Abstract
Key message
We have identified 19 QTLs for rachis architecture, a key and complex trait for grapevine production. Fifty out of 1,173 genes underlying these QTLs are candidates to be further explored.
Abstract
In the table grape industry, the rachis architecture has economic and management implications. Therefore, understanding the genetics of this trait is key for its breeding. The aim of this work was to identify genetic determinants of traits associated with the cluster architecture. Characterisations of eight traits was performed on a ‘Ruby Seedless’ × ‘Sultanina’ crossing (F1: n = 137) during three seasons, with and without gibberellic acid (GA3) applications. The genotypic effects and the genotype × GA3 interactions were significant for several traits. Rachis length (rl), lateral shoulder length and node number along the central axis were the most prominent traits. On average, the heritability of these traits was ~71 %, with heritability of rl being 76 % as estimated under different seasons. Quantitative trait loci (QTLs) analyses showed that linkage group 5 (LG5) and LG18 harboured the largest number of QTLs for these traits. According to the variance explained, the main QTL (corresponding to rl) was found on LG9. These QTLs were supported mainly by a paternal additive effect and revealed possible pleiotropic effects. Based on the grapevine reference genome, we identified 1,173 genes located under these QTL confidence intervals. Fifty of the 891 annotated genes of this list were selected for their further characterisation because of their possible participation in the rachis architecture. In conclusion, the QTLs detected indicate that these traits and their GA3 responsiveness have a clear genetic basis. Due to the percentage of the total variance explained, they are good candidates to participate in the genetic determination of the cluster architecture.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genetic determinism of the morphology of some plant organs has not received enough attention. In the case of fruit shape, there are plenty of papers describing their genetic and hormonal control in species such as tomato, Cucumis and capsicum (Peterson 1959; Grandillo et al. 1999; Chaim et al. 2001; Périn et al. 2002; Tanksley 2004). However, in species such as grapevine the fruits or berries are arranged in clusters, which in turn correspond to inflorescences. The architecture of the inflorescence determines to a large extent the fruit quality. Several authors have presented evidence showing that both the architecture and size of the inflorescences are genetically determined (Shavrukov et al. 2004; Weiss et al. 2005; Busov et al. 2008). This phenomenon has been demonstrated by studies in pea, rice, Arabidopsis and recently in rose (Nomura et al. 1997; García-Maroto et al. 2000; Nakagawa et al. 2002; Okada and Shimura, 1994; Goosey and Sharrock 2001; Kawamura et al. 2011). In general, the inflorescence architecture depends on branching pattern, number of flowers per node and the extent of terminal meristem growth in the reproductive phase where in turn developmental decisions occur (Singer 2007; Liu et al. 2013). Several QTLs associated with inflorescence architecture have been found in a number of species: Poaceae (Zhu et al. 1999; Brown et al. 2006; Upadyayula et al. 2006; Larson et al. 2013), Arabidopsis (Ungerer et al. 2002, 2003) and rose (Kawamura et al. 2011). In Arabidopsis, shoot identity gene TERMINAL FLOWER1 (TFL1) and floral meristem identity genes, such as APETALA1 (AP1) and LEAFY (LFY), regulate the pattern of inflorescence branching (Liu et al. 2013). Liu et al. (2013) have reported a conserved genetic pathway determining inflorescence architecture in Arabidopsis and rice and demonstrated that MADS-box genes in rice determine panicle branching by regulating TFL1-like genes.
In modern viticulture, the architecture of the cluster plays an important role in vineyard management and modification. According to several authors, the architecture of this organ has an important impact on the cost of grape production (Weaver and McCune 1959; Weaver et al. 1962; Vail and Marois 1991; Vail et al. 1998; Shavrukov et al. 2004). Many methods are used to modify the cluster architecture. For example, the pre-bloom application of gibberellins is used to stimulate the elongation of the main axis of the rachis, which results in less compact clusters (Weaver and McCune 1959; Weaver et al. 1962; Hed et al. 2011). Chemical thinning of the berries and removal of parts of the cluster are other methods used to reduce the number of berries per cluster, to improve the shape and to decrease its compaction. These modifications help to increase the size of the remaining berries, which is an important quality factor in table grape production. These measures also help prevent the attack of fungi such as Botrytis cinerea (Vail et al. 1998; Shavrukov et al. 2004; Hed et al. 2011).
The rachis is the dorsal spine of the cluster, which comprises a main axis and several lateral branches (shoulders). The shoulders are further divided into secondary and tertiary branches. The last branch sub-division corresponds to the pedicel, the structure that harbours each individual berry. At flowering time, this structure forms an inflorescence called the panicle (Pratt 1971). The last branch sub-division corresponds to the pedicel, the structure that harbours each individual berry. It has been proposed that genotype is an important factor in determining the size and the architecture of the grapevine cluster (Shavrukov et al. 2004). However, research on the cluster architecture and its genetic variation is currently scarce (Shavrukov et al. 2004; Fernandez et al. 2010). In fact, there have been no systematic studies undertaken to identify the defining characters and the genetic determinants of the grapevine cluster architecture.
The study of genetic variation (i.e. genetic variance or genetic diversity) and the heritability of traits of interest are important to define whether an improvement is possible and what materials to use to achieve a better outcome (Poehlman 1986; Falconer and Mackay 1996). An interesting approach to study the genetic structure of any trait is the use of quantitative trait loci (QTLs), which are segments of the genome where there exists a higher probability of finding genes that are responsible for traits of interest (Collard et al. 2005). QTL study and the observation of associated allelic diversity (molecular markers) have accelerated the identification of those genotypes with higher probabilities of expressing the desired traits. This practice is particularly useful when the trait of interest cannot be measured because it has not yet been expressed (for example, reproductive traits in the seedling stage) or it is too difficult to evaluate in a consistent way (as is the case with disease resistance) (Collard et al. 2005; Costantini et al. 2007). One of the main objectives of the use of molecular markers in applied research in agriculture has been the construction of genetic maps for several species of economic interest. Genetic maps have been used to identify QTLs related to traits of interest (Collard et al. 2005). QTL mapping is based on the principle that genes and markers segregate by recombination (“crossing over”) during meiosis, affecting the variation or the variance of quantitative traits in the resulting progeny or segregating population (where there is a mixture of parental and recombinant genotypes), allowing analysis by statistical approaches (Mackay 2001; Collard et al. 2005). Many QTLs for several traits have been identified in different grapevine progenies. The main interest in these studies has been to identify the genetic determinants related to traits associated with disease and pest resistance (Marino et al. 2003; Zyprian et al. 2003; Doucleff et al. 2004; Fischer et al. 2004; Krivanek et al. 2006; Xu et al. 2008). Some effort has been made to identify genetic determinants associated with the quality of the berry (berry size, berry seedlessness) (Doligez et al. 2002; Zyprian et al. 2003; Fischer et al. 2004; Fanizza et al. 2005; Mejía et al. 2007; Costantini et al. 2008; Cabezas et al. 2006). There have also been studies undertaken on other berry quality attributes, such as aroma profile and anthocyanin content (Eibach et al. 2003; Fournier-Level et al. 2009), fertility-associated traits (Doligez et al. 2010) and phenological stages (Costantini et al. 2008; Duchêne et al. 2012), among others. As very little is known about the genetic determinism of cluster architecture in the grapevine, the aim of this work was to analyse the genetic variation of traits associated with this organ architecture. The main traits analysed were rachis and shoulder length, rachis fresh weight and main axis internode number. These studies were performed using progeny derived from table grape varieties (n = 137) evaluated during three seasons. We compared clusters grown on plants receiving gibberellic acid at flowering time to those not receiving this hormone, which is used commercially for cluster enlargement in table grape production. The traits selected showed a clear genetic basis and can be considered good candidates for determinants of cluster architecture due to their importance in determining the genotypic variance.
Materials and methods
Plant material and treatments
This study was performed during three growing seasons (2009–2010 to 2011–2012) at the Chilean Institute of Agriculture Research, La Platina Center (33°34′20″S; 70°37′32″W; 630 m.o.s.l.), on F1 progeny (n = 137 segregants) of a controlled cross of ‘Ruby Seedless’ × ‘Sultanina’ (R × S) and obtained through embryo rescue. Note that each season begins approximately in September of year 1 and ends in May of year 2. These progenies were cultivated in an overhead trellis system (pergola type), with three plants per segregant grafted in 2006 onto adult plants of the ‘Sultanina’ variety that were used as rootstocks. Watering, fertilisation and pest/disease control followed standard conditions used in a commercial orchard. For every genotype, three inflorescences were treated by immersion before flowering (panicle length: 12–15 cm) with a solution of 10 mg L−1 of gibberellic acid (GA3) (GibGro® 20 %) plus 0.1 mL L−1 of siliconate coadjuvant (Break SL®). An equivalent number of control clusters were treated with water plus coadjuvant.
Phenotypic evaluation
To facilitate the different measurements, phenotypic evaluations were performed 2 weeks before veraison—the point at which the rachis stops its elongation (Coombe 1995). We based the phenotyping on Costes (2003) and Segura et al. (2006, 2007) works dealing with apple tree architecture and on Shavrukov et al. (2004) related to cluster architecture in grapevine. Immediately after the harvest of samples, all the traits were measured manually with the exception of rachis fresh weight and berry fresh weight, which were measured with an electronic balance. The latter was used to estimate the total number of berries per cluster and per shoulder by using the fresh weight of 50 berries. A semicircular protractor was used to measure the angles in degrees (°) between the main axis and the first and second shoulder. Peduncle diameter was measured by a vernier calliper. In summary, 23 traits and subtraits were measured in treated and control clusters during the first season (Online Resource 1 and Fig. 1). A multivariate factor analysis based on the 23 traits evaluated was performed, using principal components as the method of factor extraction, percentage of variance explained and latent root as criteria of factor extraction, and VARIMAX orthogonal rotation to interpret the factorial results. With this analysis, subsets of co-varying variables were extracted in a few conceptual and orthogonal variables called factors (Acquaah et al. 1992). To supplement this exploration, the relationships among these traits and among the genotypes were analysed by a correlogram according to Pearson’s correlation coefficient and cluster analysis, respectively. The average linkage method and the Euclidean distance measure were used according to the results of the cophenetic correlation coefficient to complete the cluster analysis. Genotypic effects and genotype × GA3 interaction were calculated and used for this purpose; analysis of variance with mixed linear models was employed, as shown below.
Schematic diagram representing the architecture of cluster at fruit set of Vitis vinifera L. 1ir, 2ir and 3ir: first, second and third internode of the main axis or rachis, respectively; 1is, 2is and 3is: first, second and third internode of the second shoulder, respectively; a: angle between the main axis and the second shoulder
During the second (2010–2011) and third (2011–2012) seasons, the multivariate first-factor and GA3 responsive traits were measured according to multivariate factor analysis as indicated for the first season. To test the effect of each genotype on each trait given by genotypic variance (random genotypic effect) and their possible interaction with the GA3 treatment (fixed effect) at each season, mixed linear models were proposed. The genotypic effect was defined according to a linear additive model:
where y ijk is the phenotypic value measured for the trait y on the cluster k of the genotype i with treatment j; μ corresponds to the overall mean; g i is the random effect of genotype i representing the effect of each genotype or genotypic effect on trait y; t j is the fixed effect of GA3 treatment with two levels (0 and 10 mg L−1); (g × t) ij is the random interaction between genotype i and treatment j; and ε ijk is the random residual error per cluster k of the genotype i with treatment j. The significance of the effects was assessed using the Akaike and Bayesian criteria and tested by likelihood ratio (Sillampää and Corander 2002; Segura et al. 2007). Broad-sense heritability (h 2) was estimated according to REML variance components according to Holland et al. (2003), Kawamura et al. (2011) and Duchêne et al. (2012).
The seasonal effect on phenotypic expression of the selected traits treatment was analysed by repeated measurement analysis of variance using the plant as the experimental unit (subject factor) and considering the genotype × season interaction.
The complete set of analyses was performed using the statistical software R (R Development Core Team 2012) and its corrplot (Wei 2011) and lme4 (Bates and Maechler 2009) packages for correlogram and linear mixed models, respectively.
Genetic evaluations
The Chilean Institute of Agriculture Research (INIA) developed a genetic map using a segregating population of R × S consistent in 137 siblings, with the aim of studying the genetics of various quality traits. The consensus genetic map was based on 272 markers (SSRs or microsatellites, AFLPs, gene-based SNPs, phenotypic marker and SCARs), with SSRs previously mapped in other crossings being the most abundant (Online Resource 2). SSRs and AFLPs were analysed by silver-stained polyacrylamide denaturant gel electrophoresis, as described by Mejía et al. (2007). Gene-based SNPs and their alleles identification was done by silver-stained SSCP gels as described by Guerrero (2007). SCARs were designed after RAPD markers and evaluated by agarose gels as described (Mejía et al. 2007). Nomenclature and primer sequences were obtained from the National Center for Biotechnology Information (NCBI) and Sefc et al. (1999), Di Gaspero et al. (2000) and Merdinoglu et al. (2005). Other newly developed SSR markers were designed from the Vitis vinifera PN40024 reference genome sequence (Jaillon et al. 2007). The genetic/linkage map of R × S was built as described by Mejía et al. (2007).
QTLs of the phenotypic means of each selected trait under each GA3 treatment, as well as some of their relationships with each other (Table 1), were identified using the consensus map via nonparametric Kruskal–Wallis test with a stringent significance level (p value) of at least 0.005. Interval mapping was followed by the multiple-QTL model (MQM) procedure using the MAP QTL 4.0 software (van Ooijen et al. 2002). The LOD genome-wide error thresholds used to declare the existence of a QTL (significance level of 0.05) were determined with the same software using 1,000 permutations. Interval mapping analysis was performed with a step size of 1 centiMorgan (cM) to find regions with potential QTLs, i.e., where the LOD score was greater than LOD thresholds. For each putative QTL given by interval mapping, the closest markers to the peak of the LOD profile were tested using the automatic cofactor selection procedure to select cofactors significantly associated with each trait at a p value of <0.02. Markers accepted as co-factors were used to perform a MQM procedure with a step size of 1 cM. Each significant QTL was characterised by its maximum LOD score, its percentage of explained variation and its confidence intervals in cM corresponding to the maximum LOD score −1 and −2 units on either side of the LOD peak. The location of each QTL detected on each linkage group in the consensus map was plotted using the program MapChart (Voorrips 2002). Allelic effects of QTLs were calculated according to Segura et al. (2007).
Search for candidate genes
In each QTL, the search for candidate genes was done in the genomic region corresponding to the confidence interval determined on the consensus map. The scrutinised sequence was limited by the most proximal SSR or SNP markers that were present both in the reference genome and in the consensus map. The genes were selected based on the information available for the annotated reference genome (Genoscope 12×) of the quasi-homozygous line 40024 derived from Pinot Noir (http://www.genoscope.cns.fr/externe/GenomeBrowser/Vitis/; Jaillon et al. 2007). The genes were classified according to their biological function registered in the database. The genes catalogued as with “unknown function” or equivalent were not considered for further analyses. In addition, a gene ontology (GO) enrichment analysis was performed considering the genes identified in the physical genomic regions associated with the confidence interval for each QTL, comparing the frequency found in each QTL vs. the complete reference genome and searching for a possible enrichment in gene functions that could be related to cluster architecture. All enrichment analyses were done with the agriGO tool (http://bioinfo.cau.edu.cn/agriGO), with the options singular enrichment analysis and complete GO. Significant GO terms (p value <0.05) were calculated using hypergeometric distribution and Yekutieli multi-test adjustment method (Du et al. 2010).
Results
During the 2009–2010, 2010–2011 and 2011–2012 seasons, a phenotypic and genetic characterisation of traits related to cluster architecture was carried out on a R × S (F1: n = 142) crossing conducted in overhead trellis systems. Only 137 of the 140 segregants of this progeny were analysed, according to the fertility of the plants. In the 2009–2010 season, a phenotypic characterisation of the 23 traits was performed with the purpose of exploring the cluster architecture, including analysis of genotypic and genotype × GA3 interaction effects, estimation of trait heritability and analysis of both the relationships among traits and relationships among the genotypes (correlation and cluster analysis respectively). During the following two seasons, only eight traits were evaluated, according to a selection based on their correlation pattern. Furthermore, to decipher their underlying genetic structure, a QTL mapping and analysis was carried out.
Phenotypic and genotypic variation
During the first season of evaluations, a trait selection was carried out according to the correlations among the 23 traits measured (Online Resource 1) to simplify the experimental procedures and measurements during the following seasons. According to mixed linear models, the phenotypes of all characters had a significant genotypic effect. Only a few of the measured traits showed response to GA3 (rw, rl, s2l and tb) (Online Resource 1). From these results, the genotypic effect of each genotype on each of the traits and the effect of genotype × GA3 were computed via the ranef() function of lme4 package of R after treatment by the best linear unbiased prediction (BLUP).
Relationships among traits
The Pearson correlation for the genotypic effects of the traits associated with the architecture of the cluster showed several correlations of high value (r > 0.6). The most important finding corresponded to the correlations among the lengths of the first and the second shoulders, the rachis fresh weight, the total number of berries per cluster and the number of nodes of the central axis (r > 0.75). In the case of genotype × GA3 effects, these variables were correlated except for the interaction given for the total number of berries. These results are summarised in a correlogram (Fig. 2) that shows a characteristic pattern of correlations between the different traits.
Correlogram for the different traits evaluated for cluster architecture of Vitis vinifera L. The correlogram was based on the Pearson’s correlation coefficient (r) of BLUP values of genotype effects of the 23 traits and the interaction genotype × GA3 of rachis fresh, rachis length, second shoulder length and total number of berries per cluster (rw GA, rl GA, s2l GA and tb GA, respectively). Each trait was ordered in terms of its eigenvalues obtained from principal component analysis. Each cell of the matrix represents the r value by colours. Right bar indicates the r values. Red negative value (−1 ≤ r < 0), blue positive value (0 < r ≤ 1), white values near to 0 (r ~ 0) (colour figure online)
To analyse these correlations, a multivariate factor analysis was performed. With respect to this analysis, 27 factors were obtained as an initial solution, of which the first five factors explained ~60 % of the total variance (up to ~90 % if considering the first 16 principal factors). According to the percentage of variance explained and latent root criteria, nine principal factors were extracted. After the VARIMAX rotation, the first nine principal factors explained 77.1 % of the total variance. The characteristic of each of these factors is shown in Online Resource 3.
The first major factor (Factor 1) explained 27.8 % of the total variance and was given (correlated) by the following variables (factor loadings > |0.5|): rachis length (rl), rachis fresh weight (rw), second shoulder length (s2l), first shoulder length (s1l), peduncle diameter (pd), number of nodes of the central axis (nr) and the number of berries on the second shoulder (bs2) (Online Resource 4). This factor accounted for approximately 80 % of the variance of these variables (communalities, Online Resource 4). Because it was more associated with the rw, nr, s2l and pd (factor loadings > |0.7|), this factor was labelled as “Traits related to the cluster size”. Using the same approach, the other factors were named as shown in Online Resource 4.
Relationships among genotypes
With the intention of representing the relationship among the traits, a cluster analysis based on genotypic and interaction effect was carried out. The combination of Euclidean distance with average linkage method had a higher cophenetic correlation coefficient (0.73). Four groups of genotypes (clusters) with at least 75 % of similarity were found. These clusters were as follows: cluster A or ‘Ruby’ type (n = 12 genotypes, including the maternal parent ‘Ruby’); cluster B or ‘Sultanina’ type (n = 91, including the paternal parent ‘Sultanina’); cluster C or ‘Intermediate’ (n = 22); and cluster D or ‘Outlier’ (n = 12). The common phenotypic characteristics of the individuals in each of these groups are summarised in Fig. 3.
Clustering of the ‘Ruby Seedless’ × ‘Sultanina’ progeny based on the different traits evaluated. Four cluster groups were identified and shown by several methods: a by a heat map with the standardised means of random effect (BLUPs) for each trait during the 2009–2010 season; and b by representative genotypes of each group. Clusters without gibberellic acid treatment (−GA3) and treated with 10 mg L−1 of gibberellic acid (+GA3) are shown. Vertical dashed lines are 40 cm long
During the 2010–2011 and 2011–2012 seasons, the traits included in the first factor and the GA3-responsive traits were measured as described for the first season according to multivariate factor analysis. These traits corresponded to rachis fresh weight and length, peduncle diameter, length of the first and second shoulders, number of nodes on the central axis and total number of berries per cluster.
Genotypic variation and heritability across the seasons
As in the first season, the subsequent seasons revealed the importance of genotypic effect and also that not all traits responded to GA3 (Online Resource 1 and Table 2). Additionally, in each season (Table 2), all traits had a significant genotypic variance (p ≤ 0.05), indicating that there is a real effect of different genotypes on the various measured traits (genotype effect). Regarding the relative contribution of genotypic effect as given by the percentages of REML variance, their effect on the different characters was on average medium to low (~39 %) (Online Resource 1 and Table 2). The heritability in turn had medium to high values (h 2 ~ 70 %). The interaction between genotype and GA3 treatment for the responsive traits contributed <20 % of the total variance. All traits showed a clear dependence on the season (Table 3), accounting for about ~35 % of phenotypic variation, regardless of whether they were treated with GA3 or not. This effect was mainly given by the interaction genotype × season, indicating that genotypes respond differentially to environmental conditions. Despite this result, the genotypic effect was significant and was always greater than the effect of the season and its interaction with genotypes. The phenotypic distribution of some traits across seasons and treatments is shown in Fig. 4.
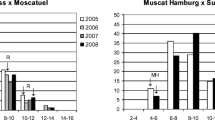
Phenotypic distribution for some traits of cluster architecture of Vitis vinifera L. across seasons. Histograms of phenotypic mean of each genotype are shown. The white boxes indicate the clusters without gibberellic acid treatment (−GA3); the black boxes indicate clusters treated with 10 mg L−1 (+GA3) gibberellic acid. Traits depicted are as follows: a rachis fresh weight (g); b rachis length (cm); c number of nodes of the central axis
Genetic structure
With the purpose of analysing the different genetic determinants associated with the cluster architecture, quantitative trait locus (QTL) analyses of the phenotypic means of each selected trait under each GA3 treatment, as well as some of their relationships measured during the 2009–2010, 2010–2011 and 2011–2012 seasons, were performed.
Genetic mapping
According to these results, a consensus linkage/recombination map was built with the aim of saturating the genome and therefore reducing both the mean intermarker distance and gaps over the different linkage groups (LGs). The main features of the framework map for the progeny R × S are summarised as follow: the consensus map consisted of 272 markers distributed on the 19 V. vinifera linkage groups, covering a total map length of 1,334.4 cM with an average intermarker distance of 4.9 cM. The longest LG corresponded to the LG18, harbouring 40 markers distributed over 136.8 cM (Fig. 5).
QTL analysis
According to nonparametric analysis (Kruskal–Wallis rank sum test per marker) and the MQM procedure, 19 QTLs were detected on LG5, LG8, LG9, LG14, LG17 and LG18, with 6, 2, 1, 3, 1 and 6 QTLs, respectively. These QTLs were grouped in ten bounded genomic regions. Each of these QTLs was closely linked with 15 markers or cofactors (Table 4 and Online Resource 5). On average, these QTLs had LOD values of ~5.3, with the results ranging from 4.1 to 6.6. Their genomic threshold was approximately 4.4. The traits with more QTLs were number of nodes of central axis (nr) and total number of berries per cluster (tb), each one with four QTLs in total, both with and without GA3 treatment. In contrast, only LG5, LG14 and LG18 harboured QTLs showing stability after treatment with GA3 for Peduncle length (pd), nr and tb, respectively. The QTLs detected by this procedure are listed in Table 4, and their confidence intervals are represented in Fig. 5. According to the variance explained, the main QTL (corresponding to rachis length) was found on LG9, close to the microsatellite VMC2D9 locus. On average, these QTLs had a moderate effect, explaining approximately 17 % of the total variance (ranging from 12 to 24 %). The majority of these QTLs were determined mainly by the male additive effect, which almost doubled the female effect. Only two QTLs had an important maternal or female effect, both detected under GA3 treatment. Although, on average, there was no visible difference between the additive and dominance effects, there were some QTLs determined mainly by a dominance effect.
Candidate genes identification
At a first glance, 1,173 genes underlying the 19 detected QTLs were identified (Online Resource 6), a significant fraction of which (282 genes corresponding to 24 %) were catalogued as “unknown protein function” (Online Resource 7). Of these 1,173 genes, 613 potentially coding genes were associated to at least a single GO term (level 3). The genes identified in the physical genomic regions associated with the confidence interval for each of the 10 QTLs for rachis architecture were scrutinised and compared to the frequency found in the complete reference genome for each Gene Ontology term. With this approach, 48 GO categories were overrepresented (Online Resource 8), which suggested a significant increase in their frequency in comparison to the whole reference genome. Indeed, significant enrichments were found in five confidence intervals, encompassing diverse functional categories such as lignin metabolic process, laccase and lyase activities, phenylpropanoid metabolic process as well as coenzyme and cofactor binding. A complete list of significantly enriched GO terms is presented in Online Resource 9. A significantly higher percentage of over-represented GO terms were observed for the gene set included in the LG18-QTL3 confidence interval, raising 19 categories under Biological Process and Molecular Function classifications (Online Resource 10). Therefore, GO analysis provided complementary evidence to support the selection of candidate genes (CGs), which was done on the 891 annotated genes. Of these genes, we selected the 50 most promising ones according to their annotation (Table 5). The selection of these genes was mainly, but not exclusively based on their biological function already described in model plant species such as Arabidopsis, rice and poplar. These genes were unevenly distributed among the 19 QTLs, with 1 to 14 genes per QTL (Table 5). These CGs were classified into six major groups, corresponding to water transport (mainly aquaporins), cell division (cyclins and related genes), cell wall remodelling [including pectinmethylesterase (PME), polygalacturonases (PG), pectate lyase (PE), expansins (EXP), laccase (LAC), among others]; genes induced by or related to hormonal responses of the auxin and gibberellin pathways; transcription factors such as TCP13, AGL8, AGL11, BIM1 and the peptide BEARSKIN1; lipid synthesis, including a phospholipid transporter and two genes directly involved in its synthesis. Finally, a gene related to floral induction (GIGANTEA) located in one of the LG18 QTLs was included in this list of candidate genes because of its possible role in organ (rachis) development.
Discussion
Genotypic effect
In grapevine, in a similar way as has been shown in other species including pea, rice, Arabidopsis and rose (Nomura et al. 1997; García-Maroto et al. 2000; Nakagawa et al. 2002; Okada and Shimura 1994; Goosey and Sharrock 2001; Kawamura et al. 2011), genotype is indicated as an important factor in determining the size and architecture of the inflorescence and later cluster (Boss and Thomas 2000; Shavrukov et al. 2004; Fernandez et al. 2010). The results of this study are consistent with that hypothesis: it was found that the genotypic effect given by genotypic variance was significant for all the traits selected to describe the architecture of the cluster, regardless of the season. Therefore, it would be possible to rely on these traits to improve the architecture of the grapevine cluster either through conventional breeding and/or by marker-assisted selection (MAS). However, the latter method requires the breeder to consider the heritability of the traits of interest. A trait with a low heritability will present an ineffective selection response (Dudley and Moll 1969; Poehlman 1986; Piepho and Möhring 2007). Typically, a heritability value >50 % is considered to be high and a value <20 % is considered to be low (Stanfield 1991). In the grapevine, broad-sense heritability (h 2) has been estimated to be approximately 96 % for the soluble solids content of berry, 88 % for the berry fresh weight and 75 % for the titratable acid of berry juice (Schneider and Staudt 1979). In addition, we have estimated an h 2 of ~92 % (confidence interval 90–96 %) for seed dry weight during the 2009–2010 and 2010–2011 seasons in a particular genetic background (unpublished data). In other perennial plant such as the rose, h 2 for the inflorescence architecture ranged between 82 and 93 % (Kawamura et al. 2011). These values exceed those for the traits associated with the architecture of the grape cluster evaluated here, which had on average an h 2 of ~65 % (Online Resource 1 and Table 2). The central axis length (rl), however, had a value of 75 %; this trait could therefore be useful for strategic breeding. In contrast, the medium to low heritability found for some traits would complicate their use in selection and improvement, especially for those with very low genetic contribution, such as first and second shoulder angle (percentage of variance explained by genotypic effect <15 %). These h 2 magnitudes and ranges were similar to those found by Segura et al. (2007) for architectural traits of the apple tree. The broad ranges found here most likely result from the low number of cluster replicates. Season effect made an important contribution to the phenotype of these traits, on average accounting for more than 30 % of the observed phenotype. This contribution was particularly significant for rachis fresh weight (rw) and number of nodes of central axis (rn), for which this effect was as important as genotypic effect (Table 3).
Gibberellic acid response
The results with regard to possible interaction of genotype with GA3 were determined by linear mixed models (Online Resource 1 and Table 2) along the different seasons. The results are consistent with what has been known traditionally for pre-bloom applications of gibberellin in grapevine, i.e. stimulating the growth of the clusters results in loose or less compact clusters—due mainly to the elongation of the rachis (Weaver and McCune 1959; Weaver et al. 1962). The significant effect of genotype × GA3 represented by its variance demonstrated the existence of genetic diversity within the tested population that segregates in its response to GA3, mainly shown by the length of the central axis, total number of berries per cluster and the rachis fresh weight. Genetic diversity in the response to GA3 in length of the grapevine rachis was previously well known, the concentration of gibberellin used for this purpose being dependent upon the grapevine variety (Winkler et al. 1974). It is also known that this response is more marked with applications performed in earlier phenological stages (Weaver and McCune 1959). These factors may be involved in the effect of genotype × GA3 interaction observed, which would be shown by the differences in response (some genotypes respond constitutively to GA3, while others do not), differences in the degree of this response and differences in the timing and concentration of GA3 used.
In this work, a profuse production of small seedless berries was observed in some of the GA3-treated clusters and especially in the genotypes of larger rachises and seeded berries (not quantified). This phenomenon was also observed by Pratt and Shaulis (1961), who found that gibberellin application to two seeded varieties (‘Fredonia’ and ‘Concord’) during flowering time increased the fruit setting by stimulating the production of parthenocarpic berries and delaying or completely inhibiting their abscission, regardless of whether pollination had occurred (emasculated plants). According to Weaver and McCune (1959), one of the side effects of pre-flowering applications of gibberellin in seeded varieties is the production of parthenocarpic berries. The earlier the treatment is performed, the more does the effect increase in intensity. Considering that seeded genotypes were evaluated within the segregating population (Mejía et al. 2007), this effect could explain in part the increase in fruit observed with the application of GA3 as measured by the total number of berries per cluster (tb). In this work, all types of berries were considered in the estimation of tb, including “shot berries” (small seedless berries with approximately 2 mm in equatorial diameter).
Mapping and analysis of QTLs
QTLs were located on linkage group 5 (LG5), LG8, LG9, LG14, LG17 and LG18 (Table 4). LG5 presented six QTLs for the traits rw, pd and some trait ratios (rl/nr, s1/rl, rw/nr), some of them with overlapping confidence intervals (co-located) (Fig. 5 and Online Resource 5). The latter could explain the correlation among these characters, which would be in part due to pleiotropic effects or close linkage. We emphasise that other QTLs have been found in LG5 for cluster weight and number of berries per cluster (Fanizza et al. 2005), and also number of clusters per shoot (Doligez et al. 2010).
With the grapevine genome sequence being available (Jaillon et al. 2007; Velasco et al. 2007), the next step would be a direct search for candidate genes to be identified within these confidence intervals. To further this aim, we focused our exercise on directed local saturation of the preliminary genetic map available (unpublished data). We focused specifically on LG5 due to the presence of co-localised QTLs for rachis length (rl), second shoulder angle (s2l) and rachis weight (rw) during the first growing season. On the previous genetic map, LG5 only had seven markers. On the current genetic map, this LG has 19 markers in total. After the identification of these markers, the search for specific sequence or single nucleotide polymorphisms (SNP markers) that can explain or be associated with the observed phenotypic variance could also be undertaken.
The presence of overlapping QTLs between GA3-treated and untreated condition found in non-responsive traits could indicate that those QTLs were stable or did not respond to GA3. This is the case for the number of nodes of central axis (nr), which did not respond to GA3 and showed the same marker and genetic region in both conditions. In contrast, those QTLs found in different genetic regions and under different treatment conditions in GA3-responding traits could correspond to QTLs interacting with GA3.
The occurrence of QTLs with an important paternal additive effect and the fact that the paternal parent ‘Sultanina’ traits were located in the cluster (“Male” group, with a 75 % similarity) with the largest number of genotypes indicate that the bulk of the segregants had a higher similarity to ‘Sultanina’ than to the maternal parent ‘Ruby Seedless’. This result could be explained by the pedigree of the parents, as ‘Ruby Seedless’, a “grandchild” of ‘Sultanina’, presents a relationship of 25 % to the pollen donor (Mejía et al. 2007), causing the additive genetic contribution given by ‘Sultanina’ to largely exceed the contribution by ‘Ruby Seedless’.
Heritability is one of the factors that affect the detection of a QTL (Bernardo 2004). As expected, most of the mapped QTLs corresponded to those characters with greater genetic determination or heritability. However, QTL detection is not always related to the heritability value (Segura et al. 2007). Another factor involved in the QTL detection is the proportion of phenotypic variance explained by the locus or genetic effect of QTL on the trait (Darvasi et al. 1993; Collard et al. 2005). Another factor to consider is the population size. Increasing the population size provides gains in statistical power for QTL detection and analysis (Darvasi et al. 1993), a condition that could explain why few QTLs were detected—the power of QTL detection is low when a population size such as our sample (<200) is used (Collard and Mackill 2008). This handicap is common to many mapping exercises in woody species; we expect to overcome it a few years from now because we are developing a mapping population of ca. 900 segregants using parents with contrasting phenotypes for cluster architecture.
One characteristic of a QTL is that the environment strongly influences both the detection and the effect (amount of variance of a character given by the segregation of a particular QTL) of the QTL (Korol et al. 1998; Collard et al. 2005). Moreover, the latter would be the rule in tree species (Asíns 2002). The comparison among conditions, according to heritability estimated and genotype × season interaction, showed an important effect of season (Tables 2 and 3). In this work, there was a tendency: if heritability was high and genotype × season interaction was low, the QTL effect will be high to moderate (percentage of variance explained), depending on the trait considered. The latter is evident in the case of rachis length vs. peduncle diameter; the first trait has in average a higher heritability (Table 2), a lower genotype × season interaction (Table 3) and a higher QTL effect (Table 4) than the second one. For grapes grown in temperate regions, the reproductive cycle and inflorescence development are completed for two consecutive growing seasons separated by a recess period between fall and spring (Galet and Morton 1988; Carmona et al. 2002; Calonje et al. 2004). Thus, there is a long period during which several environmental factors could affect the development of the inflorescences. Some of the most important climatic factors for inflorescence differentiation correspond to the light and temperature exposure of the fruiting bud during the season prior to flowering (Srinivasan and Mullins 1981; Sánchez and Dokoozlian 2005). For example, these factors have an impact on the size of primordia and dry weight of inflorescence (Sánchez and Dokoozlian 2005). The best approach to understand the environmental impact and review the stability of a particular QTL is to assess the traits of interest under different environments. In evaluating the genomes of woody plants such as vines, researchers are usually faced with a series of evaluations in different seasons and the need to increase the population size to reduce the false negative rate in QTL detection. In this study, the effect of the season made a particularly important contribution to the expression of selected traits, regardless of whether they were under the influence of GA3.
These factors together, in addition to others, would explain the evasive nature of the genetic nature of cluster architecture. According to this study, these considerations should be taken into account in future studies to improve the genetic characterisation of genomic features associated with the architecture of the cluster of grapes.
The ideal grapevine inflorescence architecture in the case of table grapes consider aspects such as the minimal requirements of hand labour and the convenience to have a “round-shaped” cluster which is more prone for long-distance transportation. Furthermore, it is desirable to have less compact clusters resulting in less contact area between the berries, which in turn increases the size of the berries and reduces fungal infections. A target architecture should consist of a long peduncle and long internodes, their shoulders and berries being low in number per cluster (<10 and <150, respectively) with open angles. We have identified QTLs that could be to some extent related to that ideal shape (number of nodes along the rachis; total number of berries and their relation to rachis length; distance between nodes) and a list of candidate genes that could be useful for breeding purposes. In any case, these genes could become selection tools only after identifying polymorphisms that could be associated with the phenotype of interest in a defined genetic background. This approach most probably will include a group of selection markers, instead of the more commonly described approach of using a single selection marker (Mejia et al. 2011).
Candidate genes for the cluster architecture
The GO term enrichment analysis in QTL confidence interval is a novel and useful approach to discover possible enrichment in gene functions that could be related to traits under study (reviewed by Drinkwater and Gould 2012). One limitation is based on the available Gene Ontology annotation for plant species such as grapevine that is quite limited in comparison to model plants such as Arabidopsis thaliana (Monclus et al. 2012). Indeed, in the case of Vitis reference genome which encompasses 26,346 genes, only 14,511 are currently associated with at least one GO term. In spite of that, the search for GO terms is very useful when the list of candidate genes is too large to be considered individually for further analyses, as happened in this case (1,173 identified genes).
We selected 50 out of the 891 genes annotated in the confidence intervals corresponding to the complete set of 19 QTLs distributed in six linkage groups (Online Resource 5). We decided this selection based on the annotated function for each gene. However, considering the poorness in the curation of the current version of the automatic annotation of the grapevine genome (A.-F. Adam-Blondon, personal communication), using this approach we could be missing some relevant genes that may have not been annotated in the targeted areas covering ca. 20.3 Mbases (Online Resources 6 and 7). This situation would not be improved until a better annotation of the grapevine genome is afforded. Most of the 50 candidate genes correspond to six different functional groups, including water transport, cell division, cell wall remodelling, hormonal response, transcription factors and lipid synthesis. Some of these genes are described in the following.
Aquaporines are key proteins for plant growth and development because of their role in water transport into the cells (Maurel and Chrispeels 2001). This could be the case during rachis elongation, a biological process that could determine the size and architecture of the cluster (Table 5). Aquaporins have been classified based on their sequences in four sub-families: plasma membrane intrinsic proteins (PIPs), tonoplast intrinsic proteins (TIPs), nodulin 26-like intrinsic proteins (NIPs) and basic plasma membrane intrinsic proteins (Chaumont et al. 2005). These proteins have been postulated as key factors for the development of plant organs in general, participating in cells elongation and expansion. Then, we could expect differential expression level or changes in functionality between alleles corresponding to each phenotype, a matter to be evaluated in further transcriptomic and gene-directed studies. We found three of these genes (two PIPs and one NIP) among the identified QTLs (Table 5).
The second group is related to cell wall structure and metabolism proteins. Any gene that could be related to cell wall would be a candidate gene for tissue elongation. The cell wall is composed of ca. 90 % polysaccharides made of three main structural components: cellulose, a matrix of intercrossed glycans (hemicellulose) and pectin-related polysaccharides (Goulao et al. 2012). The remaining components correspond to structural glycoproteins, phenolic esters and minerals. Xyloglucans are the predominant hemicelluloses in the primary cell wall of dicots. Pectins are embedded in the cellulose–hemicellulose lattice, forming hydrophilic gels that provide mechanical strength to the cell wall (Goulao et al. 2012). In Arabidopsis the gene BEARSKIN1 (BRN1) is related to transcription factors (TFs) of the VND/NST family, and these TFs have been related to the synthesis of the secondary cell wall in tissues or anatomical parts such as xylem, interfascicular fibres and anthers (Bennett et al. 2010). Pectate lyase is an enzyme involved in the degradation of the cell wall through the breakage of esterified pectins, one of the main components of the higher plants cell wall. A recent report shows that a pectate lyase present in the cell wall is associated with berry texture and cluster size (Vargas et al. 2013).
The identification of a number of TFs in various QTLs attracts our attention, considering their pivotal role in the control or coordination of many cellular events by modulating the expression of groups of genes. We identified a few TFs that we suspect could be relevant to the growth of rachis based on the available evidence mainly from Arabidopsis. This is the case of BIM1 that plays a key role in this model plant’s growth and development, participation in cellular expansion, proliferation and differentiation in young tissues (Chandler et al. 2009). Two other TFs identified were the MADS-Box AGAMOUS family AGL8 and AGL11, whose role could be related to the regulation of floral meristem development (Rounsley et al. 1995). Another member of this gene family, AGL11, has been recently demonstrated to be the key gene determining the seed content in grapevine (Mejia et al. 2011), but this gene also explains over 30 % of the phenotypic variance of berry size, a trait that we are now associating with cluster structure. In contrast, it was recently reported that the gene TERMINAL FLOWER1 (TFL1), well known as a key factor regulating the inflorescence development in Arabidopsis, would also regulate the cluster shape or architecture (Fernandez et al. 2010); however, this gene mapped in a region of LG17 where we did not find any QTL related to cluster architecture. This could be explained in different ways. For instance, the genomic organisation could be different between ‘Carignan’, the genotype harbouring the TFL1 mutation, and the table grape genetic background considered in this study. In this respect, we have recently found that the ‘Sultanina’ genome has a certain number of intra- and interchromosomal translocations in comparison to the PN-40024 grapevine reference genome (Di Genova et al. 2014). The most probable explanation, however, is that there is no trait variation associated with this gene in the studied R × S population. Alternatively, individual table grape genes can exhibit a different organisation, as we have observed in a number of cases when comparing transcripts of individuals of this mapping population vs. the reference genome (Muñoz-Espinoza et al. manuscript in preparation).
The identification of the most relevant genes related to rachis architecture would impact heavily on the future breeding of table grape cultivars, considering both the necessity to reduce the use of pesticides and the raising of the labour costs related to the hand-made cluster management.
Conclusions
The cluster architecture in a grapevine, represented by an R × S segregating population, is mainly determined by the length and the fresh weight of the rachis, the number of internodes, the peduncle diameter and the length of the lateral shoulders. There is genetic diversity in the response to GA3, mainly manifested in the length of the central axis, the number of berries and the rachis fresh weight. Furthermore, season has an important effect on the expression of these traits, regardless of whether the cluster is under the influence of GA3. The QTLs determining these traits and their response to GA3 are distributed on six LGs. The correlations found between these traits can be partially attributed to the pleiotropic effects and to a close linkage between the QTLs. A set of 50 genes underlying these QTLs are proposed as possibly related to the cluster architecture phenotype determination.
References
Acquaah G, Adams MW, Kelly JD (1992) A factor analysis of plant variables associated with architecture and seed size in dry bean. Euphytica 60:171–177. doi:10.1007/BF00039395
Asíns MJ (2002) Present and future of quantitative trait locus analysis in plant breeding. Plant Breed 121:281–291. doi:10.1046/j.1439-0523.2002.730285.x
Bates D, Maechler M (2009) lme4: Linear mixed-effects models using S4 classes. R package version 0.999375-32. http://CRAN.R-project.org. Accessed 3 March 2012
Bennett T, van den Toorn A, Sanchez-Pérez GF, Campilho A, Willemsen V, SneI B, Scheres B (2010) SOMBRERO, BEARSKIN1, and BEARSKIN2 regulate root cap maturation in Arabidopsis. Plant Cell 3:640–654
Bernardo R (2004) What proportion if declared QTL in plants are false? Theor Appl Genet 109:419–424. doi:10.1007/s00122-004-1639-3
Boss PK, Thomas MR (2000) Tendrils, inflorescence and fruitfulness: a molecular perspective. Aust J Grape Wine Res 6:168–174. doi:10.1111/j.1755-0238.2000.tb00176.x
Brown PJ, Klein PE, Bortiri E, Acharya CB, Rooney WL, Kresovich S (2006) Inheritance of inflorescence architecture in sorghum. Theor Appl Genet 113:931–942. doi:10.1007/s00122-006-0352-9
Busov VB, Brunner AM, Strauss SH (2008) Genes for control of plant stature and form. New Phytol 177:589–607. doi:10.1111/j.1469-8137.2007.02324.x
Cabezas JA, Cervera MT, Ruíz-Garcia L, Carreño J, Martínez-Zapater JM (2006) A genetic analysis of seed and berry weight in grapevine. Genome 49:1572–1585. doi:10.1139/g06-122
Calonje M, Cubas P, Martínez-Zapater JM, Carmona MJ (2004) Floral meristem identity genes are expressed during tendril development in grapevine. Plant Physiol 135:1491–1501. doi:10.1104/pp.104.040832
Carmona MJ, Cubas P, Martinez-Zapater JM (2002) VFL, the grapevine. FLORICAULA/LEAFY ortholog, is expressed in meristematic regions independently of their fate. Plant Physiol 130:68–77. doi:10.1104/pp.002428
Chaim AB, Paran I, Grube RC, Jahn M, Van Wijk R, Peleman J (2001) QTL mapping of fruit-related traits in pepper (Capsicum annuum). Theor Appl Genet 102:1016–1028
Chandler J, Cole M, Flier A, Werr W (2009) BIM, a bHLH protein involved in brassinosteroid signaling, controls Arabidopsis embryonic patterning via interaction with DORNROSCHEN and DORNROSCHEN-LIKE. Plant Mol Biol 69:57–68
Chaumont F, Moshelion M, Daniels M (2005) Regulation of plant aquaporin activity. Biol Cell 97:749–764
Collard BCY, Mackill DJ (2008) Marker assisted selection: an approach for precision plant breeding in the twenty-first century. Philos Trans R Soc Lond Ser B Biol Sci 363:557–572. doi:10.1098/rstb.2007.2170
Collard BCY, Jahufer MZZ, Brouwer J, Pang ECK (2005) An introduction to markers, quantitative trait loci (QTL) ping and marker-assisted selection for crop improvement: the basic concepts. Euphytica 142:169–196. doi:10.1007/s10681-005-1681-5
Coombe BG (1995) Growth stages of the grapevine: adoption of a system for identifying grapevine growth stages. Aust J Grape Wine Res 1:100–110. doi:10.1111/j.1755-0238.1995.tb00086.x
Costantini L, Grando MS, Feingold S, Ulanovsky S, Mejía N, Hinrichsen P, Doligez A, This P, Cabezas JA, Martinez-Zapater JM (2007) Generation of a common set of mapping markers to assist table grape breeding. Am J Enol Vitic 58:102–111
Costantini L, Battilana J, Lamaj F, Fanizza G, Grando MS (2008) Berry and phenology-related traits in grapevine (Vitis vinifera L.): from quantitative trait loci to underlying genes. BMC Plant Biol 8:38. doi:10.1186/1471-2229-8-38
Costes E (2003) Exploring within-tree architectural development of two apple tree cultivars over 6 years. Annals Bot 91:91–104. doi:10.1093/aob/mcg010
Darvasi A, Weinreb A, Minke V, Weller JI, Soller M (1993) Detecting marker-QTL linkage and estimating QTL gene effect and map location using a saturated genetic map. Genetics 134:943–951
Di Gaspero G, Peterlunger E, Testolin R, Edwards KJ, Cipriani G (2000) Conservation of microsatellite loci within genus Vitis. Theor Appl Genet 101:301–308. doi:10.1007/s001220051483
Di Genova A, Miyasaka-Almeida A, Vizoso P, Travisany D, Moraga C, Munoz C, Pinto M, Hinrichsen P, Maass A, Orellana A (2014) Whole genome comparison between table and wine grapes reveals a comprehensive catalog of structural variants. BMC Plant Biol 14(1):7
Doligez A, Bouquet A, Danglot Y, Lahogue F, Riaz S, Meredith C, Edwards K, This P (2002) Genetic mapping of grapevine (Vitis vinifera L.) applied to the detection of QTLs for seedlessness and berry weight. Theor Appl Genet 105:780–795. doi:10.1007/s00122-002-0951-z
Doligez A, Bertrand Y, Dias S, Grolier M, Ballester JF, Bouquet A, This P (2010) QTLs for fertility in table grape (Vitis vinifera L.). Tree Genet Genomes 6:413–422. doi:10.1007/s11295-009-0259-0
Doucleff M, Jin Y, Gao F, Riaz S, Krivanek AF, Walker MA (2004) A genetic linkage map of grape, utilizing Vitis rupestris and Vitis arizonica. Theor Appl Genet 109:1178–1187. doi:10.1007/s00122-004-1728-3
Drinkwater NR, Gould MN (2012) The long path from QTL to gene. PLoS Genet. doi:10.1371/journal.pgen.1002975
Du Z, Zhou X, Ling Y, Zhang Z, Su Z (2010) agriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res 38:W64–W70. doi:10.1093/nar/gkq310
Duchêne E, Butterlin G, Dumas V, Merdinoglu D (2012) Towards the adaptation of grapevine varieties to climate change: QTLs and candidate genes for developmental stages. Theor Appl Genet 124:623–635. doi:10.1007/s00122-011-1734-1
Dudley JW, Moll RH (1969) Interpretation and use of estimates of heritability and genetic variances in plant breeding. Crop Sci 9:257–262
Eibach R, Hastrich H, Töpfer R (2003) Inheritance of aroma compounds. Acta Hort 603:337–344
Falconer DS, Mackay TFC (1996) Introduction to quantitative genetics, 4th edn. Addison Wesley Longman, Harlow
Fanizza G, Lamaj F, Costantini L, Chaabane R, Grando MS (2005) QTL analysis for fruit yield components in table grapes (Vitis vinifera). Theor Appl Genet 111:658–664. doi:10.1007/s00122-005-2016-6
Fernandez L, Torregrosa L, Segura V, Bouquet A, Martinez-Zapater JM (2010) Transposon-induced gene activation as a mechanism generating cluster shape somatic variation in grapevine. Plant J 61:545–557. doi:10.1111/j.1365-313X.2009.04090.x
Fischer BM, Salakhutdinov I, Akkurt M, Eibach R, Edwards KJ, Töpfer R, Zyprian EM (2004) Quantitative trait locus analysis of fungal disease resistance factors on a molecular map of grapevine. Theor Appl Genet 108:501–515. doi:10.1007/s00122-003-1445-3
Fournier-Level A, Le Cunff L, Gomez C, Doligez A, Ageorges A, Roux C, Bertrand Y, Souquet JM, Cheynier V, This P (2009) Quantitative genetic bases of anthocyanin variation in grape (Vitis vinifera L. ssp. sativa) berry: a quantitative trait locus to quantitative trait nucleotide integrated study. Genetics 183:1127–1139. doi:10.1534/genetics.109.103929
Galet P, Morton LT (1988) The family Vitaceae and Vitis speciation. In: Pearson RC, Goheen AD (eds) Compendium of grape diseases. American Phytopathological Society, Saint Paul, pp 2–3
García-Maroto F, Ortega N, Lozano R, Carmona MJ (2000) Characterization of the potato MADS-box gene STMADS16 and expression analysis in tobacco transgenic plants. Plant Mol Biol 42:499–513. doi:10.1023/A:1006397427894
Goosey L, Sharrock R (2001) The Arabidopsis compact inflorescence genes: phase-specific growth regulation and the determination of inflorescence architecture. Plant J 26:549–559. doi:10.1046/j.1365-313x.2001.01053.x
Goulao L, Fernandes J, Lopes P, Amancio S (2012) Tackling the cell wall of the grape berry. In: Gerós M, Chaves MM, Delrot S (eds) The biochemistry of the grape berry. Bentham e-Books, Sharjah, pp 172–193
Grandillo S, Ku HM, Tanksley SD (1999) Identifying the loci responsible for natural variation in fruit size and shape in tomato. Theor Appl Genet 99:978–987
Guerrero M (2007) Mapeo integrativo físico y genético de genes candidatos relacionados al desarrollo de la baya y la semillas en Vitis vinifera L. Molecular Biotechnology Engineering Thesis. Universidad de Chile. Santiago, Chile. (In Spanish)
Hed B, Ngugi HK, Travis JW (2011) Use of gibberellic acid for management of bunch rot on Chardonnay and Vignoles grape. Plant Dis 95:269–278. doi:10.1094/PDIS-05-10-0382
Holland JB, Nyquist WE, Cervantes-Martinez CT (2003) Estimating and interpreting heritability for plant breeding: an update. Plant Breeding Rev 22:9–112. doi:10.1002/9780470650202.ch2
Jaillon O, Aury J-M, Noel B, Policriti A, Clepet C, Casagrande A, Choisne N, Aubourg S, Vitulo N, Jubin C, Vezzi A, Legeai F, Hugueney P, Dasilva C, Horner D, Mica E, Jublot D, Poulain J, Bruyere C, Billault A, Segurens B, Gouyvenoux M, Ugarte E, Cattonaro F, Anthouard V, Vico V, Del Fabbro C, Alaux M, Di Gaspero G, Dumas V, Felice N, Paillard S, Juman I, Moroldo M, Scalabrin S, Canaguier A, Le Clainche I, Malacrida G, Durand E, Pesole G, Laucou V, Chatelet P, Merdinoglu D, Delledonne M, Pezzotti M, Lecharny A, Scarpelli C, Artiguenave F, Pé E, Valle G, Morgante M, Caboche M, Adam-Blondon A-F, Weissenbach J, Quétier F, Wincker P (2007) The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449:463–468. doi:10.1038/nature06148
Kawamura K, Hibrand-Saint Oyant L, Crespel L, Thouroude T, Lalanne D, Foucher F (2011) Quantitative trait loci for flowering time and inflorescence architecture in rose. Theor Appl Genet 122:661–675. doi:10.1007/s00122-010-1476-5
Korol AB, Ronin YI, Nevo E (1998) Approximate analysis of QTL-environment interaction with no limits on the number of environments. Genetics 148:2015–2028
Krivanek AF, Riaz S, Walker MA (2006) Identification and molecular mapping of PdR1, a primary resistance gene to Pierce’s disease in Vitis. Theor Appl Genet 112:1125–1131. doi:10.1007/s00122-006-0214-5
Larson SR, Kellogg EA, Jensen KB (2013) Genes and QTLs controlling inflorescence and stem branch architecture in Leymus (Poaceae: Triticeae) Wildrye. J Hered 104:678–691. doi:10.1093/jhered/est033
Liu C, Teo ZW, Bi Y, Song S, Xi W, Yang X, Yin Z, Yu H (2013) A conserved genetic pathway determines inflorescence architecture in Arabidopsis and rice. Dev Cell 24:612–622. doi:10.1016/j.devcel.2013.02.013
Mackay TFC (2001) The genetic architecture of quantitative traits. Annu Rev Genet 35:303–339. doi:10.1146/annurev.genet.35.102401.090633
Marino R, Sevini F, Mandini A, Vecchione A, Pertot I, Serra AD, Versini G, Velasco R, Grando MS (2003) QTL mapping for disease resistance and fruit quality in grape. Acta Hort 603:527–533
Maurel C, Chrispeels MJ (2001) Aquaporins. A molecular entry into plant water relations. Plant Physiol 125:135–138
Mejia N, Soto B, Guerrero M, Casanueva X, Miccono MA, Houel C, Le Cunff L, Boursiquot JM, Hinrichsen P, Adam-Blondon AF (2011) Molecular, genetic and transcriptional evidence for a role of VvAGL11 in stenospermocarpic seedlessness in grapevine. BMC Plant Biol 11:57
Mejía N, Gebauer M, Muñoz L, Hewstone N, Muñoz C, Hinrichsen P (2007) Identification of QTLs for seedlessness, berry size, and ripening date in a seedless × seedless table grape progeny. Am J Enol Vitic 58:499–507
Merdinoglu D, Butterlin G, Bevilacqua L, Chiquet V, Adam-Blondon AF, Decroocq S (2005) Development and characterization of a large set microsatellite markers in grapevine (Vitis vinifera L.) suitable for multiplex PCR. Mol Breed 15:349–366. doi:10.1007/s11032-004-7651-0
Monclus R, Leple JC, Bastien C, Bert PF, Villar M, Marron N, Brignolas F, Jorge V (2012) Integrating genome annotation and QTL position to identify candidate genes for productivity, architecture and water-use efficiency in Populus spp. BMC Plant Biol 12:173. doi:10.1186/1471-2229-12-173
Nakagawa M, Shimamoto K, Kyozuka J (2002) Overexpression of RCN1 and RCN2, rice TERMINAL FLOWER 1/CENTRORADIALIS homologs, confers delay of phase transition and altered panicle morphology in rice. Plant J 29:743–750. doi:10.1046/j.1365-313X.2002.01255.x
Nomura T, Nakayama M, Reid JB, Takeuchi Y, Yokota T (1997) Blockage of brassinosteroid biosynthesis and sensitivity causes dwarfism in garden pea. Plant Physiol 113:31–37. doi:10.1104/pp.113.1.31
Okada K, Shimura Y (1994) Genetic analyses of signaling in flower development using Arabidopsis. Plant Mol Biol 26:1357–1377. doi:10.1007/BF00016480
Périn C, Hagen LS, Giovinazzo N, Besombes D, Dogimont C, Pitrat M (2002) Genetic control of fruit shape acts prior to anthesis in melon (Cucumis melo L.). Mol Genet Genomics 266:933–941
Peterson PA (1959) Linkage of fruit shape and color genes in Capsicum. Genetics 44:407–419
Piepho HP, Möhring J (2007) Computing heritability and selection response from unbalanced plant breeding trials. Genetics 177:1881–1888. doi:10.1534/genetics.107.074229
Poehlman JM (1986) Breeding field crops, 3rd edn. Van Nostrand Reinhold, New York
Pratt C (1971) Reproductive anatomy in cultivated grapes—a review. Am J Enol Vitic 22:92–109
Pratt C, Shaulis NJ (1961) Gibberellin-induced parthenocarpy on grapes. Proc Amer Soc Hort Sci 77:322–330
R Development Core Team (2012) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. ISBN 3-900051-07-0. http://www.R-project.org/. Accessed 3 March 2012
Rounsley S, Ditta G, Yanofsky M (1995) Diverse roles for MADS Box genes in Arabidopsis development. Plant Cell 7:1259–1269
Sánchez LA, Dokoozlian NK (2005) Bud microclimate and fruitfulness in Vitis vinifera L. Am J Enol Vitic 56:319–329
Schneider W, Staudt G (1979) Zur schätzung der heritabilität im weiteren sinn einiger merkmale von Vitis vinifera. Vitis 18:238–243
Sefc KS, Regner F, Tureschek E, Glössl J, Steinkellner H (1999) Identification of microsatellite sequences in Vitis riparia and their application for genotyping of different Vitis species. Genome 42:367–373. doi:10.1139/g98-168
Segura V, Cilas C, Laurens F, Costes E (2006) Phenotyping progenies for complex architectural traits: a strategy for 1-year-old apple trees (Malus × domestica Borkh.). Tree Genet Genomes 2:140–151. doi:10.1007/s11295-006-0037-1
Segura V, Denacé C, Durel CE, Costes E (2007) Wide range QTL analysis for complex architectural traits in a 1-year-old apple progeny. Genome 50:159–171. doi:10.1139/G07-002
Shavrukov YN, Dry IB, Thomas MT (2004) Inflorescence and bunch architecture developmentin Vitis vinifera L. Aust J Grape Wine Res 10:116–124. doi:10.1111/j.1755-0238.2004.tb00014.x
Sillampää M, Corander J (2002) Model choice in gene mapping: what and why. Trends Genet 18:301–307. doi:10.1016/S0168-9525(02)02688-4
Singer SR (2007) Inflorescence architecture—moving beyond description to development, genes and evolution. In: Ainsworth C (ed) Annual plant reviews, flowering and its manipulation, vol 20, Blackwell Publishing Ltd, Oxford, UK. doi:10.1002/9780470988602.ch5
Srinivasan C, Mullins MG (1981) Physiology of flowering en the grapevine—a review. Am J Enol Vitic 32:47–63
Stanfield WD (1991) Theory and problems of genetics, 3rd edn. McGraw-Hill, New York
Tanksley SD (2004) The genetic, developmental, and molecular bases of fruit size and shape variation in tomato. Plant Cell 16:S181–S189. doi:10.1105/tpc.018119
Ungerer MC, Halldorsdottir SS, Modliszewski JL, Mackay TF, Purugganan MD (2002) Quantitative trait loci for inflorescence development in Arabidopsis thaliana. Genetics 160:1133–1151
Ungerer MC, Halldorsdottir SS, Purugganan MD, Mackay TF (2003) Genotype-environment interactions at quantitative trait loci affecting inflorescence development in Arabidopsis thaliana. Genetics 165:353–365
Upadyayula N, da Silva HS, Bohn MO, Rocheford TR (2006) Genetic and QTL analysis of maize tassel and ear inflorescence architecture. Theor Appl Genet 112:592–606. doi:10.1007/s00122-005-0133-x
Vail ME, Marois JJ (1991) Grape cluster architecture and the susceptibility of berries to Botrytis cinerea. Phytopathology 81:188–191. doi:10.1094/Phyto-81-188
Vail ME, Wolpert JA, Gubler WD, Rademacher MR (1998) Effect of cluster tightness on Botrytis bunch rot in six Chardonnay clones. Plant Dis 82:107. doi:10.1094/PDIS.1998.82.1.107
Van Ooijen JW, Boer MP, Jansen RC, Maliepaard C (2002) MapQTL® 4.0: software for the calculation of QTL positions on genetic maps. Plant Research International, Wageningen, The Netherlands
Vargas A, Fajardo C, Borrego J, De Andres M, Ibañez J (2013) Polymorphisms in VvPel associate with variation in berry texture and bunch size in the grapevine. Aust J Grape Wine Res 19:193–207
Velasco R, Zharkikh A, Troggio M, Cartwright DA, Cestaro A, Pruss D, Pindo M, Fitzgerald LM, Vezzulli S, Reid J, Malacarne G, Iliev D, Coppola G, Wardell B, Micheletti D, Macalma T, Facci M, Mitchell JT, Perazzolli M, Eldredge G, Gatto P, Oyzerski R, Moretto M, Gutin N, Stefanini M, Chen Y, Segala C, Davenport C, Demattè L, Mraz A, Battilana J, Stormo K, Costa F, Tao Q, Si-Ammour A, Harkins T, Lackey A, Perbost C, Taillon B, Stella A, Solovyev V, Fawcett JA, Sterck L, Vandepoele K, Grando SM, Toppo S, Moser C, Lanchbury J, Bogden R, Skolnick M, Sgaramella V, Bhatnagar SK, Fontana P, Gutin A, Van de Peer Y, Salamini F, Viola R (2007) A high quality draft consensus sequence of the genome of a heterozygous grapevine variety. PLoS ONE 2:e1326
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Heredity 93:77–78. doi:10.1093/jhered/93.1.77
Weaver RJ, McCune SB (1959) Response of certain varieties of Vitis vinifera to gibberellin. Hilgardia 28:297–350
Weaver RJ, Kasimatis AN, McCune SB (1962) Studies with gibberellin on wine grapes to decrease bunch rot. Am J Enol Vitic 13:78–82
Wei T (2011) corrplot: Visualization of a correlation matrix. R package version 0.60. http://CRAN.R-project.org. Accessed 26 March 2012
Weiss J, Delgado-Benarroch L, Egea-Cortines M (2005) Genetic control of floral size and proportions. Int J Dev Biol 49:513–525. doi:10.1387/ijdb.051998jw
Winkler AJ, Cook JA, Kliewer WM, Lider LA (1974) General viticulture, 2nd edn. University of California Press, Berkeley
Xu K, Riaz S, Roncoroni NC, Jin Y, Hu R, Zhou R, Walker MA (2008) Genetic and QTL analysis of resistance to Xiphinema index in a grapevine cross. Theor Appl Genet 116:305–311. doi:10.1007/s00122-007-0670-6
Zhu H, Gilchrist L, Hayes P, Kleinhofs A, Kudrna D, Liu Z, Prom L, Steffenson B, Toojinda T, Vivar H (1999) Does function follow form? Principal QTLs for Fusarium head blight (FHB) resistance are coincident with QTLs for inflorescence traits and plant height in a doubled-haploid population of barley. Theor Appl Genet 99:1221–1232. doi:10.1007/s001220051328
Zyprian E, Eibach R, Töpfer R (2003) Comparative molecular mapping in segregating populations of grapevine. Acta Hort 603:73–77
Acknowledgments
This work was financed by GENOMA-Chile (FONDEF-CONICYT Grant G07I-1002). We also gratefully acknowledge the comments and inputs by the two anonymous reviewers.
Conflict of interest
The authors have no conflict of interest.
Ethical standards
The authors declare that the experiments comply with the current laws of Chile.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Toepfer.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Correa, J., Mamani, M., Muñoz-Espinoza, C. et al. Heritability and identification of QTLs and underlying candidate genes associated with the architecture of the grapevine cluster (Vitis vinifera L.). Theor Appl Genet 127, 1143–1162 (2014). https://doi.org/10.1007/s00122-014-2286-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-014-2286-y