Abstract
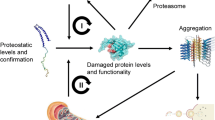
Many studies have demonstrated that the accumulation of aggregate-prone proteins due to defects in cellular quality control systems contributes to the development of neurodegenerative diseases. One form of quality control within neurons is autophagy, an intracellular pathway involved in the breakdown of cytosolic constituents. Lysosomes mediate autophagy, and their dysfunction may contribute to perturbations in cellular homeostasis and affect other organelles such as mitochondria. Mitochondrial malfunction may then further perpetuate lysosomal damage and initiate inflammatory responses. Therefore, lysosomes and mitochondria share a reciprocal relationship where dysfunction in one often affects the function of the other. These consequences of lysosome and mitochondrial impairment complete a deleterious feedback loop that concludes not only in neurodegeneration but also neuroinflammation. Herein, we discuss the primary types of autophagy and their underlying mechanisms, the regulation of lysosomal biogenesis and function, and the link between lysosomal and mitochondrial dysfunction. We conclude this chapter by assessing the role of lysosomal dysfunction in neurodegenerative diseases.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Alpha-synuclein
- TFEB
- TFE3
- Autophagy
- Lysosome
- Synucleinopathies
- Lysosomal storage disorders
- Gaucher disease
- Inflammasome
- Reactive oxygen species
- Mitophagy
- Glucocerebrosidase
1 Autophagy
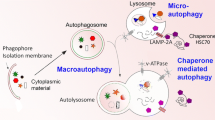
In 1963, Christian de Duve first described autophagy: a quality control system involved in the degradation of unnecessary or nonfunctional cellular components [1]. Lysosomes, with their acidic pH and lytic hydrolases, mediate autophagy in response to perturbations in cellular homeostasis that may occur as a result of organelle dysfunction, genetic mutations, or nutrient deprivation. Autophagy is particularly important in the context of neurodegeneration, as it has been shown that in the absence of any disease-associated gene products, the loss of autophagy alone is sufficient to cause neural dysfunction and eventually neuronal cell death [2, 3]. Depending on the mechanism by which cellular cargo is transported to the lysosome, autophagy can be classified into three types: microautophagy, macroautophagy, and chaperone-mediated autophagy.
1.1 Microautophagy
Little is currently known about microautophagy, especially within mammalian cells; as a result, the majority of our knowledge regarding this autophagic process stems from studies in a few species of yeast: Saccharomyces cerevisiae, Pichia pastoris, and Hansenula polymorpha [4–8]. In mammalian microautophagy, the lysosomal membrane directly invaginates and sequesters cytoplasmic constituents into vesicles intended for degradation [9–11]. This invagination may occur through either a concave retreat of the lysosomal membrane or the lysosomal wrapping mechanism (LWM) [12]. During the LWM, the lysosome elongates from a spherical to tubular shape and extends an arm-like protrusion that envelops soluble cytoplasmic components. The tip of this extension then meets and fuses with the lysosomal membrane, sealing these cellular constituents within a vesicle for breakdown inside the lysosome [13, 14]. Microautophagy, as a form of quality of control, has been implicated in the basal turnover of cellular components, but recent literature suggests that this autophagic process may possess additional functions. Microautophagy could be associated with balancing the influx of membrane components introduced by macroautophagy and may be related to multivesicular body formation as a lysosomal microautophagy-like mechanism involved in selectively delivering cytosolic proteins to late endosomes during biogenesis [11, 15–17]. As highlighted in a review article by Mijalica et al. microautophagy is a field in need of further investigation [10].
1.2 Macroautophagy
In contrast to microautophagy, macroautophagy is better understood and is described in detail in Chap. 11. In macroautophagy, cytoplasmic cargo is sequestered within double-membrane vesicles called autophagosomes for transportation to the lysosome (Fig. 12.1c) [18, 19]. Initiation of autophagy starts with the formation of a double-membrane structure exclusive to macroautophagy termed the phagophore [20–24]. Phagophore assembly is suspected to occur de novo, and within mammalian systems, this process may take place at multiple locations in the cytoplasm [25–29]. The growth of the phagophore is an area of intense debate, as the source of the membrane components used in expansion is currently unknown. Multiple studies have suggested that these building blocks arise from pre-existing membrane compartments, such as those of the mitochondria, endoplasmic reticulum (ER), Golgi apparatus, and plasma membrane. Others speculate that the membrane may originate from a phosphatidylinositol-3-phosphate-enriched portion of the ER named the omegasome [30–36]. Phagophore elongation and expansion involves sequential recruitment of several molecular complexes and subsequent delivery of membrane components to the growing phagophore (discussed in detail in Chap. 11) [37–42]. Apposition and sealing of the ends of the expanding phagophore membrane concludes autophagosome formation [39].
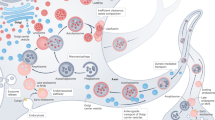
(a) TFEB is regulated at both transcriptional and posttranscriptional levels. During nutrient-rich conditions, mTORC1 phosphorylates TFEB, preventing its translocation into the nucleus and sequestering the transcription factor in the cytosol and lysosomal membrane. Under stressful conditions, mTORC1 dissociates from the lysosomal membrane without phosphorylating TFEB, allowing for its translocation to the nucleus. Within the nucleus, TFEB regulates its own transcription as well as genes involved in lipid metabolism and the CLEAR network. (b) TFEB contributes to lipid metabolism by activating transcription of PPARα and PGC-1α. These two proteins initiate pathways involved in the degradation of lipids for energy production. (c) TFEB activates CLEAR network genes involved in biogenesis of lysosomes and regulation of autophagy, which are both required for proper cellular quality control. (d) Defects in the lysosome-autophagy pathway promote accumulation of aggregate-prone proteins such as α-synuclein, huntingtin, and Tau leading to neurodegeneration
Once formed, these double-membrane vesicles may fuse with either endosomes or lysosomes resulting in the formation of chimeric organelles called amphisomes and autolysosomes, respectively [43, 44]. Amphisomes are intermediate structures that link the autophagic and endosomal pathways and are capable of further fusion with lysosomes to also produce autolysosomes [45]. Autophagosomes are dependent on microtubules for transport to both endosomes and lysosomes where their merging is facilitated by a variety of proteins including both endosomal sort complex required for transport (ESCRT) and soluble NSF attachment protein receptor (SNARE) proteins as well as Rab7 [46–48]. Of peculiar interest, proper lysosomal function and acidification were found to be important for autophagosome-lysosome and endosome-lysosome fusion as well [49]. Upon fusion of the autophagosome or amphisome with the lysosome, the inner membrane of the autolysosome is quickly degraded. The outer membrane of the autophagosome or amphisome is lost as it fuses with that of the lysosome, and cellular cargo is expelled into the acidic environment of the lysosomal lumen where it is broken down by lytic hydrolases. Lysosomal membrane permeases then release these degradation products back into the cytosol for further use in energy production or biosynthetic pathways [19, 24, 50].
1.3 Chaperone-Mediated Autophagy
Not all forms of autophagy require the formation of vesicles for transporting cargo to the lysosomal lumen. In chaperone-mediated autophagy (CMA), cellular cargo is individually targeted and directly transported across the lysosomal membrane into the lumen for degradation. This catabolic process, currently only described in mammals, was the first subtype of autophagy for which selectivity was demonstrated. Selectivity occurs through multiple steps [51–53]. The first is the recognition of substrate proteins containing a specific KFERQ pentapeptide motif by heat-shock cognate protein of 70 kDa (hsc70). All known CMA substrate proteins possess this pentapeptide motif, and several studies have demonstrated that it is both necessary and sufficient for lysosomal targeting [51, 54–57]. Recognition of protein substrates by hsc70 is regulated by the accessibility of the pentapeptide motif, as this structure may be concealed by protein folding, protein-protein interactions, and binding to subcellular membranes. Several studies have also demonstrated that posttranslational modifications of KFERQ-like motifs, which are peptide motifs that contain four out of the five required amino acid residues typically found in a KFERQ motif, regulate substrate binding as well. Since the KFERQ motif depends on its charge for proper interaction with hsc70, phosphorylation or acetylation of KFERQ-like motifs may compensate for changes in charge caused by the absence of one of the required amino acid residues typically included in a KFERQ motif. In this manner, a CMA-targeting motif may be formed upon modification of the KFERQ-like motif, allowing for association with hsc70 [52, 56, 58, 59].
Once protein substrates are bound to hsc70, this substrate/chaperone complex is delivered to the lysosomal membrane where it interacts with the cytosolic tail of lysosome-associated membrane protein type 2A (LAMP-2A), a single-span membrane protein [60]. LAMP-2A exists as a monomer at the lysosomal membrane, but, upon binding of its protein substrate, forms a multiprotein complex composed of free LAMP-2A monomers and other proteins [61]. Protein substrates may bind the monomeric form of LAMP-2A in their folded conformations; however, in order to be translocated into the lysosomal lumen, these proteins must first be unfolded. This unfolding process occurs before the assembly of the multiprotein translocation complex and may be mediated by hsc70 and its associated co-chaperones [53, 62, 63]. While numerous factors, such as the protein density and fluidity of the lysosomal membrane, influence assembly of the multiprotein complex, very few proteins have been identified as regulators of this molecular machinery [64]. Recently, work by Bandyopadhyay and colleagues have demonstrated that both GFAP and EF1α modulate the assembly and disassembly of this translocation complex in a GTP-dependent manner [65]. Furthermore, two chaperones, hsc70 and heat-shock protein 90 (hsp90), have also been implicated in this process. These chaperones function not only in the assembly and disassembly of the translocation complex but also contribute to the stabilization of LAMP-2A during multimerization [51, 61, 66].
Translocation of the unfolded protein into the lysosomal lumen occurs one by one through the LAMP-2A multiprotein complex and is dependent on the presence of an intralysosomal isoform of hsc70 (lys-hsc70) [63, 67]. This isoform of hsc70 is located in the lysosomal lumen and may enter the lysosome via fusion with hsc70-containing late endosomes [53, 66]. To date, the exact mechanism by which lys-hsc70 contributes to substrate translocation is unknown; however, it has been proposed that this protein may “pull” the substrate through the LAMP-2A translocation complex or passively “hold” the substrate, preventing its release back into the cytosol [53]. Depending on cellular conditions and cell type, the population of lysosomes containing lys-hsc70 fluctuates between 20 % and 80 %, and therefore, not all lysosomes are capable of CMA [68]. Upon entry of the protein substrate into the lysosomal lumen, substrate degradation occurs by resident hydrolases, and this process is accompanied by the dissociation of the LAMP-2A translocation complex. These resulting monomers of LAMP-2A are then free to further bind substrates and initiate new cycles of CMA [61].
2 Transcription Factor EB (TFEB)
2.1 The Lysosome
While investigating the mechanism of action of insulin within liver cells, Christian de Duve serendipitously stumbled upon a nonspecific acid phosphatase that possessed a phantasmic enzymatic activity. Tantalized by the “vanishing acts” that this acid phosphatase performed, de Duve abandoned his work on insulin and pursued this accidental finding instead. In 1955, after years of investigating this unexpected observation, de Duve described the lysosome. Roughly 20 years later, he was awarded the Nobel Prize in Physiology and Medicine for this discovery [69, 70]. Traditionally, lysosomes, Greek for “digestive body” and formed from the combination of the words lysis and soma, have been considered static cellular organelles that are not influenced by environmental cues. They are primarily implicated in the catabolism of macromolecules obtained from multiple cellular processes such as endocytosis, autophagy, and phagocytosis [10, 52, 71–74]. However, the new concept of lysosomal adaptation has subsequently broadened our perspective from merely its static role in cellular clearance [75]. Recent studies by Ballabio and colleagues have demonstrated that a majority of the 96 lysosomal genes involved in lysosomal biogenesis and function coordinately express and are influenced by environmental factors, both extracellular and intracellular, through a basic helix-loop-helix transcription factor known as transcription factor EB (TFEB), the master regulator of the coordinate lysosomal expression and regulation (CLEAR), which encompasses these lysosomal genes [76, 77]. Lysosomes have now emerged as a critical player involved in nutrient sensing, signaling, and metabolism, in addition to their established duty in cellular macromolecule degradation. At the heart of this adaptive and dynamic lysosome model is the activity of TFEB, the master regulator of lysosomal biogenesis and function that modulates the interplay between lysosome-mediated cellular processes and environmental influences.
2.2 Regulation of TFEB
TFEB is located within the cytoplasm and on the surface of the cholesterol sparse lysosomal membrane, where it is regulated through an “inside-out” signaling model initiated by the level of accumulated amino acids in the acidic lumen of the lysosome (see Fig. 12.1a). These amino acid levels are sensed by the lysosome nutrient signaling (LYNUS) machinery, which propagates a signal through a protein complex known as Ragulator to RAG GTPases that subsequently recruit mTORC1 to the lysosomal surface [78–85]. The subcellular localization of TFEB depends largely on the mTORC1-mediated phosphorylation of TFEB at two crucial serine residues: Ser142 and Ser211 [77, 84, 86, 87]. Under favorable conditions, mTORC1 phosphorylates TFEB, sequestering the transcription factor in the cytoplasm and lysosomal surface. However, during adverse cellular circumstances—starvation, stress, and lysosome dysfunction or inhibition—mTORC1 dissociates from the LYNUS complex without phosphorylating TFEB, allowing for TFEB nuclear translocation [76, 77, 81, 84, 86, 87]. Within the nucleus, TFEB activates transcription of the CLEAR network genes pertinent in the lysosomal-autophagy pathway, as well as those involved in lipid metabolism such as peroxisome proliferator-activated receptor α (PPARα), peroxisome proliferator-activated receptor-gamma coactivator-1α (PGC-1α), and their respective target genes. Furthermore, once inside the nucleus TFEB also positively regulates its own function by binding to the CLEAR motif within the promotor region of its associated gene, thus initiating its own transcription. Therefore, the activity of TFEB is regulated both at the transcriptional and posttranscriptional levels (see Fig. 12.1a), and presents a mechanism for which environmental influences, both extracellular and intracellular, can be transmitted from lysosome to nucleus [75–77].
2.3 Lipid Catabolism
TFEB is also implicated in lipid catabolism due to the intertwined features of autophagy and lipid metabolism. Through an autophagic process known as macrolipophagy, lipid droplets are transported to lysosomes via autophagosomes, where they are degraded into free fatty acids and glycerol [88, 89]. Mouse liver cells overexpressing TFEB exhibited upregulation of genes implicated in lipid catabolic processes, such as lipophagy and fatty acid oxidation, as well as downregulation of those involved in lipid biogenesis [75]. TFEB exerts its transcriptional control of lipid metabolism by inducing two key modulators of energy metabolism: PPARα and PGC-1α (see Fig. 12.1b) [75–90]. Cells stressed by starvation undergo TFEB-activated transcription of both PPARα and PGC-1α, which subsequently initiates a metabolic response where energy is produced through the breakdown of lipid reserves [75]. Furthermore, studies performed on Atg7 knockout mice, whose autophagic pathways are suppressed, demonstrate that TFEB mediates lipid metabolism through an autophagy-dependent manner [75]. Therefore, TFEB weaves together the lysosomal-autophagic pathway with lipid metabolism. It is becoming increasingly clear that the lysosome is not only a cellular garbage disposal, but it also serves as an intricate player involved in nutrient sensing, signaling, and metabolism.
2.4 TFEB, Autophagy, and Neurodegeneration
In many neurodegenerative proteopathies, the lysosomal-autophagy pathway is disrupted and the pathogenic formation of misfolded protein aggregates occurs (see Fig. 12.1c) [91–93]. Therefore, as the master regulator of the lysosomal-autophagy pathway, the role of TFEB in neurodegeneration is an area of intensive investigation. Studies utilizing a rat model of Parkinson’s disease generated by overexpressing human alpha-synuclein (α-syn) in the midbrain demonstrated that elevated α-syn levels induced TFEB retention in the cytoplasm, leading to lysosomal dysfunction, α-syn accumulation in autophagosomes, and a progressive increase in α-syn oligomers [94]. Since α-syn is structurally and functionally similar to 14-3-3 proteins, a group of proteins known to interact with and trap phosphorylated TFEB within the cytoplasm, a possible pathogenic mechanism, has been postulated where aggregates of α-syn bind to phosphorylated TFEB, preventing its nuclear translocation and eventually leading to impairment of autophagic processes [86, 87, 94–96]. Overexpression of TFEB, whether by genetic or pharmacological means, has been shown to mediate the clearance of α-syn and halt the progression of Parkinsonian neurodegeneration in both rat nigral dopaminergic neurons and human neuroglioma cells through an autophagy-dependent pathway [94, 97]. Recent research has also demonstrated that overexpression of TFEB is neuroprotective in both Huntington’s and Alzheimer’s diseases as well. In a Huntington’s disease mouse model, TFEB-mediated induction of PGC-1α was shown to rescue neurotoxicity via cellular clearance of huntingtin protein aggregation and reduction of oxidative stress [98]. Furthermore, TFEB overexpression in a mouse model of Alzheimer’s disease showed that TFEB is capable of allaying phosphorylated Tau and neurofibrillary tangle-associated neuropathology by enhancing the clearance of both hyperphosphorylated and misfolded Tau proteins [99].
2.5 Transcription Factor E3
Recently, a second member of the MiTF/TFE family was found to be a CLEAR network regulator. Like TFEB, translocation of transcription factor E3 (TFE3) from the cytosol to the nucleus is dependent upon interaction with Rag GTPases, mTORC1-dependent phosphorylation status, and nutrient availability [100]. Under conditions of starvation, TFE3 translocates from the cytosol to the nucleus where it binds to the CLEAR motif of promoters of genes belonging to the CLEAR network. Research indicates that TFE3-induced autophagy and lysosomal biogenesis are independent of TFEB. Relative protein abundance of both transcription factors, which has been shown to be different in various cell types, is speculated to be the decisive factor in taking on the role of master regulator. Just like TFEB, overexpression of TFE3 can promote lysosomal exocytosis and clearage of lysosomal substrate storage in several lysosomal storage disorders [100]. Together, these results suggest that TFEB and TFE3, both master regulators of the lysosomal-autophagy pathway, may be promising therapeutic targets for the development of a broad-spectrum neuroprotective drug.
3 Lysosomal and Mitochondrial Dysfunction: Are They Connected?
Mitochondria are eukaryotic organelles involved in a variety of cellular processes ranging from energy production to regulation of cellular calcium concentration and apoptosis. Mitochondria consist of a double-membrane structure that separates the inner-membrane space from the mitochondrial matrix [101]. Located in the inner mitochondrial membrane is the electron transport chain where oxidative phosphorylation occurs. This is also the primary site of both cellular energy and reactive oxygen species (ROS) production [102–105] (see Chap. 1). Growing evidence demonstrates that lysosomes and mitochondria share a mutual relationship, where dysfunction in one organelle often impairs the function of the other. Lysosomal damage not only directly disturbs the lysosomal-autophagy pathway but can also cause lysosomal membrane permeabilization (LMP) resulting in the release of lysosomal luminal contents into the cytosol [74, 106–108]. Whether by insufficient turnover of damaged mitochondria or via direct interaction with lysosomal cathepsins released as a result of LMP, the consequences of lysosomal dysfunction impair mitochondrial function and lead to the loss of mitochondrial membrane potential, increased ROS production, decreased generation of ATP, and eventually the discharge of mitochondrial components into the cytosol via mitochondrial membrane permeabilization [109–112]. Mitochondrial dysfunction may also occur through mutations in gene-encoding proteins involved in mitochondrial quality control and homeostasis such as PTEN-induced putative kinase 1 (PINK1/PARK6), the E3 ubiquitin ligase parkin (PARK2), and DJ-1 (PARK7). Mutations in these genes have been implicated in familial forms of Parkinson disease [101, 113–117] (see Chap. 11). While it is unclear whether lysosomal dysfunction precedes mitochondrial damage or vice versa, disruption of the intricate balance between the functions of these two organelles establishes a deleterious feedback loop [107]. A failure to degrade defective mitochondria by the lysosome results in the accumulation of dysfunctional mitochondria and the subsequent leakage of ROS. In turn, these oxidative species may perpetuate lysosome dysfunction and subsequently enhance mitochondrial stress culminating in inflammation and cell death [107, 111, 114, 118]. Therefore, aberrant quality control of both lysosomes and mitochondria has profound consequences on the pathogenesis of a multitude of diseases, especially for those distinguished by neurodegeneration [112, 119–121].
3.1 Mitophagy
Neurons are peculiarly susceptible to the subtle sequela of lysosomal and mitochondrial dysfunction due to their reduced capability for glycolysis and reliance on oxidative phosphorylation for energy production [122, 123]. In order to maintain lysosomal and mitochondrial function and homeostasis, the cell employs specialized turnover pathways that target specific organelles (see also Chap. 11). Lysosomes mediate mitochondria-specific autophagy, often referred to as mitophagy, within cells, and this catabolic mechanism is regulated by two key proteins: PINK1 and parkin. In viable mitochondria, PINK1 is continually expressed and recruited to the outer mitochondrial membrane (OMM) where this serine/threonine kinase is imported into the mitochondrial matrix in a mitochondrial membrane potential-dependent manner. Once in the mitochondrial matrix, PINK1 is immediately degraded by proteases, thus regulating its expression [101, 114, 116]. However, upon the accumulation of aberrant proteins or the loss of mitochondrial membrane potential following mitochondrial insult, PINK1 regulation is impaired, allowing for accumulation of PINK1 on the OMM and its activation via autophosphorylation [124, 125]. Activated PINK1 recruits parkin to the OMM and phosphorylates not only the ubiquitin-like domain of parkin but also ubiquitin itself. Phosphorylation of both protein species is required for full activation of the E3 ubiquitin ligase function of parkin [126–128]. Once relegated to the mitochondrial surface, parkin ubiquitinates various OMM proteins involved in mitochondrial maintenance, and this polyubiquitination labels the damaged mitochondria for turnover [129–131]. Cells with defective lysosomes, and therefore impaired mitophagy, are unable to effectively breakdown defective mitochondria resulting in their accumulation. These damaged mitochondria may leak reactive oxygen species (ROS), such as superoxide (O2 −) and hydrogen peroxide (H2O2), from complex I and III of the respiratory chain. If left unchecked, these oxidative species may ultimately cause neuronal death by not only further perpetuating organelle damage but also by activating the NLRP3 inflammasome and therefore initiating inflammatory responses [111, 112].
3.2 Oxidative Stress
ROS are important mediators of the downstream consequences stemming from the combined effects of lysosomal and mitochondrial dysfunction. The mitochondrial respiratory chain is the principal producer of ROS, primarily in the forms of O2 − and H2O2, within the cell and during abnormal lysosomal and mitochondrial function, and contributes to the elevation of oxidative stress [112]. Lysosomes are particularly vulnerable to ROS-induced damage, as these oxidative species peroxidize lysosomal membrane lipids resulting in destabilization of the membrane and, potentially, even LMP. Due to their ability to inherit cargo during fusion with autophagosomes, lysosomes may acquire large amounts of iron during the degradation of macromolecules. This accumulation of iron within lysosomes has been speculated to contribute to lysosomal susceptibility to oxidative damage [107, 132]. Simultaneously, elevated oxidative stress may damage mitochondria as well. Reactive oxygen species may peroxidize lipids in the mitochondrial membrane resulting in loss of membrane potential as well as fragmented mitochondrial morphology [112, 116, 133]. Under high levels of oxidative stress, mitochondrial membrane permeabilization may also occur and release cytochrome c into the cytosol, therefore activating apoptosis [134]. Mitochondrial DNA (mtDNA) neighbors the electron transport chain within the inner mitochondrial membrane and is prone to oxidation and release during mitochondrial damage. Once released into the cytosol, oxidized mtDNA and local ROS trigger assembly and activation of the NLRP3 inflammasome [111, 134–136]. Beyond these organelles, ROS may also regulate the activity of proteins through posttranslational modification. One example is DJ-1, a neuroprotective protein encoded by PARK7 that has been implicated in familial forms of Parkinson’s disease when mutated. DJ-1 is dependent on localization to the mitochondria for proper neuroprotective activity, and this translocation is redox regulated by oxidation of the Cys106 residue into cysteine sulfinic acid by ROS. After translocation to the mitochondria, DJ-1 protects cells from oxidative stress-induced death by modulating mitophagy and, together with other cellular processes, assumes a role in antioxidant response [137, 138].
3.3 Inflammasome Activation
Inflammasomes are intracellular multiprotein complexes that initiate an inflammatory response in response to pathogens and intracellular insults. The nucleotide-binding oligomerization domain-like receptor family, pyrin domain containing 3 (NLRP3) inflammasome is the best characterized and is closely associated with lysosomal and mitochondrial dysfunction [139]. This protein complex is formed at the interface of the mitochondria and endoplasmic reticulum in an area known as the mitochondria-associated endoplasmic reticulum membrane. The NLRP3 inflammasome is chiefly composed of three components: a NOD-like receptor (NLRP3), the adaptor protein ASC, and caspase-1. Once assembled and activated, the NLRP3 inflammasome cleaves proIL-1β and proIL-18 into their bioactive forms, where these proinflammatory messenger molecules may subsequently modulate immune and inflammatory pathways [110, 111, 139–141]. Growing evidence identifies both ROS and oxidized mtDNA as activators of the NLRP3 inflammasome. Cells with impaired mitophagy, and consequently prolonged clearance of defective mitochondria, may spontaneously secrete ROS and oxidized mtDNA into the cytosol, resulting in consistent activation of the NLRP3 inflammasome [12, 36]. Recently, elegant work by Shi and colleagues has demonstrated that autophagy may function as a negative regulator of inflammasome activation. Their data suggests that inflammasome activation concomitantly induces autophagosome formation by initiating nucleotide exchange on the G protein RalB. Inflammasomes subsequently undergo ubiquitination and are transported by adaptor proteins p62 and LC3 to autophagosomes for elimination. These results suggest that autophagy may modulate the intensity of inflammation by directly degrading active inflammasomes and therefore may result in uncontrolled inflammation during lysosomal and mitochondrial dysfunction [142]. Taken together, the inflammasome represents a link between lysosomal and mitochondrial dysfunction and inflammation, which contributes to the pathogenesis of not only neurodegenerative but also autoinflammatory diseases [143, 144].
4 Lysosomal Storage Disorders and Neurodegeneration
Lysosomal storage disorders (LSDs) are rare inborn metabolic diseases in which lysosomal function is severely compromised due to mutations in gene-encoding enzymes resident in lysosomes involved in the breakdown of specific substrates. The subsequent accumulation of substrate within lysosomes has a variety of consequences such as lysosomal enlargement, altered lysosomal pH, and diminished activity of lysosomal enzymes. Over 50 different LSDs have been described, and mutations in LSD-associated genes in patients, as well as carriers, have been linked to neurodegeneration, more particularly synucleinopathies [145]. Among LSD-associated genes, the molecular link between mutations in the glucocerebrosidase gene (GBA1) and Parkinson’s disease (PD) is the most established [146].
4.1 GBA1 and Synucleinopathies
Pathological mutations in both alleles of the GBA1 gene cause Gaucher disease (GD), the most common LSD. This disorder is characterized by lysosomal accumulation of the substrate glucosylceramide (GC), due to a deficiency in the lysosome-resident glucocerebrosidase enzyme (GCase) [147]. The cells most affected in GD patients are macrophages, which are involved in breakdown of senescent cells with GC-rich membranes such as erythrocytes. “Gaucher cells,” which are the macrophages that have lysosomes engorged with substrate, can infiltrate the spleen, liver, and bone marrow, resulting in inflammation and organomegaly [148]. GD has been historically classified into non-neuronopathic type 1, acute neuronopathic type 2, and chronic neuronopathic type 3. Today, clinicians acknowledge a broad range of clinical manifestations associated with GD and subsequently can have difficulty classifying patients into specific GD subtypes [147]. Over the last 6 years, large cohort studies have established that the presence of mutations in the GBA1 gene is a risk factor for the development of synucleinopathies including PD [146], dementia with Lewy bodies (DLB) (Fig. 12.2) [149], and, most recently, multiple system atrophy (MSA) [150]. All three synucleinopathies are characterized by the presence of inclusions of aggregated α-syn, a 14 kDa protein that is speculated to be involved in the regulation of synaptic vesicle dynamics and neurotransmitter release [151, 152]. In PD and DLB, the α-syn-positive Lewy bodies and neurites are mainly located in neurons of the substantia nigra, cerebral cortex, and hippocampus, while in MSA, the α-syn inclusions are located in glial oligodendrocytes [146, 149, 150]. The molecular link between mutations in the GBA1 gene and PD was established by molecular analyses of the GBA1 gene on a large pan-ethnic cohort comprising 5,691 patients with PD and 4,898 controls. This study revealed a strong association between GBA1 mutations and the development of PD with an odds ratio of 5.43 and earlier onset of PD symptoms in patients with GBA1 mutations [146]. These results have been replicated in multiple large cohorts with different ethnic backgrounds [114, 153, 154]. Today, GBA1 mutations are widely considered the most common genetic risk factor for PD. However, it is important to keep in mind that most patients with GD and mutant GBA1 carriers never develop synucleinopathies. These observations suggest that GBA1 mutations and subsequent dysfunctional GCase enzyme are not a direct cause of synucleinopathy development; other cellular processes affecting organelle homeostasis, such as ER-stress and lysosomal and mitochondrial function, might play a more central role in synucleinopathy pathogenesis. The presence of dysfunctional GCase could exacerbate organelle dysfunction and subsequent α-syn accumulation.
4.2 GCase and α-syn Homeostasis
Initially, the mechanistic link between dysfunctional GCase enzyme and α-syn aggregation focused on gain-of-function or loss-of function hypotheses, where the former supports the direct involvement of dysfunctional GCase enzyme in the aggregation of α-syn, and the latter supports the role of lysosomal GC substrate accumulation in α-syn aggregation [155]. Currently, in vitro and in vivo research supports a reciprocal relationship between GCase and α-syn where downregulation of GCase protein expression or enzyme activity results in accumulation of α-syn. Increases in α-syn protein expression results in reduced GCase protein expression and enzyme activity (reviewed by [114, 156]). Furthermore, three independent studies support the observation of reduced GCase activity and protein expression in postmortem brains of sporadic PD and DLB patients without GBA1 mutations, reinforcing the reciprocal relationship in relevant human samples [157–159]. The molecular mechanism of the reciprocal relationship is not fully understood although there is some evidence that an increase in α-syn protein levels inhibits ER-to-Golgi trafficking of GCase, which subsequently results in downregulation of GCase translocation to lysosomes. Less GCase in lysosomes can lead to lysosomal GC substrate accumulation and subsequent lysosomal dysfunction, which in turn may stimulate accumulation and oligomerization of α-syn throughout the cell. Buildup of α-syn aggregates could, in turn, inhibit ER-to-Golgi trafficking of GCase resulting in further decrease of this enzyme within lysosomes [160]. This reciprocal positive feedback loop could eventually lead to neurodegeneration. Evidence for this hypothesis came from a neuronopathic GD type 2 mouse model lacking GCase. Here, autophagy and proteosomal impairment lead to accumulation of fragmented mitochondria and α-syn in cultured neurons and astrocytes of the midbrain [161]. Although this gba -/- model is not reflective of PD, it suggested that the lack of GCase expression promotes α-syn accumulation through impairment of cellular turnover pathways [101]. Novel insights into maintenance of α-syn homeostasis by manipulating GCase enzyme levels are promising for the development of new treatments for synucleinopathies. Although GCase enzyme replacement therapy does not improve PD symptoms, as the recombinant enzyme does not cross the blood-brain barrier [162], molecular inhibitors of glucosylceramide synthase for GC substrate reduction therapy and molecular chaperones for enhancing GCase translocation to the lysosomes can cross the blood-brain barrier and therefore show potential as therapeutics [163–166]. Recent research indicates associations similar to that found between mutations in GBA1, and the development of synucleinopathies can be expanded to other LSD-associated genes. Large molecular cohort studies suggest that mutations in the sphingomyelin phosphodiesterase (SMPD1) and α-N-acetylglucosaminidase (NAGLU) genes, which are associated with Niemann-Pick disease A and B and mucopolysaccharidosis type III B, respectively, may be implicated in the development PD [167, 168]. These observations suggest that mutations in other lysosomal-resident enzymes might be classified as risk factors for the development of synucleinopathies.
5 Conclusion
When first described by Christian de Duve, and for many years after, lysosomes were often considered static organelles primarily involved in the degradation of cellular constituents. However, recent insights into lysosomal function and regulation have demonstrated otherwise. In fact, lysosomes are now considered dynamic organelles capable of not only cellular cleanup but also nutrient sensing and lipid catabolism. As mediators of autophagy, lysosomes also play an important role in the development of neurodegenerative diseases. Lysosomal dysfunction leads to not only the accumulation of aggregate-prone proteins but also impairs other organelles such as mitochondria. Together, dysfunction of this deleterious duo might drive a destructive feedback loop that culminates in the neuropathology often found in Parkinson’s, Alzheimer’s, and Huntington’s diseases. The association between LSDs and neurodegenerative diseases such as PD, LBD, and MSA further highlight the importance of proper lysosomal function in neuronal health. Further investigations exploring the relationship between lysosomal and mitochondrial dysfunction hold promise for the discovery of new potential drug targets.
References
De Duve C. The lysosome. Sci Am. 1963;208:64–72.
Hara T, Nakamura K, Matsui M, Yamamoto A, Nakahara Y, Suzuki-Migishima R, et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature. 2006;441(7095):885–9.
Komatsu M, Waguri S, Chiba T, Murata S, Iwata J, Tanida I, et al. Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature. 2006;441(7095):880–4.
Sakai Y, Koller A, Rangell LK, Keller GA, Subramani S. Peroxisome degradation by microautophagy in Pichia pastoris: identification of specific steps and morphological intermediates. J Cell Biol. 1998;141(3):625–36.
Kiel JA, Komduur JA, van der Klei IJ, Veenhuis M. Macropexophagy in Hansenula polymorpha: facts and views. FEBS Lett. 2003;549(1–3):1–6.
Roberts P, Moshitch-Moshkovitz S, Kvam E, O’Toole E, Winey M, Goldfarb DS. Piecemeal microautophagy of nucleus in Saccharomyces cerevisiae. Mol Biol Cell. 2003;14(1):129–41.
Dunn Jr WA, Cregg JM, Kiel JA, van der Klei IJ, Oku M, Sakai Y, et al. Pexophagy: the selective autophagy of peroxisomes. Autophagy. 2005;1(2):75–83.
Uttenweiler A, Mayer A. Microautophagy in the yeast Saccharomyces cerevisiae. Methods Mol Biol. 2008;445:245–59.
Ahlberg J, Marzella L, Glaumann H. Uptake and degradation of proteins by isolated rat liver lysosomes. Suggestion of a microautophagic pathway of proteolysis. Lab Invest. 1982;47(6):523–32.
Mijaljica D, Prescott M, Devenish RJ. Microautophagy in mammalian cells: revisiting a 40-year-old conundrum. Autophagy. 2011;7(7):673–82.
Sahu R, Kaushik S, Clement CC, Cannizzo ES, Scharf B, Follenzi A, et al. Microautophagy of cytosolic proteins by late endosomes. Dev Cell. 2011;20(1):131–9.
Marzella L, Ahlberg J, Glaumann H. In vitro uptake of particles by lysosomes. Exp Cell Res. 1980;129(2):460–6.
Sakai M, Ogawa K. Energy-dependent lysosomal wrapping mechanism (LWM) during autophagolysosome formation. Histochemistry. 1982;76(4):479–88.
Sakai M, Araki N, Ogawa K. Lysosomal movements during heterophagy and autophagy: with special reference to nematolysosome and wrapping lysosome. J Electron Microsc Tech. 1989;12(2):101–31.
de Waal EJ, Vreeling-Sindelarova H, Schellens JP, Houtkooper JM, James J. Quantitative changes in the lysosomal vacuolar system of rat hepatocytes during short-term starvation. A morphometric analysis with special reference to macro- and microautophagy. Cell Tissue Res. 1986;243(3):641–8.
Mortimore GE, Lardeux BR, Adams CE. Regulation of microautophagy and basal protein turnover in rat liver. Effects of short-term starvation. J Biol Chem. 1988;263(5):2506–12.
Muller O, Sattler T, Flotenmeyer M, Schwarz H, Plattner H, Mayer A. Autophagic tubes: vacuolar invaginations involved in lateral membrane sorting and inverse vesicle budding. J Cell Biol. 2000;151(3):519–28.
Yorimitsu T, Klionsky DJ. Autophagy: molecular machinery for self-eating. Cell Death Differ. 2005;12 Suppl 2:1542–52.
Yang Z, Klionsky DJ. Mammalian autophagy: core molecular machinery and signaling regulation. Curr Opin Cell Biol. 2010;22(2):124–31.
Ganley IG, Lam du H, Wang J, Ding X, Chen S, Jiang X. ULK1.ATG13.FIP200 complex mediates mTOR signaling and is essential for autophagy. J Biol Chem. 2009;284(18):12297–305.
Hosokawa N, Sasaki T, Iemura S, Natsume T, Hara T, Mizushima N. Atg101, a novel mammalian autophagy protein interacting with Atg13. Autophagy. 2009;5(7):973–9.
Jung CH, Jun CB, Ro SH, Kim YM, Otto NM, Cao J, et al. ULK-Atg13-FIP200 complexes mediate mTOR signaling to the autophagy machinery. Mol Biol Cell. 2009;20(7):1992–2003.
Mercer CA, Kaliappan A, Dennis PB. A novel, human Atg13 binding protein, Atg101, interacts with ULK1 and is essential for macroautophagy. Autophagy. 2009;5(5):649–62.
Parzych KR, Klionsky DJ. An overview of autophagy: morphology, mechanism, and regulation. Antioxid Redox Signal. 2014;20(3):460–73.
Noda T, Suzuki K, Ohsumi Y. Yeast autophagosomes: de novo formation of a membrane structure. Trends Cell Biol. 2002;12(5):231–5.
Kovacs AL, Palfia Z, Rez G, Vellai T, Kovacs J. Sequestration revisited: integrating traditional electron microscopy, de novo assembly and new results. Autophagy. 2007;3(6):655–62.
Itakura E, Mizushima N. Characterization of autophagosome formation site by a hierarchical analysis of mammalian Atg proteins. Autophagy. 2010;6(6):764–76.
Chen Y, Klionsky DJ. The regulation of autophagy – unanswered questions. J Cell Sci. 2011;124(Pt 2):161–70.
Feng Y, He D, Yao Z, Klionsky DJ. The machinery of macroautophagy. Cell Res. 2014;24(1):24–41.
Burman C, Ktistakis NT. Regulation of autophagy by phosphatidylinositol 3-phosphate. FEBS Lett. 2010;584(7):1302–12.
Hayashi-Nishino M, Fujita N, Noda T, Yamaguchi A, Yoshimori T, Yamamoto A. A subdomain of the endoplasmic reticulum forms a cradle for autophagosome formation. Nat Cell Biol. 2009;11(12):1433–7.
Yla-Anttila P, Vihinen H, Jokitalo E, Eskelinen EL. 3D tomography reveals connections between the phagophore and endoplasmic reticulum. Autophagy. 2009;5(8):1180–5.
Hailey DW, Rambold AS, Satpute-Krishnan P, Mitra K, Sougrat R, Kim PK, et al. Mitochondria supply membranes for autophagosome biogenesis during starvation. Cell. 2010;141(4):656–67.
Ravikumar B, Moreau K, Jahreiss L, Puri C, Rubinsztein DC. Plasma membrane contributes to the formation of pre-autophagosomal structures. Nat Cell Biol. 2010;12(8):747–57.
Ravikumar B, Moreau K, Rubinsztein DC. Plasma membrane helps autophagosomes grow. Autophagy. 2010;6(8):1184–6.
Takahashi Y, Meyerkord CL, Hori T, Runkle K, Fox TE, Kester M, et al. Bif-1 regulates Atg9 trafficking by mediating the fission of Golgi membranes during autophagy. Autophagy. 2011;7(1):61–73.
Weidberg H, Shvets E, Elazar Z. Biogenesis and cargo selectivity of autophagosomes. Ann Rev Biochem. 2011;80:125–56.
Hanada T, Noda NN, Satomi Y, Ichimura Y, Fujioka Y, Takao T, et al. The Atg12-Atg5 conjugate has a novel E3-like activity for protein lipidation in autophagy. J Biol Chem. 2007;282(52):37298–302.
Weidberg H, Shvets E, Shpilka T, Shimron F, Shinder V, Elazar Z. LC3 and GATE-16/GABARAP subfamilies are both essential yet act differently in autophagosome biogenesis. EMBO J. 2010;29(11):1792–802.
Bjorkoy G, Lamark T, Brech A, Outzen H, Perander M, Overvatn A, et al. p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J Cell Biol. 2005;171(4):603–14.
Kirkin V, Lamark T, Sou YS, Bjorkoy G, Nunn JL, Bruun JA, et al. A role for NBR1 in autophagosomal degradation of ubiquitinated substrates. Mol Cell. 2009;33(4):505–16.
Young AR, Chan EY, Hu XW, Kochl R, Crawshaw SG, High S, et al. Starvation and ULK1-dependent cycling of mammalian Atg9 between the TGN and endosomes. J Cell Sci. 2006;119(Pt 18):3888–900.
Gordon PB, Seglen PO. Prelysosomal convergence of autophagic and endocytic pathways. Biochem Biophys Res Commun. 1988;151(1):40–7.
Yang Z, Klionsky DJ. Eaten alive: a history of macroautophagy. Nat Cell Biol. 2010;12(9):814–22.
Fader CM, Colombo MI. Autophagy and multivesicular bodies: two closely related partners. Cell Death Differ. 2009;16(1):70–8.
Kochl R, Hu XW, Chan EY, Tooze SA. Microtubules facilitate autophagosome formation and fusion of autophagosomes with endosomes. Traffic. 2006;7(2):129–45.
Monastyrska I, Rieter E, Klionsky DJ, Reggiori F. Multiple roles of the cytoskeleton in autophagy. Biol Rev Camb Philos Soc. 2009;84(3):431–48.
Metcalf DJ, Garcia-Arencibia M, Hochfeld WE, Rubinsztein DC. Autophagy and misfolded proteins in neurodegeneration. Exp Neurol. 2012;238(1):22–8.
Lu Y, Dong S, Hao B, Li C, Zhu K, Guo W, et al. Vacuolin-1 potently and reversibly inhibits autophagosome-lysosome fusion by activating RAB5A. Autophagy. 2014;10(11):1895–905.
Feng Y, Yao Z, Klionsky DJ. How to control self-digestion: transcriptional, post-transcriptional, and post-translational regulation of autophagy. Trends Cell Biol. 2015;25(6):354–63.
Kaushik S, Cuervo AM. Chaperones in autophagy. Pharmacol Res. 2012;66(6):484–93.
Kaushik S, Cuervo AM. Chaperone-mediated autophagy: a unique way to enter the lysosome world. Trends Cell Biol. 2012;22(8):407–17.
Cuervo AM, Wong E. Chaperone-mediated autophagy: roles in disease and aging. Cell Res. 2014;24(1):92–104.
Dice JF, Chiang HL, Spencer EP, Backer JM. Regulation of catabolism of microinjected ribonuclease A. Identification of residues 7–11 as the essential pentapeptide. J Biol Chem. 1986;261(15):6853–9.
Chiang HL, Terlecky SR, Plant CP, Dice JF. A role for a 70-kilodalton heat shock protein in lysosomal degradation of intracellular proteins. Science. 1989;246(4928):382–5.
Dice JF. Peptide sequences that target cytosolic proteins for lysosomal proteolysis. Trends Biochem Sci. 1990;15(8):305–9.
Koga H, Martinez-Vicente M, Macian F, Verkhusha VV, Cuervo AM. A photoconvertible fluorescent reporter to track chaperone-mediated autophagy. Nat Commun. 2011;2:386.
Thompson LM, Aiken CT, Kaltenbach LS, Agrawal N, Illes K, Khoshnan A, et al. IKK phosphorylates Huntingtin and targets it for degradation by the proteasome and lysosome. J Cell Biol. 2009;187(7):1083–99.
Lv L, Li D, Zhao D, Lin R, Chu Y, Zhang H, et al. Acetylation targets the M2 isoform of pyruvate kinase for degradation through chaperone-mediated autophagy and promotes tumor growth. Mol Cell. 2011;42(6):719–30.
Cuervo AM, Dice JF. A receptor for the selective uptake and degradation of proteins by lysosomes. Science. 1996;273(5274):501–3.
Bandyopadhyay U, Kaushik S, Varticovski L, Cuervo AM. The chaperone-mediated autophagy receptor organizes in dynamic protein complexes at the lysosomal membrane. Mol Cell Biol. 2008;28(18):5747–63.
Salvador N, Aguado C, Horst M, Knecht E. Import of a cytosolic protein into lysosomes by chaperone-mediated autophagy depends on its folding state. J Biol Chem. 2000;275(35):27447–56.
Agarraberes FA, Dice JF. A molecular chaperone complex at the lysosomal membrane is required for protein translocation. J Cell Sci. 2001;114(Pt 13):2491–9.
Kaushik S, Massey AC, Cuervo AM. Lysosome membrane lipid microdomains: novel regulators of chaperone-mediated autophagy. EMBO J. 2006;25(17):3921–33.
Bandyopadhyay U, Sridhar S, Kaushik S, Kiffin R, Cuervo AM. Identification of regulators of chaperone-mediated autophagy. Mol Cell. 2010;39(4):535–47.
Xilouri M, Stefanis L. Chaperone mediated autophagy to the rescue: a new-fangled target for the treatment of neurodegenerative diseases. Mol Cell Neurosci. 2015;66(Pt A):29–36.
Agarraberes FA, Terlecky SR, Dice JF. An intralysosomal hsp70 is required for a selective pathway of lysosomal protein degradation. J Cell Biol. 1997;137(4):825–34.
Cuervo AM, Dice JF, Knecht E. A population of rat liver lysosomes responsible for the selective uptake and degradation of cytosolic proteins. J Biol Chem. 1997;272(9):5606–15.
de Duve C. The lysosome turns fifty. Nat Cell Biol. 2005;7(9):847–9.
De Duve C, Pressman BC, Gianetto R, Wattiaux R, Appelmans F. Tissue fractionation studies. 6. Intracellular distribution patterns of enzymes in rat-liver tissue. Biochem J. 1955;60(4):604–17.
Luzio JP, Parkinson MD, Gray SR, Bright NA. The delivery of endocytosed cargo to lysosomes. Biochem Soc Trans. 2009;37(Pt 5):1019–21.
Mizushima N, Levine B, Cuervo AM, Klionsky DJ. Autophagy fights disease through cellular self-digestion. Nature. 2008;451(7182):1069–75.
Saftig P. Lysosomes. Georgetown/New York: Landes Bioscience/Eurekah.com/Springer Science+Business Media; 2005;1–17.
Settembre C, Ballabio A. Lysosomal adaptation: how the lysosome responds to external cues. Cold Spring Harb Perspect Biol. 2014;6(6):1–15.
Settembre C, De Cegli R, Mansueto G, Saha PK, Vetrini F, Visvikis O, et al. TFEB controls cellular lipid metabolism through a starvation-induced autoregulatory loop. Nat Cell Biol. 2013;15(6):647–58.
Sardiello M, Palmieri M, di Ronza A, Medina DL, Valenza M, Gennarino VA, et al. A gene network regulating lysosomal biogenesis and function. Science. 2009;325(5939):473–7.
Settembre C, Di Malta C, Polito VA, Garcia Arencibia M, Vetrini F, Erdin S, et al. TFEB links autophagy to lysosomal biogenesis. Science. 2011;332(6036):1429–33.
Bar-Peled L, Schweitzer LD, Zoncu R, Sabatini DM. Ragulator is a GEF for the rag GTPases that signal amino acid levels to mTORC1. Cell. 2012;150(6):1196–208.
Sancak Y, Bar-Peled L, Zoncu R, Markhard AL, Nada S, Sabatini DM. Ragulator-Rag complex targets mTORC1 to the lysosomal surface and is necessary for its activation by amino acids. Cell. 2010;141(2):290–303.
Sancak Y, Peterson TR, Shaul YD, Lindquist RA, Thoreen CC, Bar-Peled L, et al. The Rag GTPases bind raptor and mediate amino acid signaling to mTORC1. Science. 2008;320(5882):1496–501.
Schulze H, Kolter T, Sandhoff K. Principles of lysosomal membrane degradation: cellular topology and biochemistry of lysosomal lipid degradation. Biochim Biophys Acta. 2009;1793(4):674–83.
Settembre C, Fraldi A, Medina DL, Ballabio A. Signals from the lysosome: a control centre for cellular clearance and energy metabolism. Nat Rev Mol Cell Biol. 2013;14(5):283–96.
Zoncu R, Bar-Peled L, Efeyan A, Wang S, Sancak Y, Sabatini DM. mTORC1 senses lysosomal amino acids through an inside-out mechanism that requires the vacuolar H(+)-ATPase. Science. 2011;334(6056):678–83.
Settembre C, Zoncu R, Medina DL, Vetrini F, Erdin S, Erdin S, et al. A lysosome-to-nucleus signalling mechanism senses and regulates the lysosome via mTOR and TFEB. EMBO J. 2012;31(5):1095–108.
Martina JA, Puertollano R. Rag GTPases mediate amino acid-dependent recruitment of TFEB and MITF to lysosomes. J Cell Biol. 2013;200(4):475–91.
Martina JA, Chen Y, Gucek M, Puertollano R. MTORC1 functions as a transcriptional regulator of autophagy by preventing nuclear transport of TFEB. Autophagy. 2012;8(6):903–14.
Roczniak-Ferguson A, Petit CS, Froehlich F, Qian S, Ky J, Angarola B, et al. The transcription factor TFEB links mTORC1 signaling to transcriptional control of lysosome homeostasis. Sci Signal. 2012;5(228):ra42.
Singh R, Cuervo AM. Autophagy in the cellular energetic balance. Cell Metab. 2011;13(5):495–504.
Singh R, Kaushik S, Wang Y, Xiang Y, Novak I, Komatsu M, et al. Autophagy regulates lipid metabolism. Nature. 2009;458(7242):1131–5.
Finck BN, Kelly DP. PGC-1 coactivators: inducible regulators of energy metabolism in health and disease. J Clin Invest. 2006;116(3):615–22.
Harris H, Rubinsztein DC. Control of autophagy as a therapy for neurodegenerative disease. Nat Rev Neurol. 2012;8(2):108–17.
Wong E, Cuervo AM. Autophagy gone awry in neurodegenerative diseases. Nat Neurosci. 2010;13(7):805–11.
Taylor JP, Hardy J, Fischbeck KH. Toxic proteins in neurodegenerative disease. Science. 2002;296(5575):1991–5.
Decressac M, Mattsson B, Weikop P, Lundblad M, Jakobsson J, Bjorklund A. TFEB-mediated autophagy rescues midbrain dopamine neurons from alpha-synuclein toxicity. Proc Natl Acad Sci U S A. 2013;110(19):E1817–26.
Ostrerova N, Petrucelli L, Farrer M, Mehta N, Choi P, Hardy J, et al. alpha-Synuclein shares physical and functional homology with 14-3-3 proteins. J Neurosci. 1999;19(14):5782–91.
Perez RG, Waymire JC, Lin E, Liu JJ, Guo F, Zigmond MJ. A role for alpha-synuclein in the regulation of dopamine biosynthesis. J Neurosci. 2002;22(8):3090–9.
Kilpatrick K, Zeng Y, Hancock T, Segatori L. Genetic and chemical activation of TFEB mediates clearance of aggregated alpha-synuclein. PLoS ONE. 2015;10(3), e0120819.
Tsunemi T, Ashe TD, Morrison BE, Soriano KR, Au J, Roque RA, et al. PGC-1alpha rescues Huntington’s disease proteotoxicity by preventing oxidative stress and promoting TFEB function. Sci Transl Med. 2012;4(142):142ra97.
Polito VA, Li H, Martini-Stoica H, Wang B, Yang L, Xu Y, et al. Selective clearance of aberrant tau proteins and rescue of neurotoxicity by transcription factor EB. EMBO Mol Med. 2014;6(9):1142–60.
Martina JA, Diab HI, Lishu L, Jeong AL, Patange S, Raben N, et al. The nutrient-responsive transcription factor TFE3 promotes autophagy, lysosomal biogenesis, and clearance of cellular debris. Sci Signal. 2014;7(309):ra9.
Osellame LD, Duchen MR. Quality control gone wrong: mitochondria, lysosomal storage disorders and neurodegeneration. Br J Pharmacol. 2014;171(8):1958–72.
Chance B, Sies H, Boveris A. Hydroperoxide metabolism in mammalian organs. Physiol Rev. 1979;59(3):527–605.
Mitchell P. Coupling of phosphorylation to electron and hydrogen transfer by a chemi-osmotic type of mechanism. Nature. 1961;191:144–8.
Mitchell P, Moyle J. Chemiosmotic hypothesis of oxidative phosphorylation. Nature. 1967;213(5072):137–9.
Liu Y, Fiskum G, Schubert D. Generation of reactive oxygen species by the mitochondrial electron transport chain. J Neurochem. 2002;80(5):780–7.
Appelqvist H, Waster P, Kagedal K, Ollinger K. The lysosome: from waste bag to potential therapeutic target. J Mol Cell Biol. 2013;5(4):214–26.
Repnik U, Hafner Cesen M, Turk B. Lysosomal membrane permeabilization in cell death: concepts and challenges. Mitochondrion. 2014;19(Pt A):49–57.
Ballabio A, Gieselmann V. Lysosomal disorders: from storage to cellular damage. Biochim Biophys Acta. 2009;1793(4):684–96.
Boya P, Andreau K, Poncet D, Zamzami N, Perfettini JL, Metivier D, et al. Lysosomal membrane permeabilization induces cell death in a mitochondrion-dependent fashion. J Exp Med. 2003;197(10):1323–34.
Heid ME, Keyel PA, Kamga C, Shiva S, Watkins SC, Salter RD. Mitochondrial reactive oxygen species induces NLRP3-dependent lysosomal damage and inflammasome activation. J Immunol. 2013;191(10):5230–8.
Zhou R, Yazdi AS, Menu P, Tschopp J. A role for mitochondria in NLRP3 inflammasome activation. Nature. 2011;469(7329):221–5.
Lee J, Giordano S, Zhang J. Autophagy, mitochondria and oxidative stress: cross-talk and redox signalling. Biochem J. 2012;441(2):523–40.
Puschmann A. Monogenic Parkinson’s disease and parkinsonism: clinical phenotypes and frequencies of known mutations. Parkinsonism Relat Disord. 2013;19(4):407–15.
Siebert M, Sidransky E, Westbroek W. Glucocerebrosidase is shaking up the synucleinopathies. Brain. 2014;137(Pt 5):1304–22.
Kitada T, Asakawa S, Hattori N, Matsumine H, Yamamura Y, Minoshima S, et al. Mutations in the parkin gene cause autosomal recessive juvenile parkinsonism. Nature. 1998;392(6676):605–8.
Ryan BJ, Hoek S, Fon EA, Wade-Martins R. Mitochondrial dysfunction and mitophagy in Parkinson’s: from familial to sporadic disease. Trends Biochem Sci. 2015;40(4):200–10.
Valente EM, Abou-Sleiman PM, Caputo V, Muqit MM, Harvey K, Gispert S, et al. Hereditary early-onset Parkinson’s disease caused by mutations in PINK1. Science. 2004;304(5674):1158–60.
van der Burgh R, Boes M. Mitochondria in autoinflammation: cause, mediator or bystander? Trends Endocrinol Metab. 2015;26(5):263–71.
Boland B, Kumar A, Lee S, Platt FM, Wegiel J, Yu WH, et al. Autophagy induction and autophagosome clearance in neurons: relationship to autophagic pathology in Alzheimer’s disease. J Neurosci. 2008;28(27):6926–37.
Narendra D, Tanaka A, Suen DF, Youle RJ. Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J Cell Biol. 2008;183(5):795–803.
Settembre C, Fraldi A, Jahreiss L, Spampanato C, Venturi C, Medina D, et al. A block of autophagy in lysosomal storage disorders. Hum Mol Genet. 2008;17(1):119–29.
Bolanos JP, Almeida A, Moncada S. Glycolysis: a bioenergetic or a survival pathway? Trends Biochem Sci. 2010;35(3):145–9.
Herrero-Mendez A, Almeida A, Fernandez E, Maestre C, Moncada S, Bolanos JP. The bioenergetic and antioxidant status of neurons is controlled by continuous degradation of a key glycolytic enzyme by APC/C-Cdh1. Nat Cell Biol. 2009;11(6):747–52.
Okatsu K, Oka T, Iguchi M, Imamura K, Kosako H, Tani N, et al. PINK1 autophosphorylation upon membrane potential dissipation is essential for Parkin recruitment to damaged mitochondria. Nat Commun. 2012;3:1016.
Jin SM, Youle RJ. The accumulation of misfolded proteins in the mitochondrial matrix is sensed by PINK1 to induce PARK2/Parkin-mediated mitophagy of polarized mitochondria. Autophagy. 2013;9(11):1750–7.
Kane LA, Lazarou M, Fogel AI, Li Y, Yamano K, Sarraf SA, et al. PINK1 phosphorylates ubiquitin to activate Parkin E3 ubiquitin ligase activity. J Cell Biol. 2014;205(2):143–53.
Kazlauskaite A, Kondapalli C, Gourlay R, Campbell DG, Ritorto MS, Hofmann K, et al. Parkin is activated by PINK1-dependent phosphorylation of ubiquitin at Ser65. Biochem J. 2014;460(1):127–39.
Koyano F, Okatsu K, Kosako H, Tamura Y, Go E, Kimura M, et al. Ubiquitin is phosphorylated by PINK1 to activate parkin. Nature. 2014;510(7503):162–6.
Gegg ME, Cooper JM, Chau KY, Rojo M, Schapira AH, Taanman JW. Mitofusin 1 and mitofusin 2 are ubiquitinated in a PINK1/parkin-dependent manner upon induction of mitophagy. Hum Mol Genet. 2010;19(24):4861–70.
Wang X, Winter D, Ashrafi G, Schlehe J, Wong YL, Selkoe D, et al. PINK1 and Parkin target Miro for phosphorylation and degradation to arrest mitochondrial motility. Cell. 2011;147(4):893–906.
Sarraf SA, Raman M, Guarani-Pereira V, Sowa ME, Huttlin EL, Gygi SP, et al. Landscape of the PARKIN-dependent ubiquitylome in response to mitochondrial depolarization. Nature. 2013;496(7445):372–6.
Kurz T, Terman A, Gustafsson B, Brunk UT. Lysosomes in iron metabolism, ageing and apoptosis. Histochem Cell Biol. 2008;129(4):389–406.
Grohm J, Plesnila N, Culmsee C. Bid mediates fission, membrane permeabilization and peri-nuclear accumulation of mitochondria as a prerequisite for oxidative neuronal cell death. Brain Behav Immun. 2010;24(5):831–8.
Giorgio M, Migliaccio E, Orsini F, Paolucci D, Moroni M, Contursi C, et al. Electron transfer between cytochrome c and p66Shc generates reactive oxygen species that trigger mitochondrial apoptosis. Cell. 2005;122(2):221–33.
Shimada K, Crother TR, Karlin J, Dagvadorj J, Chiba N, Chen S, et al. Oxidized mitochondrial DNA activates the NLRP3 inflammasome during apoptosis. Immunity. 2012;36(3):401–14.
Nakahira K, Haspel JA, Rathinam VA, Lee SJ, Dolinay T, Lam HC, et al. Autophagy proteins regulate innate immune responses by inhibiting the release of mitochondrial DNA mediated by the NALP3 inflammasome. Nat Immunol. 2011;12(3):222–30.
Ariga H, Takahashi-Niki K, Kato I, Maita H, Niki T, Iguchi-Ariga SM. Neuroprotective function of DJ-1 in Parkinson’s disease. Oxid Med Cell Longev. 2013;2013:683920.
Canet-Aviles RM, Wilson MA, Miller DW, Ahmad R, McLendon C, Bandyopadhyay S, et al. The Parkinson’s disease protein DJ-1 is neuroprotective due to cysteine-sulfinic acid-driven mitochondrial localization. Proc Natl Acad Sci U S A. 2004;101(24):9103–8.
Menu P, Vince JE. The NLRP3 inflammasome in health and disease: the good, the bad and the ugly. Clin Exp Immunol. 2011;166(1):1–15.
Martinon F, Mayor A, Tschopp J. The inflammasomes: guardians of the body. Annu Rev Immunol. 2009;27:229–65.
Marchi S, Patergnani S, Pinton P. The endoplasmic reticulum-mitochondria connection: one touch, multiple functions. Biochim Biophys Acta. 2014;1837(4):461–9.
Shi CS, Shenderov K, Huang NN, Kabat J, Abu-Asab M, Fitzgerald KA, et al. Activation of autophagy by inflammatory signals limits IL-1beta production by targeting ubiquitinated inflammasomes for destruction. Nat Immunol. 2012;13(3):255–63.
van der Burgh R, Boes M. Mitochondria in autoinflammation: cause, mediator or bystander? Trends Endocrinol Metab: TEM. 2015;26(5):263–71.
Heneka MT, Kummer MP, Latz E. Innate immune activation in neurodegenerative disease. Nat Rev Immunol. 2014;14(7):463–77.
Shachar T, Lo Bianco C, Recchia A, Wiessner C, Raas-Rothschild A, Futerman AH. Lysosomal storage disorders and Parkinson’s disease: Gaucher disease and beyond. Mov Disord. 2011;26(9):1593–604.
Sidransky E, Nalls MA, Aasly JO, Aharon-Peretz J, Annesi G, Barbosa ER, et al. Multicenter analysis of glucocerebrosidase mutations in Parkinson’s disease. N Engl J Med. 2009;361(17):1651–61.
Beutler E, Grabowski GA. Gaucher disease. In: Scriver CBA, Beaudet AL, Sly WS, Valle D, editors. The metabolic & molecular bases of inherited disease. New York: McGraw-Hill; 2001. p. 3635–68.
Sidransky E. Gaucher disease: complexity in a “simple” disorder. Mol Genet Metab. 2004;83(1–2):6–15. Review.
Nalls MA, Duran R, Lopez G, Kurzawa-Akanbi M, McKeith IG, Chinnery PF, et al. A multicenter study of glucocerebrosidase mutations in dementia with Lewy bodies. JAMA Neurol. 2013;70(6):727–35.
Mitsui J, Matsukawa T, Sasaki H, Yabe I, Matsushima M, Durr A, et al. Variants associated with Gaucher disease in multiple system atrophy. Ann Clin Transl Neurol. 2015;2(4):417–26.
Ito H, Nakayama K, Jin C, Suzuki Y, Yazawa I. alpha-Synuclein accumulation reduces GABAergic inhibitory transmission in a model of multiple system atrophy. Biochem Biophys Res Commun. 2012;428(3):348–53 [Research Support, Non-U.S. Gov’t].
Vargas KJ, Makani S, Davis T, Westphal CH, Castillo PE, Chandra SS. Synucleins regulate the kinetics of synaptic vesicle endocytosis. J Neurosci. 2014;34(28):9364–76.
Malec-Litwinowicz M, Rudzinska M, Szubiga M, Michalski M, Tomaszewski T, Szczudlik A. Cognitive impairment in carriers of glucocerebrosidase gene mutation in Parkinson disease patients. Neurol Neurochir Pol. 2014;48(4):258–61.
Pulkes T, Choubtum L, Chitphuk S, Thakkinstian A, Pongpakdee S, Kulkantrakorn K, et al. Glucocerebrosidase mutations in Thai patients with Parkinson’s disease. Parkinsonism Relat Disord. 2014;20(9):986–91.
Westbroek W, Gustafson AM, Sidransky E. Exploring the link between glucocerebrosidase mutations and parkinsonism. Trends Mol Med. 2011;17(9):485–93.
Sardi SP, Cheng SH, Shihabuddin LS. Gaucher-related synucleinopathies: the examination of sporadic neurodegeneration from a rare (disease) angle. Prog Neurobiol. 2015;125:47–62.
Gegg ME, Burke D, Heales SJ, Cooper JM, Hardy J, Wood NW, et al. Glucocerebrosidase deficiency in substantia nigra of parkinson disease brains. Ann Neurol. 2012;72(3):455–63.
Murphy KE, Halliday GM. Glucocerebrosidase deficits in sporadic Parkinson disease. Autophagy. 2014;10(7):1350–1.
Chiasserini D, Paciotti S, Eusebi P, Persichetti E, Tasegian A, Kurzawa-Akanbi M, et al. Selective loss of glucocerebrosidase activity in sporadic Parkinson’s disease and dementia with Lewy bodies. Mol Neurodegener. 2015;10(1):15.
Mazzulli JR, Xu YH, Sun Y, Knight AL, McLean PJ, Caldwell GA, et al. Gaucher disease glucocerebrosidase and alpha-synuclein form a bidirectional pathogenic loop in synucleinopathies. Cell. 2011;146(1):37–52 [Research Support, N.I.H., Extramural Research Support, N.I.H., Intramural].
Osellame LD, Rahim AA, Hargreaves IP, Gegg ME, Richard-Londt A, Brandner S, et al. Mitochondria and quality control defects in a mouse model of Gaucher disease – links to Parkinson’s disease. Cell Metab. 2013;17(6):941–53.
Zimran A, Elstein D. Management of Gaucher disease: enzyme replacement therapy. Pediatr Endocrinol Rev. 2014;12 Suppl 1:82–7.
Cabrera-Salazar MA, Deriso M, Bercury SD, Li L, Lydon JT, Weber W, et al. Systemic delivery of a glucosylceramide synthase inhibitor reduces CNS substrates and increases lifespan in a mouse model of type 2 Gaucher disease. PLoS ONE. 2012;7(8):e43310 [Comparative Study Research Support, Non-U.S. Gov’t].
Bendikov-Bar I, Maor G, Filocamo M, Horowitz M. Ambroxol as a pharmacological chaperone for mutant glucocerebrosidase. Blood Cells Mol Dis. 2013;50(2):141–5 [Research Support, Non-U.S. Gov’t].
Luan Z, Li L, Higaki K, Nanba E, Suzuki Y, Ohno K. The chaperone activity and toxicity of ambroxol on Gaucher cells and normal mice. Brain Dev. 2013;35(4):317–22 [Research Support, Non-U.S. Gov’t].
Patnaik S, Zheng W, Choi JH, Motabar O, Southall N, Westbroek W, et al. Discovery, structure-activity relationship, and biological evaluation of noninhibitory small molecule chaperones of glucocerebrosidase. J Med Chem. 2012;55(12):5734–48.
Gan-Or Z, Ozelius LJ, Bar-Shira A, Saunders-Pullman R, Mirelman A, Kornreich R, et al. The p.L302P mutation in the lysosomal enzyme gene SMPD1 is a risk factor for Parkinson disease. Neurology. 2013;80(17):1606–10 [Research Support, N.I.H., Extramural Research Support, Non-U.S. Gov’t].
Winder-Rhodes SE, Garcia-Reitbock P, Ban M, Evans JR, Jacques TS, Kemppinen A, et al. Genetic and pathological links between Parkinson’s disease and the lysosomal disorder Sanfilippo syndrome. Mov Disord. 2012;27(2):312–5 [Research Support, Non-U.S. Gov’t].
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing
About this chapter
Cite this chapter
Nguyen, M., Sidransky, E., Westbroek, W. (2016). The Deleterious Duo of Neurodegeneration: Lysosomes and Mitochondria. In: Reeve, A., Simcox, E., Duchen, M., Turnbull, D. (eds) Mitochondrial Dysfunction in Neurodegenerative Disorders. Springer, Cham. https://doi.org/10.1007/978-3-319-28637-2_12
Download citation
DOI: https://doi.org/10.1007/978-3-319-28637-2_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-28635-8
Online ISBN: 978-3-319-28637-2
eBook Packages: MedicineMedicine (R0)