Abstract
Regulation of gene expression is associated with RNA interference (RNAi) at the posttranscriptional level and chromatin modification in transcriptional silencing during plant development. Dicer-like (DCL) and Argonaute (AGO) proteins, along with RNA-dependent RNA polymerase (RDR), play integral roles in these processes. Many members of the DCL, AGO, and RDR gene families have been characterized in Arabidopsis, rice, maize, and tomato. However, little is known about these genes in grapevine. Here, we implemented a comprehensive bioinformatics analysis of the DCL, AGO, and RDR gene families in grapevine. In this study, a total of four VvDCL, thirteen VvAGO, and five VvRDR genes were identified in grapevine. These genes were categorized into four groups for each family based on phylogenetic analysis. Twenty-two genes were distributed across 11 chromosomes. We detected five segmental or tandem duplication events, which suggested their conserved functions during evolution. In each subfamily, protein motif composition was relatively conserved. The digital EST expression analyses provided a first glimpse of the expression patterns of DCL, AGO, and RDR gene in grapevine. It is worth mentioning that one gene, VvAGO10a, only expressed in the stem, which suggested that VvAGO10a may function in the regulation of siRNAs in the grapevine stem. Gene expression patterns during berry development and under different abiotic stresses were investigated utilizing microarray data. Thus, these detailed investigations provide a foundation for transcriptional silencing-associated gene families in grapevine for future functional genomics studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There are two types of small RNA molecules (approximately 21–24 nucleotides) in multicellular eukaryotes: microRNAs (miRNAs) and short interfering RNAs (siRNAs). These two types act to modify chromatin and genome structure as sequence-specific guides to silence and regulate genes, transposons, and viruses (Xie and others 2004; Alves and others 2006). miRNAs (approximately 21–22 nucleotides nt), which are found in plants, are often phylogenetically conserved; siRNAs are processed from precursors which contain extensive or exclusive double-stranded RNA (dsRNA) structures (Hannon 2002). The target RNAs are suppressed by siRNAs and miRNAs that function posttranscriptionally. siRNAs guide sequence-specific nucleolytic activity of the RNA-induced silencing complex (RISC) to complementary target sequences (Hannon 2002). Several silencing-associated protein factors, such as proteins of the Dicer-like (DCL) family, the Argonaute (AGO) family, and RNA-dependent RNA polymerases (RDR) have been identified in plants (Wang and Metzlaff 2005).
Dicer-like (DCL) proteins are critical components of the miRNA and siRNA biogenesis pathways, which play a key role in processing long double-stranded RNAs into mature small RNAs. DCL proteins generally contain six types of domains: DExD, Helicase-C, DUF283, PAZ, RNase III, and double-stranded RNA-binding (dsRB). The DExD and Helicase-C domains are found at the N-terminal and C-terminal of the proteins, respectively. Duf283 is located at the N-terminal of the first RNase III domain (Dlakić 2006; Margis and others 2006). There are two RNase III domains (RNase IIIa, RNase IIIb) present in Dicer proteins. A long α-helix generally helps the PAZ domain of Dicer directly connect to the RNase IIIa domain. The 3′ end of a dsRNA containing a two-base overhang binds specifically to the PAZ domain (MacRae and others 2006). The dsRBa domain, along with the PAZ, RNase IIIa and IIIb domains, recognizes and processes the substrate RNA (Margis and others 2006). In monocot and dicot plants, the remarkable expansion of DCL family members may mirror the deployment of the RNA silencing strategy in antiviral defense. Four DCL proteins (DCL1–DCL4) found in Arabidopsis are suggested to specialize in small RNA biogenesis (Margis and others 2006). AtDCL1 is not only associated with miRNA production, but also plays a role in the production of small RNAs from endogenous inverted repeats. AtDCL2 generates siRNAs from natural cis-acting antisense transcripts and functions in antiviral defense. AtDCL3 generates siRNAs to guide chromatin modification, whereas AtDCL4 is associated with tasiRNA metabolism and acts during posttranscriptional silencing (Liu and others 2007).
Argonaute (AGO) is a strongly conserved protein family, the members of which are catalytic components of the RNA-induced gene silencing complex (RISC). RISCs also contain small RNAs and play important roles in gene silencing at the transcriptional and posttranscriptional levels, as well as disease resistance in plants. AGO proteins usually contain three conserved domains: PAZ at the N-terminus, MID and PIWI at the C-terminus. A 3′ two-nucleotide overhang, which results from RNA digestion by RNase III, is anchored to a specific binding pocket in the PAZ domain. The MID domain specifically binds the 5′ phosphate of the small RNAs. Therefore, small RNAs are anchored onto AGO proteins (Peters and Meister 2007). The 5′ end of the siRNA is bound to the target RNA by the PIWI domain, and the spatial structure and the catalytic site show a high level of homology to RNase H (Höck and Meister 2008; Qian and others 2011). In Arabidopsis, the AGO family comprises 10 members (Fagard and others 2000; Carmell and others 2002), of which two have been unambiguously associated with different forms of RNA silencing. It is therefore likely that, as in animals, the functional diversification of RNA silencing is directly linked to the variation in AGO family members. AGO1 is associated with the miRNA pathway and the transgene-silencing pathway (Fagard and others 2000), and AGO4 is associated with endogenous siRNAs affecting epigenetic silencing (Zilberman and others 2003; Zilberman and others 2004). Moreover, AGO7 and ZLL/AGO10 have a function in the transition from the juvenile to adult phases of plant growth (Hunter and others 2003) and meristem maintenance (Moussian and others 1998; Lynn and others 1999), respectively.
RNA-dependent RNA polymerase (RDR) proteins in plants rely on aberrant RNA molecules to expand RNA interference signals. These proteins can enhance the genomic level of transposons to silence viral genes in infected cytoplasm. RDR proteins share a same sequence motif which is not related to the catalytic domain of DNA-dependent RNA polymerases (Iyer and others 2003). At least three active RDR genes are known in Arabidopsis, and these have been named RDR1, RDR2, and RDR6 (also known as SDE1/SGS2). RDR1 orthologs are necessary components of the cytoplasmic RNA silencing pathway, which silences transgenes and viruses. Compared with miRNAs, the endogenous siRNA requirement for RDR2 showed complete insensitivity to each of the rdr mutations tested. RDR2 appears to interact with DCL3 both physically and functionally and is presumably localized in the nucleus (Xie and others 2004). RDR2 works with DCL3 to form chromatin-associated siRNAs (24 nt) that function through AGO4. RDR6 plays a role in antiviral defense (Wassenegger and Krczal 2006; Kasschau and others 2007), and the direct involvement of these proteins in viral siRNA biogenesis has recently been investigated (Diaz-Pendon and others 2007; Qi and others 2009).
To date, a total of 20 genes in Arabidopsis (Arabidopsis thaliana), 32 genes in rice (Oryza sativa), 28 genes in maize (Zea mays), 28 genes in tomato (Solanum lycopersicum), and 38 genes in foxtail millet (Setaria italica) have been identified in the DCL, AGO, and RDR gene families (Kapoor and others 2008; Qian and others 2011; Bai and others 2012; Yadav and others 2014). While grapevine ranks first among fruit crops in the world in terms of both production and economic importance (Cramer and others 2007), none of the DCL, AGO, and RDR genes has been investigated in grapevine. Therefore, research on the RNAi machinery components in grapevine is very important. In this study, a comprehensive set of DCL, AGO, and RDR genes was identified and analyzed in the grapevine genome. The results presented will provide basic genomic information for these gene families and provide insights into the probable physiological function of these genes in grapevine growth and development.
Materials and Methods
Identification of Grapevine DCL, AGO, and RDR Genes
The latest Vitis vinifera genome assembly and protein sequences were downloaded from Phytozome v7.0 (http://www.phytozome.net). The published Arabidopsis DCL, AGO, and RDR gene sequences were used as queries to search for the orthologous grapevine genes. The Hidden Markov Model (HMM) profile of the DCL, AGO, and RDR protein domains was from the Pfam database (http://pfam.janelia.org/) and was used to search for grapevine DCL, AGO, and RDR genes with the BlastP program (P value = 0.001). Significant hits were then used as query sequences to search against the National Centre for Biotechnology Information (http://www.ncbi.nlm.nih.gov/BLAST) using the TBLASTN program (P value = 0.001). The Pfam database (http://www.sanger.ac.uk/resources/software/) was finally used to confirm each predicted VvDCL, VvAGO, or VvRDR protein sequence as a DCL, AGO, or RDR protein, respectively. A complete sequence alignment using MEGA 4.0 was performed to eliminate similar and deficient sequences (Tamura and others 2007). Non-overlapping DCL, AGO, and RDR protein sequences were screened for further analysis. In addition, length, molecular weight, and isoelectric points (pI) of the predicted proteins were predicted with ExPasy (http://au.expasy.org/tools/pi_tool.html) (He and others 2012).
Phylogenetic Analysis of Grapevine DCL, AGO, and RDR Genes
Amino acid sequences of all the DCL, AGO, and RDR genes of tomato, Arabidopsis, foxtail millet, maize, and rice were downloaded from Phytozome v7.0, and multiple-sequence alignments were merged with Clustal-X (version 1.83) (Thompson and others 1997). Phylogenetic trees for all complete DCL, AGO, and RDR protein sequences were also constructed using the neighbor-joining (NJ) method with bootstrapping as implemented in MEGA v4.0 (Tamura and others 2007). The genes used in this study were named according to their phylogenetic relationship with other members of the same gene family in the previous species (Bai and others 2012).
Chromosomal Localization, Gene Duplication, and Conserved Motif Analysis
The physical locations of DCL, AGO, and RDR gene loci were obtained from the Phytozome v7.0 database. All genes were located on the 11 chromosomes in grapevine based on their locus positions from the top to the bottom. The DCL, AGO, and RDR gene loci on the grapevine chromosomes were drawn on a schematic diagram using MapInspect software (He and others 2012). Two genes located in the same clade of the phylogenetic tree were treated as paralogs in the same species for the detection of tandem and segmental duplications (Zhao and others 2011). The online web server Synteny (http://pipeline.lbl.gov/blockview/blockview/StartView.html) was used to indentify regions where a locus showed tandem and segmental duplications. Genes were designated as segmental duplications provided that they were co-paralogs and were located on duplicated chromosomal blocks as proposed by Wei and others (2007). Paralogs were regarded as tandem duplicated genes if two genes were separated by five or fewer genes(Wang and others 2010).
The multiple expectation maximization for motif elicitation (MEME) web server displayed motifs in all of the predicted DCL, AGO, and RDR proteins (http://meme.sdsc.edu/meme4_3_0/cgi-bin/meme.cgi) (Bailey and others 2009). Parameters were set such that optimum motif widths were ≥6 and ≤200 and that maximum number of motifs was 20. Motifs that did not belong to structural domain in each protein family were rejected. The Pfam database was used to annotate the motifs identified by MEME (Qian and others 2011).
Expression Profile Analysis of Grapevine DCL, AGO, and RDR Genes in Silico
The dbEST database was used to analyze the expression profiles of the VvDCLs, VvAGOs, and VvRDRs by searching the annotated grapevine ESTs (http://compbio.dfci.harvard.edu/tgi/plant.html) (Guo and others 2008). Probe-set IDs were obtained using the new Vitis Affymetrix (Santa Clara, CA, USA) Gene Chip® oligonucleotide microarray version 1.0 (Cramer and others 2007) which was used to prepare a compendium of transcriptome profiles for during berry development and different stress responses at different time stages in grapevine. Berry development dates presented on the array representing the VvDCLs, VvAGOs, and VvRDRs genes were extracted using the VMatch tool available at PLEXdb (http://www.plantgdb.org) (Deluc and others 2007; Wise and others 2007). Data were stored in the Gene Expression Omnibus database at the NCBI under the series accession number GSE36177 which was standardized with GC-RMA algorithm. Expression values were calculated based on the log2 of the ratios between the treatments (water-deficit and salinity) and the control. The values were derived from three biological replicates for each treatment. Expression data for DCL, AGO, and RDR genes were collected using the unique probe-set IDs. The heatmaps were drawn with Cluster 3.0 (Chen and others 2011).
Results
Identification and Structural Organization Analysis of Grapevine DCL, AGO, and RDR Genes
The Hidden Markov model (HMM) profile analyses identified four genes encoding DCL proteins (VvDCLs), thirteen encoding Argonaute proteins (VvAGOs), and five genes for RDRs (VvRDRs) in the grapevine genome database. The candidate genes identified in this study are listed in Table 1.
In this study, we predicted DCL family in grapevine with totally four members. The DCL proteins in grapevine range from 1340 to 1688 amino acids (aa) in length. The PIs of four VvDCL genes were determined to be above 6.0, and VvDCL2 had a PI of 7.21. From the Pfam and Phytozome databases, four DCL gene loci were confirmed as VvDCL genes in the grapevine genome based on analysis of all six types of conserved domains (DExD, Helicase-C, DUF283, PAZ, RNase III, and dsRB) from the putative polypeptide sequences. However, all DCL family proteins contained two RNase III domains except VvDCL1, which contained only one. Both DCL2 and DCL3 clades lacked a dsRB domain. VvDCL4 lacked a PAZ domain, but contained two dsRB domains.
Thirteen Argonaute (AGO) genes were identified in grapevine in the current study. These genes coded for approximately 100 kDa basic proteins with pIs ranging from 8.79 to 9.60. At the level of gene structure, the number of introns varied from two to 22. The lengths of the VvAGO ORFs ranged from 1,329 bp for VvAGO12 to 3,117 bp for VvAGO1, encoding potential proteins of 442 and 1,038 amino acids, respectively (Table 1). Searches of the NCBI databases for conserved domains revealed that all VvAGOs shared an N-terminus PAZ domain and a C-terminus PIWI domain, characteristic of other plant AGO proteins. Furthermore, previous studies have shown that the PIWI domain shares extensive homology with RNaseH, binds the siRNA 5′ end to the target RNA (Hock and Meister 2008), and cleaves target RNAs that exhibit sequence complementary to small RNAs (Baumberger and Baulcombe 2005; Rivas and others 2005). Previous studies of Argonaute proteins in Arabidopsis, rice, maize, and tomato showed that the PIWI domain folds in a manner similar to that of RNase H enzymes and exhibits endonuclease activity owing to an active site usually containing an aspartate–aspartate-histidine (DDH) motif and a conserved histidine at position 798 (H798) (Kapoor and others 2008). This DDH motif/H798 histidine was found in Arabidopsis AGO1 and is critical for the endonuclease activity of AGO1 in vitro (Baumberger and Baulcombe 2005; Kapoor and others 2008). To readily infer whether VvAGOs possess these conserved catalytic residues and could potentially act as slicer components of silencing effector complexes, we aligned the PIWI domains of all the AGOs using CLUSTAL-X as described in Kapoor and others (2008) (Fig. 1).
Amino acid alignment of PIWI domains of grapevine and Arabidopsis AGO proteins with Clustal-X (1.83). Amino acid positions corresponding to the beginning and end of the PIWI domains in each protein are shown. The conserved DDH triad residues corresponding to D760, D845, and H986 of Arabidopsis AGO1 are shown in gray, and the conserved H residues corresponding to H798 of Arabidopsis AGO1 are boxed
Seven VvAGO proteins (VvAGO1, VvAGO5, VvAGO7, VvAGO9, VvAGO10, VvAGO12, and VvAGO13) possessed conserved DDH/H798 residues. D760 and D845 were conserved in AGO proteins from grapevine and Arabidopsis. H986 was also conserved in most grapevine VvAGOs, but it was replaced by aspartate in VvAGO2a, VvAGO2b, VvAGO3, AtAGO2, and AtAGO3. Interestingly, these proteins all belong to the ZIPPY/AGO7 clade (see below). Similarly, compared with previous research that identified a variety of H798 sites in monocots (Kapoor and others 2008; Qian and others 2011). The AGO4 group of proteins in grapevine (VvAGO4, VvAGO6, VvAGO8, VvAGO11) had H798 sites that were replaced by proline (Table 2).
In accord with previous studies, all five of the RDR genes present in the grapevine genome encode predicted proteins that share a common RdRP domain corresponding to the catalytic β′ subunit of DNA-dependent RNA polymerases (Iyer and others 2003). The lengths of the VvRDR ORFs ranged from 2,760 bp for VvRDR1a to 3,384 bp for VvRDR2, encoding predicted polypeptides of 919 and 1,127 amino acids, respectively. Remarkably, VvRDR3 encodes an open reading frame of 2790 bp containing 17 introns (Table 1), which was the highest number of introns predicted in grapevine RDR genes. Two of the grapevine RDR genes (VvRDR1a and VvRDR1b) contained two predicted RdRP domains; the others contained a unique RdRP domain.
Phylogenetic Analysis of DCL, AGO, and RDR Genes in Grapevine, Rice, Tomato, and Arabidopsis
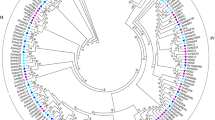
To explore the evolutionary relationships between monocots and dicots in AGO, DCL, and RDR families, we chose three monocots (foxtail millet, maize, and rice) and three dicots (grapevine, Arabidopsis, and tomato) to construct an unrooted tree for each from alignments of the full-length protein sequences (Fig. 2). From phylogenetic analysis of the six plants, dicot and monocot AGO genes revealed that AGO genes clustered into four clades, AGO1, MEL1/AGO5, ZIPPY/AGO7, and AGO4 with well-supported bootstrap values (Fig. 2a). In the AGO1 clade, three grapevine genes were named VvAGO1, VvAGO10a, and VvAGO10b based on their high sequence homologies with AtAGO1 and SlAGO10. The MEL/AGO5 clade comprised proteins similar to AtAGO5 and contained only one grapevine protein, which was named VvAGO5. No grapevine AGO shared a high level of sequence similarity with OsAGO11-14, so the second grapevine protein in the MEL/AGO5 clade was named VvAGO9. The third clade comprised proteins similar to SlAGO7 and AtAGO7 and contained one grapevine protein, which was named VvAGO7. Based on sequence comparisons, three grapevine proteins clustered with two Arabidopsis proteins (AtAGO2, AtAGO3) and two rice proteins (OsAGO2, OsAGO3) in the ZIPPY/AGO7 subfamily, and the respective genes were designated VvAGO2a, VvAGO2b, and VvAGO3, based on their high sequence similarities with AtAGO2 and AtAGO3, respectively. The AGO4 clade comprised 18 proteins, and the four grapevine genes were named VvAGO4, VvAGO8, VvAGO6, and VvAGO11. In this subgroup, VvAGO4 and VvAGO11 were highly similar to AtAGO4 and AtAGO16. In addition, no grapevine AGO displayed a high similarity with OsAGO17 and OsAGO18.
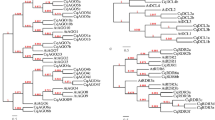
Phylogenetic analysis of Dicer-like, Argonaute, and RDR genes of grapevine and five other plant species. Unrooted neighbor-joining (NJ) phylogenetic trees of grapevine, tomato, Arabidopsis, foxtail millet, maize, and rice proteins; a AGO, b DCL, and c RDR. And the phylogenetic trees were constructed using MEGA v4.0 software. The grapevine DCLs, AGOs, or RDRs have been highlighted in purple for each group. Each gene family is divided into four clades (Kapoor and others 2008), which are shown in different colors. Sequences of tomato, Arabidopsis, foxtail millet, maize, and rice were downloaded from the Phytozome v9.1 database (Supplementary Table S1)
Phylogenetic relationships were analyzed by conserved structural alignments. The four candidate VvDCL genes were assigned to four groups in the unrooted tree, and the respective genes were designated VvDCL1, VvDCL2, VvDCL3, and VvDCL4 (Fig. 2b). Interestingly, in DCL1, DCL3, and DCL4 subgroups, VvDCL1 VvDCL3 and VvDCL4 were closely allied with SlDCL1, SlDCL3, and SlDCL4 with high similarity, respectively. Moreover, the other three subfamilies each contained one identified VvDCL protein, which have been named based on sequence similarity with their counterparts in rice and Arabidopsis.
The unrooted phylogenetic tree, generated from aligned full-length protein sequences of all five VvRDRs, five AtRDRs, five OsRDRs, five ZmRDRs, 11 SiRDRs, and six SlRDRs, grouped grapevine, Arabidopsis, rice, maize, foxtail millet, and tomato RDR proteins into four subfamilies, RDR1, RDR2, RDR3, and RDR4 (Fig. 2c). The predicted product of the newly identified VvRDR2 locus grouped closely with the highly similar SlRDR2 and AtRDR2 proteins. Other grapevine RDR genes were designated VvRDR1a, VvRDR1b, VvRDR2, VvRDR3, and VvRDR6 based on their similarities with tomato, and Arabidopsis RDR proteins.
Chromosomal Location and Gene Duplication of Grapevine DCL, AGO, and RDR Genes
To investigate the evolution of multiple DCLs, AGOs, and RDRs in grapevine, we analyzed their genomic distribution by localizing the genes on grapevine chromosomes (Fig. 3; Table 1). Among the nineteen grapevine chromosomes, four DCL genes, 13 AGO genes, and five RDR genes were found to be distributed unevenly across 11 chromosomes of the grapevine genome (Fig. 3). Four VvDCL genes were distributed on three chromosomes; VvDCL2 and VvDCL3 were on chromosome four, and VvDCL1 and VvDCL4 were on chromosomes 11 and 15, respectively. In this family, none of the VvDCL genes seems to have undergone tandem duplications or segmental duplications.
Genomic distribution of DCL, AGO, and RDR genes on the chromosomes of grapevine. Chromosome numbers are shown at the top of each chromosome. Different gene families are indicated by different colors. Tandem gene duplications are indicated by filled dots. Genes involved in segmental duplication are joined by dashed lines
Thirteen VvAGO genes were distributed on nine chromosomes; three (VvAGO2a, VvAGO2b, and VvAGO3) were detected on chromosome 10, two were located on chromosomes six and eight, respectively, and chromosomes 1, 5, 11, 12, 13, and 17 each contained a single VvAGO gene. VvAGOs were not present on 2, 3, 4, 7, 9, 14, 15, 16, 18, and 19 chromosomes. Alignment analysis of the protein sequences showed that there are high degrees of similarity among VvAGOs belonging to the same clade, which is consistent with previous findings in tomato (Bai and others 2012). Three pairs of grapevine Argonaute genes, VvAGO10a/-10b, VvAGO4/-8, and VvAGO5/-9, were found to be located in duplicated genomic segments and shared 71.4, 73.4, and 48.6 % identity at the amino acid level between the pair partners. The members of each pair of two genes were located on different chromosomes but belong to same clade. While the VvAGO2a/-2b/-3 genes were detected on chromosome 10, they appear to have undergone tandem duplication on the basis of 61.7–83.3 % amino acid similarity.
Localization of VvRDR genes on the grapevine chromosomes indicated that the five VvRDRs are distributed on four chromosomes. VvRDR1a and VvRDR1b are located on chromosome one, and VvRDR2, VvRDR3, and VvRDR6 are on chromosomes 17, 11, and four, respectively. VvRDR1a and VvRDR1b, which share 83.4 % amino acid identity, appear to represent a tandem duplication on chromosome one. The other RDR protein genes were not found to be located in duplicated genome segments.
Conserved Motif Analysis of the Grapevine DCL, AGO, and RDR Genes
Based on the MEME motif analysis, the majority of protein motifs were conserved in the DCL family, and the order of these motifs were maintained between the four DCL subfamilies from Arabidopsis, rice, tomato, and grapevine. The results are shown in Fig. 4. Motifs 10, 11, and 12 of the RNase III domain were absent only in VvDCL1. In VvDCL2, motif 1, belonging to the RNase III domain, was not found, as was motif 7 of the dsRB domain. VvDCL3 and VvDCL4 contain the same general types of motifs as compared with the other genes in the DCL3 and DCL4 subgroups, respectively (Fig. 4b).
Distribution of conserved motifs in grapevine DCL, AGO, and RDR proteins. Schematic representation of the predicted conserved motifs in the DCL, AGO, and RDR proteins from grapevine was elucidated using the MEME motif search tool for each protein family. Motifs were rejected that did not belong to the structural domains for each protein family. Different color boxes represent different motifs. Box length and position correspond to motif length and position in the individual protein sequences. Details of the individual motifs are provided in Supplementary Table S2
The MEME analyses of the AGO family identified 11 conserved motifs among all the AGO proteins. Although the motif configurations identified by MEME reflected conservation and specificity within the AGO families of Arabidopsis, rice, tomato, and grapevine, some variability is present between different subfamilies in the individual members of the AGO family we detected (Fig. 4a). For example, the absence of motif 2 was only found in AGO4 subfamilies, except for AtAGO6 and VvAGO6. Nine AGO members shared the absence of motif 10 that is present in most PIWI domains of the other AGO subfamilies, which was detected by Pfam analyses. In four AGO subfamilies, a subset of AGO members shared duplication of one or two motifs. For example, distinct copies of the newly duplicated motifs 1 and 3 are located in the DUF1785 domain of OsAGO13 in the MEL/AGO5 subgroup, and the PAZ domain of OsAGO15 in the AGO4 subgroup, respectively. However, the other duplicated motifs are located in PIWI domain.
In the RDR protein family, 14 motifs were characterized as being conserved by MEME analysis (Fig. 4c). Among them, motifs 10, 11, 13, and 14 were conserved and identified as major motifs of the RDR domain. However, the protein motif patterns of the individual RDR family members did not always follow the same rules in all subgroups. For example, motifs 7, 8, and 9 shared distinct diversification in the RDR class III subfamily, which are different from all other RDR family subfamily members. Motif 6 was completely absent in RDRI subfamily members from grapevine, and motifs 10 and 11 were duplicated in proteins from Arabidopsis in subgroups RDR3 and RDR6, respectively.
Expression Patterns of Grapevine DCL, AGO, and RDR Genes
We analyzed the expression of small RNAs in grapevine by searching EST databases. These small RNAs could be divided into nine classes, depending on the organ or tissue in which they were expressed (Fig. 5). The twenty-two grapevine genes involved in small RNA regulation that were investigated in this study were not expressed in all nine tissues or organs examined. Rather, expression for each gene was selective to specific groups of tissues or organs. The most widely expressed genes in grapevine were VvAGO1, 8, and 10a which were expressed in six tissues or organs each. There were three genes (VvDCL4, VvAGO10b, and VvRDR1b) that showed significant levels of expression in only one tissue or organ. The remaining genes were found to be expressed in two or more tissues or organs. It is worth mentioning that we detected expression of VvAGO10a only in the stem. Generally, the highest levels of gene expression were in mature tissues and organs, such as the fruit, flower, inflorescence, and leaf; lower levels were found in stem, seed, pericarp, and embryo. Our results showed that siRNAs appear to regulate function more during the mature growth stages in grapevine.
Expression analysis of VvDCL, VvAGO, and VvRDR genes in different tissues and organs in grapevine. Searches for EST sequences corresponding to the grapevine VvDCL, VvAGO, and VvRDR genes were made in the TIGR gene index database (http://compbio.dfci.harvard.edu/tgi/plant.html) using the BLASTn program with default parameters and the sequences of VvDCL, VvAGO, and VvRDR genes as query sequences. The empty cells represent a lack of ESTs for that gene under that condition in the database
To analyze expression profile of DCL, AGO, and RDR genes during berry development stages in grapevine, microarray datasets were utilized. Of all the genes, the expression profile of only one (VvRDR6) was not obtained from the NCBI database, and two genes (VvAGO9 and VvAGO10b) have a common probe-set ID in the Affymetrix GeneChip® Grapevine Genome Arrays. Grapevine berries undergo a complex series of physical and biochemical changes during development. These changes can be divided into three major phases with more detailed descriptive designations known as the modified E-L system, to define more precisely the growth stages through an entire grapevine lifecycle (Guo and others 2008). Expression profile of almost all of the genes in different families or subfamilies was showed from Fig. 6, we suggested that VvDCLs, VvAGOs, and VvRDRs play important roles in grapevine berry development. All genes (VvAGO4, VvAGO6, VvAGO8, and VvAGO11) in AGO4 subfamily showed high expression during the three phases. In addition,VvDCL2, VvRDR3, VvAGO2b, and VvAGO10a displayed low expression, but these genes had a greater change in the amplitude during fruit development.
Expression profiles of grapevine DCL, AGO, and RDR genes during seven stages of berry development from small pea size berries (E-L stages 31–33 as defined by the modified E-L system) to veraison (E-L stages 34 and 35) and mature berries (E-L stages 36 and 38). The microarray data were extracted using the VMatch tool available at PLEXdb (http://www.plantgdb.org) (Supplementary Table S3). The y-axis indicates the relative expression level, and error bars represent standard deviation calculated based on three biological replicates
The relative transcription of grapevine DCL, AGO, and RDR genes at different growth stages under different abiotic stress conditions was investigated using microarray data. Gene expression levels for 21 genes is shown in Fig. 7 under two abiotic stress conditions (salt and water-deficit) for four different time points (4, 8, 12, and 16 days). Most of genes were low expression in different stages of abiotic stress conditions relative to control. Salt and water-deficit stress caused upregulation of three genes on 16 days, including VvAGO2a, VvAGO4, and VvRDR2 compared with control. However, salt and water-deficit stress caused downregulation of seven genes (VvDCL1, VvAGO8, VvAGO9, VvAGO11, VvAGO5, VvRDR1a, and VvRDR3) on 16 days compared with control. However, VvRDR1a and VvRDR3 showed downregulation expression on 16 days in salt condition.
Expression profiles of grapevine DCL, AGO, and RDR genes in response to different abiotic stresses. Heatmap showing hierarchical clustering of DCL, AGO, and RDR genes expression under salt stress and water-deficit stress for various periods of time. The Affymetrix microarray data were obtained from the NCBI Gene Expression Omnibus (GEO) database under the series accession number GSE31677 (Supplementary Table S4). Gene expression values were calculated based on the ratios between the treatments (water-deficit and salinity) and the control. The color bar in each panel represents log2 expression values, blue represents a low level, and red indicates a high level of transcript abundance. Abiotic stress stages used for expression profiling are mentioned at the top of each column. Numbers (0, 4, 8, 12, and 16) represent days in the stress treatments. A gene cluster dendrogram is shown to the left of each expression heatmap
Discussion
RNA silencing plays an important role in regulating gene expression at the posttranscriptional level, chromatin modification during vegetative and reproductive development, and gene expression profiles during various abiotic stress conditions in plants. In recent years, the molecular basis of these complicated and interconnected pathways has been clarified. In the present investigation, we identified gene families in grapevine and analyzed their chromosomal distribution in the genome, conducted a phylogenetic analysis for each gene family and defined gene multiplicity. EST expression data analysis and microarray-based expression analysis shed light on the contribution of the activity of these proteins.
Dicer-Like Proteins in Grapevine
Dicer-like (DCL) proteins are the key enzymes involved in miRNA and siRNA biogenesis pathways in processing long double-stranded RNAs into mature small RNAs (Millar and Waterhouse 2005; Chapman and Carrington 2007; Großhans and Filipowicz 2008). In this study, based on phylogenetic analysis of DCL genes from three monocots (foxtail millet, maize, and rice) and three dicots (grapevine, Arabidopsis, and tomato), we found that VvDCL were more closely related with SlDCL genes and monocots and dicots were distinct displayed on different branches in every subfamilies. This suggested that DCL genes existed before the divergence of monocots and dicots. Also, the DCL gene families in grape and tomato more likely descended from a common dicot ancestor. Therefore, we inferred that the two dicot species had similarities in evolution, and they may have an analogous mode of selection and evolution.
The MEME analysis revealed that the majority of DCL genes in the same subfamily have similar protein motifs. However, the RNase III domain was shown to have a few C-terminal motifs rearranged, one (motif 1) had shifted to the N-terminal, and others (motif 9,motif 10,motif 11, and motif 12)were still located at the C-terminal region. MEME analyses clearly demonstrated that these motifs are well conserved and that the conserved motifs play crucial roles in subfamily-specific functions. In view of these similarities, we suggest a similar evolutionary alliance in the functional diversification of the grapevine DCL gene family with tomato, rice, and Arabidopsis. However, few reports of the biochemical and genetic analysis of DCL genes in grapevine are available. Therefore, these data offer a basis for continued research on the grapevine DCL gene family.
Argonaute Proteins in Grapevine
Argonaute (AGO) RNA-binding proteins are involved in RNA silencing. Here, we investigated all four subfamilies of AGO genes in the grapevine genome. Based on the phylogenetic analysis of AGO genes from grapevine, Arabidopsis, tomato, foxtail millet, maize, and rice, we found that VvAGO genes are more similar to the SlAGO and AtAGO genes, and monocots and dicots were clear displayed on different branches in every subfamilies. This implies that the AGO genes underwent diversification before the divergence of monocots and dicots. It is interesting to note that VvAGO2a and VvAGO2b are appeared on same branch. Therefore, we infer that pairs of paralogous VvAGO2a and VvAGO2b may generate by recent duplication event.
Gene duplication is believed to play a significant role in gene family evolution and expansion (Taylor and Raes 2004) and can be divided into tandem and segmental duplication events. Gene duplication events were found in the AGO gene family, and three gene pairs (VvAGO10a/-10b, VvAGO4/-8, and VvAGO5/-9) were assigned to segmental duplication events; previous studies showed similar duplications for two gene pairs in rice and three pairs in maize, indicating that segmental duplication could be an important mechanism in the expansion of the AGO gene family. While one group (VvAGO2a/-2b/-3) was considered to be the result of a tandem duplication event, the result is congruent with the SlAGO2a/-2b/-3 duplications, and the genes have a close phylogenetic relationship. It has been suggested that they may share the same function in evolution in dicotyledonous plants. The protein motif schemes of the individual AGO family members clearly demonstrated structural similarities among the proteins within the four species examined in this study. Furthermore, motif analysis displayed the genes that shared similar structures in each subfamily, suggesting that the motifs might share major functional roles in these proteins.
The endonuclease function of AGO proteins includes RNAi residues in the PIWI domain that contain three conserved metal-chelating amino acids (DDH). In Arabidopsis, although the catalytic residues are conserved, many AGO proteins are endonucleolytically inactive. Also, in the model plant Arabidopsis, there are six AGO genes which do not contain the conserved catalytic residues found in most AGO proteins. We confirmed that seven genes lack the conserved catalytic residues in the grapevine PIWI domains. The absence of conserved catalytic residues could lead to loss of function of target RNA processing by endonucleolytic cleavage in these AGO proteins (Kapoor and others 2008). It will be interesting to find out to what extent selective recruitment of siRNA and miRNA plays a role in the diversification of AGO protein function and on RNA silencing pathways.
RNA-Dependent RNA Polymerase Proteins in Grapevine
In this study, five grapevine RDR gene family members were identified. The phylogenetic analysis showed that VvRDR2 and VvRDR6 are similar to SlRDR2 and AtRDR6, and they share sequence similarities of 67.08 and 61.44 % at the amino acid level, respectively. This finding indicates that VvRDR2 and VvRDR6 should share similar functions with SlRDR2 and AtRDR6 in plant growth and development. One pair of tandemly duplicated genes (VvRDR1a/-1b) was found that shared 83.4 % identity at the amino acid level. This result is incongruent with the RDR families in rice, tomato, and maize, which are without gene duplication events. The gene duplication event suggests that isoform duplication may play a role in gene expansion in the RDR gene family in grapevine. From MEME analysis of the conserved motifs in RDR proteins of grapevine and other species, it has been suggested that functional diversification in RDR protein families is similar, and the RDR gene families of grapevine and other species diverged from the same common ancestor. The observed conservation and widespread distribution throughout the subfamily suggest that the motifs are important in protein function.
Expression Profiles of VvDCL, VvAGO, and VvRDR genes
In silico gene expression data from EST databases can provide information that is important for continued research. In this study, an EST database was used to determine potential gene expression profiles. Most VvDCL, VvAGO, and VvRDR genes were expressed in most organs and tissues. Interestingly, VvAGO10b had no specific expression in the EST database except in stem tissues. One explanation is that the gene shows a tissue-specific expression pattern and may function in the regulation of siRNAs in the grapevine stem. In addition, a few of genes that we confirmed had apparent rearrangements in the gene or partial motif absences in the gene structure. These changes in structure might lead to distinct expression patterns, and the genes were also determined to be part of pseudogenes (Qian and others 2011). For example, in the ZIPPY/AGO7subfamily, VvAGO7 had been appeared obvious rearrangement in domain and lacked motif 5 and 6, which suggested that this was a result that lead to VvAGO7 not expression in fruit.
Expression profiles of grapevine DCL, AGO, and RDR genes during berry development were displayed. It was worth noting that the AGO4 subfamily had higher expression during seven stages of berry development. This suggests that the genes in AGO4 subfamily involved in fruit physiology and development are interesting targets for functional characterization. Microarray expression analyses showed that grapevine VvDCL, VvAGO, and VvRDR genes exhibited different expression levels under two different abiotic stress treatments at various stages. Some genes, such as VvDCL2, VvDCL4, VvAGO11, and VvAGO7, clearly showing increased levels of expression at specific times during stress treatments had a high possibility to contribute to grapevine RNAi regulation by only expressing under specific conditions or in specific time. Although three genes (VvAGO2a, VvAGO4, and VvRDR2) showed upregulation under salt and water-deficit stress on 16 days, the variation of candidate genes at 16 days and different stresses suggested the genes played a significant role of during growing and development in the specific timing. These data would expedite further functional characterization of VvDCL, VvAGO, and VvRDR genes in response to stress treatments which may deepen the understanding of complex stress responsive network equipped by plants.
Semi-quantitative and quantitative real-time PCR analysis of DCL, AGO, and RDR genes in rice, maize, and tomato (Kapoor and others 2008; Qian and others 2011a; Bai and others 2012) demonstrated that different growth environments may also affect gene expression. The comparative and phylogenetic analyses of the VvDCL, VvAGO, and VvRDR gene families, and the expression and structural analysis of the VvDCL, VvAGO, and VvRDR genes will lay the foundation for further functional genetic studies in grapevine.
References
Alves VAF, Castelo A, Vianna M, Taromaru E, Namiyama G, Lorincz A, Dores G (2006) Performance of the DNA-Citoliq liquid-based cytology system compared with conventional smears. Cytopathology 17:86–93
Bai M, Yang GS, Chen WT, Mao ZC, Kang HX, Chen GH, Yang YH, Xie BY (2012) Genome-wide identification of Dicer-like, Argonaute and RNA-dependent RNA polymerase gene families and their expression analyses in response to viral infection and abiotic stresses in Solanum lycopersicum. Gene 501:52–62
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren JY, Li WW, Noble WS (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37:W202–W208
Baumberger N, Baulcombe D (2005) Arabidopsis ARGONAUTE1 is an RNA Slicer that selectively recruits microRNAs and short interfering RNAs. Proc Natl Acad Sci USA 102:11928–11933
Carmell MA, Xuan Z, Zhang MQ, Hannon GJ (2002) The Argonaute family: tentacles that reach into RNAi, developmental control, stem cell maintenance, and tumorigenesis. Genes Dev 16:2733–2742
Chapman EJ, Carrington JC (2007) Specialization and evolution of endogenous small RNA pathways. Nat Rev Genet 8:884–896
Chen Q, Yang L, Ahmad P, Wan X, Hu X (2011) Proteomic profiling and redox status alteration of recalcitrant tea (Camellia sinensis) seed in response to desiccation. Planta 233:583–592
Cramer GR, Ergül A, Grimplet J, Tillett RL, Tattersall EA, Bohlman MC, Vincent D, Sonderegger J, Evans J, Osborne C (2007) Water and salinity stress in grapevines: early and late changes in transcript and metabolite profiles. Funct Integr Genomics 7:111–134
Deluc LG, Grimplet J, Wheatley MD, Tillett RL, Quilici DR, Osborne C, Schooley DA, Schlauch KA, Cushman JC, Cramer GR (2007) Transcriptomic and metabolite analyses of Cabernet Sauvignon grape berry development. Bmc Genomics 8(1):429
Diaz-Pendon JA, Li F, Li W-X, Ding S-W (2007) Suppression of antiviral silencing by cucumber mosaic virus 2b protein in Arabidopsis is associated with drastically reduced accumulation of three classes of viral small interfering RNAs. Plant Cell Online 19:2053–2063
Dlakić M (2006) DUF283 domain of Dicer proteins has a double-stranded RNA-binding fold. Bioinformatics 22:2711–2714
Fagard M, Boutet S, Morel J-B, Bellini C, Vaucheret H (2000) AGO1, QDE-2, and RDE-1 are related proteins required for post-transcriptional gene silencing in plants, quelling in fungi, and RNA interference in animals. Proc Natl Acad Sci 97:11650–11654
Großhans H, Filipowicz W (2008) The expanding world of small RNAs. Nature 451:414–416
Guo AY, Zhu QH, Gu X, Ge S, Yang J, Luo J (2008) Genome-wide identification and evolutionary analysis of the plant specific SBP-box transcription factor family. Gene 418:1–8
Hannon GJ (2002) RNA interference. Nature 418:244–251
He H, Dong Q, Shao Y, Jiang H, Zhu S, Cheng B, Xiang Y (2012) Genome-wide survey and characterization of the WRKY gene family in Populus trichocarpa. Plant Cell Rep 31:1199–1217
Hock J, Meister G (2008) The Argonaute protein family. Genome Biol 9:210
Hunter C, Sun H, Poethig RS (2003) The Arabidopsis heterochronic gene ZIPPY is an ARGONAUTE family member. Curr Biol 13:1734–1739
Iyer LM, Koonin EV, Aravind L (2003) Evolutionary connection between the catalytic subunits of DNA-dependent RNA polymerases and eukaryotic RNA-dependent RNA polymerases and the origin of RNA polymerases. BMC Struct Biol 3:1
Kapoor M, Arora R, Lama T, Nijhawan A, Khurana JP, Tyagi AK, Kapoor S (2008) Genome-wide identification, organization and phylogenetic analysis of Dicer-like, Argonaute and RNA-dependent RNA Polymerase gene families and their expression analysis during reproductive development and stress in rice. BMC genomics 9:451
Kasschau KD, Fahlgren N, Chapman EJ, Sullivan CM, Cumbie JS, Givan SA, Carrington JC (2007) Genome-wide profiling and analysis of Arabidopsis siRNAs. PLoS Biol 5:e57
Liu B, Chen Z, Song X, Liu C, Cui X, Zhao X, Fang J, Xu W, Zhang H, Wang X (2007) Oryza sativa dicer-like4 reveals a key role for small interfering RNA silencing in plant development. The Plant Cell Online 19:2705–2718
Lynn K, Fernandez A, Aida M, Sedbrook J, Tasaka M, Masson P, Barton MK (1999) The PINHEAD/ZWILLE gene acts pleiotropically in Arabidopsis development and has overlapping functions with the ARGONAUTE1 gene. Development 126:469–481
MacRae IJ, Zhou K, Li F, Repic A, Brooks AN, Cande WZ, Adams PD, Doudna JA (2006) Structural basis for double-stranded RNA processing by Dicer. Science 311:195–198
Margis R, Fusaro AF, Smith NA, Curtin SJ, Watson JM, Finnegan EJ, Waterhouse PM (2006) The evolution and diversification of Dicers in plants. FEBS Lett 580:2442–2450
Millar AA, Waterhouse PM (2005) Plant and animal microRNAs: similarities and differences. Funct Integr Genomics 5:129–135
Moussian B, Schoof H, Haecker A, Jürgens G, Laux T (1998) Role of the ZWILLE gene in the regulation of central shoot meristem cell fate during Arabidopsis embryogenesis. EMBO J 17:1799–1809
Peters L, Meister G (2007) Argonaute proteins: Mediators of RNA silencing. Mol Cell 26:611–623
Qi X, Bao FS, Xie Z (2009) Small RNA deep sequencing reveals role for Arabidopsis thaliana RNA-dependent RNA polymerases in viral siRNA biogenesis. PLoS ONE 4:e4971
Qian Y, Cheng Y, Cheng X, Jiang H, Zhu S, Cheng B (2011) Identification and characterization of Dicer-like, Argonaute and RNA-dependent RNA polymerase gene families in maize. Plant Cell Rep 30:1347–1363
Rivas FV, Tolia NH, Song JJ, Aragon JP, Liu J, Hannon GJ, Joshua-Tor L (2005) Purified Argonaute2 and an siRNA form recombinant human RISC. Nat Struct Mol Biol 12:340–349
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Taylor JS, Raes J (2004) Duplication and divergence: the evolution of new genes and old ideas. Annu Rev Genet 38:615–643
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Wang MB, Metzlaff M (2005) RNA silencing and antiviral defense in plants. Curr Opin Plant Biol 8:216–222
Wang L, Guo K, Li Y, Tu Y, Hu H, Wang B, Cui X, Peng L (2010) Expression profiling and integrative analysis of the CESA/CSL superfamily in rice. BMC Plant Biol 10:282
Wassenegger M, Krczal G (2006) Nomenclature and functions of RNA-directed RNA polymerases. Trends Plant Sci 11:142–151
Wei F, Coe E, Nelson W, Bharti AK, Engler F, Butler E, Kim H, Goicoechea JL, Chen M, Lee S (2007) Physical and genetic structure of the maize genome reflects its complex evolutionary history. PLoS Genet 3:e123
Wise RP, Caldo RA, Hong L, Shen L, Cannon E, Dickerson JA (2007) Barleybase/plexdb. In: eds. Plant bioinformatics. Springer, pp. 347-363
Xie Z, Johansen LK, Gustafson AM, Kasschau KD, Lellis AD, Zilberman D, Jacobsen SE, Carrington JC (2004) Genetic and functional diversification of small RNA pathways in plants. PLoS Biol 2:e104
Yadav CB, Muthamilarasan M, Pandey G, Prasad M (2014) Identification, characterization and expression profiling of Dicer-like, Argonaute and RNA-dependent RNA polymerase gene families in foxtail millet. Plant Mol Biol Rep. doi:10.1007/s11105-014-0736-y
Zhao Y, Zhou YQ, Jiang HY, Li XY, Gan DF, Peng XJ, Zhu SW, Cheng BJ (2011) Systematic analysis of sequences and expression patterns of drought-responsive members of the HD-Zip gene family in maize. Plos One 6:e28488
Zilberman D, Cao X, Jacobsen SE (2003) ARGONAUTE4 control of locus-specific siRNA accumulation and DNA and histone methylation. Science 299:716–719
Zilberman D, Cao X, Johansen LK, Xie Z, Carrington JC, Jacobsen SE (2004) Role of Arabidopsis ARGONAUTE4 in RNA-Directed DNA Methylation Triggered by Inverted Repeats. Curr Biol 14:1214–1220
Acknowledgments
This work was supported by the National Natural Science Foundation (31370561), Specialized Research Fund for the Doctoral Program of Higher Education (20133418110005), Anhui Provincial Natural Science Foundation (1308085MC36), and Anhui Agricultural University disciplinary construction Foundation (XKTS2013001). We thank to the members of the Laboratory of Modern Biotechnology and especially thank Guo Wei and YuanHua Shao for their assistance in this study.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, H., Zhao, K., Wang, J. et al. Comprehensive Analysis of Dicer-Like, Argonaute, and RNA-dependent RNA Polymerase Gene Families in Grapevine (Vitis Vinifera). J Plant Growth Regul 34, 108–121 (2015). https://doi.org/10.1007/s00344-014-9448-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00344-014-9448-7