002/ps.4883
Silva DE, Do Nascimento JM, da Silva RTL, Ferla JJ, Ruffatto K et al (2021) Feeding preference and biological traits of Panonychus ulmi on leaves of apple and grapevine. Oecologia Aust 25:80–89. https://doi.org/10.4257/OECO.2021.2501.08
Simmons GS, Varela L, Daugherty M, Cooper M, Lance D et al (2021) Area-wide Eradication of the invasive European grapevine moth Lobesia botrana in California, USA. In: Hendrichs J, Pereira R, Vreysen MJB (eds) Area-wide integrated pest management, 1st edn. CRC Press, Boca Raton, FL, pp 581–596. https://doi.org/10.1201/9781003169239-31
Simpson AJG, Reinach FC, Arruda P, Abreu FA, Acencio M et al (2000) The genome sequence of the plant pathogen Xylella fastidiosa. Nature 406:151–157. https://doi.org/10.1038/35018003
Slater SC, Goldman BS, Goodner B, Setubal JC, Farrand SK et al (2009) Genome sequences of three Agrobacterium biovars help elucidate the evolution of multichromosome genomes in bacteria. J Bacteriol 191:2501–2511. https://doi.org/10.1128/JB.01779-08
Smit SJ, Vivier MA, Young PR (2020) Comparative (within species) genomics of the Vitis vinifera L. terpene synthase family to explore the impact of genotypic variation using phased diploid genomes. Front Genet 11:421. https://doi.org/10.3389/fgene.2020.00421
Smith LM, Stafford EM (1948) The bud mite and the erineum mite of grapes. Hilgardia 18:317–334. https://doi.org/10.3733/hilg.v18n07p317
Smith HM, Smith BP, Morales NB, Moskwa S, Clingeleffer PR et al (2018a) SNP markers tightly linked to root knot nematode resistance in grapevine (Vitis cinerea) identified by a genotyping-by-sequencing approach followed by Sequenom MassARRAY validation. PLoS ONE 13:e0193121. https://doi.org/10.1371/journal.pone.0193121
Smith HM, Clarke CW, Smith BP, Carmody BM, Thomas MR et al (2018b) Genetic identification of SNP markers linked to a new grape phylloxera resistant locus in Vitis cinerea for marker-assisted selection. BMC Plant Biol 18:1–13. https://doi.org/10.1186/s12870-018-1590-0
Smith HM, Dunlevy JD, Morales NB, Smith AA, Clingeleffer PR (2019) Developing next-generation grapevine rootstocks with long-term resistance to phylloxera and root knot nematode. In: Beames KS, Robinson EMC, Dry PR, Johnson DL (eds) Australian wine industry technical conference. Adelaide, SA, pp 107–110
Sofia J, Mota M, Gonçalves MT, Rego C (2018) Response of four Portuguese grapevine cultivars to infection by Phaeomoniella chlamydospora. Phytopathol Mediterr 57:506–518. https://doi.org/10.14601/Phytopathol_Mediterr-23485
Sosnowski MR, Lardner R, Wicks TJ, Scott ES (2007) The influence of grapevine cultivar and isolate of Eutypa lata on wood and foliar symptoms. Plant Dis 91:924–931. https://doi.org/10.1094/PDIS-91-8-0924
Sosnowski M, Ayres MR, McCarthy M, Wicks T, Scott E (2016) Investigating potential resistance grapevine trunk diseases. Wine Vitic J 31:41–45
Spagnolo A, Magnin-Robert M, Alayi TD, Cilindre C, Mercier L et al (2012) Physiological changes in green stems of Vitis vinifera L. cv. Chardonnay in response to esca proper and apoplexy revealed by proteomic and transcriptomic analyses. J Proteome Res 11:461–475. https://doi.org/10.1021/pr200892g
Spigno G, Marinoni L, Garrido GD (2017) State of the art in grape processing by-products. In: Galanakis CM (ed) Handbook of grape processing by-products. Elsevier Academic Press, Cambridge, MA, pp 1–27. https://doi.org/10.1016/B978-0-12-809870-7.00001-6
Staudt G, Weischer B (1992) Resistance to transmission of grapevine fanleaf virus by Xiphinema index in Vitis rotundifolia and Vitis munsoniana. Vitic. Enol. Sci. 47:56–61
Staudt G, Kassemeyer HH (1995) Evaluation of downy mildew resistance in various accessions of wild Vitis species. VITIS J Grapevine Res 34:225–228. https://doi.org/10.5073/VITIS.1995.34.225-228
St.Clair DA (2010) Quantitative disease resistance and quantitative resistance loci in breeding. Annu Rev Phytopathol 48:247–268. https://doi.org/10.1146/annurev-phyto-080508-081904
Stam R, McDonald BA (2018) When resistance gene pyramids are not durable—the role of pathogen diversity. Mol Plant Pathol 19:521–524. https://doi.org/10.1111/mpp.12636
Su H, Jiao YT, Wang FF, Liu YE, Niu WL et al (2018) Overexpression of VpPR10.1 by an efficient transformation method enhances downy mildew resistance in V. vinifera. Plant Cell Rep 37:819–832. https://doi.org/10.1007/s00299-018-2271-z
Su K, Guo Y, Zhong W, Lin H, Liu Z et al (2021) High-density genetic linkage map construction and white rot resistance Quantitative Trait Loci mapping for genus Vitis based on restriction site-associated DNA sequencing. Phytopathology 111:659–670. https://doi.org/10.1094/PHYTO-12-19-0480-R
Sunitha S, Rock CD (2020) CRISPR/Cas9-mediated targeted mutagenesis of TAS4 and MYBA7 loci in grapevine rootstock 101–14. Transgenic Res 29:355–367. https://doi.org/10.1007/s11248-020-00196-w
Szitenberg A, Salazar-Jaramillo L, Blok VC, Laetsch DR, Joseph S et al (2017) Comparative genomics of apomictic root-knot nematodes: hybridization, ploidy, and dynamic genome change. Genome Biol Evol 9:2844–2861. https://doi.org/10.1093/gbe/evx201
Tait G, Vezzulli S, Sassù F, Antonini G, Biondi A et al (2017) Genetic variability in Italian populations of Drosophila suzukii. BMC Genet 18:87. https://doi.org/10.1186/s12863-017-0558-7
Tamba CL, Ni Y-L, Zhang Y-M (2017) Iterative sure independence screening EM-Bayesian LASSO algorithm for multi-locus genome-wide association studies. PLOS Comput Biol 13:e1005357. https://doi.org/10.1371/journal.pcbi.1005357
Tasin M, Bäckman AC, Bengtsson M, Ioriatti C, Witzgall P (2006) Essential host plant cues in the grapevine moth. Naturwissenschaften 93:141–144. https://doi.org/10.1007/s00114-005-0077-7
Tasin M, Lucchi A, Ioriatti C, Mraihi M, De cristofaro A et al (2011) Oviposition response of the moth Lobesia botrana to sensory cues from a host plant. Chem Senses 36:633–639. https://doi.org/10.1093/chemse/bjr027
Taylor CE, Raski DJ (1964) On the transmission of grape fanleaf by Xiphinema index. Nematologica 10:489–495. https://doi.org/10.1163/187529264X00510
Tegli S, Bertelli E, Surico G (2000) Sequence analysis of ITS ribosomal DNA in five Phaeoacremonium species and development of a PCR-based assay for the detection of P. chlamydosporum and P. aleophilum in grapevine tissue. Phytopathol Mediterr 39:134–149. https://doi.org/10.14601/Phytopathol_Mediterr-1555
Teh SL, Fresnedo-Ramírez J, Clark MD, Gadoury DM, Sun Q et al (2017) Genetic dissection of powdery mildew resistance in interspecific half-sib grapevine families using SNP-based maps. Mol Breed 37:1–16. https://doi.org/10.1007/s11032-016-0586-4
Teh SL, Rostandy B, Awale M, Luby JJ, Fennell A et al (2019) Genetic analysis of stilbenoid profiles in grapevine stems reveals a major mQTL hotspot on chromosome 18 associated with disease-resistance motifs. Hortic Res 6:121. https://doi.org/10.1038/s41438-019-0203-x
Téliz D, Landa BB, Rapoport HF, Camacho FP, Jiménez-Díaz RM et al (2007) Plant-parasitic nematodes infecting grapevine in Southern Spain and susceptible reaction to root-knot nematodes of rootstocks reported as moderately resistant. Plant Dis 91:1147–1154. https://doi.org/10.1094/PDIS-91-9-1147
Tello J, Torres-Pérez R, Grimplet J, Ibáñez J (2016) Association analysis of grapevine bunch traits using a comprehensive approach. Theor Appl Genet 129:227–242. https://doi.org/10.1007/s00122-015-2623-9
Tello J, Forneck A (2019) Use of DNA markers for grape Phylloxera population and evolutionary genetics: from RAPDS to SSRs and beyond. Insects 10:317. https://doi.org/10.3390/insects10100317
Tello J, Mammerler R, Čajić M, Forneck A (2019) Major outbreaks in the nineteenth century shaped grape phylloxera contemporary genetic structure in Europe. Sci Rep 9:1–11. https://doi.org/10.1038/s41598-019-54122-0
Jaillon O, Aury JM, Noel B, Policriti A, Clepet C, et al (2007) The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449:463–467. https://doi.org/10.1038/nature06148
Thiéry D, Moreau J (2005) Relative performance of European grapevine moth (Lobesia botrana) on grapes and other hosts. Oecologia 143:548–557. https://doi.org/10.1007/s00442-005-0022-7
Thind TS (2019) Anthracnose. In: Wilcox WF, Gubler WD, Uyemoto JK (eds) Compendium of grape diseases, disorders, and pests. APS Press, St. Paul, MN, pp 17–19
This P, Boursiquot J (1999) Essai de définition du cépage. Progrès Agric Vitic 116:359–361
This P, Jung A, Boccacci P, Borrego J, Botta R et al (2004) Development of a standard set of microsatellite reference alleles for identification of grape cultivars. Theor Appl Genet 109:1448–1458. https://doi.org/10.1007/s00122-004-1760-3
This P, Lacombe T, Thomas MR (2006) Historical origins and genetic diversity of wine grapes. Trends Genet 22:511–519. https://doi.org/10.1016/j.tig.2006.07.008
Tixier M-S (2018) Predatory mites (Acari: Phytoseiidae). In: Agro-ecosystems and conservation biological control: A review and explorative approach for forecasting plant-predatory mite interactions and mite dispersal. Front Ecol Evol 6. https://doi.org/10.3389/fevo.2018.00192
Thomas MR, Scott NS (1993) Microsatellite repeats in grapevine reveal DNA polymorphisms when analysed as sequence-tagged sites (STSs). Theor Appl Genet 86:985–990. https://doi.org/10.1007/BF00211051
Tian S, Yin X, Fu P, Wu W, Lu J (2020) Ectopic expression of grapevine gene VaRGA1 in arabidopsis improves resistance to downy mildew and Pseudomonas syringae pv. Tomato DC3000 but increases susceptibility to Botrytis cinerea. Int J Mol Sci 21. https://doi.org/10.3390/ijms21010193
Tibbs Cortes L, Zhang Z, Yu J (2021) Status and prospects of genome-wide association studies in plants. Plant Genome 14. https://doi.org/10.1002/tpg2.20077
Toffolatti S, Venturini G, Maffi D, Vercesi A (2012) Phenotypic and histochemical traits of the interaction between Plasmopara viticola and resistant or susceptible grapevine varieties. BMC Plant Biol 12:124. https://doi.org/10.1186/1471-2229-12-124
Toffolatti SL, Maddalena G, Salomoni D, Maghradze D, Bianco PA et al (2016) Evidence of resistance to the downy mildew agent Plasmopara viticola in the Georgian Vitis vinifera germplasm. VITIS J Grapevine Res 55:121–128. https://doi.org/10.5073/vitis.2016.55.121-128
Toffolatti SL, De Lorenzis G, Costa A, Maddalena G, Passera A et al (2018) Unique resistance traits against downy mildew from the center of origin of grapevine (Vitis vinifera). Sci Rep 8:12523. https://doi.org/10.1038/s41598-018-30413-w
Toffolatti SL, De Lorenzis G, Brilli M, Moser M, Shariati V et al (2020) Novel aspects on the interaction between grapevine and Plasmopara viticola: dual-RNA-seq analysis highlights gene expression dynamics in the pathogen and the plant during the battle for infection. Genes (basel) 11:261. https://doi.org/10.3390/genes11030261
Tomoiaga L, Chedea VS (2020) The behaviour of some grapevine varieties to the Guignardia bidwellii fungus attack. Bull Univ Agric Sci Vet Med Cluj-Napoca Hortic 77:122. https://doi.org/10.15835/buasvmcn-hort:2019.0042
Topalović O, Hussain M, Heuer H (2020) Plants and associated soil microbiota cooperatively suppress plant-parasitic nematodes. Front Microbiol 11:313. https://doi.org/10.3389/fmicb.2020.00313
Töpfer R, Hausmann L, Harst M, Maul E, Zyprian E et al (2011) New horizons for grapevine breeding. Fruit, Veg Cereal Sci Biotechnol 5:79–100
Travadon R, Rolshausen PE, Gubler WD, Cadle-Davidson L, Baumgartner K (2013) Susceptibility of cultivated and wild Vitis spp. to wood infection by fungal trunk pathogens. Plant Dis 97:1529–1536. https://doi.org/10.1094/PDIS-05-13-0525-RE
Travadon R, Lawrence DP, Rooney-Latham S, Gubler WD, Wilcox WF et al (2015) Cadophora species associated with wood-decay of grapevine in North America. Fungal Biol 119:53–66. https://doi.org/10.1016/j.funbio.2014.11.002
Trdá L, Fernandez O, Boutrot F, Héloir M, Kelloniemi J et al (2014) The grapevine flagellin receptor VvFLS2 differentially recognizes flagellin-derived epitopes from the endophytic growth-promoting bacterium Burkholderia phytofirmans and plant pathogenic bacteria. New Phytol 201:1371–1384. https://doi.org/10.1111/nph.12592
Trivellone V, Jermini M, Linder C, Cara C, Delabays N et al (2013) Rôle de la flore du vignoble sur la distribution de Scaphoideus titanus. Rev Suisse Vitic Arboric Hortic 45:222–228
Troggio M, Malacarne G, Coppola G, Segala C, Cartwright DA et al (2007) A dense single-nucleotide polymorphism-based genetic linkage map of grapevine (Vitis vinifera L.) anchoring pinot noir bacterial artificial chromosome contigs. Genetics 176:2637–2650. https://doi.org/10.1534/genetics.106.067462
Tröndle D, Schröder S, Kassemeyer H-H, Kiefer C, Koch MA et al (2010) Molecular phylogeny of the genus Vitis (Vitaceae) based on plastid markers. Am J Bot 97:1168–1178. https://doi.org/10.3732/ajb.0900218
Trotel-Aziz P, Couderchet M, Vernet G, Aziz A (2006) Chitosan stimulates defense reactions in grapevine leaves and inhibits development of Botrytis cinerea. Eur J Plant Pathol 114:405–413. https://doi.org/10.1007/s10658-006-0005-5
Tsai C-W, Rowhani A, Golino DA, Daane KM, Almeida RPP (2010) Mealybug transmission of Grapevine Leafroll Viruses: an analysis of virus–vector specificity. Phytopathology 100:830–834. https://doi.org/10.1094/PHYTO-100-8-0830
Tumber KP, Alston JM, Fuller KB (2014) Pierce’s disease costs California $104 million per year. Calif Agric 68:20–29. https://doi.org/10.3733/ca.v068n01p20
Uecker FA (1988) A world list of Phomopsis names with notes on nomenclature, morphology and biology. Mycol Mem 1–231
Ullrich C, Kleespies R, Enders M, Koch E (2009) Biology of the black rot pathogen, Guignardia bidwellii, its development in susceptible leaves of grapevine Vitis vinifera., J Für Kult 61:82–90
Umina PA, Corrie AM, Herbert KS, White VL, Powell KS et al (2007) The use of DNA markers for pest management-clonal lineages and population biology of grape phylloxera. Acta Hortic 733:183–195. https://doi.org/10.17660/ActaHortic.2007.733.20
Úrbez-Torres JR, Peduto F, Smith RJ, Gubler WD (2013) Phomopsis dieback: a grapevine trunk disease caused by Phomopsis viticola in California. Plant Dis 97:1571–1579. https://doi.org/10.1094/PDIS-11-12-1072-RE
Valenzano D, Bari G, Valeria M, de Lillo E (2019) Off-host survival of Eriophyoidea and remarks on their dispersal modes. Exp Appl Acarol 79:21–33. https://doi.org/10.1007/s10493-019-00417-w
Valenzano D, Tumminello MT, Gualandri V, de Lillo E (2020) Morphological and molecular characterization of the Colomerus vitis erineum strain (Trombidiformes: Eriophyidae) from grapevine erinea and buds. Exp Appl Acarol 80:183–201. https://doi.org/10.1007/s10493-020-00470-w
Van Damme M, Huibers RP, Elberse J, Van Den Ackerveken G (2008) Arabidopsis DMR6 encodes a putative 2OG-Fe(II) oxygenase that is defense-associated but required for susceptibility to downy mildew. Plant J 54:785–793. https://doi.org/10.1111/j.1365-313X.2008.03427.x
van Heerden CJ, Burger P, Vermeulen A, Prins R (2014) Detection of downy and powdery mildew resistance QTL in a ‘Regent’ × ‘RedGlobe’ population. Euphytica 200:281–295. https://doi.org/10.1007/s10681-014-1167-4
Van Leeuwen T, Tirry L, Yamamoto A, Nauen R, Dermauw W (2015) The economic importance of acaricides in the control of phytophagous mites and an update on recent acaricide mode of action research. Pestic Biochem Physiol 121:12–21. https://doi.org/10.1016/j.pestbp.2014.12.009
Van Leeuwen T, Witters J, Nauen R, Duso C, Tirry L (2010) The control of eriophyoid mites: state of the art and future challenges. Exp Appl Acarol 51:205–224. https://doi.org/10.1007/s10493-009-9312-9
Van Niekerk JM, Groenewald JZ, Farr DF, Fourie PH, Halleen F et al (2005) Reassessment of Phomopsis species on grapevines. Australas Plant Pathol 34:27–39. https://doi.org/10.1071/AP04072
Van Schie CCN, Takken FLW (2014) Susceptibility genes 101: how to be a good host. Annu Rev Phytopathol 52:551–581. https://doi.org/10.1146/annurev-phyto-102313-045854
Vandelle E, Ariani P, Regaiolo A, Danzi D, Lovato A et al (2021) The grapevine e3 ubiquitin ligase Vriatl156 confers resistance against the downy mildew pathogen Plasmopara viticola. Int J Mol Sci 22:1–19. https://doi.org/10.3390/ijms22020940
Vanegas F, Bratanov D, Powell KS, Weiss J, Gonzalez F (2018) A novel methodology for improving plant pest surveillance in vineyards and crops using UAV-based hyperspectral and spatial data. Sensors (switzerland) 18:1–21. https://doi.org/10.3390/s18010260
Varela A, Ibañez VN, Alonso R, Zavallo D, Asurmendi S et al (2021) Vineyard environments influence Malbec grapevine phenotypic traits and DNA methylation patterns in a clone-dependent way. Plant Cell Rep 40:111–125. https://doi.org/10.1007/s00299-020-02617-w
Vargas AM, Velez MD, de Andres MT, Laucou V, Lacombe T et al (2007) Corinto bianco: a seedless mutant of Pedro Ximenes. Am J Enol Vitic 58:540–543
Varnier A, Sanchez L, Vatsa P, Boudesocque L, Garcia-brugger A et al (2009) Bacterial rhamnolipids are novel MAMPs conferring resistance to Botrytis cinerea in grapevine. Plant Cell Environ 32:178–193. https://doi.org/10.1111/j.1365-3040.2008.01911.x
Velasco R, Zharkikh A, Troggio M, Cartwright DA, Cestaro A et al (2007) A high quality draft consensus sequence of the genome of a heterozygous grapevine variety. PLoS ONE 2. https://doi.org/10.1371/journal.pone.0001326
Venuti S, Copetti D, Foria S, Falginella L, Hoffmann S et al (2013) Historical introgression of the downy mildew resistance gene Rpv12 from the Asian species Vitis amurensis into grapevine varieties. PLoS ONE 8:e61228. https://doi.org/10.1371/journal.pone.0061228
Vezzulli S, Malacarne G, Masuero D, Vecchione A, Dolzani C et al (2019a) The Rpv3-3 haplotype and stilbenoid induction mediate downy mildew resistance in a grapevine interspecific population-supplementary material. Front Plant Sci 10:1–23. https://doi.org/10.3389/fpls.2019.00234
Vezzulli S, Doligez A, Bellin D (2019b) Molecular mapping of grapevine genes. In: Cantu D, Walker M (eds) The grape genome. Springer, Cham, Switzerland, pp 103–136. https://doi.org/10.1007/978-3-030-18601-2_7
Vezzulli S, Zulini L, Stefanini M (2019c) Genetics-assisted breeding for downy/powdery mildew and phylloxera resistance at FEM. BIO Web Conf 12:01020. https://doi.org/10.1051/bioconf/20191201020
Vigl Egarter L, Schmid A, Moser F, Balotti A, Gartner E et al (2018) Upward shifts in elevation - a winning strategy for mountain viticulture in the context of climate change? In: E3S web of conferences. p 2006. https://doi.org/10.1051/e3sconf/20185002006
Vigne E, Komar V, Fuchs M (2004) Field safety assessment of recombination in transgenic grapevines expressing the coat protein gene of Grapevine Fanleaf Virus. Transgenic Res 13:165–179. https://doi.org/10.1023/B:TRAG.0000026075.79097.c9
Villate L, Morin E, Demangeat G, Van Helden M, Esmenjaud D (2012) Control of Xiphinema index populations by fallow plants under greenhouse and field conditions. Phytopathology 102:627–634. https://doi.org/10.1094/PHYTO-01-12-0007
Viret O, Spring JL, Gindro K (2018) Stilbenes: biomarkers of grapevine resistance to fungal diseases. OENO One 52:235–240. https://doi.org/10.20870/oeno-one.2018.52.3.2033
Wairich A, Malabarba J, Buffon V, Porto DD, Togawa R et al (2021) Structural and molecular characterization of the Rpv3 locus towards the development of KASP markers for downy mildew resistance in grapevine (Vitis spp.). bioRxiv 2021.02.25.432814. https://doi.org/10.1101/2021.02.25.432814
Waite H, Morton L (2007) Hot water treatment, trunk diseases and other critical factors in the production of high-quality grapevine planting material. Phytopathol Mediterr 46:5–17. https://doi.org/10.14601/Phytopathol_Mediterr-1857
Walker A-S (2016) Diversity within and between species of Botrytis. In: Fillinger S, Elad Y (eds) Botrytis – the fungus, the pathogen and its management in agricultural systems. Springer, Cham, Switzerland, pp 91–125. https://doi.org/10.1007/978-3-319-23371-0_6
Walker A, Tenscher A (2017) Breeding Pierce’s disease resistant winegrapes. In: Research progress reports Pierce’s disease and other designated pests and diseases of winegrapes. California Department of Food and Agriculture, San Diego, CA, pp 167–177
Walker MA, Tenscher A (2019) Breeding Pierce’s disease resistant winegrapes. In: Department of Food and Agriculture (ed) Research progress reports: Pierce’s disease and other designated pests and diseases of winegrapes. Sacramento, CA, pp 104–115
Walker MA, Heinitz C, Riaz S, Uretsky J (2019) Grape taxonomy and germplasm. In: Cantu D, Walker MA (eds) The grape genome. Springer, Cham, Switzerland, pp 25–38. https://doi.org/10.1007/978-3-030-18601-2_2
Walker MA, Tenscher AC, Riaz S, Romero N (2021a) Grapevine plant named ‘Camminare noir’
Walker MA, Tenscher AC, Riaz S, Romero N (2021b) Grapevine plant named ‘Ambulo blanc’
Walsh DB, Bolda MP, Goodhue RE, Dreves AJ, Lee J et al (2011) Drosophila suzukii (Diptera: Drosophilidae): invasive pest of ripening soft fruit expanding its geographic range and damage potential. J Integr Pest Manag 2:G1–G7. https://doi.org/10.1603/IPM10010
Walton VM, Daane KM, Pringle KL (2004) Monitoring Planococcus ficus in South African vineyards with sex pheromone-baited traps. Crop Prot 23:1089–1096. https://doi.org/10.1016/j.cropro.2004.03.016
Walton VM, Dreves AJ, Gent DH, James DG, Martin RR et al (2007) Relationship between rust mites Calepitrimerus vitis (nalepa), bud mites Colomerus vitis (pagenstecher) (acari: Eriophyidae) and short shootsyndrome in Oregon vineyards. Int J Acarol 33:307–318. https://doi.org/10.1080/01647950708683691
Walton VM, Daane KM, Bentley WJ, Millar JG, Larsen TE et al (2006) Pheromone-based mating disruption of Planococcus ficus (Hemiptera: Pseudococcidae) in California vineyards. J Econ Entomol 99:1280–1290.
Walton VM, Dreves AJ, Coop LB, Jones GV, Skinkis PA (2010) Developmental parameters and seasonal phenology of Calepitrimerus vitis (Acari: Eriophyidae) in wine grapes of Western Oregon. Environ Entomol 39:2006–2016. https://doi.org/10.1603/EN09197
Wan Y, He P, Wang Y (2007a) Inheritance of downy mildew resistance in two interspecific crosses between Chinese wild grapes and European grapes. VITIS J Grapevine Res 46:156–157. https://doi.org/10.5073/vitis.2007.46.156-157
Wan Y, Schwaninger H, He P, Wang Y (2007b) Comparison of resistance to powdery mildew and downy mildew in Chinese wild grapes. VITIS J Grapevine Res 46:132–136. https://doi.org/10.5073/vitis.2007.46.132-136
Wan Y, Schwaninger H, Li D, Simon CJ, Wang Y et al (2008a) A review of taxonomic research on Chinese wild grapes. VITIS J Grapevine Res 47:81–88. https://doi.org/10.5073/vitis.2008.47.81-88
Wan Y, Schwaninger H, Li D, Simon CJ, Wang Y et al (2008b) The eco-geographic distribution of wild grape germplasm in China. VITIS J Grapevine Res 47:77–80. https://doi.org/10.5073/vitis.2008.47.77-80
Wan Y, Schwaninger HR, Baldo AM, Labate JA, Zhong G-YY et al (2013) A phylogenetic analysis of the grape genus (Vitis L.) reveals broad reticulation and concurrent diversification during neogene and quaternary climate change. BMC Evol Biol 13:141. https://doi.org/10.1186/1471-2148-13-141
Wan D-Y, Guo Y, Cheng Y, Hu Y, Xiao S et al (2020) CRISPR/Cas9-mediated mutagenesis of VvMLO3 results in enhanced resistance to powdery mildew in grapevine (Vitis vinifera). Hortic Res 7:116. https://doi.org/10.1038/s41438-020-0339-8
Wang Y, Liu Y, He P, Chen J, Lamikanra O et al (1995) Evaluation of foliar resistance to Uncinula necator in Chinese wild Vitis spp. species. VITIS J Grapevine Res 34:159–164
Wang Y, Liu Y, He P, Lamikanra O, Lu J (1998) Resistance of Chinese Vitis species to Elsinoe ampelina (de Bary) Shear. Hortic Sci 33:123–126. https://doi.org/10.21273/HORTSCI.33.1.123
Wang W, Wen Y, Berkey R, Xiao S (2009) Specific targeting of the arabidopsis resistance protein RPW8.2 to the interfacial membrane encasing the fungal haustorium renders broad-spectrum resistance to powdery mildew. Plant Cell 21:2898–2913. https://doi.org/10.1105/tpc.109.067587
Wang S-B, Feng J-Y, Ren W-L, Huang B, Zhou L et al (2016a) Improving power and accuracy of genome-wide association studies via a multi-locus mixed linear model methodology. Sci Rep 6:19444. https://doi.org/10.1038/srep19444
Wang Y, Liu X, Ren C, Zhong GY, Yang L et al (2016b) Identification of genomic sites for CRISPR/Cas9-based genome editing in the Vitis vinifera genome. BMC Plant Biol 16:3–9. https://doi.org/10.1186/s12870-016-0787-3
Wang J, Yao W, Wang L, Ma F, Tong W et al (2017a) Overexpression of VpEIFP1, a novel F-box/Kelch-repeat protein from wild Chinese Vitis pseudoreticulata, confers higher tolerance to powdery mildew by inducing thioredoxin z proteolysis. Plant Sci 263:142–155. https://doi.org/10.1016/j.plantsci.2017.07.004
Wang L, Xie X, Yao W, Wang J, Ma F et al (2017b) RING-H2-type E3 gene VpRH2 from Vitis pseudoreticulata improves resistance to powdery mildew by interacting with VpGRP2A. J Exp Bot 68:1669–1687. https://doi.org/10.1093/jxb/erx033
Wang M, Zhu Y, Han R, Yin W, Guo C et al (2018a) Expression of Vitis amurensis VaERF20 in Arabidopsis thaliana improves resistance to Botrytis cinerea and Pseudomonas syringae pv. Tomato DC3000. Int J Mol Sci 19. https://doi.org/10.3390/ijms19030696
Wang X, Tu M, Wang D, Liu J, Li Y et al (2018b) CRISPR/Cas9-mediated efficient targeted mutagenesis in grape in the first generation. Plant Biotechnol J 16:844–855. https://doi.org/10.1111/pbi.12832
Wang FP, Zhao PP, Zhang L, Zhai H, Du YP (2019) Functional characterization of WRKY46 in grape and its putative role in the interaction between grape and phylloxera (Daktulosphaira vitifoliae). Hortic Res 6. https://doi.org/10.1038/s41438-019-0185-8
Wang Y, Zhang R, Liang Z, Li S (2020a) Grape-RNA: a database for the collection, evaluation, treatment, and data sharing of grape RNA-seq datasets. Genes (basel) 11(3):351. https://doi.org/10.3390/genes11030315
Wang L, Ahmad B, Liang C, Shi X, Sun R et al (2020b) Bioinformatics and expression analysis of histone modification genes in grapevine predict their involvement in seed development, powdery mildew resistance, and hormonal signaling. BMC Plant Biol 20:412. https://doi.org/10.1186/s12870-020-02618-7
Wang L, Liu W, Wang Y (2020c) Heterologous expression of Chinese wild grapevine VqERFs in Arabidopsis thaliana enhance resistance to Pseudomonas syringae pv. tomato DC3000 and to Botrytis cinerea. Plant Sci 293:110421. https://doi.org/10.1016/j.plantsci.2020.110421
Wang X, Tu M, Wang Y, Yin W, Zhang Y et al (2021) Whole-genome sequencing reveals rare off-target mutations in CRISPR/Cas9-edited grapevine. Hortic Res 8:114. https://doi.org/10.1038/s41438-021-00549-4
Welter LJ, Göktürk-Baydar N, Akkurt M, Maul E, Eibach R et al (2007) Genetic mapping and localization of Quantitative Trait Loci affecting fungal disease resistance and leaf morphology in grapevine (Vitis vinifera L). Mol Breed 20:359–374. https://doi.org/10.1007/s11032-007-9097-7
Welter LJ, Grando SM, Zyprian EM (2011) Basics of grapevine genetic analysis. In: Adam-Blondon A-F, Martinez-Zapater JM, Kole C (eds) Genetics, genomics, and breeding of grapes. CRC Press, Boca Raton, FL, pp 165–187. https://doi.org/10.1201/b10948-11
Wen Z, Yao L, Singer SD, Muhammad H, Li Z et al (2017) Constitutive heterologous overexpression of a TIR-NB-ARC-LRR gene encoding a putative disease resistance protein from wild Chinese Vitis pseudoreticulata in Arabidopsis and tobacco enhances resistance to phytopathogenic fungi and bacteria. Plant Physiol Biochem 112:346–361. https://doi.org/10.1016/j.plaphy.2017.01.017
Wermelinger B, Candolfi MP, Baumgärtner J (1992) A model of the European red mite (Acari, Tetranychidae) population dynamics and its linkage to grapevine growth and development. J Appl Entomol 114:155–166. https://doi.org/10.1111/j.1439-0418.1992.tb01110.x
Wiedemann-Merdinoglu S, Prado E, Coste P, Dumas V, Butterlin G et al (2006) Genetic analysis of resistance to downy mildew from Muscadinia rotundifolia. In: 9th International conference on grape genetics and breeding, Udine, Italy
Wilcox WF, Gubler WD, Uyemoto JK (2015) Diseases caused by biotic factors. In: Wilcox WF, Gubler WD, Uyemoto JK (eds) Compendium of grape diseases. APS Press, St. Paul, MN, pp 17–146
Wilcox WF, Hoffman LE (2019) Black rot. In: Wilcox WF, Gubler WD, Uyemoto JK (eds) Compendium of grape diseases, disorders, and pests. APS Press, St. Paul, MN, pp 28–33
Wilcox WF, Mahaffee W, Gubler WD (2019a) Botrytis bunch rot and blight. In: Wilcox WF, Gubler WD, Uyemoto JK (eds) Compendium of grape diseases, disorders, and pests. APS Press, St. Paul, MN, pp 39–45
Wilcox WF, Ellis MA, Rawnsley B, Rossman A, Pscheidt J (2019b) Phomopsis cane and leaf spot. In: Wilcox WF, Gubler WD, Uyemoto JK (eds) Compendium of grape diseases, disorders, and pests. APS Press, St. Paul, MN, pp 68–71
Williams BR, Edwards CE, Kwasniewski MT, Miller AJ (2020) Epigenomic patterns reflect irrigation and grafting in the grapevine clone. bioRxiv. https://doi.org/10.1101/2020.09.09.290072
Wilmink J, Breuer M, Forneck A (2021a) Grape phylloxera genetic structure reveals root–leaf migration within commercial vineyards. Insects 12:697. https://doi.org/10.3390/insects12080697
Wilmink J, Breuer M, Forneck A (2021b) Effect of temperature on host plant‐specific leaf‐ and root‐feeding performances: a comparison of grape phylloxera biotypes C and G. Entomol Exp Appl eea.13102. https://doi.org/10.1111/eea.13102
Wilson H, Daane KM (2017) Review of ecologically-based pest management in California vineyards. Insects 8:1–13. https://doi.org/10.3390/insects8040108
Wong DCJ, Sweetman C, Drew DP, Ford CM (2013) VTCdb: a gene co-expression database for the crop species Vitis vinifera (grapevine). BMC Genomics 14:1–17. https://doi.org/10.1186/1471-2164-14-882
Wu J, Zhang Y, Zhang H, Huang H, Folta KM et al (2010) Whole genome wide expression profiles of Vitis amurensis grape responding to downy mildew by using Solexa sequencing technology. BMC Plant Biol 10:234. https://doi.org/10.1186/1471-2229-10-234
Wu J, Folta KM, Xie Y, Jiang W, Lu J et al (2017) Overexpression of Muscadinia rotundifolia CBF2 gene enhances biotic and abiotic stress tolerance in Arabidopsis. Protoplasma 254:239–251. https://doi.org/10.1007/s00709-015-0939-6
Würschum T (2012) Mapping QTL for agronomic traits in breeding populations. Theor Appl Genet 125:201–210. https://doi.org/10.1007/s00122-012-1887-6
Wyss U (2014) Xiphinema index, maintenance and feeding in monoxenic cultures. In: Maramorosch K, Mahmood F (eds) Rearing animal and plant pathogen vectors. CRC Press, Boca Raton, FL, pp 235–267. https://doi.org/10.1201/b16804-14
Xie H, Konate M, Sai N, Tesfamicael KG, Cavagnaro T et al (2017) Global DNA methylation patterns can play a role in defining terroir in grapevine (Vitis vinifera cv. Shiraz). Front Plant Sci 8:01860. https://doi.org/10.3389/fpls.2017.01860
Xiong J-S, Ding J, Li Y (2015) Genome-editing technologies and their potential application in horticultural crop breeding. Hortic Res 2:15019. https://doi.org/10.1038/hortres.2015.19
Xu K, Riaz S, Roncoroni NC, Jin Y, Hu R et al (2008) Genetic and QTL analysis of resistance to Xiphinema index in a grapevine cross. Theor Appl Genet 116:305–311. https://doi.org/10.1007/s00122-007-0670-6
Xu Y, Wang Y (2014) Introduction of application of Chinese wild grapes species. Acta Hortic 1046:49–56. https://doi.org/10.17660/ActaHortic.2014.1046.4
Xu W, Ma F, Li R, Zhou Q, Yao W et al (2019) VpSTS29/STS2 enhances fungal tolerance in grapevine through a positive feedback loop. Plant Cell Environ 42:2979–2998. https://doi.org/10.1111/pce.13600
Yan X, Qiao H, Zhang X, Guo C, Wang M, et al. (2017) Analysis of the grape (Vitis vinifera L.) thaumatin-like protein (TLP) gene family and demonstration that TLP29 contributes to disease resistance. Sci Rep 7:4269. https://doi.org/10.1038/s41598-017-04105-w
Yang S, Fresnedo-Ramírez J, Wang M, Cote L, Schweitzer P et al (2016) A next-generation marker genotyping platform (AmpSeq) in heterozygous crops: a case study for marker-assisted selection in grapevine. Hortic Res 3:16002. https://doi.org/10.1038/hortres.2016.2
Yang Y, Hu X, Liu P, Chen L, Peng H, et al. (2021) A new root-knot nematode, Meloidogyne vitis sp. nov. (Nematoda: Meloidogynidae), parasitizing grape in Yunnan. PLoS ONE 16:e0245201. https://doi.org/10.1371/journal.pone.0245201
Yin K, Gao C, Qiu J-L (2017) Progress and prospects in plant genome editing. Nat Plants 3:17107. https://doi.org/10.1038/nplants.2017.107
Yin L, Clark MD, Burkness EC, Hutchison WD (2019) Grape phylloxera (Hemiptera: Phylloxeridae), on cold-hardy hybrid wine grapes (Vitis spp.): a review of pest biology, damage, and management practices. J Integr Pest Manag 10:1–9. https://doi.org/10.1093/jipm/pmz011
Yin L, Karn A, Cadle-Davidson L, Zou C, Underhill A et al (2021) Fine mapping of leaf trichome density revealed a 747-kb region on chromosome 1 in cold-hardy hybrid wine grape populations. Front Plant Sci 12:1–15. https://doi.org/10.3389/fpls.2021.587640
Young ND (1996) QTL mapping and quantitative disease resistance in plants. Annu Rev Phytopathol 34:479–501. https://doi.org/10.1146/annurev.phyto.34.1.479
Yu J, Pressoir G, Briggs WH, Vroh Bi I, Yamasaki M et al (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208. https://doi.org/10.1038/ng1702
Yu Y, Xu W, Wang J, Wang L, Yao W et al (2013) The Chinese wild grapevine (Vitis pseudoreticulata) E3 ubiquitin ligase Erysiphe necator-induced RING finger protein 1 (EIRP1) activates plant defense responses by inducing proteolysis of the VpWRKY11 transcription factor. New Phytol 200:834–846. https://doi.org/10.1111/nph.12418
Yu YH, Jiao ZL, Bian L, Wan YT, Yu KK et al (2019a) Overexpression of Vitis vinifera VvbZIP60 enhances Arabidopsis resistance to powdery mildew via the salicylic acid signaling pathway. Sci Hortic 256:1086402. https://doi.org/10.1016/j.scienta.2019.108640
Yu Y, Bian L, Jiao Z, Yu K, Wan Y et al (2019b) Molecular cloning and characterization of a grapevine (Vitis vinifera L.) serotonin N-acetyltransferase (VvSNAT2) gene involved in plant defense. BMC Genomics 20:1–13. https://doi.org/10.1186/s12864-019-6085-3
Yun HK, Park KS, Rho JH, Choi YJ, Kang KK (2006) Evaluating the resistance of grapevines against anthracnose by pathogen inoculation, vineyard inspection, and bioassay with culture filtrate from Elsinoe ampelina. J Am Pomol Soc 60:97–103
Zanettin G, Bullo A, Pozzebon A, Burgio G, Duso C (2021) Influence of vineyard inter-row groundcover vegetation management on arthropod assemblages in the vineyards of North-Eastern Italy. Insects 12:349. https://doi.org/10.3390/insects12040349
Zdunić G, Lukšić K, Nagy ZA, Mucalo A, Hančević K et al (2020) Genetic structure and relationships among wild and cultivated grapevines from Central Europe and part of the Western Balkan peninsula. Genes (basel) 11:962. https://doi.org/10.3390/genes11090962
Zecca G, Abbott JR, Sun WB, Spada A, Sala F et al (2012) The timing and the mode of evolution of wild grapes (Vitis). Mol Phylogenet Evol 62:736–747. https://doi.org/10.1016/j.ympev.2011.11.015
Zecca G, Labra M, Grassi F (2020) Untangling the evolution of American wild grapes: admixed species and how to find them. Front Plant Sci 10:1814. https://doi.org/10.3389/fpls.2019.01814
Zeilinger AR, Turek D, Cornara D, Sicard A, Lindow SE et al (2018) Bayesian vector transmission model detects conflicting interactions from transgenic disease-resistant grapevines. Ecosphere 9(11):e02494. https://doi.org/10.1002/ecs2.2494
Zendler D, Schneider P, Töpfer R, Zyprian E (2017) Fine mapping of Ren3 reveals two loci mediating hypersensitive response against Erysiphe necator in grapevine. Euphytica 213:68. https://doi.org/10.1007/s10681-017-1857-9
Zendler D, Malagol N, Schwandner A, Töpfer R, Hausmann L, Zyprian E (2021a) High-throughput phenotyping of leaf discs infected with grapevine downy mildew using shallow convolutional neural networks. Agronomy 11(9):1768. https://doi.org/10.3390/agronomy11091768
Zendler D, Töpfer R, Zyprian E (2021b) Confirmation and fine mapping of the resistance locus Ren9 from the grapevine cultivar ‘Regent.’ Plants 10:1–20. https://doi.org/10.3390/plants10010024
Zhang J, Hausmann L, Eibach R, Welter LJ, Töpfer R et al (2009) A framework map from grapevine V3125 (Vitis vinifera ‘Schiava grossa’ × ‘Riesling’) × rootstock cultivar ‘Börner’ (Vitis riparia × Vitis cinerea) to localize genetic determinants of phylloxera root resistance. Theor Appl Genet 119:1039–1051. https://doi.org/10.1007/s00122-009-1107-1
Zhang J, Zhou J-M (2010) Plant immunity triggered by microbial molecular signatures. Mol Plant 3:783–793. https://doi.org/10.1093/mp/ssq035
Zhang K, Zhang N, Cai L (2013) Typification and phylogenetic study of Phyllosticta ampelicida and P. vaccinii. Mycologia 105:1030–1042. https://doi.org/10.3852/12-392
Zhang W, Manawasinghe IS, Zhao W, Xu J, Brooks S et al (2017) Multiple gene genealogy reveals high genetic diversity and evidence for multiple origins of Chinese Plasmopara viticola population. Sci Rep 7:1–11. https://doi.org/10.1038/s41598-017-17569-7
Zhang Y, Yao JL, Feng H, Jiang J, Fan X et al (2019) Identification of the defense-related gene VdWRKY53 from the wild grapevine Vitis davidii using RNA sequencing and ectopic expression analysis in Arabidopsis. Hereditas 156:14. https://doi.org/10.1186/s41065-019-0089-5
Zhang Y, Fan X, Li Y, Sun H, Jiang J et al (2020) Restriction site-associated DNA sequencing reveals the molecular genetic diversity of grapevine and genes related to white rot disease. Sci Hortic 261:108907. https://doi.org/10.1016/j.scienta.2019.108907
Zhao H, Zhao K, Wang J, Chen X, Chen Z et al (2015) Comprehensive analysis of dicer-like, argonaute, and RNA-dependent RNA polymerase gene families in grapevine (Vitis vinifera). J Plant Growth Regul 34:108–121. https://doi.org/10.1007/s00344-014-9448-7
Zhao C, Rispe C, Nabity PD (2019a) Secretory RING finger proteins function as effectors in a grapevine galling insect. BMC Genomics 20:1–12. https://doi.org/10.1186/s12864-019-6313-x
Zhao C, Zhang Y, Du J, Guo X, Wen W et al (2019b) Crop phenomics: current status and perspectives. Front Plant Sci 10:714. https://doi.org/10.3389/fpls.2019.00714
Zhou X, Stephens M (2014) Efficient multivariate linear mixed model algorithms for genome-wide association studies. Nat Methods 11:407–409. https://doi.org/10.1038/nmeth.2848
Zhou Y, Massonnet M, Sanjak JS, Cantu D, Gaut BS (2017) Evolutionary genomics of grape (Vitis vinifera ssp. vinifera) domestication. Proc Natl Acad Sci USA 114(44):11715–11720. https://doi.org/10.1073/pnas.1709257114
Zhou J, Li D, Wang G, Wang F, Kunjal M et al (2020) Application and future perspective of CRISPR/Cas9 genome editing in fruit crops. J Integr Plant Biol 62:269–286. https://doi.org/10.1111/jipb.12793
Zhu YM, Hoshino Y, Nakano M, Takahashi E, Mii M (1997) Highly efficient system of plant regeneration from protoplasts of grapevine (Vitis vinifera L.) through somatic embryogenesis by using embryogenic callus culture and activated charcoal. Plant Sci 123:151–157. https://doi.org/10.1016/S0168-9452(96)04557-8
Zhu C, Gore M, Buckler ES, Yu J (2008) Status and prospects of association mapping in plants. Plant Genome 1(1):5–20. https://doi.org/10.3835/plantgenome2008.02.0089
Zhu Z, Shi J, Cao J, He M, Wang Y (2012) VpWRKY3, a biotic and abiotic stress-related transcription factor from the Chinese wild Vitis pseudoreticulata. Plant Cell Rep 31:2109–2120. https://doi.org/10.1007/s00299-012-1321-1
Zhu X, Jiu S, Li X, Zhang K, Wang M et al (2018) In silico identification and computational characterization of endogenous small interfering RNAs from diverse grapevine tissues and stages. Genes Genomics 40:801–817. https://doi.org/10.1007/s13258-018-0679-z
Zilberman D (2017) An evolutionary case for functional gene body methylation in plants and animals. Genome Biol 18:87. https://doi.org/10.1186/s13059-017-1230-2
Zinelabidine LH, Cunha J, Eiras-Dias JE, Cabello F, Martinez-Zapater JM et al (2015) Pedigree analysis of the Spanish grapevine cultivar “Hebén.” VITIS J Grapevine Res 54:81–86. https://doi.org/10.5073/vitis.2015.54.special-issue.81-86
Zini E, Dolzani C, Stefanini M, Gratl V, Bettinelli P et al (2019) R-Loci arrangement versus downy and powdery mildew resistance level: a Vitis hybrid survey. Int J Mol Sci 20:3526. https://doi.org/10.3390/ijms20143526
Zipfel C (2008) Pattern-recognition receptors in plant innate immunity. Curr Opin Immunol 20:10–16. https://doi.org/10.1016/j.coi.2007.11.003
Zou C, Karn A, Reisch BI, Nguyen A, Sun Y et al (2020) Haplotyping the Vitis collinear core genome with rhAmpSeq improves marker transferability in a diverse genus. Nat Commun 11:413. https://doi.org/10.1038/s41467-019-14280-1
Zuo J, Niu Q, Møller SG, Chua N (2001) Chemical-regulated, site-specific DNA excision in transgenic plants. Nat Biotechnol 19:157–161. https://doi.org/10.1038/84428
Zyprian E, Töpfer R (2005) Development of microsatellite-derived markers for grapevine genotyping and genetic mapping. NCBI, GeneBank
Zyprian E, Ochßner I, Schwander F, Šimon S, Hausmann L et al (2016) Quantitative Trait Loci affecting pathogen resistance and ripening of grapevines. Mol Genet Genomics 291(4):1573–1594. https://doi.org/10.1007/s00438-016-1200-5
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Vezzulli, S. et al. (2022). Genomic Designing for Biotic Stress Resistant Grapevine. In: Kole, C. (eds) Genomic Designing for Biotic Stress Resistant Fruit Crops. Springer, Cham. https://doi.org/10.1007/978-3-030-91802-6_4
Download citation
DOI: https://doi.org/10.1007/978-3-030-91802-6_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-91801-9
Online ISBN: 978-3-030-91802-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)