Abstract
Branched-chain amino acids (BCAAs), viz., l-isoleucine, l-leucine, and l-valine, are essential amino acids that cannot be synthesized in higher organisms and are important nutrition for humans as well as livestock. They are also valued as synthetic intermediates for pharmaceuticals. Therefore, the demand for BCAAs in the feed and pharmaceutical industries is increasing continuously. Traditional industrial fermentative production of BCAAs was performed using microorganisms isolated by random mutagenesis. A collection of these classical strains was also scientifically useful to clarify the details of the BCAA biosynthetic pathways, which are tightly regulated by feedback inhibition and transcriptional attenuation. Based on this understanding of the metabolism of BCAAs, it is now possible for us to pursue strains with higher BCAA productivity using rational design and advanced molecular biology techniques. Additionally, systems biology approaches using augmented omics information help us to optimize carbon flux toward BCAA production. Here, we describe the biosynthetic pathways of BCAAs and their regulation and then overview the microorganisms developed for BCAA production. Other chemicals, including isobutanol, i.e., a second-generation biofuel, can be synthesized by branching the BCAA biosynthetic pathways, which are also outlined.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Branched-chain amino acids (BCAAs)
- Corynebacterium glutamicum
- Escherichia coli
- Feedback inhibition
- Isobutanol
- l-Isoleucine
- l-Leucine
- l-Valine
- Metabolic engineering
- Transcriptional attenuation
1 Introduction
Branched-chain amino acids (BCAAs), namely, l-isoleucine, l-leucine, and l-valine, are essential amino acids [1] that are not synthesized in mammals, but have critical roles in physiological functions and metabolism [2]. BCAAs are used in dietary products, pharmaceuticals, and cosmetics and serve as a precursor of antibiotics and herbicides. Moreover, they are expected to play a leading role in future feed additives [3]. As other amino acids, BCAAs have been manufactured by fermentation using mutated or metabolically engineered microorganisms that originated from Corynebacterium glutamicum or Escherichia coli [4, 5]. The amount of the annual production of l-isoleucine, l-leucine, and l-valine in 2001 was approximately 400, 500, and 500 tons, respectively [6, 7], and has been increasing continuously. For example, the nonfeed market for l-valine reached around 1,000–1,500 tons per annum with an estimated 5–8 % annual increase [8].
Historically, BCAAs were produced chemically, and their enantiomers were separated enzymatically after chemical derivatization or via chromatographic separation followed by crystallization. However, as the demand for BCAAs increased, fermentative methods gathered attention for economic reasons as well as from an environmental perspective. In the early stages, most BCAA production strains were isolated by random mutagenesis. However, the random mutagenesis approach distributes genetic alterations throughout the chromosome, which are difficult to identify and may cause unexpected effects. Recently, amino acid-producing strains have been developed by rational genetic manipulation to avoid this problem. Highly productive strains can be created in a genetically defined manner, e.g., by specifically overexpressing biosynthetic genes responsible for a target amino acid. Systems metabolic engineering, which analyzes metabolism in a genome scale [9], also helps us to optimize carbon flux toward the biosynthesis of a target amino acid [10].
In this chapter, we will describe the biosynthetic pathways of BCAAs and their regulation in microorganisms, in particular, C. glutamicum, which is a nonpathogenic Gram-positive bacterium of the family Actinomycetes and is widely used for the industrial production of amino acids and nucleotides [11]. The development of microbial strains for production of BCAAs will then be outlined. Finally, the extrapolated application of the BCAA biosynthetic pathways to other chemicals, including isobutanol, i.e., a second-generation biofuel, will be shown.
2 Regulation of BCAA Biosynthesis
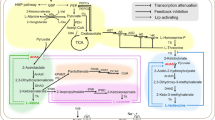
The biosynthetic pathways of BCAAs of C. glutamicum are summarized in Fig. 1. C. glutamicum has one acetohydroxy acid synthase (AHAS) encoded by ilvBN [12], which catalyzes the initial step of the BCAA biosynthesis. AHAS produces 2-acetolactate and 2-aceto-2-hydroxybutyrate for the l-leucine/l-valine and l-isoleucine biosynthesis, respectively. The starting materials for this reaction are pyruvate and 2-ketobutyrate. Pyruvate is provided from the glycolytic pathway, while 2-ketobutyrate is synthesized from l-threonine by threonine dehydratase (TDH) encoded by ilvA. TDH is feedback inhibited by l-isoleucine [13], although the activity can be restored by l-valine [14]. AHAS consists of large and small subunits, which are encoded by ilvB and ilvN, respectively. The small subunit of AHAS is responsible for the multivalent regulation by all three BCAAs [12]. Unlike C. glutamicum, some bacteria have several isoforms of AHAS. For instance, E. coli has three isozymes, AHAS I, II, and III [3], which are encoded by ilvBN, ilvGM, and ilvIH, respectively. The expression of these genes is regulated differently, and ilvGM is attenuated by all BCAAs, while ilvBN is affected only by l-leucine and l-valine [15]. The activity of AHAS I and III is inhibited strongly by l-valine and weakly by l-isoleucine, while l-leucine has no effect [16]. E. coli is very sensitive to l-valine, and its growth is inhibited at an extremely low concentration [17].
The BCAA biosynthetic pathways and their regulation in C. glutamicum. The genes and enzymes are shown in italic font and in parentheses, respectively. Dotted lines and gray lines indicate feedback inhibition and transcriptional attenuation, respectively. Abbreviations: AHAIR acetohydroxy acid isomeroreductase, AHAS acetohydroxy acid synthase, AK aspartate kinase, ASADH aspartate semialdehyde dehydrogenase, BCAT branched-chain amino acid aminotransferase, DHAD dihydroxy acid dehydratase, HDH homoserine dehydrogenase, HK homoserine kinase, IPMD isopropylmalate dehydrogenase, IPMI isopropylmalate isomerase, IPMS isopropylmalate synthase, TDH threonine dehydratase, TrAT tyrosine-repressible transaminase, TS threonine synthase
The ilvC gene encodes acetohydroxy acid isomeroreductase (AHAIR) and is transcripted as an operon with the ilvBN genes [18] in C. glutamicum. AHAIR converts 2-acetolactate to 2,3-dihydroxyisovalerate in the l-valine and l-leucine biosynthesis and 2-aceto-2-hydroxybutyrate to 2,3-dihydroxy-3-methylvalerate in the l-isoleucine biosynthesis by using NADPH as a cofactor. The expression of ilvBNC operon is controlled by transcriptional attenuation, which is mediated by all BCAAs [19].
Dihydroxy-acid dehydratase (DHAD) encoded by ilvD [20] is the enzyme for the next biosynthetic step in which 2-ketoisovalerate and 2-keto-3-methylvalerate are formed from 2,3-dihydroxyisovalerate and 2,3-dihydroxy-3-methylvalerate, respectively. This enzyme is inhibited by either l-valine or l-leucine [21]. Transcriptional regulation of the ilvD gene is unknown.
Branched-chain amino acid transaminase (BCAT) or transaminase B encoded by ilvE [20] is the last player in the l-isoleucine and l-valine biosynthesis. This enzyme transfers the amine moiety of l-glutamate to 2-ketoisovalerate and 2-keto-3-methylvalerate to afford l-valine and l-isoleucine, respectively [22].
The specific pathway of the l-leucine biosynthesis starts from the reaction by isopropylmalate synthase (IPMS) encoded by leuA, which generates 2-isopropylmalate from 2-ketoisovalerate and acetyl-CoA. This enzyme in C. glutamicum is subjected to strong feedback inhibition, and its expression is also regulated by l-leucine [23]. Then, 2-isopropylmalate is isomerized to 3-isopropylmalate by isopropylmalate isomerase (IPMI). IPMI consists of large and small subunits, which are encoded by leuC and leuD, respectively. Next, 3-isopropylmalate is converted to 2-keto-4-methylvalerate by isopropylmalate dehydrogenase (IPMD), which is encoded by leuB. In C. glutamicum, leuB is strongly repressed by l-leucine [24], while the leuABCD genes form an operon in E. coli, and their expression is controlled by the transcriptional attenuation mediated by l-leucine [25]. As the other two BCAAs, l-leucine is formed from 2-keto-4-methylvalerate by the catalysis of BCAT, which is activated by the substrate itself [26]. Additionally, the same reaction is catalyzed by tyrosine-repressible transaminase (TrTA) encoded by tyrB [27].
3 Production of BCAAs by Metabolically Engineered Microorganisms
In this section, we describe the BCAA-producing strains that have been developed to date. Table 1 shows the recent representative strains for BCAA production.
3.1 l-Valine
Microbial production of l-valine was first reported by Udaka and Kinoshita [50] and Sugisaki [51] independently. Udaka and Kinoshita screened a large number of microorganisms, and the selected bacteria, namely, Paracolobacterum coliforme and Brevibacterium ammoniagenes, produced l-valine in 23 % molar yield from glucose. Sugisaki isolated Aerobacter cloacae and A. aerogenes, which produced l-valine in 20 % molar yield from glucose. These initial findings provoked a hunt for better l-valine producers, and auxotrophic mutants or amino acid analog-resistant mutants were included within the research scope. One example of the auxotrophic mutants is the isoleucine auxotrophic mutant of Micrococcus glutamicus [52], which produced more than 10 g/L l-valine when the culture was supplemented with a small amount of dl-isoleucine and dl-valine. An amino acid analog-resistant mutant accumulating l-valine was then found while validating a method to isolate amino acid-producing microorganisms using E. coli ATCC 4157 as a model microbe [53]. This l-valine-producing mutant was originated from a norvaline-resistant strain, which was further mutated to acquire l-leucine auxotrophy. The resultant strain accumulated more than 2 g/L l-valine. An amino acid analog-resistant mutant was also developed using Serratia marcescens [54]. Among several BCAA analogs tested, α-aminobutyric acid conferred mutants that were able to accumulate more than 8 g/L l-valine. An enzymatic analysis revealed that in these mutants, expression of AHAS was derepressed and/or AHAS was only weakly affected by feedback inhibition. Three l-glutamate-producing bacteria, B. lactofermentum, C. acetoacidphilum, and Arthrobacter citreus, were also led to l-valine production mutants using a histidine analog, 2-thiazolealanine [55]. The most efficient strain, one from B. lactofermentum, achieved 31 g/L production in 72 h. In this strain also, AHAS was desensitized from the feedback inhibition by l-valine as well as l-isoleucine and l-leucine. Moreover, the expression of this enzyme was partially derepressed [56]. An interesting example is the α-aminobutyric acid-resistant mutant of the biotin-auxotrophic B. flavum MJ-233 [57]. In the reaction using this mutant, carbon resources were converged to l-valine production when the medium lacked biotin and the bacteria did not grow [58]. This “living cell reaction” process using this strain allowed for accumulation of 300 mM l-valine within one day in 80 % molar yield from glucose with 96 % purity out of the total amino acids.
Genetically defined strains for l-valine production have been developed mostly by using C. glutamicum along with a few examples using other bacteria, including E. coli. The following mainly describes the strains derived from these two bacteria.
The general basic strategy in fermentative production of chemicals is to direct common intermediates for various end products solely to a target compound. In the case of l-valine, this can be achieved by deleting or repressing the transcription of ilvA and panB [20, 28–30, 59], which are the first genes for the l-isoleucine and d-pantothenate biosynthesis, respectively (Figs. 1 and 2). Limitation of d-pantothenate biosynthesis leads to reduction of CoA supply, which is an additional benefit for l-valine production because consumption of pyruvate by pyruvate dehydrogenase complex (PDHC) is suppressed due to the decreased concentration of the reaction partner [20].
The biosynthetic routes branched from the BCAA biosynthetic pathways for production of isobutanol, 3-methyl-1-butanol, 2-methyl-1-butanol, and d-pantothenate. Abbreviations: ADH alcohol dehydrogenase, AHAIR acetohydroxy acid isomeroreductase, KIVD 2-ketoisovalerate decarboxylase, KPHMT ketopantoate hydroxymethyltransferase, PS pantothenate synthetase
Radmacher et al. [20] prepared C. glutamicum ΔilvA ΔpanBC whose biosynthesis of l-valine was strengthened by introduction of the plasmid containing ilvBNCD or ilvBNCE. The best strain C. glutamicum ΔilvA ΔpanBC (pJC1ilvBNCD) accumulated 92 mM l-valine along with a small amount of l-alanine (1.3 mM) in 48 h. Though AHAS was not desensitized from the feedback inhibition by l-valine in this strain, it was able to produce l-valine efficiently. This is because the feedback inhibition of AHAS of C. glutamicum by l-valine is not very strong, and the maximum inhibition of the enzymatic activity does not exceed 50 % [12]. Nonetheless, desensitization of AHAS from the feedback inhibition was investigated [28]. The native gene of the regulatory subunit of AHAS (ilvN) was replaced with the feedback-resistant mutant ilvNM13, which was designed based on the precedent structural information on the feedback-resistant homologs of E. coli and Streptomyces cinnamonensis. This plasmid-free strain C. glutamicum ΔilvA ΔpanB ilvNM13 produced 90 mM l-valine in the 48-h cultivation, while less than 40 mM l-valine formed in the culture of the parent strain C. glutamicum ΔilvA ΔpanB. l-Valine production by these strains was strengthened by a plasmid-carrying ilvBNC, and the resultant strain C. glutamicum ΔilvA ΔpanB ilvNM13 (pECKAilvBNC) produced 130 mM l-valine. It should be mentioned that the effect of the mutation on ilvN was less significant in the strains with the plasmid-borne ilvBNC, and the parent C. glutamicum ΔilvA ΔpanB (pECKAilvBNC) was able to produce up to 120 mM l-valine.
Formation of l-alanine was a prevailing issue in the l-valine-producing strains [59]. Pyruvate is a common precursor for the biosynthesis of l-valine and l-alanine. Therefore, suppression of l-alanine formation is beneficial for l-valine production. Two genes, alaT or avtA, are responsible for the conversion of pyruvate to l-alanine, which encode aminotransferases using l-glutamate or l-valine as an amine source, respectively [59]. Deletion of either gene was performed on C. glutamicum ΔilvA ΔpanBC (pJC1ilvBNCD), and there was no detrimental effect on l-valine production in both cases. While the avtA-deficient mutant showed only slight decrease of l-alanine accumulation, the alaT-deficient mutant improved greatly, and l-alanine formation decreased to 0.16 mM from 1.2 mM in the parent strain. It should be mentioned that the effect of deletion of these genes may be dependent on strains, and in other cases, effectiveness of avtA deletion has been demonstrated as described below.
In place of the complete deletion of the side-path genes or strong overexpression of the rate-limiting enzymes for production of target compounds, their expression can be modified by tuning promoter activity. Holátko et al. [29] prepared C. glutamicum ilvNM13 ΔpanB P-ilvAM1CG P-ilvDM7 P-ilvEM6, in which expression of ilvA was downregulated by the mutated promoter, while that of ilvD and ilvE was upregulated. This strain produced 136 mM l-valine in 48 h of flask fermentation. Modulation of promoter activities allows for overexpression of genes without using plasmids, which ensures genetic stability as well as no need for antibiotic markers. Additionally, the bradytrophic property is advantageous in that the strains do not require nutrition supplementation.
Recently, a strain with mixed strategies was reported [30], where expression of ilvA is partially limited by a mutated weak promoter, and production of l-alanine was suppressed by deletion of avtA. The resultant strain overexpressing the feedback-free l-valine biosynthetic genes, C. glutamicum MPilvA ΔavtA (pDXW-8-ilvEBN r C), produced 266 mM l-valine in 27 % molar yield from glucose. It is noteworthy that the side products, such as l-lactate and l-glutamate, were controlled by maintaining the dissolved oxygen (DO) at 15 % saturation. The same strategy was applied to B. flavum JV16, which is an α-aminobutyric acid-resistant and Leu–Ile–Met-auxotrophic strain generated by random mutagenesis from B. flavum DSM20411. The resultant strain B. flavum JV16 avtA::Cm (pDXW-8-ilvEBN r C) produced 331 mM l-valine in 39 % molar yield from glucose [30]. It should be mentioned that there is another example of an l-valine-producing strain using this subspecies of C. glutamicum [38]. B. flavum ATCC14067, which was transformed with the plasmid pDXW-8-ilvEBN r C, was able to produce 325 mM l-valine in 37 % molar yield from glucose at elevated temperatures as high as 37°C after the 48-h fed-batch fermentation.
An alternative strategy for producing l-valine to deleting ilvA and panB was reported by Blombach et al. [32]. Deletion of aceE, which encodes the E1p subunit of PDHC, resulted in inability to grow solely on glucose, while acetate supplementation compensated for it [60]. Blombach et al. [32] investigated accumulation of organic acids and amino acids in the culture medium of this strain and found that it started to produce pyruvate (30–35 mM), l-alanine (25–30 mM), and l-valine (30–35 mM) after depletion of acetate. Pyruvate accumulated as a direct consequence of inactivation of PDHC, while l-alanine and l-valine were the drain-off compounds of pyruvate. When ilvBNCE was overexpressed using the plasmid, the resultant strain C. glutamicum ΔaceE (pJC4ilvBNCE) produced 210 mM l-valine in the overall 50 % molar yield from glucose along with 5 mM pyruvate in the fed-batch process. This strain is advantageous over the aforementioned ΔilvA ΔpanB strains because it does not require supplementation of l-isoleucine or d-pantothenate.
Blombach et al. [34] performed further improvement of this aceE-deficient strain. They deleted the pqo gene encoding pyruvate:quinone oxidoreductase (PQO), which converts pyruvate to acetate and carbon dioxide. This resulted in an increase of 30 % molar yield. It is likely that PQO in the combination with acetate kinase and phosphotransacetylase bypasses the PDHC reaction to provide acetyl-CoA when the cell density is high. Therefore, inactivation of PQO led to the increase of l-valine production by cutting off the supply of the carbon resources for growth purposes in the late phase. They continued to observe pyruvate in the culture of C. glutamicum ΔaceE Δpqo (pJC4ilvBNCE), which indicated that l-valine production is limited by the downstream reactions from pyruvate to l-valine. They then tested to improve supply of NADPH. In the total reactions from glucose to l-valine, the downstream part (pyruvate to l-valine) requires two equivalents of NADPH at the AHAIR reaction as well as for regeneration of l-glutamate consumed by BCAT, whereas the glycolytic pathway provides only NADH, not NADPH. To compensate for the shortage of NADPH supply in l-valine production, they deleted pgi so that the carbon flux from glucose is directed to the pentose phosphate pathway, which produces 2 mol of NADPH from 1 mol of glucose. This strain, C. glutamicum ΔaceE Δpqo Δpgi (pJC4ilvBNCE), achieved more than 400 mM l-valine excretion in the 75 % molar yield from glucose, and pyruvate was not observed anymore. From the viewpoint of carbon usage, the deletion of pyc encoding pyruvate carboxylase was also beneficial, and C. glutamicum ΔaceE Δpqo Δpgi Δpyc (pJC4ilvBNCE) reached 86 % molar yield, although the final concentration of l-valine was about 240 mM.
Addition of acetate inhibits production of l-valine during growth of the ΔaceE strains because acetate represses the genes of the phosphoenolpyruvate:sugar phosphotransferase system (PTS), ptsG, ptsI, and ptsH via the DeoR-type regulator, SugR, and prevents uptake of glucose [61–63]. Therefore, Blombach et al. [33] deleted the sugR gene to remove repression of the PTS genes. Indeed, C. glutamicum ΔaceE Δpqo ΔsugR (pJC4ilvBNCE) consumed glucose five times faster than the parent strain and produced l-valine even in the growth phase in the presence of acetate, although the overall production was 40 % lower. Alternatively, they tested ethanol instead of acetate as the secondary carbon source to avoid the repression of the PTS genes. Under this condition, C. glutamicum ΔaceE Δpqo (pJC4ilvBNCE) was able to produce l-valine during its growth phase. These strains are effective producers of l-valine, but they accumulated 14–26 mM l-alanine and pyruvate. This accumulation of the by-products may be overcome by supply of NADPH in the strategy mentioned above.
Recently, Chen et al. [37] prepared a strain, which lacked both ilvA and aceE and alaT from C. glutamicum ATCC13869 to direct as much pyruvate as possible for production of l-valine. They also overexpressed the brnF and brnE genes encoding the BCAA exporter and the lrp gene encoding the global regulator Lrp that activates the expression of brnFE in addition to the l-valine biosynthetic genes (ilvBNC). The resultant strain C. glutamicum ATCC13869 ΔaceE ΔalaT ΔilvA (pJYW-4-ilvBNC 1 -lrp 1 -brnFE) produced 435 mM l-valine after a 96-h fermentation under the fed-batch condition.
Other strains of interest are the H+-ATPase-defective strains [31], which are known to increase the intracellular concentration of pyruvate [64, 65]. The native atpGDC genes were replaced with the inactivated atpG*DC genes containing a single point mutation, and the resultant strain was transformed with the plasmid pVK7ilvN53C, which allows for overexpression of feedback-resistant AHAS as well as AHAIR. C. glutamicum atpG*DC (pVK7ilvN53C) produced 96 mM l-valine in 72 h under the flask-shaking condition.
A completely different strategy is to exploit oxygen deprivation conditions, where most of the glucose is expected to be utilized for product formation and not for growth in C. glutamicum. Hasegawa et al. [35] used the strain without the ldhA gene, which encodes lactate dehydrogenase and is responsible for the production of lactate, i.e., the main fermentation product under oxygen deprivation conditions. Mere overexpression of the l-valine biosynthetic ilvBNCDE genes did not result in the efficient l-valine production because of poor glucose uptake caused by the redox imbalance. They solved this issue by converting the cofactor dependence from NADPH to NADH through mutagenesis of AHAIR (ilvC TM) and introduction of NADH-dependent exogenous leucine dehydrogenase in place of NADPH-dependent endogenous BCAT using the plasmid, pCRB-DLD. In addition, the feedback-resistant mutant of AHAS (ilvBN GE) was overexpressed. The resultant strain C. glutamicum R ΔldhA (pCRB-BNGECTM, pCRB-DLD) produced 1,470 mMFootnote 1 l-valine in 63 % molar yield from glucose in 24 h, and the concentration of l-valine reached 1,940 mM1 in 48 h. This strain produced succinate as a major by-product and left room for improvement [36]. To minimize the carbon flux to succinate, the phosphoenolpyruvate carboxylase gene ppc was deleted. While succinate production was suppressed, this resulted in the elevated NADH/NAD+ ratio. However, this redox imbalance was overcome by deletion of three genes, ctfA, pta, and ackA, associated with acetate synthesis, which produces excess NADH. Additionally, five glycolytic genes, gapA, pyk, pfkA, pgi, and tpi, were overexpressed. Moreover, l-alanine production was suppressed by deleting avtA. The resultant strain C. glutamicum R ilvN GE C TM, gapA, pyk, pfkA, pgi, and tpi ΔldhA Δppc Δpta ΔackA ΔctfA ΔavtA (pCRB-BNGECTM, pCRB-DLD) produced 1,280 mM1 l-valine in 88 % molar yield from glucose in the 24-h fed-batch fermentation.
Compared with C. glutamicum, development of the l-valine-producing strains of E. coli lags behind, which is probably because of the more complicated regulatory mechanisms for l-valine biosynthesis. However, using E. coli as a base strain is advantageous because of its rapid growth rate and rich genetic information. Early examples include the strains that acquire resistance against the feedback inhibition of AHAS III [66] or overexpress ygaZH encoding the l-valine exporter [67]. The strain with the mutation in isoleucine-tRNA synthetase [68] and the lipoic acid auxotroph with the inactivated H+-ATPase [69] were also reported to accumulate l-valine. However, these strains were prepared by classical random mutagenesis and their genotypes cannot be defined. It was only recently that the rationally designed strain of E. coli was reported to produce l-valine [39]. In this example, Park et al. first removed the product regulation by disarming the feedback inhibition of AHAS III and removing the transcriptional attenuation of the ilvBN and ilvGMEDA operons by replacing their attenuator leader regions with tac promoters. Then the ilvA, panB, and leuA genes were knocked out to converge the carbon resources to l-valine production. Next, AHAS I, which has higher affinity to pyruvate than other two isozymes, was overexpressed as well as ilvCED by the plasmid pKBRilvBNCED. The strain was further improved by the plasmid pTrc184ygaZHlrp harboring lrp and ygaZH that encodes the positive regulator for the ilvIH operon and the l-valine exporter, respectively. In silico gene knockout simulation was then performed, and they identified three candidates to be deleted: aceF, pfkA, and mdh. When these genes were deleted, the resultant strain E. coli ilvH (G41A, C50T), Ptac-ilvBN, Ptac-ilvGMED, and ΔilvA ΔpanB ΔleuA ΔaceF ΔpfkA Δmdh (pKBRilvBNCED, pTrc184ygaZHlrp) improved the yield and produced 64 mM l-valine in 58 % molar yield from glucose.
3.2 l-Isoleucine
An l-isoleucine producer was initially reported by Hayashibe and Uemura [70], which is an α-aminobutyric acid-resistant B. subtilis No. 14 that was isolated during the investigation of threonine metabolism and produced 4.3 g/L l-isoleucine. Screening of the α-aminobutyric acid-resistant microorganisms led to other l-isoleucine-producing strains, such as A. aerogenes IAM 1019 (2.4 g/L), Pseudomonas aureofaciens IAM 1001 (3.0 g/L), S. marcescens (2.7 g/L), and Erwinia carotovora E30 (1.3 g/L) [70]. Alternatively, d-threonine was used as a natural amino acid analog for the screening of the bacteria belonging to genera Serratia and Pseudomonas [71]. The most efficient strain among them, S. marcescens No. 1, produced more than 8 g/L l-isoleucine in a 40-h incubation. More microorganisms for l-isoleucine production were isolated using other BCAA analogs [72], including thiaisoleucine for E. coli [73, 74], Salmonella typhimurium [72, 75], and Saccharomyces cerevisiae [76], cyclopentaneglycine for S. typhimurium [75], glycyl isoleucine for E. coli [74], isoleucine hydroxamate for S. marcescens [77], and ketomycin for Bacillus subtilis [78]. As is the case for l-valine production, the “living cell reaction” process was effective for l-isoleucine production [58, 79, 80], which was performed using an α-aminobutyric acid-resistant B. flavum MJ-233 [57] to produce 200 mM l-isoleucine per day.
For rational metabolic engineering to enhance l-isoleucine production, it is important to address the fact that the biosynthetic pathway of l-isoleucine shares the genes and enzymes with those of l-valine and l-leucine (Fig. 1) and to converge the common intermediates toward the l-isoleucine synthesis. The specific issue for l-isoleucine production is the supply of l-threonine, which is one of the precursors for l-isoleucine biosynthesis. Additionally, removal of the tight feedback regulation for TDH (Fig. 1) is key to the efficient production of l-isoleucine.
As mentioned above, l-isoleucine production requires a supply of l-threonine. Therefore, the biosynthetic pathway of l-threonine and its regulation will be outlined briefly (Fig. 1). The biosynthesis of l-threonine starts from l-aspartate, and the pathway consists of five enzymatic reactions [81], which are catalyzed by aspartate kinase, aspartate semialdehyde dehydrogenase, homoserine dehydrogenase, homoserine kinase, and threonine synthase. E. coli has three aspartate kinase isozymes, I, II, and III, encoded by thrA, metL, and lysC, respectively, while C. glutamicum has only one aspartate kinase encoded by lysC. Aspartate kinases I and III of E. coli are under control of the feedback inhibition by l-threonine and l-lysine, respectively, whereas aspartate kinase II is not affected by feedback inhibition [82]. Instead, aspartate kinase II of E. coli is regulated by l-methionine through repression of metBL operon [83]. Aspartate kinase of C. glutamicum is subjected to the feedback inhibition by both l-lysine and l-threonine [13].
Homoserine dehydrogenase and homoserine kinase are encoded in the hom and thrB genes, respectively, in C. glutamicum. They form an operon and its expression is repressed by l-methionine [84]. Additionally, both enzymes are feedback inhibited by l-threonine [13]. In E. coli, thrA encodes a bifunctional enzyme that works as homoserine dehydrogenase in addition to aspartate kinase [85]. This gene forms an operon with thrB and thrC, the latter of which encodes threonine synthase. Expression of the thrABC operon is controlled by transcriptional attenuation by l-threonine as well as l-isoleucine [82]. Moreover, the activity of homoserine dehydrogenase and homoserine kinase is feedback inhibited [82].
For l-isoleucine production, it is advantageous to use l-lysine-producing strains to supply l-threonine because the biosynthetic pathway of l-threonine shares a significant part with that of l-lysine. Colón et al. [40] reported accumulation of 114 mM l-isoleucine using an l-lysine-producing strain C. glutamicum ATCC 21799 (termed as C. lactofermentum in the original paper) by overexpressing the wild-type ilvA. Morbach et al. [13, 86] used another l-lysine-producing strain of C. glutamicum generated by random mutagenesis. They introduced multiple copies of the feedback-resistant hom and thrB in the chromosome or replaced the chromosomal native lysC and hom genes with the feedback-resistant ones. Along with overexpression of ilvA, the former strain produced 96 mM l-isoleucine in the batch culture and the latter 138 mM in the fed-batch culture. It should be mentioned that overexpression of hom was possible only after overexpression of ilvA because otherwise accumulation of l-threonine and l-homoserine caused instability of the strains [42]. Yin et al. [45] identified the feedback-resistant mutants of TDH and AHAS from the l-isoleucine-producing strain C. glutamicum JHI3-156. When the both feedback-resistant enzymes were overexpressed in the same strain, this resultant strain produced 234 mM l-isoleucine in the fed-batch condition. A comparative proteomic study on this strain further identified up- and downregulated proteins, which were related to cell growth, l-isoleucine biosynthesis, and stress response [87].
Introduction of exogenous genes is also a useful strategy. For example, an E. coli-derived ilvA was introduced to an l-threonine-producing B. flavum strain, a relative bacterium to C. glutamicum, to yield 153 mM l-isoleucine [88]. Wang et al. [43] introduced the E. coli K-12-originated thrABC genes to C. glutamicum. After deletion of alaT, the resultant strain produced 100 mM l-isoleucine along with low concentrations of l-lysine, l-alanine, and l-valine. Guillouet et al. [89] reported the advantage of the tdcB gene encoding the catabolic TDH of E. coli over the feedback-resistant ilvA. The strain with the tdcB gene accumulated four times more l-isoleucine than the one with the ilvA gene and yielded 30 mM l-isoleucine in the batch culture [41].
As is the case for l-valine, export of the accumulated product is important for efficient production of l-isoleucine. Therefore, overexpression of the global regulator Lrp and the BCAA exporter BrnFE was performed in C. glutamicum JHI3-156 to produce 205 mM l-isoleucine [46]. Recently, Zhao et al. [47] reported increased l-isoleucine production using C. glutamicum IWJ001, which was identified as an l-isoleucine-producing strain by random mutagenesis. They found that the biosynthetic enzymes for l-isoleucine were significantly upregulated when the fusA and frr genes, which encode ribosome elongation factor G and ribosome recycling factor, respectively, were overexpressed. Together with overexpression of ilvA, ilvB, ilvN, and ppnk (a polyphosphate/ATP-dependent NAD kinase), the resultant strain produced 217 mM l-isoleucine in 72 h of fed-batch fermentation.
Optimization of the fermentation conditions is also an issue to be addressed. Peng et al. [44] optimized DO and pH of the fermentation conditions using C. glutamicum JHI3-156 (termed as B. lactofermentum in the original paper) and finally achieved 203 mM l-isoleucine in the batch culture.
E. coli is also the target of manipulation for l-isoleucine bio-production. Hashiguchi et al. [90] strengthened the downstream reactions by introducing the plasmid with the ilvA, ilvGM, ilvD, and ilvE genes to an l-threonine-producing E. coli K-12 mutant, and the resultant strain produced 78 mM l-isoleucine. This strain coproduced l-valine, but they solved this problem by introducing the gene of the feedback-resistant aspartate kinase III, which successfully reduced l-valine production and increased the final concentration of l-isoleucine to 94 mM [48].
3.3 l-Leucine
In the classical examples, accumulation of l-leucine was observed in the revertants of S. marcescens from the l-isoleucine auxotrophic mutant, which had been generated by resistance to α-aminobutyric acid [91]. Mechanistic investigations [91, 92] revealed that the resistance to α-aminobutyric acid was acquired by derepression of both l-isoleucine/l-valine and l-leucine biosynthetic enzymes. Interestingly, the reversion from the l-isoleucine auxotrophy was not due to desensitization of the feedback inhibition of TDH, but due to that of IPMS, which allowed for overproduction of l-leucine as well as supply of 2-ketobutyrate, i.e., a precursor for biosynthesis of l-isoleucine in place of TDH.
Another l-leucine-producing mutant was obtained from the glutamic acid-producing bacterium, B. lactofermentum [93]. This strain was screened from an l-methionine/l-isoleucine double auxotroph of B. lactofermentum 2256 using 2-thiazolealanine as an amino acid metabolism competitor. The optimization of the culture conditions allowed production of 30 g/L l-leucine [94, 95]. In this strain, IPMS is both desensitized and derepressed while AHAS remained intact [56]. Therefore, another screening was performed using β-hydroxyleucine, which obtained AHAS mutants desensitized from all of the BCAAs [96]. These strains showed improved productivity of 34 g/L l-leucine. A further campaign was conducted using high concentrations of d-α-aminobutyric acid, and a mutant with higher activities of AHAS and IPMS was isolated [97]. This mutant produced more l-leucine and showed a better l-leucine/l-valine ratio than the parent strain.
Mutants of E. coli have been developed for l-leucine production more recently. Some 4-azaleucine-resistant strains derived from E. coli K-12 were desensitized to the feedback inhibition of IPMS, and the most efficient strain produced 5.2 g/L l-leucine [25]. Alternatively, the l-isoleucine/l-valine double auxotroph with mutation in ilvE was isolated [98]. This strain was then supplemented with the plasmid to overexpress tyrB, which works only on the l-leucine biosynthesis among the BCAAs. The resultant strain produced 2.7 g/L l-leucine with no detectable l-valine or l-isoleucine. Lowering the promoter activity of the sucAB gene encoding α-ketoglutarate dehydrogenase is likely to reduce the carbon flux into TCA cycle and decrease consumption of acetyl-CoA [99]. When this was combined with the feedback-free IPMS and the inactivated BCAT, the resultant strain produced 11.4 g/L l-leucine.
l-Leucine-producing strains were also found from amino acid auxotrophs of C. glutamicum. One of the best producers was the l-phenylalanine/l-histidine double auxotroph, which accumulated 16.0 g/L l-leucine. Another example is the S-(2-aminoethyl)-l-cysteine-resistant mutant of C. glutamicum [100], though this strain was found to be unstable and generated several types of revertants during the fermentation.
Recently, a genetically defined strain which produces l-leucine was reported [49]. This strain contains three copies of the P tuf -leuA_B018 module in the chromosome. This gene, leuA_B018, encodes IPMS, which is disarmed from feedback inhibition by l-leucine. Moreover, in this module, the native promoter was replaced by the strong tuf promoter, which is free from transcriptional attenuation. Acetyl-CoA, i.e., another substrate for the IPMS reaction, was increased by replacing the native promoter of gltA (encoding citrate synthase) to P dapA-L1 . The transcriptional repressor-encoding gene ltbR was deleted to enhance expression of the downstream genes leuBCD for l-leucine production. Furthermore, mutations were introduced to the regulatory site of AHAS (encoded by ilvN) to be desensitized from feedback inhibition and increase the carbon flux toward l-leucine production. IolT1, which is regulated by IolR, catalyzes glucose uptake in a PTS-independent manner. Therefore, the iolR gene was also deleted to enhance glucose uptake. This strain, C. glutamicum P tuf -leuA_B018 P dap-L1 -gltA ilvN_fbr ΔltbR::P tuf -leuA_B018 ΔleuA::P tuf -leuA_B018 ΔiolR, produced up to 181 mM l-leucine in the culture solution along with precipitate of l-leucine after 72 h of the fed-batch fermentation.
4 Production of Chemicals Using the BCAA Biosynthetic Pathways
Recently, the instability of oil prices and environmental concerns has been driving the microbial production of biofuels. Such research includes ethanol fermentation as well as production of higher alcohols, which are expected to be next-generation biofuels due to their high energy density [101, 102]. Isobutanol is one representative example of such alcohols.
Isobutanol can be synthesized using the l-valine biosynthetic pathway (Fig. 2). 2-Ketoisovalerate, i.e., an intermediate of the l-valine biosynthesis, is a starting material for isobutanol production. It is first converted to isobutyraldehyde by 2-ketoisovalerate decarboxylase (KIVD), and then isobutanol is formed from the aldehyde by reduction using alcohol dehydrogenase (ADH). Like l-valine production, overexpression of the genes encoding AHAS, AHAIR, and DHAD is effective to enhance availability of 2-ketoisovalarate for isobutanol production [103]. The bacterial strains developed for isobutanol production are reported for E. coli [104, 105], B. subtilis [106], and C. glutamicum [107], and the yields range between 74 and 297 mM. Although less efficient, S. cerevisiae is also used as a host for isobutanol production. The engineered strain overexpressing the l-valine biosynthetic genes accumulated 2.4 mM isobutanol [108], and the one whose l-isoleucine biosynthesis was eliminated produced 3.0 mM [109]. One of the difficulties in microbial production of isobutanol is that this alcohol is very toxic for microbes. To address this issue, Yamamoto et al. [110] performed isobutanol production in a growth-uncoupled manner under oxygen-deprived conditions. Moreover, they continuously extracted isobutanol from the aqueous reaction phase by layering oleyl alcohol and achieved as high as 981 mM of volumetric productivity.
In a similar manner to isobutanol, 3-methyl-1-butanol and 2-methyl-1-butanol can be produced from 2-keto-4-methylvalerate and 2-keto-3-methylvalerate, i.e., intermediates for the l-leucine and l-isoleucine biosynthesis, respectively, by the catalysis of KIVD and ADH (Fig. 2) [104, 111]. Such examples include the E. coli strain reported by Connor and Liao [112], where accumulation of 17 mM of 3-methyl-1-butanol was achieved when expression of the l-valine and l-leucine biosynthetic genes was enhanced to increase availability of 2-keto-4-methylvalerate for 3-methyl-1-butanol production. They also performed random mutagenesis and obtained an efficient strain that was able to produce 128 mM of 3-methyl-1-butanol in the biphasic fermentation using oleyl alcohol [113]. Cann and Liao [114] engineered E. coli to produce 2-methyl-1-butanol. In addition to strengthening the l-isoleucine biosynthetic pathway, the flux to l-threonine was optimized, and the resultant strain produced 14 mM of 2-methyl-1-butanol. Recently, production of isobutanol (17 mM), 3-methyl-1-butanol (8.5 mM), and 2-methyl-1-butanol (12 mM) was performed by the strains engineered for each alcohol with relevant genes from S. cerevisiae using C. crenatum as a parent strain [115].
Another important chemical for which the BCAA fermentative pathway can be applied is d-pantothenate. In C. glutamicum, it is produced from 2-ketoisovalerate by three enzymatic reactions catalyzed by ketopantoate hydroxymethyltransferase (KPHMT), AHAIR, and pantothenate synthetase (PS) [116] (Fig. 2). Hüser et al. [117] reported a d-pantothenate-producing strain. In this strain, ilvA was deleted, and the transcription of ilvE was attenuated to decrease the carbon flux to the competing BCAA biosynthesis. Additionally, overexpression of ilvBNCD was performed to increase availability of 2-ketoisovalerate as well as overexpression of panBC to direct more 2-ketoisovalerate to the d-pantothenate biosynthetic pathway. This strain produced 8 mM of d-pantothenate.
5 Conclusion
In this chapter, the biosynthetic pathway of BCAAs and its regulatory system were described, and the microbial strains engineered for BCAA production were summarized. The classical mutagenic strains provided a heritage of information about the complicated feedback inhibition and transcriptional attenuation in the BCAA biosynthesis. They now work as a guide for the rational design of BCAA-producing strains using advanced molecular biology techniques.
Generally, genetic modifications for producing target BCAAs are divided into four stages [3]: (1) to disarm feedback inhibition by the target BCAA, (2) to remove transcriptional attenuation by the target BCAA, (3) to minimize carbon flux to the competing pathways to reduce formation of by-products, and (4) to enhance expression of genes encoding biosynthetic enzymes of a BCAA to converge carbon resources as much as possible to production of it. Additionally, engineering of the export and import systems as well as modification of the transcriptional factors or the global regulators may be beneficial to improve production of the target amino acid further.
Disarming feedback inhibition is the most important step for efficient production of BCAAs because it has a very strong inhibitory effect on the microorganism growth and therefore the fermentative production of BCAAs. For example, the minimum inhibitory concentration of l-valine to the growth of E. coli K-12 is reported to be as low as 2 mg/L [17]. The complicated feedback inhibition system of the BCAA biosynthetic pathways was clarified by the early-stage studies on the auxotrophic strains and the amino acid analog-resistant strains. The key players of feedback inhibition are AHAS for all BCAAs, TDH for l-isoleucine, and IPMS for l-leucine, which can be now desensitized by rational mutation.
Production of BCAAs is also affected by transcriptional attenuation of the relevant genes. Derepression of the genes coding for AHAS is a common tactic because they are related to biosynthesis of all BCAAs. Removal of transcriptional attenuation of leuA is also important for l-leucine production. For l-isoleucine production, increasing supply of l-threonine by relieving transcriptional attenuation of the relevant genes may be beneficial. These can be performed by replacing the native promoters with others, e.g., tac promoter.
Minimization of carbon flux to competing pathways is required for efficient BCAA production. This can be achieved by deleting the corresponding genes. Pyruvate, which is the common intermediate for BCAA production, is also used for production of other competing by-products such as lactate, acetate, and succinate. Therefore, the genes responsible for production of these by-products are the target of deletion to increase availability of pyruvate for the BCAA biosynthesis. Additionally, for production of one of the BCAAs, the genes specifically relevant to the other two BCAAs are to be deleted, or their expression should be suppressed. For example, the ilvA gene, which is involved only in the l-isoleucine biosynthesis, is the target of deletion or attenuation for l-valine production.
Finally, overexpression of the genes responsible for biosynthesis of the target amino acid enhances its productivity. In particular, overexpression of AHAS, AHAIR, and DHAD, which catalyze the common reactions for all BCAAs, is effective for their production. It is beneficial to overexpress the ilvA and leuA genes for production of l-isoleucine and l-leucine, respectively, because they code for the enzymes responsible for their specific biosynthetic pathways. In addition, enhanced expression of the exporters of BCAAs as well as their positive regulators is also beneficial.
Currently, in silico studies of carbon flux simulation are emerging as a powerful tool to design strategies to maximize efficiency of biosynthetic pathways for production of the target compound [118, 119]. Application of these techniques will become an essential part of creating more sophisticated microbial strains, which can realize our goals.
Notes
- 1.
The concentration values are corrected by the dilution factors caused by addition of ammonia solution to maintain the pH of the reaction solutions.
References
Wu G (2009) Amino acids: metabolism, functions, and nutrition. Amino Acids 37(1):1–17
Monirujjaman M, Ferdouse A (2014) Metabolic and physiological roles of branched-chain amino acids. Adv Mol Biol 2014:364976
Park JH, Lee SY (2010) Fermentative production of branched chain amino acids: a focus on metabolic engineering. Appl Microbiol Biotechnol 85(3):491–506
Wendisch VF, Bott M, Eikmanns BJ (2006) Metabolic engineering of Escherichia coli and Corynebacterium glutamicum for biotechnological production of organic acids and amino acids. Curr Opin Microbiol 9(3):268–274
Zahoor A, Lindner SN, Wendisch VF (2012) Metabolic engineering of Corynebacterium glutamicum aimed at alternative carbon sources and new products. Comput Struct Biotechnol J 3, e201210004
Eggeling L, Pfefferle W, Sahm H (2001) Amino acids. In: Ratledge C, Kristiansen B (eds) Basic biotechnology. Cambridge University Press, Cambridge, pp 281–303
Ikeda M (2003) Amino acid production processes. Adv Biochem Eng Biotechnol 79:1–35
Oldiges M, Eikmanns BJ, Blombach B (2014) Application of metabolic engineering for the biotechnological production of L-valine. Appl Microbiol Biotechnol 98(13):5859–5870
Park JH, Lee SY (2008) Towards systems metabolic engineering of microorganisms for amino acid production. Curr Opin Biotechnol 19(5):454–460
Vertes AA, Inui M, Yukawa H (2012) Postgenomic approaches to using corynebacteria as biocatalysts. Annu Rev Microbiol 66:521–550
Shen XH, Zhou NY, Liu SJ (2012) Degradation and assimilation of aromatic compounds by Corynebacterium glutamicum: another potential for applications for this bacterium? Appl Microbiol Biotechnol 95(1):77–89
Eggeling I, Cordes C, Eggeling L, Sahm H (1987) Regulation of acetohydroxy acid synthase in Corynebacterium glutamicum during fermentation of α-ketobutyrate to L-isoleucine. Appl Microbiol Biotechnol 25(4):346–351
Morbach S, Sahm H, Eggeling L (1995) Use of feedback-resistant threonine dehydratases of Corynebacterium glutamicum to increase carbon flux towards L-isoleucine. Appl Environ Microbiol 61(12):4315–4320
Changeux JP (1963) Allosteric interactions on biosynthetic L-threonine deaminase from Escherichia coli K-12. Cold Spring Harb Symp Quant Biol 28:497–504
Umbarger HE (1996) Biosynthesis of the branched-chain amino acids. In: Neidhardt FC, Curtiss RI, Ingraham JL, Lin EC, Low KB Jr, Magasanik B et al (eds) Escherichia coli and Salmonella: cellular and molecular biology, 2nd edn. ASM Press, Washington, DC, pp 442–457
De Felice M, Squires CH, Levinthal M (1978) A comparative study of the acetohydroxy acid synthase isoenzymes of Escherichia coli K-12. Biochim Biophys Acta 541:9–17
Kutukova EA, Livshits VA, Altman IP, Ptitsyn LR, Zyiatdinov MH, Tokmakova IL et al (2005) The yeaS (leuE) gene of Escherichia coli encodes an exporter of leucine, and the Lrp protein regulates its expression. FEBS Lett 579(21):4629–4634
Keilhauer C, Eggeling L, Sahm H (1993) Isoleucine synthesis in Corynebacterium glutamicum: molecular analysis of the ilvB-ilvN-ilvC operon. J Bacteriol 175(17):5595–5603
Morbach S, Junger C, Sahm H, Eggeling L (2000) Attenuation control of ilvBNC in Corynebacterium glutamicum: evidence of leader peptide formation without the presence of a ribosome binding site. J Biosci Bioeng 90(5):501–507
Radmacher E, Vaitsikova A, Burger U, Krumbach K, Sahm H, Eggeling L (2002) Linking central metabolism with increased pathway flux: L-valine accumulation by Corynebacterium glutamicum. Appl Environ Microbiol 68(5):2246–2250
Leyval D, Uy D, Delaunay S, Goergen JL, Engasser JM (2003) Characterisation of the enzyme activities involved in the valine biosynthetic pathway in a valine-producing strain of Corynebacterium glutamicum. J Biotechnol 104(1–3):241–252
Inoue K, Kuramitsu S, Aki K, Watanabe Y, Takagi T, Nishigai M et al (1988) Branched-chain amino acid aminotransferase of Escherichia coli: overproduction and properties. J Biochem 104(5):777–784
Pátek M, Krumbach K, Eggeling L, Sahm H (1994) Leucine synthesis in Corynebacterium glutamicum: enzyme activities, structure of leuA, and effect of leuA inactivation on lysine synthesis. Appl Environ Microbiol 60(1):133–140
Pátek M, Hochmannova J, Jelinkova M, Nesvera J, Eggeling L (1998) Analysis of the leuB gene from Corynebacterium glutamicum. Appl Microbiol Biotechnol 50(1):42–47
Gusyatiner MM, Lunts MG, Kozlov YI, Ivanovskaya LV, Voroshilova EB (2002) DNA coding for mutant isopropylmalate synthase L-leucine-producing microorganism and method for producing L-leucine. US Patent 6403342 B1
Groeger U, Sahm H (1987) Microbial production of L-leucine from α-ketoisocaproate by Corynebacterium glutamicum. Appl Microbiol Biotechnol 25(4):352–356
Berg CM, Liu L, Vartak NB, Whalen WA, Wang BM (1990) The branched chain amino acid transaminase genes and their production in Escherichia coli. In: Chipman D, Barak Z, Schloss JV (eds) Biosynthesis of branched chain amino acids. VCH Verlagsgesellschaft, Weinheim, pp 131–162
Elišáková V, Pátek M, Holátko J, Nešvera J, Leyval D, Goergen JL et al (2005) Feedback-resistant acetohydroxy acid synthase increases valine production in Corynebacterium glutamicum. Appl Environ Microbiol 71(1):207–213
Holátko J, Elišáková V, Prouza M, Sobotka M, Nesvera J, Patek M (2009) Metabolic engineering of the L-valine biosynthesis pathway in Corynebacterium glutamicum using promoter activity modulation. J Biotechnol 139(3):203–210
Hou X, Chen X, Zhang Y, Qian H, Zhang W (2012) L-Valine production with minimization of by-products’ synthesis in Corynebacterium glutamicum and Brevibacterium flavum. Amino Acids 43(6):2301–2311
Wada M, Hijikata N, Aoki R, Takesue N, Yokota A (2008) Enhanced valine production in Corynebacterium glutamicum with defective H+-ATPase and C-terminal truncated acetohydroxyacid synthase. Biosci Biotechnol Biochem 72(11):2959–2965
Blombach B, Schreiner ME, Holatko J, Bartek T, Oldiges M, Eikmanns BJ (2007) L-Valine production with pyruvate dehydrogenase complex-deficient Corynebacterium glutamicum. Appl Environ Microbiol 73(7):2079–2084
Blombach B, Arndt A, Auchter M, Eikmanns BJ (2009) L-Valine production during growth of pyruvate dehydrogenase complex-deficient Corynebacterium glutamicum in the presence of ethanol or by inactivation of the transcriptional regulator SugR. Appl Environ Microbiol 75(4):1197–1200
Blombach B, Schreiner ME, Bartek T, Oldiges M, Eikmanns BJ (2008) Corynebacterium glutamicum tailored for high-yield L-valine production. Appl Microbiol Biotechnol 79(3):471–479
Hasegawa S, Uematsu K, Natsuma Y, Suda M, Hiraga K, Jojima T et al (2012) Improvement of the redox balance increases L-valine production by Corynebacterium glutamicum under oxygen deprivation conditions. Appl Environ Microbiol 78(3):865–875
Hasegawa S, Suda M, Uematsu K, Natsuma Y, Hiraga K, Jojima T et al (2013) Engineering of Corynebacterium glutamicum for high-yield L-valine production under oxygen deprivation conditions. Appl Environ Microbiol 79(4):1250–1257
Chen C, Li Y, Hu J, Dong X, Wang X (2015) Metabolic engineering of Corynebacterium glutamicum ATCC13869 for L-valine production. Metab Eng 29:66–75
Hou X, Ge X, Wu D, Qian H, Zhang W (2012) Improvement of L-valine production at high temperature in Brevibacterium flavum by overexpressing ilvEBN r C genes. J Ind Microbiol Biotechnol 39(1):63–72
Park JH, Lee KH, Kim TY, Lee SY (2007) Metabolic engineering of Escherichia coli for the production of L-valine based on transcriptome analysis and in silico gene knockout simulation. Proc Natl Acad Sci U S A 104(19):7797–7802
Colón GE, Nguyen TT, Jetten MS, Sinskey AJ, Stephanopoulos G (1995) Production of isoleucine by overexpression of ilvA in a Corynebacterium lactofermentum threonine producer. Appl Microbiol Biotechnol 43(3):482–488
Guillouet S, Rodal AA, An G-H, Gorret N, Lessard PA, Sinskey AJ (2001) Metabolic redirection of carbon flow toward isoleucine by expressing a catabolic threonine dehydratase in a threonine-overproducing Corynebacterium glutamicum. Appl Microbiol Biotechnol 57(5–6):667–673
Morbach S, Sahm H, Eggeling L (1996) L-Isoleucine production with Corynebacterium glutamicum: further flux increase and limitation of export. Appl Environ Microbiol 62(12):4345–4351
Wang J, Wen B, Wang J, Xu Q, Zhang C, Chen N et al (2013) Enhancing L-isoleucine production by thrABC overexpression combined with alaT deletion in Corynebacterium glutamicum. Appl Biochem Biotechnol 171(1):20–30
Peng Z, Fang J, Li J, Liu L, Du G, Chen J et al (2010) Combined dissolved oxygen and pH control strategy to improve the fermentative production of L-isoleucine by Brevibacterium lactofermentum. Bioprocess Biosyst Eng 33(3):339–345
Yin L, Hu X, Xu D, Ning J, Chen J, Wang X (2012) Co-expression of feedback-resistant threonine dehydratase and acetohydroxy acid synthase increase L-isoleucine production in Corynebacterium glutamicum. Metab Eng 14(5):542–550
Yin L, Shi F, Hu X, Chen C, Wang X (2013) Increasing L-isoleucine production in Corynebacterium glutamicum by overexpressing global regulator Lrp and two-component export system BrnFE. J Appl Microbiol 114(5):1369–1377
Zhao J, Hu X, Li Y, Wang X (2015) Overexpression of ribosome elongation factor G and recycling factor increases L-isoleucine production in Corynebacterium glutamicum. Appl Microbiol Biotechnol 99(11):4795–4805
Hashiguchi K, Matsui H, Kurahashi O (1999) Effects of a feedback-resistant aspartokinase III gene on L-isoleucine production in Escherichia coli K-12. Biosci Biotechnol Biochem 63(11):2023–2024
Vogt M, Haas S, Klaffl S, Polen T, Eggeling L, van Ooyen J et al (2014) Pushing product formation to its limit: metabolic engineering of Corynebacterium glutamicum for L-leucine overproduction. Metab Eng 22:40–52
Udaka S, Kinoshita S (1959) The fermentative production of L-valine by bacteria. J Gen Appl Microbiol 5(4):159–174
Sugisaki Z (1959) Studies on L-valine fermentation. Part I. Production of L-valine by Aerobacter bacteria. J Gen Appl Microbiol 5:138–149
Nakayama K, Kitada S, Kinoshita S (1961) L-Valine production using microbial auxotroph. J Gen Appl Microbiol 7:52–69
Karlström O (1965) Methods for the production of mutants suitable as amino acid fermentation organisms. Biotechnol Bioeng 7:245–268
Kisumi M, Komatsubara S, Chibata I (1971) Valine accumulation by α-aminobutyric acid-resistant mutants of Serratia marcescens. J Bacteriol 106(2):493–499
Tsuchida T, Yoshinaga F, Kubota K, Momose H (1975) Production of L-valine by 2-thiazolealanine resistant mutants derived from glutamic acid producing bacteria. Agric Biol Chem 39(6):1319–1322
Tsuchida T, Momose H (1975) Genetic changes of regulatory mechanisms occurred in leucine and valine producing mutants derived from Brevibacterium lactofermentum 2256. Agric Biol Chem 39(11):2193–2198
Yukawa H, Terasawa M (1986) L-Isoleucine production by ethanol utilizing microorganism. Process Biochem 21:196–199
Terasawa M, Inui M, Goto M, Shikata K, Imanari M, Yukawa H (1990) Living cell reaction process for L-isoleucine and L-valine production. J Ind Microbiol 5(5):289–293
Marienhagen J, Eggeling L (2008) Metabolic function of Corynebacterium glutamicum aminotransferases AlaT and AvtA and impact on L-valine production. Appl Environ Microbiol 74(24):7457–7462
Schreiner ME, Fiur D, Holatko J, Patek M, Eikmanns BJ (2005) E1 enzyme of the pyruvate dehydrogenase complex in Corynebacterium glutamicum: molecular analysis of the gene and phylogenetic aspects. J Bacteriol 187(17):6005–6018
Engels V, Wendisch VF (2007) The DeoR-type regulator SugR represses expression of ptsG in Corynebacterium glutamicum. J Bacteriol 189(8):2955–2966
Gaigalat L, Schluter JP, Hartmann M, Mormann S, Tauch A, Puhler A et al (2007) The DeoR-type transcriptional regulator SugR acts as a repressor for genes encoding the phosphoenolpyruvate:sugar phosphotransferase system (PTS) in Corynebacterium glutamicum. BMC Mol Biol 8:104
Tanaka Y, Teramoto H, Inui M, Yukawa H (2008) Regulation of expression of general components of the phosphoenolpyruvate: carbohydrate phosphotransferase system (PTS) by the global regulator SugR in Corynebacterium glutamicum. Appl Microbiol Biotechnol 78(2):309–318
Aoki R, Wada M, Takesue N, Tanaka K, Yokota A (2005) Enhanced glutamic acid production by a H+-ATPase-defective mutant of Corynebacterium glutamicum. Biosci Biotechnol Biochem 69(8):1466–1472
Sekine H, Shimada T, Hayashi C, Ishiguro A, Tomita F, Yokota A (2001) H+-ATPase defect in Corynebacterium glutamicum abolishes glutamic acid production with enhancement of glucose consumption rate. Appl Microbiol Biotechnol 57(4):534–540
Livshits VA, Doroshenko VG, Gorshkova NV, Belaryeva AV, Ivanovskaya LV, Khourges EM, Akhverdian VZ, Gusyatiner MM, Kozlov YI (2004) Mutant ilvH gene and method for producing L-valine. US Patent US6737255 B2
Tabolina EA, Rybak KV, Khourges EM, Voroshilova EB, Gusyatiner MM (2005) Method for producing L-amino acid using bacteria belonging to the genus Escherichia. US Patent 2005/0239175 A1
Livshits VA, Debabov VG, Fedorovva AO, Pavlovva ZN, Shakulov RS, Bachina TA, Khourges EM (1997) Strains of Escherichia coli which produce isoleucine or valine and a method for their production. US Patent 5658766
Tomita F, Yokota A, Hashiguchi K, Ishigooka M, Kurahashi O (2001) Methods for producing L-valine and L-leucine. US Patent 6214591 B1
Hayashibe M, Uemura T (1961) Release from the feedback inhibition controlling the biosynthesis of isoleucine. Nature 191:1417–1418
Kisumi M (1962) Studies on the isoleucine fermentation. I. Screening of organisms and investigation of cultural conditions. J Biochem 52:390–399
Umbarger HE (1971) Metabolite analogs as genetic and biochemical probes. Adv Genet 16:119–140
Szentirmai A, Szentirmai M, Umbarger HE (1968) Isoleucine and valine metabolism of Escherichia coli. XV. Biochemical properties of mutants resistant to thiaisoleucine. J Bacteriol 95(5):1672–1679
Vonder Haar RA, Umbarger HE (1972) Isoleucine and valine metabolism in Escherichia coli. XIX. Inhibition of isoleucine biosynthesis by glycyl-leucine. J Bacteriol 112(1):142–147
O’Neill JP, Freundlich M (1972) Effect of cyclopentaneglycine on metabolism in Salmonella typhimurium. J Bacteriol 111(2):510–515
Betz JL, Hereford LM, Magee PT (1971) Threonine deaminases from Saccharomyces cerevisiae mutationally altered in regulatory properties. Biochemistry 10(10):1818–1824
Kisumi M, Komatsubara S, Sugiura M, Chibata I (1971) Properties of isoleucine hydroxamate-resistant mutants of Serratia marcescens. J Gen Microbiol 69(3):291–297
Krupe H, Poralla K (1972) Properties of mutants of Bacillus subtilis which are resistant to the isoleucine antagonist ketomycin. Arch Mikrobiol 85(3):253–258
Terasawa M, Kakinuma N, Shikata K, Yukawa H (1989) New process for L-isoleucine production. Process Biochem 24:60–61
Terasawa M, Inui M, Goto M, Kurusu Y, Yukawa H (1991) Depression of by-product formation during L-isoleucine production by a living-cell reaction process. Appl Microbiol Biotechnol 35(3):348–351
Park JH, Lee SY (2010) Metabolic pathways and fermentative production of L-aspartate family amino acids. Biotechnol J 5(6):560–577
Patte JC (1996) Biosynthesis of threonine and lysine. In: Neidhardt FC, Curtiss RI, Ingraham JL, Lin EC, Low KB Jr, Magasanik B et al (eds) Escherichia coli and Salmonella: cellular and molecular biology, 2nd edn. ASM Press, Washington, DC, pp 528–541
Greene RC, Smith AA (1984) Insertion mutagenesis of the metJBLF gene cluster of Escherichia coli K-12: evidence for an metBL operon. J Bacteriol 159(2):767–769
Peoples OP, Liebl W, Bodis M, Maeng PJ, Follettie MT, Archer JA et al (1988) Nucleotide sequence and fine structural analysis of the Corynebacterium glutamicum hom-thrB operon. Mol Microbiol 2(1):63–72
Katinka M, Cossart P, Sibilli L, Saint-Girons I, Chalvignac MA, Le Bras G et al (1980) Nucleotide sequence of the thrA gene of Escherichia coli. Proc Natl Acad Sci U S A 77(10):5730–5733
Morbach S, Kelle R, Winkels S, Sahm H, Eggeling L (1996) Engineering the homoserine dehydrogenase and threonine dehydrogenase control points to analyse flux towards L-isoleucine in Corynebacterium glutamicum. Appl Microbiol Biotechnol 45(5):612–620
Yin L, Hu X, Wang X (2014) Proteomic analysis of L-isoleucine production by Corynebacterium glutamicum. J Pure Appl Microbiol 8(2):899–908
Hashiguchi K, Kojima H, Sato K, Sano K (1997) Effects of an Escherichia coli ilvA mutant gene encoding feedback-resistant threonine deaminase on L-isoleucine production by Brevibacterium flavum. Biosci Biotechnol Biochem 61(1):105–108
Guillouet S, Rodal AA, An G-H, Lessard PA, Sinskey AJ (1999) Expression of the Escherichia coli catabolic threonine dehydratase in Corynebacterium glutamicum and its effect on isoleucine production. Appl Environ Microbiol 65(7):3100–3107
Hashiguchi K, Takesada H, Suzuki E, Matsui H (1999) Construction of an L-isoleucine overproducing strain of Escherichia coli K-12. Biosci Biotechnol Biochem 63(4):672–679
Kisumi M, Komatsubara S, Chibata I (1973) Leucine accumulation by isoleucine revertants of Serratia marcescens resistant to α-aminobutyric acid lack of both feedback inhibition and repression. J Biochem 73(1):107–115
Kisumi M, Komatsubara S, Chibata I (1977) Pathway for isoleucine formation from pyruvate by leucine biosynthetic enzymes in leucine-accumulating isoleucine revertants of Serratia marcescens. J Biochem 82(1):95–103
Tsuchida T, Yoshinaga F, Kubota K, Momose H, Okumura S (1974) Production of L-leucine by a mutant of Brevibacterium lactofermentum 2256. Agric Biol Chem 38(10):1907–1911
Akashi K, Ikeda S, Shibai H, Kobayashi K, Hirose Y (1978) Determination of redox potential levels critical for cell respiration and suitable for L-leucine production. Biotechnol Bioeng 20(1):27–41
Tsuchida T, Yoshinaga F, Kubota K, Momose H, Okumura S (1975) Cultural conditions for L-leucine production by strain No. 218, a mutant of Brevibacterium lactofermentum 2256. Agric Biol Chem 39(5):1149–1153
Tsuchida T, Momose H (1986) Improvement of an L-leucine-producing mutant of Brevibacterium lactofermentum 2256 by genetically desensitizing it to α-acetohydroxy acid synthetase. Appl Environ Microbiol 51(5):1024–1027
Ambe-Ono Y, Sato K, Totsuka K, Yoshihara Y, Nakamori S (1996) Improved L-leucine production by an α-aminobutyric acid resistant mutant of Brevibacterium lactofermentum. Biosci Biotechnol Biochem 60(8):1386–1387
Gusyatiner MM, Voroshilova EB, Rostova YG, Ivanovskaya LV, Lunts MG, Khourges EM (2004) Method for producing L-leucine. US Patent 2004/0091980
Katashkina J, Lunts M, Doroshenko V, Fomina S, Skorokhodova A, Ivanovskaya L, Mashko S (2006) Method for producing an L-amino acid using a bacterium with an optimized level of gene expression. US Patent 2006/0063240
Azuma T, Nakanishi T, Hagino H (1987) Properties of revertants appearing in L-leucine fermentation culture broth. Agric Biol Chem 51(12):3245–3249
Weber C, Farwick A, Benisch F, Brat D, Dietz H, Subtil T (2010) Trends and challenges in the microbial production of lignocellulosic bioalcohol fuels. Appl Microbiol Biotechnol 87:1303–1315
Dellomonaco C, Fava F, Gonzalez R (2012) The path to next generation biofuels:successes and challenges in the era of synthetic biology. Microb Cell Fact 9:3
Blombach B, Eikmanns BJ (2011) Current knowledge on isobutanol production with Escherichia coli, Bacillus subtilis and Corynebacterium glutamicum. Bioeng Bugs 2:346–350
Atsumi S, Hanai T, Liao JC (2008) Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature 451(7174):86–89
Bastian S, Liu X, Meyerowitz JT, Snow CD, Chen MM, Arnold FH (2011) Engineered ketol-acid reductoisomerase and alcohol dehydrogenase enable anaerobic 2-methylpropan-1-ol production at theoretical yield in Escherichia coli. Metab Eng 13(3):345–352
Li S, Huang D, Li Y, Wen J, Jia X (2012) Rational improvement of the engineered isobutanol-producing Bacillus subtilis by elementary mode analysis. Microb Cell Fact 11:101
Blombach B, Riester T, Wieschalka S, Ziert C, Youn JW, Wendisch VF et al (2011) Corynebacterium glutamicum tailored for efficient isobutanol production. Appl Environ Microbiol 77(10):3300–3310
Chen X, Nielsen KF, Borodina I, Kielland-Brandt MC, Karhumaa K (2011) Increased isobutanol production in Saccharomyces cerevisiae by overexpression of genes in valine metabolism. Biotechnol Biofuels 4:21
Ida K, Ishii J, Matsuda F, Kondo T, Kondo A (2015) Eliminating the isoleucine biosynthetic pathway to reduce competitive carbon outflow during isobutanol production by Saccharomyces cerevisiae. Microb Cell Fact 14:62
Yamamoto S, Suda M, Niimi S, Inui M, Yukawa H (2013) Strain optimization for efficient isobutanol production using Corynebacterium glutamicum under oxygen deprivation. Biotechnol Bioeng 110(11):2938–2948
Cann AF, Liao JC (2010) Pentanol isomer synthesis in engineered microorganisms. Appl Microbiol Biotechnol 85(4):893–899
Connor MR, Liao JC (2008) Engineering of an Escherichia coli strain for the production of 3-methyl-1-butanol. Appl Environ Microbiol 74(18):5769–5775
Connor MR, Cann AF, Liao JC (2010) 3-Methyl-1-butanol production in Escherichia coli: random mutagenesis and two-phase fermentation. Appl Microbiol Biotechnol 86(4):1155–1164
Cann AF, Liao JC (2008) Production of 2-methyl-1-butanol in engineered Escherichia coli. Appl Microbiol Biotechnol 81(1):89–98
Su H, Jiang J, Lu Q, Zhao Z, Xie T, Zhao H et al (2015) Engineering Corynebacterium crenatum to produce higher alcohols for biofuel using hydrolysates of duckweed (Landoltia punctata) as feedstock. Microb Cell Fact 14:16
Kalinowski J, Bathe B, Bartels D, Bischoff N, Bott M, Burkovski A et al (2003) The complete Corynebacterium glutamicum ATCC 13032 genome sequence and its impact on the production of L-aspartate-derived amino acids and vitamins. J Biotechnol 104(1–3):5–25
Hüser AT, Chassagnole C, Lindley ND, Merkamm M, Guyonvarch A, Elišáková V et al (2005) Rational design of a Corynebacterium glutamicum pantothenate production strain and its characterization by metabolic flux analysis and genome-wide transcriptional profiling. Appl Environ Microbiol 71(6):3255–3268
Nishio Y, Ogishima S, Ichikawa M, Yamada Y, Usuda Y, Masuda T et al (2013) Analysis of L-glutamic acid fermentation by using a dynamic metabolic simulation model of Escherichia coli. BMC Syst Biol 7:92
Trinh CT, Li J, Blanch HW, Clark DS (2011) Redesigning Escherichia coli metabolism for anaerobic production of isobutanol. Appl Environ Microbiol 77(14):4894–4904
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Yamamoto, K., Tsuchisaka, A., Yukawa, H. (2016). Branched-Chain Amino Acids. In: Yokota, A., Ikeda, M. (eds) Amino Acid Fermentation. Advances in Biochemical Engineering/Biotechnology, vol 159. Springer, Tokyo. https://doi.org/10.1007/10_2016_28
Download citation
DOI: https://doi.org/10.1007/10_2016_28
Published:
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-56518-5
Online ISBN: 978-4-431-56520-8
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)