Abstract
Autophagy delivers cytoplasmic components and organelles to lysosomes for degradation. This pathway serves to degrade nonfunctional or unnecessary organelles and aggregate-prone and oxidized proteins to produce substrates for energy production and biosynthesis. Macroautophagy delivers large aggregates and whole organelles to lysosomes by first enveloping them into autophagosomes that then fuse with lysosomes. Chaperone-mediated autophagy (CMA) degrades proteins containing the KFERQ-like motif in their amino acid sequence, by transporting them from the cytosol across the lysosomal membrane into the lysosomal lumen. Autophagy is especially important for the survival and homeostasis of postmitotic cells like neurons, because these cells are not able to dilute accumulating detrimental substances and damaged organelles by cell division. Our current knowledge on the autophagic pathways and molecular mechanisms and regulation of autophagy will be summarized in this review. We will describe the physiological functions of macroautophagy and CMA in neuronal cells. Finally, we will summarize the current evidence showing that dysfunction of macroautophagy and/or CMA contributes to neuronal diseases. We will give an overview of our current knowledge on the role of autophagy in aging neurons, and focus on the role of autophagy in four types of neurodegenerative diseases, i.e., amyotrophic lateral sclerosis and frontotemporal dementia, prion diseases, lysosomal storage diseases, and Parkinson’s disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
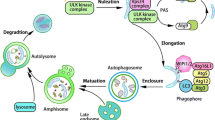
Autophagy is a catabolic process in which cytoplasmic material and/or organelles are transported to lysosomes for degradation by lysosomal acid hydrolases [19, 55, 58, 81, 180]. The delivery of cytoplasmic material to lysosomes can occur via three different routes (Fig. 1). In macroautophagy, the cytoplasm and/or organelle is first engulfed by a flat membrane cistern that forms a double-membrane vesicle, called the autophagosome, which then fuses with endosomal and lysosomal vesicles. In a poorly defined process called microautophagy, lysosomes internalize small portions of cytoplasm in a process that resembles endocytosis. In chaperone-mediated autophagy (CMA), cytosolic soluble proteins containing a certain sequence tag (the KFERQ-like motif) in their amino acid sequence are recognized by the A isoform of the lysosomal associated membrane protein 2 (LAMP-2A), and transported through the lysosomal membrane in a process assisted by chaperones [108].
A schematic representation of the three different forms of autophagy. In macroautophagy, the cytoplasmic cargo is first segregated into autophagosomes. Autophagosomes are formed by phagophores, also called isolation membranes, that expand and capture the cargo inside the forming autophagosome, which then fuses with a lysosome. In microautophagy, lysosomal membrane invaginates and engulfs cytosol. In chaperone-mediated autophagy, protein substrates are unfolded by the cytosolic chaperone HSC70 prior to transport through the lysosomal membrane. The lysosomal associated membrane protein LAMP-2A is thought to function as a receptor and/or channel for CMA. Intralysosomal HSC70 assists the transport across the lysosomal membrane
Autophagy is functioning under basal conditions, but during nutrient starvation it is upregulated in order to supply nutrients by degradation of redundant cytoplasmic material. Clearance of aggregate-prone proteins and damaged organelles by macroautophagy is imperative for postmitotic cells like neurons, which cannot dilute these components by cell division [27]. The role of autophagy in neurons is highlighted by a recent systems biology study revealing the pivotal role of dysregulated autophagy in neurodegenerative dementia [25], and the proposed role of autophagy in the turnover of repressor element 1-silencing transcription factor (REST), a central player in the repression of genes promoting cell death and in the induction of genes mediating stress resistance in neurons [135].
Macroautophagy: formation of the phagophore
Autophagosomes are formed by phagophores (also called isolation membranes), flat membrane cisterns that elongate and curve into double-membraned vesicles. The ATG proteins (autophagy-related proteins) were first discovered using yeast mutants. After the initial discoveries in yeast, most of these autophagy proteins were discovered in mammalian cells; some of them had already been described and named before their role in autophagy was determined. Autophagosome biogenesis is initiated by sequential function of several protein complexes, including ULK1/ULK2 complex, PI3KC3 complex, ATG9 and its binding partners, ATG12-ATG5 complex, and LC3-phosphatidyl ethanolamine complex [58, 81, 180]. These complexes and proteins are summarized in Table 1 and Fig. 2.
A schematic representation of phagophore formation in ER-associated, DFCP1-positive omegasome. ULK1/2-ATG13-FIP200-ATG101 (induction) complex is stable regardless of nutrient status. Under nutrient-rich conditions mTORC1 associates with the ULK complex and inactivates it by phosphorylation of ULK1/2 and ATG13. During nutrient starvation autophagy begins when mTORC1 dissociates from the ULK complex, which results in partial dephosphorylation of ULK1/2 and ATG13, and induction of macroautophagy. ULK1/ULK2 becomes activated by phosphorylation of two other serine residues by AMPK, which in turn makes it capable of phosphorylating ATG13 and FIP200 (these phosphorylations are symbolized by stars). This initiates the recruitment of PI3KC3 complex to the omegasome where it produces PI3P, which in turn recruits PI3P-binding proteins including WIPIs to the growing phagophore. ATG12-ATG5 complex binds ATG16L1 and associates with the phagophore. The localization ATG12-ATG5 determines the site of LC3 lipidation to LC3-II, which associates with the phagophore membrane. ATG9 is not depicted in this figure since it is currently unclear at which stage it functions. The function of the omegasome marker DFCP1 is also unclear
Mammalian target of rapamycin complex 1 (mTORC1) is a central regulatory hub in nutrient sensing and metabolism. Inactivation of mTORC1 initiates the activation of ULK1/ULK2 complex and autophagosome formation (see below for details) (Fig. 2). The next protein complex to be activated is the class III phosphatidyl inositol 3-kinase (PI3KC3) complex, for which beclin 1 acts as a scaffold. Beclin 1, encoded by the BECN1 gene, functions at an early step during autophagosome formation. Beclin 1 can form a complex with several proteins, but during autophagosome formation it interacts with PI3KC3, its regulatory subunit PIK3R4, activating molecule in beclin 1-regulated autophagy (AMBRA1), and ATG14 that targets the complex to the endoplasmic reticulum (ER) [148]. Localized synthesis of phosphatidyl inositol 3-phosphate (PI3P) in the ER recruits PI3P-binding proteins called double FYVE-containing protein 1 (DFCP1) and WIPI1 or WIPI2 (WD repeat domain, phosphoinositide interacting). This recruitment is connected to the formation of specialized ER subdomains called omegasomes (Fig. 2) [6]. Autophagosome biogenesis occurs inside these omega-shaped subdomains. ATG9 is a membrane spanning protein that is needed for autophagosome formation. ATG9 recycles between the trans-Golgi network and endosomes in an ULK-dependent manner, and has been suggested to assist in the delivery of membrane to phagophores [219, 260]. Recent studies in yeast showed that Atg1 (the yeast equivalent to ULK1) phosphorylates Atg9, and that this phosphorylation is required for autophagosome formation [178]. Another group showed that the amount of Atg9 protein directly correlates with the frequency of autophagosome formation and rate of autophagic degradation [100]. It remains to be shown whether the mammalian ATG9 has similar functions. During phagophore elongation the membrane recruits ATG12 that is covalently conjugated to ATG5. These conjugates form large oligomers by additionally recruiting ATG16L1. The ATG12-ATG5 conjugate determines the site where the protein called microtubule associated protein 1 light chain 3 (MAP1-LC3) is conjugated to phosphatidyl ethanolamine (PE), giving rise to the lipidated form of LC3, called LC3-II [67, 94]. The lipidation is needed for the recruitment of LC3 to the phagophore [104]. LC3 is the most widely used marker of autophagosomes in mammalian cells. Soon after sealing of the autophagosome, ATG12-ATG5/ATG16L1 dissociates from the membrane, while part of LC3-II stays on the inner limiting membrane and is delivered to the lysosome together with the cytoplasmic cargo. Before lipidation, newly synthesized LC3 is cleaved by ATG4, which is also able to delipidate LC3-II after a sealed autophagosome has formed. The conjugation reactions of ATG12 to ATG5 and LC3 to PE are similar to protein ubiquitination. The equivalents to the ubiquitin ligases E1 and E2 are ATG7 and ATG10 for ATG12-ATG5 conjugation, and ATG7 and ATG3 for LC3-PE conjugation [180]. Several recent reviews have been published on phagophore and autophagosome formation [58, 73, 211, 219].
One of the most debated questions in macroautophagy concerns the origin of phagophore and autophagosome membranes [54]. As described above recent results indicate that ER plays a role in phagophore biogenesis. In starved cells, phagophores locate between two cisterns of rough ER [84, 256]. The omegasome marker protein DFCP1 has been localized by immunoelectron microscopy in delicate membrane tubules extending from the ER to the phagophore [235]. Recent studies also suggest that coat protein II (COPII) coated vesicles budding from ER exit sites deliver membrane for phagophores [76, 223]. During starvation, the outer mitochondrial membrane may also participate in phagophore biogenesis [80], and a recent paper showed that autophagosomes form in ER-mitochondria contact sites [82]. Further, plasma membrane may contribute to the formation of autophagosome precursors, structures that give rise to phagophores [187]. Exit from the Golgi complex may be necessary for the delivery of membrane during the elongation of phagophores [77, 222, 238]. Recycling endosomes may also contribute membrane to forming autophagosomes [134]. Taken together, it is possible that phagophores have more than one membrane source, possibly depending on the cargo or the signals that initiate autophagy.
After their formation, autophagosomes undergo a maturation process that includes fusion with endosomal and lysosomal vesicles (Fig. 3) [52]. These fusion events are controlled by several RAB and SNARE proteins [16, 31, 120]. Autophagosomes that have fused with endosomes are called amphisomes, and fusions with lysosomes produce autolysosomes. During maturation, the autophagosome limiting membrane acquires lysosomal membrane proteins, for example LAMP-1, LAMP-2 and the vacuolar proton pump V-ATPase, the activity of which acidifies the contents of autophagosomes. Lysosomal hydrolases that degrade the autophagosome cargo are also delivered into autophagosomes via the fusion of lysosomes with autophagosomes. Lysosomal membrane protein transporters then carry the degradation products back to cytoplasm, where the cell can use them for energy production and/or biosynthetic reactions. It is currently unclear what happens after the degradation in autolysosomes is complete, or alternatively, in the case that the degradation is not complete, but different possibilities have been reported. (1) New lysosomes may reform from autolysosomes as budding tubular extensions, which then separate and form new lysosomes [261]. (2) The second possibility is exocytosis of autolysosome contents by fusion of the limiting membrane with the plasma membrane and extrusion of the contents into the extracellular space. These two alternatives are not mutually exclusive. Recent studies suggest that autophagosome exocytosis increases if autophagic degradation capacity is compromised by inhibiting autophagosome–lysosome fusion. This process may depend on the transcription factor EB (TFEB) [51, 122, 150] and/or other transcription factors like TFE3 [143]. Impaired acidification of lysosomes may also increase exocytosis of autolysosome contents [186].
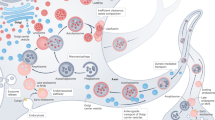
Autophagosome formation and maturation are spatially controlled in neuronal cells. Autophagosomes form in the neurite tips, where ubiquitinated protein aggregates bind different cargo receptors that also bind the autophagosome protein LC3-II. This directs phagophore growth and leads to engulfment of harmful aggregates to autophagosomes (in the drawing in axon terminals). Autophagosomes as well as endocytosed material in multivesicular bodies (MVB) and late endosomes (LE) are transported by dynein motors towards the neuron soma. During the transport, autophagosomes fuse with multivesicular bodies and late endosomes forming amphisomes, and these finally fuse with lysosomes to create autolysosomes. Fusion with lysosomes leads to acidification of the cytosolic cargo by proton pump V-ATPase in lysosomal membrane (acidification is illustrated by fading of the pink and blue color). Lysosomes move bidirectionally by means of kinesin and dynein motors. RE/EE recycling endosome/early endosome
Nutritional and transcriptional control of macroautophagy
Macroautophagy induction during starvation is controlled by a sophisticated system of amino acid sensing, which is coupled to transcription-dependent and -independent activation of autophagosome formation. This system allows the cell to respond to starvation by increased allocation of amino acids from proteins among the sequestered autophagic cargo. The central part of the amino acid sensing machinery is the conserved protein complex named mammalian Target Of Rapamycin (mTOR) complex 1 (mTORC1), which is a potent repressor of autophagy in all eukaryotes (Fig. 4) [155]. mTORC1 consists of (1) the Ser/Thr protein kinase mTOR; (2) regulatory associated protein of mTOR (RAPTOR) that acts as a scaffold for the complex; and (3) the additional subunits PRAS40 and mLST8. mTORC1 acts on several intracellular pathways and its direct substrates include the translational regulator 4EBP, ribosomal S6 kinase (S6K1), death-associated protein 1 (DAP1, a negative regulator of autophagy [119]), and AMBRA1, a protein needed for autophagosome formation [164]. The primary autophagy regulating targets of mTORC1 are ULK1 and/or Unc-51-like kinase 2 (ULK2) [103]. This linkage of mTOR and ULK1/ULK2 connects autophagy regulation to the availability of nutrients. The ULK1/ULK2 complex contains ULK1 or ULK2, 200 kDa FAK family kinase-interacting protein/RB1-inducible coiled-coil 1 (FIP200/RB1CC1), ATG13 and C12orf44/ATG101, and functions during autophagy induction. Under starvation conditions, ULK1/ULK2 phosphorylation by mTORC1 at serine 757 is reduced, and consequently, ULK1/ULK2 becomes activated by phosphorylation of two other serine residues by AMP activated protein kinase (AMPK), which makes it in turn capable of phosphorylating ATG13 and FIP200 (Fig. 2). This initiates autophagy by the formation of an active ULK kinase complex. Under nutrient-rich conditions, mTORC1 is active and inhibits autophagy by phosphorylating ATG13 and ULK1/ULK2 subunits and thereby repressing ULK1/ULK2 kinase activity and ultimately autophagy [112]. mTORC1 activity in turn is negatively regulated by the tuberous sclerosis complex (TSC) 1/2. Neurons deficient in TSC2 were recently shown to have an mTORC1-independent mechanism for autophagy regulation [46]. Unlike non-neuronal cells, neurons deficient in TSC2 show increased accumulation of autolysosomes and increased autophagic flux, despite mTORC1-dependent inhibition of ULK1. In the absence of TSC2, ULK1 was activated by AMPK. These findings suggest that, in TSC2 knockdown neurons, AMPK activation is the dominant regulator of autophagy, and that neuronal TSC1/2 complex is required for coordinated regulation of autophagy by AMPK [46].
mTORC1 is the main regulator of starvation-induced autophagy. mTORC1 senses the nutrient status of the cell via different pathways. Amino acid sensing occurs on the lysosomal surface. When the amount of amino acids is high (left side of the figure) V-ATPase signals to Ragulator complex, which leads to activation of the small GTPase RagA or B by GDP exchange to GTP. Mammals express four Rag proteins—RagA, RagB, RagC, and RagD—that form heterodimers consisting of RagA or RagB with RagC or RagD. GTP-bound RagA or RagB and mTORC1 are recruited to the lysosomal surface. mTORC1 then interacts with Rheb–GTP, and phosphorylates its downstream effectors leading to inhibited autophagy and enhanced cell growth. Also TFEB is phosphorylated and remains in the cytosol. When amino acids are missing or low (right side of the figure), V-ATPase prevents the guanine nucleotide exchange activity of the Ragulator complex, and mTORC1 remains inactive and resides in the cytosol. Downstream effectors of mTORC1 are not phosphorylated, leading to general translational halt and induction of autophagy (see Fig. 2). Also TFEB is not phosphorylated and thus it translocates to the nucleus, where it activates the transcription of autophagy and lysosomal genes
The precise mechanism for amino acid sensing is not yet resolved, but recent findings indicate that it is initiated by the accumulation of amino acids within the lysosomal lumen. This induces V-ATPase to form a complex with Ragulator, a guanine nucleotide exchange factor, and small heterodimeric Rag (A + C or B + D) guanosine triphosphatases (GTPases) (Fig. 4). Rags are heterodimers consisting of RagA or RagB, which are highly similar to each other, bound to RagC or RagD, respectively. Ragulator and Rag GTPases locate on the lysosomal surface, forming a docking site for mTORC1. mTORC1 then becomes activated to inhibit autophagy (Fig. 4 ) [270].
In addition to this transcription-independent regulation of autophagy by mTORC1, autophagy is also regulated by a transcription-dependent mechanism [202]. This mechanism depends on TFEB, which is also critical for the coordinated expression of lysosomal genes [202]. In the presence of nutrients, mTORC1 phosphorylates TFEB at serine 211. This causes retention of TFEB in the cytoplasm. In the absence of nutrients, when mTORC1 is inactive, TFEB translocates into the nucleus [192, 209]. After binding to its target sequence in the promotor-region of lysosomal and autophagic genes TFEB induces the expression of these genes. Thus, TFEB directly links the enhanced formation of autophagosomes and the subsequent degradation of autophagocytosed cargo in autolysosomes by increased lysosomal biogenesis (Fig. 4).
mTORC1-dependent and TFEB-induced induction of autophagy are linked, connecting nutrition sensing at the lysosomal membrane by mTORC1 and V-ATPase, and induction of autophagic and lysosomal gene expression by TFEB (Fig. 4). Puertollano and coworkers showed that, upon inhibition of mTORC1, TFEB transiently associates with mTORC1 and then rapidly translocates to the nucleus [142]. Under normal conditions, most TFEB is retained in the cytosol by a phosphorylation-dependent interaction with members of the YWHA (14-3-3) protein family, whereas a small amount associates with lysosomal membranes. TFEB bound to the cytosolic side of the lysosomal membrane becomes phosphorylated by mTORC1 and cycles back to the cytosol [142]. In the presence of nutrients, active Rag GTPases A and B recruit TFEB and mTORC1 to lysosomal membranes [144, 145], where mTORC1 comes in close contact with Ras homolog enriched in brain (Rheb), an essential GTPase for amino acid activation of mTORC1 [133]. Rheb is a target of growth-factor-dependent signaling and thereby further modulates mTORC1 activity [72, 93]. This nutrient sensing at lysosomal membranes allows the cell to coordinate availability of amino acids with the induction of a number of autophagy genes and genes encoding lysosomal hydrolases and membrane proteins.
In addition to amino acid starvation, other cellular stress factors can induce macroautophagy, including hypoxia, glucose deprivation, endoplasmic reticulum stress and reactive oxygen species (ROS). In experimental neuronal models of hypoxia–ischemia (oxygen and glucose deprivation) autophagy was found to increase dramatically, ultimately leading to cell death [177, 210]. In agreement with these results, a recent study showed that hexokinase, a central enzyme in glucose metabolism, directly activates macroautophagy by inhibition of mTORC1 [191]. Papadakis et al. [177] examined why hippocampal CA1 neurons are more vulnerable to ischemic insults than CA3 neurons and how ischemic preconditioning affects these neurons. Hamartin, the product of TSC1, was shown to confer neuroprotection against ischemia both in vitro and in vivo in mice by inducing neuronal macroautophagy in an mTORC1- and presumably Rheb-dependent manner [177]. Another study showed that macroautophagy induction under hypoxic conditions was independent of mTORC1, but at least in part dependent on hypoxia-inducible factor (HIF), a key transcription factor that allows adaptation to and survival in reduced oxygen concentrations. HIF was shown to induce the expression of Bnip3, whose gene product (BNIP3, BCL2/adenovirus E1B 19 kDa interacting protein 3) is proposed to induce macroautophagy via beclin 1 (reviewed in [149]).
Another cellular stressor leading to activation of macroautophagy is endoplasmic reticulum stress, induced by unfolded or misfolded proteins [63, 165, 170, 200]. Several different pathways have been implicated, including direct induction of autophagic genes by the unfolded protein response (UPR) sensors. In mammals, serine/threonine-protein kinase/endoribonuclease protein (IRE1), protein kinase-like ER kinase (PERK) and activating transcription factor 6 (ATF6) [170] function as UPR sensors. Furthermore, additional signaling pathways have been described to induce macroautophagy, including negative regulation of the Akt-mTOR pathway by UPR and calcium signaling via Ca2+-dependent protein kinase C theta (PKCθ) activity [200]. Finally, ROS were shown to be a common inducer of autophagy in neurons. Since ROS are highly toxic for cells, macroautophagy clearly plays a neuroprotective role in this regard, by removing ROS-generating organelles including mitochondria and endoplasmic reticulum. However, how precisely ROS induce macroautophagy is poorly understood. In cancer cells both Akt-mTOR pathway and BNIP3-dependent pathways were described to contribute to autophagy induction [24, 265]. Since ROS are highly reactive, direct interaction with autophagy-related proteins are also likely to contribute: the redox-sensitive cysteine protease ATG4 was shown to be directly influenced by ROS. A cysteine residue near the catalytic center becomes oxidized, thereby inhibiting ATG4 activity and the delipidation of LC3-II [204]. Intriguingly, mitophagy, selective macroautophagy of mitochondria, was demonstrated to be mediated by mitochondria-generated ROS and a direct interaction of Rheb and LC3-II [151].
Localization and movement of autophagosomes in neuronal cells
Neuronal cells are morphologically and functionally specialized. In agreement with this specialization, the formation and maturation of autophagosomes are spatially controlled in neurons. Autophagosomes were identified in neurite tips by electron microscopy already in 1973 [23]. Live cell imaging showed that both endocytic and autophagic vesicles move along axons towards the cell soma [90]. Later studies demonstrated that autophagosomes positive for green fluorescent protein (GFP) tagged LC3 move in axons bidirectionally, with net movement towards the cell soma where the majority of lysosomes are located. This movement depends on microtubules and on the motor proteins dynein and kinesin [107, 255]. Dynein mediates the movement towards cell soma, while kinesin assists movement to the opposite direction [138]. A recent study showed that the scaffolding protein JIP1 regulates autophagosome transport in neuronal cells. JIP1 binds LC3 and the dynein activator protein dynactin, and these interactions are needed for autophagosome transport towards the cell soma [66]. Interestingly, JIP1 also binds and activates kinesin-1 in a phosphorylation-dependent manner. Phosphodeficient JIP1 activates transport towards the cell soma, while phosphomimetic JIP1 aberrantly activates transport to the opposite direction. In primary neurons, autophagosomes constitutively form in the tips of neurites and then travel towards the cell soma, undergoing fusion with endosomal and lysosomal vesicles and turning acidic during the transport [66, 138] (Fig. 3). These autophagosomes may contain organelles like mitochondria and/or aggregate-prone proteins as cargo. The small GTPase RAB7 is needed for the maturation of autophagosomes to autolysosomes, most likely for the fusion with lysosomes [78, 97]. In Purkinje neurons, insulin-like growth factor I enhances autophagosome maturation by increasing the interaction of RAB7 with RILP, a RAB7 effector protein that is needed for association of RAB7 with dynein motor proteins [8]. Fusion of autophagosomes with lysosomes initiates the degradation of autophagic cargo. Inhibition of lysosomal proteolysis with drug treatment or due to genetic deficiency of lysosomal enzymes (discussed later) slows down the axonal transport of autolysosomes, endosomes and lysosomes. As a consequence, these organelles accumulate in axonal swellings while other organelles like mitochondria are still able to move normally [127]. Accumulation of autolysosomes in axonal swellings is a typical finding in Alzheimer’s disease [168].
Basal and induced macroautophagy and selective and non-selective macroautophagy in neurons
Basal macroautophagy occurs at low level without a specific stimulus. It is important for cellular homeostasis, because it prevents metabolic and oxidative stress by degrading long-lived proteins, damaged organelles like mitochondria, and aggregated or aggregate-prone proteins. These functions are especially important for postmitotic cells like neurons, which are not able to dilute the harmful constituents by cell division. Stress such as starvation enhances macroautophagy several-fold in isolated tissues and cultured cells [158]. This allows cells to mobilize energy from internal resources and to survive the stress. Induced macroautophagy is inhibited by mTORC1 that senses cellular energy and nutrient levels and regulates cell growth, while basal macroautophagy is less sensitive to mTORC1 activity [4]. Although both basal and induced autophagosomes may form in a similar manner [160], the proteins mediating their fusion events with endosomes and lysosomes may differ. The small GTPase RAB7 is essential for the maturation of starvation-induced, but not basal, autophagosomes [97]. Recent results also show that the histone deacetylase HDAC6 and valosin-containing protein (VCP/p97) are essential for the maturation of basal, but not starvation-induced, autophagosomes [124, 234]. In agreement with these results, proteomic analysis also revealed that basal and induced autophagosomes contain proteomes that only partially overlap [44].
In mammals, the induction of macroautophagy is organ dependent. Fasting causes a more profound induction of macroautophagy in heart and muscle when compared to brain [156]. This was suggested to be due to nutrient supply from other organs to brain during food deprivation. A more recent study, however, detected a significant increase in autophagosomes in cerebral cortical and cerebellar Purkinje cells in nutrient-deprived mice [5]. Interestingly, another recent study showed that physical exercise induces macroautophagy in the cerebral cortex of adult mice [85]. These observations suggest that macroautophagy may participate in mediating the known beneficial effects of calorie restriction and exercise on brain function.
Recent studies support the idea that basal macroautophagy is selective, acting as a quality control mechanism, while induced macroautophagy is less selective, acting mainly to support survival by producing nutrients by degrading randomly segregated cytoplasmic components and organelles [124]. As discussed below, in neuronal cells defective macroautophagy leads to accumulation of protein aggregates and neurodegeneration. Selective macroautophagy of aggregates is called aggrephagy [121]. Dysfunction of mitochondria can cause oxidative stress and is also linked to aging and neurodegeneration [50, 188, 233, 269]. Cells have developed a mechanism to remove damaged mitochondria by selective macroautophagy, called mitophagy [14, 225, 228]. This process is induced by mitochondrial depolarization, which stabilizes the serine/threonine kinase PINK1 on the mitochondrial surface. PINK1 then recruits the E3 ubiquitin ligase parkin/PARK2 [162], which leads to ubiquitination of mitochondrial proteins. Ubiquitination recruits mitophagy adaptor proteins, which in turn recruit autophagy proteins and initiate mitophagy [50]. Loss of the PINK1/PARK2 pathway causes one form of familial Parkinson’s disease. Mitochondrial depolarization increases the interaction of PARK2 with AMBRA1, a component of the PI3KC3 complex. AMBRA1 favors the interaction of beclin 1 with PI3KC3 and the production of phosphatidylinositol 3-phosphate by the PI3KC3. This in turns enhances mitophagy [239].
Protein aggregates or damaged organelles are hallmarks of many neurological diseases. They are selectively engulfed by a mechanism involving one or more cargo receptors, including sequestosome 1 (SQSTM1/p62), NBR1, NDP52, optineurin (OPTN), NIX/BNIP3L, and FUNDC1 [113, 121, 196, 205] (Fig. 5). The cargo receptors contain a cargo-binding domain and another domain that binds LC3 or another member of the ATG8 family (LC3A, LC3B, LC3C, GABARAP, GAPARAPL1, GATE16/GAPARAPL2), which enables the recruitment of autophagic membranes to the cargo. BNIP3L and FUNDC1 are integral membrane proteins of the mitochondrial outer membrane and interact with LC3 in order to mediate hypoxia-induced mitophagy [132, 169]. Additional scaffolding proteins like HDAC6 and ALFY may be needed for the selective macroautophagic segregation of protein aggregates [95, 190, 213]. Similar to the proteasomal system, ubiquitination serves as a general cargo recognition signal for selective macroautophagy, and many of the cargo receptors have a ubiquitin-binding domain. Unlike in proteasomal degradation, selective macroautophagy degrades the cargo together with the ubiquitin tag, as well as the associated cargo receptors and scaffolding proteins. Lysine 63-linked polyubiquitin has been suggested to be the main signal for macroautophagic degradation of protein aggregates and mitochondria [125, 172, 224]. Lysine 63-linked ubiquitination increases the formation of aggresomes, inclusion bodies containing aggregated proteins, as well as the macroautophagic degradation of the ubiquitinated proteins. HDAC6 and the motor protein dynein participate in the aggresome formation by transport of ubiquitinated cargo, i.e., proteins or mitochondria, along microtubules towards their minus ends (in neurons towards the cell soma) [69, 95, 125, 171, 172]. HDAC6 regulates microtubule dynamics and microtubule-mediated transport by deacetylating tubulin [41]. During proteasome inhibition, HDAC6 promotes the localization of the E3 ubiquitin ligase TRIM50 to protein aggregates, which promotes their transport to aggresomes [69]. HDAC6 also controls the fusion of autophagosomes with lysosomes by recruiting the machinery that assembles filamentous actin, which in turn stimulates autophagosome–lysosome fusion [124].
Mechanism and regulation of CMA
The cytosolic chaperone heat shock cognate protein 70 (HSC70) binds to the canonical KFERQ-like motif of substrate proteins, mediating their binding to the C-terminal cytosolic tail of the lysosomal membrane protein LAMP-2A. This binding mediates the assembly of monomeric LAMP-2A into a high molecular weight complex, which was suggested to act as a translocation pore [11]. In addition to HSC70 and LAMP-2A, other proteins such as co-chaperones are present in the translocation complex [108]. How the translocation pore accommodates its substrates remains an open question. After translocation of substrate proteins the complex was proposed to dissociate into monomers. Intralysosomal form of HSC70 is also needed for the transport of the CMA substrates across the lysosomal membrane [36].
The selectivity of CMA is mediated by the KFERQ-like motif which binds to cytosolic HSC70. This motif is present in ~30 % of cytosolic proteins. Similar to macroautophagy, also the level of CMA changes according to conditions, allowing adaptation to long-term needs of the cells. The extent of macroautophagy and CMA are tightly linked; a block in macroautophagy has been shown to upregulate CMA and vice versa [109, 147]. The extent of CMA depends on the amount of its receptor LAMP-2A on the lysosomal membrane [47]. The levels of available LAMP-2A can be modulated by changes in its biosynthesis, degradation, localization in the lysosomal membranes, and conformation [11]. In addition, the stability of the translocation complex was shown to be regulated by GTP [12]. Phosphorylation and acetylation where shown to modify amino acid charge of the KFERQ-like motifs, which may modify the efficiency of recognition by HSC70. Partial unfolding of the protein may reveal the KFERQ-like motif in case it is normally concealed in the interior of a protein and this may facilitate CMA-mediated degradation [136, 185].
Similar to macroautophagy, CMA is activated by starvation of nutrients, but the timing of these two autophagy pathways differ. Macroautophagy is activated as the first response to starvation, and CMA is upregulated later, when the activity of macroautophagy decreases. In cultured cells, removal of serum from the culture medium for more than 10 h increases the activity of CMA and reduces the levels of KFERQ-containing proteins [173]. It has been suggested that the switch from macroautophagy to CMA is beneficial since it allows more selective degradation of proteins that are not necessary for cells during nutrient deprivation. CMA may also allow the cells to adjust their proteome and adapt to the changed conditions during long starvation [38].
Physiological functions of autophagy in neuronal cells
A growing number of physiological functions have been described for both macroautophagy and CMA [55, 173]. In postmitotic cells like neurons, autophagy prevents the accumulation of oxidized proteins, harmful aggregates and nonfunctional, potentially harmful organelles like mitochondria. The importance of autophagy for neurons is highlighted by a recent systems biology study that revealed a pivotal role of dysregulated autophagy in neurodegenerative dementia [25].
The role of macroautophagy in neurons has been studied using mice with central nervous system (CNS)-specific, or neuronal cell type-specific, knockout of autophagy proteins (ATG5, ATG7 or FIP200/RB1CC1) that are indispensable for autophagosome formation [83, 117, 128]. These studies report shortened life span and progressive motor and behavioral defects in the mutant mice. Histological examination of CNS-specific knockout mice revealed neurodegenerative changes in brain that were most prominent in the cerebral cortical region and in cerebellar Purkinje cells. In Atg5 and Atg7 mutant mice, an increase in the number of TUNEL-positive cells was detected in the cerebral cortex and within the granular cell layer of the cerebellum [83, 117]. In the cerebellum of FIP200 mutant mice, neuronal loss, spongiosis and neurite degeneration were observed [128]. Ubiquitin-positive inclusions and protein aggregates were reported in Atg5 mutant mice in large neurons in the thalamus, midbrain, pons, medulla and dorsal root ganglion, as well as in neurons in the cerebral cortex, hippocampus, striatum and olfactory bulb [83], while in Atg7 mutant mice these were encountered in the cerebral cortex, cerebellar Purkinje cells, hippocampal pyramidal neurons, thalamus, hypothalamus, amygdala and pontine nuclei [117], and in FIP200 mutant mice in Purkinje cells and white matter in the cerebellum [128]. Furthermore, inclusion bodies were only observed in neurons, not in glial cells of Atg5 mutant mice [83]. Notably, the proteasomal activity was not affected in Atg7 and FIP200 mutant mice, indicating that ubiquitin-positive aggregates did not accumulate due to impaired proteasome function [117, 128]. In addition, abnormal mitochondria were detected in Purkinje cells of FIP200 mutant mice, possibly due to defective removal via mitophagy [128]. Taken together, these studies indicate that macroautophagy is necessary for the elimination of aggregate-prone proteins in neurons even in the absence of disease-causing mutations in these cargo proteins, and that loss of macroautophagic activity leads to severe neurodegeneration.
To circumvent the early lethal effects of the classical knockout mice, neuronal cell type-specific knockout mice have also been generated. Knockout of Atg7 specifically in dopamine neurons of midbrain caused axonal and dendritic dystrophy and loss of dopamine neurons [3, 65]. Inclusions positive for ubiquitin and α-synuclein were detected in the degenerating neurons, similar to the findings in Atg5 and Atg7 CNS-specific knockout mice [83, 117]. Purkinje cell-specific conditional Atg5 or Atg7 knockout mice were used to show that macroautophagy is essential for the prevention of axonal degeneration and maintenance of homeostasis in axon terminals [118, 167]. Interestingly, the Atg7 knockout Purkinje cells did not show atrophy of dendrites or dendritic spines. However, the axonal damage was followed by Purkinje cell death. Deletion of Atg7 in mouse pro-opiomelanocortin (POMC) neurons led to an age-related accumulation of ubiquitin and SQSTM1 positive aggregates in the hypothalamus, and disrupted maturation of POMC-containing axonal projections [33]. Higher body weight, increased adiposity and glucose intolerance were also observed in these mutant mice [33, 184]. The results suggest that macroautophagy in the hypothalamus may be able to control and reduce obesity. Additionally, in Purkinje cells and POMC neurons, macroautophagy may be specifically important for the homeostasis of axons. Knockout mice for Atg7 in ventral midbrain dopamine neurons were used to demonstrate that macroautophagy plays a role in the regulation of presynaptic transmission. The axonal profiles of the Atg7 knockout dopamine neurons were one-third larger than the control axonal profiles. The mutant neurons released ~50 % more dopamine after stimulus, and the synaptic recovery was also faster than in the control neurons [89, 231]. Induction of macroautophagy by rapamycin treatment reduced the axon profile size of dopamine neurons by ~30 % in control mice, but had no effect on the Atg7 knockout axons. Rapamycin treatment also decreased the dopaminergic synaptic vesicle density and inhibited dopamine release by 25 % in control mice, but not in the Atg7 knockout mice. These findings illustrate that macroautophagy can reduce presynaptic activity in vivo, likely by degrading synaptic vesicles [89, 231].
A recent study showed that macroautophagy negatively regulates axon growth in cortical neurons [10]. Silencing of the autophagy protein ATG7 caused elongation of axons, while activation of macroautophagy by rapamycin treatment suppressed axon growth. The regulation was shown to be mediated via the Rho A-ROCK pathway. Finally, both defective and excessive macroautophagy have been shown to contribute to neurite degeneration in various model systems. These studies are summarized in a recent review [254].
Autophagy can protect neuronal cells during stress or injury
Several studies have shown that macroautophagy can protect neuronal cells under stress conditions. The class III histone deacetylase sirtuin1 (SIRT1) mediates the beneficial metabolic effects of caloric restriction. SIRT1 was shown to protect neurons against the toxicity of mutant huntingtin, while reduction of SIRT1 increased huntingtin toxicity [99]. Another group showed that SIRT1 overexpression prevented prion peptide neurotoxicity by inducing macroautophagy, while preventing autophagy by knockdown of Atg5 abolished the SIRT1 induced neuroprotection [98]. Further, the macrolide drug FK506 delayed the accumulation of prion protein in brains of mice inoculated with Fukuoka-1 prion [161]. This was accompanied by an increased expression of autophagy proteins in brain tissue. Cell culture experiments revealed that FK506 upregulated macroautophagy and enhanced degradation of prions in autolysosomes. Finally, in Purkinje cells and retinal ganglion cells suffering from axonal dystrophy or traumatic injury, induction of macroautophagy in axons apparently serves as a stress response that protects the cell soma [193, 194, 241, 262].
Macroautophagic segregation and degradation, or autophagy proteins, can have both antiviral and proviral roles, depending on the tissue and virus type [49, 259]. It has been proposed that in postmitotic cells like neurons, macroautophagy, rather than an interferon response, plays the predominant role in controlling viral infection [258]. Several studies support a vital role for macroautophagy in antiviral defense in neurons. Because neurons do not divide and thus cannot replace dead cells, they need non-cytolytic mechanisms to control viral replication. Deficiency in the autophagy gene Atg5 or autophagic cargo receptor gene Sqstm1 accelerated Sindbis virus-induced cell death in mouse brain without affecting virus replication [176]. The results suggest that SQSTM1 targets the viral particles for autophagic degradation. Yordy and coworkers studied neurotropic herpes simplex type 1 virus infection in dorsal root ganglion (DRG) neurons. They showed that in neuronal DRG cells—in contrast to mucosal epithelial cells and other renewable cells—little type I interferon was produced, nor was interferon treatment able to block viral replication or to induce interferon-primed cell death. Instead, in DRG neurons macroautophagy was required to restrict viral replication [257]. Viruses have also developed ways to resist the macroautophagic defense system. Herpes simplex virus produces the protein ICP34.5 that binds to the autophagy protein beclin 1 and prevents its function in macroautophagy. Viral mutants that lack this beclin 1 inhibitor were less infective in a mouse intracerebral infection model [175]. Viruses can also exploit the macroautophagy pathway or individual autophagy proteins for their replication [49]. This was first discovered for poliovirus that uses macroautophagy for both replication and nonlytic release [96, 189]. Positive-stranded RNA viruses like poliovirus utilize autophagosome maturation into acidic autolysosomes to promote their replication.
There are also situations where macroautophagy may enhance neuronal degeneration. One example is degeneration as observed in the rat optic nerve. In this case, inhibition of macroautophagy by 3-methyladenine (an inhibitor of PI3KC3) attenuated acute axonal degeneration after a lesion caused by crushing the optic nerve [114]. Another example of possibly harmful macroautophagy is cerebral ischemia and ischemic brain damage. The induction of macroautophagy after cerebral ischemia has been observed in numerous studies, but the role of macroautophagy in this pathological process is under debate: inhibition of macroautophagy has been reported to have both beneficial and harmful effects for neurons [7, 244, 252]. The effect of macroautophagy in neurons may be influenced by various physiological and pathological conditions, such as the age of the patient, affected brain region, severity of the insult, and stage of ischemia. A recent study showed that macroautophagy may have different roles during cerebral ischemia and subsequent reperfusion [267]. During permanent ischemia, inhibition of macroautophagy reduced brain and cell damage. However, during reperfusion, inhibition of macroautophagy increased both cell injury in vitro and brain injury in vivo. The neuroprotection during reperfusion was mediated by increased mitophagy [267].
Increasing evidence suggest that CMA can also protect neurons from stress and injury. CMA is upregulated during oxidative stress, and oxidized proteins are more readily transported into lysosomes than non-oxidized proteins [110]. Transcriptional upregulation of LAMP-2A levels was shown to be the mechanism of the increased CMA. Cells unable to upregulate CMA during oxidative stress accumulate oxidative damage and show increased cell death [147]. It has been shown that LAMP-2A levels and CMA activity decline with age and this may contribute to the accumulation of oxidized proteins that is observed in most tissues in old animals [173, 264].
In addition to macroautophagy, CMA may also contribute to neuronal cell survival after hypoxic stress. A recent study showed that LAMP-2A was upregulated in the ischemic hemisphere after brain ischemia induced by occlusion of the middle cerebral artery. In addition, hypoxia increased LAMP-2A levels and activated CMA in neuronal cell cultures. Activation of CMA by drug treatment reduced hypoxia-mediated cell death, while silencing LAMP-2A increased it [48].
CMA was also shown to act neuroprotectively by regulating the activity of the myocyte enhancer factor 2D (MEF2D), a transcription factor required for neuronal survival [253]. MEF2D was shown to continuously shuttle from the nucleus to the cytoplasm where it interacts with HSC70 and is degraded by CMA. Inhibition of CMA was shown to lead to the accumulation of inactive MEF2D in the cytoplasm and depletion of active MEF2D from the nucleus. It was also shown that wild-type α-synuclein disrupts CMA-mediated degradation of MEF2D, which could explain why both wild-type and mutant α-synuclein can cause Parkinson’s disease.
Role of macroautophagy in neuronal development and neurogenesis
Emerging evidence supports a role for macroautophagy in neuronal development and neurodevelopmental disorders [126]. AMBRA1 is a component of the PI3KC3 complex that functions during autophagosome formation (Fig. 2). Ambra1 is a highly conserved vertebrate-specific gene, and it is highly expressed in the neuroepithelium during early neurogenesis. Functional knockout of AMBRA1 in mice showed that this protein plays a role in the development of the nervous system. Embryos deficient in Ambra1 died before birth and showed severe defects in the neural tube, including impaired macroautophagy, accumulation of ubiquitinated proteins, and unbalanced cell proliferation followed by excessive apoptosis [60]. Interestingly, mice heterozygous for Ambra1 showed an autism-like phenotype that was only detected in female animals [45]. Autism is considered to be associated with defects in neurodevelopment. Vazquez and colleagues [243] studied the role of autophagy proteins in neurogenesis of neural stem cells. The expression of autophagy genes Atg7, Becn1, Ambra1 and Lc3 increased in mouse embryonic olfactory bulb during the initial neuronal differentiation. In addition, macroautophagy was increased during neuronal differentiation in cultured stem cells derived from mouse olfactory bulb, and inhibition of macroautophagy by drugs or deletion of autophagy proteins decreased neurogenesis. The results also suggest that neural stem cells activate macroautophagy in order to fulfill their energy demands that are high due to remodeling of the cell shape and cytoskeleton [243].
Aburto and colleagues [1] investigated the role of macroautophagy in otic neurogenesis in chicken inner ear. They showed that autophagy genes Becn1, Atg5 and Lc3B were expressed and autophagic vesicles were present in the otic neurogenic zone. Inhibiting macroautophagy by antisense morpholinos or by drugs caused aberrant morphology, accumulation of apoptotic cells, impaired neurogenesis and poor axonal outgrowth. The results also suggest that macroautophagy provides the energy for the clearance of dying neuroepithelial cells and migration of otic neuronal precursors [1].
Role of autophagy in neuronal aging and neuronal diseases
In this review we will give a short overview of our current knowledge on the role of autophagy in aging neurons, and focus on the role of autophagy in four types of neurodegenerative diseases, i.e., amyotrophic lateral sclerosis and frontotemporal dementia, prion diseases, lysosomal storage diseases, and Parkinson’s disease. Other neuronal diseases where autophagy has been shown to play a role are discussed in the other review articles of this cluster.
Autophagy in aging neurons
As stated above, removal of harmful oxidated and aggregate-prone proteins and damaged organelles that produce oxidative damage is particularly important for postmitotic neuronal cells. Removal of damaged mitochondria by mitophagy is vital for cell survival, because damaged mitochondria can produce ROS and induce harmful events including protein carbonylation, lipid peroxidation and DNA damage. Further, damaged mitochondria can trigger apoptosis by releasing cytochrome c [236]. Thus, is not surprising that both macroautophagy and CMA have been shown to delay aging [27, 38]. Promotion of macroautophagy has been shown to extend the lifespan of many model organisms including yeast, worms, flies, and mice [86]. In these studies, macroautophagy has been increased either by genetic manipulations (e.g., overexpression of SIRT1) or by drugs (e.g., rapamycin, resveratrol and spermidine). Further, studies with several model organisms have also shown that single-gene mutations, e.g., in the insulin and mTOR signaling pathways, can extend lifespan, and many of these mutations also induce macroautophagy [74].
The neurological phenotypes of macroautophagy-deficient animals, described above, reflect at least in part the phenotype of normal aging, including accumulation of the aging pigment lipofuscin, aggregates consisting of ubiquitinated substrates and autophagy cargo receptors, and dysfunctional mitochondria. These observations are highly suggestive of a pivotal role for autophagy in the prevention of aging [13, 221]. The rate of macroautophagy decreases with age in several cell types including the brain cells [131], even though differences between various brain regions are described, e.g., for the expression of Ambra1 in mouse brain [208]. In addition to these transcriptional alterations, decreased lysosome–autophagosome fusion events also contribute to the inefficient clearance of macroautophagic substrates in aging tissues. Extremely reactive hydroxyl radicals are formed during the degradation of iron-containing metalloproteins and ROS-generating mitochondria in lysosomes under acidic and reducing conditions (partly due to the presence of the reducing amino acid cysteine). Hydroxyl radicals generate aldehydes which react with free amino groups within proteins, crosslinking them tightly, which ultimately leads to the formation of non-degradable, insoluble lipofuscin aggregates (reviewed in [21]), which can fill up to 75 % of some brain neurons [232]. These abundant lipofuscin aggregates are likely to contribute to the impairment of the lysosome–autophagosome fusion process and the degradative capacity of lysosomes and autolysosomes.
As stated above, both macroautophagy and CMA have been shown to delay aging [27, 38]. Similar to macroautophagy, CMA levels decrease with aging [35]. In the liver, the decrease in CMA was assigned to decreased levels of the CMA receptor LAMP-2A, due to apparent loss of its stability with aging [111, 195]. Whether this mechanism is also responsible for decreased CMA in neurons remains to be determined. CMA can be involved in neurodegeneration via two alternative mechanisms [38]. The first possibility is that CMA contributes to the degradation of the pathogenic protein, and the age-related or other type of decline in CMA causes or contributes to disease. The second possibility is that the pathogenic protein blocks CMA, for example by irreversibly binding to the translocation complex. In both scenarios, the decline of CMA leads to the accumulation of all CMA substrates, including physiological and pathological proteins.
Amyotrophic lateral sclerosis and frontotemporal dementia
Frontotemporal dementia (FTD) and amyotrophic lateral sclerosis (ALS) are diseases in which a large subset of patients typically accumulate cytosolic aggregates consisting of the Tar-DNA binding protein 43 (TDP-43). TDP-43 is degraded by the ubiquitin proteasome system (UPS) and macroautophagy. In addition, it was also shown to be degraded by CMA, mediated through interaction between HSC70 and ubiquitylated TDP-43 [91].
Pathogenic mutations in several autophagy-related genes were identified in a small subgroup of patients suffering from FTD and ALS, including mutations in cargo receptors like UBQLN2, a gene encoding the ubiquitin-like protein ubiquilin 2 (UBQLN2). Like the autophagic cargo receptors described above, this protein directly interacts with LC3 and contains an ubiquitin-binding domain (UBA) that binds polyubiquitinated proteins [198]. UBQLN2 mutations were found in patients presenting with ALS/FTD symptoms, and mutated ubiquilin 2 co-localized with aggregated proteins in neurons of the hippocampus and the spinal cord [43]. Another cargo receptor protein mutated in rare familial ALS cases is optineurin (OPTN). Sequencing of a cohort of 371 FTD patients failed to detect any mutations, indicating that OPTN mutations are only seen in ALS [197]. Similar to UBQLN2, OPTN directly binds to LC3 via an interacting motif and ubiquitin via an independent ubiquitin interacting motif [247, 268]. In addition to their autophagy cargo receptor function, both UBQLN2 and OPTN act in the degradation of substrates via the ubiquitin–proteasome system. The third cargo receptor protein found to be mutated in a subset of ~3 % of FTD and ALS patients is SQSTM1 [56, 199], which preferentially binds Lys63-polyubiquitinated aggregates [17, 18, 207]. It seems to act predominantly in the macroautophagic degradation, since most autophagy gene knockout mouse models show a profound accumulation of ubiquitin and SQSTM1-positive aggregates especially in neurons [33, 118, 128].
Another gene mutated in TDP-43 proteinopathies is VCP, which encodes the valosin-containing protein (VCP). VCP is a member of the AAA+-ATPase (ATPase associated with diverse cellular activities) protein super family. Mutations in VCP have been documented on both arms of the ALS and FTD disease continuum [75, 101], indicating essential function in neurons. In the proteasomal degradation pathway of misfolded proteins, VCP binds to polyubiquitinated proteins and extracts them from the membrane of the endoplasmic reticulum in an ATP-dependent manner to facilitate delivery to the proteasome [152]. In addition, mutations in VCP lead to an accumulation of nondegradative autophagosomes due to impaired fusion of autophagosomes with lysosomes [102]. As described earlier, VCP was shown to be essential for the fusion of ubiquitin-containing, basal autophagosomes with proteolytically active acidified vesicles [234]. These data suggest important functions for VCP in both macroautophagy and proteasomal degradation. With regard to TDP-43 proteinopathies, another intriguing function of VCP is its role in the degradation of stress granules, which are commonly observed in neurons of patients with neurodegenerative diseases. Stress granules are composed of cytoplasmic mRNA and RNA-binding proteins (such as TDP-43), which become sequestered by macroautophagy. VCP was shown to be essential for this process and it was suggested that it might act in the disassembly and/or targeting of stress granules to macroautophagy [22]. However, the mechanisms that lead to inclusions of TDP-43 in VCP mutant neurons need to be determined. Interestingly, VCP mutations directly affect mTOR-mediated nutrient sensing. Mutations in VCP or inhibition of VCP lead to a failure in mTOR activation upon nutrient stimulation, even though the precise mechanism remains unclear [30]. However, this additional regulatory function implicates a complex interplay between degradative pathways in the cell.
Rare mutations in charged multivesicular protein 2B (CHMP2B) have been described in FTD patients [22, 179, 214] and also in some ALS patients [34]. CHMP2B is part of the endosomal sorting complex required for transport-III (ESCRT-III), which is needed for efficient sorting of ubiquitinated integral membrane proteins into the internal vesicles of multivesicular bodies (MVBs), and the formation of MVBs in general. Moreover, MVBs fuse with autophagosomes and were shown to be essential for the degradation of macroautophagic cargo and the macroautophagic clearance of protein aggregates, including TDP-43 aggregates [59, 123, 206]. These findings suggest that CHMP2B is an essential player in the macroautophagic pathway.
One-fifth of familial ALS cases bear gain-of-function mutations in SOD1, leading to accumulation of aggregated superoxide dismutase 1 (SOD1) in the cytosol of motor neurons. In neuronal cells, SOD1 was shown to be degraded by both the proteasomal pathway and macroautophagy [106]. Even though SOD1 apparently does not directly act in the autophagic pathway, autophagic alterations were observed in patients and mouse models with mutated SOD1 [70, 71, 79, 154]. How aggregated SOD1 leads to inhibition of autophagic flux is still unknown, but impairment of vesicular trafficking, heterotypic organelle fusion and lysosomal function might contribute to the pathology of ALS [266]. Of note, induction of macroautophagy by rapamycin augmented neuronal cell death and decreased the life span of SOD1 mutant mice, suggesting that increasing the uptake of macroautophagic cargo in the context of impaired clearance is harmful [266]. In contrast, a previous study in which lithium was applied to induce macroautophagy in ALS mice and patients, revealed a beneficial effect of this treatment [61]. It is unknown whether this positive effect was due to different signaling routes induced by lithium and rapamycin, or to side effects of lithium unrelated to macroautophagy induction [62]. Interestingly, when trehalose was used to induce macroautophagy in an mTOR-independent manner, the treatment prolonged the life span of mutant SOD1 transgenic mice and attenuated the progression of the disease [28]. The complexity of macroautophagy modulation in the context of ALS is highlighted by a recent study using a transgenic ALS mouse model expressing mutant SOD1 in a beclin 1 heterozygous background. Surprisingly, even though lower beclin 1 levels limited the autophagic flux, these mice had a significantly increased lifespan, suggesting that beclin 1 has a pathogenic role in ALS [163]. Nevertheless, beclin 1 was shown to have a functional role in the macroautophagic clearance of mutant SOD1 in cultured motoneurons. The molecular mechanism underlying the unexpected results with beclin 1 heterozygous ALS mice was suggested to include an abnormal interaction of mutant SOD1 with beclin1, mediated by the macroautophagy inhibitory protein BCL2L1, which may alter macroautophagy stimulation [163].
Similar to ALS, most patients with familial FTD harbor mutations in genes that are apparently unrelated to autophagy, including the frequent mutations in GRN (encoding progranulin), MAPT (encoding tau) and hexanucleotide repeat expansion in C9ORF72. In the case of tau, a proteolytic tau-fragment was shown to interact with HSC70 facilitating its binding to the CMA receptor LAMP-2A. However, in contrast to typical CMA substrates, these forms of tau are not translocated into the lysosomal lumen by CMA. Instead, association of tau to lysosomes through the CMA machinery seems to contribute to the cleavage of this mutant form of tau and the organization of the protein into oligomeric complexes [242]. It remains to be determined whether this association of tau with CMA is relevant in the pathogenesis of FTD.
Rare FTD cases also bear mutations in TARDBP gene (encoding TDP-43 and causing frontotemporal lobar degeneration of type FTLD-TDP) and FUS (encoding fused in sarcoma protein) [230]. Whether autophagy is impaired in these patients is currently unknown. However, the findings that certain pathogenic TDP-43 species can bind SQSTM1 and become degraded via macroautophagy further strengthen findings that autophagy might play a role in the degradation of the aggregating TDP-43 species [20, 227]. Moreover, in contrast to the SOD1 mouse model (see above), a mouse model for FTLD-TDP showed beneficial effects of rapamycin treatment on TDP-43 aggregation, neuronal survival and motor dysfunction [240]. These results support a role for dysfunctional macroautophagy in the pathogenesis of TDP-43 proteinopathies.
Transmissible spongiform encephalopathies (prion diseases)
Prion diseases are transmissible neurodegenerative disorders characterized by an accumulation of misfolded pathogenic isoforms (PrPSc) of normal prion protein (PrPC) [183]. Autophagy has been suggested to mediate the degradation of PrPSc and/or PrPC at least to some extent [2, 88, 250]. As mentioned above, induction of macroautophagy by SIRT1 overexpression prevented prion peptide neurotoxicity, while prevention of macroautophagy by knockdown of Atg5 abolished the SIRT1-induced neuroprotection [98]. In a recent study, macroautophagy induction by caffeine was shown to reduce neurotoxicity of human PrPSc in neuronal cells [157]. Both prion disease mouse models and human patients exhibit increased steady-state numbers of autophagosomes in neurons [130, 203, 212]. Several approaches have been used to test the effect of macroautophagy induction in prion diseases. Imatinib, an autophagy-inducing cancer drug, delayed the disease onset and appearance of PrPSc in an induced scrapie mouse model [263], and rapamycin and lithium modestly extended survival in scrapie infected mice [87]. In a genetic prion disease model, rapamycin delayed the disease onset and prevented plaque formation [32]. In contrast, trehalose did not change the course of the disease in prion-infected mice [2]. Taken together, these results suggest that macroautophagy is involved in the clearance of prion proteins in neurons, and that induction of macroautophagy is a potential treatment for prion diseases.
Lysosomal storage disorders
Lysosomal storage disorders (LSDs) are caused by deficiencies in lysosomal hydrolases or membrane proteins [181]. LSDs are characterized by the intralysosomal accumulation of different types of substrates and an impairment of endocytic trafficking pathways [9]. Since autophagic cargo is degraded by lysosomal hydrolases, a defect in the autophagic flux is also observed in many LSDs. This is demonstrated by the observations showing that a deficiency of the major lysosomal membrane protein LAMP-2 in mice [226] and man [166] leads to an accumulation of autophagic vacuoles due to an impaired lysosomal/autophagosomal fusion process [53]. After the fusion of lysosomes and autophagosomes, an efficient lysosomal degradation is necessary for successful completion of the autophagic flux. For example, deficiencies in the major lysosomal proteinases, cathepsin D or cathepsins B and L, in mice cause a massive neuronal accumulation of autophagic structures [57, 115], leading to neuronal ceroid lipofuscinosis (NCL) and severe neurological symptoms. In humans, deficiency of cathepsin D causes type 10 NCL (CLN10) and that of cathepsin F results in CLN13.
The findings in cathepsin-deficient mice, but also in other NCL models (e.g., CLN2, CLN3, and CLN6 deficient mice) underline the major role of macroautophagy in the pathogenesis of many NCL diseases [26, 153, 229]. A block of autophagic flux is a characteristic feature in several other lysosomal storage disorders, due to either defective autophagosome–lysosome fusion or defective lysosomal degradation, or both. As a consequence, polyubiquitinated protein aggregates and dysfunctional mitochondria accumulate in neurons, which is thought to cause cell death [17]. As an example of this model, an impaired basal macroautophagy associated with severe neurodegeneration has been revealed in the brain of mucolipidosis type II (ML-II) mouse model [116]. ML-II (also called I-cell disease) results from defective intracellular targeting of mannose-6 phosphate tagged lysosomal hydrolases due to a deficiency of the N-acetylglucosamine-1-phosphotransferase. Autophagic dysfunction is also evident in mucolipidosis type IV patients and mice [40, 246]. Mutations in the lysosomal cation channel mucolipin 1, the protein mutated in mucolipidosis type IV, led to increased autophagosome formation and decreased autophagosome–lysosome fusion in mucolipidosis IV patient fibroblasts. Another study showed that the impaired autophagic clearance in mucolipidosis type IV fibroblasts was caused at least in part by impaired CMA, which was due to reduced level of LAMP-2A in the lysosomal membrane [245]. However, in mucolipin 1-deficient mouse neurons, autophagic dysfunction was assigned to impaired macroautophagic clearance [40]. These at first glance conflicting data may be due to different roles of macroautophagy and CMA in different cell types.
It is not understood how accumulation of lysosomal cargo or mutations in lysosomal membrane proteins cause the defects in the fusion of lysosomes and autophagosomes, but different mechanisms have been proposed. One possibility is that the defects are at least partly due to a secondary accumulation of cholesterol inside endosomes and lysosomes, which is observed in many LSDs [237]. It is possible that in LSDs with neuronal involvement, this secondary accumulation of cholesterol in endosomal and lysosomal contents and/or membranes contributes to the reduced fusion capacity. How this cholesterol imbalance occurs is not understood, but it is possible that an impaired lysosomal cholesterol export contributes to it. Interestingly, Fraldi and colleagues [64] showed that in two severe neurodegenerative LSDs, SNAREs (soluble N-ethyl-maleimide-sensitive factor attachment protein receptors) were abnormally localized in the lysosomal membrane, with a preference for the cholesterol-rich membrane domains. It was proposed that this causes an impaired SNARE sorting and recycling, preventing efficient lysosomal fusion. The role of a defective cholesterol homeostasis is also supported by the observations in Niemann–Pick type C (NPC) disease, in which cholesterol accumulates in endosomes and lysosomes due to mutations in NPC1 or NPC2, proteins that function in lysosomal cholesterol export. An early neuronal autophagic vacuolation is observed in NPC disease, which is connected with neurodegeneration [129]. In agreement with this, a recent study with induced pluripotent stem cells showed that NPC1 function is needed for macroautophagy [140].
One possibility to overcome the problems associated with an impaired autophagic flux in LSDs was revealed in studies analyzing glycogen storage disease type II (Pompe disease), an LSD due to lack of α-glucosidase (acid maltase) activity affecting mainly skeletal muscle and heart. It was shown that increasing the expression of TFEB significantly improved the pathological alterations in this specific LSD. TFEB increased lysosome biogenesis and exocytosis of autophagosomes, which both contributed to the removal of the excessive accumulation of autophagic cargo in cells [68, 216]. Increased expression of TFEB can be anticipated to work in a similar beneficial manner in LSDs with CNS involvement. Interestingly, the effect of two drugs, 2-hydroxypropyl-β-cyclodextrin and genistein, used for therapeutic intervention in mucopolysaccharidoses and NPC disease, was recently shown to depend at least in part on the TFEB-mediated modulation of the expression of lysosomal and autophagic genes [159, 215]. Clearly, more experiments are needed to evaluate, whether this approach is also beneficial in neurological LSDs, and whether the increased appearance of storage material in the extracellular space due to autophagosomal exocytosis is well tolerated in neuronal tissues.
Damaged lysosomes, which may occur in LSDs, may also be a substrate for macroautophagy [92, 139], and this autophagy of lysosomes was shown to be important for the recovery of lysosomal degradation capacity. Therefore, it is possible that lysosomal storage causes trouble from both ends, i.e., both by inhibition of lysosome–autophagosome fusion and by reduced autophagic clearance of storage lysosomes.
Parkinson’s disease
Increasing evidence shows that lysosomal and autophagosomal function and the development of Parkinson’s disease (PD) are closely linked [137]. Cytosolic α-synuclein accumulation and associated neurotoxicity are major hallmarks of this disease [218], and mutations within the α-synuclein gene are linked to PD [182]. Both CMA and macroautophagy have been linked to PD pathogenesis [38, 141]. α-Synuclein contains the KFERQ-like motif that acts as a zip code for CMA, and CMA has been reported to participate in the degradation of α-synuclein [249]. Mutated α-synuclein was suggested to inhibit the transport of CMA substrates into lysosomes, possibly by blocking the binding sites in the CMA receptor LAMP-2A [146]. This block can lead to abnormal cytoplasmic accumulation of α-synuclein and other CMA substrates [37]. Neurons expressing mutated α-synuclein also show accumulation of autophagosomes, defective dopamine release and higher susceptibility to autophagic cell death [220]. An increase in macroautophagy accompanied α-synuclein accumulations in neuronal cells, and cell death was reduced in mutant α-synuclein expressing cells when ATG5 was silenced, suggesting that macroautophagy contributed to cell death (Fig. 4d in [251]). The upregulated macroautophagy was proposed to be a consequence of the impaired CMA. However, other studies have shown that macroautophagy is beneficial for cells expressing mutant α-synuclein. Induction of macroautophagy by overexpression of beclin 1 and rapamycin treatment could rescue α-synuclein degradation in neuronal cell lines overexpressing α-synuclein [217]. Lentivirus-mediated overexpression of beclin 1 also decreased the PD pathology in α-synuclein transgenic mice. These results suggest that macroautophagy also contributes to degradation of α-synuclein, and that induction of macroautophagy may be a potential treatment for diseases associated with α-synuclein accumulation. In agreement with this proposition, increased expression of wild-type α-synuclein negatively influences the rate of macroautophagy in cell lines and mice. This effect is mediated by impaired activity of RAB1a and mislocalization of the autophagy protein ATG9, which are needed for the formation of autophagosomes [248].
CMA may also be involved in PD caused by mutations in other proteins. Leucine-rich repeat kinase 2 (LRRK2), the mutations of which cause familial PD, was recently shown to be a CMA substrate [174]. Mutant LRRK2 was observed to interfere the function of the CMA translocation complex on the lysosomal membrane. Thus, mutant LRRK2 may inhibit the degradation of CMA substrates including α-synuclein. Similar observations were published for another PD-associated CMA substrate protein, the ubiquitin C-terminal hydrolase L1 [105].
Neurons are heavily dependent on mitochondrial ATP production. Dysfunctional mitochondria have been reported to accumulate in the brains of PD patients, and mitochondrial defects, implicated as reduced bioenergetic capacity, oxidative stress and reduced stress resistance, are observed in several PD models [29]. In agreement with these observations, mutations in mitophagy and mitochondrial quality control genes PARK2, PINK1 and DJ-1 have been associated with autosomal recessive PD [29, 50, 188].
Mutations that lead to impaired lysosomal degradation capacity have also been linked to synucleinopathies. Mutations in the vesicular P type ATPase ATP13A2 are associated with autosomal recessive familial PD. Disease-causing mutations impair lysosomal acidification and proteolytic activity, thereby also decreasing autophagic flux and clearance of α-synuclein aggregates [42]. Mutations within the gene encoding the lysosomal sphingolipid degrading hydrolase β-glucocerebrosidase influence lysosomal protein degradation and clearance of α-synuclein [39, 201] and are one of the genetic risk factors of synucleinopathies including PD and Lewy body disease. Overexpression of β-glucocerebrosidase in the central nervous system of mice expressing mutant α-synuclein decreased the levels of α-synuclein, suggesting that this enzyme may be linked to the clearance of α-synuclein. A recent study showed that parkin/PARK2 and β-glucocerebrosidase interact with each other, and that parkin mediates ubiquitination of mutant β-glucocerebrosidase, which facilitates its degradation in proteasomes. Further, mutant β-glucocerebrosidase competes with other substrates of parkin such as PARIS and ARTS proteins, whose accumulation triggers apoptosis [15].
Conclusions and open questions
The importance of macroautophagy as a central degradative pathway in nutrient homeostasis, clearance of damaged organelles and aggregate-prone substrates in postmitotic cells like neurons is highlighted by mice with neuron-specific autophagy deficiencies. An impaired autophagic function has also been reported in many neurodegenerative diseases. There is also increasing evidence showing that CMA dysfunction contributes to neurodegeneration. However, even though major advances have been made to describe functions and dysfunctions of macroautophagy and CMA in neurodegenerative disorders, we are far from understanding their exact roles in different diseases. In a disease context it is not always clear whether autophagy is beneficial or detrimental in an affected neuron. Thus, it is not always certain whether an induction or inhibition of autophagy would serve as a therapeutic option. The conflicting results in many of the initial studies applying autophagy modulating agents underline this lack of knowledge and call for more detailed and thorough investigation of the roles of autophagy in neurons.
Abbreviations
- AAA+-ATPase:
-
ATPase associated with diverse cellular activities
- ALS:
-
Amyotrophic lateral sclerosis
- AMBRA1:
-
Activating molecule in beclin 1-regulated autophagy
- AMPK:
-
AMP-activated protein kinase
- ATF6:
-
Activating transcription factor 6
- ATG:
-
Autophagy-related
- BECN1:
-
Beclin 1
- BNIP3:
-
BCL2/adenovirus E1B 19 kDa interacting protein 3
- CHMP2B:
-
Charged multivesicular protein 2B
- CMA:
-
Chaperone-mediated autophagy
- CNS:
-
Central nervous system
- DAP1:
-
Death-associated protein 1
- DFCP1:
-
Double FYVE-containing protein 1
- DRG:
-
Dorsal root ganglion
- ESCRT-III:
-
Endosomal sorting complex required for transport-III
- ER:
-
Endoplasmic reticulum
- ERK2:
-
Extracellular signal-regulated kinase 2
- FIP200:
-
200 kDa FAK family kinase-interacting protein
- FTD:
-
Frontotemporal dementia
- GFP:
-
Green fluorescent protein
- GTPase:
-
Guanosine triphosphatase
- HIF:
-
Hypoxia-inducible factor
- HSC70:
-
Heat shock cognate protein of 70 kDa
- IRE1:
-
Serine/threonine-protein kinase/endoribonuclease protein
- LAMP:
-
Lysosomal associated membrane protein
- LC3/MAP1-LC3:
-
Microtubule associated protein 1 light chain 3
- LRRK2:
-
Leucine-rich repeat kinase 2
- LSD:
-
Lysosomal storage disorder
- MEF2D:
-
Myocyte enhancer factor 2D
- ML-II:
-
Mucolipidosis type II
- NCL:
-
Neuronal ceroid lipofuscinosis
- NPC:
-
Niemann–Pick C
- mTOR:
-
Mammalian target of rapamycin
- mTORC1:
-
mTOR complex 1
- MVB:
-
Multivesicular body
- OPTN:
-
Optineurin
- PD:
-
Parkinson’s disease
- PERK:
-
Protein kinase-like ER kinase
- PI3P:
-
Phosphatidyl inositol 3-phosphate
- PI3KC3:
-
Phosphatidyl inositol 3-kinase class 3
- PKC:
-
Protein kinase C
- POMC:
-
Pro-opiomelanocortin
- RAPTOR:
-
Regulatory associated protein of mTOR)
- RB1CC1:
-
RB1-inducible coiled-coil 1
- REST:
-
Repressor element 1 silencing transcription factor
- Rheb:
-
Ras homolog enriched in brain
- ROS:
-
Reactive oxygen species
- SNARE:
-
Soluble N-ethyl-maleimide-sensitive factor attachment protein receptor
- SOD:
-
Superoxide dismutase
- SQSTM1:
-
Sequestosome 1
- TDP-43:
-
Tar-DNA binding protein 43
- TFEB:
-
Transcription factor EB
- TSC:
-
Tuberous sclerosis complex
- UBA:
-
Ubiquitin-binding domain
- UBQLN2:
-
Ubiquilin 2
- ULK1:
-
Unc-51-like kinase 1
- UPR:
-
Unfolded protein response
- V-ATPase:
-
Vacuolar ATPase
- VCP:
-
Valosin-containing protein
- WIPI:
-
WD repeat domain, phosphoinositide interacting
References
Aburto MR, Sanchez-Calderon H, Hurle JM, Varela-Nieto I, Magarinos M (2012) Early otic development depends on autophagy for apoptotic cell clearance and neural differentiation. Cell Death Dis 3:e394. doi:10.1038/cddis.2012.132
Aguib Y, Heiseke A, Gilch S, Riemer C, Baier M, Schatzl HM, Ertmer A (2009) Autophagy induction by trehalose counteracts cellular prion infection. Autophagy 5(3):361–369
Ahmed I, Liang YD, Schools S, Dawson VL, Dawson TM, Savitt JM (2012) Development and characterization of a new Parkinson’s disease model resulting from impaired autophagy. J Neurosci 32(46):16503–16509. doi:10.1523/Jneurosci.0209-12.2012
Alers S, Loffler AS, Wesselborg S, Stork B (2012) Role of AMPK-mTOR-Ulk1/2 in the regulation of autophagy: cross talk, shortcuts, and feedbacks. Mol Cell Biol 32(1):2–11. doi:10.1128/Mcb.06159-11
Alirezaei M, Kemball CC, Flynn CT, Wood MR, Whitton JL, Kiosses WB (2010) Short-term fasting induces profound neuronal autophagy. Autophagy 6(6):702–710
Axe EL, Walker SA, Manifava M, Chandra P, Roderick HL, Habermann A et al (2008) Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum. J Cell Biol 182(4):685–701
Baek SH, Noh AR, Kim KA, Akram M, Shin YJ, Kim ES et al (2014) Modulation of mitochondrial function and autophagy mediates carnosine neuroprotection against ischemic brain damage. Stroke 45(8):2438–2443. doi:10.1161/STROKEAHA.114.005183
Bains M, Zaegel V, Mize-Berge J, Heidenreich KA (2011) IGF-I stimulates Rab7-RILP interaction during neuronal autophagy. Neurosci Lett 488(2):112–117. doi:10.1016/j.neulet.2010.09.018
Ballabio A, Gieselmann V (2009) Lysosomal disorders: from storage to cellular damage. Biochim Biophys Acta 1793(4):684–696. doi:10.1016/j.bbamcr.2008.12.001
Ban BK, Jun MH, Ryu HH, Jang DJ, Ahmad ST, Lee JA (2013) Autophagy negatively regulates early axon growth in cortical neurons. Mol Cell Biol 33(19):3907–3919. doi:10.1128/MCB.00627-13
Bandyopadhyay U, Kaushik S, Varticovski L, Cuervo AM (2008) The chaperone-mediated autophagy receptor organizes in dynamic protein complexes at the lysosomal membrane. Mol Cell Biol 28(18):5747–5763. doi:10.1128/MCB.02070-07
Bandyopadhyay U, Sridhar S, Kaushik S, Kiffin R, Cuervo AM (2010) Identification of regulators of chaperone-mediated autophagy. Mol Cell 39(4):535–547. doi:10.1016/j.molcel.2010.08.004
Bartlett BJ, Isakson P, Lewerenz J, Sanchez H, Kotzebue RW, Cumming RC et al (2011) p62, Ref(2)P and ubiquitinated proteins are conserved markers of neuronal aging, aggregate formation and progressive autophagic defects. Autophagy 7(6):572–583
Batlevi Y, La Spada AR (2011) Mitochondrial autophagy in neural function, neurodegenerative disease, neuron cell death, and aging. Neurobiol Dis 43(1):46–51. doi:10.1016/j.nbd.2010.09.009
Bendikov-Bar I, Rapaport D, Larisch S, Horowitz M (2014) Parkin-mediated ubiquitination of mutant glucocerebrosidase leads to competition with its substrates PARIS and ARTS. Orphanet J Rare Dis 9(1):86. doi:10.1186/1750-1172-9-86
Bento CF, Puri C, Moreau K, Rubinsztein DC (2013) The role of membrane-trafficking small GTPases in the regulation of autophagy. J Cell Sci 126(Pt 5):1059–1069. doi:10.1242/jcs.123075
Bjorkoy G, Lamark T, Brech A, Outzen H, Perander M, Overvatn A et al (2005) p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J Cell Biol 171(4):603–614. doi:10.1083/jcb.200507002
Bjorkoy G, Lamark T, Johansen T (2006) p62/SQSTM1: a missing link between protein aggregates and the autophagy machinery. Autophagy 2(2):138–139
Boya P, Reggiori F, Codogno P (2013) Emerging regulation and functions of autophagy. Nat Cell Biol 15(7):713–720
Brady OA, Meng P, Zheng Y, Mao Y, Hu F (2011) Regulation of TDP-43 aggregation by phosphorylation and p62/SQSTM1. J Neurochem 116(2):248–259. doi:10.1111/j.1471-4159.2010.07098.x
Brunk UT, Terman A (2002) The mitochondrial-lysosomal axis theory of aging: accumulation of damaged mitochondria as a result of imperfect autophagocytosis. Eur J Biochem 269(8):1996–2002
Buchan JR, Kolaitis RM, Taylor JP, Parker R (2013) Eukaryotic stress granules are cleared by autophagy and Cdc48/VCP function. Cell 153(7):1461–1474. doi:10.1016/j.cell.2013.05.037
Bunge MB (1973) Fine structure of nerve fibers and growth cones of isolated sympathetic neurons in culture. J Cell Biol 56(3):713–735
Byun YJ, Kim SK, Kim YM, Chae GT, Jeong SW, Lee SB (2009) Hydrogen peroxide induces autophagic cell death in C6 glioma cells via BNIP3-mediated suppression of the mTOR pathway. Neurosci Lett 461(2):131–135. doi:10.1016/j.neulet.2009.06.011
Caberlotto L, Nguyen TP (2014) A systems biology investigation of neurodegenerative dementia reveals a pivotal role of autophagy. BMC Syst Biol 8(1):65. doi:10.1186/1752-0509-8-65
Cao Y, Espinola JA, Fossale E, Massey AC, Cuervo AM, MacDonald ME, Cotman SL (2006) Autophagy is disrupted in a knock-in mouse model of juvenile neuronal ceroid lipofuscinosis. J Biol Chem 281(29):20483–20493. doi:10.1074/jbc.M602180200
Carroll B, Hewitt G, Korolchuk VI (2013) Autophagy and ageing: implications for age-related neurodegenerative diseases. Essays Biochem 55:119–131. doi:10.1042/bse0550119
Castillo K, Nassif M, Valenzuela V, Rojas F, Matus S, Mercado G et al (2013) Trehalose delays the progression of amyotrophic lateral sclerosis by enhancing autophagy in motoneurons. Autophagy 9(9):1308–1320
Celardo I, Martins LM, Gandhi S (2014) Unravelling mitochondrial pathways to Parkinson’s disease. Br J Pharmacol 171(8):1943–1957. doi:10.1111/bph.12433
Ching JK, Elizabeth SV, Ju JS, Lusk C, Pittman SK, Weihl CC (2013) mTOR dysfunction contributes to vacuolar pathology and weakness in valosin-containing protein associated inclusion body myopathy. Hum Mol Genet 22(6):1167–1179. doi:10.1093/hmg/dds524
Chua CE, Gan BQ, Tang BL (2011) Involvement of members of the Rab family and related small GTPases in autophagosome formation and maturation. Cell Mol Life Sci 68(20):3349–3358. doi:10.1007/s00018-011-0748-9
Cortes CJ, Qin K, Cook J, Solanki A, Mastrianni JA (2012) Rapamycin delays disease onset and prevents PrP plaque deposition in a mouse model of Gerstmann–Straussler–Scheinker disease. J Neurosci 32(36):12396–12405. doi:10.1523/JNEUROSCI.6189-11.2012
Coupe B, Ishii Y, Dietrich MO, Komatsu M, Horvath TL, Bouret SG (2012) Loss of autophagy in pro-opiomelanocortin neurons perturbs axon growth and causes metabolic dysregulation. Cell Metab 15(2):247–255. doi:10.1016/j.cmet.2011.12.016
Cox LE, Ferraiuolo L, Goodall EF, Heath PR, Higginbottom A, Mortiboys H et al (2010) Mutations in CHMP2B in lower motor neuron predominant amyotrophic lateral sclerosis (ALS). PLoS One 5(3):e9872. doi:10.1371/journal.pone.0009872
Cuervo AM, Dice JF (2000) Age-related decline in chaperone-mediated autophagy. J Biol Chem 275(40):31505–31513
Cuervo AM, Dice JF, Knecht E (1997) A population of rat liver lysosomes responsible for the selective uptake and degradation of cytosolic proteins. J Biol Chem 272(9):5606–5615
Cuervo AM, Stefanis L, Fredenburg R, Lansbury PT, Sulzer D (2004) Impaired degradation of mutant alpha-synuclein by chaperone-mediated autophagy. Science 305(5688):1292–1295
Cuervo AM, Wong E (2014) Chaperone-mediated autophagy: roles in disease and aging. Cell Res 24(1):92–104. doi:10.1038/cr.2013.153
Cullen V, Sardi SP, Ng J, Xu YH, Sun Y, Tomlinson JJ et al (2011) Acid beta-glucosidase mutants linked to Gaucher disease, Parkinson disease, and Lewy body dementia alter alpha-synuclein processing. Ann Neurol 69(6):940–953. doi:10.1002/ana.22400
Curcio-Morelli C, Charles FA, Micsenyi MC, Cao Y, Venugopal B, Browning MF et al (2010) Macroautophagy is defective in mucolipin-1-deficient mouse neurons. Neurobiol Dis 40(2):370–377. doi:10.1016/j.nbd.2010.06.010
d’Ydewalle C, Bogaert E, Van Den Bosch L (2012) HDAC6 at the intersection of neuroprotection and neurodegeneration. Traffic 13(6):771–779. doi:10.1111/j.1600-0854.2012.01347.x
Dehay B, Ramirez A, Martinez-Vicente M, Perier C, Canron MH, Doudnikoff E et al (2012) Loss of P-type ATPase ATP13A2/PARK9 function induces general lysosomal deficiency and leads to Parkinson disease neurodegeneration. Proc Natl Acad Sci USA 109(24):9611–9616. doi:10.1073/pnas.1112368109
Deng HX, Chen W, Hong ST, Boycott KM, Gorrie GH, Siddique N et al (2011) Mutations in UBQLN2 cause dominant X-linked juvenile and adult-onset ALS and ALS/dementia. Nature 477(7363):211–215. doi:10.1038/nature10353
Dengjel J, Hoyer-Hansen M, Nielsen MO, Eisenberg T, Harder LM, Schandorff S et al (2012) Identification of autophagosome-associated proteins and regulators by quantitative proteomic analysis and genetic screens. Mol Cell Proteomics 11(3):M111 014035. doi:10.1074/mcp.M111.014035
Dere E, Dahm L, Lu D, Hammerschmidt K, Ju A, Tantra M et al (2014) Heterozygous ambra1 deficiency in mice: a genetic trait with autism-like behavior restricted to the female gender. Front Behav Neurosci 8:181. doi:10.3389/fnbeh.2014.00181
Di Nardo A, Wertz MH, Kwiatkowski E, Tsai PT, Leech JD, Greene-Colozzi E et al (2014) Neuronal Tsc1/2 complex controls autophagy through AMPK-dependent regulation of ULK1. Hum Mol Genet 23(14):3865–3874. doi:10.1093/hmg/ddu101
Dice JF (2007) Chaperone-mediated autophagy. Autophagy 3(4):295–299
Dohi E, Tanaka S, Seki T, Miyagi T, Hide I, Takahashi T et al (2012) Hypoxic stress activates chaperone-mediated autophagy and modulates neuronal cell survival. Neurochem Int 60(4):431–442. doi:10.1016/j.neuint.2012.01.020
Dong X, Levine B (2013) Autophagy and viruses: adversaries or allies? J Innate Immun 5(5):480–493. doi:10.1159/000346388
Dupuis L (2014) Mitochondrial quality control in neurodegenerative diseases. Biochimie 100:177–183. doi:10.1016/j.biochi.2013.07.033
Ejlerskov P, Rasmussen I, Nielsen TT, Bergstrom AL, Tohyama Y, Jensen PH, Vilhardt F (2013) Tubulin polymerization-promoting protein (TPPP/p25alpha) promotes unconventional secretion of alpha-synuclein through exophagy by impairing autophagosome–lysosome fusion. J Biol Chem 288(24):17313–17335. doi:10.1074/jbc.M112.401174
Eskelinen EL (2005) Maturation of autophagic vacuoles in mammalian cells. Autophagy 1:1–10
Eskelinen EL, Illert AL, Tanaka Y, Schwarzmann G, Blanz J, Von Figura K, Saftig P (2002) Role of LAMP-2 in lysosome biogenesis and autophagy. Mol Biol Cell 13(9):3355–3368. doi:10.1091/mbc.E02-02-0114
Eskelinen EL, Reggiori F, Baba M, Kovacs AL, Seglen PO (2011) Seeing is believing: the impact of electron microscopy on autophagy research. Autophagy 7(9):935–956
Eskelinen EL, Saftig P (2009) Autophagy: a lysosomal degradation pathway with a central role in health and disease. Biochim Biophys Acta Mol Cell Res 1793(4):664–673. doi:10.1016/j.bbamcr.2008.07.014
Fecto F, Siddique T (2012) UBQLN2/P62 cellular recycling pathways in amyotrophic lateral sclerosis and frontotemporal dementia. Muscle Nerve 45(2):157–162. doi:10.1002/mus.23278
Felbor U, Kessler B, Mothes W, Goebel HH, Ploegh HL, Bronson RT, Olsen BR (2002) Neuronal loss and brain atrophy in mice lacking cathepsins B and L. Proc Natl Acad Sci USA 99(12):7883–7888. doi:10.1073/pnas.112632299
Feng Y, He D, Yao Z, Klionsky DJ (2014) The machinery of macroautophagy. Cell Res 24(1):24–41. doi:10.1038/cr.2013.168
Filimonenko M, Stuffers S, Raiborg C, Yamamoto A, Malerod L, Fisher EM et al (2007) Functional multivesicular bodies are required for autophagic clearance of protein aggregates associated with neurodegenerative disease. J Cell Biol 179(3):485–500. doi:10.1083/jcb.200702115
Fimia GM, Stoykova A, Romagnoli A, Giunta L, Di Bartolomeo S, Nardacci R et al (2007) Ambra 1 regulates autophagy and development of the nervous system. Nature 447:1121–1125
Fornai F, Longone P, Cafaro L, Kastsiuchenka O, Ferrucci M, Manca ML et al (2008) Lithium delays progression of amyotrophic lateral sclerosis. Proc Natl Acad Sci USA 105(6):2052–2057. doi:10.1073/pnas.0708022105
Fornai F, Longone P, Ferrucci M, Lenzi P, Isidoro C, Ruggieri S et al (2008) Autophagy and amyotrophic lateral sclerosis: the multiple roles of lithium. Autophagy 4(4):527–530
Fouillet A, Levet C, Virgone A, Robin M, Dourlen P, Rieusset J et al (2012) ER stress inhibits neuronal death by promoting autophagy. Autophagy 8(6):915–926. doi:10.4161/auto.19716
Fraldi A, Annunziata F, Lombardi A, Kaiser HJ, Medina DL, Spampanato C et al (2010) Lysosomal fusion and SNARE function are impaired by cholesterol accumulation in lysosomal storage disorders. EMBO J 29(21):3607–3620. doi:10.1038/emboj.2010.237
Friedman LG, Lachenmayer ML, Wang J, He LQ, Poulose SM, Komatsu M et al (2012) Disrupted autophagy leads to dopaminergic axon and dendrite degeneration and promotes presynaptic accumulation of alpha-synuclein and LRRK2 in the brain. J Neurosci 32(22):7585–7593. doi:10.1523/Jneurosci.5809-11.2012
Fu MM, Nirschl JJ, Holzbaur EL (2014) LC3 binding to the scaffolding protein JIP1 regulates processive dynein-driven transport of autophagosomes. Dev Cell 29(5):577–590. doi:10.1016/j.devcel.2014.04.015
Fujita N, Itoh T, Omori H, Fukuda M, Noda T, Yoshimori T (2008) The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol Biol Cell 19(5):2092–2100
Fukuda T, Ewan L, Bauer M, Mattaliano RJ, Zaal K, Ralston E et al (2006) Dysfunction of endocytic and autophagic pathways in a lysosomal storage disease. Ann Neurol 59(4):700–708. doi:10.1002/ana.20807
Fusco C, Micale L, Egorov M, Monti M, D’Addetta EV, Augello B et al (2012) The E3-ubiquitin ligase TRIM50 interacts with HDAC6 and p62, and promotes the sequestration and clearance of ubiquitinated proteins into the aggresome. PLoS One 7(7):e40440. doi:10.1371/journal.pone.0040440
Gal J, Strom AL, Kilty R, Zhang F, Zhu H (2007) p62 accumulates and enhances aggregate formation in model systems of familial amyotrophic lateral sclerosis. J Biol Chem 282(15):11068–11077. doi:10.1074/jbc.M608787200
Gal J, Strom AL, Kwinter DM, Kilty R, Zhang J, Shi P et al (2009) Sequestosome 1/p62 links familial ALS mutant SOD1 to LC3 via an ubiquitin-independent mechanism. J Neurochem 111(4):1062–1073. doi:10.1111/j.1471-4159.2009.06388.x
Garami A, Zwartkruis FJ, Nobukuni T, Joaquin M, Roccio M, Stocker H et al (2003) Insulin activation of Rheb, a mediator of mTOR/S6K/4E-BP signaling, is inhibited by TSC1 and 2. Mol Cell 11(6):1457–1466
Ge L, Baskaran S, Schekman R, Hurley JH (2014) The protein-vesicle network of autophagy. Curr Opin Cell Biol 29C:18–24. doi:10.1016/j.ceb.2014.02.005
Gelino S, Hansen M (2012) Autophagy—an emerging anti-aging mechanism. J Clin Exp Pathol Suppl 4:006
Gitcho MA, Strider J, Carter D, Taylor-Reinwald L, Forman MS, Goate AM, Cairns NJ (2009) VCP mutations causing frontotemporal lobar degeneration disrupt localization of TDP-43 and induce cell death. J Biol Chem 284(18):12384–12398. doi:10.1074/jbc.M900992200
Graef M, Friedman JR, Graham C, Babu M, Nunnari J (2013) ER exit sites are physical and functional core autophagosome biogenesis components. Mol Biol Cell 24(18):2918–2931. doi:10.1091/mbc.E13-07-0381
Guo YJ, Chang CM, Huang R, Liu B, Bao L, Liu W (2012) AP1 is essential for generation of autophagosomes from the trans-Golgi network. J Cell Sci 125(7):1706–1715. doi:10.1242/Jcs.093203
Gutierrez MG, Munafo DB, Beron W, Colombo MI (2004) Rab7 is required for the normal progression of the autophagic pathway in mammalian cells. J Cell Sci 117:2687–2697
Hadano S, Otomo A, Kunita R, Suzuki-Utsunomiya K, Akatsuka A, Koike M et al (2010) Loss of ALS2/Alsin exacerbates motor dysfunction in a SOD1-expressing mouse ALS model by disturbing endolysosomal trafficking. PLoS One 5(3):e9805. doi:10.1371/journal.pone.0009805
Hailey DW, Rambold AS, Satpute-Krishnan P, Mitra K, Sougrat R, Kim PK, Lippincott-Schwartz J (2010) Mitochondria supply membranes for autophagosome biogenesis during starvation. Cell 141(4):656–667. doi:10.1016/j.cell.2010.04.009
Hale AN, Ledbetter DJ, Gawriluk TR, Rucker EB (2013) Autophagy. Regulation and role in development. Autophagy 9:951–972
Hamasaki M, Furuta N, Matsuda A, Nezu A, Yamamoto A, Fujita N et al (2013) Autophagosomes form at ER-mitochondria contact sites. Nature 495(7441):389–393. doi:10.1038/nature11910
Hara T, Nakamura K, Matsui M, Yamamoto A, Nakahara Y, Suzuki-Migishima R et al (2006) Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441(7095):885–889
Hayashi-Nishino M, Fujita N, Noda T, Yamaguchi A, Yoshimori T, Yamamoto A (2009) A subdomain of the endoplasmic reticulum forms a cradle for autophagosome formation. Nat Cell Biol 11(12):1433–1437
He CC, Sumpter R, Levine B (2012) Exercise induces autophagy in peripheral tissues and in the brain. Autophagy 8(10):1548–1551. doi:10.4161/Auto.21327
He LQ, Lu JH, Yue ZY (2013) Autophagy in ageing and ageing-associated diseases. Acta Pharmacol Sin 34(5):605–611. doi:10.1038/aps.2012.188
Heiseke A, Aguib Y, Riemer C, Baier M, Schatzl HM (2009) Lithium induces clearance of protease resistant prion protein in prion-infected cells by induction of autophagy. J Neurochem 109(1):25–34. doi:10.1111/j.1471-4159.2009.05906.x
Heiseke A, Aguib Y, Schatzl HM (2010) Autophagy, prion infection and their mutual interactions. Curr Issues Mol Biol 12(2):87–97
Hernandez D, Torres CA, Setlik W, Cebrian C, Mosharov EV, Tang GM et al (2012) Regulation of presynaptic neurotransmission by macroautophagy. Neuron 74(2):277–284. doi:10.1016/j.neuron.2012.02.020
Hollenbeck PJ (1993) Products of endocytosis and autophagy are retrieved from axons by regulated retrograde organelle transport. J Cell Biol 121(2):305–315
Huang CC, Bose JK, Majumder P, Lee KH, Huang JT, Huang JK, Shen CKJ (2014) Metabolism and mis-metabolism of the neuropathological signature protein TDP-43. J Cell Sci 127(Pt 14):3024–3038. doi:10.1242/jcs.136150
Hung YH, Chen LM, Yang JY, Yang WY (2013) Spatiotemporally controlled induction of autophagy-mediated lysosome turnover. Nat Commun 4:2111. doi:10.1038/ncomms3111
Inoki K, Li Y, Xu T, Guan KL (2003) Rheb GTPase is a direct target of TSC2 GAP activity and regulates mTOR signaling. Genes Dev 17(15):1829–1834. doi:10.1101/gad.1110003
Itakura E, Mizushima N (2010) Characterization of autophagosome formation site by a hierarchical analysis of mammalian Atg proteins. Autophagy 6(6):764–776
Iwata A, Riley BE, Johnston JA, Kopito RR (2005) HDAC6 and microtubules are required for autophagic degradation of aggregated huntingtin. J Biol Chem 280(48):40282–40292
Jackson WT, Giddings TH, Taylor MP, Mulinyawe S, Rabinovitch M, Kopito RR, Kirkegaard K (2005) Subversion of cellular autophagosomal machinery by RNA viruses. PLoS Biol 3(5):861–871. doi:10.1371/journal.pbio.0030156
Jäger S, Bucci C, Tanida I, Ueno T, Kominami E, Saftig P, Eskelinen EL (2004) Role for Rab7 in maturation of late autophagic vacuoles. J Cell Sci 117(20):4837–4848. doi:10.1242/Jcs.01370
Jeong JK, Moon MH, Lee YJ, Seol JW, Park SY (2013) Autophagy induced by the class III histone deacetylase Sirt1 prevents prion peptide neurotoxicity. Neurobiol Aging 34(1):146–156. doi:10.1016/j.neurobiolaging.2012.04.002
Jiang M, Wang J, Fu J, Du L, Jeong H, West T et al (2012) Neuroprotective role of Sirt1 in mammalian models of Huntington’s disease through activation of multiple Sirt1 targets. Nat Med 18(1):153–158. doi:10.1038/nm.2558
Jin M, He D, Backues SK, Freeberg MA, Liu X, Kim JK, Klionsky D (2014) Transcriptional regulation by pho23 modulates the frequency of autophagosome formation. Curr Biol 24(12):1314–1322. doi:10.1016/j.cub.2014.04.048
Johnson JO, Mandrioli J, Benatar M, Abramzon Y, Van Deerlin VM, Trojanowski JQ et al (2010) Exome sequencing reveals VCP mutations as a cause of familial ALS. Neuron 68(5):857–864. doi:10.1016/j.neuron.2010.11.036
Ju JS, Fuentealba RA, Miller SE, Jackson E, Piwnica-Worms D, Baloh RH, Weihl CC (2009) Valosin-containing protein (VCP) is required for autophagy and is disrupted in VCP disease. J Cell Biol 187(6):875–888. doi:10.1083/jcb.200908115
Jung CH, Jun CB, Ro SH, Kim YM, Otto NM, Cao J et al (2009) ULK-Atg13-FIP200 complexes mediate mTOR signaling to the autophagy machinery. Mol Biol Cell 20(7):1992–2003. doi:10.1091/mbc.E08-12-1249
Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T et al (2000) LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J 19:5720–5728
Kabuta T, Furuta A, Aoki S, Furuta K, Wada K (2008) Aberrant interaction between Parkinson disease-associated mutant UCH-L1 and the lysosomal receptor for chaperone-mediated autophagy. J Biol Chem 283(35):23731–23738. doi:10.1074/jbc.M801918200
Kabuta T, Suzuki Y, Wada K (2006) Degradation of amyotrophic lateral sclerosis-linked mutant Cu, Zn-superoxide dismutase proteins by macroautophagy and the proteasome. J Biol Chem 281(41):30524–30533. doi:10.1074/jbc.M603337200
Katsumata K, Nishiyama J, Inoue T, Mizushima N, Takeda J, Yuzaki M (2010) Dynein- and activity-dependent retrograde transport of autophagosomes in neuronal axons. Autophagy 6(3):378–385
Kaushik S, Cuervo AM (2012) Chaperone-mediated autophagy: a unique way to enter the lysosome world. Trends Cell Biol 22(8):407–417. doi:10.1016/j.tcb.2012.05.006
Kaushik S, Massey AC, Mizushima N, Cuervo AM (2008) Constitutive activation of chaperone-mediated autophagy in cells with impaired macroautophagy. Mol Biol Cell 19(5):2179–2192. doi:10.1091/mbc.E07-11-1155
Kiffin R, Christian C, Knecht E, Cuervo AM (2004) Activation of chaperone-mediated autophagy during oxidative stress. Mol Biol Cell 15(11):4829–4840. doi:10.1091/mbc.E04-06-0477
Kiffin R, Kaushik S, Zeng M, Bandyopadhyay U, Zhang C, Massey AC, Cuervo AM (2007) Altered dynamics of the lysosomal receptor for chaperone-mediated autophagy with age. J Cell Sci 120(Pt 5):782–791. doi:10.1242/jcs.001073
Kim J, Kundu M, Viollet B, Guan KL (2011) AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat Cell Biol 13(2):132–141. doi:10.1038/ncb2152
Knaevelsrud H, Simonsen A (2010) Fighting disease by selective autophagy of aggregate-prone proteins. FEBS Lett 584:2635–2645
Knoferle J, Koch JC, Ostendorf T, Michel U, Planchamp V, Vutova P et al (2010) Mechanisms of acute axonal degeneration in the optic nerve in vivo. Proc Natl Acad Sci USA 107(13):6064–6069. doi:10.1073/pnas.0909794107
Koike M, Nakanishi H, Saftig P, Ezaki J, Isahara K, Ohsawa Y et al (2000) Cathepsin D deficiency induces lysosomal storage with ceroid lipofuscin in mouse CNS neurons. J Neurosci 20(18):6898–6906
Kollmann K, Damme M, Markmann S, Morelle W, Schweizer M, Hermans-Borgmeyer I et al (2012) Lysosomal dysfunction causes neurodegeneration in mucolipidosis II ‘knock-in’ mice. Brain 135(Pt 9):2661–2675. doi:10.1093/brain/aws209
Komatsu M, Waguri S, Chiba T, Murata S, Iwata J, Tanida I et al (2006) Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 441(7095):880–884
Komatsu M, Wang QJ, Holstein GR, Friedrich VL Jr, Iwata J, Kominami E et al (2007) Essential role for autophagy protein Atg7 in the maintenance of axonal homeostasis and the prevention of axonal degeneration. Proc Natl Acad Sci USA 104(36):14489–14494
Koren I, Reem E, Kimchi A (2010) DAP1, a novel substrate of mTOR, negatively regulates autophagy. Curr Biol 20(12):1093–1098. doi:10.1016/j.cub.2010.04.041
Kramer H (2013) Route to destruction: autophagosomes SNARE lysosomes. J Cell Biol 201(4):495–497. doi:10.1083/jcb.201304065
Lamark T, Johansen T (2012) Aggrephagy: selective disposal of protein aggregates by macroautophagy. Inte J Cell Biol 2012:736905. doi:10.1155/2012/736905
Lee HJ, Cho ED, Lee KW, Kim JH, Cho SG, Lee SJ (2013) Autophagic failure promotes the exocytosis and intercellular transfer of alpha-synuclein. Exp Mol Med 45:e22. doi:10.1038/emm.2013.45
Lee JA, Beigneux A, Ahmad ST, Young SG, Gao FB (2007) ESCRT-III dysfunction causes autophagosome accumulation and neurodegeneration. Curr Biol 17(18):1561–1567. doi:10.1016/j.cub.2007.07.029
Lee JY, Koga H, Kawaguchi Y, Tang W, Wong E, Gao YS et al (2010) HDAC6 controls autophagosome maturation essential for ubiquitin-selective quality-control autophagy. EMBO J 29(5):969–980. doi:10.1038/emboj.2009.405
Lee JY, Nagano Y, Taylor JP, Lim KL, Yao TP (2010) Disease-causing mutations in parkin impair mitochondrial ubiquitination, aggregation, and HDAC6-dependent mitophagy. J Cell Biol 189(4):671–679. doi:10.1083/jcb.201001039
Lee KM, Hwang SK, Lee JA (2013) Neuronal autophagy and neurodevelopmental disorders. Exp Neurobiol 22(3):133–142. doi:10.5607/en.2013.22.3.133
Lee S, Sato Y, Nixon RA (2011) Lysosomal proteolysis inhibition selectively disrupts axonal transport of degradative organelles and causes an Alzheimer’s-like axonal dystrophy. J Neurosci 31(21):7817–7830. doi:10.1523/JNEUROSCI.6412-10.2011
Liang CC, Wang C, Peng X, Gan B, Guan JL (2010) Neural-specific deletion of FIP200 leads to cerebellar degeneration caused by increased neuronal death and axon degeneration. J Biol Chem 285(5):3499–3509. doi:10.1074/jbc.M109.072389
Liao G, Yao Y, Liu J, Yu Z, Cheung S, Xie A et al (2007) Cholesterol accumulation is associated with lysosomal dysfunction and autophagic stress in Npc1−/− mouse brain. Am J Pathol 171(3):962–975. doi:10.2353/ajpath.2007.070052
Liberski PP, Sikorska B, Bratosiewicz-Wasik J, Gajdusek DC, Brown P (2004) Neuronal cell death in transmissible spongiform encephalopathies (prion diseases) revisited: from apoptosis to autophagy. Int J Biochem Cell Biol 36(12):2473–2490. doi:10.1016/j.biocel.2004.04.016
Lipinski MM, Zheng B, Lu T, Yan Z, Py BF, Ng A et al (2010) Genome-wide analysis reveals mechanisms modulating autophagy in normal brain aging and in Alzheimer’s disease. Proc Natl Acad Sci USA 107(32):14164–14169. doi:10.1073/pnas.1009485107
Liu L, Feng D, Chen G, Chen M, Zheng Q, Song P et al (2012) Mitochondrial outer-membrane protein FUNDC1 mediates hypoxia-induced mitophagy in mammalian cells. Nat Cell Biol 14(2):177–185. doi:10.1038/ncb2422
Long X, Ortiz-Vega S, Lin Y, Avruch J (2005) Rheb binding to mammalian target of rapamycin (mTOR) is regulated by amino acid sufficiency. J Biol Chem 280(25):23433–23436. doi:10.1074/jbc.C500169200
Longatti A, Lamb CA, Razi M, Yoshimura S, Barr FA, Tooze SA (2012) TBC1D14 regulates autophagosome formation via Rab11-and ULK1-positive recycling endosomes. J Cell Biol 197(5):659–675. doi:10.1083/jcb.201111079
Lu T, Aron L, Zullo J, Pan Y, Kim H, Chen Y et al (2014) REST and stress resistance in ageing and Alzheimer’s disease. Nature 507(7493):448–454. doi:10.1038/nature13163
Lv L, Li D, Zhao D, Lin R, Chu Y, Zhang H et al (2011) Acetylation targets the M2 isoform of pyruvate kinase for degradation through chaperone-mediated autophagy and promotes tumor growth. Mol Cell 42(6):719–730. doi:10.1016/j.molcel.2011.04.025
Lynch-Day MA, Mao K, Wang K, Zhao M, Klionsky DJ (2012) The role of autophagy in Parkinson’s disease. Cold Spring Harb Perspect Med 2(4):a009357. doi:10.1101/cshperspect.a009357
Maday S, Wallace KE, Holzbaur EL (2012) Autophagosomes initiate distally and mature during transport toward the cell soma in primary neurons. J Cell Biol 196(4):407–417. doi:10.1083/jcb.201106120
Maejima I, Takahashi A, Omori H, Kimura T, Takabatake Y, Saitoh T et al (2013) Autophagy sequesters damaged lysosomes to control lysosomal biogenesis and kidney injury. EMBO J 32(17):2336–2347. doi:10.1038/emboj.2013.171
Maetzel D, Sarkar S, Wang H, Abi-Mosleh L, Xu P, Cheng AW et al (2014) Genetic and chemical correction of cholesterol accumulation and impaired autophagy in hepatic and neural cells derived from Niemann–Pick type C patient-specific iPS cells. Stem Cell Rep 2(6):866–880. doi:10.1016/j.stemcr.2014.03.014
Manzoni C, Lewis PA (2013) Dysfunction of the autophagy/lysosomal degradation pathway is a shared feature of the genetic synucleinopathies. FASEB J 27(9):3424–3429. doi:10.1096/fj.12-223842
Martina JA, Chen Y, Gucek M, Puertollano R (2012) MTORC1 functions as a transcriptional regulator of autophagy by preventing nuclear transport of TFEB. Autophagy 8(6):903–914. doi:10.4161/auto.19653
Martina JA, Diab HI, Lishu L, Jeong AL, Patange S, Raben N, Puertollano R (2014) The nutrient-responsive transcription factor TFE3 promotes autophagy, lysosomal biogenesis, and clearance of cellular debris. Sci Signal 7(309):ra9. doi:10.1126/scisignal.2004754
Martina JA, Puertollano R (2013) Rag GTPases mediate amino acid-dependent recruitment of TFEB and MITF to lysosomes. J Cell Biol 200(4):475–491. doi:10.1083/jcb.201209135
Martina JA, Puertollano R (2013) RRAG GTPases link nutrient availability to gene expression, autophagy and lysosomal biogenesis. Autophagy 9(6):928–930. doi:10.4161/auto.24371
Martinez-Vicente M, Talloczy Z, Kaushik S, Massey AC, Mazzulli J, Mosharov EV et al (2008) Dopamine-modified alpha-synuclein blocks chaperone-mediated autophagy. J Clin Invest 118(2):777–788. doi:10.1172/JCI32806
Massey AC, Kaushik S, Sovak G, Kiffin R, Cuervo AM (2006) Consequences of the selective blockage of chaperone-mediated autophagy. Proc Natl Acad Sci USA 103(15):5805–5810. doi:10.1073/pnas.0507436103
Matsunaga K, Morita E, Saitoh T, Akira S, Ktistakis NT, Izumi T et al (2010) Autophagy requires endoplasmic reticulum targeting of the PI3-kinase complex via Atg14L. J Cell Biol 190(4):511–521. doi:10.1083/jcb.200911141
Mazure NM, Pouyssegur J (2010) Hypoxia-induced autophagy: cell death or cell survival? Curr Opin Cell Biol 22(2):177–180. doi:10.1016/j.ceb.2009.11.015
Medina DL, Fraldi A, Bouche V, Annunziata F, Mansueto G, Spampanato C et al (2011) Transcriptional activation of lysosomal exocytosis promotes cellular clearance. Dev Cell 21(3):421–430. doi:10.1016/j.devcel.2011.07.016
Melser S, Chatelain EH, Lavie J, Mahfouf W, Jose C, Obre E et al (2013) Rheb regulates mitophagy induced by mitochondrial energetic status. Cell Metab 17(5):719–730. doi:10.1016/j.cmet.2013.03.014
Meyer H, Bug M, Bremer S (2012) Emerging functions of the VCP/p97 AAA-ATPase in the ubiquitin system. Nat Cell Biol 14(2):117–123. doi:10.1038/ncb2407
Micsenyi MC, Sikora J, Stephney G, Dobrenis K, Walkley SU (2013) Lysosomal membrane permeability stimulates protein aggregate formation in neurons of a lysosomal disease. J Neurosci 33(26):10815–10827. doi:10.1523/JNEUROSCI.0987-13.2013
Mizuno Y, Amari M, Takatama M, Aizawa H, Mihara B, Okamoto K (2006) Immunoreactivities of p62, an ubiqutin-binding protein, in the spinal anterior horn cells of patients with amyotrophic lateral sclerosis. J Neurol Sci 249(1):13–18. doi:10.1016/j.jns.2006.05.060
Mizushima N, Klionsky DJ (2007) Protein turnover via autophagy: implications for metabolism. Annu Rev Nutr 27:19–40. doi:10.1146/annurev.nutr.27.061406.093749
Mizushima N, Yamamoto A, Matsui M, Yoshimori T, Ohsumi Y (2004) In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol Biol Cell 15:1101–1111
Moon JH, Lee JH, Park JY, Kim SW, Lee YJ, Kang SJ et al (2014) Caffeine prevents human prion protein-mediated neurotoxicity through the induction of autophagy. Int J Mol Med 34(2):553–558. doi:10.3892/ijmm.2014.1814
Mortimore GE, Schworer CM (1977) Induction of autophagy by amino-acid deprivation in perfused rat liver. Nature 270(5633):174–176
Moskot M, Montefusco S, Jakobkiewicz-Banecka J, Mozolewski P, Wegrzyn A, Di Bernardo D et al (2014) The phytoestrogen genistein modulates lysosomal metabolism and transcription factor EB (TFEB) activation. J Biol Chem 289(24):17054–17069. doi:10.1074/jbc.M114.555300
Musiwaro P, Smith M, Manifava M, Walker SA, Ktistakis NT (2013) Characteristics and requirements of basal autophagy in HEK293 cells. Autophagy 9(9):1407–1417
Nakagaki T, Satoh K, Ishibashi D, Fuse T, Sano K, Kamatari YO et al (2013) FK506 reduces abnormal prion protein through the activation of autolysosomal degradation and prolongs survival in prion-infected mice. Autophagy 9(9):1386–1394
Narendra D, Tanaka A, Suen DF, Youle RJ (2008) Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J Cell Biol 183(5):795–803. doi:10.1083/jcb.200809125
Nassif M, Valenzuela V, Rojas-Rivera D, Vidal R, Matus S, Castillo K et al (2014) Pathogenic role of BECN1/beclin 1 in the development of amyotrophic lateral sclerosis. Autophagy 10(7):1256–1271
Nazio F, Strappazzon F, Antonioli M, Bielli P, Cianfanelli V, Bordi M et al (2013) mTOR inhibits autophagy by controlling ULK1 ubiquitylation, self-association and function through AMBRA1 and TRAF6. Nat Cell Biol 15(4):406. doi:10.1038/Ncb2708
Nijholt DA, de Graaf TR, van Haastert ES, Oliveira AO, Berkers CR, Zwart R et al (2011) Endoplasmic reticulum stress activates autophagy but not the proteasome in neuronal cells: implications for Alzheimer’s disease. Cell Death Differ 18(6):1071–1081. doi:10.1038/cdd.2010.176
Nishino I, Fu J, Tanji K, Yamada T, Shimojo S, Koori T et al (2000) Primary LAMP-2 deficiency causes X-linked vacuolar cardiomyopathy and myopathy (Danon disease). Nature 406(6798):906–910. doi:10.1038/35022604
Nishiyama J, Miura E, Mizushima N, Watanabe M, Yuzaki M (2007) Aberrant membranes and double-membrane structures accumulate in the axons of Atg5-null Purkinje cells before neuronal death. Autophagy 3(6):591–596
Nixon RA, Wegiel J, Kumar A, Yu WH, Peterhoff C, Cataldo A, Cuervo AM (2005) Extensive involvement of autophagy in Alzheimer disease: an immuno-electron microscopy study. J Neuropathol Exp Neurol 64:113–122
Novak I, Kirkin V, McEwan DG, Zhang J, Wild P, Rozenknop A et al (2010) Nix is a selective autophagy receptor for mitochondrial clearance. EMBO Rep 11(1):45–51. doi:10.1038/embor.2009.256
Ogata M, Hino S, Saito A, Morikawa K, Kondo S, Kanemoto S et al (2006) Autophagy is activated for cell survival after endoplasmic reticulum stress. Mol Cell Biol 26(24):9220–9231. doi:10.1128/MCB.01453-06
Olzmann JA, Chin LS (2008) Parkin-mediated K63-linked polyubiquitination: a signal for targeting misfolded proteins to the aggresome-autophagy pathway. Autophagy 4(1):85–87
Olzmann JA, Li L, Chudaev MV, Chen J, Perez FA, Palmiter RD, Chin LS (2007) Parkin-mediated K63-linked polyubiquitination targets misfolded DJ-1 to aggresomes via binding to HDAC6. J Cell Biol 178(6):1025–1038. doi:10.1083/jcb.200611128
Orenstein SJ, Cuervo AM (2010) Chaperone-mediated autophagy: molecular mechanisms and physiological relevance. Semin Cell Dev Biol 21(7):719–726. doi:10.1016/j.semcdb.2010.02.005
Orenstein SJ, Kuo SH, Tasset I, Arias E, Koga H, Fernandez-Carasa I et al (2013) Interplay of LRRK2 with chaperone-mediated autophagy. Nat Neurosci 16(4):394–406. doi:10.1038/nn.3350
Orvedahl A, Alexander D, Talloczy Z, Sun Q, Wei Y, Zhang W et al (2007) HSV-1 ICP34.5 confers neurovirulence by targeting the beclin 1 autophagy protein. Cell Host Microbe 1(1):23–35
Orvedahl A, MacPherson S, Sumpter R, Talloczy Z, Zou ZJ, Levine B (2010) Autophagy protects against sindbis virus infection of the central nervous system. Cell Host Microbe 7(2):115–127. doi:10.1016/j.chom.2010.01.007
Papadakis M, Hadley G, Xilouri M, Hoyte LC, Nagel S, McMenamin MM et al (2013) Tsc1 (hamartin) confers neuroprotection against ischemia by inducing autophagy. Nat Med 19(3):351–357. doi:10.1038/nm.3097
Papinski D, Schuschnig M, Reiter W, Wilhelm L, Barnes CA, Maiolica A et al (2014) Early steps in autophagy depend on direct phosphorylation of Atg9 by the Atg1 kinase. Mol Cell 53(3):471–483. doi:10.1016/j.molcel.2013.12.011
Parkinson N, Ince PG, Smith MO, Highley R, Skibinski G, Andersen PM et al (2006) ALS phenotypes with mutations in CHMP2B (charged multivesicular body protein 2B). Neurology 67(6):1074–1077
Parzych KR, Klionsky DJ (2014) An overview of autophagy: morphology, mechanism and regulation. Antioxid Redox Signal 20(3):460–473
Platt FM, Boland B, van der Spoel AC (2012) The cell biology of disease: lysosomal storage disorders: the cellular impact of lysosomal dysfunction. J Cell Biol 199(5):723–734. doi:10.1083/jcb.201208152
Polymeropoulos MH, Lavedan C, Leroy E, Ide SE, Dehejia A, Dutra A et al (1997) Mutation in the alpha-synuclein gene identified in families with Parkinson’s disease. Science 276(5321):2045–2047
Prusiner SB, Scott MR, DeArmond SJ, Cohen FE (1998) Prion protein biology. Cell 93(3):337–348. doi:10.1016/S0092-8674(00)81163-0
Quan W, Kim HK, Moon EY, Kim SS, Choi CS, Komatsu M et al (2012) Role of hypothalamic proopiomelanocortin neuron autophagy in the control of appetite and leptin response. Endocrinology 153(4):1817–1826. doi:10.1210/en.2011-1882
Quintavalle C, Di Costanzo S, Zanca C, Tasset I, Fraldi A, Incoronato M et al (2014) Phosphorylation-regulated degradation of the tumor-suppressor form of PED by chaperone-mediated autophagy in lung cancer cells. J Cell Physiol 229(10):1359–1368. doi:10.1002/jcp.24569
Ramachandran N, Munteanu I, Wang PX, Ruggieri A, Rilstone JJ, Israelian N et al (2013) VMA21 deficiency prevents vacuolar ATPase assembly and causes autophagic vacuolar myopathy. Acta Neuropathol 125(3):439–457. doi:10.1007/s00401-012-1073-6
Ravikumar B, Moreau K, Jahreiss L, Puri C, Rubinsztein DC (2010) Plasma membrane contributes to the formation of pre-autophagosomal structures. Nat Cell Biol 12(8):747–757. doi:10.1038/ncb2078
Redmann M, Dodson M, Boyer-Guittaut M, Darley-Usmar V, Zhang J (2014) Mitophagy mechanisms and role in human diseases. Int J Biochem Cell Biol 53C:127–133. doi:10.1016/j.biocel.2014.05.010
Richards AL, Jackson WT (2013) How positive-strand RNA viruses benefit from autophagosome maturation. J Virol 87(18):9966–9972. doi:10.1128/JVI.00460-13
Richter-Landsberg C, Leyk J (2013) Inclusion body formation, macroautophagy, and the role of HDAC6 in neurodegeneration. Acta Neuropathol 126(6):793–807. doi:10.1007/s00401-013-1158-x
Roberts DJ, Tan-Sah VP, Ding EY, Smith JM, Miyamoto S (2014) Hexokinase-II positively regulates glucose starvation-induced autophagy through TORC1 inhibition. Mol Cell 53(4):521–533. doi:10.1016/j.molcel.2013.12.019
Roczniak-Ferguson A, Petit CS, Froehlich F, Qian S, Ky J, Angarola B et al (2012) The transcription factor TFEB links mTORC1 signaling to transcriptional control of lysosome homeostasis. Sci Signal 5(228):ra42. doi:10.1126/scisignal.2002790
Rodriguez-Muela N, Boya P (2012) Axonal damage, autophagy and neuronal survival. Autophagy 8(2):286–288. doi:10.4161/auto.8.2.18982
Rodriguez-Muela N, Germain F, Marino G, Fitze PS, Boya P (2012) Autophagy promotes survival of retinal ganglion cells after optic nerve axotomy in mice. Cell Death Differ 19(1):162–169. doi:10.1038/cdd.2011.88
Rodriguez-Navarro JA, Kaushik S, Koga H, Dall’Armi C, Shui G, Wenk MR et al (2012) Inhibitory effect of dietary lipids on chaperone-mediated autophagy. Proc Natl Acad Sci USA 109(12):E705–E714. doi:10.1073/pnas.1113036109
Rogov V, Dotsch V, Johansen T, Kirkin V (2014) Interactions between autophagy receptors and ubiquitin-like proteins form the molecular basis for selective autophagy. Mol Cell 53(2):167–178. doi:10.1016/j.molcel.2013.12.014
Rollinson S, Bennion J, Toulson G, Halliwell N, Usher S, Snowden J et al (2012) Analysis of optineurin in frontotemporal lobar degeneration. Neurobiol Aging 33(2):425 e421–422. doi:10.1016/j.neurobiolaging.2010.10.002
Rothenberg C, Srinivasan D, Mah L, Kaushik S, Peterhoff CM, Ugolino J et al (2010) Ubiquilin functions in autophagy and is degraded by chaperone-mediated autophagy. Hum Mol Genet 19(16):3219–3232. doi:10.1093/hmg/ddq231
Rubino E, Rainero I, Chio A, Rogaeva E, Galimberti D, Fenoglio P et al (2012) SQSTM1 mutations in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Neurology 79(15):1556–1562. doi:10.1212/WNL.0b013e31826e25df
Sakaki K, Wu J, Kaufman RJ (2008) Protein kinase C theta is required for autophagy in response to stress in the endoplasmic reticulum. J Biol Chem 283(22):15370–15380. doi:10.1074/jbc.M710209200
Sardi SP, Clarke J, Viel C, Chan M, Tamsett TJ, Treleaven CM et al (2013) Augmenting CNS glucocerebrosidase activity as a therapeutic strategy for parkinsonism and other Gaucher-related synucleinopathies. Proc Natl Acad Sci USA 110(9):3537–3542. doi:10.1073/pnas.1220464110
Sardiello M, Palmieri M, di Ronza A, Medina DL, Valenza M, Gennarino VA et al (2009) A gene network regulating lysosomal biogenesis and function. Science 325(5939):473–477. doi:10.1126/science.1174447
Schatzl HM, Laszlo L, Holtzman DM, Tatzelt J, DeArmond SJ, Weiner RI et al (1997) A hypothalamic neuronal cell line persistently infected with scrapie prions exhibits apoptosis. J Virol 71(11):8821–8831
Scherz-Shouval R, Shvets E, Fass E, Shorer H, Gil L, Elazar Z (2007) Reactive oxygen species are essential for autophagy and specifically regulate the activity of Atg4. EMBO J 26(7):1749–1760. doi:10.1038/sj.emboj.7601623
Schreiber A, Peter M (2013) Substrate recognition in selective autophagy and the ubiquitin-proteasome system. Biochim Biophys Acta 1843(1):163–181. doi:10.1016/j.bbamcr.2013.03.019
Scotter EL, Vance C, Nishimura AL, Lee YB, Chen HJ, Urwin H et al (2014) Differential roles of the ubiquitin proteasome system and autophagy in the clearance of soluble and aggregated TDP-43 species. J Cell Sci 127(Pt 6):1263–1278. doi:10.1242/jcs.140087
Seibenhener ML, Babu JR, Geetha T, Wong HC, Krishna NR, Wooten MW (2004) Sequestosome 1/p62 is a polyubiquitin chain binding protein involved in ubiquitin proteasome degradation. Mol Cell Biol 24(18):8055–8068. doi:10.1128/MCB.24.18.8055-8068.2004
Sepe S, Nardacci R, Fanelli F, Rosso P, Bernardi C, Cecconi F et al (2013) Expression of Ambra1 in mouse brain during physiological and Alzheimer type aging. Neurobiol Aging 35(1):96–108. doi:10.1016/j.neurobiolaging.2013.07.001
Settembre C, Zoncu R, Medina DL, Vetrini F, Erdin S, Erdin S et al (2012) A lysosome-to-nucleus signalling mechanism senses and regulates the lysosome via mTOR and TFEB. EMBO J 31(5):1095–1108. doi:10.1038/emboj.2012.32
Shi R, Weng J, Zhao L, Li XM, Gao TM, Kong J (2012) Excessive autophagy contributes to neuron death in cerebral ischemia. CNS Neurosci Ther 18(3):250–260. doi:10.1111/j.1755-5949.2012.00295.x
Shibutani ST, Yoshimori T (2014) A current perspective of autophagosome biogenesis. Cell Res 24(1):58–68. doi:10.1038/cr.2013.159
Sikorska B, Liberski PP, Giraud P, Kopp N, Brown P (2004) Autophagy is a part of ultrastructural synaptic pathology in Creutzfeldt-Jakob disease: a brain biopsy study. Int J Biochem Cell Biol 36(12):2563–2573. doi:10.1016/j.biocel.2004.04.014
Simonsen A, Birkeland HC, Gillooly DJ, Mizushima N, Kuma A, Yoshimori T et al (2004) Alfy, a novel FYVE-domain-containing protein associated with protein granules and autophagic membranes. J Cell Sci 117:4239–4251
Skibinski G, Parkinson NJ, Brown JM, Chakrabarti L, Lloyd SL, Hummerich H et al (2005) Mutations in the endosomal ESCRTIII-complex subunit CHMP2B in frontotemporal dementia. Nat Genet 37(8):806–808. doi:10.1038/ng1609
Song W, Wang F, Lotfi P, Sardiello M, Segatori L (2014) 2-Hydroxypropyl-beta-cyclodextrin promotes transcription factor EB-mediated activation of autophagy: implications for therapy. J Biol Chem 289(14):10211–10222. doi:10.1074/jbc.M113.506246
Spampanato C, Feeney E, Li L, Cardone M, Lim JA, Annunziata F et al (2013) Transcription factor EB (TFEB) is a new therapeutic target for Pompe disease. EMBO Mol Med 5(5):691–706. doi:10.1002/emmm.201202176
Spencer B, Potkar R, Trejo M, Rockenstein E, Patrick C, Gindi R et al (2009) Beclin 1 gene transfer activates autophagy and ameliorates the neurodegenerative pathology in alpha-synuclein models of Parkinson’s and Lewy body diseases. J Neurosci 29(43):13578–13588. doi:10.1523/JNEUROSCI.4390-09.2009
Spillantini MG, Schmidt ML, Lee VM, Trojanowski JQ, Jakes R, Goedert M (1997) Alpha-synuclein in Lewy bodies. Nature 388(6645):839–840. doi:10.1038/42166
Stanley RE, Ragusa MJ, Hurley JH (2014) The beginning of the end: how scaffolds nucleate autophagosome biogenesis. Trends Cell Biol 24(1):73–81. doi:10.1016/j.tcb.2013.07.008
Stefanis L, Larsen KE, Rideout HJ, Sulzer D, Greene LA (2001) Expression of A53T mutant but not wild-type alpha-synuclein in PC12 cells induces alterations of the ubiquitin-dependent degradation system, loss of dopamine release, and autophagic cell death. J Neurosci 21(24):9549–9560
Sulzer D, Mosharov E, Talloczy Z, Zucca FA, Simon JD, Zecca L (2008) Neuronal pigmented autophagic vacuoles: lipofuscin, neuromelanin, and ceroid as macroautophagic responses during aging and disease. J Neurochem 106(1):24–36. doi:10.1111/j.1471-4159.2008.05385.x
Takahashi Y, Meyerkord CL, Hori T, Runkle K, Fox TE, Kester M et al (2011) Bif-1 regulates Atg9 trafficking by mediating the fission of Golgi membranes during autophagy. Autophagy 7(1):61–73. doi:10.4161/auto.7.1.14015
Tan D, Cai Y, Wang J, Zhang J, Menon S, Chou HT et al (2013) The EM structure of the TRAPPIII complex leads to the identification of a requirement for COPII vesicles on the macroautophagy pathway. Proc Natl Acad Sci USA 110(48):19432–19437. doi:10.1073/pnas.1316356110
Tan JM, Wong ES, Kirkpatrick DS, Pletnikova O, Ko HS, Tay SP et al (2008) Lysine 63-linked ubiquitination promotes the formation and autophagic clearance of protein inclusions associated with neurodegenerative diseases. Hum Mol Genet 17(3):431–439. doi:10.1093/hmg/ddm320
Tanaka A (2010) Parkin-mediated selective mitochondrial autophagy, mitophagy: Parkin purges damaged organelles from the vital mitochondrial network. FEBS Lett 584(7):1386–1392. doi:10.1016/j.febslet.2010.02.060
Tanaka Y, Guhde G, Suter A, Eskelinen EL, Hartmann D, Lullmann-Rauch R et al (2000) Accumulation of autophagic vacuoles and cardiomyopathy in LAMP-2-deficient mice. Nature 406(6798):902–906. doi:10.1038/35022595
Tanji K, Zhang HX, Mori F, Kakita A, Takahashi H, Wakabayashi K (2012) p62/sequestosome 1 binds to TDP-43 in brains with frontotemporal lobar degeneration with TDP-43 inclusions. J Neurosci Res 90(10):2034–2042. doi:10.1002/Jnr.23081
Tatsuta T, Langer T (2008) Quality control of mitochondria: protection against neurodegeneration and ageing. EMBO J 27(2):306–314. doi:10.1038/sj.emboj.7601972
Thelen M, Damme M, Schweizer M, Hagel C, Wong AM, Cooper JD et al (2012) Disruption of the autophagy-lysosome pathway is involved in neuropathology of the nclf mouse model of neuronal ceroid lipofuscinosis. PLoS One 7(4):e35493. doi:10.1371/journal.pone.0035493
Thomas M, Alegre-Abarrategui J, Wade-Martins R (2013) RNA dysfunction and aggrephagy at the centre of an amyotrophic lateral sclerosis/frontotemporal dementia disease continuum. Brain 136(Pt 5):1345–1360. doi:10.1093/brain/awt030
Torres CA, Sulzer D (2012) Macroautophagy can press a brake on presynaptic neurotransmission. Autophagy 8(10):1540–1541. doi:10.4161/Auto.21330
Treff W (1974) Das involutionsmuster des nucleus dentatus. In: Platt D (ed) Altern. Schattauer, Stuttgart, pp 37–54
Trempe JF, Fon EA (2013) Structure and function of Parkin, PINK1, and DJ-1, the three musketeers of neuroprotection. Front Neurol 4:38. doi:10.3389/fneur.2013.00038
Tresse E, Salomons FA, Vesa J, Bott LC, Kimonis V, Yao TP et al (2010) VCP/p97 is essential for maturation of ubiquitin-containing autophagosomes and this function is impaired by mutations that cause IBMPFD. Autophagy 6(2):217–227
Uemura T, Yamamoto M, Kametaka A, Sou YS, Yabashi A, Yamada A et al (2014) A cluster of thin tubular structures mediates transformation of the endoplasmic reticulum to autophagic isolation membrane. Mol Cell Biol 34(9):1695–1706. doi:10.1128/MCB.01327-13
Wager K, Russell C (2013) Mitophagy and neurodegeneration: the zebrafish model system. Autophagy 9(11):1693–1709. doi:10.4161/auto.25082
Walkley SU, Vanier MT (2009) Secondary lipid accumulation in lysosomal disease. Biochim Biophys Acta 1793(4):726–736. doi:10.1016/j.bbamcr.2008.11.014
van der Vaart A, Griffith J, Reggiori F (2010) Exit from the golgi is required for the expansion of the autophagosomal phagophore in yeast Saccharomyces cerevisiae. Mol Biol Cell 21(13):2270–2284. doi:10.1091/mbc.E09-04-0345
Van Humbeeck C, Cornelissen T, Hofkens H, Mandemakers W, Gevaert K, De Strooper B, Vandenberghe W (2011) Parkin interacts with Ambra1 to induce mitophagy. J Neurosci 31(28):10249–10261. doi:10.1523/JNEUROSCI.1917-11.2011
Wang IF, Guo BS, Liu YC, Wu CC, Yang CH, Tsai KJ, Shen CK (2012) Autophagy activators rescue and alleviate pathogenesis of a mouse model with proteinopathies of the TAR DNA-binding protein 43. Proc Natl Acad Sci USA 109(37):15024–15029. doi:10.1073/pnas.1206362109
Wang QJ, Ding Y, Kohtz DS, Mizushima N, Cristea IM, Rout MP et al (2006) Induction of autophagy in axonal dystrophy and degeneration. J Neurosci 26(31):8057–8068. doi:10.1523/JNEUROSCI.2261-06.2006
Wang Y, Martinez-Vicente M, Kruger U, Kaushik S, Wong E, Mandelkow EM et al (2009) Tau fragmentation, aggregation and clearance: the dual role of lysosomal processing. Hum Mol Genet 18(21):4153–4170. doi:10.1093/hmg/ddp367
Vazquez P, Arroba AI, Cecconi F, de la Rosa EJ, Boya P, de Pablo F (2012) Atg5 and Ambra1 differentially modulate neurogenesis in neural stem cells. Autophagy 8(2):187–199. doi:10.4161/auto.8.2.18535
Wei K, Wang P, Miao CY (2012) A double-edged sword with therapeutic potential: an updated role of autophagy in ischemic cerebral injury. CNS Neurosci Ther 18(11):879–886. doi:10.1111/cns.12005
Venugopal B, Mesires NT, Kennedy JC, Curcio-Morelli C, Laplante JM, Dice JF, Slaugenhaupt SA (2009) Chaperone-mediated autophagy is defective in mucolipidosis type IV. J Cell Physiol 219(2):344–353. doi:10.1002/jcp.21676
Vergarajauregui S, Connelly PS, Daniels MP, Puertollano R (2008) Autophagic dysfunction in mucolipidosis type IV patients. Hum Mol Genet 17(17):2723–2737. doi:10.1093/hmg/ddn174
Wild P, Farhan H, McEwan DG, Wagner S, Rogov VV, Brady NR et al (2011) Phosphorylation of the autophagy receptor optineurin restricts Salmonella growth. Science 333(6039):228–233. doi:10.1126/science.1205405
Winslow AR, Chen CW, Corrochano S, Acevedo-Arozena A, Gordon DE, Peden AA et al (2010) Alpha-synuclein impairs macroautophagy: implications for Parkinson’s disease. J Cell Biol 190(6):1023–1037. doi:10.1083/jcb.201003122
Vogiatzi T, Xilouri M, Vekrellis K, Stefanis L (2008) Wild type alpha-synuclein is degraded by chaperone-mediated autophagy and macroautophagy in neuronal cells. J Biol Chem 283(35):23542–23556. doi:10.1074/jbc.M801992200
Wong E, Cuervo AM (2010) Autophagy gone awry in neurodegenerative diseases. Nat Neurosci 13(7):805–811. doi:10.1038/nn.2575
Xilouri M, Vogiatzi T, Vekrellis K, Park D, Stefanis L (2009) Abberant alpha-synuclein confers toxicity to neurons in part through inhibition of chaperone-mediated autophagy. PLoS One 4(5):e5515. doi:10.1371/journal.pone.0005515
Xu F, Gu JH, Qin ZH (2012) Neuronal autophagy in cerebral ischemia. Neurosci Bull 28(5):658–666. doi:10.1007/s12264-012-1268-9
Yang Q, She H, Gearing M, Colla E, Lee M, Shacka JJ, Mao Z (2009) Regulation of neuronal survival factor MEF2D by chaperone-mediated autophagy. Science 323(5910):124–127. doi:10.1126/science.1166088
Yang Y, Coleman M, Zhang L, Zheng X, Yue Z (2013) Autophagy in axonal and dendritic degeneration. Trends Neurosci 36(7):418–428. doi:10.1016/j.tins.2013.04.001
Yang Y, Feng LQ, Zheng XX (2011) Microtubule and kinesin/dynein-dependent, bi-directional transport of autolysosomes in neurites of PC12 cells. Int J Biochem Cell Biol 43(8):1147–1156. doi:10.1016/j.biocel.2011.04.007
Yla-Anttila P, Vihinen H, Jokitalo E, Eskelinen EL (2009) 3D tomography reveals connections between the phagophore and endoplasmic reticulum. Autophagy 5(8):1180–1185
Yordy B, Iijima N, Huttner A, Leib D, Iwasaki A (2012) A Neuron-specific role for autophagy in antiviral defense against herpes simplex virus. Cell Host Microbe 12(3):334–345. doi:10.1016/j.chom.2012.07.013
Yordy B, Iwasaki A (2011) Autophagy in the control and pathogenesis of viral infection. Curr Opin Virol 1(3):196–203. doi:10.1016/j.coviro.2011.05.016
Yordy B, Tal MC, Hayashi K, Arojo O, Iwasaki A (2013) Autophagy and selective deployment of Atg proteins in antiviral defense. Int Immunol 25(1):1–10. doi:10.1093/intimm/dxs101
Young AR, Chan EY, Hu XW, Kochl R, Crawshaw SG, High S et al (2006) Starvation and ULK1-dependent cycling of mammalian Atg9 between the TGN and endosomes. J Cell Sci 119(Pt 18):3888–3900
Yu L, McPhee CK, Zheng L, Mardones GA, Rong Y, Peng J et al (2010) Termination of autophagy and reformation of lysosomes regulated by mTOR. Nature 465(7300):942–946. doi:10.1038/nature09076
Yue Z (2007) Regulation of neuronal autophagy in axon: implication of autophagy in axonal function and dysfunction/degeneration. Autophagy 3(2):139–141
Yun SW, Ertmer A, Flechsig E, Gilch S, Riederer P, Gerlach M et al (2007) The tyrosine kinase inhibitor imatinib mesylate delays prion neuroinvasion by inhibiting prion propagation in the periphery. J Neurovirol 13(4):328–337. doi:10.1080/13550280701361516
Zhang C, Cuervo AM (2008) Restoration of chaperone-mediated autophagy in aging liver improves cellular maintenance and hepatic function. Nat Med 14(9):959–965
Zhang H, Kong X, Kang J, Su J, Li Y, Zhong J, Sun L (2009) Oxidative stress induces parallel autophagy and mitochondria dysfunction in human glioma U251 cells. Toxicol Sci 110(2):376–388. doi:10.1093/toxsci/kfp101
Zhang X, Li L, Chen S, Yang D, Wang Y, Wang Z, Le W (2011) Rapamycin treatment augments motor neuron degeneration in SOD1(G93A) mouse model of amyotrophic lateral sclerosis. Autophagy 7(4):412–425
Zhang X, Yan H, Yuan Y, Gao J, Shen Z, Cheng Y et al (2013) Cerebral ischemia-reperfusion-induced autophagy protects against neuronal injury by mitochondrial clearance. Autophagy 9(9):1321–1333
Zhu G, Wu CJ, Zhao Y, Ashwell JD (2007) Optineurin negatively regulates TNFalpha-induced NF-kappaB activation by competing with NEMO for ubiquitinated RIP. Curr Biol 17(16):1438–1443. doi:10.1016/j.cub.2007.07.041
Zhu J, Wang KZ, Chu CT (2013) After the banquet: mitochondrial biogenesis, mitophagy and cell survival. Autophagy 9(11):1663–1676
Zoncu R, Bar-Peled L, Efeyan A, Wang S, Sancak Y, Sabatini DM (2011) mTORC1 senses lysosomal amino acids through an inside-out mechanism that requires the vacuolar H(+)-ATPase. Science 334(6056):678–683. doi:10.1126/science.1207056
Acknowledgments
Work in the laboratories P.S. and M.D. is supported through Grants of the Deutsche Forschungsgemeinschaft (DFG: SFB877, SPP1580, GRK1459, Cluster of Excellence: Inflammation at Interfaces), the VERUM foundation and the Interuniversity Attraction Poles Program IUAP P7/16 of the Belgian Federal Science Policy Office. E.L.E. and T.S. are supported by the Academy of Finland, Biocentrum Helsinki, and Sigrid Juselius Foundation. We thank Nicolas Yeung (University of Helsinki) for his contribution to designing Fig. 1.
Author information
Authors and Affiliations
Corresponding author
Additional information
P. Saftig and E.-L. Eskelinen have equal contribution.
Rights and permissions
About this article
Cite this article
Damme, M., Suntio, T., Saftig, P. et al. Autophagy in neuronal cells: general principles and physiological and pathological functions. Acta Neuropathol 129, 337–362 (2015). https://doi.org/10.1007/s00401-014-1361-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-014-1361-4