Abstract
The brain is a complex and energy-demanding organ, which like any organ is subject to the ravages of time. Mitochondria are synonymous with their role in energy production, which is particularly critical to high-energy-demanding cells such as neurons.
Here we discuss ageing of the brain, initially setting the scene by introducing the core concepts associated with brain ageing; discussing the physiological, genetic and cognitive changes which occur over time; and subsequently introducing the roles that mitochondria play in the ‘normal’ brain ageing process.
The final section of the chapter discusses the role of both inherited and somatic mitochondrial DNA variation in neurodegeneration, initially in the context of primary mitochondrial disorders (such as Leber’s hereditary optic neuropathy, myoclonic epilepsy and ragged-red fibres and mitochondrial encephalomyopathy lactic acidosis and stroke-like episodes) and subsequently in the context of common, but more complex, neurodegenerative diseases such as Parkinson’s disease, Alzheimer’s disease, Huntington’s disease, amyotrophic lateral sclerosis, Friedreich’s ataxia, hereditary spastic paraplegia and multiple sclerosis.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 The ‘Normal’ Ageing Brain
Like any organ, the human brain is susceptible to the endless march of time with the effects of ageing often manifesting as diverse pathologies. To understand the role of mitochondrial DNA (mtDNA) variation in brain ageing, we must first understand what brain ageing entails.
1.1 Physiological Changes
At the organ level, brain weight and volume decline at a rate of around 5 % per decade after 40 years of age [1], with rates increasing in individuals over 70 years [2]. Volume loss is associated with an increase in ventricular volume and cerebrospinal fluid (CSF) spaces. Longitudinal magnetic resonance imaging (MRI) and comparative cross-sectional studies have shown that age-related brain changes are regional [2, 3] with the hippocampus and frontal lobes most affected. Estimates indicate that, over an age range of 30–90 years, the cerebral cortex, hippocampus and cerebral white matter are subjected to volumetric losses of 14 %, 35 % and 26 %, respectively [4, 5]. The occipital cortex appears least affected.
The exact mechanism of reduction is up for debate, with studies indicating that grey matter shrinkage [6] and changing neuronal morphology [7], rather than the dogmatic theory of neuronal loss, are the principle components contributing to decreasing brain volume. Additional ageing hallmarks include dendritic sprouting [8, 9], a compensatory mechanism to maintain synapse number and cell number [10], a loss of dendritic arbour [11] and a decline in white matter [12].
The effects of ‘normal’ ageing are also seen at the cellular level. Lipofuscin, the pigmented residues of lysosomal digestion, accumulates in neurons; it is a hallmark of ageing and characteristic of a failure to eliminate the products of peroxidation-induced cell damage [13]. Neurofibrillary tangles and plaques, hallmarks of Alzheimer’s disease (AD), are identifiable in hippocampal, amygdala and entorhinal cortex neurons in healthy aged brains [14] although at subclinical levels. Amyloid β-peptide (Aβ)-containing plaques, another hallmark of AD, form in the grey matter of ‘normal’ aged brains, although present only in small numbers compared to the diseased brain. Advanced age increases the frequency of vacuoles, immunoreactive for neurofibrillary tangles (155K and 210K neurofilament polypeptides) [15], in the cell bodies of pyramidal and hippocampal cells leading to granulovacuolar degeneration [16]. Large (30 μm long) rod-shaped, paracrystalline structures, termed Hirano bodies, accumulate in hippocampal pyramidal cells [17]. Similar to granulovacuoles, these bodies appear to be composed of cytoskeletal proteins (including actin) [18] as well as the constituents of neurofibrillary tangles [19]. As we age, a reduction of oxygen through a reduction in blood supply causes cerebral microinfarcts (cellular necrosis). Typically associated with dementia, these microinfarcts, difficult to detect by conventional means (MRI), are surprisingly common in aged brains (typically >80 years) [20].
1.2 Genetic and Transcriptomic Changes
Genetic factors, such as apolipoprotein E haplotype (APOE), have been identified largely through the investigation of pathological ageing such as in AD [21]. However, these studies are mixed, with some indicating age-related defects in elderly ε4-haplotype carriers [22] and others showing no significant effect [23]. Studies in mice indicate that genetic loci (Hipp1a, Ch1 and Hipp5a, Ch5) can modulate hippocampal structure and volume during normal ageing (potentially mediated through the genes: retinoid X receptor gamma, Rxrg, and fibroblast growth factor receptor 3, Fgfr3, respectively) supporting the role of genetics in brain ageing [24].
At the transcriptomic level, work in mice indicates that the gene expression landscape of the brain, particularly genes involved in stress and inflammatory response, changes dramatically over time and can be modulated by dietary restriction [25]. This supports earlier work investigating the role of ageing in mice at the protein level [26]. In humans, a reduction in synaptic plasticity, modulated by a reduction in α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA) receptor expression, is a hallmark of ageing [27]. In addition reductions of key brain transcripts, including N-methyl-D-aspartate (NMDA) receptors (required for learning and memory) [28], GABA-A receptor subunits (required for neurotransmission) [29] and genes involved in long-term potentiation (e.g. CALM1, calmodulin 1, required for calcium signalling) [30], begin at around 40 years of age.
1.3 Cognitive Changes
Cognitive performance declines with age, although the differentiation between ‘normal ageing,’ ‘non-pathological ageing’ and neurodegenerative disease confounds quantitative measurement. The most observable phenotype associated with ‘normal’ brain ageing is a loss of memory, with no significant loss of visuospatial, cognitive function or language [31]. Memory is classified as four distinct classes: episodic (memory of situation, e.g. a memory of a birthday party), semantic (memory of facts, e.g. Washington, DC, is the capital of the USA), procedural (memory of motor skills, e.g. how to walk) and working (transient memory, for new and stored information) [32]. Studies have investigated the effects of ageing on memory using neuropsychological testing and neuroimaging, demonstrating that brain ageing has a profound effect on both episodic and semantic memory.
Episodic memory, typically affected in AD, is thought to decline from middle age onwards, affecting both recall and recognition in elderly life [33]. Semantic memory also decreases from middle age and then declines rapidly during elderly life [33], driven in part by slower cognitive reaction times, lower attention levels and slower processing speeds in aged individuals [6, 34]. In addition to memory, ageing affects an individual’s orientation and attention. Orientation, or the awareness of one’s surroundings, has been shown to decline mildly with advancing age [35], although this is inconsistent between studies, possibly reflecting the interaction of other variables [36]. Attention, or our ability to focus our brain’s resources on a task, declines with age; moreover secondary phenotypes associated with advancing age [37], i.e. impaired hearing or vision, compound valid attention testing. Indeed, it is worth noting that differentiating these separate cognitive functions, which are intrinsically linked, makes data interpretation difficult [38],
1.4 The Drivers of Brain Ageing
Dopamine, a neurotransmitter involved in motor control, arousal, cognition and reward, declines from early adulthood at a rate of around 10 % per decade, subsequently affecting both cognitive and motor function in aged individuals [33]. Serotonin (a neurotransmitter) and brain-derived neurotrophic factor (BNDF, a neuronal growth factor) are both expressed to maintain synaptic plasticity and both decline with age. We also know that sex hormones, particularly oestrogen, can affect dopaminergic efficacy – purportedly relaying a protective effect in neurodegenerative disorders such as AD [39]. As we age, cerebrovascular efficiency falls, which in turn impairs our ability to metabolise glucose [33], although subsequent data indicates that impaired glucose metabolism may be an effect of cellular atrophy, rather than a cause [40]. Finally, it is becoming clearer that mitochondria play an integral role in brain ageing, from modulating calcium influx and homeostasis [41] to direct dysfunction and the increased proliferation of reactive oxygen species (ROS) [42].
2 Mitochondrial Function in the ‘Normal’ Ageing Brain
Even in the absence of a primary pathogenic defect, changes in mitochondrial function are intrinsically linked to ‘normal’ brain ageing, attributed principally to increased mitochondrial dysfunction, a direct loss of cellular energy or an increase in ROS production, which increases mitochondrial decay [43]. It is worth noting that, from a bioenergetic perspective, the brain is an extremely heterogeneous and dynamic organ; for example, there is a reciprocal relationship between neurons, which have aerobic terminals, and astrocytes, which are more dependent on glycolysis [44]. This complexity, which makes energetic interpretations difficult, must be considered when investigating mitochondrial function in the brain.
Aged animal brains show a significant decline in mitochondrial function, characterised as a loss of phosphorylation capacity, decreased mitochondrial membrane potential [45], decreased respiration, ATP synthesis and activation of the mitochondrial permeability transition pore. The aged brain shows a characteristic reduction in two key respiratory chain (RC) enzymes: NADH/ubiquinone oxidoreductase (complex I, CI) and cytochrome c oxidase (complex IV, CIV) [46]; however, the extent and localisation of the RC deficit are variable [47]. This reduction in RC activity appears to trigger a corresponding proliferation of mitochondria with age, presumably an attempt to attenuate a loss of cellular energy [48] and neuronal mtDNA [49].
The brain, a high-energy organ, is particularly vulnerable to oxidative damage, in part due to a lack of antioxidant enzymes and the increased abundance of polyunsaturated fats, transition metals and the high rate of oxygen consumption [50]. Markers of oxidative stress, carbonylation (protein oxidation), lipid oxidation and the oxidation of the mitochondrial genome [51] are increased with age; however, whether increased oxidative stress markers are a result of a loss of antioxidant enzyme activity or an increase in pro-stress factors is contentious [52]. The coexistence of mitochondrial dysfunction and increased oxidative stress in the brain likely acts as a trigger for apoptotic activation, yet the role of apoptosis in the ‘normal’ ageing brain is controversial, with studies indicating that mitochondrially mediated apoptosis is not a cardinal element of ‘normal’ brain ageing [53] and is rather restricted to disease [54].
2.1 Mitochondrial DNA in the ‘Normal’ Ageing Brain
The risk of mtDNA mutation is high, primarily due to its localisation near to the site of oxidative stress [55], and is not restricted by tissue type or disease [56]. The result is a spectrum of mtDNA variation, ranging from the ‘inherited’ population-level polymorphisms which classify mtDNA haplogroups [57] to the acquired or somatic variation that accumulates with age (either large-scale deletions or single base-pair substitutions).
Despite strong links to mitochondrial functionality [58, 59], there is little evidence for the role of inherited mtDNA variation in ‘normal’ brain ageing. However, there are several reports linking inherited variation to the progression of neurodegenerative disorders such as Parkinson’s and Alzheimer’s disease (discussed later).
Conversely, there is mounting evidence indicating that acquired mtDNA variation accumulates in the brain during ‘normal’ ageing and much work has focused on the aptly named ‘common deletion’ (a 5 Kb deletion with known disease associations [60]), with reports showing that age-related oxidative damage increases the levels of the common deletion in several regions of the ‘healthy’ brain, particularly in the cortex, cerebellum, putamen [61, 62] and more recently substantia nigra and basal ganglia [60], and appears to correlate with regional differences in energy demand [63]. In addition to the common deletion, several studies have reported an increase in random and variable mtDNA deletions, typically smaller in size (>50 bp), which can be differentially distributed amongst brain regions [62, 64]. Studies of acquired single nucleotide variation in the brain are sparse; however studies have shown that healthy individuals harbour an increase in the frequency of low-level heteroplasmic variation in cytochrome c oxidase subunit I (MTCO1), the core catalytic subunit of complex IV of the RC [65] and the non-coding, regulatory, D-loop region [66].
3 Mitochondrial DNA Mutation and Neurodegeneration
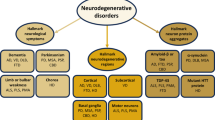
As discussed, mitochondria are the cellular powerhouses, supplying critical ATP through oxidative phosphorylation. Secondary roles in apoptosis, responses to cellular stress and modulation of reactive oxygen species make mitochondrial function a central component of almost all cellular survival (see Chap. 1). The central nervous system (CNS), a highly energy-demanding system, is dependent on efficient mitochondrial function. A disruption of this function, through mitochondrial DNA mutation, is in many cases associated with neurodegenerative disease (Fig. 3.1).
3.1 Mitochondrial DNA Mutations
Mitochondrial DNA, a circular 16.5 Kb molecule encoding the 13 core RC polypeptides, has a significantly higher mutation rate compared to its nuclear counterpart (around 3× or 2.7 × 10−5 base pairs per generation) [67], due in part to an absence of histone and its proximity to the inner mitochondrial membrane – the site of ROS production. However, mitochondria do contain antioxidant and DNA repair enzymes, including oxoguanine glycosylase (OGG1, responsible for 8-oxoguanine base excision) and (MUTYH, involved in oxidative DNA damage repair); furthermore mitochondrial gene expression is adaptive and isolated from the nucleus, allowing the post-translational machinery to generate individual gene products in response to ROS through retrograde signalling [68]. This retrograde signalling stimulates an adaptive nuclear response to mtDNA impairment, which modulates the expression of over 40 nuclear genes [69].
This high mutation rate and lack of effective mtDNA repair mechanisms lead to the formation of two classes of mtDNA variants: (1) inherited mtDNA variants, typically single base-pair exchanges that are present at varying population frequencies, and (2) acquired or somatic mtDNA variants, which can be either single base-pair exchanges or deletions of mtDNA.
3.2 Primary Mitochondrial Disorders and Neurodegeneration
Since the 1980s mitochondrial dysfunction has been a hallmark of metabolic disease [70] and subsequently over 300 mtDNA mutations have been associated with disease [71]. More recently, the phenotypic spectrum has broadened to include neurodegenerative disease [72], typically as a function of a primary mtDNA defect.
Leber’s hereditary optic neuropathy (LHON) is the commonest cause of maternally inherited blindness and a cardinal example of how mtDNA mutations cause neurodegeneration [73]. MtDNA mutations (m.3460G>A, m.11778G>A and m.14484T>C in ~90 % of cases) in NADH-dehydrogenase subunits (complex I) reduce cellular energy and increase oxidative stress and ROS production to cause a selective loss of retinal ganglion cells (RGC) in the retina whilst sparing the retinal pigmented epithelium and photoreceptor layer [74]. Atrophy of neuronal cell bodies and demyelination of the optic nerve are associated with impaired activity of the EAAT1 (excitatory amino acid transporter) glutamate transporter and increased mtDNA mutation-mediated ROS production triggering the apoptotic cascade [75]. RGC selectivity is attributed to high energy demand; however, atypical individuals may experience secondary neuronal loss.
Myoclonic epilepsy with ragged-red fibres (MERRF [76]) is typically attributed to m.8344A>G in the mitochondrial tRNA lysine gene (MT-TK) and presents as a progressive degeneration of the olivocerebellar pathway [77, 78], with severe neuronal loss in the inferior olivary complex, Purkinje cells and dentate nucleus. Biochemical studies indicate that m.8344A>G significantly impairs mitochondrial protein synthesis, hampering oxidative phosphorylation, decreasing ATP production and mitochondrial membrane potential and finally culminating in apoptotic cell death [79].
In addition, mtDNA mutations are associated with cerebellar or sensory ataxia [80], where a secondary loss of Purkinje or other cerebellar cells is seen in syndromes typically associated with a primary mtDNA defect, including Kearns-Sayre syndrome (KSS [81]), mitochondrial encephalopathy and stroke-like episodes (MELAS [82]) and neuropathy, ataxia and retinitis pigmentosa (NARP [83]).
3.3 Mitochondrial DNA Mutation in Complex Neurodegeneration
There is increasing evidence that an increase in oxidative stress and apoptosis is linked to the aetiopathogenesis of several age-related neurodegenerative diseases. Much research has focused on the role of mitochondrial impairment and oxidative stress in the onset and progression of the major ageing brain disorders, particularly Parkinson’s disease (PD) and AD; however, research has also linked mitochondrial mutations to Huntington’s disease (HD) and amyotrophic lateral sclerosis (ALS) (see Fig. 3.1).
3.3.1 Parkinson’s Disease
Parkinson’s disease is a degenerative disorder of the central nervous system, primarily affecting the motor system through a selective loss of dopaminergic neurons in the substantia nigra pars compacta. Early disease is typically movement related, manifesting as tremor, rigidity, slowness of movement initiation and gait instability, progressing to cognitive impairment in advanced stages.
The link between mitochondrial dysfunction and postmortem PD tissue is well recognised [84]. The substantia nigra pars compacta (SNpc), the most vulnerable tissue in PD patients, suffers an age-related reduction of complex I activity [85] and exposure to potent inhibitors of respiratory chain activity such as rotenone [86], 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP) [87] and certain pesticides [88] manifest with neurological abnormalities similar to PD. The combined reduction in cellular energy availability and an increase in ROS may lower the threshold for neuronal apoptosis killing the energy-demanding dopaminergic neurons in the SNpc.
Despite a consistent link between mitochondrial proteomics and PD, the underlying genetic mechanisms remain complex and conflicting, due in part to the complex interactions of both inherited and acquired mtDNA mutations.
The role of inherited mtDNA variation, mediated primarily through mtDNA haplogroups, in the aetiopathology of PD is well studied, although the results are often conflicting and contentious.
Evolutionarily, inherited mtDNA mutations have created stable population subgroups termed ‘mitochondrial haplogroups,’ groups of phylogenetically similar mtDNA molecules, separated by a collection of stable and frequent polymorphic variants [57]. Many of these subdivisions occurred over 50,000 years ago, developing as humans migrated into new geographical regions. Over 95 % of Europeans belong to one of ten major haplogroups, H, J, T, U, K (a subgroup of U), M, I, V, W and X, with each haplogroup defined by specific mtDNA sequence variants within the population. The role of inherited mtDNA variants and their defined haplogroups has evolved; once thought to be benign population variants, mtDNA haplogroups are now known to directly affect mitochondrial function [58, 89] and have become associated with a broad spectrum of human disease [90–92]. In addition, there is evidence to suggest that inherited mtDNA may influence neuronal cytosolic pH and calcium regulation, which over time could influence neuronal function and viability [93].
In early work, the common European haplogroup J, driven largely by the presence of a single nucleotide substitution in MT-ND3 (m.10398G>A), was shown to be underrepresented in PD, suggestive of a protective effect, and is interesting given the purported effects of haplogroup J on ‘normal’ ageing, where it is overrepresented in healthy centenarians [94]. This is supported by a moderately powered study which found that a phylogenetically linked cluster containing haplogroup J (JTUK) reduced PD risk [95]. Smaller studies have reported a contradictory increase in PD risk with haplogroup J [96], but sample number and genetic heterogeneity, critically misunderstood in the highly mutable mtDNA [97], may account for this disparity. More recently, a much larger study and meta-analysis confirmed the role of haplogroup J/K in reducing the risk of PD [98], an effect which was replicated in a follow-up study later [99]. A meta-analysis of all available mtDNA-PD data was able to identify that the phylogenetically linked H/V haplogroups increased the risk of PD [99].
It is intriguing that ostensibly common, and once thought benign, variation in the mitochondrial genome could convey a direct effect on the pathoaetiology of PD. More likely, haplogroup J/K/H/V are genetic tags of further mtDNA variation not directly investigated in these studies. For example, haplogroup K contains the lowest frequency of nonsynonymous complex I gene variants (when compared to the other haplogroups), raising the possibility that natural selection against genetic variation of complex I reduces the risk of PD in haplogroup K individuals. Similarly, an association with haplogroup H, which is linked to infection survival, raises the possibility that natural selection has led to the emergence of variants which predispose to age-related neurodegenerative disease through antagonistic pleiotropy. Similar to HD, where an increased CAG(n) repeat length in the Huntingtin gene causes a neurodegenerative disorder, but is associated with a lower incidence of cancer, recent mutations in mtDNA may be tolerated in humans because they increase the chance of surviving early-life insults such as sepsis, but are pathogenic in later life through the increased generation of reactive oxygen species. In both of these instances, advances in genotype and sequencing technology will allow researchers to understand the role of inherited mtDNA variants, by enabling more and more sequences to be collected and analysed.
More speculative is the exact role of acquired or age-related somatic, mtDNA variants. Much early work focused on the role of mtDNA deletions, large and small lesions in the genome. These deletions typically remove large proportions of the heavily coded mtDNA molecule, dramatically affecting mitochondrial protein synthesis and replication, which due to their obvious pathogenic nature appear heteroplasmic (a mixed population of mtDNA molecules within a cell or tissue).
Early studies failed to find a direct association between mtDNA deletion levels in the brains of PD patients, showing no significant increase when compared to age-matched healthy control tissue [100–102]. Advances in mtDNA isolation technology, enabling capture from single cells, and improvements in the PD diagnostic efficacy improved deletion detection. Studies taking advantage of these technologies show that the frequency of mtDNA deletions in individual neurons in the SNpc of PD patients clonally accumulates with age, increasing after 65 years old [103], and is associated with a decrease in cytochrome c oxidase activity [104]. Both of these reports suggest that the SNpc is particularly vulnerable to free radical-mediated mtDNA damage and that mtDNA deletions may play a role in the pathoaetiology of PD.
The identification of mtDNA deletions in non-PD-related disease [105] and healthy individuals [106] questions the exact contribution of these lesions to the development of PD. In addition, the observation of mtDNA deletions in newborn brains [107] raises the question of whether the clonal expansion mtDNA deletions are the cause of, or are driven by, SNpc failure.
Studies of acquired mtDNA point mutations, typically heteroplasmic, in PD are extremely limited and are often focused on idiopathic cases with no evidence of mitochondrial dysfunction. Studies have identified heteroplasmic variation in PD cases [108, 109]; however, without supporting biochemical or functional data, these variants and their role in the development of PD are difficult to interpret.
3.3.2 Alzheimer’s Disease
Alzheimer’s disease is a progressive loss of neurons and synapses in the cerebral cortex and subcortical regions and is the commonest cause of dementia in the elderly [110]. The disease is typically associated with plaques or neurofibrillary tangles (aggregates of hyper-phosphorylated tau protein).
Like PD, there are several lines of evidence linking mitochondrial dysfunction and oxidative damage in the aetiopathogenesis of AD. Animal experiments using mice which overexpress precursor amyloid protein (APP) demonstrated mitochondrial dysfunction and ATP production through a reduction in mitochondrial protein synthesis [111] or an interaction with mitochondrial matrix proteins [112]. Mitochondrial dysfunction, primarily a reduction in neuronal cell energy, promotes tau phosphorylation, an indication that mitochondrial vitality and oxidative stress are linked to the neurofibrillary tangles seen in AD [113].
Unlike PD, the mitochondrial genetics of AD remain nebulous. Studies have shown that inherited mtDNA variants, again mediated through mtDNA haplogroups, may affect the pathoaetiology of AD; however, like PD these studies are conflicting. Reports show that haplogroups H, U, K, T, I, W and X (seven of the ten typical European haplogroups) associate with an increased risk of developing AD, with effect sizes ranging from OR>2 [114–120]. If correct, the only haplogroups not associated with AD are haplogroups J and V, which is seemingly unlikely, especially given the reciprocal relationship between mtDNA haplogroups.
It is possible that geographical variation could affect these results, with the relative contribution of specific mtDNA variants varying in different ethnic groups; however, this would imply that the overall effect of inherited mtDNA on AD is small, or restricted to very rare pathogenic variants, and would certainly require significantly larger studies to fully investigate. A more recent and comprehensive study supports this hypothesis, investigating the role of inherited mtDNA variants in a large AD cohort concluding that there is no clear role for inherited mtDNA variation in the pathoaetiology of AD [121]. Further, when mtDNA from AD patients is transferred to Rho0 cell lines (cells devoid of native mtDNA), a biochemical defect is observed, suggesting an effect rare inheritable mtDNA abnormalities [122]; however, to date, studies have failed to find an overrepresentation of inherited mtDNA mutations in AD patients [123].
Despite strong evidence that amyloid beta (Aβ) enters mitochondria and increases ROS production [124], likely increasing mtDNA mutation formation, few studies have estimated the contribution of acquired mtDNA variation in AD. The role of the ‘common’ mtDNA deletion, a recurring pathogenic ~5 Kb mtDNA deletion, [125] was assessed in AD, but no significant association was found [126]. The same authors did identify a significantly increased acquired mtDNA mutational burden, two to threefold higher than controls, in the parietal gyrus, hippocampus and cerebellum of AD patient brains – consistent with the hypothesis of oxidative-induced cell death seen in AD [49] and supported by a more recent study [65]. Additionally, it has been hypothesised that somatic mutations in the mtDNA control region (displacement or D-loop) impair mitochondrial transcription and translation in AD brains [127].
3.3.3 Huntington’s Disease
Huntington’s disease is a comparatively early-onset neurodegenerative disorder (physical symptoms can begin at any age from infancy to old age, but usually begin between 35 and 44 years of age) and is caused by autosomal dominant repeat expansions in the Huntington gene. Early pathology begins in the striatum but rapidly progresses to other brain regions.
Mitochondrial dysfunction appears to play an important role in the progression of HD, characterised by metabolic deficits and a decrease in core mitochondrial enzymes [128]. In addition, mutant huntingtin protein (Htt) interacts with the mitochondrial membrane, detrimentally affecting calcium metabolism [129], and Htt cytotoxicity triggers mitochondrial transcriptional changes and initiates a cascade of energy failure ultimately leading to cell death [130]. At the mtDNA level, increased oxidative stress resulting from Htt toxicity induces mtDNA lesion formation and a reduction in mtDNA copy number, which in turn triggers the vicious circle of ROS-mediated mtDNA damage [130]; however, this appears more as a consequence rather than a cause of disease progression. HD patients have higher frequencies of mtDNA deletions when compared to age-matched controls; however, much of this is based on measurements in peripheral blood [131, 132], with only one study investigating cortical tissue [133].
Studies of inherited mtDNA variation are conflicted. Early work investigating the role of mtDNA haplogroups in HD concluded that they do not affect disease development or progression [134]; however, this study was relatively small compared to typical genetic association studies. Contrary to this, a more recent study demonstrated that HD patients with a haplogroup H (the commonest European haplogroup, ~40–45 % population) background showed significantly higher ATP concentrations than non-H HD patients [135]. This is puzzling, given the synonymous nature of the haplogroup H defining mtDNA variant (m.7028C, Ala375Ala in MTCOI), and likely reflects either a statistical power inadequacy or haplogroup tagging or rare mtDNA variants, which only analysis of whole mtDNA genome sequencing could identify.
3.3.4 Amyotrophic Lateral Sclerosis
Amyotrophic lateral sclerosis (ALS, often termed Lou Gehrig’s disease in the USA, Charcot disease or motor-neurone disease in the UK) is a typically sporadic disease (~95 % of cases), causing progressive loss of neurons in the motor cortex and spinal cord. Clinical hallmarks of ALS include muscle stiffness and twitching, gradually progressing to atrophy [136].
Although the underlying mechanism is not fully understood, reports link nDNA mutations in superoxide dismutase 1 (SOD1), one of three superoxide dismutases responsible for destroying free superoxide radicals, to mitochondrial dysfunction in ALS patients [137]. Mutant SOD1 inhibits voltage-dependent anion selective protein channel 1 (VDAC1) [138], reducing energy production and driving an increase in oxidative damage in transgenic mice motor neurons [139]. ALS patients were found to harbour significantly higher levels (>30×) of the mtDNA common deletion, compared to matched controls, in the motor cortex [140], and spinal cord [141]. It is likely, given the reported SOD1-mediated reduction in mtDNA repair enzymes in ALS patients [142], that mtDNA lesions and point mutations are proliferating; although this has yet to be fully investigated.
3.3.5 Friedreich’s Ataxia
Friedreich’s ataxia (FA) is the commonest form of hereditary ataxia, pathophysiologically characterised as a reduction of the mitochondrial protein frataxin in the peripheral sensory nerves, dorsal root ganglia, posterior columns, spinocerebellar and corticospinal tracts of the spinal cord, gracile and cuneate nuclei, dorsal nuclei of Clarke and dentate nucleus [143]. This deficiency causes an accumulation of iron in mitochondria and a reciprocal depletion in neuronal cytosol, which in turn leads to respiratory chain dysfunction and an increase in oxidative stress [144]. At the mtDNA level, frataxin-deficient yeast hybrid cell lines show a progressive loss of mtDNA and a reciprocal rise in mtDNA lesion formation [145, 146].
3.3.6 Hereditary Spastic Paraplegia
Hereditary spastic paraplegia (HSP) is not a single entity, rather a group of neurodegenerative disorders phenotypically characterised by progressive lower extremity weakness and spasticity (progressive stiffness and contraction) [147]. Mitochondrial dysfunction is linked to two forms of HSP: SPG13, the mitochondrial chaperonin Hsp60 [148], and SPG7, a nuclear-encoded mitochondrial metalloprotease protein which increases ROS [149].
Investigations into the effects of inherited mtDNA mutation on HSP are limited. An mtDNA mutation in MTATP6 (m.9176T>C) was identified in a large HSP pedigree [150] and a mutation in the mitochondrial 12 s rRNA (m.1432A>G) was identified in an Amish family who demonstrated abnormal mRNA maturation [151]. A single study investigating the role of mtDNA haplogroups appears negative, but is largely inconclusive due to low sample size and genetic heterogeneity [152].
3.3.7 Multiple Sclerosis
Multiple sclerosis (MS) is one of the commonest neuroinflammatory diseases in the world [153] and is phenotypically characterised by autonomic, visual, motor and sensory deficits [154]. Until relatively recently, MS pathology was restricted to the formation of lesions or plaques and the destruction of myelin in the spinal cord; however, improvements in imaging technology confirm that neurodegeneration begins in the grey and white matter of MS patients [155, 156].
Mitochondrial dysfunction appears a common component of MS. N-Acetylaspartate and N-acetylaspartylglutamate (NAA), which correspond to MS relapses, are markers for neuronal integrity produced by mitochondria and indicate mitochondrial functionality [157]. RC chain function, particularly complex I, is reduced in MS tissue and complexes I and III are reduced in non-lesional MS motor cortex tissue [158, 159]. Further, there are established links between inherited mtDNA mutation and MS, with female primary LHON (discussed previously) mutation carriers presenting with an MS-like phenotype – termed ‘Harding’s disease’ [160]. In support, m.3460G>A (a primary LHON mutation) was identified in a small cohort of neuromyelitis optica cases, where spinal nerve degeneration is a predominant feature [161], indicating that inherited mtDNA variants could modulate MS pathogenesis. However, subsequent studies focusing on the role of LHON mutations (m.3460G>A, m.11778G>A and m.14484T>C) refute a link to MS [162, 163].
Early attempts to link mtDNA haplogroups to MS indicate that European super-haplogroup ‘UK’ increases MS risk, with a reciprocal risk reduction for individuals on a super-haplogroup ‘JT’ background [164, 165]. These associations were confirmed in a much larger investigation into inherited mtDNA mutations in late-onset disease, with haplogroup U increasing MS risk and haplogroup J/K reducing MS risk [99].
Attempts to identify somatic or acquired mtDNA mutations, focusing on the identification of mtDNA deletions in normal-appearing grey and white matter or paraspinal muscle, failed to find a significant difference between MS cases and controls [166, 167].
4 Conclusion
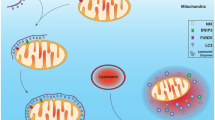
The modulation of mitochondrial function is a key component to neurodegeneration, both during ‘normal’ ageing and disease (Fig. 3.2). Changes in mitochondrial function as we age and when disease manifests are often mediated by mitochondrial DNA variation, which can be either inherited (typically single base-pair exchanges) or acquired as we age (either single base-pair exchanges or mtDNA deletions).
Despite strong evidence of mitochondrial involvement in several neurodegenerative diseases, the ubiquity of mitochondria makes identifying disease-causing mtDNA mutations, and how they interact with disease-specific pathologies, difficult. Nevertheless, if we are to fully understand if this phenomenon is the cause or correlation, it is important that studies of mtDNA in neurodegenerative disease continue.
As the evidence demonstrates, mitochondrial function, mediated through genomic flux, can play a role in neuronal loss, making mitochondria a useful target for therapeutic strategies in an increasingly aged population.
References
Svennerholm L, Bostrom K, Jungbjer B. Changes in weight and compositions of major membrane components of human brain during the span of adult human life of Swedes. Acta Neuropathol. 1997;94(4):345–52.
Scahill RI, Frost C, Jenkins R, Whitwell JL, Rossor MN, Fox NC. A longitudinal study of brain volume changes in normal aging using serial registered magnetic resonance imaging. Arch Neurol Chicago. 2003;60(7):989–94.
Trollor JN, Valenzuela MJ. Brain ageing in the new millennium. Aust N Z J Psychiatry. 2001;35(6):788–805.
Esiri MM, Wilcock GK, Morris JH. Neuropathological assessment of the lesions of significance in vascular dementia. J Neurol Neurosurg Psychiatry. 1997;63(6):749–53.
Jernigan TL, Archibald SL, Fennema-Notestine C, Gamst AC, Stout JC, Bonner J, et al. Effects of age on tissues and regions of the cerebrum and cerebellum. Neurobiol Aging. 2001;22(4):581–94.
Raz N. The ageing brain: structural changes and their implications for cognitive ageing. In: Dixon R, Bäckman L, Nilssonn L, editors. New frontiers in cognitive ageing. New York: Oxford University Press; 2004.
Burke SN, Barnes CA. Neural plasticity in the ageing brain. Nat Rev Neurosci. 2006;7(1):30–40.
Kolb B, Gibb R, Robinson TE. Brain plasticity and behavior. Curr Dir Psychol Sci. 2003;12(1):1–5.
Kolb B, Whishaw IQ. Brain plasticity and behavior. Annu Rev Psychol. 1998;49:43–64.
Anderton BH. Ageing of the brain. Mech Ageing Dev. 2002;123(7):811–7.
Flood DG, Buell SJ, Horwitz GJ, Coleman PD. Dendritic extent in human dentate gyrus granule cells in normal aging and senile dementia. Brain Res. 1987;402(2):205–16.
Head D, Buckner RL, Shimony JS, Williams LE, Akbudak E, Conturo TE, et al. Differential vulnerability of anterior white matter in nondemented aging with minimal acceleration in dementia of the Alzheimer type: evidence from diffusion tensor imaging. Cereb Cortex. 2004;14(4):410–23.
Gray DA, Woulfe J. Lipofuscin and aging: a matter of toxic waste. Sci Aging Knowledge Environ. 2005;2005(5):re1.
Braak H, Braak E. Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol. 1991;82(4):239–59.
Kahn J, Anderton BH, Probst A, Ulrich J, Esiri MM. Immunohistological study of granulovacuolar degeneration using monoclonal-antibodies to neurofilaments. J Neurol Neurosurg Psychiatry. 1985;48(9):924–6.
Xu M, Shibayama H, Kobayashi H, Yamada K, Ishihara R, Zhao P, et al. Granulovacuolar degeneration in the hippocampal cortex of aging and demented patients – a quantitative study. Acta Neuropathol. 1992;85(1):1–9.
Gibson PH, Tomlinson BE. Numbers of Hirano bodies in the hippocampus of normal and demented people with Alzheimer’s disease. J Neurol Sci. 1977;33(1–2):199–206.
Galloway PG, Perry G, Gambetti P. Hirano body filaments contain actin and actin-associated proteins. J Neuropathol Exp Neurol. 1987;46(2):185–99.
Dickson DW, Liu WK, Kress Y, Ku J, DeJesus O, Yen SH. Phosphorylated tau immunoreactivity of granulovacuolar bodies (GVB) of Alzheimer’s disease: localization of two amino terminal tau epitopes in GVB. Acta Neuropathol. 1993;85(5):463–70.
Smith EE, Schneider JA, Wardlaw JM, Greenberg SM. Cerebral microinfarcts: the invisible lesions. Lancet Neurol. 2012;11(3):272–82.
Saunders AM, Strittmatter WJ, Schmechel D, George-Hyslop PH, Pericak-Vance MA, Joo SH, et al. Association of apolipoprotein E allele epsilon 4 with late-onset familial and sporadic Alzheimer’s disease. Neurology. 1993;43(8):1467–72.
Deary IJ, Whiteman MC, Pattie A, Starr JM, Hayward C, Wright AF, et al. Apolipoprotein E gene variability and cognitive functions at age 79: a follow-up of the Scottish Mental Survey of 1932. Psychol Aging. 2004;19(2):367–71.
Jorm AF, Mather KA, Butterworth P, Anstey KJ, Christensen H, Easteal S. APOE genotype and cognitive functioning in a large age-stratified population sample. Neuropsychology. 2007;21(1):1–8.
Lu L, Airey DC, Williams RW. Complex trait analysis of the hippocampus: mapping and biometric analysis of two novel gene loci with specific effects on hippocampal structure in mice. J Neurosci. 2001;21(10):3503–14.
Lee CK, Weindruch R, Prolla TA. Gene-expression profile of the ageing brain in mice. Nat Genet. 2000;25(3):294–7.
Tsugita A, Kawakami T, Uchida T, Sakai T, Kamo M, Matsui T, et al. Proteome analysis of mouse brain: two-dimensional electrophoresis profiles of tissue proteins during the course of aging. Electrophoresis. 2000;21(9):1853–71.
Henley JM, Wilkinson KA. AMPA receptor trafficking and the mechanisms underlying synaptic plasticity and cognitive aging. Dialogues Clin Neurosci. 2013;15(1):11–27.
Magnusson KR, Brim BL, Das SR. Selective vulnerabilities of N-methyl-D-aspartate (NMDA) receptors during brain aging. Front Aging Neurosci. 2010;2:11.
Limon A, Reyes-Ruiz JM, Miledi R. Loss of functional GABA(A) receptors in the Alzheimer diseased brain. Proc Natl Acad Sci U S A. 2012;109(25):10071–6.
Lu T, Pan Y, Kao SY, Li C, Kohane I, Chan J, et al. Gene regulation and DNA damage in the ageing human brain. Nature. 2004;429(6994):883–91.
Mayeux R, Small SA, Tang MX, Tycko B, Stern Y. Memory performance in healthy elderly without Alzheimer’s disease: effects of time and apolipoprotein-E. Neurobiol Aging. 2001;22(4):683–9.
Parkin A. Human-memory and amnesia – Cermak, Ls. Q J Exp Psychol A. 1983;35(Aug):544–5.
Nyberg L, Bäckman L. Cognitive ageing: a view from brain imaging. In: Dixon R, Bäckman L, Nilsson L, editors. New frontiers in cognitive ageing. New York: Oxford University Press; 2004.
Lustig C, Buckner RL. Preserved neural correlates of priming in old age and dementia. Neuron. 2004;42(5):865–75.
Sweet JJ, Suchy Y, Leahy B, Abramowitz C, Nowinski CJ. Normative clinical relationships between orientation and memory: age as an important moderator variable. Clin Neuropsychol. 1999;13(4):495–508.
Hopp GA, Dixon RA, Grut M, Bäckman L. Longitudinal and psychometric profiles of two cognitive status tests in very old adults. J Clin Psychol. 1997;53(7):673–86.
Kensinger EA. Cognition in aging and age related disease. In: Hof PR, Mobbs CV, editors. Handbook of the neuroscience of aging. London: Elsevier; 2009. p. 249–56.
Cabeza R. Cognitive neuroscience of aging: contributions of functional neuroimaging. Scand J Psychol. 2001;42(3):277–86.
Compton J, van Amelsvoort T, Murphy D. HRT and its effect on normal ageing of the brain and dementia. Br J Clin Pharmacol. 2001;52(6):647–53.
Ibanez V, Pietrini P, Furey ML, Alexander GE, Millet P, Bokde AL, et al. Resting state brain glucose metabolism is not reduced in normotensive healthy men during aging, after correction for brain atrophy. Brain Res Bull. 2004;63(2):147–54.
Toescu EC, Verkhratsky A, Landfield PW. Ca2+ regulation and gene expression in normal brain aging. Trends Neurosci. 2004;27(10):614–20.
Melov S. Modeling mitochondrial function in aging neurons. Trends Neurosci. 2004;27(10):601–6.
Balaban RS, Nemoto S, Finkel T. Mitochondria, oxidants, and aging. Cell. 2005;120(4):483–95.
Magistretti PJ, Pellerin L. Cellular mechanisms of brain energy metabolism and their relevance to functional brain imaging. Philos Trans R Soc B. 1999;354(1387):1155–63.
LaFrance R, Brustovetsky N, Sherburne C, Delong D, Dubinsky JM. Age-related changes in regional brain mitochondria from Fischer 344 rats. Aging Cell. 2005;4(3):139–45.
Navarro A, Boveris A. The mitochondrial energy transduction system and the aging process. Am J Physiol Cell Physiol. 2007;292(2):C670–86.
Navarro A. Mitochondrial enzyme activities as biochemical markers of aging. Mol Aspects Med. 2004;25(1–2):37–48.
Barrientos A, Casademont J, Cardellach F, Estivill X, Urbano-Marquez A, Nunes V. Reduced steady-state levels of mitochondrial RNA and increased mitochondrial DNA amount in human brain with aging. Mol Brain Res. 1997;52(2):284–9.
Hirai K, Aliev G, Nunomura A, Fujioka H, Russell RL, Atwood CS, et al. Mitochondrial abnormalities in Alzheimer’s disease. J Neurosci. 2001;21(9):3017–23.
Halliwell B, Gutteridge JMC. Free radicals in biology and medicine. New York: Oxford University Press; 1999.
Navarro A, Boveris A. Brain mitochondrial dysfunction in aging, neurodegeneration, and Parkinson’s disease. Front Aging Neurosci. 2010;2:34.
Ceballospicot I, Nicole A, Clement M, Bourre JM, Sinet PM. Age-related-changes in antioxidant enzymes and lipid-peroxidation in brains of control and transgenic mice overexpressing copper-zinc superoxide-dismutase. Mutat Res. 1992;275(3–6):281–93.
Pollack M, Leeuwenburgh C. Apoptosis and aging: role of the mitochondria. J Gerontol A Biol Sci Med Sci. 2001;56(11):B475–82.
Swerdlow RH. Mitochondria and cell bioenergetics: increasingly recognized components and a possible etiologic cause of Alzheimer’s disease. Antioxid Redox Signal. 2012;16(12):1434–55.
Swerdlow RH. Brain aging, Alzheimer’s disease, and mitochondria. BBA Mol Basis Dis. 2011;1812(12):1630–9.
Payne BAI, Wilson IJ, Yu-Wai-Man P, Coxhead J, Deehan D, Horvath R, et al. Universal heteroplasmy of human mitochondrial DNA. Hum Mol Genet. 2013;22(2):384–90.
Torroni A, Huoponen K, Francalacci P, Petrozzi M, Morelli L, Scozzari R, et al. Classification of European mtDNAs from an analysis of three European populations. Genetics. 1996;144(4):1835–50.
Gomez-Duran A, Pacheu-Grau D, Martinez-Romero I, López-Gallardo E, López-Pérez MJ, Montoya J, et al. Oxidative phosphorylation differences between mitochondrial DNA haplogroups modify the risk of Leber’s hereditary optic neuropathy. Biochim Biophys Acta. 2012;1822(8):1216–22.
Pello R, Martin MA, Carelli V, Nijtmans LG, Achilli A, Pala M, et al. Mitochondrial DNA background modulates the assembly kinetics of OXPHOS complexes in a cellular model of mitochondrial disease. Hum Mol Genet. 2008;17(24):4001–11.
Meissner C, Bruse P, Mohamed SA, Schulz A, Warnk H, Storm T, et al. The 4977 bp deletion of mitochondrial DNA in human skeletal muscle, heart and different areas of the brain: a useful biomarker or more? Exp Gerontol. 2008;43(7):645–52.
Cortopassi GA, Arnheim N. Detection of a specific mitochondrial-DNA deletion in tissues of older humans. Nucleic Acids Res. 1990;18(23):6927–33.
Corraldebrinski M, Horton T, Lott MT, Shoffner JM, Beal MF, Wallace DC. Mitochondrial-DNA deletions in human brain – regional variability and increase with advanced age. Nat Genet. 1992;2(4):324–9.
Pickrell AM, Fukui H, Wang X, Pinto M, Moraes CT. The striatum is highly susceptible to mitochondrial oxidative phosphorylation dysfunctions. J Neurosci. 2011;31(27):9895–904.
Fukui H, Moraes CT. Mechanisms of formation and accumulation of mitochondrial DNA deletions in aging neurons. Hum Mol Genet. 2009;18(6):1028–36.
Lin MT, Simon DK, Ahn CH, Kim LM, Beal MF. High aggregate burden of somatic mtDNA point mutations in aging and Alzheimer’s disease brain. Hum Mol Genet. 2002;11(2):133–45.
Williams SL, Mash DC, Zuchner S, Moraes CT. Somatic mtDNA mutation spectra in the aging human putamen. Plos Genet. 2013;9(12):e1003990.
Schneider S, Excoffier L. Estimation of past demographic parameters from the distribution of pairwise differences when the mutation rates vary among sites: application to human mitochondrial DNA. Genetics. 1999;152(3):1079–89.
Mercer TR, Neph S, Dinger ME, Crawford J, Smith MA, Shearwood AM, et al. The human mitochondrial transcriptome. Cell. 2011;146(4):645–58.
Epstein CB, Waddle JA, Hale 4th W, Davé V, Thornton J, Macatee TL, et al. Genome-wide responses to mitochondrial dysfunction. Mol Biol Cell. 2001;12(2):297–308.
Scholte HR. The biochemical basis of mitochondrial diseases. J Bioenerg Biomembr. 1988;20(2):161–91.
Brandon MC, Lott MT, Nguyen KC, Spolim S, Navathe SB, Baldi P, et al. MITOMAP: a human mitochondrial genome database – 2004 update. Nucleic Acids Res. 2005;33:D611–3.
Morais VA, De Strooper B. Mitochondria dysfunction and neurodegenerative disorders: cause or consequence. J Alzheimers Dis. 2010;20:S255–63.
Kirches E. LHON: mitochondrial mutations and more. Curr Genomics. 2011;12(1):44–54.
Carelli V, Ross-Cisneros FN, Sadun AA. Mitochondrial dysfunction as a cause of optic neuropathies. Prog Retin Eye Res. 2004;23(1):53–89.
Beretta S, Mattavelli L, Sala G, Tremolizzo L, Schapira AH, Martinuzzi A, et al. Leber hereditary optic neuropathy mtDNA mutations disrupt glutamate transport in cybrid cell lines. Brain. 2004;127:2183–92.
Fukuhara N, Tokiguchi S, Shirakawa K, Tsubaki T. Myoclonus epilepsy associated with ragged-red fibres (mitochondrial abnormalities): disease entity or a syndrome? Light-and electron-microscopic studies of two cases and review of literature. J Neurol Sci. 1980;47(1):117–33.
Shoffner JM, Wallace DC. A mitochondrial transfer rnalys mutation causes myoclonic epilepsy and ragged-red fiber disease. Prog Neuropathol. 1991;7:161–7.
Wallace DC, Zheng XX, Lott MT, Shoffner JM, Hodge JA, Kelley RI, et al. Familial mitochondrial encephalomyopathy (MERRF): genetic, pathophysiological, and biochemical characterization of a mitochondrial DNA disease. Cell. 1988;55(4):601–10.
Rommelaere G, Michel S, Malaisse J, Charlier S, Arnould T, Renard P. Hypersensitivity of A8344G MERRF mutated cybrid cells to staurosporine-induced cell death is mediated by calcium-dependent activation of calpains. Int J Biochem Cell B. 2012;44(1):139–49.
Zeviani M, Simonati A, Bindoff LA. Ataxia in mitochondrial disorders. Handb Clin Neurol. 2012;103:359–72.
Kearns TP. External ophthalmoplegia, pigmentary degeneration of the retina, and cardiomyopathy: a newly recognized syndrome. Trans Am Ophthalmol Soc. 1965;63:559–625.
Goto Y, Horai S, Matsuoka T, Koga Y, Nihei K, Kobayashi M, et al. Mitochondrial myopathy, encephalopathy, lactic acidosis, and stroke-like episodes (MELAS): a correlative study of the clinical features and mitochondrial DNA mutation. Neurology. 1992;42(3 Pt 1):545–50.
Holt IJ, Harding AE, Petty RK, Morgan-Hughes JA. A new mitochondrial disease associated with mitochondrial DNA heteroplasmy. Am J Hum Genet. 1990;46(3):428–33.
Winklhofer KF, Haass C. Mitochondrial dysfunction in Parkinson’s disease. BBA Mol Basis Dis. 2010;1802(1):29–44.
Schapira AH, Cooper JM, Dexter D, Jenner P, Clark JB, Marsden CD. Mitochondrial complex I deficiency in Parkinson’s disease. Lancet. 1989;1(8649):1269.
Sherer TB, Betarbet R, Testa CM, Seo BB, Richardson JR, Kim JH, et al. Mechanism of toxicity in rotenone models of Parkinson’s disease. J Neurosci. 2003;23(34):10756–64.
Forno LS, Delanney LE, Irwin I, Langston JW. Similarities and differences between Mptp-induced parkinsonism and Parkinson’s disease. Neuropathologic considerations. Adv Neurol. 1993;60:600–8.
Betarbet R, Sherer TB, MacKenzie G, Garcia-Osuna M, Panov AV, Greenamyre JT. Chronic systemic pesticide exposure reproduces features of Parkinson’s disease. Nat Neurosci. 2000;3(12):1301–6.
Gomez-Duran A, Pacheu-Grau D, Lopez-Gallardo E, Díez-Sánchez C, Montoya J, López-Pérez MJ, et al. Unmasking the causes of multifactorial disorders: OXPHOS differences between mitochondrial haplogroups. Hum Mol Genet. 2010;19(17):3343–53.
Hudson G, Carelli V, Spruijt L, Gerards M, Mowbray C, Achilli A, et al. Clinical expression of Leber hereditary optic neuropathy is affected by the mitochondrial DNA-haplogroup background. Am J Hum Genet. 2007;81(2):228–33.
Fuku N, Park KS, Yamada Y, Nishigaki Y, Cho YM, Matsuo H, et al. Mitochondrial haplogroup N9a confers resistance against type 2 diabetes in Asians. Am J Hum Genet. 2007;80(3):407–15.
Chinnery PF, Elliott HR, Syed A, Rothwell PM, Oxford Vascular Study. Mitochondrial DNA haplogroups and risk of transient ischaemic attack and ischaemic stroke: a genetic association study. Lancet Neurol. 2007;9(5):498–503.
Kazuno AA, Munakata K, Nagai T, Shimozono S, Tanaka M, Yoneda M, et al. Identification of mitochondrial DNA polymorphisms that alter mitochondrial matrix pH and intracellular calcium dynamics. PLoS Genet. 2006;2(8):1167–77.
Rose G, Passarino G, Carrieri G, Altomare K, Greco V, Bertolini S, et al. Paradoxes in longevity: sequence analysis of mtDNA haplogroup J in centenarians. Eur J Hum Genet. 2001;9(9):701–7.
Pyle A, Foltynie T, Tiangyou W, Lambert C, Keers SM, Allcock LM, et al. Mitochondrial DNA haplogroup cluster UKJT reduces the risk of PD. Ann Neurol. 2005;57(4):564–7.
Ross OA, McCormack R, Maxwell LD, Duguid RA, Quinn DJ, Barnett YA, et al. mt4216C variant in linkage with the mtDNA TJ cluster may confer a susceptibility to mitochondrial dysfunction resulting in an increased risk of Parkinson’s disease in the Irish. Exp Gerontol. 2003;38(4):397–405.
Biffi A, Anderson CD, Nalls MA, Rahman R, Sonni A, Cortellini L, et al. Principal-component analysis for assessment of population stratification in mitochondrial medical genetics. Am J Hum Genet. 2010;86(6):904–17.
Hudson G, Nalls M, Evans JR, Breen DP, Winder-Rhodes S, Morrison KE, et al. Two-stage association study and meta-analysis of mitochondrial DNA variants in Parkinson disease. Neurology. 2013;80(22):2042–8.
Hudson G, Gomez-Duran A, Wilson IJ, Chinnery PF. Recent mitochondrial DNA mutations increase the risk of developing common late-onset human diseases. PLoS Genet. 2014;10(5):e1004369.
Ikebe S, Tanaka M, Ohno K, Sato W, Hattori K, Kondo T, et al. Increase of deleted mitochondrial DNA in the striatum in Parkinson’s disease and senescence. Biochem Biophys Res Commun. 1990;170(3):1044–8.
Schapira AH, Holt IJ, Sweeney M, Harding AE, Jenner P, Marsden CD. Mitochondrial DNA analysis in Parkinson’s disease. Mov Disord. 1990;5(4):294–7.
Lestienne P, Nelson I, Riederer P, Reichmann H, Jellinger K. Mitochondrial DNA in postmortem brain from patients with Parkinson’s disease. J Neurochem. 1991;56(5):1819.
Kraytsberg Y, Kudryavtseva E, McKee AC, Geula C, Kowall NW, Khrapko K. Mitochondrial DNA deletions are abundant and cause functional impairment in aged human substantia nigra neurons. Nat Genet. 2006;38(5):518–20.
Bender A, Krishnan KJ, Morris CM, Taylor GA, Reeve AK, Perry RH, et al. High levels of mitochondrial DNA deletions in substantia nigra neurons in aging and Parkinson disease. Nat Genet. 2006;38(5):515–7.
Kakiuchi C, Ishiwata M, Kametani M, Nelson C, Iwamoto K, Kato T. Quantitative analysis of mitochondrial DNA deletions in the brains of patients with bipolar disorder and schizophrenia. Int J Neuropsychopharmacol. 2005;8(4):515–22.
Corral-Debrinski M, Horton T, Lott MT, Shoffner JM, Beal MF, Wallace DC. Mitochondrial DNA deletions in human brain: regional variability and increase with advanced age. Nat Genet. 1992;2(4):324–9.
Nadasi E, Melegh B, Seress L, Kosztolányi G. Mitochondrial DNA deletions in newborn brain samples. Orv Hetil. 2004;145(25):1321–5.
Shoffner JM, Brown MD, Torroni A, Lott MT, Cabell MF, Mirra SS, et al. Mitochondrial DNA variants observed in Alzheimer disease and Parkinson disease patients. Genomics. 1993;17(1):171–84.
Parker Jr WD, Parks JK. Mitochondrial ND5 mutations in idiopathic Parkinson’s disease. Biochem Biophys Res Commun. 2005;326(3):667–9.
Burns A, Iliffe S. Alzheimer’s disease. BMJ. 2009;338:b158.
Anandatheerthavarada HK, Biswas G, Robin MA, Avadhani NG. Mitochondrial targeting and a novel transmembrane arrest of Alzheimer’s amyloid precursor protein impairs mitochondrial function in neuronal cells. J Cell Biol. 2003;161(1):41–54.
Manczak M, Anekonda TS, Henson E, Park BS, Quinn J, Reddy PH. Mitochondria are a direct site of A beta accumulation in Alzheimer’s disease neurons: implications for free radical generation and oxidative damage in disease progression. Hum Mol Genet. 2006;15(9):1437–49.
Melov S, Adlard PA, Morten K, Johnson F, Golden TR, Hinerfeld D, et al. Mitochondrial oxidative stress causes hyperphosphorylation of tau. PLoS One. 2007;2(6):e536.
Lakatos A, Derbeneva O, Younes D, Keator D, Bakken T, Lvova M, et al. Association between mitochondrial DNA variations and Alzheimer’s disease in the ADNI cohort. Neurobiol Aging. 2010;31(8):1355–63.
Santoro A, Balbi V, Balducci E, Pirazzini C, Rosini F, Tavano F, et al. Evidence for sub-haplogroup h5 of mitochondrial DNA as a risk factor for late onset alzheimer’s disease. PLoS One. 2010;5(8):e12037.
Kruger J, Hinttala R, Majamaa K, Remes AM. Mitochondrial DNA haplogroups in early-onset alzheimer’s disease and frontotemporal lobar degeneration. Mol Neurodegener. 2010;5:8.
Maruszak A, Canter JA, Styczynska M, Zekanowski C, Barcikowska M. Mitochondrial haplogroup H and Alzheimer’s disease – is there a connection? Neurobiol Aging. 2009;30(11):1749–55.
van der Walt JM, Dementieva YA, Martin ER, Scott WK, Nicodemus KK, Kroner CC, et al. Analysis of European mitochondrial haplogroups with Alzheimer disease risk. Neurosci Lett. 2004;365(1):28–32.
Carrieri G, Bonafe M, De Luca M, Rose G, Varcasia O, Bruni A, et al. Mitochondrial DNA haplogroups and APOE4 allele are non-independent variables in sporadic Alzheimer’s disease. Hum Genet. 2001;108(3):194–8.
Chagnon P, Gee M, Filion M, Robitaille Y, Belouchi M, Gauvreau D. Phylogenetic analysis of the mitochondrial genome indicates significant differences between patients with Alzheimer disease and controls in a French-Canadian founder population. Am J Med Genet. 1999;85(1):20–30.
Hudson G, Sims R, Harold D, Chapman J, Hollingworth P, Gerrish A, et al. No consistent evidence for association between mtDNA variants and Alzheimer disease. Neurology. 2012;78(14):1038–42.
Swerdlow RH, Parks JK, Cassarino DS, Maguire DJ, Maguire RS, Bennett Jr JP, et al. Cybrids in Alzheimer’s disease: a cellular model of the disease? Neurology. 1997;49(4):918–25.
Tanaka N, Goto Y, Akanuma J, Kato M, Kinoshita T, Yamashita F, et al. Mitochondrial DNA variants in a Japanese population of patients with Alzheimer’s disease. Mitochondrion. 2010;10(1):32–7.
Lustbader JW, Cirilli M, Lin C, Xu HW, Takuma K, Wang N, et al. ABAD directly links Abeta to mitochondrial toxicity in Alzheimer’s disease. Science. 2004;304(5669):448–52.
Cortopassi GA, Shibata D, Soong NW, Arnheim N. A pattern of accumulation of a somatic deletion of mitochondrial-DNA in aging human tissues. Proc Natl Acad Sci U S A. 1992;89(16):7370–4.
Chang SW, Zhang DK, Chung HD, Zassenhaus HP. The frequency of point mutations in mitochondrial DNA is elevated in the Alzheimer’s brain. Biochem Biophys Res Commun. 2000;273(1):203–8.
Coskun PE, Beal MF, Wallace DC. Alzheimer’s brains harbor somatic mtDNA control-region mutations that suppress mitochondrial transcription and replication. Proc Natl Acad Sci U S A. 2004;101(29):10726–31.
Damiano M, Galvan L, Deglon N, Brouillet E. Mitochondria in Huntington’s disease. Biochim Biophys Acta. 2010;1802(1):52–61.
Panov AV, Gutekunst CA, Leavitt BR, Hayden MR, Burke JR, Strittmatter WJ, et al. Early mitochondrial calcium defects in Huntington’s disease are a direct effect of polyglutamines. Nat Neurosci. 2002;5(8):731–6.
Siddiqui A, Rivera-Sanchez S, Castro MD, Acevedo-Torres K, Rane A, Torres-Ramos CA, et al. Mitochondrial DNA damage is associated with reduced mitochondrial bioenergetics in Huntington’s disease. Free Radic Biol Med. 2012;53(7):1478–88.
Banoei MM, Houshmand M, Panahi MS, Shariati P, Rostami M, Manshadi MD, et al. Huntington’s disease and mitochondrial DNA deletions: event or regular mechanism for mutant huntingtin protein and CAG repeats expansion?! Cell Mol Neurobiol. 2007;27(7):867–75.
Chen CM, Wu YR, Cheng ML, Liu JL, Lee YM, Lee PW, et al. Increased oxidative damage and mitochondrial abnormalities in the peripheral blood of Huntington’s disease patients. Biochem Biophys Res Commun. 2007;359(2):335–40.
Horton TM, Graham BH, Corral-Debrinski M, Shoffner JM, Kaufman AE, Beal MF, et al. Marked increase in mitochondrial DNA deletion levels in the cerebral cortex of Huntington’s disease patients. Neurology. 1995;45(10):1879–83.
Oliveira JM, Chen S, Almeida S, Riley R, Gonçalves J, Oliveira CR, et al. Mitochondrial-dependent Ca2+ handling in Huntington’s disease striatal cells: effect of histone deacetylase inhibitors. J Neurosci. 2006;26(43):11174–86.
Arning L, Haghikia A, Taherzadeh-Fard E, Saft C, Andrich J, Pula B, et al. Mitochondrial haplogroup H correlates with ATP levels and age at onset in Huntington disease. J Mol Med JMM. 2010;88(4):431–6.
Kiernan MC, Vucic S, Cheah BC, Turner MR, Eisen A, Hardiman O, et al. Amyotrophic lateral sclerosis. Lancet. 2011;377(9769):942–55.
Al-Chalabi A, Leigh PN. Recent advances in amyotrophic lateral sclerosis. Curr Opin Neurol. 2000;13(4):397–405.
Israelson A, Arbel N, Da Cruz S, Ilieva H, Yamanaka K, Shoshan-Barmatz V, et al. Misfolded mutant SOD1 directly inhibits VDAC1 conductance in a mouse model of inherited ALS. Neuron. 2010;67(4):575–87.
Warita H, Hayashi T, Murakami T, Manabe Y, Abe K. Oxidative damage to mitochondrial DNA in spinal motoneurons of transgenic ALS mice. Brain Res Mol Brain Res. 2001;89(1–2):147–52.
Dhaliwal GK, Grewal RP. Mitochondrial DNA deletion mutation levels are elevated in ALS brains. Neuroreport. 2000;11(11):2507–9.
Wiedemann FR, Manfredi G, Mawrin C, Beal MF, Schon EA. Mitochondrial DNA and respiratory chain function in spinal cords of ALS patients. J Neurochem. 2002;80(4):616–25.
Murakami T, Nagai M, Miyazaki K, Morimoto N, Ohta Y, Kurata T, et al. Early decrease of mitochondrial DNA repair enzymes in spinal motor neurons of presymptomatic transgenic mice carrying a mutant SOD1 gene. Brain Res. 2007;1150:182–9.
Marmolino D. Friedreich’s ataxia: past, present and future. Brain Res Rev. 2011;67(1–2):311–30.
Karthikeyan G, Santos JH, Graziewicz MA, Copeland WC, Isaya G, Van Houten B, et al. Reduction in frataxin causes progressive accumulation of mitochondrial damage. Hum Mol Genet. 2003;12(24):3331–42.
Wilson RB, Roof DM. Respiratory deficiency due to loss of mitochondrial DNA in yeast lacking the frataxin homologue. Nat Genet. 1997;16(4):352–7.
Houshmand M, Panahi MSS, Nafisi S, Soltanzadeh A, Alkandari FM. Identification and sizing of GAA trinucleotide repeat expansion, investigation for D-loop variations and mitochondrial deletions in Iranian patients with Friedreich’s ataxia. Mitochondrion. 2006;6(2):82–8.
Lo Giudice T, Lombardi F, Santorelli FM, Kawarai T, Orlacchio A. Hereditary spastic paraplegia: clinical-genetic characteristics and evolving molecular mechanisms. Exp Neurol. 2014;261:518–39.
Hansen JJ, Durr A, Cournu-Rebeix I, Georgopoulos C, Ang D, Nielsen MN, et al. Hereditary spastic paraplegia SPG13 is associated with a mutation in the gene encoding the mitochondrial chaperonin Hsp60. Am J Hum Genet. 2002;70(5):1328–32.
Almontashiri NA, Chen HH, Mailloux RJ, Tatsuta T, Teng AC, Mahmoud AB, et al. SPG7 variant escapes phosphorylation-regulated processing by AFG3L2, elevates mitochondrial ROS, and is associated with multiple clinical phenotypes. Cell Rep. 2014;7(3):834–47.
Verny C, Guegen N, Desquiret V, Chevrollier A, Prundean A, Dubas F, et al. Hereditary spastic paraplegia-like disorder due to a mitochondrial ATP6 gene point mutation. Mitochondrion. 2011;11(1):70–5.
Crosby AH, Patel H, Chioza BA, Proukakis C, Gurtz K, Patton MA, et al. Defective mitochondrial mRNA maturation is associated with spastic ataxia. Am J Hum Genet. 2010;87(5):655–60.
Sanchez-Ferrero E, Coto E, Corao AI, Díaz M, Gámez J, Esteban J, et al. Mitochondrial DNA polymorphisms/haplogroups in hereditary spastic paraplegia. J Neurol. 2012;259(2):246–50.
Berer K, Krishnamoorthy G. Microbial view of central nervous system autoimmunity. Febs Lett. 2014;588(22):4207–13.
Compston A, Coles A. Multiple sclerosis. Lancet. 2008;372(9648):1502–17.
Kutzelnigg A, Lucchinetti CF, Stadelmann C, Brück W, Rauschka H, Bergmann M, et al. Cortical demyelination and diffuse white matter injury in multiple sclerosis. Brain. 2005;128:2705–12.
Ingle GT, Stevenson VL, Miller DH, Thompson AJ. Primary progressive multiple sclerosis: a 5-year clinical and MR study. Brain. 2003;126(Pt 11):2528–36.
Signoretti S, Marmarou A, Tavazzi B, Lazzarino G, Beaumont A, Vagnozzi R. N-acetylaspartate reduction as a measure of injury severity and mitochondrial dysfunction following diffuse traumatic brain injury. J Neurotrauma. 2001;18(10):977–91.
Mahad D, Campbell G, Ziabreva I, Rosenstengel C, Siegfried Schroeder HW. Mitochondrial dysfunction as a cause of axonal degeneration in the progressive stage of multiple sclerosis. J Neurol Neurosurg Psychiatry. 2008;79(3):343–4.
Dutta R, McDonough J, Yin XG, Peterson J, Chang A, Torres T, et al. Mitochondrial dysfunction as a cause of axonal degeneration in multiple sclerosis patients. Ann Neurol. 2006;59(3):478–89.
Harding AE, Sweeney MG, Miller DH, Mumford CJ, Kellar-Wood H, Menard D, et al. Occurrence of a multiple sclerosis-like illness in women who have a Leber’s hereditary optic neuropathy mitochondrial DNA mutation. Brain. 1992;115(Pt 4):979–89.
Ghezzi A, Baldini S, Zaffaroni M, Leoni G, Koudriavtseva T, Casini AR, et al. Devic’s neuromyelitis optica and mitochondrial DNA mutation: a case report. Neurol Sci. 2004;25:S380–2.
Cock H, Mandler R, Ahmed W, Schapira AH. Neuromyelitis optica (Devic’s syndrome): no association with the primary mitochondrial DNA mutations found in Leber hereditary optic neuropathy. J Neurol Neurosurg Psychiatry. 1997;62(1):85–7.
Kalman B, Mandler RN. Studies of mitochondrial DNA in Devic’s disease revealed no pathogenic mutations, but polymorphisms also found in association with multiple sclerosis. Ann Neurol. 2002;51(5):661–2.
Otaegui D, Saenz A, Martinez-Zabaleta M, Villoslada P, Fernández-Manchola I, Alvarez de Arcaya A, et al. Mitochondrial haplogroups in Basque multiple sclerosis patients. Mult Scler. 2004;10(5):532–5.
Ban M, Elson J, Walton A, Turnbull D, Compston A, Chinnery P, et al. Investigation of the role of mitochondrial DNA in multiple sclerosis susceptibility. PLoS One. 2008;3(8):e2891.
Blokhin A, Vyshkina T, Komoly S, Kalman B. Lack of mitochondrial DNA deletions in lesions of multiple sclerosis. Neuromolecular Med. 2008;10(3):187–94.
Campbell GR, Reeve AK, Ziabreva I, Reynolds R, Turnbull DM, Mahad DJ. No excess of mitochondrial DNA deletions within muscle in progressive multiple sclerosis. Mult Scler J. 2013;19(14):1858–66.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing
About this chapter
Cite this chapter
Hudson, G. (2016). The Ageing Brain, Mitochondria and Neurodegeneration. In: Reeve, A., Simcox, E., Duchen, M., Turnbull, D. (eds) Mitochondrial Dysfunction in Neurodegenerative Disorders. Springer, Cham. https://doi.org/10.1007/978-3-319-28637-2_3
Download citation
DOI: https://doi.org/10.1007/978-3-319-28637-2_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-28635-8
Online ISBN: 978-3-319-28637-2
eBook Packages: MedicineMedicine (R0)